Abstract
Hair cells play key roles in hearing and balance, and hair cell loss would result in hearing loss or vestibular dysfunction. Cellular and molecular research in hair cell biology provides us a better understanding of hearing and deafness. Zebrafish, owing to their hair cell-enriched organs, have been widely applied in hair cell-related research worldwide. Similar to mammals, zebrafish have inner ear hair cells. In addition, they also have lateral line neuromast hair cells. These different types of hair cells vary in morphology and function. However, systematic analysis of their molecular characteristics remains lacking. In this study, we analyzed the GFP+ cells isolated from Tg(Brn3c:mGFP) larvae with GFP expression in all hair cells using single-cell RNA-sequencing (scRNA-seq). Three subtypes of hair cells, namely macula hair cell (MHC), crista hair cell (CHC), and neuromast hair cell (NHC), were characterized and validated by whole-mount in situ hybridization analysis of marker genes. The hair cell scRNA-seq data revealed hair cell-specific genes, including hearing loss genes that have been identified in humans and novel genes potentially involved in hair cell formation and function. Two novel genes were discovered to specifically function in NHCs and MHCs, corresponding to their specific expression in NHCs and MHCs. This study allows us to understand the specific genes in hair cell subpopulations of zebrafish, which will shed light on the genetics of both human vestibular and cochlear hair cell function.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hair cells, getting their name from the hair-like structure on the surface of the cell body [1], detect mechanical vibrations through the cilia. Mechanoelectrical transduction (MET) channels on the tip of stereocilia open in response to stimuli, which cause the depolarization of the hair cells [2, 3] and the release of the neurotransmitters into the synaptic cleft between the hair cell and auditory neuron [4,5,6]. Inner ear mechanosensory hair cells, including cochlear hair cells and vestibular hair cells, play crucial roles in hearing and balance in mammals. The cochlear hair cells function as receptors of acoustic vibration, and they can be divided into outer hair cells (OHCs) and inner hair cells (IHCs) [7]. There are two types of vestibular sensory epithelia: maculae and cristae. The maculae are located in the utricle and saccule, and the cristae lie at the ends of the three semicircular canals. Both of the vestibular sensory epithelia are composed of hair cells and supporting cells. The vestibular hair cells can be divided into type I and type II hair cells [8]. The macula hair cells can sense linear acceleration and gravitational equivalent, and crista hair cells can sense angular acceleration and deceleration [8, 9]. In mammals, the cochlear hair cells get mechanical signals from the tectorial membrane, but the crista hair cells and macula hair cells sense the vibration from the endolymph and otolith, respectively.
As vertebrates, zebrafish have many organs similar to humans, which make them an excellent animal model for biological and medical research [10]. Zebrafish have an ear-like structure, otic vesicle; however, they do not have the cochlea. In the otic vesicle of zebrafish, there are five clusters of hair cells, three clusters of crista hair cells, and two clusters of macula hair cells. In the cristae, including anterior crista (AC), lateral crista (LC), and posterior crista (PC), hair cells put their kinocilia into the endolymph and detect the endolymph flow caused by head rotation. In the maculae, including the utricular macula (UM) and saccular macula (SM), the cilia of hair cells are in contact with the otoliths, which would produce movement if acceleration occurs. Therefore, the macula hair cells can sense linear acceleration and gravity. It has been demonstrated that the utricular macula is responsible for vestibular function [11] and the saccular macula is the hearing organ [12] in zebrafish. Unlike mammals, zebrafish have neuromast hair cells in their lateral line system. These hair cells are located on the surface of the skin, and are sensors of the surrounding water, which help them detect prey and avoid predators. Because they are easy to be administered to and imaged, the neuromast hair cells have been widely used in hair cell-related biomedical research, for example, in ototoxic drug screening.
The zebrafish model has been widely utilized in the hearing research field [13,14,15,16,17,18]. However, the systematic analysis of molecular and morphological differences among these hair cells is so far lacking. In this study, scRNA-seq was used to analyze zebrafish hair cells’ gene expression, and uncover the molecular differences among the different subtypes of hair cells.
Results
The morphological characteristics of the three types of hair cells in zebrafish
Hair cells play crucial roles in hearing and balance, and they are variable in morphology and function. The typical feature of hair cells that differ from other cells is the cilia, including kinocilia and stereocilia, on the surface of the cell body. The cilia of hair cells can detect mechanical vibration by stereocilia bundle deflection, leading to the tension of the tip links, opening of the MET channels, and causing depolarization of hair cells. In zebrafish, except for the inner ear, there are hair cells in their lateral line system, and the hair cells can be divided into three types according to their morphology and location, namely crista hair cells, macula hair cells, and neuromast hair cells (Fig. 1A). The first two types are both located in the inner ear, and function importantly in hearing and balance, and the third type is key component of the lateral line system, which are distributed on the surface of the skin, and help the fish to sense the environmental water. Although the cilia are the common structure of all hair cells, there are also differences among the three types of hair cells. The crista hair cells and macula hair cells have straight kinocilia, which seem inflexible, and the kinocilia of neuromast hair cells can bend themselves to detect the water movement (Fig. 1B–D). For the length of kinocilia, taking the zebrafish larva of 3 dpf as an example, the crista hair cells have the longest kinocilia, which are almost 30 μm in length, and the kinocilia of macula hair cells and neuromast hair cells are approximately 10 μm and 20 μm, respectively (Fig. 1E). For the size of cell bodies, the macula hair cells are the biggest ones, and the cell bodies of the neuromast hair cells are the smallest among them (Fig. 1F).
The morphological feature of the three different types of hair cells in zebrafish. A The diagram of the zebrafish larva with hair cell labeling. UO, utricular otolith; SO, saccular otolith; UMHC, utricular macula hair cell; SMHC, saccular macula hair cell; ACHC, anterior crista hair cell; LCHC, lateral crista hair cell; PCHC, posterior crista hair cell. B–D The fluorescent image at high magnification of the three different types of hair cells. The diagrams of the different hair cells are presented in the boxes drawn with white dotted lines at the bottom left. E–F The comparison of the kinocilia and the cell bodies of the three different types of hair cells. *P < 0.05, ***P < 0.001, ****P < 0.0001
Single-cell RNA-sequencing reveals different subpopulations of hair cells in zebrafish
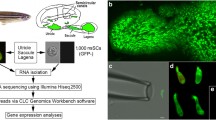
To further distinguish the different types of hair cells at the molecular level, we used single-cell RNA-sequencing to analyze the gene expression patterns. Here, the transgenic zebrafish line Tg(Brn3c:mGFP) [19], in which the hair cells and retinal ganglion cells (RGCs) were labeled by the membrane-targeted green fluorescent protein (mGFP) (Fig. 2A), was used as the animal model. The zebrafish larvae at 6 dpf were dissociated into the single cells by trypsin, and the GFP-positive cells were sorted using the fluorescence activated cell sorting (FACS) method (Figs. S1, S2). After high-throughput sequencing based on the 10 × Genomics system, the gene expression data were obtained. Based on Seurat [20] analysis, the cells were divided into 21 clusters. By analyzing the gene expression patterns of the top genes in each cluster, the four clusters of cells from the cluster 0, 5, 7, 12, were annotated as hair cells, and the cells from the cluster 2, 3, and 10 were identified as retinal ganglion cells. Both hair cells and retinal ganglion cells are GFP-expressing cells in the Tg(Brn3c:mGFP) zebrafish [19]. The rest were considered as other cells, which were GFP-negative and brought into the GFP-positive cells due to inevitable technical problems, including red blood cells, lymphocytes, muscle cells, and so on (Figs. 2B, S3). The number of cells and marker genes detected in each cluster was shown (Figs. 2 C, D). For each cluster, the top marker genes were listed, and the expression patterns of these genes were also presented (Fig. 2E).
Single-cell RNA-sequencing (scRNA-seq) of zebrafish hair cells. A The fluorescent image of the Tg(Brn3c:mGFP) transgenic zebrafish larva at 3 dpf. The GFP-expressing cells in the optic tectum, which were circled by the orange dotted line, are the retinal ganglion cells (RGCs), and the other GFP-expressing cells in the otic vesicle and lateral line system are hair cells. B The UMAP analysis of the zebrafish scRNA-seq data. The four clusters of cells circled by the red dotted line were annotated as hair cells. C, D The number of cells and marker genes detected in each cluster. E The dotplot of the top genes in each cluster
Validation of the gene expression in the subtypes of hair cells using whole-mount in situ hybridization
Through our analysis, as shown above, the cells from cluster 0, 5, 7, 12 were considered as hair cells. In our subsequent study, we focused on these cells, and further analysis showed that these hair cells can be divided into three subpopulations, namely macula hair cells (cluster 5), crista hair cells (cluster 12), and neuromast hair cells (cluster 0, 7) (Fig. 3A). We further analyzed the top marker genes expressed in the four clusters, and found that some genes were mainly expressed in one of the clusters specifically, for example, tectb in cluster 5, zpld1a in cluster 12, calm1b in cluster 0 and 7 (Figs. 3B, S3). Functional enrichment analysis showed that many of the genes expressed in the four clusters of cells above have hair cell-related biological function (Fig. 3C–F), which indicated that these cells were hair cells. To further confirm the hair cell clustering and annotation, we performed the whole-mount in situ hybridization (WISH). As shown in Fig. 3G, the cells with high tectb gene expression were mainly gathering in the cluster 5, and the tectb gene was proved to be expressed specifically in the macula hair cells, including the utricular hair cells and saccular hair cells, by WISH. The zpld1a gene was mainly expressed in the crista hair cells, which were thought to be located in the cluster 12 according to our analysis (Fig. 3H), and this is consistent with the previous studies [21]. The cells from the cluster 0 and 7 were considered as neuromast hair cells because of the expression patterns of their marker genes, such as calm1b (Fig. 3I). As for cells from cluster 0 and 7, although they are both neuromast hair cells, they are different from each other. The cells from cluster 0 were classified to be mature neuromast hair cells, and the cluster 7 were classified to be young neuromast hair cells, according to the expression of the mature and young hair cell markers, s100s [22] and prox1a [23], respectively, in the two clusters (Fig. S4).
The hair cell classification in zebrafish. A The four clusters of hair cells can be further divided into three types, namely macula hair cells (cluster 5), crista hair cells (cluster 12), and neuromast hair cells (cluster 0 and 7). B The dotplot of top genes in the four clusters of hair cells. C–F The cnetplot of the genes expressed in the four clusters of hair cells. G-I The results of the WISH. The genes presented here are expressed in the macula hair cells (G), crista hair cells (H), and neuromast hair cells (I), respectively. The images at the bottom right in G and H show the dorsal and enlarged view of the gene expression. The red arrowheads and arrows indicate the inner ear hair cells and neuromast hair cells, respectively
On the other hand, we analyzed the marker gene of supporting cells, klf17 [24], and found that it was mainly expressed in the cells of cluster 1 (Fig. S5), which suggests that this cluster of cells are supporting cells. Likewise, we also analyzed the genes that were reported to be expressed in mantle cells, such as tnfsf10, ponzr6, pkhd1l1, fat1b, crb3b, cts12, ovgp1, and cldne [17], and found that cells expressing these genes with high level were clustering in cluster 9 (Fig. S6). For cluster 14, these cells are closer to the supporting cells (cluster 1) in UMAP clustering (Fig. 1B); however, they express some of the genes that have been proved to be expressed in hair cells specifically, such as myo6b [25], myo7aa [26] (Fig. S7), which raises a possibility that they are supporting cells that can differentiate into hair cells. Therefore, we concluded that the cells from cluster 1 and 9 were supporting cells and mantle cells, respectively, and the cells from the cluster 14 might be hair cell progenitors.
The molecular properties of the three types of hair cells
Given that different types of hair cells were clustered into different populations, we made a comparison among the distinguishable hair cells at the molecular level. As shown in Fig. 2D, the number of marker genes detected in the cluster 0, 5, 7, 12 was 2352, 1023, 2029, 928, respectively. The two types of inner ear hair cells, macula hair cells and crista hair cells (cluster 5 and 12), share 568 genes, and 1677 marker genes were expressed in both the mature and young neuromast hair cells (cluster 0 and 7). However, the neuromast hair cells and inner ear hair cells only share a small number of marker genes, which reveals their differences on the gene expression level (Fig. 4A). Furthermore, the top marker genes expressed in the different clusters were obviously different (Fig. 4B). We also performed gene enrichment analysis through gene ontology (GO) term, and found that these different types of hair cells vary dramatically in biological process, molecular function, and cellular component. For example, the neuromast hair cells have more energy metabolism-related activity, which raises the possibility that they require more energy to work compared to the inner ear hair cells (Fig. 4C–E).
The molecular difference among the different hair cells. A The marker genes that are expressed in different hair cells and the relationship among them. B The heatmap of the top marker genes in each cluster of hair cells. C–E The GO analysis reveals the different properties among the different hair cells in biological process (C), molecular function (D), and cellular component (E)
The neuromast hair cells were enriched with MET gene expression
The MET channels are required for functional hair cells and they are complex comprising of several components, such as TMC1, TMC2, TMIE, LHFPL5, and CIB2 [2, 27,28,29,30,31]. Hair cells with functional MET channels are crucial for normal hearing and balance. In addition, the tip links play important roles in the stereocilia deflection and MET channel-gating, and it was proved to be composed of two cadherins, CDH23 and PCDH15 [32,33,34]. To determine whether these key molecules are also expressed in zebrafish hair cells and what differences there are among these different hair cells, we analyzed the expression patterns of the genes encoding these important proteins. As illustrated in Fig. 5A–F, the orthologs of mammalian MET complex components were expressed in most of the neuromast hair cells; however, only a small proportion of inner ear hair cells expressed these genes. The genes encoding the tip link components, cdh23 and pcdh15a, were expressed in a few hair cells (Fig. 5G–H).
The zebrafish hair cell scRNA-seq reveals potential hearing loss genes
Hearing is one of the most important sensory functions, and it relies on functional hair cells. Healthy hair cells are the receptors of acoustic signals; therefore, genes responsible for hair cell development, survival, and function are also important for normal hearing. 119 genes have been identified as non-syndromic hearing loss (NSHL) genes in humans (https://hereditaryhearingloss.org/). Among these genes, 96 human genes have orthologous genes in zebrafish (Fig. 6A), which demonstrates that zebrafish are quite similar to humans in terms of NSHL genes. Furthermore, 51 human NSHL genes have 57 orthologous genes which are expressed in zebrafish hair cells (cluster 0, 5, 7, and 12), and the detailed information was shown in Fig. 6B. Except for the orthologs of identified human NSHL genes, there are still more than 3000 genes being specifically expressed in the zebrafish hair cells (Fig. 6C). Some of these genes are thought to be crucial for hair cell function, and it is quite possible that they are potential hearing loss genes. In other words, these data may provide us clear direction for our scientific research in hearing loss gene identification. Under the guidance of this idea, we randomly selected some genes for further analysis, and found that these genes were specifically expressed in either or both of the zebrafish inner ear and neuromast hair cells (Fig. 6D), indicating that these genes function in hair cell-related biological processes, and may even be essential for hearing function.
The zebrafish hair cell scRNA-seq reveals potential hearing loss genes. A The relationship between the known human NSHL genes and zebrafish hair cell-enriched genes. B The list of the human NSHL genes, which have orthologs expressed in zebrafish hair cells. C Venn diagram of the zebrafish orthologs of human NSHL genes and marker genes expressed in each cluster of hair cells in zebrafish. The numbers indicate the number of the genes. D The expression patterns of the hair cell-enriched genes in zebrafish embryos
Functional analysis of the candidate genes involved in hair cell development
Our scRNA-seq analysis revealed genes that are specifically expressed in zebrafish hair cells, some of which are orthologs of human NSHL genes; however, many genes have not been reported to play roles in hair cell development or function. To investigate the function of hair cell-enriched genes, we took the capgb and mb gene as examples, which were mainly expressed in neuromast hair cells and macula hair cells, respectively (Figs. 6D, S8). Here, we used morpholino-mediated gene knockdown to down-regulate the gene expression (Fig. S9). As illustrated in Fig. 7A, B, the capgb-morphants exhibited decreased hair cells in their lateral line neuromasts compared to the littermate control, and this abnormality can be rescued by supplying with wild-type capgb-mRNA. Furthermore, the capgb-morphants had less response to the acoustic stimuli in the startle response test (Fig. 7C, D), which indicated that their hair cells had lost their function to some extent. Likewise, in another experiment, the mb gene knockdown resulted in reduced macula hair cells, which can be rescued by coinjecting with mb-mRNA (Fig. 7E, F), and the mb-morphants showed severely abnormal balance ability in the vestibulo-ocular reflex (VOR) test (Fig. 7G, H).
Functional analysis of the candidate genes involved in hair cell development. A, B The capgb gene knockdown resulted in decreased neuromast hair cells in zebrafish. C, D The capgb-morphants showed less response to the acoustic stimuli. E, F The mb gene knockdown resulted in reduced macula hair cells in zebrafish. G, H The mb-morphants showed damaged balance ability. *P < 0.05, **P < 0.01, ****P < 0.0001
Discussion
In mammals, inner ear hair cells can detect the mechanical signals and transform them into physiological signals, which were then transmitted to the brain through the auditory neurons [35]. Inner ear hair cells can be divided into cochlear hair cells and vestibular hair cells, which play important roles in hearing and balance, respectively. The two types of hair cells differ from each other in both structure and function [7].
As an excellent animal model, zebrafish have hundreds of hair cells, making it a good model to study hair cell function. Similar to mammals, zebrafish have inner ears, consisting of semicircular canals and otolith; however, zebrafish have no cochlea in their inner ear. Therefore, inner ear hair cells can be divided into macula hair cells and crista hair cells in zebrafish. Except for inner ear hair cells, zebrafish have a third type of hair cells, neuromast hair cells, in their lateral line system. The three different hair cells differ from each other in morphology; however, the molecular difference among them is unknown.
Here, we used the single-cell RNA-sequencing method to analyze the gene expression patterns in zebrafish hair cells, and uncovered the molecular difference among the different types of hair cells. In brief, after quality control, the scRNA-seq data received dimensional reduction processing using the UMAP method, and all of the cells sequenced were classified as 21 clusters. After annotating according to the known marker genes and gene expression patterns in the database (http://zfin.org/), we identified four clusters of cells as macula hair cells, crista hair cells, and neuromast hair cells. Further validation using WISH confirmed our cell clustering and annotation in our following analysis. As for other types of cells, such as supporting cells, mantle cells, and epithelial cells, they are GFP-negative (Fig. S3), but they cannot be filtered out in our FACS sorting. The possible mechanism involved is that they interact with the GFP-positive cells, namely hair cells and retinal ganglion cells, and they were also sorted, coupling with GFP-positive cells.
Furthermore, we performed a comparison among these different subtypes of hair cells, and found that the neuromast hair cells have stronger expression of MET components and energy metabolism-related activity compared to the inner ear hair cells. Moreover, we analyzed the relationship between the known human NSHL genes and zebrafish hair cell marker genes revealed by the scRNA-seq, and found that 42.86% of human NSHL genes have orthologous genes expressed in zebrafish hair cells, which further reflects the reliability of our data. On the other hand, there are plenty of genes in our hair cell scRNA-seq gene pool that have not been reported to function in hair cells, which raises the possibility that these genes have potential value in hair cell function and even hearing loss gene identification. We randomly picked some of the marker genes for further analysis, and found that mb and capgb gene were specifically expressed in zebrafish inner ear macula hair cells and lateral line neuromast hair cells, respectively. Gene functional analysis also demonstrated that these two genes were required for hair cell development or function; for that knockdown of these two genes can lead to hair cell loss and hair cell dysfunction.
As far as we know, myoglobin, encoded by the Mb gene, is a single-chain heme protein containing 153 amino acids. It mainly exists in cardiac and skeletal muscle, and recent studies showed that it also exists in a variety of non-muscle tissues, such as the brain, kidney, gill, and liver [36]. Myoglobin functions in transporting and storing oxygen in muscle cells, and it can also promote the removal of reactive oxygen species (ROS) and NO in cancer cells [37]. However, the role of myoglobin in hair cells was not reported before, and it is the first time that the mb gene was proved to be expressed in zebrafish inner ear hair cells and important for hair cell development and function in our current study. Further investigation was needed to uncover the molecular mechanisms involved in hair cell function.
CAPG, encoding a gelsolin-like capping protein, as a proto-oncogene [38], is involved in the migration and invasion of various cancer cells [39,40,41,42]. Recently, a rare homozygous deletion of the chromosome 2p11.2 region was found in a Tunisian patient with autism, intellectual disability, and hearing impairment, and the affected genes mainly included ELMOD3, CAPG, and SH2D6 [43]. In our study, we found that the capgb gene was mainly expressed in zebrafish neuromast hair cells, and essential for hearing, indicating its crucial role in auditory function. Nevertheless, how CAPG contributes to hair cell development, as well as hearing, merits further investigation.
To sum up, in this study, we identified three subpopulations of hair cells, which corresponded with macula hair cells, crista hair cells, and neuromast hair cells in zebrafish, through scRNA-seq. In addition, it also uncovered thousands of genes in zebrafish hair cells, which would be helpful for our research in hair cell biology in the future.
Based on differential gene expression, Lush., et al. [17] found that hair cells can be subdivided into young hair cells and mature hair cells, and these two different hair cells have distinct distribution and gene expression that the young hair cells form a ring and express atoh1b. Differently, our current work revealed the molecular difference among three different types of hair cells and specific marker genes for the first time, which could provide further insights into understanding the mechanisms underlying hearing and balance.
Materials and methods
Zebrafish embryos
Zebrafish were maintained at 28.5 °C. Two zebrafish lines were used in the study, including the wild-type AB and the transgenic line Tg(Brn3c:mGFP), which was described in the previous work [19]. All animal procedures were performed according to protocols approved by the Animal Care and Use Committee of Nantong University and were consistent with the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Single-cell RNA-sequencing
The Tg(Brn3c:mGFP) transgenic zebrafish larvae at 6 days post-fertilization (dpf) were anesthetized and then treated with 0.25% trypsin to dissociate into single cells, which were divided into GFP-positive and GFP-negative cells through the fluorescent activated cell sorter, and the RNA expressed in GFP-positive cells were then obtained and sequenced using the 10 × Genomics platform.
Single-cell sequencing analysis
The basic procedure for single-cell sequencing analysis was carried out as previously described [44]. Briefly, Seurat V4.0.1 [20] was used for the integrated analysis of the single-cell sequencing data, including data filtering, data normalization, cell clustering, and cluster-level marker gene identification. First, the Seurat object was created by “CreateSeuratObject” function and the raw data were primarily filtered through the parameter settings “min.cells = 5,min.features = 200”, which required that only the genes expressed in at least five cells were considered and only the cells with minimal 200 genes detected were kept. Then, the data were further filtered through the “subset” function in Seurat with the parameter settings “nFeature_RNA < 5000 & percent.mt < 10”, which required the cells containing no more than 10% reads from mitochondrial genes and up to 5000 detected genes. Second, “SCTransform” function in Seurat was applied to data normalization, and the data were regressed by several factors(percentage of mitochondrial expression, total number of UMIs, and total number of genes detected) through a second non-regularized linear regression (variable.features.n = 1000,vars.to.regress = c(“nFeature_RNA”,“nCount_RNA”,“percent.mt”)). For cell clustering, the principal component analysis (PCA) was first applied for extracting the top principal components (top 50 PCs), and then, the clusters were identified through shared k-nearest neighbor graph construction and cluster modularity function optimization (k.param = 20, resolution = 0.25). To identify marker genes for each cell type, “FindAllMarkers” function in Seurat was used with the parameter settings “only.pos = TRUE, min.pct = 0.25,logfc.threshold = 0.25”. The default Wilcoxon rank-sum test in “FindAllMarkers” function was selected for differential expressed gene detection between clusters, where the p values were further adjusted by Bonferroni correction.
For GO enrichment analysis, the R package clusterProfiler [45] was used, and the marker genes identified by Seurat were used as input. The three GO categories: BP (Biological Process), CC (Cellular Component), and MF (Molecular Function) were analyzed respectively against the marker genes for each cell cluster, and the p values were corrected by Benjamini & Hochberg (BH) method, where the significantly enriched GO terms were defined as the ones with BH adjusted p value less than 0.05. The GO enrichment results were further visualized through the enrichplot method integrated in clusterProfiler.
Whole-mount in situ hybridization
The whole-mount in situ hybridization procedure was similar to our previous description [46, 47]. In brief, the gene-specific primers targeting the coding sequence of interest were designed and synthesized. After amplification using the PCR method, the fragments were subcloned into the pGEM-T Easy vector (Promega). The DIG-labeling RNA probe was transcribed in vitro using the linearized recombinant plasmid as a template. For the hybridization step, the pretreated zebrafish embryos were incubated with the digoxigenin-labeling RNA probe overnight first, and then, alkaline phosphatase-conjugated primary antibody against digoxigenin was used to detect the RNA probe following three washes. Subsequently, for color reaction, the nonspecific binding was washed out and the substrate of alkaline phosphatase, NBT/BCIP solution, was added to the reaction system. Finally, the samples were imaged and the gene-specific mRNA expression was visualized through the microscope. The primers used for probe synthesis are as follows:
tectb-F: GCCGTGTTTAGCCAAAGAGT;
tectb-R: ACAACTTCTCCCCGTCACAT;
zpld1a-F: CCACGGTTGTGCTGTTTAGC;
zpld1a-R: GTTGGTGGCTTTTACAGCGG;
calm1b-F: AAGTGTTGTGTCGTGGGGTT;
calm1b-R: GCAGGGGAGATTTCACTGGT;
capgb-F: ACCTGGTGCTGGATAACAGG;
capgb-R: ATCTGAGCTTTGCCGTGTCT;
C5H11orf1-F: TGATGTCCAATAAAAGCCAGGT;
C5H11orf1-R: CACACATTGAGGCTCTGAAGT;
zdhhc16b-F: CCTGTGGAATTATGGGATGG;
zdhhc16b-R: ATGCTGCAGTGATGAGTTCG;
tekt3-F: AGATTTCAGCGCTGTCCGAT;
tekt3-R: AAGCAGCACGTTCACTCTGA;
mb-F: TGATCTGGTTCTGAAGTGCTG;
mb-R: GGCAAATCCGATCTCCTTGT;
s100s-F: CCAAGATGCCACGCTCAAAG;
s100s-R: CCCGCTAACACTTCTCTCGG.
Morpholino-mediated gene knockdown
The gene-specific morpholinos (MOs) were synthesized by the Gene Tools, LLC. For gene knockdown, the 2–3 nL of 0.3 mM of gene-specific morpholinos were microinjected into the 1–2-cell-stage zebrafish embryos. Here, the morpholinos were used to block the splicing of the pre-mRNA, and therefore down-regulate the target gene expression. The sequences of the morpholinos used in this study were as follows:
capgb-MO: TCTGGAGGAACAAAGATGAGATGGT;
mb-MO: ATCAGAGAGTCCTGCTTTACCCTGA.
Reverse transcription-polymerase chain reaction (RT-PCR)
The RT-PCR was performed following the standard procedure. Briefly, the total RNA was extracted from the zebrafish embryos and then reverse-transcribed into cDNA. The PCR experiment was performed to detect the gene expression using the cDNA as a template. Herein, the primers used for PCR were listed as follows:
capgb-F: TCTGACAGCATGCCGGAGC;
capgb-R: TAACATTGGTGATCTGAGCTTTGC;
mb-F: GACTTTTCCAAAGCCACAGGC;
mb-R: TCCTGAGACCCTAACGAACCA.
mRNA rescue experiment
The rescue experiment was performed by coinjecting the specific mRNA with the morpholino. The mRNA was transcribed in vitro using the linearized recombinant plasmid as a template, which contains the coding sequence of specific gene. The primers used in the PCR to amplify the specific genes were as follows:
capgb-mRNA-F: ACCTAGGTGCAGGAACAGG;
capgb-mRNA-R: ACAACTGGTTGAGTGCAGTTTA;
mb-mRNA-F: GCCCCGATATTGAAGACAGGT;
mb-mRNA-R: TGACTCCCATTTGAGATCTGGT.
Startle response test
The startle response test in zebrafish was carried out following the procedure [48, 49]. In this test, 20 zebrafish larvae at 5 dpf were put into the culture dish for free swimming. When acoustic stimuli occurred, the behavior of the zebrafish was recorded by a high-speed camera (500 fps). In response to the stimuli, the zebrafish with normal auditory function would have a characteristic C-bend motion lasting less than 10 ms; however, the zebrafish with damaged auditory function would not. Herein, both the distance moved and the number of the C-bend motions were used to quantify the startle response.
VOR test
As previously described [49, 50], the VOR test was performed according to standard procedure. Briefly, the zebrafish larva at 5 dpf was fixed in the chamber in a head-up position with 5% methylcellulose. The rotary platform, with the chamber unit fixed, rotated back and forth at a speed of 30 rpm. The eye movements of the zebrafish were recorded by an infrared camera, and the periodical changes of the projection area of the eyes were used to evaluate the VOR.
Imaging and statistical analysis
For the confocal fluorescence microscopic analysis, the zebrafish were embedded in the 0.8% low melt agarose following anesthetization with MS-222, and the imaging was carried out by the Nikon A1 microscope. The Olympus MVX10 microscope was used for bright-field imaging in the WISH experiments. All data were presented as mean with SD, and the unpaired Student’s t tests were used to determine statistical significance. A value of P < 0.05 was considered statistically significant.
Availability of data and materials
All the high-throughput sequencing data generated in this study have been deposited in the Gene Expression Omnibus database under accession number GSE221471 and are available at the following URL: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85337. All the experimental materials generated in this study are available from the corresponding authors upon reasonable request.
Change history
18 May 2023
Text in the Availability of data and materials has been updated.
12 May 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00018-023-04780-1
References
Taylor RR, Filia A, Paredes U, Asai Y, Holt JR, Lovett M, Forge A (2018) Regenerating hair cells in vestibular sensory epithelia from humans. Elife. https://doi.org/10.7554/eLife.34817
Kurima K, Ebrahim S, Pan B, Sedlacek M, Sengupta P, Millis BA, Cui R, Nakanishi H, Fujikawa T, Kawashima Y, Choi BY, Monahan K, Holt JR, Griffith AJ, Kachar B (2015) TMC1 and TMC2 localize at the site of mechanotransduction in mammalian inner ear hair cell stereocilia. Cell Rep 12(10):1606–1617
Wagner EL, Shin JB (2019) Mechanisms of hair cell damage and repair. Trends Neurosci 42(6):414–424
Nordang L, Cestreicher E, Arnold W, Anniko M (2000) Glutamate is the afferent neurotransmitter in the human cochlea. Acta Otolaryngol 120(3):359–362
Dalet A, Bonsacquet J, Gaboyard-Niay S, Calin-Jageman I, Chidavaenzi RL, Venteo S, Desmadryl G, Goldberg JM, Lysakowski A, Chabbert C (2012) Glutamate transporters EAAT4 and EAAT5 are expressed in vestibular hair cells and calyx endings. PLoS ONE 7(9):e46261
Highstein SM, Holstein GR, Mann MA, Rabbitt RD (2014) Evidence that protons act as neurotransmitters at vestibular hair cell-calyx afferent synapses. Proc Natl Acad Sci USA 111(14):5421–5426
Dallos P (1992) The active cochlea. J Neurosci 12(12):4575–4585
Burns JC, Stone JS (2017) Development and regeneration of vestibular hair cells in mammals. Semin Cell Dev Biol 65:96–105
Day BL, Fitzpatrick RC (2005) The vestibular system. Curr Biol 15(15):R583–R586
Patton EE, Zon LI, Langenau DM (2021) Zebrafish disease models in drug discovery: from preclinical modelling to clinical trials. Nat Rev Drug Discov. https://doi.org/10.1038/s41573-021-00210-8 (1474-1784 (Electronic))
Riley BB, Moorman SJ (2000) Development of utricular otoliths, but not saccular otoliths, is necessary for vestibular function and survival in zebrafish. J Neurobiol 43(4):329–337
Yao Q, DeSmidt AA, Tekin M, Liu X, Lu Z (2016) Hearing assessment in zebrafish during the first week postfertilization. Zebrafish 13(2):79–86
Smith ET, Pacentine I, Shipman A, Hill M, Nicolson T (2020) Disruption of tmc1/2a/2b genes in zebrafish reveals subunit requirements in subtypes of inner ear hair cells. J Neurosci 40(23):4457–4468
Pacentine IV, Nicolson T (2019) Subunits of the mechano-electrical transduction channel, Tmc1/2b, require Tmie to localize in zebrafish sensory hair cells. PLoS Genet 15(2):e1007635
Erickson T, Pacentine IV, Venuto A, Clemens R, Nicolson T (2019) The lhfpl5 Ohnologs lhfpl5a and lhfpl5b are required for mechanotransduction in distinct populations of sensory hair cells in zebrafish. Front Mol Neurosci 12:320
Navajas Acedo J, Voas MG, Alexander R, Woolley T, Unruh JR, Li H, Moens C, Piotrowski T (2019) PCP and Wnt pathway components act in parallel during zebrafish mechanosensory hair cell orientation. Nat Commun 10(1):3993
Lush ME, Diaz DC, Koenecke N, Baek S, Boldt H, St Peter MK, Gaitan-Escudero T, Romero-Carvajal A, Busch-Nentwich EM, Perera AG, Hall KE, Peak A, Haug JS, Piotrowski T (2019) scRNA-Seq reveals distinct stem cell populations that drive hair cell regeneration after loss of Fgf and Notch signaling. Elife. https://doi.org/10.7554/eLife.44431
Kozlovskaja-Gumbriene A, Yi R, Alexander R, Aman A, Jiskra R, Nagelberg D, Knaut H, McClain M, Piotrowski T (2017) Proliferation-independent regulation of organ size by Fgf/Notch signaling. Elife. https://doi.org/10.7554/eLife.21049
Xiao T, Roeser T, Staub W, Baier H (2005) A GFP-based genetic screen reveals mutations that disrupt the architecture of the zebrafish retinotectal projection. Development 132(13):2955–2967
Huo L, Jiao Li J, Chen L, Yu Z, Hutvagner G, Li J (2021) Single-cell multi-omics sequencing: application trends, COVID-19, data analysis issues and prospects. Brief Bioinform. https://doi.org/10.1093/bib/bbab229
Yang CH, Cheng CH, Chen GD, Liao WH, Chen YC, Huang KY, Hwang PP, Hwang SP, Huang CJ (2011) Zona pellucida domain-containing protein β-tectorin is crucial for zebrafish proper inner ear development. PLoS ONE 6(8):e23078
Kozak EL, Palit S, Miranda-Rodriguez JR, Janjic A, Bottcher A, Lickert H, Enard W, Theis FJ, Lopez-Schier H (2020) Epithelial planar bipolarity emerges from notch-mediated asymmetric inhibition of Emx2. Curr Biol 30(6):1142-1151 e6
Pistocchi A, Feijoo CG, Cabrera P, Villablanca EJ, Allende ML, Cotelli F (2009) The zebrafish prospero homolog prox1 is required for mechanosensory hair cell differentiation and functionality in the lateral line. BMC Dev Biol 9:58
Aman A, Piotrowski T (2008) Wnt/beta-catenin and Fgf signaling control collective cell migration by restricting chemokine receptor expression. Dev Cell 15(5):749–761
Seiler C, Ben-David O, Sidi S, Hendrich O, Rusch A, Burnside B, Avraham KB, Nicolson T (2004) Myosin VI is required for structural integrity of the apical surface of sensory hair cells in zebrafish. Dev Biol 272(2):328–338
Ernest S, Rosa FM (2015) A genomic region encompassing a newly identified exon provides enhancing activity sufficient for normal myo7aa expression in zebrafish sensory hair cells. Dev Neurobiol 75(9):961–983
Wu Z, Muller U (2016) Molecular identity of the mechanotransduction channel in hair cells: not quiet there yet. J Neurosci 36(43):10927–10934
Qiu X, Muller U (2018) Mechanically gated ion channels in mammalian hair cells. Front Cell Neurosci 12:100
Xiong W, Grillet N, Elledge HM, Wagner TF, Zhao B, Johnson KR, Kazmierczak P, Muller U (2012) TMHS is an integral component of the mechanotransduction machinery of cochlear hair cells. Cell 151(6):1283–1295
Giese APJ, Tang YQ, Sinha GP, Bowl MR, Goldring AC, Parker A, Freeman MJ, Brown SDM, Riazuddin S, Fettiplace R, Schafer WR, Frolenkov GI, Ahmed ZM (2017) CIB2 interacts with TMC1 and TMC2 and is essential for mechanotransduction in auditory hair cells. Nat Commun 8(1):43
Wang Y, Li J, Yao X, Li W, Du H, Tang M, Xiong W, Chai R, Xu Z (2017) Loss of CIB2 causes profound hearing loss and abolishes mechanoelectrical transduction in mice. Front Mol Neurosci 10:401
Sollner C, Rauch GJ, Siemens J, Geisler R, Schuster SC, Muller U, Nicolson T (2004) Tubingen screen, consortium, mutations in cadherin 23 affect tip links in zebrafish sensory hair cells. Nature 428(6986):955–959
Sakaguchi H, Tokita J, Muller U, Kachar B (2009) Tip links in hair cells: molecular composition and role in hearing loss. Curr Opin Otolaryngol Head Neck Surg 17(5):388–393
Choudhary D, Narui Y, Neel BL, Wimalasena LN, Klanseck CF, De-la-Torre P, Chen C, Araya-Secchi R, Tamilselvan E, Sotomayor M (2020) Structural determinants of protocadherin-15 mechanics and function in hearing and balance perception. Proc Natl Acad Sci USA 117(40):24837–24848
Zhang S, Dong Y, Qiang R, Zhang Y, Zhang X, Chen Y, Jiang P, Ma X, Wu L, Ai J, Gao X, Wang P, Chen J, Chai R (2021) Characterization of strip1 expression in mouse cochlear hair cells. Front Genet 12:625867
Cossins AR, Williams DR, Foulkes NS, Berenbrink M, Kipar A (2009) Diverse cell-specific expression of myoglobin isoforms in brain, kidney, gill and liver of the hypoxia-tolerant carp and zebrafish. J Exp Biol 212(5):627–638
Quinting T, Heymann AK, Bicker A, Nauth T, Bernardini A, Hankeln T, Fandrey J, Schreiber T (2021) Myoglobin protects breast cancer cells due to Its ROS and NO scavenging properties. Front Endocrinol (Lausanne) 12:732190
Chen ZF, Huang ZH, Chen SJ, Jiang YD, Qin ZK, Zheng SB, Chen T (2021) Oncogenic potential of macrophage-capping protein in clear cell renal cell carcinoma. Mol Med Rep. https://doi.org/10.3892/mmr.2020.11718
Jiang S, Yang Y, Zhang Y, Ye Q, Song J, Zheng M, Li X (2022) Overexpression of CAPG Is associated with poor prognosis and immunosuppressive cell infiltration in ovarian cancer. Dis Markers 2022:9719671
Lang Z, Chen Y, Zhu H, Sun Y, Zhang H, Huang J, Zou Z (2019) Prognostic and clinicopathological significance of CapG in various cancers: Evidence from a meta-analysis. Pathol Res Pract 215(12):152683
Nader JS, Boissard A, Henry C, Valo I, Verrièle V, Grégoire M, Coqueret O, Guette C, Pouliquen DL (2020) Cross-species proteomics identifies CAPG and SBP1 as crucial invasiveness biomarkers in rat and human malignant mesothelioma. Cancers (Basel) 12(9):2430
Prescher N, Hänsch S, Knobbe-Thomsen CB, Stühler K, Poschmann G (2021) The migration behavior of human glioblastoma cells is influenced by the redox-sensitive human macrophage capping protein CAPG. Free Radic Biol Med 167:81–93
Lahbib S, Leblond CS, Hamza M, Regnault B, Lemee L, Mathieu A, Jaouadi H, Mkaouar R, Youssef-Turki IB, Belhadj A, Kraoua I, Bourgeron T, Abdelhak S (2019) Homozygous 2p11.2 deletion supports the implication of ELMOD3 in hearing loss and reveals the potential association of CAPG with ASD/ID etiology. J Appl Genet 60(1):49–56
Shi J, Fok KL, Dai P, Qiao F, Zhang M, Liu H, Sang M, Ye M, Liu Y, Zhou Y, Wang C, Sun F, Xie G, Chen H (2021) Spatio-temporal landscape of mouse epididymal cells and specific mitochondria-rich segments defined by large-scale single-cell RNA-seq. Cell Discov 7(1):34
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287
Qian F, Wang X, Yin Z, Xie G, Yuan H, Liu D, Chai R (2020) The slc4a2b gene is required for hair cell development in zebrafish. Aging (Albany NY) 12(19):18804–18821
Zhang L, Gao Y, Zhang R, Sun F, Cheng C, Qian F, Duan X, Wei G, Sun C, Pang X, Chen P, Chai R, Yang T, Wu H, Liu D (2020) THOC1 deficiency leads to late-onset nonsyndromic hearing loss through p53-mediated hair cell apoptosis. PLoS Genet 16(8):e1008953
Wang C, Zhong Z, Sun P, Zhong H, Li H, Chen F (2017) Evaluation of the hair cell regeneration in zebrafish larvae by measuring and quantifying the startle responses. Neural Plast 2017:8283075
Gong J, Qian P, Hu Y, Guo C, Wei G, Wang C, Cai C, Wang H, Liu D, Claudin H (2021) Is essential for hair cell morphogenesis and auditory function in zebrafish. Front Cell Dev Biol 9:663995
Sun P, Zhang Y, Zhao F, Wu JP, Pun SH, Peng C, Du M, Vai MI, Liu D, Chen F (2018) An assay for systematically quantifying the vestibulo-ocular reflex to assess vestibular function in zebrafish larvae. Front Cell Neurosci 12:257
Funding
This work was supported in part by grants from the National Natural Science Foundation of China, http://www.nsfc.gov.cn (2018YFA0801004 and 81870359 received by Dong Liu; 31900484 to Gangcai Xie); Natural Science Foundation of Jiangsu Province, http://kjjh.jspc.org.cn (BK20180048 and BRA2019278 received by Dong Liu; BK20190924 to Gangcai Xie; BK20190920 to Guanyun Wei). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
FQ, DL, GX, and RC conceived and designed the experiments, and wrote the manuscript. FQ, YG, XW, JG, CG, XW, XZ, JZ, CW, MX, YH, and GY performed the experiments. FQ, GW, JK, GX, DL, and RC analyzed the data. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval and consent to participate
All zebrafish experimentation was carried out in accordance with the NIH Guidelines for the care and use of laboratory animals (http://oacu.od.nih.gov/regs/index.htm) and ethically approved by the Administration Committee of Experimental Animals, Jiangsu Province, China (Approval ID: 20180405-039).
Consent for publication
The authors declare the consent for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
18_2022_4410_MOESM1_ESM.pdf
Supplementary file1 Fig. S1. The procedure of single cell preparation and cell sorting using the FACS method. Fig. S2. The FACS sorting data. Cells of Tg(Brn3c:mGFP) zebrafish at 6 dpf were sorted. Fig. S3. The expression pattern of GFP, tectb and zpld1a gene in sequenced cells. The feature plot and violin plot are shown. Fig. S4. The expression patterns of the cluster 0- and cluster 7-enriched genes. The feature plot and violin plot are shown for each gene. Fig. S5. The expression pattern of the cluster 1-enriched gene. The feature plot and violin plot are shown for each gene. Fig. S6. The expression patterns of the cluster 9-enriched genes. The feature plot and violin plot are shown for each gene. Fig. S7. The expression patterns of some hair cell marker genes. The feature plot and violin plot are shown for each gene. Fig. S8. The expression pattern of capgb and mb gene. The feature plot and violin plot are shown for each gene. Fig. S9. The verification of gene knockdown using RT-PCR. The gene expression was reduced in capgb-morphants and mb-morphants compared to the control (Ctrl). The white dotted boxes indicate the gene expression detected. (PDF 1521 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qian, F., Wei, G., Gao, Y. et al. Single-cell RNA-sequencing of zebrafish hair cells reveals novel genes potentially involved in hearing loss. Cell. Mol. Life Sci. 79, 385 (2022). https://doi.org/10.1007/s00018-022-04410-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04410-2