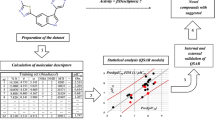
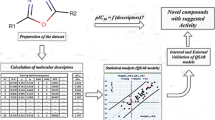
Abstract
Quantitative structure–activity relationship (QSAR) studies were performed on a series of 21 thiazolidine-2,4-dione derivatives to find the structural requirements for PIM-2 kinase inhibitory activity by two-dimensional (2D-QSAR), group-based (G-QSAR) and three-dimensional (3D-QSAR) studies. In the present study, widely used technique viz. stepwise forward–backward (SW-FB) has been applied for the development of 2D- and G-QSAR as variable selection method. The statistically significant best 2D-QSAR model was developed by partial least squares regression (PLSR) having r 2 = 0.78, q 2 = 0.63 with pred_r 2 = 0.78. The statistically significant best G-QSAR model was developed by PLSR method having r 2 = 0.89, q 2 = 0.79 and pred_r 2 = 0.82. The 3D-QSAR studies were performed by k-nearest neighbor molecular field analysis along with genetic algorithm method which showed q 2 = 0.64 and pred_r 2 = 0.94. A docking study revealed the binding orientations of these inhibitors at active site amino acid residues (PHE 43, ASP 124, ASP 182 and GLU 83) of PIM-2 enzyme (PDB ID: 3IWI). The results of this study may be useful to (medicinal) chemists to design more potent thiazolidine-2,4-dione analogs as PIM-2 kinase inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PIM kinases play a key role in regulation of signaling pathways via Janus kinase (JAK)/signal transducer and activator of transcription (STAT) pathway including proliferation, migration and metabolism. The PIM kinases act as weak oncogenes when expressed as transgenes. However, their oncogenic potential increases significantly when co-expressed with a strong oncogene such as c-Myc, a transcription factor which plays an important role in cell growth and differentiation (Forshell et al., 2011; Möröy et al., 1991; Van Lohuizen et al., 1989; Zhang et al., 2008).
Evidence showed that PIM-1 and PIM-2 kinases are overexpressed not only in hematologic malignancies such as multiple myeloma, lymphomas and leukemia but also in solid tumors such as prostate cancer. PIM-3 over expression caused for solid tumors in pancreatic, prostate, colon and other organelles (Brault et al., 2010; Nawijn et al., 2011). Therefore, the PIM kinases may be considered as a potential target for cancer therapy. Furthermore, no severe side effects have been observed after inhibiting all these kinases in an experiment on mice (Mikkers et al., 2004). Novel substituted benzylidene-1,3-thiazolidine-2,4-diones (TZDs) have been identified as potent and highly selective inhibitors of the PIM kinases (Dakin et al., 2012).
Heterocyclic compounds play an important role in cancer chemotherapy particularly five-membered ring heterocycles which contain three carbon atoms, one nitrogen atom and one sulfur atom, known as thiazoles are of considerable interest in different areas of medicinal chemistry (Asati et al., 2014). TZD, one of the most important heterocyclic systems, has therapeutic importance and when combined with other heterocyclic rings it may produce better anticancer activity.
In the literature, several TZDs have been synthesized and evaluated for their anticancer activity. In the present study, QSAR analysis was performed for 21 previously synthesized 5-benzylidenethiazolidine-2,4-dione analogs (Lee et al., 2014) to establishing quantitative relationship between biological activity of derivatives and their structural/physicochemical properties. The aim of the present work is to generate best predictive and validated 2D-, 3D- and G-QSAR models which may help (medicinal) chemists in designing and development of novel thiazolidine-2,4-dione derivatives. In this work, widely used technique viz. stepwise forward–backward (SW-FB) with partial least square (PLS) analysis has been applied for the development of 2D- and G-QSAR models as variable selection method. The k-nearest neighbor (kNN) analysis with genetic algorithm (GA) has been applied for the development of 3D-QSAR model. The generated models may provide insights into the influence of various interactive fields on the activity and thus can help in designing and forecasting the inhibitory activity of novel anticancer agents.
Materials and methods
Data set
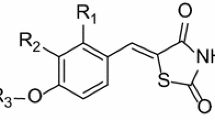
Molecular modeling studies (2D-, 3D- and G-QSAR) were performed using the VLife Molecular Design Suite (MDS 4.4). Selected data set and their biological activity are given in Table 1 where R 1 and R 2 are various substituents. Biological data presented as IC50 (μM) were converted into log (1/IC50) or pIC50 for computational work.

Molecular modeling for 2D-QSAR
The molecular structures of all the 21 compounds were built in 2D builder module using VLife MDS 4.4 software. These 2D structures were converted to 3D and saved in .mol2 format for the development of QSAR models. All the compounds were batch optimized using standard Merck molecular force field (MMFF) and Gasteiger–Marsili charge followed by taking into account distance-dependent dielectric constant at 1.0, convergence criterion or root-mean-square (RMS) gradient at 0.01 kcal/mol Å and the iteration limit to 10,000 (Halgren, 1996; Ghosh and Bagchi, 2009; Sahu et al., 2011). The most stable structure for each compound was generated and used for the calculation of various 2D descriptors such as physicochemical and Baumann alignment-independent topological descriptors (Baumann, 2002). The energy-minimized geometry was used for the calculation of the various 2D descriptors including individual, Chi, ChiV, path count, ChiChain, ChiVChain, chain path count, cluster, path cluster, kappa, element count, estate number, estate contribution, semi-empirical and polar surface area. Alignment-independent descriptors used “T-attribute” which characterize the topology of the molecule. In this study to calculate alignment-independent descriptors, we have used following attributes, 2 (double bonded atom), 3 (triple bonded atom), T (any), C, N, O, S, H, F, Cl, Br and I; the distance range of 0–7 was considered as independent variables. Preprocessing of the independent variables (i.e., descriptors) was done by removing the invariables (descriptors that are constant for all the molecules), which resulted in 212 descriptors in the descriptor pool.
In order to evaluate the QSAR model externally, data sets were divided into training and test set using random selection method, manual data selection method and sphere exclusion methods. Training sets are used as known biological activity data in QSAR model development. Test set is used to evaluate the QSAR model which has been developed by training set. Data set selected randomly given best 2D-QSAR results included five compounds; namely, 4, 6, 8, 16 and 18 were used as test set for model 1 while remaining 16 molecules were used as the training set (Table 1). In order to assess the similarity of the distribution pattern of the molecules in the generated sets, statistical parameters (with respect to the biological activity), i.e., mean, maximum, minimum and standard deviation were calculated for the training and test sets (Table 2). For the prediction statistics to be reliable, the test set must include at least four compounds (Golbraikh and Tropsha, 2002).
Molecular modeling for group-based QSAR (G-QSAR)
G-QSAR addresses the challenges of QSAR model interpretation and the inverse QSAR problems. Group-based QSAR allows establishing a correlation of chemical group/fragment variation at different molecular sites of interest with the biological activity. Group-based QSAR method (Ajmani et al., 2009) comprises of three steps: (a) generation of molecule fragments using a set of predefined chemical rules, (b) calculation of descriptors for the generated fragments and (c) building statistical models using the calculated fragment descriptors and their interactions.
G-QSAR method deals with the molecular fragments rather than the whole molecule. The fragment descriptors and their interactions are related to the biological activity, resulting in model(s) that highlight important substitution site(s) along with their chemical nature and interactions. The suggested important fragments can be used as the building blocks to design novel molecules (Ajmani et al., 2010). For group-based QSAR analysis, various 2D descriptors were calculated for various groups present at different substitution sites of the molecules (i.e., fragment R 1 and R 2). The removal of the invariable group descriptors resulted in a total of 280 group descriptors which can be used for further analysis. Since the same descriptors are calculated for various groups at different sites, the following nomenclature is used for naming a descriptor at a particular position; for example R 1-XAMostHydrophobic descriptor signifies most hydrophobic value on the van der Waals (vdW) surface present at substitution site R 1 (Ajmani et al., 2010).
In G-QSAR, five compounds, namely 2, 4, 11, 16 and 18, were used as test for the development of model 2. For determining the distribution patterns of molecules different parameters such as mean, maximum, minimum and standard deviation were calculated for the training and test sets (Table 2). Partial least squares regression (PLSR) methodology coupled with feature selection method, viz. SW-FB, was used for model development. This was further validated for statistical significance and predictive ability by internal and external validation for building quantitative G-QSAR models (Hasegawa et al., 1999; Wold, 1995).
Molecular modeling for 3D-QSAR
For 3D-QSAR, molecular modeling has been done by a process including alignment of molecules, descriptor generation with the selection of training and test set and development of model.
Alignment of molecules
The 3D-QSAR studies were performed using VLife Molecular Design Suite software. In this study, molecules of the data set are aligned by template-based method (Ajmani et al., 2006). In this method, a template structure is defined and used as a basis for alignment of a set of molecules. The template structure, i.e., 5-(3-pyrazin-2-yl benzylidene)-1,3-thiazolidine-2,4-dione is used for alignment by considering common elements of the series and is shown in Fig. 1. The reference molecule compound 19 has been chosen in such a way that it is the most active among the series of molecules. Reference molecule is chosen on which the other molecules of the data set get aligned considering the chosen template. After optimizing, the template structure and reference molecule were used to superimpose all molecules. The superimposition of all molecules based on minimizing root-mean-square deviation (RMSD), shown in Fig. 2.
Generation of field descriptors
The results of molecular field analysis (MFA) may provide predictive and sufficiently reliable information to medicinal chemist for design and development of novel anticancer agents. This approach is effective for the analysis of data sets, where activity information is available but the structure of the receptor site is unknown. It attempts to postulate and represent the essential features of a receptor site from the aligned common features of the molecules that bind to it. The MFA calculates probe interaction energies on a rectangular grid around a bundle of active molecules. The atomic coordinates of the contributing models were used to compute field values on each point of a 3D grid. Fields of molecules were represented using grids and energy associated with each grid point which can serve as input for the calculation of 3D-QSAR. These energies are added to the study table to form new columns headed according to the probe type. The molecular field is created using methyl group as probe, which represent steric, electrostatic and hydrophobic fields, respectively. For calculation of field descriptor values electrostatic, steric and hydrophobic fields with cutoff values 10.0 and 30.0 kcal/mol, respectively, were selected and charge was selected as Gasteiger and Marsili (Shen et al., 2003). The dielectric constant was set to 1.0 considering the distance-dependent dielectric function. Probe setting was carbon atom with charge 1.0. This resulted in calculation of 4711 field descriptors (electrostatic, steric and hydrophobic) for all the compounds in separate columns after removing descriptors having zero values or same values. Data set selected manually given best result in which five compounds, namely 2, 4, 8, 12 and 18, were used as test set while the remaining 16 molecules as the training set. The unicolumn statistics has been given in Table 2.
Model development
3D-QSAR model development was performed using k-nearest neighbor MFA (kNN-MFA) methodology which relies on a simple distance learning approach whereby an unknown member is classified according to the majority of its kNNs in the training set (Ajmani et al., 2006; Sharaf et al., 1986). The nearness is measured by an appropriate distance metrics (e.g., a molecular similarity measure calculated using field interactions of molecular structures). Given a suitable distance metric a kNN algorithm only requires that a suitable value of k be chosen. In many cases, setting k to 1 provides reasonably good predictive performance for classification purposes. In general, optimal values of k are obtained via trial and error. A more systematic approach is to use a cross-validation scheme to obtain the best value of k for a given data set. The kNN-MFA models were developed by the using of GA approach where cross-correlation limit set to 1.0 and the term selection criterion set as r 2. Some parameters such as population and number of generations were set up to 100 and 500 correspondingly. As some additional parameters, variance cutoff was set at 0.0, and scaling to autoscaling; additionally, kNN parameter setting was done within the range of 2–5 and the prediction method was selected as the distance-based weighted average.
Model evaluation and validation
This is done to test the internal stability and predictive ability of the QSAR models. Internal validation was carried out using “leave-one-out” (q 2, LOO) method. The cross-validated coefficient, q 2, was calculated using the following Eq. 1:
where y i and \(\hat{y}_{i}\) are the actual and predicted activity of the ith molecule in the training set, respectively, and y mean is the average activity of all molecules in the training set. For external validation, activity of each molecule in the test set was predicted using the model generated from the training set. The pred_r 2 value is calculated by using Eq. 2
where y i and \(\hat{y}_{i}\) are the actual and predicted activity of the ith molecule in the test set, respectively, and y mean is the average activity of all molecules in the training set.
The robustness of the QSAR models for experimental training sets was examined by comparing these models to those derived for random data sets. Random sets were generated by rearranging biological activities of the training set molecules. The significance of the models hence obtained was derived based on calculated Z score. Z score can be defined as the absolute difference between the value of the model and the activity field, divided by the square root of the mean square error of the data set Eq. 3
The developed QSAR models were evaluated using the following statistical measures: r 2 (coefficient of determination), q 2 (cross-validated r 2 by LOO), pred_r 2 (r 2 for external test set), F test (Fischer’s value for statistical significance), Z score (Z score calculated by q 2 in the randomization test), best_ran_q 2 (highest q 2 value in the randomization test), a (statistical significance parameter obtained by the randomization test), r 2_se (SEE, standard error of estimate of the model) q 2_se (CV_SE, standard error of cross-validation) and pred_r 2se (pred_SE, standard error of external test set prediction). However, a QSAR model is considered to be predictive, if the following conditions are satisfied: r 2 > 0.6, q 2 > 0.6 and pred_r 2 > 0.5.
Molecular docking
Molecular docking study was performed by using AutoDock Vina which gives the binding mode prediction accurately. It has been tested against a virtual screening benchmark and was found to be better than other programs. Some important attributes for ligand and macromolecular receptor are essential before doing the docking and following the proceeding sequence (Trott and Olson, 2010). For ligand, (1) add all hydrogens, compute Gasteiger charges and merge non-polar H, (2) ensure total charge corresponds to tautomeric state, (3) choose torsion tree root and rotatable bonds. For macromolecule, (1) add all hydrogens, compute Gasteiger charges and merge non-polar H, (2) assign Stouten atomic solvation parameters, (3) create flexible residues PDBQT in addition to the rigid PDBQT file.
The final evaluation was performed to get docking score. Each docking produced multiple docked conformations of the ligand as well as corresponding binding energy scores which were computed using AutoDock scoring function (Seeliger and Groot, 2010). The conformations were ranked based on the scores; a lower scoring conformation was ranked higher. Since an experimentally derived conformation of the bound ligand (true conformation) is available, for each docked conformation of the ligand, a RMSD value was also computed. The RMSD value measures the distance between the docked conformation and the true conformation (Chang et al., 2010; Osterberg et al., 2002). The conformations were also ranked based on the RMSD values, a conformation with lower RMSD value was ranked higher. AutoDock generated different file format that loaded in python molecular viewer (PyMOL) and visualized different binding site properties which may provide valuable insights for structure-based drug design. A RMSD value inferior or close to 2 Å was considered as a successful docking (Stigliani et al., 2012).
Results and discussion
In this study, biological activity (pIC50) was used as dependent variable, and various physiochemical, topological and 3D descriptors have been taken as independent variable. A data set of 21 compounds was divided into training (16 molecules) and test sets (five molecules) using random selection method. The care was taken in such a way that biological activities of all compounds in test set lie within the maximum and minimum value range of biological activities of training set of compounds. A unicolumn statistics for training set and test set were generated to check correctness of selection criteria for trainings and test set molecules (Table 2). After regression analysis, the best models were selected, on the basis of r 2, q 2 and pred_r 2 values (Table 3).
The model generated in 2D-QSAR study through PLSR analysis coupled with SW-FB variable selection method has been presented in Table 4. It reveals that Baumann’s alignment-independent topological descriptors have a major contribution in explaining the variation in activity. In general, a descriptor T_X_Y_Z can be defined as a count of fragments formed with atom types X and Y separated by topological distance of Z bonds. The definitions for the descriptors that were found in the developed 2D-QSAR models are given below.
-
H-DonorCount Number of hydrogen bond donor atoms.
-
T_C_O_5 This is the count of number of carbon atoms (single double or triple bonded) separated from any oxygen atom (single or double bonded) by five bond distance in a molecule.
-
T_2_C_6 This is the count of number of double bounded atoms (i.e., any double bonded atom, T_2) separated from carbon atom by 6 bonds.
The descriptors obtained in the best 2D model suggested that number of hydrogen bond donor groups (~40 %) such as –OH contributed significant role in anticancer activity. The positive contribution of this showed that increase in the values would be beneficial for the anticancer activity of thiazolidine-2,4-dione derivatives. Alignment-independent descriptor T_C_O_5 (~25 %) with positive contribution showed that increase in the number of carbon atoms separated from oxygen atom would be beneficial for activity. Another alignment-independent descriptor T_2_C_6 (negative ~35 %) showed that minimum number of double bounded atoms separated from carbon is beneficial for activity. The inter-correlation matrix between four descriptors with the biological activity for the models 1 is presented in Table 5. Comparative observed and predicted activities of thiazolidine-2,4-dione derivatives by the best 2D-QSAR model are presented in Table 6. Contribution plot of descriptor for models 1 is depicted in Fig. 3a. The graph of actual versus predicted activity for the PLSR analysis is shown in Fig. 3b.
In order to gain insight into the influential molecular part(s) responsible for the variation in activity, G-QSAR model was developed by fragment descriptors and their interactions (Tables 3, 4). Contribution plot of descriptors for model 2 is presented in Fig. 4a. The graph of actual versus predicted activity for the PLSR analysis is shown in Fig. 4b. Inter-correlation matrix between descriptors with the biological activity for the model 2 is presented in Table 5. Comparative observed and predicted activities of thiazolidine-2,4-dione derivatives by best G-QSAR model are presented in Table 6. The definitions for the descriptors in the developed G-QSAR models are given below.
-
R2-Average-vePotential This descriptor signifies the average of the total positive electrostatic potential on van der Waal’s surface area of the molecule at R 2.
-
R1-PolarSurfaceAreaExcludingPandS This descriptor signifies total polar surface area excluding phosphorous and sulfur at R 1.
-
R1-T_T_O_2 The descriptor T_T_O_2 indicates that the presence of substituent with direct attachment of carbon on aromatic ring (i.e., –COOH) is favorable for the activity at R 1.
-
R1-MomInertiaY This descriptor signifies moment of inertia at Y-axis of R 1.
The most contributing descriptors in part R1 with positive coefficient values are PolarSurfaceAreaExcludingPandS (~32 %), so increasing the polar surface area at R1 portion may influence the anticancer activity. Other descriptors at R1 position with negative coefficient values are MomInertiaY (negative ~22 %) and T_T_O_2 (negative ~22 %). These descriptors indicated that direct attachment of carbon group to R1 position may decrease the anticancer activity. The descriptor such as R2 Average-vePotential (negative ~26 %) was found to be inversely proportional to the activity. This indicates that group such as –NH(CH2)3NEt2 which decreases in the average of the total positive electrostatic potential on van der Waal’s surface area at R2 may lead to an increase in the activity.
The 3D-QSAR models were selected on the basis of statistical parameters and the values of the best model 3 having q 2 = 0.64 and pred_r 2 = 0.94 (Table 3). The comparative observed and predicted activities of thiazolidine-2,4-dione derivatives by best QSAR models is presented in Table 6. The graphs of actual verses predicted activity for the series are plotted in Fig. 5 for model 3 which showed good correlation coefficient. Inter-correlation matrix between descriptors with the biological activity for the models 3 is presented in Table 5.
In 3D-QSAR studies, 3D data points generated around thiazolidine-2,4-dione pharmacophore in model 3 are S_556 (−0.0929 −0.0812), S_1381 (−0.3207 −0.3034) and E_849 (0.2828 0.3184), that is, steric and electrostatic interaction field at lattice points 556, 1381 and 849, respectively (Figs. 6, 7). This helps in identification of various molecular features responsible for variation of activity and hence aid in design of novel anticancer agents. The steric descriptor such as S_556 and S_1381 with negative coefficients showed a region where bulky substituents are disfavored for activity. Electrostatic field descriptor such as E_849 with positive coefficient represents regions where electropositive (electron-withdrawing) groups are favorable for activity. The model is validated by predicting the biological activities of the test molecules, as indicated in Table 6.
Binding mode analysis by molecular docking
The intermolecular interaction between ligand and target PIM-2 kinase was confirmed by docking study. The most active compound 19 showed significant interaction with receptor and bound with different amino acid residues such as PHE-43, GLU-83, ASP-182 and ASP-124 of PIM-2 kinase through hydrogen bonds (Fig. 8). The hydrogen bond distance from PHE-43 to carbonyl, GLU-83 to carbonyl, GLU-83 to hydrogen, ASP-182 to carbonyl and ASP-124 to –NH groups were found to be 3.0, 3.1, 1.9, 3.3 and 2.0 Å, respectively. The conformations of the most active compound were ranked on the basis of RMSD values (Fig. 9). The lowest RMSD value was 2.266 Å with −7.9 kcal/mol binding affinity which showed good inhibitory activity against PIM-2 kinase.
Conclusions
In the present study, an attempt has been made to identify the structural requirements of thiazolidine-2,4-dione derivatives for potential anticancer activity. From 2D-, 3D- and G-QSAR analysis, different models have been generated and the best model may be used for the design of novel anticancer agent(s). The descriptors generated in 2D-QSAR equation highlighted the importance of thiazolidine-2,4-dione scaffold for anticancer activity of compounds. The 2D-QSAR model indicates that the descriptors are statistically significant and having high correlation coefficient and reliable predictability. These models generated various descriptors such as H-DonorCount, T_2_C_6 and T_C_O_5 which contribute to biological activity. The negative coefficient value of descriptors indicated that a lower value leads to better anticancer activity. Positive coefficient value of descriptors indicates that a higher value leads to better anticancer activity. The results obtained from G-QSAR generated various descriptors such as R2-Average-vePotential, R1-PolarSurfaceAreaExcludingPandS, R1-T_T_O_2 and R1-MomInertiaY which contribute to biological activity. The G-QSAR study revealed that increase in total polar surface area at R 1 position and decrease in Average-vePotential at R 2 increase the anticancer activity. The results obtained from 3D-QSAR studies were used to optimize the electrostatic, steric and hydrophobic requirements around the thiazolidine-2,4-dione scaffold for enhancing the anticancer activity. The 3D-QSAR studies suggested that the substituents with less bulky and more electropositive groups around thiazolidine-2,4-dione core increase anticancer activity. The results of the present work may be useful to (medicinal) chemists to understand the relationship of structural/physiochemical parameters with biological activity. It may be helpful to select suitable substituent(s) for the design and development of more potent, effective and selective anticancer agents.
References
Ajmani S, Jadhav K, Kulkarni SA (2006) Three-dimensional QSAR using the k-nearest neighbor method and its interpretation. J Chem Inf Model 46:24–31
Ajmani S, Jadhav K, Kulkarni SA (2009) Group-based QSAR (GQSAR): mitigating interpretation challenges in QSAR. QSAR Comb Sci 28:36–41
Ajmani S, Jadhav K, Kulkarni SA (2010) A comprehensive structure–activity analysis of protein kinase B-alpha (Akt1) inhibitors. J Mol Graph Model 28:683–694
Asati V, Mahapatra DK, Bharti SK (2014) Thiazolidine-2,4-diones as multi-targeted scaffold in medicinal chemistry: potential anticancer agents. Eur J Med Chem 87:814–833
Baumann K (2002) An alignment-independent versatile structure descriptor for QSAR and QSPR based on the distribution of molecular features. J Chem Inf Comput Sci 42:26–35
Brault L, Gasser C, Bracher F, Huber K, Knapp S, Schwaller J (2010) PIM serine/threonine kinases in the pathogenesis and therapy of hematologic malignancies and solid cancers. Haematologica 95:1004–1015
Chang MW, Ayeni C, Breuer S, Torbett BE (2010) Virtual screening for HIV protease inhibitors: a comparison of AutoDock 4 and Vina. PLoS One 5(8):e11955
Dakin LA, Block MH, Chen H, Code E, Dowling JE, Feng X, Ferguson AD, Green I, Hird AW, Howard T, Keeton EK, Lamb ML, Lyne PD, Pollard H, Read J, Wu AJ, Zhang T, Zheng X (2012) Discovery of novel benzylidene-1,3-thiazolidine-2,4-diones as potent and selective inhibitors of the PIM-1, PIM-2, and PIM-3 protein kinases. Bioorg Med Chem Lett 22:4599–4604
Forshell LP, Li Y, Forshell TZ, Rudelius M, Nilsson L, Keller U, Nilsson J (2011) The direct Myc target PIM-3 cooperates with other PIM kinases in supporting viability of Myc-induced B-cell lymphomas. Oncotarget 2:448–460
Ghosh P, Bagchi MC (2009) QSAR modeling for quinoxaline derivatives using genetic algorithm and simulated annealing based feature selection. Curr Med Chem 16:4032–4048
Golbraikh A, Tropsha A (2002) Predictive QSAR modeling based on diversity sampling of experimental datasets for the training and test set selection. J Comput Aided Mol Des 16:357–369
Halgren TA (1996) Merck molecular force field. III. Molecular geometries and vibrational frequencies for MMFF94. J Comput Chem 17:553–586
Hasegawa K, Kimura T, Funatsu K (1999) GA strategy for variable selection in QSAR studies: enhancement of comparative molecular binding energy analysis by GA-based PLS method. Quant Struct Act Relatsh 18:262–272
http://vina.scripps.edu/ (Accessed on 2 Sep 2014)
Lee J, Park J, Hong VS (2014) Synthesis and evaluation of 5-(3-(pyrazin-2-yl)benzylidene)thiazolidine-2,4-dione derivatives as pan–PIM kinases inhibitors. Chem Pharm Bull 62(9):906–914
Mikkers H, Nawijn M, Allen J, Brouwers C, Verhoeven E, Jonkers J, Berns A (2004) Mice deficient for all PIM kinases display reduced body size and impaired responses to hematopoietic growth factors. Mol Cell Biol 24:6104–6115
Möröy T, Verbeek S, Ma A, Achacoso P, Berns A, Alt F (1991) E mu N and E mu L-myc cooperate with E mu pim-1 to generate lymphoid tumors at high frequency in double-transgenic mice. Oncogene 6:1941–1948
Nawijn MC, Alendar A, Berns A (2011) For better or for worse: the role of Pim oncogenes in tumorigenesis. Nat Rev Cancer 11:23–34
Osterberg F, Morris GM, Sanner MF, Olson AJ, Goodsell DS (2002) Automated docking to multiple target structures: incorporation of protein mobility and structural water heterogeneity in autodock. Proteins Struct Funct Genet 46:34–40
Sahu NK, Shahi S, Sharma MC, Kohli DV (2011) QSAR studies on imidazopyridazine derivatives as PfPK7 inhibitors. Mol Simul 37:752–765
Seeliger D, Groot BL (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des 24:417–422
Sharaf MA, Illman DL, Kowalski BR (1986) Chemometrics. Wiley, New York
Shen M, Xiao Y, Golbraikh A, Gombar VK, Tropsha A (2003) Development and validation of k-nearest-neighbor QSPR models of metabolic stability of drug candidates. J Med Chem 46:3013–3020
Stigliani JL, Bernardes-Génisson V, Bernadoua J, Pratviela G (2012) Cross-docking study on InhA inhibitors: a combination of Autodock Vina and PM6-DH2 simulations to retrieve bio-active conformations. Org Biomol Chem 10:6341–6345
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comp Chem 31:455–461
Van Lohuizen M, Verbeek S, Krimpenfort P, Domen J, Saris C, Radaszkiewicz T, Berns A (1989) Predisposition to lymphomagenesis in pim-1 transgenic mice: cooperation with c-myc and N-myc in murine leukemia virus-induced tumors. Cell 56:673–682
VLife Sciences Technology Pvt. Ltd., Pune. www.vlifesciences.com
Wold S (1995) PLS for multivariate linear modelling. In: van de Waterbeemd H (ed) QSAR: chemometric methods in molecular design. Wiley-VCH, Weinheim, pp 195–218
Zhang Y, Wang Z, Li X, Magnuson NS (2008) Pim kinase-dependent inhibition of c-Myc degradation. Oncogene 27:4809–4819
Acknowledgments
The authors gratefully acknowledge Indian Council of Medical Research (ICMR), New Delhi, for providing Senior Research Fellowship (SRF) to Mr. Vivek Asati (SRF No. 45/28/2013-PHA/BMS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Asati, V., Bharti, S.K. QSAR studies for some thiazolidine-2,4-dione derivatives as PIM-2 kinase inhibitors. Med Chem Res 25, 1329–1339 (2016). https://doi.org/10.1007/s00044-016-1577-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-016-1577-z