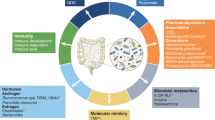
Abstract
Current scientific advances have considerably added to our understanding of the complex association between the microbiome and cancer. Host and microbiota have co-evolved into a “super-organism,” and several physiological processes and multifactorial disease conditions are influenced by host–microbiome interactions. In the past, microbial communities have been suggested to influence the development, progression, metastasis formation, and treatment response of multiple cancer types. However, a better molecular understanding of cancer-modulating interactions and influences on cancer treatment is considered of major scientific relevance and clinical importance. Here, we discuss the scientific evidence on the role of gut microbiota in cancer progression and its treatment and highlight the latest knowledge leveraged to target specific microbes contributing to tumorigenesis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Microbes represent wide ecological adaptations dominating all four spheres of the earth and in that way captivate microbiologists to explore their niches. A narrative of the terminology has been proposed explaining that microbial taxa linked to a host organism or a dominant environment refer to “microbiota,” whereas the “microbiome” is the exclusive assemblage of the microbes and their associated genes (Ursell et al. 2012). Next-generation sequencing (NGS) techniques have garnered ample attention for providing a comprehensive view of human microbiome. The human microbiome is a multi-kingdom repertoire of bacteria, archaea, fungi, protists, and viruses residing in and on the human body (Ursell et al. 2012). Microbiome performs vital functions such as regulating barrier, maintaining homeostasis, preventing pathogenic infection, and regulating metabolism and vitamin synthesis (Li et al. 2019). Oral, gut, and skin microbiomes comprise greatly enriched and diverse microbial consortia, whereas lungs, bladder, prostate, liver, pancreas, and vagina harbor low diverse microbial populations (Cho and Blaser 2012). The host and its microbiome exist in symbiosis as a super-organism by offering a nutrient-rich microenvironment (Schwabe and Jobin 2013). Despite the presence of trillions of beneficial microbes inhabiting human body, dysbiosis can still occur that might lead to the development of cancers and inflammatory diseases. Microbial diversity and abundance differ in different organs; thus, many diseases, including cancer, occur in specific locations within an organ. Chances of cancer are high in those locations where microbial densities are high (Human Microbiome Project Consortium 2012; Cullin et al. 2021).
Investigation of evidence of microbial influence on biology of cancer is in its infancy, and a better comprehensive view of cancer-modeling interfaces and its influence on cancer treatment are reflected as of great scientific relevance. Composition and functional repertoire of microbial communities can be characterized and used for defining pathology and physiology of human cancers (Cullin et al. 2021). Human bodies are continuously exposed to variety of microbes and their byproducts including some tumor-promoting metabolites such as high levels of polyamines, sulfides, and N-nitroso compounds (Louis et al. 2014). These metabolites while circulating in the body may lead to cancer progression at locations distinct from the specific microbial residence (Rajagopala et al. 2017). Microorganisms can also migrate to different locations and get associated with the development of tumors. However, the microbes impact the process of carcinogenesis by inflammation and immune system-independent mechanisms and the most decipherable link is via the immune system as the microbes themselves play a significant role in activating and regulation of host immunity (Rajagopala et al. 2017). Interaction of immune system and microorganisms can occur at (1) mucosal layers (via microbial metabolites) or (2) locally at lymphoid organs. Remote/local microbial signals may impact both innate and adaptive immune responses, leading to systemic immunity modulation and anti-tumor innate immune responses (Cullin et al. 2021). Microorganisms can modulate metabolism, inflammation, carcinogenesis, and genotoxicity through multiple mechanisms, and targeting these mechanisms could envision cancer prevention strategies. Genetic modification of microbiota producing/lacking particular enzymes could be utilized for expressing tumor-reducing phytochemicals or reducing tumor-promoting elements (Shwabe and Jobin 2013).
By means of proliferation, escaping cell growth suppression, activation of metastatic pathways, angiogenesis induction, and autophagy resistance cancer cells evade the immune system. All these processes have been explored in detail for decades, but role of microbiome in both cancer progression and treatment is still partly unknown. Microbiome data analysis may assist the advancement of novel cancer diagnostic strategies encompassing cancer detection (identification of microbial DNA/RNA in peripheral blood), surveying metastatic cancer progression, assessing prognosis, and applying artificial intelligence algorithms in foreseeing patient treatment responses. The understanding of the microbiome and cancer needs to be broadened with enhanced feasibility of cancer diagnosis based on microbial profile. Exploring various effects of the microbiome on carcinogenesis will provide new opportunities for diagnostic, preventive, and therapeutic strategies and would represent the next frontier of medical research.
2 Cancer Triggering Microbes and Their Cancer-Promoting Mechanisms
Hepatitis B (HBV), Epstein–Barr (EBV), hepatitis C (HCV), Kaposi sarcoma herpesvirus (KSV), human immunodeficiency virus-1 (HIV), human papillomaviruses (HPV), human T cell lymphotropic virus type 1 (HTLV), Opisthorchis viverrini, Clonorchis sinensis, Schistosoma haematobium, and Helicobacter pylori have been recognized as causes of distinct human cancer (IARC 2009). They participate in cancer progression through different mechanisms, such as B-cell differentiation, cell-cycle disruption, immune hyperactivation (HBV, EBV, HIV, and HCV), dysregulation of T cell (HTLV, EBV), and direct oncogenesis (HCV). Merkel cell polyomavirus (MCV) and Simian virus 40 (SV40) are involved in Merkel cell carcinoma (MCC) and mesothelioma, respectively (Pagano et al. 2004; Weitzman and Weitzman 2014).
In the case of H. pylori, CagA dissociates E-cadherin/b-catenin complex, leading to accumulation of b-catenin in both nucleoplasm and cytoplasm (Fig. 18.1). Further, this b-catenin forms complex with transcription factors (TCF/LEF) to activate target gene expression (Hamway et al. 2020). It binds to gastric epithelial cells by HopQ binding to CEACAM, whereby virulence factor CagA is directly injected into the epithelial cells via the T4SS. CagA activates Wnt/β-catenin pathways resulting in dysregulated cellular turnover and apoptosis (Cullin et al. 2021). On the other hand, F. nucleatum secretes Fap2 protein that interacts with TIGIT and hinders natural killer cell-facilitated immunosurveillance of cancer (Fig. 18.1). Another important adhesion, FadA, allows cellular internalization and induction of proinflammatory cascades mediated by NF-κB and IL-6. Fap2 interacts with d-galactose-β (1–3)-N-acetyl-D-galactosamine (Gal-GalNAc) carbohydrate moieties at the tumor surface to enhance cellular proliferation via Wnt/β-catenin pathway and increase proinflammatory cytokine production, leading to cancer cell invasion and therapy resistance (Zhang et al. 2020; Cullin et al. 2021).
The intestinal dysbiosis favors growth of adherent-invasive Escherichia coli (AIEC) and activation of pks island during inflammation, which increases IL-6-inducing CEACAM6 expression, hence increasing invasiveness of AIEC (Cullin et al. 2021). Once internalized, it secretes genotoxin colibactin, which induces inter-strand crosslink and double-stranded breaks with pro-tumoral cellular transformation (Fig. 18.1). Likewise, the enterotoxigenic Bacteroides fragilis (ETBF) produce BFT that can disrupt the intestinal environment by causing inflammation and increased permeability. BFT targets intestinal cell tight junction and causes cleavage of E-cadherin, which causes increased intestinal permeability and induces chronic intestinal inflammation via NF_KB signaling, leading to colorectal tumorigenesis (Fig. 18.1) (Pleguezuelos-Manzano et al. 2020).
3 Mechanisms of Microbial Carcinogenesis
Mechanism of microbial carcinogenesis involves (a) inflammation: Bacterial translocation may increase due to change in microbiome and host defense system, which leads to inflammation mediated by microorganism-associated molecular patterns (MAMPs) that activate Toll-like receptors (TLRs) in various cells like macrophage, myofibroblasts, and tumor cells; (b) genotoxin effect: Bacterial genotoxins like colibactin and cytolethal distending toxin (CTD), which when delivered to host cell nucleus cause DNA damage in various organs (Cullin et al. 2021). Reactive oxygen species (ROS) and reactive nitrogen species (RNS) released from inflammatory cells like macrophage and hydrogen sulfide (H2S) from bacterial microbiota may also be genotoxic, thus triggering carcinogenesis; (c) metabolic effect: Genotoxins like acetaldehyde, dietary nitrosamine in metabolism of bile acids, and hormones like estrogen and testosterone may activate due to metabolic actions of microbiome. The microbiota also mediates tumor suppressive effect by inactivation of carcinogenesis via generation of short-chain fatty acids and activation of cancer-preventing phytochemicals (Schwabe and Jobin 2013) (Fig. 18.2).
4 Risk Factors of Specific Cancers
Numerous propensities especially lifestyle and diet stand as the risk factors for the changes in the microbial diversity of the gut, thus affecting the microenvironment of host’s cells (Moskal et al. 2016). It is still not clear whether the shift in population pattern causes carcinogenesis or is an outcome of the emergence of tumors. Hence, this important variation will be the epicenter of research in cancer–microbiome area in the upcoming future. Different types of cancer and their association with microbiome are discussed below.
4.1 Oral and Gastric Cancer
Oral cavity is inhabited by a variety of bacterial species, which play a leading role in the development of oral diseases (Al-Hebshi et al. 2017). Dysbiosis in the oral microflora destabilizes the defense mechanisms of the host, resulting in chronic periodontal disease (Bullon et al. 2014; Johannsen et al. 2014), which is related to changes in the oral microflora that is caused by the outgrowth of certain pathogenic microbes. The two notable pathogenic members of the oral microbiome that are known to induce tumorogenesis in the oral cavity are Fusobacterium nucleatum and Prevotella gingivalis (Mager et al. 2005). Repeated periodontitis is considered to increase risk for the development of oral squamous cell carcinoma (OSCC) and Prevotella intermedia and Porphyromonas gingivalis are associated with the occurrence of periodontitis (Mysak et al. 2014; Zhang et al. 2017; Hsiao et al. 2018, Li et al. 2019). Human papillomavirus (HPV) type 16 has also been recognized as a causative agent for oropharyngeal cancer (Bray et al. 2018).
Infection caused by P. gingivalis has been related to increase in oral cancer, oro-digestive cancer, and propagation of oral cancer stem cells. P. gingivalis infection tempers with many anti-apoptotic pathways and also spreads inter-cellularly with the help of actin-based membrane protrusions and interferes with various cell signaling pathways (Mao et al. 2007; Yilmaz et al. 2004). Firstly, P. gingivalis activates Jak1/Akt/Stat3 signaling, which controls intrinsic mitochondrial apoptosis pathways (Mao et al. 2007; Yilmaz et al. 2004). It inhibits gingival epithelial cell apoptosis induced by ATP ligation of P2X7 receptors (Yilmaz et al. 2008). P. gingivalis also causes reduction in amount of p53 and accelerates the progression through the S-phase of the cell cycle (Kuboniwa et al. 2008). It promotes the expression of the B7-H1, which accelerates regulatory T-cell production and B7-DC receptors in primary gingival epithelial cells (Groeger et al. 2011). This may be the possible reason that B7-H1 expression contributes to immune evasion by oral cancers. AR2/NF-KB pathways are activated by P. gingivalis infection, which in turn induces expression of promatrix metalloproteinase (proMMP-9) (Inaba et al. 2014). P. gingivalis produces gingipains and cysteine proteinases, which play a key role in engaging the PAR2 receptor and also cleave the MMP-9 pro-enzyme into an active form. The active form of MMP-9 along with extracellular matrix degrades the basement membrane. This promotes carcinoma, cell migration, and invasion. The P. gingivalis might contribute to oral squamous cell carcinoma metastasis. The another pathogenic member Fusobacterium nucleatum works by activating inflammatory cytokines like TNF-α, IL-1β, and IL-6. Several antiapoptotic pathways are modulated by F. nucleatum, and it also activates p38, which results in the secretion of MMP-9 and MMP-13 (collagenase 3).
Role of human papilloma virus in carcinogenesis has been confirmed since the discovery of HPV16. HPV16 expresses E6 and E7 proteins, which further lead to the inactivation of tumor suppressor proteins p53 and Rb (Wiest et al. 2002). Inactivation of p53 and Rb causes dysregulation of host DNA synthesis, as a consequence cell cycle control is lost (D'Souza et al. 2007). This leads to destabilization of the genome, chromosomal aberrations, and abnormal centrosome numbers. HPV acts as an independent risk factor for the development of oral squamous cell carcinoma (Duensing et al. 2000).
4.1.1 GI Tract Cancer
Gastric cancer has been acknowledged as the model to study bacterial cancers. Helicobacter pylori in the gastrointestinal tract has been classified as a class I carcinogen by the International Agency for Research on Cancer (International Agency for Research on Cancer 1994). Strong host immune response is generated due to infection with H. pylori, which results in various gastric problems such as gastric inflammation, dysplasia, achlorhydria, and epithelial atrophy (Blaser and Atherton 2004). The key hallmarks of cancer driven by H. pylori include inflammation that promotes tumor, downregulation of antitumor immune destruction, and increase in proliferative signaling (Asano et al. 2015). H. pylori is able to adhere to gastric epithelial cells with the help of multiple adhesins, including SabA and BabA (Ilver et al. 1998). Once it firmly associates with the gastric epithelium, it deposits CagA and other virulence factors on human cells by utilizing its type IV secretion system. SHP2, a host oncoprotein, gets dysregulated by CagA after entering the cell leading to uncontrolled cell growth and motility (Murata-Kamiya et al. 2010). Other virulence factors such as VacA are responsible for making pores in host membranes, causing cell death and higher rates of cell turnover (Cover and Blanke 2005). Neutrophils and macrophages start producing reactive oxygen species (ROS) due to H. pylori infection (Tsugawa et al. 2012; Sung et al. 2022; Kuo et al. 2022). Inflammatory cytokines, including IL-1β, tumor necrosis factor-α, and IL-8, are also produced by H. pylori infection, ultimately leading to induction of Th1 immune behavior in the gut (Handa et al. 2011; Wei et al. 2010). NF-κB levels rise in patients with infection, which further drive inflammatory mechanisms of tumor initiation and succession. H. pylori is protected from ROS due to the presence of catalase and superoxide dismutase proteins but bystander effect damages host tissues (Chaturvedi et al. 2011). VacA, virulence factor of the pathogen, obstructs nuclear factor of activated T-cell (NFAT) activation in T cells. Blocking of NFAT leads to a deficiency of IL-2-driven T-cell proliferation, which would gradually result in expulsion of H. pylori (Jain et al. 2011). γ-glutamyltranspeptidase (GGT) produced by H. pylori also has the ability to block T-cell proliferation. Hence, a perfect storm is generated by the combination of cellular damage, innate procarcinogenic signals, and reduced immune surveillance in patients, which results in cancer.
4.2 Hepatic and Pancreatic Cancer
Hepatocellular carcinoma (HCC) is responsible for 80–90% of liver cancers and stands as the third leading cause of cancer-related deaths (Mima et al. 2017; Tong et al. 2011). Fox et al. (2010) observed the presence of Helicobacter spp. in gastric mucosa, leading to an increase in the chances of tumor progression in the hepatobiliary tract. H. pylori seizes in the host’s cells by attaching to gastric epithelial cells by HopQ protein, which binds to carcinoembryonic antigen-related cell adhesion molecules (CEACAM) and destructs the antitumor immune system, further inducing tumorigenic inflammation (Schwabe and Jobin 2013). Murphy et al. (2014) studied the association of fifteen H. pylori proteins with hepatobiliary carcinoma using a multiplex serology panel and found an increase in antibodies against these proteins, predicting an increase in HCC and biliary tract cancer. Helicobacter hepaticus also promotes HCC by producing toxins that promote anti-apoptotic factors and activate nuclear factor kappa B (NF- κB) and WNT signaling pathways, thus promoting tumor-inducing metabolites and suppressing antitumor immunity (Fox et al. 2010; Beyoğlu and Idle 2022). Other experimental studies have shown the enrichment of Methylophilaceae, Fusobacterium, Prevotella, Actinomyces, and Novosphingobium along with Helicobacter pylori in extrahepatic carcinoma tissue specimens of 100 patients (Avilés-Jiménez et al. 2016).
Similarly, highly lethal pancreatic cancer is also influenced by gut microbiota, which induces oncogenic metabolites for tumor development. Pancreatic cancer has been found to be associated with periodontal diseases and gum inflammation (Michaud et al. 2007; Stolzenberg-Solomon et al. 2003). In a comparative study of salivary microbiome of pancreatic cancer patients and healthy people, Neisseria elongata and Streptococcus mitis were found to be associated with cancer (Farrell et al. 2012; Herremans et al. 2022). Another oral bacteria, Fusobacterium spp., was found to be present in the tumor tissues of 283 pancreatic ductal adenocarcinoma patients (Mitsuhashi et al. 2015). Fusobacterium increases the inflammatory cytokines and reactive oxygen species (ROS), which leads to epigenetic alteration of mismatch proteins such as mutL homolog 1 and CpG island methylator phenotype (tumor suppressor gene), thus promoting carcinogenesis (Kostic et al. 2013; Schetter et al. 2010). Helicobacter spp. has also been found to be associated with pancreatic cancer (Nilsson et al. 2002; Trikudanathan et al. 2011). Evidence from different studies suggests that the accumulation of specific microbes can lead to the activation of carcinogenic factors producing hepatocellular and pancreatic carcinoma. Prevotella sp., Bacteroides, Ruminococcus, Faecalibacterium, and Ruminiclostridium are involved in progression of hepatic cancer (Hu et al. 2020; Yu et al. 2017). The gut microbiota may impact not only the formation of tumor but also the adequacy of chemotherapies and immunotherapies for HCC and pancreatic medical procedure, leading to low survival rates of cancer patients (Mima et al. 2017).
4.3 Colorectal Cancer
Colorectal carcinoma (CRC) being the third most malignant tumor with high occurrence in Western countries stands as the fourth most common cause for cancer-related deaths with mortality rate of 9.2% (Mármol et al. 2017; Zhou et al. 2020b, b). 70% of the human microbiome is present in the colon, which makes it the most vigorously colonized part of the gastrointestinal system (Saus et al. 2019). The modulation of colonic flora can be responsible for the dysplasia and can cause colonic inflammation and biosynthesis of carcinogenic molecules, leading to CRC development (Arthur et al. 2012; Rubinstein et al. 2013). Fusobacterium nucleatum, Peptostreptococcus stomatis, Peptostreptococcus anaerobius, and Bacteroides fragilis along with twenty other microbial gene markers were found to be associated with CRC (Yu et al. 2017; Zhou et al. 2020b, b). On the other hand, the commonly found Escherichia coli can have the pathogenicity island (pks), which can synthesize colibactin, a genotoxin causing oncogenic mutations and DNA damage (Pleguezuelos-Manzano et al. 2020) (Fig. 18.1c). A polyamine catabolic enzyme, spermine, is highly inducible by inflammatory stimuli of enterotoxigenic Bacteroides fragilis, which results in DNA damage and increase in ROS, leading to colorectal carcinoma (Goodwin et al. 2011) (Fig. 18.1d). It is also known to produce another inflammatory toxin that causes diarrhea and colonizes in host, which can further induce CRC (Wu et al. 2009). Enterococcus faecalis also have capability to produce enterotoxins and ROS, causing inflammation and epithelial damage (Saus et al. 2019). The microbes alter the signaling pathway by producing toxic proteins or biochemicals, which creates unfavorable microenvironment for cells and promotes carcinogenesis. For instance, FasA surface protein produced by Fusobacterium nucleatum adheres to the epithelial walls of colon and interacts with E-cadherin (Zhou et al. 2022). The complex then alters the β-catenin and Wnt signaling pathways. Increase in FadA protein leads to increase in the inflammatory and oncogenic genes (Kostic et al. 2013; Rubinstein et al. 2013). Other bacterial species such as Bacteroides, Parabacteroides, Lachnospiraceae bacterium, and Alistipes spp. has high prevalence in CRC patients and are found to play role in development of colon carcinoma (Feng et al. 2015).
5 Role of Microbiome in Treatment of Cancer and Future Applications
Since certain microbial signatures are known to promote the development of cancer and affect the safety, tolerability, and effectiveness of treatments, there is growing evidence that the gut microbiome is connected to cancer in a variety of ways. From over past few years, tremendous progress has been made in the field of cancer treatment, but somehow the tumor microenvironment has a significant impact on how well tumor immunotherapy works. Studies have demonstrated that a variety of tumor microenvironment cells, including T cells, fibroblasts, natural killer (NK) cells, dendritic cells (DCs), and others, play a key role in tumor immunotherapy (Ferlazzo et al. 2004). NK cells additionally induce cDC1 to enter the tumor microenvironment (TME) and assist tumor immune control (Poutahidis and Erdman 2016; Zhang et al. 2018; Böttcher et al. 2018; Zhou et al. 2020b, b). Chemotherapy or immune checkpoint inhibitor resistance is linked to altered gut flora immune checkpoint inhibitors (ICIs). The antitumor effects of chemotherapy medicines or ICIs may be enhanced by altering the microbiota through the use of antibiotics, probiotics, fecal microbiota transplants, or nanotechnologies (Cheng et al. 2020).
5.1 Immunotherapy
The term “immunotherapy or immuno-anticancer treatments” refers to a variety of therapeutic modalities intended to stimulate a patient’s immune system or recruit external immunological cells to combat cancer. Immunotherapy achieves the goal of eradicating cancers by inhibiting negative immune regulatory factors, stimulating the immune system, and improving immune cell identification, which results in the death of immune cells to tumors (Beatty and Gladney 2015).
In comparison with conventional cancer treatments, the gut microbiota has a stronger anti-cancer effect and further activates the host immune system. Additionally, there is growing acknowledgment for the role that the interaction between the gut microbiota and cancer ICIs plays in antitumor immune therapy, which is a form of targeted therapy for cancer immunotherapy (Qiu et al. 2021). Numerous species, including Bifidobacterium breve and Bifidobacterium longum, have been shown to improve dendritic cell activity and therefore trigger CD8+ T-cell priming and accumulation in tumor microenvironments (Tanoue et al. 2019; Sivan et al. 2015). In the tumor microenvironment, molecular cell refinement and immunological control of therapeutic targets are growing, and clinical application is expanding as well, such would be the resistance to the programmed cell death ligand 1, which plays a very crucial role in anti-tumor immunotherapy. The reduced programmed cell death ligand 1 activity that is resistant to the tumor can have favorable therapeutic effects and may be used for modifying the inert tumor microenvironment in future too (Qiu et al. 2021). Resistance to PD-1/PD-L1 plays many roles in tumor immunotherapy. Using an acceptable and selective combination of immunotherapy in a constrained tumor microenvironment reactivates the anti-tumor immune response in the host. This refers to the possibility of immune toxicity and immunotherapy to increase antitumor immunity (Ngiow and Young 2020). Antibodies that are able to act against the ligands of the PD-1 and CTLA-4 may inhibit affinity of T lymphocytes with their suppressive two or more ligands, which will act on the tumor cells, which in turn will activate the anti-tumor response in the immune system against the tumor cells (Cullin et al. 2021) (Fig. 18.3).
5.2 Chemotherapy
Chemotherapy refers to the treatment that requires anti-cancer drugs with high and powerful chemical content to treat any form of cancer by targeting the fast multiplying or growing cancer cells in the body. Chemotherapy causes DNA and non-DNA damage that is ROS-mediated, which allows bacteria to pass the intestinal epithelium. In turn, this causes an inflammatory reaction and may result in systemic infections (Kalasabail et al. 2021). The gut microbiota can influence how the body reacts to chemotherapy by enhancing medication efficacy, encouraging chemoresistance, and/or mediating adverse effects and toxicity.
Gut microbiota uses various mechanisms to regulate or modify the potency of the anti-cancer drugs. Many species of bacteria that are present in the gut impact the competency of the drugs that are used in chemotherapy and immunotherapy, which also includes immune checkpoint inhibitors by various mechanisms (Cullin et al. 2021).
Fusobacterium nucleatum is one such bacterium found in the gut microbiota and with the use of antibiotics/drugs can also reduce the intensity of cancer. F. nucleatum functions via myeloid differentiation primary response 88(MYD88) and Toll-like receptors (TLR4), which further results in the selective deprivation of miR-18a and miR-4802, and this in turn initiates autophagy. This process can further assist in chemoresistance in cancer patients (Fig. 18.3).
An earlier research demonstrated that cyclophosphamide can alter the composition of the gut microbiota by causing some gram-positive bacteria to translocate into the secondary lymphoid organs, which in turn triggers the production of “pathogenic” T helper 17 (pTh17) cells and improves the host immune response brought on by memory T helper 1 (Th1) cells. Researchers using Caenorhabditis elegans models have also identified bacteria that play role in chemotherapeutic effectiveness, particularly in the metabolism of ribonucleotides and vitamins B6 and B9 (Scott et al. 2017; García-González et al. 2017), which further increase the efficacy of fluoropyrimidine, an antimetabolite by inhibiting bacterial deoxynucleotide metabolism (Cheng et al. 2020). Efficacy of chemotherapy drugs can be modulated by bacteria via different mechanisms. The efficiency of 5-FU can be modulated via B6, B9, and ribonucleotide metabolism. The efficiency of 5-FU was promoted by inhibiting bacterial deoxynucleotide metabolism (Fig. 18.3).
Cyclophosphamide (CTX), one of the most often prescribed chemotherapy medications for the treatment of solid tumors and lymphomas, stimulates immunogenic cancer cell apoptosis and immunomodulatory effects (Qiu et al. 2021). Daillère et al. (2016) showed the incorporation of E. hirae in mice treated with antibiotics and reverses cyclophosphamide chemoresistance, which promotes pTh-17 production and T-helper (Th)-1 cell differentiation by increasing CD8+ T cells and CD4+ T regulatory cell (Treg) in the intratumoral region (Fig. 18.3). As Barnesiella intestinihominis builds up in the colon, it activates Th1 and polyfunctional CD8+ cytotoxic T cells in the body, which encourages interferon (IFN)-producing γδT cells to invade tumors (Daillère et al. 2016; Cheng et al. 2020) on treatment with cyclophosphamide (Cheng et al. 2020) (Fig. 18.3). Non-enterotoxigenic B. fragilis and Erysipelotrichaceae are examples of immunogenic bacteria that promote migratory dendritic cells (DCs), which then promote follicular T helper (TFH) cells through interleukin (IL) 1 and IL-12. The increased IgG2b response from stimulated TFH cells then interacts with B cells to boost the antitumor effector or the memory CD8+ T-cell activity. The response is further enhanced in the presence of gut commensal and boosts antineoplastic drugs like oxaliplatin and cisplatin (Roberti et al. 2020).
5.3 Radiotherapy
Radiation therapy is a part of the treatment plans for more than 50% of patients with cancer, and it is thought to make up around 40% of therapeutic protocols with efficacy in more than 90% of cases particularly those in their first stage of diseases (Poonacha et al. 2022). The predominantly most significant impediment to malignant cancer cure in patients is radiation enteropathy and radiation toxicity (Hauer-Jensen et al. 2014). RT also affects the gut microbiota; however, only a few studies have attempted to examine the connection between the microbiota and radiotherapy response (Tonneau et al. 2021). In melanoma, lung, and cervical cancer models, an oral vancomycin-induced decrease in gram-positive gut commensals mediated by IFNg and CD8 T-cell-dependent pathways was linked to improved radiation efficiency (Uribe-Herranz et al. 2020). Cui et al. (2017) showed the benefits of fecal microbiota transplantation (FMT) against total body irradiation-induced acute radiation enteritis in the mouse model with an increase in microbiota diversity. Irradiated mice that received (FMT) survived longer and had enhanced digestive system performance and increased levels of peripheral white blood cells. An evaluation of FMT in the treatment of chronic radiation enteritis showed a radical shift in diversifying the microbiome composition and reducing gastrointestinal symptoms (Ding et al. 2020).
5.4 Targeting Microbiome for Therapeutic Modulation of Carcinogenesis
Gut microbiota influences the shape of the tumor microenvironment by acting on the host immune system. TLR4 signaling in tumor cells recruit neutrophils, and they release tumor necrosis factor (TNF), which causes metastasis. Gut microbiota reduces the number of neutrophils, and its metabolites inosine promotes differentiation of Th1 cells in the presence of exogenous interferon-γ. An additional therapeutic option might be the introduction of anaerobic bacteria since they colonize all tumor sites regardless of oxygenation level and can even eradicate hypoxic cancers (Riehl et al. 2019; Poonacha et al. 2022). Through anaerobic action, butyrate-producing bacteria can degrade polysaccharides to create short-chain fatty acids (SCFAs), which are crucial for reducing the proliferation of cancer cells (Wang et al. 2019; Wagner et al. 2018). SCFAs like butyrate along with metabolites of tryptophan lessen pro-inflammatory cytokines and encourage the release of anti-inflammatory cytokines and affect the class conversion of B-cells, activating dendritic cells and macrophages, which affect differentiation of memory T cells, which helps limit radiation-related toxicity (Fish et al. 2016; Badgeley et al. 2021; Cheng et al. 2020) (Fig. 18.3). An introduction of Bifidobacterium infantis, a commensal bacterium to mice, and B. infantis with monoclonal antibody along with RT shows improvement in tumor cell proliferation, decreases blood supply, and enhanced cell apoptosis of the tumor. Furthermore, mice that received the combination therapy showed a higher survival rate than mice that received either the RT or a bacterial antibody alone (Du et al. 2018). Similarly, the symbiotic introduction of Lactobacillus rhamnosus (ATCC 7469) exopolysaccharides (EPS) in rat models with a dose of ionizing radiation was tested to slow colorectal development, enhanced the modulation of signaling growth factors involved in inflammation, and also reduced the progression of colorectal carcinoma in mice when compared to untreated control or those given L. rhamnosus or irradiation alone (Ruiz-Ruiz et al. 2017). These findings regarding the significance of the unique characteristics of the patient’s pre-existing microbiota suggest the possibility of incorporating microbiota analysis into personalized treatment protocols to predict how therapy will affect the patient and how the gut microbiota can be used as a biodosimeter to check the biological response for treatment planning (González-Mercado et al. 2021; Shi et al. 2020; Ding et al. 2020).
References
Al-Hebshi NN, Nasher AT, Maryoud MY, Homeida HE, Chen T, Idris AM, Johnson NW (2017) Inflammatory bacteriome featuring fusobacterium nucleatum and Pseudomonas aeruginosa identified in association with oral squamous cell carcinoma. Sci Rep 7(1):1–10. https://doi.org/10.1038/s41598-017-02079-3
Arthur JC, Perez-Chanona E, Mühlbauer M, Tomkovich S, Uronis JM, Fan TJ et al (2012) Intestinal inflammation targets cancer-inducing activity of the microbiota. Science 338(6103):120–123
Asano N, Iijima K, Koike T, Imatani A, Shimosegawa T (2015) Helicobacter pylori-negative gastric mucosa-associated lymphoid tissue lymphomas: a review. World J Gastroenterol: WJG 21(26):8014. https://doi.org/10.3748/wjg.v21.i26.8014
Avilés-Jiménez F, Guitron A, Segura-Lopez F, Mendez-Tenorio A, Iwai S, Hernández-Guerrero A, Torres J (2016) Microbiota studies in the bile duct strongly suggest a role for helicobacter pylori in extrahepatic cholangiocarcinoma. Clin Microbiol Infect 22(2):178–e11. https://doi.org/10.1016/j.cmi.2015.10.008
Badgeley A, Anwar H, Modi K, Murphy P, Lakshmikuttyamma A (2021) Effect of probiotics and gut microbiota on anti-cancer drugs: mechanistic perspectives. Biochim Biophys Acta Rev Cancer 1875(1):188494. https://doi.org/10.1016/j.bbcan.2020.188494
Beatty GL, Gladney WL (2015) Immune escape mechanisms as a guide for cancer immunotherapy tailoring cancer immunotherapy. Clin Cancer Res 21(4):687–692. https://doi.org/10.1158/1078-0432.CCR-14-1860
Beyoğlu D, Idle JR (2022) The gut microbiota–a vehicle for the prevention and treatment of hepatocellular carcinoma. Biochem Pharmacol 115225:115225
Blaser MJ, Atherton JC (2004) Helicobacter pylori persistence: biology and disease. J Clin Invest 113(3):321–333. https://doi.org/10.1172/JCI20925
Böttcher JP, Bonavita E, Chakravarty P, Blees H, Cabeza-Cabrerizo M, Sammicheli S et al (2018) NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172(5):1022–1037. https://doi.org/10.1016/j.cell.2018.01.004
Braakhuis BJ, Snijders PJ, Keune WJH, Meijer CJ, Ruijter-Schippers HJ, Leemans CR, Brakenhoff RH (2004) Genetic patterns in head and neck cancers that contain or lack transcriptionally active human papillomavirus. J Natl Cancer Inst 96(13):998–1006. https://doi.org/10.1093/jnci/djh183
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424. https://doi.org/10.3322/caac.21492
Bullon P, Newman HN, Battino M (2014) Obesity, diabetes mellitus, atherosclerosis and chronic periodontitis: a shared pathology via oxidative stress and mitochondrial dysfunction? Periodontology 64(1):139–153. https://doi.org/10.1111/j.1600-0757.2012.00455.x
Chaturvedi R, Asim M, Romero Gallo J, Barry DP, Hoge Sablet TD, Delgado AG, Wroblewski LE, Piazuelo MB, Yan F, Israel DA, Casero RA, Correa P, Gobert AP, Polk DB, Peek RM, Wilson KT (2011) Spermine oxidase mediates the gastric cancer risk associated with helicobacter pylori CagA. Gastroenterology 141(5):1696–1708. https://doi.org/10.1053/j.gastro.2011.07.045
Cheng WY, Wu CY, Yu J (2020) The role of gut microbiota in cancer treatment: friend or foe? Gut 69(10):1867–1876. https://doi.org/10.1136/gutjnl-2020-321153
Cho I, Blaser MJ (2012) The human microbiome: at the interface of health and disease. Nat Rev Genet 13(4):260–270
Cover TL, Blanke SR (2005) Helicobacter pylori VacA, a paradigm for toxin multi functionality. Nat Rev Microbiol 3(4):320–332. https://doi.org/10.1038/nrmicro1095
Cui M, Xiao H, Li Y, Zhou L, Zhao S, Luo D, Fan S (2017) Faecal microbiota transplantation protects against radiation-induced toxicity. EMBO Mol Med 9(4):448–461. https://doi.org/10.15252/emmm.201606932
Cullin N, Antunes CA, Straussman R, Stein-Thoeringer CK, Elinav E (2021) Microbiome and cancer. Cancer Cell 39(10):1317–1341. https://doi.org/10.1016/j.ccell.2021.08.006
Daillère R, Vétizou M, Waldschmitt N, Yamazaki T, Isnard C, Poirier-Colame V et al (2016) Enterococcus hirae and Barnesiella intestinihominis facilitate cyclophosphamide-induced therapeutic immunomodulatory effects. Immunity 45(4):931–943. https://doi.org/10.1016/j.immuni.2016.09.009
Ding X, Li Q, Li P, Chen X, Xiang L, Bi L et al (2020) Fecal microbiota transplantation: a promising treatment for radiation enteritis? Radiother Oncol 143:12–18. https://doi.org/10.1016/j.radonc.2020.01.011
D'Souza G, Kreimer AR, Viscidi R, Pawlita M, Fakhry C, Koch WM, Gillison ML (2007) Case–control study of human papillomavirus and oropharyngeal cancer. N Engl J Med 356(19):1944–1956. https://doi.org/10.1056/NEJMoa065497
Du SX, Jia YR, Ren SQ, Gong XJ, Tang H, Wan-Shui W, Li-Ming S (2018) The protective effects of bacillus licheniformis preparation on gastrointestinal disorders and inflammation induced by radiotherapy in pediatric patients with central nervous system tumor. Adv Med Sci 63(1):134–139. https://doi.org/10.1016/j.advms.2017.09.005
Duensing S, Lee LY, Duensing A, Basile J, Piboonniyom SO, Gonzalez S et al (2000) The human papillomavirus type 16 E6 and E7 oncoproteins cooperate to induce mitotic defects and genomic instability by uncoupling centrosome duplication from the cell division cycle. Proc Natl Acad Sci 97(18):10002–10007. https://doi.org/10.1073/pnas.170093297
Farrell JJ, Zhang L, Zhou H, Chia D, Elashoff D, Akin D et al (2012) Variations of oral microbiota are associated with pancreatic diseases including pancreatic cancer. Gut 61(4):582–588. https://doi.org/10.1136/gutjnl-2012-302312
Feng Q, Liang S, Jia H, Stadlmayr A, Tang L, Lan Z et al (2015) Gut microbiome development along the colorectal adenoma–carcinoma sequence. Nat Commun 6(1):1–13. https://doi.org/10.1038/ncomms7528
Ferlazzo G, Pack M, Thomas D, Paludan C, Schmid D, Strowig T, Bougras G, Muller W, Moretta L, Münz C (2004) Distinct roles of IL-12 and IL-15 in human natural killer cell activation by dendritic cells from secondary lymphoid organs. Proc Natl Acad Sci 101(47):16606–16611. https://doi.org/10.1073/pnas.0407522101
Fish BL, Gao F, Narayanan J, Bergom C, Jacobs ER, Cohen EP et al (2016) Combined hydration and antibiotics with lisinopril to mitigate acute and delayed high-dose radiation injuries to multiple organs. Health Phys 111(5):410. https://doi.org/10.1097/HP.0000000000000554
Fox JG, Feng Y, Theve EJ, Raczynski AR, Fiala JL, Doernte AL et al (2010) Gut microbes define liver cancer risk in mice exposed to chemical and viral transgenic hepatocarcinogens. Gut 59(01):88–97. https://doi.org/10.1136/gut.2009.204115
García-González AP, Ritter AD, Shrestha S, Andersen EC, Yilmaz LS, Walhout AJ (2017) Bacterial metabolism affects the C. elegans response to cancer chemotherapeutics. Cell 169(3):431–441. https://doi.org/10.1016/j.cell.2017.03.046
González-Mercado VJ, Henderson WA, Sarkar A, Lim J, Saligan LN, Berk L et al (2021) Changes in gut microbiome associated with co-occurring symptoms development during chemo-radiation for rectal cancer: a proof of concept study. Biol Res Nurs 23(1):31–41. https://doi.org/10.1177/1099800420942830
Goodwin AC, Shields CED, Wu S, Huso DL, Wu X, Murray-Stewart TR et al (2011) Polyamine catabolism contributes to enterotoxigenic Bacteroides fragilis-induced colon tumorigenesis. Proc Natl Acad Sci 108(37):15354–15359. https://doi.org/10.1073/pnas.1010203108
Groeger S, Domann E, Gonzales JR, Chakraborty T, Meyle J (2011) B7-H1 and B7-DC receptors of oral squamous carcinoma cells are upregulated by Porphyromonas gingivalis. Immunobiology 216(12):1302–1310. https://doi.org/10.1016/j.imbio.2011.05.005
Hamway Y, Taxauer K, Moonens K, Neumeyer V, Fischer W, Schmitt V et al (2020) Cysteine residues in helicobacter pylori Adhesin HopQ are required for CEACAM–HopQ interaction and subsequent CagA translocation. Microorganisms 8(4):465
Handa O, Naito Y, Yoshikawa T (2011) Redox biology and gastric carcinogenesis: the role of helicobacter pylori. Redox Rep 16(1):1–7. https://doi.org/10.1179/174329211X12968219310756
Hauer-Jensen M, Denham JW, Andreyev H (2014) Radiation enteropathy—pathogenesis, treatment and prevention. Nat Rev Gastroenterol Hepatol 11(8):470–479. https://doi.org/10.1038/nrgastro.2014.46
Herremans KM, Riner AN, Cameron ME, McKinley KL, Triplett EW, Hughes SJ, Trevino JG (2022) The oral microbiome, pancreatic cancer and human diversity in the age of precision medicine. Microbiome 10(1):1–14
Hsiao JR, Chang CC, Lee WT, Huang CC, Ou CY, Tsai ST et al (2018) The interplay between oral microbiome, lifestyle factors and genetic polymorphisms in the risk of oral squamous cell carcinoma. Carcinogenesis 39(6):778–787. https://doi.org/10.1093/carcin/bgy053
Hu H, Lin A, Kong M, Yao X, Yin M, Xia H et al (2020) Intestinal microbiome and NAFLD: molecular insights and therapeutic perspectives. J Gastroenterol 55(2):142–158
Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214
IARC (2009) IARC monographs on the evaluation of carcinogenic risks to humans 100, B. https://publications.iarc.fr/Book-And-Report-Series/IarcMonographs-On-The-Identification-Of-Carcinogenic-Hazards-To-Humans/biological-Agents-2012
IARC, Working group on the evaluation of carcinogenic risks to humans, International Agency for Research on Cancer, World Health Organization (1994) Schistosomes, liver flukes and helicobacter pylori, vol 61. International Agency for Research on Cancer, Lyon
Ilver D, Arnqvist A, Ogren J, Frick IM, Kersulyte D, Incecik ET et al (1998) Helicobacter pylori adhesin binding fucosylated histo-blood group antigens revealed by retagging. Science 279(5349):373–377. https://doi.org/10.1126/science.279.5349.373
Inaba, H., Sugita, H., Kuboniwa, M., Iwai, S., Hamada, M., Noda, T. et al. (2014). Porphyromonas gingivalis promotes invasion of oral squamous cell carcinoma through induction of pro MMP 9 and its activation. Cell Microbiol, 16(1), 131–145. ://doi.org/https://doi.org/10.1111/cmi.12211
Jain P, Luo ZQ, Blanke SR (2011) Helicobacter pylori vacuolating cytotoxin A (VacA) engages the mitochondrial fission machinery to induce host cell death. Proc Natl Acad Sci 108(38):16032–16037. https://doi.org/10.1073/pnas.1105175108
Johannsen A, Susin C, Gustafsson A (2014) Smoking and inflammation: evidence for a synergistic role in chronic disease. Periodontol 64(1):111–126. https://doi.org/10.1111/j.1600-0757.2012.00456.x
Kalasabail S, Engelman J, Zhang LY, El-Omar E, Yim HCH (2021) A perspective on the role of microbiome for colorectal cancer treatment. Cancers 13(18):4623. https://doi.org/10.3390/cancers13184623
Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M et al (2013) Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe 14(2):207–215. https://doi.org/10.1016/j.chom.2013.07.007
Kuboniwa M, Hasegawa Y, Mao S, Shizukuishi S, Amano A, Lamont RJ, Yilmaz Ö (2008) P. gingivalis accelerates gingival epithelial cell progression through the cell cycle. Microbes Infect 10(2):122–128. https://doi.org/10.1016/j.micinf.2007.10.011
Kuo YC, Yu LY, Wang HY, Chen MJ, Wu MS, Liu CJ, Lin YC, Shih SC, Hu KC (2022) Effects of helicobacter pylori infection in gastrointestinal tract malignant diseases: from the oral cavity to rectum. World J Gastrointest Oncol 14(1):55–74. https://doi.org/10.4251/wjgo.v14.i1.55
Li W, Deng Y, Chu Q, Zhang P (2019) Gut microbiome and cancer immunotherapy. Cancer Lett 447:41–47
Louis P, Hold GL, Flint HJ (2014) The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol 12(10):661–672
Mager DL, Haffajee AD, Devlin PM, Norris CM, Posner MR, Goodson JM (2005) The salivary microbiota as a diagnostic indicator of oral cancer: a descriptive, non-randomized study of cancer-free and oral squamous cell carcinoma subjects. J Transl Med 3(1):1–8. https://doi.org/10.1186/1479-5876-3-27
Mao S, Park Y, Hasegawa Y, Tribble GD, James CE, Handfield M, Stavropoulos MF, Yilmaz Ö, Lamont RJ (2007) Intrinsic apoptotic pathways of gingival epithelial cells modulated by Porphyromonas gingivalis. Cell Microbiol 9:1997–2007. https://doi.org/10.1111/j.1462-5822.2007.00931.x
Mármol I, Sánchez-de-Diego C, Pradilla Dieste A, Cerrada E, Rodriguez Yoldi MJ (2017) Colorectal carcinoma: a general overview and future perspectives in colorectal cancer. Int J Mol Sci 18(1):197. https://doi.org/10.3390/ijms18010197
Michaud DS, Joshipura K, Giovannucci E, Fuchs CS (2007) A prospective study of periodontal disease and pancreatic cancer in US male health professionals. J Natl Cancer Inst 99(2):171–175. https://doi.org/10.1093/jnci/djk021
Mima K, Nakagawa S, Sawayama H, Ishimoto T, Imai K, Iwatsuki M et al (2017) The microbiome and hepatobiliary-pancreatic cancers. Cancer Lett 402:9–15. https://doi.org/10.1016/j.canlet.2017.05.001
Mitsuhashi K, Nosho K, Sukawa Y, Matsunaga Y, Ito M, Kurihara H et al (2015) Association of Fusobacterium species in pancreatic cancer tissues with molecular features and prognosis. Oncotarget 6(9):7209. https://doi.org/10.18632/oncotarget.3109
Moskal A, Freisling H, Byrnes G, Assi N, Fahey MT, Jenab M et al (2016) Main nutrient patterns and colorectal cancer risk in the European prospective investigation into cancer and nutrition study. Br J Cancer 115(11):1430–1440. https://doi.org/10.1038/bjc.2016.334
Murata-Kamiya N, Kikuchi K, Hayashi T, Higashi H, Hatakeyama M (2010) Helicobacter pylori exploits host membrane phosphatidylserine for delivery, localization, and pathophysiological action of the CagA oncoprotein. Cell Host Microbe 7(5):399–411. https://doi.org/10.1016/j.chom.2010.04.005
Murphy G, Michel A, Taylor PR, Albanes D, Weinstein SJ, Virtamo J et al (2014) Association of seropositivity to helicobacter species and biliary tract cancer in the ATBC study. Hepatology 60(6):1963–1971. https://doi.org/10.1002/hep.27193
Mysak J, Podzimek S, Sommerova P, Lyuya-Mi Y, Bartova J, Janatova T et al (2014) Porphyromonas gingivalis: major periodontopathic pathogen overview. J Immunol Res 2014:1. https://doi.org/10.1155/2014/476068
Ngiow SF, Young A (2020) Re-education of the tumor microenvironment with targeted therapies and immunotherapies. Front Immunol 11:1633. https://doi.org/10.3389/fimmu.2020.01633
Nilsson HO, Stenram U, Ihse I, Wadström T (2002) Re: helicobacter pylori seropositivity as a risk factor for pancreatic cancer. J Natl Cancer Inst 94(8):632–633. https://doi.org/10.1016/S1473-3099(08)70066-5
Pagano JS, Blaser M, Buendia MA, Damania B, Khalili K, Raab-Traub N, Roizman B (2004) Infectious agents and cancer: criteria for a causal relation. In: Seminars in cancer biology, vol 14. Academic, Cambridge, MA, pp 453–471
Pleguezuelos-Manzano C, Puschhof J, Rosendahl Huber A, van Hoeck A, Wood HM, Nomburg J et al (2020) Mutational signature in colorectal cancer caused by genotoxic pks+ E. coli. Nature 580(7802):269–273. https://doi.org/10.1038/s41586-020-2080-8
Poonacha KNT, Villa TG, Notario V (2022) The interplay among radiation therapy, antibiotics and the microbiota: impact on cancer treatment outcomes. Antibiotics 11(3):331. https://doi.org/10.3390/antibiotics11030331
Poutahidis T, Erdman SE (2016) Commensal bacteria modulate the tumor microenvironment. Cancer Lett 380(1):356–358. https://doi.org/10.1016/j.canlet.2015.12.028
Qiu Q, Lin Y, Ma Y, Li X, Liang J, Chen Z et al (2021) Exploring the emerging role of the gut microbiota and tumor microenvironment in cancer immunotherapy. Front Immunol 11:612202. https://doi.org/10.3389/fimmu.2020.612202
Rajagopala SV, Vashee S, Oldfield LM, Suzuki Y, Venter JC, Telenti A, Nelson KE (2017) The human microbiome and cancer. The human microbiome and cancer. Cancer Prev Res 10(4):226–234
Riehl TE, Alvarado D, Ee X, Zuckerman A, Foster L, Kapoor V et al (2019) Lactobacillus rhamnosus GG protects the intestinal epithelium from radiation injury through release of lipoteichoic acid, macrophage activation and the migration of mesenchymal stem cells. Gut 68(6):1003–1013. https://doi.org/10.1136/gutjnl-2018-316226
Roberti MP, Yonekura S, Duong CP, Picard M, Ferrere G, Tidjani Alou M et al (2020) Chemotherapy-induced ileal crypt apoptosis and the ileal microbiome shape immunosurveillance and prognosis of proximal colon cancer. Nat Med 26(6):919–931. https://doi.org/10.1038/s41591-020-0882-8
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe 14(2):195–206. https://doi.org/10.1016/j.chom.2013.07.012
Ruiz-Ruiz JC, Matus-Basto AJ, Acereto-Escoffié P, Segura-Campos MR (2017) Antioxidant and anti-inflammatory activities of phenolic compounds isolated from Melipona beecheii honey. Food Agric Immunol 28(6):1424–1437. https://doi.org/10.1080/09540105.2017.1347148
Saus E, Iraola-Guzmán S, Willis JR, Brunet-Vega A, Gabaldón T (2019) Microbiome and colorectal cancer: roles in carcinogenesis and clinical potential. Mol Asp Med 69:93–106. https://doi.org/10.1016/j.mam.2019.05.001
Schetter AJ, Heegaard NH, Harris CC (2010) Inflammation and cancer: interweaving microRNA, free radical, cytokine and p53 pathways. Carcinogenesis 31(1):37–49. https://doi.org/10.1093/carcin/bgp272
Schwabe RF, Jobin C (2013) The microbiome and cancer. Nat Rev Cancer 13(11):800–812. https://doi.org/10.1038/nrc3610
Scott TA, Quintaneiro LM, Norvaisas P, Lui PP, Wilson MP, Leung KY et al (2017) Host-microbe co-metabolism dictates cancer drug efficacy in C. elegans. Cell 169(3):442–456. https://doi.org/10.1016/j.cell.2017.03.040
Shi W, Shen L, Zou W, Wang J, Yang J, Wang Y et al (2020) The gut microbiome is associated with therapeutic responses and toxicities of neoadjuvant chemoradiotherapy in rectal cancer patients—a pilot study. Front Cell Infect Microbiol 10:562463. https://doi.org/10.3389/fcimb.2020.562463
Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM et al (2015) Commensal Bifidobacterium promotes antitumor immunity and facilitates anti–PD-L1 efficacy. Science 350(6264):1084–1089. https://doi.org/10.1126/science.aac4255
Stolzenberg-Solomon RZ, Dodd KW, Blaser MJ, Virtamo J, Taylor PR, Albanes D (2003) Tooth loss, pancreatic cancer, and helicobacter pylori. Am J Clin Nutr 78(1):176–181. https://doi.org/10.1093/ajcn/78.1.176
Sung C-E, Lin F-G, Huang R-Y, Fang W-H, Cheng W-C, Tsai Y-WC, Chen W-L (2022) Periodontitis, helicobacter pylori infection, and gastrointestinal tract cancer mortality. J Clin Periodontol 49(3):210–220. https://doi.org/10.1111/jcpe.13590
Tanoue T, Morita S, Plichta DR, Skelly AN, Suda W, Sugiura Y et al (2019) A defined commensal consortium elicits CD8 T cells and anti-cancer immunity. Nature 565(7741):600–605. https://doi.org/10.1038/s41586-019-0878-z
Tong CM, Ma S, Guan XY (2011) Biology of hepatic cancer stem cells. J Gastroenterol Hepatol 26(8):1229–1237. https://doi.org/10.1093/ajcn/78.1.176
Tonneau M, Elkrief A, Pasquier D, Del Socorro TP, Chamaillard M, Bahig H, Routy B (2021) The role of the gut microbiome on radiation therapy efficacy and gastrointestinal complications: a systematic review. Radiother Oncol 156:1–9
Trikudanathan G, Philip A, Dasanu CA, Baker WL (2011) Association between helicobacter pylori infection and pancreatic cancer. A cumulative meta-analysis. JOP 12(1):26–31. https://doi.org/10.6092/1590-8577/3379
Tsugawa H, Suzuki H, Saya H, Hatakeyama M, Hirayama T, Hirata K et al (2012) Reactive oxygen species-induced autophagic degradation of helicobacter pylori CagA is specifically suppressed in cancer stem-like cells. Cell Host Microbe 12(6):764–777. https://doi.org/10.1016/j.chom.2012.10.014
Uribe-Herranz M, Rafail S, Beghi S, Gil-de-Gómez L, Verginadis I, Bittinger K et al (2020) Gut microbiota modulate dendritic cell antigen presentation and radiotherapy-induced antitumor immune response. J Clin Invest 130(1):466–479. https://doi.org/10.1172/JCI124332
Ursell LK, Metcalf JL, Parfrey LW, Knight R (2012) Defining the human microbiome. Nutr Rev 70(suppl_1):S38–S44
Wagner BD, Grunwald GK, Zerbe GO, Mikulich-Gilbertson SK, Robertson CE, Zemanick ET, Harris JK (2018) On the use of diversity measures in longitudinal sequencing studies of microbial communities. Front Microbiol 9:1037. https://doi.org/10.3389/fmicb.2018.01037
Wang Z, Wang Q, Wang X, Zhu L, Chen J, Zhang B et al (2019) Gut microbial dysbiosis is associated with development and progression of radiation enteritis during pelvic radiotherapy. J Cell Mol Med 23(5):3747–3756. https://doi.org/10.1111/jcmm.14289
Wei J, Nagy TA, Vilgelm A, Zaika E, Ogden SR, Romero Gallo J et al (2010) Regulation of p53 tumor suppressor by helicobacter pylori in gastric epithelial cells. Gastroenterology 139(4):1333–1343. https://doi.org/10.1053/j.gastro.2010.06.018
Weitzman MD, Weitzman JB (2014) What’s the damage? The impact of pathogens on pathways that maintain host genome integrity. Cell Host Microbe 15(3):283–294
Wiest T, Schwarz E, Enders C, Flechtenmacher C, Bosch FX (2002) Involvement of intact HPV16 E6/E7 gene expression in head and neck cancers with unaltered p53 status and perturbed pRb cell cycle control. Oncogene 21(10):1510–1517. https://doi.org/10.1038/sj.onc.1205214
Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR et al (2009) A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med 15(9):1016–1022. https://doi.org/10.1038/nm.2015
Yilmaz O, Jungas T, Verbeke P, Ojcius DM (2004) Activation of the phosphatidylinositol 3-kinase/Akt pathway contributes to survival of primary epithelial cells infected with the periodontal pathogen Porphyromonas gingivalis. Infect Immun 72(7):3743–3751. https://doi.org/10.1128/IAI.72.7.3743-3751.2004
Yilmaz O, Yao L, Maeda K, Rose TM, Lewis EL, Duman M, Lamont RJ, Ojcius DM (2008) ATP scavenging by the intracellular pathogen Porphyromonas gingivalis inhibits P2X7-mediated host-cell apoptosis. Cell Microbiol 10(4):863–875. https://doi.org/10.1111/j.1462-5822.2007.01089.x
Yu J, Feng Q, Wong SH, Zhang D, Yi Liang Q, Qin Y et al (2017) Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66(1):70–78. https://doi.org/10.1136/gutjnl-2015-309800
Zhang S, Li C, Liu J, Geng F, Shi X, Li Q et al (2020) Fusobacterium nucleatum promotes epithelial-mesenchymal transition through regulation of the lncRNA MIR4435-2HG/miR-296-5p/Akt2/SNAI1 signaling pathway. FEBS J 287(18):4032–4047
Zhang Y, Yu G, Chu H, Wang X, Xiong L, Cai G et al (2018) Macrophage-associated PGK1 phosphorylation promotes aerobic glycolysis and tumorigenesis. Mol Cell 71(2):201–215. https://doi.org/10.1016/j.molcel.2018.06.023
Zhang Y, Zhen M, Zhan Y, Song Y, Zhang Q, Wang J (2017) Population-genomic insights into variation in Prevotella intermedia and Prevotella nigrescens isolates and its association with periodontal disease. Front Cell Infect Microbiol 7:409. https://doi.org/10.3389/fcimb.2017.00409
Zhou Y, Fei M, Zhang G, Liang WC, Lin W, Wu Y et al (2020a) Blockade of the phagocytic receptor MerTK on tumor-associated macrophages enhances P2X7R-dependent STING activation by tumor-derived cGAMP. Immunity 52(2):357–373
Zhou Z, Ge S, Li Y, Ma W, Liu Y, Hu S et al (2020b) Human gut microbiome-based knowledge base as a biomarker screening tool to improve the predicted probability for colorectal cancer. Front Microbiol 11:596027. https://doi.org/10.3389/fmicb.2020.596027
Zhou Z, Wang Y, Ji R, Zhang D, Ma C, Ma W et al (2022) Vanillin derivatives reverse fusobacterium nucleatum-induced proliferation and migration of colorectal cancer through E-cadherin/β-catenin pathway. Front Pharmacol 13:841918
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Bharti, M. et al. (2023). Emerging Role of Gut Microbiome in Cancer Immunotherapy. In: Sobti, R., Kuhad, R.C., Lal, R., Rishi, P. (eds) Role of Microbes in Sustainable Development. Springer, Singapore. https://doi.org/10.1007/978-981-99-3126-2_18
Download citation
DOI: https://doi.org/10.1007/978-981-99-3126-2_18
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-3125-5
Online ISBN: 978-981-99-3126-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)