Abstract
The microbiota plays a crucial role in regulating various physiological functions and pathological conditions within the human body. An important aspect of COVID-19 pathogenesis is comprehending how various infections in the body, including COVID-19, affect and influence the microbiome. We may develop better diagnostics and strategies against COVID-19 infection by examining the association between the intestinal and respiratory microbiota. To take a broader scientific approach, we must answer several key questions, such as how microbiome diversity and composition vary from person to person, how accumulated microbiota can benefit individuals over time, and what factors contribute to microbiota development. Analyzing the signaling molecules that mediate biological mechanisms for immune responses between the host and microbiota and among microbiota may provide valuable insight. Several potential therapies to improve the microbiome or target specific microbiota include phage therapy, FMT, prebiotics, probiotics, and synbiotics. Modulating the neonatal microbiome has been a challenging goal to increase efficacy recently. However, much more research is required to engineer microbiome therapeutics. This chapter provides an overview of existing challenges and strategies to make the necessary modifications to restore the naive gut.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Microbiome
The word “microbiome” is the collection of genomes from all the microorganisms in an environment. The variation in the microbiota depends on the location of specific microbiota in the body. For instance, the gut microbiota differs from the skin microbiota (Sanapala and Pola 2021). The microbiome colonizes the human body in different roles and varies between individuals. The microbiome plays a major role in various physicochemical activities like forming tight junctions between the cells to maintain the integrity of the tissues, enhancing the production of T cells to boost the immune system, and controlling the body’s metabolism (Pola and Padi 2021). The human microbiome is currently associated with several disorders, including inflammatory bowel disease, type 2 diabetes, Parkinson’s disease, hypothyroidism, colorectal cancer, COPD, and rheumatoid arthritis. Other respiratory diseases are directly or indirectly associated with specific microorganism patterns (Yamamoto et al. 2021).
Microbiome formation is influenced by genetics (Mohan and Sudhakar 2022), delivery mode, infant feeding, nutrition, antibiotic administration, age, medication, vaccination (health complications), and geographical and seasonal differences. Diseased conditions in the human body impact microbiome composition. The areas where the microbiota plays a key role in humans are nutrition, immunity, behavior, and disease (Harper et al. 2021). The beneficial microbiota in the gut can help digest food that humans can’t break down. The harmful microbiota can damage the immune system and make the body prone to various conditions like gastrointestinal diseases. As an example, the presence of the microbiome genus Pseudomonadales and Streptococcus is linked to human upper respiratory tract infections in patients.
Consuming certain live bacterial strains confers health benefits on the human body. These bacteria are known as probiotics, and the administration of probiotics helps reduce many bacterial infections. Some probiotics have also been engineered to kill pathogenic bacteria, known as “smart microbes” (Ronda et al. 2019). For instance, Lactococcus lactis was modified to create molecules that would attack Enterococcus faecium, a bacterium linked to the onset of meningitis in infants. Researchers have laid the groundwork for a future technique to alter microbes in an individual’s gut.
The production of smart microbes as probiotics can be engineered using MAGIC (metagenomic alteration of gut microbiome by in situ conjugation). This method includes oral ingestion of bacteria that can transfer DNA with specific traits to the bacterial microbiome already present in the body (Baghbani et al. 2020). This method needs a lot of improvements to make use of it to treat infectious diseases in humans, increase the persistence of DNA in the gut, and ensure that the DNA can only be transferred to targeted, nonpathogenic strains of the microbiome.
2 Microbiomes of Infectious Disease
Microbiome research focuses on the microbial communities’ behavior, interactions, and functions within a specified environment. The microbiota has enzymes that cannot be coded by the human genome but are necessary to fulfill some physiologic tasks, like the digestive enzymes that break down substances like polysaccharides and polyphenols and hydrolytic enzymes to regulate and balance cellular metabolism. The microbiota’s function within a healthy host involves influencing diverse pathogens through colonization resistance, allowing the host’s immune system to participate in immune cell differentiation, promoting the proliferation of granulocyte/monocyte progenitors, activating innate lymphoid cells and myeloid cells, triggering proinflammatory T- and B-cell responses, and initiating pre-inflammatory T- and B-cell secretion of SIGA (secretory IgA). Both secretory IgA and gut inflammation modify the microbiota composition, resulting in shifts in microbial proportions and increased pathogen growth.
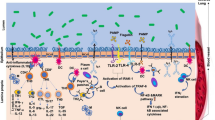
Firstly, the microbiota induces alpha-defensin, beta-defensin, C-type lectins, secretory IgA, and other AMPs (antimicrobial peptides) that affect the immune system (innate and adaptive) through intestinal epithelia and paneth cell receptors; macrophage cell receptors such as TLRs or NLRs; and CCR6. TLRs are important for developing the mucosal and intestinal immune systems, decreasing inflammatory responses, and promoting immunological tolerance to the necessary microbial components. NLRs regulate IL-18 levels, immune responses (Konatala et al. 2021), dysbiosis, and intestinal hyperplasia. When antigens of the microbiota bind to these receptors, they start a chain reaction of signaling pathways that release antimicrobial compounds like defensins and stimulate T cells like T-helper 1 and 17 to make IL-1, IL-15, IL-17, IL-22, etc. They also stimulate B cells to produce antibodies (Harper et al. 2021) (Fig. 8.1).
3 Effects of the Microbiome in the COVID-19 Infection
COVID-19 is a respiratory disease ranging from mild (cough and/or fever) to severe pneumonia (ARDS and multiple organ failure). Angiotensin-converting enzyme 2 (ACE2) is the receptor to which the viral spike binds. This receptor is expressed on the respiratory and gastrointestinal epithelium, leading to changes in its microbiome composition during the infection (Liu et al. 2022).
SARS-CoV-2 is an enveloped, single-stranded RNA virus consisting of structural proteins like the nucleocapsid, membrane, envelope, and spike proteins. The viral particles mediated by the S glycoprotein attach and fuse to the host cell membrane and get inserted into the virion membrane in multiple copies with a crown-like appearance. The S protein of coronaviruses is cleaved into S1 and S2 subunits by proprotein convertases during their biosynthesis or after reaching their target site.
ACE2 is an 805-amino acid carboxypeptidase, and the downregulation of ACE2 leads to a severe disturbance in the renin-angiotensin-aldosterone system. As shown in Fig. 8.2, the virus bound to the ACE2 receptor induces conformational changes in both S1 and S2 subunits. The conformational changes in the S1 subunit expose the S2 cleavage site of the S2 subunit. During insufficient transmembrane protease, serine 2 (TMPRSS2), the virus-ACE2 complex triggers clathrin-mediated endocytosis, where cathepsins perform S2′ cleavage in endolysosomes. Cleavage of the S2′ site in the presence of TMPRSS2 exposes the fusion peptide, and the separation of S1 from S2 induces conformational changes in the S2 subunit and triggers fusion to the membrane. The fusion between the viral membrane and cellular membranes forms a fusion pore through which viral RNA is released into the host cell cytoplasm.
The other molecules that serve as receptors in SARS-CoV-2 infection are C-type lectins, DC-SIGN and L-SIGN, AXL and TIM1, phosphatidylserine receptors TIM and TAM, and CD147 (a transmembrane glycoprotein). Lectins bind to the surface (glycans) of the virion, promoting viral entry and intracellular adhesion. TIM and TAM bind to phosphatidylserine on the virion membrane to promote the entry of enveloped viruses. Increased viral entry was observed with higher levels of CD147 in SARS-CoV-2 and is a potential risk factor as it was upregulated in diabetic and obese patients. A remarkable modification of the nasopharyngeal microbiome is noticed in this study and assumed to have a proportional dysbiosis effect from the onset, treatment, and reduction of COVID-19 infection (Hoque et al. 2021) (Fig. 8.3).
Mechanism of SARS-COV-2 entry into cells (Jackson et al. 2022)
4 Variations in the Upper and Lower Respiratory Tract Microbiomes in the COVID-19 Patients
Upper vs. lower respiratory tract sampling, collection time, point or stage of infection, treatment with broad-spectrum antibiotics, invasive mechanical ventilation, and prolonged hospitalization are the vital factors that can determine the range and composition of the microbiome at specific sites. The respiratory tract microbiome of COVID-19 patients was analyzed in two experimental studies (using nasopharyngeal swabs and sputum) and detected using next-generation sequencing.
The oral and upper respiratory microbiomes include Candida albicans and human alpha-herpesvirus, which are the most common, and coinfection of COVID-19 with other viruses like Influenza A/B, enteroviruses, Aspergillus respiratory syncytial virus, and Veillonella species was found in some of the patients. Certain research studies identified microbiome makeup within COVID-19 throat samples, encompassing Haemophilus parainfluenzae, Neisseria cinerea, rhinovirus, Streptococcus mitis, Streptococcus bovis, Leptotrichia buccalis, and Rothia mucilaginosa. Additionally, bronchoalveolar samples exhibited Acinetobacter, Pseudomonas, Enterococcus, Lactobacillus, and Chryseobacterium. Some studies show that the diversity of pharyngeal microbiome (e.g., Bacteroidetes, Proteobacteria) decreased in older adult patients than in younger adult patients suffering from COVID-19 (Gaibani et al. 2021).
The lower respiratory microbiome in COVID-19 patients showed potentially pathogenic microorganisms like Candida albicans, human influenza virus, and human alpha-herpesvirus identified in the nasopharyngeal microbiome. Some studies described that the main microbiome composition in the throat of COVID-19 patients includes Streptococcus bovis, Leptotrichia buccalis, Haemophilus parainfluenzae, and Neisseria cinerea.
Whole-genome sequencing of BALF samples showed dysbiosis in oral and upper respiratory bacteria, which includes bacteria like Acinetobacter, Sphingobium, Enterobacterales, Escherichia coli, Enterococcus, Rothia, and Lactobacillus. They also identified fungi like Cryptococcus, Cladosporium, and Alternaria in the patients. A study based on 16S rRNA sequences indicates that Acinetobacter was the most common bacterial genus, followed by Chryseobacterium, Burkholderia, Brevundimonas, Sphingobium, and Enterobacterales in the lung tissues of deceased COVID-19 patients. Cryptococcus was identified as a prevalent fungus along with other species like Issatchenkia, Wallemia, Cladosporium, and Alternaria. Patients with both moderate and severe COVID-19 showed dysbiotic microbiomes and those treated with various antibiotics, under mechanical ventilation, and prolonged hospitalization.
The severity of SARS-CoV-2 infection-induced changes in the microbiome’s diversity at the entry site of infection was high. Still, patients with mild illness did not show any significant changes in their diversity compared to healthy patients or patients with other viral respiratory tract infections. The patients admitted to the hospital had reduced microbiome diversity and increased dysbiosis of the oropharyngeal and nasopharyngeal microbiome. A detailed study of patients admitted to an ICU for different medical conditions showed differences in the oral and gut microbiomes, which were found to have major microbial dysbiosis disturbing an individual’s complete respiratory microbiome equilibrium (de Castilhos et al. 2022).
In healthy individuals, bacteria in the oral cavity include Actinobacteria (e.g., Corynebacterium spp., Propionibacterium spp.), Firmicutes (e.g., Staphylococcus spp.), and Proteobacteria. The nasopharyngeal system contains members of Firmicutes, Staphylococcus; Bacteroidetes, Corynebacterium; Proteobacteria, Prevotella; and commensal bacteria like Streptococcus, Neisseria, and Haemophilus spp. in the URT (upper respiratory tract) (Fig. 8.4).
Acinetobacter spp., Clostridium hiranonis, Enterobacteriaceae, Pseudomonas alcaligenes, and Sphingobacterium spp. were found in the lungs of critically ill COVID-19 patients. In contrast, Haemophilus influenzae, Streptococcus spp., and Veillonella dispar were found in the lungs of COVID-19-negative patients. The lung microbiota profile of critically ill patients with COVID-19 was dominated by the phyla Bacteroidetes (9%), Firmicutes (37%), and Proteobacteria (48%) (Hanada et al. 2018).
5 Impact of COVID-19 Infection on the Intestinal Microbiome
There are evident changes in the microbiome composition of the intestinal and respiratory microbiota during the infection. Some studies on COVID-19 infection have shown that the gut microbiota has more importance than the respiratory microbiota, as any change in the composition of the gut microbiota would adversely affect lung function and lead to gut dysbiosis. Another research study explains that any imbalance in the gut microbiome can result in illnesses of human health, indirectly affecting immunity (Jackson et al. 2022).
Macronutrient metabolism includes metabolites like short-chain fatty acids, alcohol, branched-chain fatty acids (acetate, propionate, and butyrate), amines, indoles, sulfur compounds, phenols, and glycerol and choline derivatives of the gut microbiome that influence human health. The commensals of gut microbiota like Bacteroides and Bifidobacteria secrete metabolites and immune signaling molecules that bind to receptors in innate cells such as dendritic cells and macrophages to modulate their functions. This helps regulate the development and function of the innate and adaptive immune systems, secreting antimicrobial peptides that can kill microbial pathogens directly or indirectly by modulating the host defense systems by competing for nutrients and the habitat site and maintaining the homeostasis of the body. The disruptions caused by the microbiota may alter the mechanisms of colonization resistance and affect the outcomes of infection. For example, dietary fiber can modify the microbiota’s structure and function by producing SCFAs, such as acetate, butyrate, and propionate. They bind to the G-protein-coupled receptor (GPR43) and can stimulate AMPs, REGIIIγ and β-defensins, as shown in Fig. 8.5. SCFAs can diffuse through the membrane, acidify cytoplasm, and inhibit the growth of some pathogens. Alterations to the microbiota (due to antibiotics or a high-fat diet) can result in lowered B-cell-modulated production of IgA68 and increased permeability, leading to susceptibility to infection.
The studies showed that patients treated with antibiotics during hospitalization had a further depletion of bacterial species (symbionts) beneficial to host immunity. Figure 8.6 shows an increase in opportunistic bacterial and fungal pathogens: Coprobacillus, Streptococcus, Actinomycetes, C. albicans, C. auris, Enterococcus, and Aspergillus niger. The lack of microbiota and mycobiota, Intestinibacter, Eubacterium, Fusicatenibacter, Ruminococcus, Clostridium ramosum (Firmicutes phylum), Basidiomycota, Ascomycota, and Penicillium citrinu, can lead to the severity of the infection (Yamamoto et al. 2021).
Modifying gut virome, mycobiome, and the bacterial microbiome from healthy to COVID-19-infected patients (Zuo et al. 2021)
An increase in the number of OPs revealed the levels of C-reactive protein, TNF-alpha, and IL-18, which are proinflammatory cytokines produced by the intestinal microbiota in the sera of COVID-19 patients compared with those in influenza patients and healthy individuals.
6 The Intestinal Dysbiosis Associated with COVID-19 Severity
The concept of dysbiosis has been broadly defined as the change in the composition of the resident commensal microbiota compared to those found in healthy individuals. This alteration causes the disruption of symbiosis between the host and microbes, with adverse consequences. The composition of a healthy human gut microbiome is still a question with a complicated answer that has yet to be addressed. Recent advancements by a working group of the International Life Sciences Institute North America show some of the challenges, like the high degree of intra- and interindividual variation in the human microbiome, the lack of potential biomarkers to define and measure microbiome-host interactions, and the extreme consequences of dysbiosis on human physiology and disease.
Dysbiosis is described as an increase in potential pathogenic species and a decrease in beneficial organisms (e.g., Bifidobacterium and Faecalibacterium species) or a change in alpha diversity due to the production of harmful microbially derived compounds (e.g., hydrogen sulfide produced by sulfate-reducing bacteria). Higher alpha diversity is a marker of health in the GI tract but, conversely, a marker of dysbiosis in the vaginal microbiome.
Reductions were observed in the endurance of bacteria and archaea such as Lactobacillus, Bifidobacterium, Clostridium butyricum, Lachnospiraceae, Prevotella, Roseburia, Ruminococcus bromii, Faecalibacterium, and Bacteroides. This shows a reduction in probiotic bacteria, neutrophil concentration, and IL-6 concentration compared to healthy individuals (Yamamoto et al. 2021).
7 Dysbiosis in the Fecal Microbiome of COVID-19 Patients
The microbiome and mycobiome dysbiosis in the gut of COVID-19 patients was observed to have an increase in the number of opportunistic pathogens (OPs) like Actinomyces, Rothia, Enterococcus, Enterobacter, Streptococcus, and Klebsiella species that pose a threat of a reduction in host immunity (Chakravorty et al. 2007). The intestinal instability, prolonged dysbiosis, and high viral transcription and replication began during hospitalization and lasted even after 2 weeks of recovery. Fecal samples with less SARS-CoV-2 infectivity show improved levels of bacteria like Parabacteroides, Lachnospiraceae, and Bifidobacterium, which produce SCFAs, GOS, and FOS vital for boosting host immunity.
Roseburia, Faecalibacterium, Coprococcus, and Parabacteroides showed lower abundance in COVID-19 patients than fungal pathogens like Candida and Aspergillus spp., which were found enriched in healthy individuals. Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria phyla were observed to have enhanced gut and oral microbiota in both COVID-19 patients and healthy individuals.
Bacterial microbiota identified in the gut samples of COVID-19 patients are Bacteroidales, Enterobacterales, Clostridiales, Lactobacillales, and Bifidobacteriales, whereas the oral samples showed Lactobacillales, Micrococcales, Enterobacterales, and Selenomonadales (Libertucci and Young 2018).
8 Therapeutic Modulation of the Microbiome and Its Effects
Prebiotics and probiotics are used as therapies to prevent the colonization of pathogens, whereas fecal microbiome transplantation is used to clear the pathogens. This helps modulate the microbiota to prevent colonization and promote the clearance of pathogens. Some other treatments include the mechanisms of action that are different and unclear, and patients have yet to be studied (Harper et al. 2021).
8.1 Fecal Microbiota Transplantation in COVID-19 Patients
The transfer of a fecal suspension from a healthy donor to a recipient (a patient with dysbiosis) modulates the microbiota, which would indirectly help recover the health of a diseased individual by improving the functioning of the microbiota and decreasing the infection by replacing or restoring the functions in the body. It has proven highly effective against the decolonization of drug-resistant organisms to reduce or clear up the disease.
Studies have shown that using filtrates from healthy donor stools (fecal filtrate transfer), which includes transferring bacterial components and metabolites from donor to recipient, is more beneficial for treating infections. Evidence provided from studies has shown that outcomes of FMT were associated with alterations to the enteric virome and bacterial microbiota showing greater treatment success. When a diet high in resistant starch was eaten, Ruminococcus bromii, Oscillibacter, Firmicutes, Bacteroidetes, and Eubacterium rectale increased significantly.
Inulins found in various fruits, vegetables, and wheat have been shown to stimulate the growth of Bifidobacterium spp. and Faecalibacterium by increasing butyrate production. Fructooligosaccharide (FOS) and galactosaccharide (GOS) administration reduces the release of corticosterone, increases cecal acetate and propionate concentrations simultaneously, and reduces proinflammatory cytokines with an increase in interleukin (IL)-10, IL-8, and other anti-inflammatory cytokines (Cryan et al. 2019).
8.2 Prebiotics
Prebiotics stimulate the promotion of indigenous microbiota by participating in fermentation. They can inhibit or limit the development of the pathogen through the digestion of insoluble fiber sources to increase SCFA production, lactic acid, and peptidoglycan, which stimulate the innate immune system against pathogenic microorganisms. It has also been shown that pH alteration can affect the population of acid-sensitive species, regulate the virulence expression of pathogens, and inhibit the binding of pathogens to epithelial receptors.
The different types of prebiotics used for treating diseases or infections are oligosaccharide carbohydrates (OSCs), fructooligosaccharides, galactooligosaccharides (GOS), and resistant starch (RS). For example, fructooligosaccharide (FOS) increases the level of interleukin-4 (IL-4), a myeloid dendritic cell that improves the immune response in volunteers. Combining inulin and FOS can enhance antibody responses toward viral vaccines (diarrhea, measles).
The GOS improved the level of IL-8, IL-10, and C-reactive protein in the blood and the function of NK cells. SCFA increases the production of mucins and antimicrobial peptides, reduces pH, increases intestinal motility, and has anti-inflammatory properties significant for the health of the gut epithelium (Shin et al. 2022).
8.3 Probiotics
Probiotics are live microorganisms that help boost the health of the host when administered in an adequate amount. They are live microbial feed supplements that improve intestinal microbial balance by modulating the intestinal microbiota. The benefits of probiotic consumption include regulation of the intestinal microbiota, stimulation of the immune system, promoting the synthesis and bioavailability of nutrients, and reducing the symptoms of lactose intolerance and the risk of other diseases.
Probiotic products generally contain one or more microbial strains like Lactobacillus, Enterococcus, Bifidobacterium, Lactococcus, Streptococcus, Saccharomyces cerevisiae, and Bacillus. The FDA (Food and Drug Administration) must regulate the microorganisms used in these products for human consumption, and they must be GRAS (generally recognized as safe) organisms. The QPS (Qualified Presumption of Safety) criteria are used to maintain the safety assessment of bacterial supplements, safe usage, and the risk of acquired resistance to antibiotics (Shin et al. 2022). Probiotic microorganisms can produce enzymes like esterase, protease, and lipase and coenzymes A, Q, NAD, and NADP, which show the metabolism of antibiotics (bacitracin and lactacin), anticarcinogens, and immunomodulatory properties.
Most of these bacteria are probiotics, the good bacteria colonizing within your digestive tract that serve a beneficial purpose (produce vitamins, absorb nutrients from your food, and even help regulate your mood). Prebiotics are the natural dietary fiber (nonliving and nondigestible by humans) that nourishes our probiotic bacteria. The probiotics introduce good bacteria into the gut, and the prebiotics act as fertilizer for the good bacteria’s growth, improving the ratio of good to bad bacteria. This directly correlates to your health and overall well-being (gut-brain axis) (Fig. 8.7).
Common medications have both a beneficial and harmful effect on gut microbes, according to new research. Antibiotics and gastric acid suppressants disrupt beneficial bacterial populations in the gut, while statins and ACE inhibitors are linked to improved bacterial composition and function (Shin et al. 2022) (Fig. 8.8).
The combination of these medications may provide the viable bacterial strain with a greater potential to fill a niche and restore its community structure and function by restoring the metabolome (Cryan et al. 2019)
Certain bacterial strains present in probiotics can also have an effect on central neuronal processes like neural communication, neurogenesis, the expression of neuropeptides, neurological inflammation, and even behavior. It helped with a wide range of conditions, from autism to melancholy and anxiety. “Psychobiotics” and “chobiotics,” which aim to treat neurological and psychiatric diseases by altering the composition of the gut microbiota, emerged in response to the prevalence of these conditions. Also included are models based on flux balance analysis (FBA) used to foretell and comprehend how microorganisms will behave in a certain setting (Fig. 8.9).
8.4 The Gut-Associated Peptide Reg3g Connects the Microbiota of the Small Intestine to the Control of Energy Homeostasis, Blood Sugar Levels, and Gastrointestinal Activity
The composition of the gut microbiome and its adaptation to various environments in various metabolic diseases are key to understanding the necessary changes to maintain homeostasis. Fermentable fiber-rich inulin diets with gastric sleeve surgery (which Acyl homoserine lactones (Acyl-HSL) are acyl homoserine lactones that gram-negative bacteria produce, limiting amount of food you can eat) boost the production of the antibacterial peptide Reg3g in the intestines and the bloodstream. New approaches to therapy may take advantage of its role as a gut hormone that links the microbiota of the intestines to the remainder of the host’s physiology (Lazar et al. 2018).
8.5 Microbe-Microbe Interactions
Bacteria use signaling molecules to share information and adapt their gene expression to their surroundings, which is especially important in highly competitive ecosystems with many coexisting species. The term for this phenomenon is “quorum sensing” (QS). Quantum signaling (QS) depends on the density of molecular language that controls cell phenotypic expression and behavior in response to external cues. This intercellular communication is divided into two categories. Interspecific communication among bacterial and eukaryotic/host cells is facilitated by interspecific interaction that utilizes a universal chemical language, the first form of cell-to-cell communication (Falcao et al. 2004).
Autoinducers (AIs) are tiny organic compounds that act like hormones and are part of this system. Acyl homoserine lactones (Acyl-HSL) are acyl homoserine lactones that gram-negative bacteria produce. Peptide compounds (AIP) are not diffusible in gram-positive bacteria. These cell-to-cell communication networks in bacteria were first reported as a way for bacteria to control the release of virulence genes, which are found in pathogens and play a crucial role in infection through cell density (Villapol 2020). This method allows commensal bacteria to control how much of the host they colonize (Fig. 8.10).
Cathelicidins, defensins, and AIs are all antimicrobial peptides made by eosinophils. They all work as signaling molecules within and between species.
Quorum sensing processes include measuring some of these processes, including bioluminescence, pathogenicity factor expression, biofilm formation, and conjugation. For instance, enteric pathogens use quorum sensing to control the expression of genes that encode virulence traits like mobility and type 3 secretion.
Thus, the use of quorum sensing ensures effective host colonization by detecting the presence of normal gut flora (Villapol 2020). This phenomenon sheds light on mitigating microbial infection against virulence factors and biofilm formation controlled by quorum sensing (Seibert et al. 2022) (Fig. 8.11).
9 Conclusion
9.1 The Microbiome Is an Extraordinary Helper: We Must Nurture Our Bodies’ Microbes
-
The makeup of the gut microbiome has an impact on host metabolism and general health.
-
Take antibiotics exactly as prescribed. Antibiotic usage over an extended period is linked to a change in antibiotic-driven gut microbiome composition that upsets the body’s normal microbial equilibrium.
-
Therapeutic modulation of the microbiota is the future of diagnostics for infectious diseases.
-
Identification of the imbalance (if any) in the changing microbiome of humans (testing for the microbiome).
-
Developing noninvasive microbiome-based diagnostics (Peter et al. 2019).
-
Increase the fiber in your diet. All plant foods, such as vegetables, fruits, and whole grains, contain fiber (Sudhakar and Padi 2022). Avoid these foods to keep your gut microorganisms happy (Atiatorme et al. 2022). These are foods that are heavy in sugar, fat, or processing.
References
Atiatorme E, Bramhachari PV, Kariali E, Sudhakar P (2022) A paradigm shift in the role of the microbiomes in environmental health and agriculture sustainability. In: Veera Bramhachari P (ed) Understanding the microbiome interactions in agriculture and the environment. Springer, Singapore, pp 83–101
Baghbani T, Nikzad H, Azadbakht J et al (2020) Dual and mutual interaction between microbiota and viral infections: a possible treatment for COVID-19. Microb Cell Factories 19:217
Chakravorty S, Helb D, Burday M, Connell N, Alland D (2007) A detailed analysis of 16S ribosomal RNA gene segments for the diagnosis of pathogenic bacteria. J Microbiol Methods 69(2):330–339
Cryan JF, Cowan M, Sandhu CS, Bastiaanssen KVS et al (2019) The microbiota-gut-brain axis. Physiol Rev 99(4):1877–2013
de Castilhos J, Zamir E, Hippchen T, Rohrbach R, Schmidt S, Hengler S et al (2022) Severe dysbiosis and specific Haemophilus and Neisseria signatures as hallmarks of the oropharyngeal microbiome in critically ill coronavirus disease 2019 (COVID-19) patients. Clin Infect Dis 75(1):e1063–e1071
Falcao JP, Sharp F, Sperandio V (2004) Cell-to-cell signaling in intestinal pathogens. Curr Issues Intest Microbiol 5:9–17
Gaibani P, Viciani E, Bartoletti M et al (2021) The lower respiratory tract microbiome of critically ill patients with COVID-19. Sci Rep 11:10103
Hanada S, Pirzadeh M, Carver KY, Deng JC (2018) Respiratory viral infection-induced microbiome alterations and secondary bacterial pneumonia. Front Immunol 9:2640
Harper A, Vijayakumar V, Ouwehand AC, ter Haar J, Obis D, Espadaler J, Binda S, Desiraju S, Day R (2021) Viral infections, the microbiome, and probiotics. Front Cell Infect Microbiol 10:596166. https://doi.org/10.3389/fcimb.2020.596166
Hoque MN, Sarkar MMH, Rahman MS, Akter S, Banu TA, Khan MS et al (2021) SARS-CoV-2 infection reduces the human nasopharyngeal commensal microbiome with the inclusion of pathobionts. Sci Rep 11(1):1–17
Jackson CB, Farzan M, Chen B et al (2022) Mechanisms of SARS-CoV-2 entry into cells. Nat Rev Mol Cell Biol 23:3–20
Shin J, Kramer NB, Shao Y, Abbott S et al (2022) The gut peptide Reg3g links the small intestine microbiome to the regulation of energy balance, glucose levels, and gut function. Cell Metab 34(11):1765–1778.e6
Konatala A, Parackel F, Sudhakar P (2021) Recent advancements in microbiome–immune homeostasis and their involvement in cancer immunotherapy. In: Microbiome in human health and disease. Springer, Singapore, pp 239–258
Lazar V, Ditu L-M, Pircalabioru GG, Gheorghe I, Curutiu C, Holban AM, Picu A, Petcu L, Chifiriuc MC (2018) Aspects of gut microbiota and immune system interactions in infectious diseases, immunopathology, and cancer. Front Immunol 9:1830
Libertucci J, Young VB (2018) The role of the microbiota in infectious diseases. Nat Microbiol 4(1):35–45
Liu TFD, Philippou E, Kolokotroni O, Siakallis G, Rahima K, Constantinou C (2022) Gut and airway microbiota and their role in COVID-19 infection and pathogenesis: a scoping review. Infection 50(4):815–847
Mohan SM, Sudhakar P (2022) Metagenomic approaches for studying plant–microbe interactions. In: Understanding the microbiome interactions in agriculture and the environment. Springer, Singapore, pp 243–254
Peter AE, Sudhakar P, Sandeep BV, Rao BG (2019) Antimicrobial and anti-quorum sensing activities of medicinal plants. In: Bramhachari P (ed) Implication of quorum sensing and biofilm formation in medicine, agriculture and food industry. Springer, Singapore, pp 189–213
Pola S, Padi DL (2021) Overview of human gut microbiome and its role in immunomodulation. In: Bramhachari PV (ed) Microbiome in human health and disease. Springer, Singapore, pp 69–82
Ronda C, Chen SP, Cabral V et al (2019) Metagenomic engineering of the mammalian gut microbiome in situ. Nat Methods 16:167–170
Sanapala P, Pola S (2021) Modulation of systemic immune responses through genital, skin, and oral microbiota: unveiling the fundamentals of human microbiomes. In: Bramhachari PV (ed) Microbiome in human health and disease. Springer, Singapore, pp 13–34
Seibert B, Cáceres CJ, Carnaccini S, Cardenas-Garcia S, Gay LC, Ortiz L et al (2022) Pathobiology and dysbiosis of the respiratory and intestinal microbiota in 14 months old golden syrian hamsters infected with SARS-CoV-2. PLoS Pathog 18(10):e1010734
Sudhakar P, Padi D (2022) Modifications in environmental microbiome and the evolution of viruses through genetic diversity. In: Veera Bramhachari P (ed) Understanding the microbiome interactions in agriculture and the environment. Springer, Singapore, pp 103–112
Villapol S (2020) Gastrointestinal symptoms associated with COVID-19: impact on the gut microbiome. Transl Res 226:57–69
Yamamoto S, Saito M, Tamura A, Prawisuda D, Mizutani T, Yotsuyanagi H (2021) The human microbiome and COVID-19: a systematic review. PLoS One 16(6):e0253293
Zuo T, Wu X, Wen W, Lan P (2021) Gut microbiome alterations in COVID-19: science direct. Genomics Proteomics Bioinformatics 19(5):679–688
Acknowledgments
The authors would like to express their gratitude to the authorities of Andhra University for providing the necessary infrastructure and facilities for conducting this research. They would also like to acknowledge the support of the principal of Andhra University College of Science and Technology, Andhra University, Visakhapatnam, and the head of the Department of Biotechnology, AUCST, Andhra University, Visakhapatnam.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
The authors declare that no financial or personal relationships with other people or organizations could inappropriately influence (bias) their work. Additionally, they have no competing interests related to this research.
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mukala, N., Pola, S., Konatala, A. (2023). Influence of Intestinal Microbiomes on COVID Progression and Its Effects by Immunotherapeutic Modulation. In: Veera Bramhachari, P. (eds) Human Microbiome in Health, Disease, and Therapy. Springer, Singapore. https://doi.org/10.1007/978-981-99-5114-7_8
Download citation
DOI: https://doi.org/10.1007/978-981-99-5114-7_8
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-5113-0
Online ISBN: 978-981-99-5114-7
eBook Packages: MedicineMedicine (R0)