Abstract
The process of finding a medication lead starts with identifying the target of a certain disease, followed by hit identification, and lead optimization. However, the traditional approaches to drug discovery required a lot of human labor, money, and time over an extended period of time. The advancement in the technology of AI and ML techniques becomes an important part in the initial phase to the market analysis reports and tremendously accelerates the research work in the drug discovery. Moreover, AI/ML minimizes the human error as compared to the conventional methods and reduces the risk and expenditure in clinical trials. By assessing clinically pertinent data that directs the discovery of new potential targets, applications of artificial intelligence (AI) in data and chemical synthesis process are directly involved in drug development optimization. In this chapter, we described the role of the AI/ML in the drug discovery process with the application and limitations and future approaches. We anticipate that as more scientists become aware of its potential, the usage of AI/ML in the drug discovery will continue to expand in the search of new drugs.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
9.1 Introduction
An average of $1.3 billion is spent on research and development for individual medicine (Kolluri et al. 2022). For non-oncology drugs, the median period from conception to approval spans from 5.9 to 7.2 years, whereas for oncology drugs, the median time is 13.1 with 13.8% overall probability of success for drug-development (DiMasi et al. 2016). Hence, lowering the success rate and overall costs resulting to lengthy timelines for the modern medication R & D process is a significant issue for both business and academics. Furthermore, the ongoing attrition of drug candidates is the cause of the modern pharmaceutical industry’s excessive expenditure. Recent data indicate that animal toxicity (11%), poor pharmacokinetics (39%), and ineffectiveness (30%) account for 80% of the causes of attrition of the drug development process. Unpredictably, the issues raised above are directly connected to the discovery of drugs prior to clinical trials, showing that there is space for improvement (Wong et al. 2019). Since it is practically impossible to synthesis and evaluates all the potential compounds through tests. However, the overall procedure is typically decided by knowledge-based judgments, which might be highly prejudiced.
In the past 10 years, machine learning (ML) and artificial intelligence (AI) techniques have been well-known, thanks to the significant developments in computer technology. Artificial intelligence has the tendency to gather and process massive amounts of data required for research purposes. This helps in finding broad patterns of illness targets using a data-driven method which is a difficult task to recognize due to the complexity of the disease mechanism. In this area, a number of innovative researches have demonstrated the potential use of AI and ML techniques in drug-target identification as well as their capacity to learn and uncover disease patterns with the corresponding targets without relying on biological proficiency.
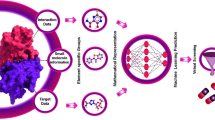
The process of finding a medication lead starts with identifying the target of a certain disease, followed by hit identification, and lead optimization (Fig. 9.1). However, the traditional approaches to drug discovery required a lot of human labor, money, and time over an extended period of time. Additionally, research cannot be done with absolute certainty regarding the possibility that a given drug candidate’s trial will be successful. The stages of drug discovery where AI is effectively cost-reducible start with the target identification and then identification of the lead or hit molecules through hit identification; furthermore, it helps in lead optimization and even aids in post-marketing surveillance reports.
AI/ML and deep learning systems have the potential to upsurge the probability of accomplishment ratio in drug development process. Moreover, these techniques provide significant progress in a number of R & D fields that includes novel target identification, deep learning and understanding of the target’s role in the disease, insights protein structures prediction, and the molecular compound design and optimization. AI further extends their support into the discovery of small molecule by involving in different field deals with new biology, better or distinctive chemistry, and in vivo and in vitro study with higher chances of success and less time making it less expensive as well. In this chapter, artificial intelligence, machine learning, and deep learning in the drug development process with its application will be discussed.
9.2 Artificial Intelligence
9.2.1 Concept of Modernization
The use of computers and computational techniques in research and engineering could be considerably improved with the development of modern artificial intelligence (AI). By assessing clinically pertinent data that directs the discovery of new potential targets, applications of artificial intelligence (AI) in data and chemical synthesis process are directly involved in drug development optimization. The creation and improvement of potential medications’ molecular structures can be done using an AI in drug design. Additionally, medication design methodologies comprehend how proteins’ specific forms impact their activities in health and malfunction in sickness.
AI is commonly combined with better patient monitoring process performed during clinical trials and medical devices that access specific patient data and advise medical decisions in the organization, optimization, and operation and acquire crucial patient’s data for clinical studies (Zhavoronkov et al. 2020). Additionally, it is increasingly feasible to utilize AI approaches to enhance healthcare research and services. However, one such application is risk-based guidance with deep-learning models used to anticipate preventable hospital readmissions (Farghali et al. 2021).
9.2.2 Models
9.2.2.1 AI-Guided Target Identification
A very popular and effective approach to finding new drugs is target-based drug discovery. For the treatment of any particular diseases, one should identify the target responsible for the agonist or antagonist actions. However, because of the choice of targets that are weakly related to the disease or have an unsupported theory, many therapeutic candidates in clinical trials have poor efficacy or elevated toxicity (Kim et al. 2020). Consequently, choosing appropriate targets requires a clearly distinct model for the relationship between the ailment and biological components. To interpret the connections, a variety of omics data types including genomics, proteomics, and metabolomics are required for better results.
The three kinds of conventional target identification techniques include machine learning, network-based models, and statistical analysis (Brown 2007). The most common and traditional methods for target identification have been statistical analyses of omics data for many years. These techniques were developed using the genome-wide study of associations (GWAS) and its emphases on finding genetic differences between samples from healthy and diseased people. By using association tests for the disease’s gene expression, such as the Chi-squared test, Fisher’s exact test, or t-test, it is possible to pinpoint potential target genes. Numerous study used different types of data such as for the tumor samples from the Gene Expression Omnibus (GEO) project, miRNA expression data from NCI-60 cancer cell lines, and TNBC and non-TNBC data from the Cancer Cell Line Encyclopedia (CCLE) project to identify three kinase (PKC, CDK6, and MET) targets for triple-negative breast cancer (TNBC) (Chen and Butte 2016). They performed a two-stage bioinformatics investigation that involved a patient-based Kaplan Meier survival test and cell-based gene expression analysis. The disease-related genetic variations can be found using GWAS (Zhu et al. 2016). In order to find the genes linked to a complex human feature, Zhu et al. introduced a technique called Summary data-based Mendelian Randomization (SMR).
9.2.2.2 Network-Based Approaches
Network-based approaches are frequently employed to depict the intricate relationships between the many biological components. Networks are made up of nodes, which stand in biological components, and edges, which show how the nodes interact. Furthermore, this method uses a heterogeneous network to manage the various omics data types. Consequently, a network-based method to target identification is used in numerous investigations.
This network identifies gene sets linked to disease pathways by capturing genes with identical biological process function. Network analysis was utilized by Petyuk et al. to pinpoint a late-onset Alzheimer’s target; to determine the gene-protein expression association contours, they built a co-expression network using peptide and transcript data (Petyuk et al. 2018; Mohamed et al. 2020). To add order or path towards network edges, they also created causal predictive networks.
Recently, target identification has also been accomplished using the knowledge graph. Entities, relations, and semantic data are represented in knowledge graphs as a machine-interpretable graph. Based on tensor factorization, a knowledge graph’s entity and relationship are encoded into three embedding vectors and efficient through learning by decreasing wrong facts and maximizing accurate facts.
9.2.2.3 Machine Learning-Based Approaches
Finding broad spectrum of illness targets using a data-driven method remains difficult due to the complexity of the disease mechanism. Using the classifiers, we can determine whether a gene is associated with the therapeutic target or not. Through gene-disease association data, Open Targets platform that can be classified into four types such as Random Forest (RF), Support vector machine (SVM), Neural Net, and Gradient Boosting Machine (GBM) (Ferrero et al. 2017). When the four classifiers are performed similarly, the results will be an AUC of 0.75 and an accuracy of about 70%. By combining these regression models with gene expression data from GEO and Array Express, Mamoshina and co-workers created an age prediction system. They used feature importance analysis to determine which genes were most closely connected with age prediction, and found that five well-known medication targets among the top 20 genes (Mamoshina et al. 2018).
9.2.2.4 AI-Guided Hit Identification
The important milestone in preclinical drug discovery is the identification of drug-target interactions. The molecular interaction among the drug and the chosen target determines the desired effects of the treatment, but unwanted interactions that were not specifically targeted during drug development might also result in side effects and the need to reposition the drug (Keiser et al. 2009). In order to maximize the effectiveness of the initial phases of drug development, numerous computational models are used to detect drug-target interaction and estimate binding affinities, which also has the benefit of delivering unique drug candidates (Ballester and Mitchell 2012; Stepniewska-Dziubinska et al. 2018). There are three basic types of hit identification computational approaches: the first focuses on the structure of the protein, the second on the structure of the ligand, and the third on the chemogenomic methods that describe similarity and feature-based methods (Fig. 9.2).
9.2.2.5 Structure-Based Approaches
The target protein’s 3D structures, which are produced by X-ray crystallography (XRC) and proton nuclear magnetic resonance spectroscopy (protein NMR), are utilized by structure-based approaches. A key strategy in structure-based techniques is a molecular docking simulation, which is carried out in two parts (Imrie et al. 2018). The first phase is the search for ligands in conformational space, which thoroughly simulates potential binding poses. Following a conformational search, a scoring function ranks potential ligand poses on the targeted protein structure and calculates binding affinity in the second phase. The evaluation of docking simulations is influenced by the scoring function’s quality. Traditionally, binding affinity posture is predicted using empirical or knowledge-based scoring systems.
Data-driven machine learning scoring functions (MLSF) are created by employing support vector machines (SVM) and a random forest score (RF-Score) towards correcting the bias of classical scoring systems. In order to evaluate binding affinity, numerous deep learning-based scoring functions (DLSF) have recently been created. They have used a variety of deep learning approaches with the given pose, including a 3D convolutional neural network (3D-CNN) and a graph convolutional network (GCN). Each voxel has features that describe internal characteristics including ionization, hydrophobicity, aromaticity, and hydrogen bonds, among others. Deep learning today uses the convolutional neural network (CNN) as a key tool for pattern recognition. 3D-CNN is considered to find a three-dimensional spatial feature, binding pose, and affinity patterns for 3D voxel-based approaches. The potential net performed better than the RF-Score using GCN for the non-covalent (O’Boyle et al. 2011; Ma et al. 2015). Additionally, a number of recent researches recommended an examination of feature weights, which helped to better expand the compound’s design.
9.2.2.6 Ligand-Based Approaches
The foundation of ligand-based approaches is the idea that molecules with comparable structural characteristics would interact with the same target. The main strategies in ligand-based methodologies are the quantitative structure-activity relationship (QSAR) models, three-dimensional QSAR, two-dimensional study fingerprint regions for the arrangement of the atoms (2D-QSAR), and estimation of quantitative associations (weights) between structure and its bioactivity (Yadav et al. 2020). A compound’s structural and physicochemical characteristics have several connections to its biological activity, for instance, the partition coefficient is closely connected to the hydrophobic effect, which results towards receptor affinity. Many quantitative representations of a chemical can be utilized for prediction, ranging from a straightforward atom count to the Lipinski rule of five (Yadav et al. 2022). In order to develop quantitative molecular descriptors of chemicals, there are numerous tools available. To produce molecular descriptors for bioinformatics and cheminformatics, there are three open-source programs: RDKit, OpenBabel, and chemical development kit (CDK). QSAR creates a model to predict the bioactivity of compounds based on the quantitative descriptors that are generated. On-target bioactivities and ADME attributes were included in the benchmark datasets (Kaggle datasets) for QSAR prediction that is Merck Molecular Activity Challenge (MMAC) issued in 2012. Deep learning-based QSAR calculations have done better than earlier RF QSAR predictions as an advanced deep learning technique methodology.
9.2.2.7 Chemogenomic Approaches
Target proteins and chemicals are both used in chemogenomic techniques. The excellence and variety of chemogenomic techniques are taken advantage of by the exponential growth of data on proteins, compounds, and drug-target interactions (DTI). Chemogenomic techniques are often divided into two groups: similarity methods and feature-based techniques.
-
(a)
Similarity-Based Approaches
In order to predict DTIs, similarity-based approaches focus on resemblances between the obtained protein and chemical structures. To develop most suitable similarity index between the proteins and chemicals, a variety of methods can be used such as the topological similarity in graphs and networks, normalized Smith-Waterman scores, Tanimoto coefficient, and pretense distance between protein domains. The bipartite local model (BLM) is a noteworthy study that makes use of the graph-based approach (Bleakley and Yamanishi 2009; Ding et al. 2013). BLM creates a bipartite graph connecting medications and targets, which expects the drug target interactions, and then aggregates both to get a final prediction.
-
(b)
Feature-Based Approaches
Target and compound feature vectors, which are fixed-length vectors describing significant physicochemical qualities, are used in feature-based approaches. Drug and target vectors are concatenated, and machine learning models are trained to categorize DTIs using feature vectors of interaction and labels. Moreover, the drug-target characteristics features can be analyzed by protein-protein and drug-drug interaction networks to improve prediction performance (Li et al. 2016; Lee and Nam 2018). Furthermore, applying a deep learning model to feature-based methods has been suggested in numerous papers as a way to improve the results for drug design. However, feature-based approaches have a number of drawbacks, one of which is the information that is lost during feature engineering.
9.2.2.8 AI-Guided ADMET Prediction
Optimizing pharmacokinetic parameters with absorption, distribution, metabolism, excretion, and toxicity is one of the crucial parts in the drug discovery process (ADMET). In order to effectively direct the stages of drug discovery, it is necessary to examine compound’s ADMET properties for the detailed understanding of complex biological mechanism (Gola et al. 2006). AI helps in understanding this complex human biological process to determine the results in a faster way with accuracy. The collection of bioactivity and data as well as sophisticated machine learning techniques, the pharmaceutical industries, as well as academic institutions have been drawn to in silico ADMET property predictions.
9.2.2.9 AI-Guided Lead Optimization
The phrase “finding a needle in a haystack” is used to describe the process of identifying a molecule that provides the appropriate pharmacological characteristics or has activity against biological targets. Researchers believe that there are roughly 1030–1060 chemical possibilities in the space of synthesizable compounds, although there are now only about 160 million chemicals listed in Chemical Abstracts Service. Too many resources and computational resources would be needed to fully count this enormous expanse.
These methods make use of deep learning techniques that have shown measurable effectiveness in the fields of machine translation and synthetic image synthesis. Knowledge of the chemical space distribution and performing targeted optimization to get the desired pharmacophore features are key aspects of performing deep generative models in the lead optimization areas (Brown et al. 2019). Although every method has its particular unique advantages, however the AI-guided methods provide number of advantages over other conventional approaches such as it is entirely data-driven and it can decrease human bias. Furthermore, using gradient-based optimization, the chemical space is explicitly modeled as a continuous function (Noorbakhsh-Sabet et al. 2019).
9.3 Applications
Healthcare professionals ought to be prepared for the approaching era of artificial intelligence and welcome the new capabilities that will enable more effective and efficient care. In this article, we examine machine learning’s uses, difficulties, ethical issues, and viewpoints in the fields of medicine, translational research, and public health.
9.3.1 Disease Prediction and Diagnosis
Although artificial intelligence is increasingly being used in healthcare, research still mostly focuses on cardiovascular, nervous system, and cancer related as these are the major causes for the ill health and death. Early diagnosis of a variety of diseases can now be accomplished by refining the extraction of clinical understandings and performing these from the well-trained and verified system. For instance, the Food and Drug Administration (FDA) of the United States has approved the use of diagnostic software intended to find wrist fractures in adult patients. More over, 6% of the adult population in the United States suffers from depression. Image heatmap pattern recognition was 74% accurate at predicting severe depressive illness.
Artificial intelligence has the ability to provide prompt and accurate disease diagnosis, according to several researches. For the classification of complicated and multifactorial diseases, supervised approaches are useful tools for capturing nonlinear interactions. Abedi V. et al. discovered in a research involving 260 individuals that the model can identify acute cerebral ischemia more accurately than skilled emergency medical peoples (Abedi et al. 2017). However, the noisy data and experimental constraints diminish the therapeutic value of the models; deep learning methods can solve these constraints by lowering the dimensionality of the data by layered auto-encoding analyses.
9.3.2 Clinical Trials and In Silico-Based Prediction
With the AI method, researchers can partially replace animals or people in a clinical trial and create virtual patients with particular traits to improve the results of such investigations.
The deep learning techniques can be used in pharmacokinetics and pharmacodynamics from the initial preclinical stage to the later post-marketing analysis, and they are especially useful for pediatric or orphan disease trials. In one study, researchers created a sizable in silico randomized, placebo-controlled Phase III clinical trial study in which they treated artificial Crohn’s disease patients using virtual therapies. However, with variable drug efficacy results revealed a favorable association between the baseline disease activity score and the decline in disease activity score. The investigational medicine GED-0301 did not receive a high score from the model, and this prediction was confirmed when the business that was conducting the phase III study on GED-0301 halted it after failing to pass an interim futility review. The design and discovery stages of a biomedical product, the identification of biomarkers, the optimization of dose, or the length of the proposed intervention can all benefit significantly from AI-guided in silico clinical trials.
9.3.3 Drug Discovery and Repurposing
Around 25% of altogether medications have been found as a result of unintentional bringing together of various areas. Due to the factors such as high costs of drug research, low success rates in clinical trials, the application of AI and ML is growing significantly and three-dimensional structural data that can aid in the characterization of pharmacological targets, and are used in the drug discovery process. The AI in drug repurposing process not only provides the new targets for the existing drugs but also reduces the expenditure cost.
For example, the DSP-1181 is the first AI-created medication to enter in clinical trials; it is a long-acting, powerful serotonin 5-HT1A receptor agonist. Exscientia is the biotech company which discovered DSP-1181 in collaboration with Sumitomo Dainippon Pharma of Japan, which noted that the time from screening to the conclusion of preclinical testing was less than 12 months as compared to 4 years utilizing conventional procedures. Researchers at the Massachusetts Institute of Technology (MIT) discovered the medication halicin, which is effective against bacterial type (Escherichia coli), using a machine learning algorithm (Stokes et al. 2020).
Moreover, artificial intelligence is used in deep learning of the mechanism of medication toxicity, for example, terbinafine toxicity. The antifungal terbinafine may cause liver damage in some patients, which has very negative health effects. In another example, the machine learning method was performed to determine potential biochemical routes of the terbinafine drug to identify the biotransformation mechanism by the liver. The student discovered that the terbinafine metabolism is a two-step process through the AI/ML algorithm data.
In other examples, sildenafil, a drug first created in 1989 to treat angina, was later discovered to be effective in treating erectile dysfunction and was given the name Viagra. Thalidomide was initially created to treat morning sickness, but it caused serious birth problems, including limb deformity, and was removed off the market. A few years later, scientists learned that thalidomide has an anti-angiogenesis effect and began using it to treat leprosy and multiple myeloma.
To advance the knowledge of understudied biological systems, AlphaFold AI technique revealed the possible predictions of five SARS-CoV-2 targets in 2020, including the SARS-CoV-2’s membrane protein, Nsp2, Nsp4, Nsp6, and papain-like proteinase (C terminal domain). The antiviral medications such as atazanavir, remdesivir, efavirenz, ritonavir, and dolutegravir were computationally identified by the MT-DTI technique.
9.4 Machine Learning
Machine learning, a well-known branch of artificial intelligence, used a large number of databases to identify different patterns of variable interactions. The ML can generate novel ideas, uncover previously unknown relationships, and be found to be helpful in obtaining a fruitful path for the drug development and research. Many fields, including data production and analytics, have adopted machine learning (ML). Algorithm-based approaches, like ML, have a strong mathematics and computational theory foundation. Many potential technologies have made use of ML models, including support vector machine-based improved search engines, deep learning (DL) assisted driverless automobiles, and advanced dialogue recognition technology.
Deep learning is a branch of machine learning that creates automated predictions from training datasets by simulating the functioning of the human intellect with numerous layers of artificial neuronal networks (Patel et al. 2020). Deep learning-based models frequently have several parameters and layers; as a result, model over-fitting may result in subpar prediction accuracy. Over fitting can be avoided by enlarging the training sample, reducing the number of hidden layers to obtain the balanced data. The example of the deep neural network application is to reduce the time it took to diagnose new outpatient cerebral hemorrhages by 96% with an accuracy of 84%.
9.4.1 Classifications
The machine learning methods are categorized into two types such as supervised and unsupervised methods (Table 9.1). In supervised learning, labels for fresh samples are determined using training examples with established labels. The regression and classification are useful applications of supervised learning. Examples of applications for supervised learning techniques include the identification of lung nodules from chest X-rays, risk estimation models for anticoagulation therapy, automated defibrillator implantation in cardiomyopathy, categorization of stroke and stroke mimics, identification of arrhythmia in electrocardiograms, and the designing of the in silico clinical trials. In addition to processing labeled input in supervised learning, generative deep neural networks (DNNs) can also be used to analyze unlabeled data. One of the most popular generative network topologies for unsupervised learning is the deep auto-encoder network (DEAN).
Unsupervised learning does not require labeled data and can find unseen patterns in the data that are frequently used for data exploration and the production of innovative ideas. Prior to recognizing patterns in high-dimensional data, the data are typically translated into a lower dimension using unsupervised learning methods. The unsupervised learning utilized to review failed clinical trials with drugs such as spironolactone, enalapril, and sildenafil versus placebo to revisit patients with heterogeneous conditions who had heart failure. The examination was done with three different studies to determine the patient’s recovery without any human intervention (Carracedo-Reboredo et al. 2021).
The reinforcement learning method uses trial-and-error to increase accuracy while combining supervised and unsupervised learning. In all stage of the drug discovery process, large amounts of data are essential for the creation, development, and feasibility of efficacious ML algorithms. In precision medicine and therapies within drug discovery, the dependence on large, high-quality datasets and recognized, well-defined training sets is very crucial for the study.
Apart from these classifications, other model classifications frequently used are binary, multiclass, multi-label, and imbalanced. The binary is a two-label classification that employs algorithms like logistic regression, k-nearest neighbors, choices trees, support vector machines, and naive Bayes, while the multiclass involves more than two labels using techniques such choices trees, support vector machines, naive Bayes, random forests, and gradient boosting. In contrast to multiclass, which predicts a single class label for each example, multi-label classifies jobs that have more than two labels. The imbalanced classification model is used to classify the class labels with unevenly distributed jobs.
The deep learning (DL) is a type of machine learning algorithms which is known for using higher level characteristics such as neural networks that are developed from a model of the human brain to enable computers to read, create, and learn complicated hierarchical representations. The input data are transformed into a more compounded output data as a result of this process. There are various kinds of DL architectures, and depending on how the training set is organized, each one may recognize patterns and extract high-level features in a particular way. In this chapter, we briefly discuss on the common architectures, such as the CNN, RNN, and generative networks.
Convolutional neural network (CNN) is one of the most widely used DL designs in various industries, including natural language processing, image and speech identification, and many other natural language processing (NLP). Another sample type of DL architecture is the recurrent neural network (RNN), which was specifically designed to handle sequence data, and has been successfully applied to NLP.
9.4.2 ML Algorithms Used in Drug Discovery
The use of multiple ML algorithms in drug discovery has considerably benefited pharmaceutical businesses. There are different types of ML algorithms models available for forecasting the chemical, biological, and physical properties of molecules in drug advancement method. All phases from the drug identification to the market surveillances of the drug discovery process can benefit from the use of ML algorithms. As an illustration, ML algorithms have been applied to discover novel therapeutic uses, forecast drug-protein interactions, identify medication efficacy, assure the presence of safety biomarkers, and enhance the bioactivity of molecules.
9.4.2.1 Naive Bayes
Machine learning algorithms seek out the most promising theory from a set of relevant data, in particular, for the class of an unknown data sample. According on the description provided by the vector values of each sample’s variables, Bayesian classifiers assign each sample to the most likely class. The technique assumes that the variables are independent in its most basic form, making it easier to apply Bayes’ Theorem (Madhukar et al. 2019). While the assumption that not all variables are equally significant is impractical, this family of classifiers known as NB (Naive Bayes) that comes from it achieves excellent results, despite the fact that sometimes their set of characteristics exhibits high interdependence. This algorithm provides a straightforward method which is quick and efficient that can handle noisy data. Although it provides better results even though the data volume is very high in terms of the number of samples because of the tiny datasets. It responds each variable as a definite one and employs frequency tables to extract information. However, it is not the best technique for large dimensional issues with many features and requires some kind of transformation when dealing with numerical variables.
9.4.2.2 Naive Bayes in Drug Discovery
The identification of potential drug targets has been done using this approach in drug discovery. They specifically created a Bayesian model that incorporates many data sources, such as data of known side effects or gene expression, and they achieved a model with 90% accuracy on more than 2000 compounds. There are reports that used an experimental approach on machine learning and molecular docking study to identify the potential inhibitors of DNA topoisomerase I enzyme of mycobacterium tuberculosis (MtTOP1) species and evaluated in vitro confirmation of their computational findings (Ekins et al. 2017). The AUC values for these predictions were 74%. In this, the drug prediction models are used in accordance with the ATM (Anatomical Therapeutic Chemical) system using the datasets from STITCH and ChEMB. The different types of molecular descriptors were analyzed for the structural information, and interactions with similar targets are displayed with an accuracy of 65%.
9.4.2.3 Support Vector Machines
Support Vector Machines (SVM) were first presented by Vapnik in the late 1970s. Due to the robustness and capacity to generalize in high-dimensional domains, particularly in bioinformatics, these are among the most extensively utilized approaches (Fernandez-Lozano et al. 2014). Sets of points in a particular space are used in machine learning to figure out how to handle brand new observations. These points are used by kernel-based approaches to determine how comparable the new observations are and to reach a conclusion.
9.4.2.4 Support Vector Machines in Drug Discovery
The SVM is one of the most often used models in bioinformatics because of its capacity to handle challenging issues that are complicated, nonlinear, high dimensional, and noisy. They have been utilized to classify pharmaceuticals based on their KEGG categorization, with an accuracy score of 83.9%. A brand new method for predicting intricate drug-target interaction networks using interaction matrices with function values of 80% was put forward. Additionally, by calculating several molecular descriptors and chemical indices using ChEMBL datasets with values near 70% in validation, it is able to predict the stability in human liver microsomes. The method used in expression data is an intriguing new method to anticipate a drug’s impact on a tumor line by learning more about the genes involved in the drug’s response in various tumor types (GEO).
For the prediction of HDAC1 inhibitors, SVMs were also applied to 3D-QSAR descriptors using a feature selection strategy described (Hu et al. 2016). The 2D-QSAR used to predict the compounds that inhibit the P-gp membrane protein target in the cancer study and wrapper feature selection models along with metaheuristic as a genetic algorithm produced promising results that were later confirmed by molecular docking approaches. Multiple Kernel Learning (MKL), which generates various linear combinations of SVMs with various parameters or kernels in an effort solve the problems, is an illustration of a sophisticated application of SVMs. Additionally, this enables the integration of many heterogeneous data sources, although at the expense of raising the computing cost.
9.4.2.5 Tree-Based Models
A decision tree is a hierarchical structure made up of nodes and the connections between them or branches. The method employed for classification issues is distinguished among methods for other sorts of problems, such as regression, survival, or outlier’s detection. In this, the root nodes, internal nodes, and terminal nodes were found within a decision tree’s hierarchical structure. The root node is found at the top of the tree model with one or more branches emerging from it but no branches reaching towards it. Regarding internal nodes, two or more branches originate from them and reach the next level of the hierarchy. There is no branches originating from the terminal nodes since they are located at the bottom of the hierarchy.
The out of bag error is equivalent to the error that the algorithm would make when the cross-validation is performed. The bagging approach in which the random forest (RF) divides the dataset into one-third part for validation and two-third part for training sets and analyzes to determine generalization error internally from each individual decision tree. Finally, because each decision tree is trained using various samples and characteristics, it is easy to estimate the importance of each attribute, while ignoring the others and lowering the problem’s dimensionality. Because of this, issues with very high dimensionality and noise are particularly well-suited for this technique.
9.4.2.6 Random Forest in Drug Discovery
When it comes to greater performance, speed, and generalizability, the RF model is the greatest among all other models. This model is deemed to be more suited and offers protein interactions with greater than 90% accuracy. They made use of the Open Babel descriptors and the GO and KEGG protein enrichment scores for the validation.
9.4.2.7 Artificial Neural Networks
The artificial neuron is a useful component of the network that accepts input from other components and processes it in some way to provide an output that can be processed by other components before talking about ANN. The artificial neurons may communicate with one another, just like natural neurons, and their connections are represented by weights, which are merely values that attempt to capture the synaptic force of a connection between two neurons. The net value, which sums together all the forces received by an artificial neuron or processing element, is considered first. The output of the processing element is determined by applying a trigger function after the net value calculation. The network of neurons can be created where the outputs of one neuron are used as the input for other neurons. It is important to realize that ANNs require input nodes, or neurons that receive data from the external world; these neurons are referred to as the network’s input layer. Additionally, the network involves output nodes, which are located in the hidden layer and transmit ANN results. The network’s hidden nodes, which transport data between neurons, are arranged into one or more hidden layers.
9.4.2.8 ANN in Drug Discovery
ACD (Available Chemicals Directory) and CMC (Comprehensive Medicinal Chemistry) data were used to train ANN and tree-based algorithms for drugs and non-drugs, respectively. The 2D descriptors provide the detailed information of the functional groups availability inside the molecules structures. However, 1D descriptors provide the information regarding the molecule’s molecular weight and hydrogen bond numbers for each available compound. An ANN with both 1D and 2D descriptors produced the greatest results, with an accuracy of 89%.
To forecast the initial carcinogenesis of substances suggested to be medications includes the calculation of six distinct types of descriptors with a deep learning model and an accuracy of 86% using 1003 chemicals from the Carcinogenic Potency Database. AUC of 76% can be achieved by beginning an experimental phase in the lab, generating a set of 2130 compounds of potential novel medications of interest for cardiotoxicity, computing each compound’s DRAGON 3456 descriptors, and including the analysis in a feature selection procedure.
9.4.2.9 De novo Molecular Design
Recent developments in ML have greatly improved the field of de novo or inverse molecular design. In a very short period of time, many intriguing strategies have been proposed. Recurrent neural networks (RNNs), generative adversarial networks (GANs), and auto-encoders, in particular, have been applied to the optimization of devices and the rational design of organic and inorganic materials. ReLeaSE is a deep reinforcement learning-based technology that produces chemical compounds and focuses on chemical collections with anticipated physical, chemical, and/or bioactivity features (RL). Both generating (G) and predictive (P) neural networks are used in the ReLeaSE method’s main workflow. The generative model G serves as an agent in this system by creating new, chemically viable compounds, whereas the predictive model P serves as a critic. P assigns a numerical reward (or penalty) to each created molecule in order to estimate the agent’s behavior.
9.4.2.10 Synthesis Planning
Recent advances in research, synthesis planning have made use of ML-based methodologies. Without human support, full syntheses of crucial chemicals for medicine were planned using the computer application Chematica. In order to identify the successful synthetic paths, the reaction guidelines are merged into graphs that connect lots of potential molecules with the chemical reaction knowledge. Retrosynthetic paths can be found using Monte Carlo tree search and symbolic AI without the aid of human expert rules and widely used today in the research organizations. Practically, all organic chemistry-related reported reactions were used to train this neural network. However, the synthetic chemists judged computer-generated pathways to be comparable to approaches described in the literature and with practical results.
9.5 Applications
9.5.1 CNS Disorder
Futuristic CNS drug discovery study will increasingly rely on AI/ML, mostly in the fields such as patient subtyping, identification of crucial disease drivers, estimation of cell type-specific drug response, sovereign novel drugs design, and with better BBB (blood brain barrier) permeability tests. The role of AI/ML is now being constrained by structural limitations in data and algorithms. However, in the long run, we will be able to create CNS disease treatments that are more potent because of ongoing and new breakthroughs in AI/ML approaches to neuropharmacology (Carpenter and Huang 2018).
9.5.2 Discovering Novel Antimicrobial Agents
Several reported works showed how ML may be used in the context of antibiotic discovery to learn small molecule structural properties from screenings that contain prevailing antimicrobial activity to advance novel antimicrobials. By first creating a genetic library of hypomorph knockdowns for these crucial genes and then screening 50,000 chemical compounds against these hypomorphs, Johnson et al. done a screening for finding biochemical inhibitors of key genes in M. tuberculosis.
The supervised ML classification evaluates the novel classes of chemical inhibitors for existing drug targets and recent discovered targets, validated in wild-type cells against standard antibiotics. A deep learning ML model used screening of several molecules with different structural features for antimicrobial activity against E. coli in order to predict antimicrobial functions. To predict the inhibition of Escherichia coli growth, the scientists used a training set of 2335 molecules for a DNN model. The model was then run on more than 107 million molecules from various chemical libraries.
9.5.3 Epidemic COVID
In order to find effective medications for 65 human proteins (targets) that had shown to interact with SARS-CoV-2 proteins, Kowalewski and Ray created machine learning (ML) models (Kowalewski and Ray 2020). They infer it from inhalation treatments to directly target the injured cells because the virus is known to target the respiratory tract, including nose epithelial cells, upper airway, and lungs. In order to rank the chemicals and identify medications that share the identical chemical space, they gathered 14 million compounds from ZINC databases and used machine learning algorithms to obtain vapor pressure and mammalian toxicity. The objective of the study was to create a short- and long-term pipeline for use in the future. They also developed models that might forecast drug efficacy using SVM and RF.
9.6 Drug Discovery Process
9.6.1 AI and Machine Learning in Precision Drug Discovery
A new approach to disease prevention and treatment called precision medicine considers a person’s unique gene, lifestyle, and environmental variations. Based on the genetic profiles of the patients, this technique aids scientists and medical professionals in more precisely preventing and treating disease. Powerful supercomputer infrastructure and innovative algorithms that can autonomously learn in an unheard of fashion from the trained set of data are needed to make the strategy more comprehensive. Medical professionals’ cognitive abilities and biomedical data are used by artificial intelligence to achieve results.
With technological advancements, the future of healthcare will change as a result of the creation of large digital datasets obtained through next-generation sequencing (NGS), use of image processing algorithms, patient-related health records, and data resulting from significant clinical trials. Oncology can benefit greatly from machine learning, which is frequently used in precision medicine. Complex neural networks are used to generate diagnostic images and genetic data, which are then used to forecast the likelihood of disease and treatment outcomes (Dlamini et al. 2020). In radiomic field of machines that produces diagnostic images to discover malignant tumors that are undetectable by the human sight, the implementation of AI and ML technologies in healthcare is done to enhance illness management and deliver high-quality medical care (Fig. 9.3).
By highlighting diverse uses of AI in oncology healthcare, such as next-generation sequencing (NGS), advancements in medical imaging, digital pathology, and drug discovery, we present information on AI and precision oncology towards clinical environment for cancer management.
9.6.1.1 NGS and Molecular Profiling
The NGS technique utilizes RNA sequencing to discover novel RNA variants and splice sites, or quantify mRNAs for gene expression analysis. Genomic profiling is conceivable and offers promise for the future of precision oncology to the implementation of NGS, which is quickly evolving the field of genomic sequencing for clinical use. Advanced NGS methods can sequence DNA and RNA on a wide scale with high-throughput data and at a lower cost. Numerous sequencing techniques, such as whole-genome, whole-exome, RNA, target, and whole-transcriptome shotgun sequencing, as well as methylation sequencing, are made possible by NGS. DNA or RNA samples from blood samples, tumor samples, cell lines, formalin-fixed paraffin-embedded (FFPE) blocks, and liquid biopsies can all be used for sequencing. As part of the Human Genome Project, the first whole-genome sequencing was carried out at significant expense and over a lengthy period of time. To detect changes in the cellular transcriptome and changed molecular pathways, RNA sequencing is frequently employed in cancer research and diagnosis (Jiang et al. 2017).
The advantages of RNA profiling of cancer models for treatment results have been demonstrated in clinical studies that sequence RNA using precise oncology protocols. RNA profiling is applied to RNA extracted from blood or a tumor sample. According to a study, RNA profiling should be a standard of care for oncology patients because it may have potential clinical benefits, particularly for cancers that are challenging to treat in children and young adults. The study also illustrates the impact of precision oncology. According to the study’s findings, about 70% of the gene expression data acquired from RNA sequencing may have clinical applications.
Identification of gene expression signatures to decipher the underlying molecular pathways of cancer and the detection of RNA mutations with implications for alternative splicing are the two most significant and often used applications of RNA sequencing. However, many NGS approaches have drawbacks such as labor-intensiveness, the introduction of sequencing coverage mistakes, and expense. Acquiring pertinent data from NGS datasets is becoming more and more time-effective because of the developments in AI and computational approaches, with some platforms enabling real-time viewing.
9.6.1.2 Biomarkers
Molecular biomarkers are often used in the cancer diagnostics in the early detection of the diseases. Different biomarkers are used, for example, circulating cancer antigen is used to detect ovarian cancer at early stage, carcinoembryonic antigen is used to monitor relapse of colorectal cancer, and estrogen receptor 1 (ESR1) is used for the prognosis prediction and treatment outcomes in breast cancer. Cancer management can be improved by locating biomarkers in the early disease prevention and prognosis prediction for successful treatment. By locating germline DNA alterations and doing full transcriptome analyses by RNA sequencing, novel molecular biomarkers for various malignancies can be uncovered and utilized to detect the diseases. The potential of RNA sequencing in the development of biomarkers for diagnosis and as a prognostic predictor has been demonstrated in large consortia studies like the Cancer Genome Atlas (TCGA). Aside from pathogenic mutations and changed expression or activity of proteins that regulate significant cellular complexes, these investigations also clarified predicted biomarkers that fuel transformation. Additionally, Shallow full genome sequencing was used to identify copy number variants (CNV) in breast cancer utilizing FFPE samples to diagnoses for breast cancer, lung cancer, and neuroblastoma.
9.6.1.3 Medical Imaging
Applications of AI in radiology are essential for many modalities with enhanced quality, including X-rays, ultrasounds, computed tomography (CT/CAT), magnetic resonance imaging (MRI), positron-emission tomography (PET), and digital pathology. Images are analyzed quickly and accurately using highly specialized algorithms. Accurate diagnosis depends in large part on the ability to distinguish between normal and aberrant medical images. Early cancer detection is extremely important because it will result in a better prognosis and treatments. The future of AI in medical imaging will be focused on increasing speed and lowering costs. AI has already contributed to medical imaging by improving image quality, computer-aided image interpretation, and radiomics (Lewis et al. 2019). The main advancements and breakthroughs of artificial intelligence in healthcare have been widely used for clinical purposes in medical imaging.
9.6.1.4 Radiographic Imaging
In order to accurately diagnose and treat patients, which can take time and be subject to human error and variability, it is necessary to extract pertinent quantitative data from medical images, such as size, symmetry, location, volume, and form. For routine clinical treatment, automated medical imaging analysis is highly necessary. The radiographic imaging includes three stages: the first one is the image segmentation, which detects the image of interest and defines its boundaries; the second one is the image registration, which establishes the spatial three-dimensional relationship between images; and image visualization, which displays pertinent information for precise interpretation, is necessary to analyze the medical images accurately. However, despite of the advancement in the medical imaging, there are still some complications with data complexity, object complexity, and validation issues.
The deep learning-based algorithms for an automated detection system for chest radiography are the recent advancement. However, the chest radiograph analyses for thoracic disease are difficult and error-prone, and the highly skilled radiographers are required to analyze the images. These AI methods were created to differentiate between common thoracic disorders, including pulmonary malignant tumors.
Imaging in medicine using AI extends beyond radiology. The advent of digital pathology will soon revolutionize pathology laboratories. The gold standard for pathology for many years has been microscopic examination of stained cells and tissues. By reducing labor-intensive microscopic tasks, boosting efficiency, and maintaining the quality for better clinical treatment, technological and AI advancements will transform pathology. Digital pathology that incorporates AI improves workflow, enables doctors to analyze images for precise interpretation, and lowers subjectivity by standardizing processes. Additionally, digital pathology enables reduced fluctuation in color information and larger-scale image viewing. This makes it possible to successfully find distinctive markers linked to disease-specific biomarkers for diagnosis, prognosis, and treatment (Bera et al. 2019).
9.6.2 Repurposed Drug/Drug Discovery by AI/ML Approach
About 25% of all medications have been found as a result of unintentional bringing of various areas. Pharmaceutical companies prefer targeted drug discovery over conventional blind screening because it has a clear mechanism and a better success rate and is less expensive. Due to the following factors such as high costs of drug research, growing accessibility of three-dimensional structural data that can aid in the characterization of pharmacological targets, and shockingly low success rates in clinical trials, machine learning is currently used in the drug discovery process. Cross-domain linkage can be accomplished using machine learning as a bridge. By identifying contextual cues like a discussion of a drug’s indication or side effects, it may recognize a newly approved drug.
Despite these innovative methods for drug development, there are still significant obstacles, such as data access and the fact that various datasets are typically kept in a number of separate repositories. Additionally, clinical trial raw data and other preclinical study raw data are often unavailable. The utilization of pharmacological information to gain knowledge into mechanism of action by employing methods like similarity metrics across all diseases to uncover shared pathways is just one example of how artificial intelligence has been successful when applied to available data. Another illustration is the use of NLP to find hidden or unexpected relationships that may be significant in the identification of probable pharmacological side effects based on scholarly articles.
Few organizations have started to make use of these developments to accelerate the release of COVID-19 medications and better understand how the immune system combats the illness. Pharmaceutical companies GlaxoSmithKline (GSK) and Vir Biotechnology teamed together at the beginning of April to accelerate coronavirus treatment development using CRISPR and artificial intelligence. Additionally, in the academic world, the Human Immunomics Initiative, launched recently by the Harvard T. Chan School of Public Health and the Human Vaccines Project, employs artificial intelligence to accelerate the production of antibodies for a variety of illnesses, including COVID-19. A team from Southern Illinois University (SIU) recently developed an information visualization tool that shows users the locations of known COVID-19 instances using GPS data. A contact following application powered by Bluetooth technology has also been developed in cooperation between Google and Apple. These techniques might be successful in collecting a lot of precise data. Businesses that have developed wellness profiles for people based on a fundamental understanding of the infection are conducting research into various medicine delivery methods that have been successfully licensed. The two most well-known examples of this in relation to COVID-19 to date are hydroxychloroquine (recommended for the treatment of malaria) and remdesivir (for the treatment of Ebola). The effectiveness dataset for these drugs may therefore be a decent input for an AI model. The businesses using artificial intelligence (AI) to repurpose currently available drugs for COVID-19 are listed in Table 9.2.
9.7 Limitations of AI/ML Approaches
The use of routine clinical NGS sequencing for cancer diagnosis and management faces significant challenges with data interpretation. Large servers and knowledgeable bioinformaticians are needed for the management and interpretation of big data. The provided datasets for diagnosis contain details on variants that can be classified as benign, likely benign, variant of unknown importance, likely pathogenic, and pathogenic variants. It is crucial to classify all variations into groups and understand their clinical importance. Data acquired can be helpful for cancer management in addition to diagnosis.
However, the drawbacks of whole-genome and exome sequencing include high costs, a heavy computing burden, and challenging data interpretation. In the following 10 years, further development of NGS platforms may result in cost reductions without a reduction in quality.
Despite the advantages of AI, there are still several obstacles to its implementation in the healthcare industry. Big data and costs are on the rise as a result of automated computation. Due to their reliance on specialized computational requirements for rapid data processing, AI systems can be costly. Additional quality procedures are also necessary for these systems. The targeted users must receive training and gain a knowledge of the technology in order to implement AI-based solutions for everyday clinical practice. Rigby emphasized the moral dilemma presented by AI in healthcare. It is crucial to resolve the ethical problem of using patient data without authorization or justification in light of the big data boom. Additionally, in order to safeguard patient privacy and safety, ethical norms and guidelines are necessary.
Despite appearing to be effective and acceptable in the de novo lead creation approach, the connecting mechanism has some drawbacks. The first restriction is that for proper linking: the linking fragments must be precisely positioned in the cavity. De novo design is additionally assumed to be totally automated, but still requires some arduous manual labor. Furthermore, it is not always simple to manufacture the chemicals created using this method in a lab. Thus, new software that includes de novo compound design and considers synthesis parameters is required.
Although the connecting approach in the de novo lead generation method appears to be effective and acceptable, there are certain restrictions. The first restriction is that for proper linking, the linking fragments must be precisely positioned in the cavity. De novo design is additionally assumed to be totally automated, but still requires some arduous manual labor. Furthermore, it is not always simple to manufacture the chemicals created using this method in a lab. Thus, new software that includes de novo compound design and considers synthesis parameters is required.
Overall, the complexity of small molecule drug discovery will increase. DL ought should be able to manage that complexity since it was made for complicated simulation. Additionally, using DL techniques, we should not limit ourselves to making the conventional predictions about biological activities, ADMET properties, or pharmacokinetic simulations. Instead, it might be possible to systematically integrate all the data and information and reach a new level of AI in drug discovery.
9.8 Conclusions and Future Perspectives
The ultimate goal of machine learning is to create algorithms that can learn continually from fresh information and data in order to find solutions to a wide range of problems. Complex algorithms have appealing prospects for precision medicine, but they also present computing difficulties. To realize this potential, unique solutions are needed for at least three technical problems:
-
1.
The quantity and size of data inputs, outputs, and attributes. This problem can be partially solved by leveraging CPU clusters, data sharing systems, cloud computing, and deep learning techniques.
-
2.
Variety—diverse types of data (picture, video, and text). This problem can be partially solved by integrating data from many sources using novel deep learning techniques.
-
3.
Velocity—the pace of streaming data. To solve this problem, online learning techniques can be developed.
Machine learning techniques used nowadays are very similar to real-world situations. As a result of the quick improvements in technology, algorithms will take on duties that were previously the domain of humans. Radiologists and anatomical pathologists will lose a lot of their jobs as a result of machine learning’s capacity to turn data into insight. Clinical medicine, however, has always required physicians to manage enormous amounts of data, from the history and physical examination to the laboratory and imaging examinations, as well as the more recent genetic data. Effective medical professionals have always been able to handle this complexity.
We anticipate that as more scientists become aware of its potential, the usage of ML in VS for drug discovery will continue to expand in the search of new drugs. Drug discovery will undoubtedly become more effective and less expensive, thanks to the combined efforts of computer science and medicinal chemistry.
References
Abedi V, Goyal N, Tsivgoulis G et al (2017) Novel screening tool for stroke using artificial neural network. Stroke 48(6):1678–1681. https://doi.org/10.1161/STROKEAHA.117.017033
Ballester PJ, Mitchell JBO (2012) A machine learning approach to predicting protein-ligand binding affinity with applications to molecular docking. Bioinformatics 26(9):1169–1175. https://doi.org/10.1093/bioinformatics/btq112
Bera K, Schalper KA, Rimm DL, Velcheti V, Madabhushi A (2019) Diagnosis and precision oncology. Nat Rev Clin Oncol 16(11):703–715. https://doi.org/10.1038/s41571-019-0252-y
Bleakley K, Yamanishi Y (2009) Supervised prediction of drug-target interactions using bipartite local models. Bioinformatics 25(18):2397–2403. https://doi.org/10.1093/bioinformatics/btp433
Brown D (2007) Unfinished business: target-based drug discovery. Drug Discov Today 12(23–24):1007–1012. https://doi.org/10.1016/j.drudis.2007.10.017
Brown N, Fiscato M, Segler MHS, Vaucher AC (2019) GuacaMol: benchmarking models for de novo molecular design. J Chem Inf Model 59(3):1096–1108. https://doi.org/10.1021/acs.jcim.8b00839
Carpenter KA, Huang X (2018) Machine learning-based virtual screening and its applications to Alzheimer’s drug discovery: a review. Curr Pharm Des 24(28):3347–3358. https://doi.org/10.2174/1381612824666180607124038
Carracedo-Reboredo P, Liñares-Blanco J, Rodríguez-Fernández N et al (2021) A review on machine learning approaches and trends in drug discovery. Comput Struct Biotechnol J 19:4538–4558
Chen B, Butte AJ (2016) Leveraging big data to transform target selection and drug discovery. Clin Pharmacol Ther 99(3):285–297. https://doi.org/10.1002/cpt.318
DiMasi JA, Grabowski HG, Hansen RW (2016) Innovation in the pharmaceutical industry: new estimates of R & D costs. J Health Econ 47:20–33. https://doi.org/10.1016/j.jhealeco.2016.01.012
Ding H, Takigawa I, Mamitsuka H, Zhu S (2013) Similarity-basedmachine learning methods for predicting drug-target interactions: a brief review. Brief Bioinform 15(5):734–747. https://doi.org/10.1093/bib/bbt056
Dlamini Z, Francies FZ, Hull R, Marima R (2020) Artificial intelligence (AI) and big data in cancer and precision oncology. Comput Struct Biotechnol J 18:2300–2311
Ekins S, Godbole AA, Kéri G et al (2017) Machine learning and docking models for mycobacterium tuberculosis topoisomerase I. Tuberculosis 103:52–60. https://doi.org/10.1016/j.tube.2017.01.005
Farghali H, Canová NK, Arora M (2021) The potential applications of artificial intelligence in drug discovery and development. Physiol Res 70:715–722. https://doi.org/10.33549/physiolres.934765
Fernandez-Lozano C, Gestal M, González-Díaz H et al (2014) Markov mean properties for cell death-related protein classification. J Theor Biol 349:12–21. https://doi.org/10.1016/j.jtbi.2014.01.033
Ferrero E, Dunham I, Sanseau P (2017) In silico prediction of novel therapeutic targets using gene-disease association data. J Transl Med 15(1):1–16. https://doi.org/10.1186/s12967-017-1285-6
Gola J, Obrezanova O, Champness E, Segall M (2006) ADMET property prediction: the state of the art and current challenges. QSAR Comb Sci 25(12):1172–1180. https://doi.org/10.1002/qsar.200610093
Hu B, Kuang ZK, Feng SY et al (2016) Three-dimensional biologically relevant Spectrum (BRS-3D): shape similarity profile based on PDB ligands as molecular descriptors. Molecules 21(11). https://doi.org/10.3390/molecules21111554
Imrie F, Bradley AR, Van Der Schaar M, Deane CM (2018) Protein family-specific models using deep neural networks and transfer learning improve virtual screening and highlight the need for more data. J Chem Inf Model 58(11):2319–2330. https://doi.org/10.1021/acs.jcim.8b00350
Jiang F, Jiang Y, Zhi H et al (2017) Artificial intelligence in healthcare: past, present and future. Stroke Vasc Neurol 2(4):230–243. https://doi.org/10.1136/svn-2017-000101
Keiser MJ, Setola V, Irwin JJ et al (2009) Predicting new molecular targets for known drugs. Nature 462(7270):175–181. https://doi.org/10.1038/nature08506
Kim H, Kim E, Lee I et al (2020) Artificial intelligence in drug discovery: a comprehensive review of data-driven and machine learning approaches. Biotechnol Bioprocess Eng 25:895–930
Kolluri S, Lin J, Liu R, Zhang Y, Zhang W (2022) Machine learning and artificial intelligence in pharmaceutical research and development: a review. AAPS J 24(1):19. https://doi.org/10.1208/s12248-021-00644-3
Kowalewski J, Ray A (2020) Predicting novel drugs for SARS-CoV-2 using machine learning from a >10 million chemical space. Heliyon 6(8):e04639. https://doi.org/10.1016/j.heliyon.2020.e04639
Lee I, Nam H (2018) Identification of drug-target interaction by a random walk with restart method on an interactome network. BMC Bioinformatics 19(Suppl 8):208. https://doi.org/10.1186/s12859-018-2199-x
Lewis SJ, Gandomkar Z, Brennan PC (2019) Artificial Intelligence in medical imaging practice: looking to the future. J Med Radiat Sci 66(4):292–295. https://doi.org/10.1002/jmrs.369
Li ZC, Huang MH, Zhong WQ et al (2016) Identification of drug-target interaction from interactome network with “guilt-by-association” principle and topology features. Bioinformatics 32(7):1057–1064. https://doi.org/10.1093/bioinformatics/btv695
Ma J, Sheridan RP, Liaw A, Dahl GE, Svetnik V (2015) Deep neural nets as a method for quantitative structure-activity relationships. J Chem Inf Model 55(2):263–274. https://doi.org/10.1021/ci500747n
Madhukar NS, Khade PK, Huang L et al (2019) A Bayesian machine learning approach for drug target identification using diverse data types. Nat Commun 10(1):1–14. https://doi.org/10.1038/s41467-019-12928-6
Mamoshina P, Volosnikova M, Ozerov IV et al (2018) Machine learning on human muscle transcriptomic data for biomarker discovery and tissue-specific drug target identification. Front Genet 9:1–10. https://doi.org/10.3389/fgene.2018.00242
Mohamed SK, Nováček V, Nounu A (2020) Discovering protein drug targets using knowledge graph embeddings. Bioinformatics 36(2):603–610. https://doi.org/10.1093/bioinformatics/btz600
Noorbakhsh-Sabet N, Zand R, Zhang Y, Abedi V (2019) Artificial intelligence transforms the future of health care. Am J Med 132:795–801
O’Boyle NM, Banck M, James CA et al (2011) Open babel: an open chemical toolbox. J Cheminform 3(33):1–14
Patel L, Shukla T, Huang X, Ussery DW, Wang S (2020) Machine learning methods in drug discovery. Molecules 25(22). https://doi.org/10.3390/MOLECULES25225277
Petyuk VA, Chang R, Ramirez-Restrepo M et al (2018) The human brainome: network analysis identifies HSPA2 as a novel Alzheimer’s disease target. Brain 141(9):2721–2739. https://doi.org/10.1093/brain/awy215
Stepniewska-Dziubinska MM, Zielenkiewicz P, Siedlecki P (2018) Development and evaluation of a deep learning model for protein–ligand binding affinity prediction. Bioinformatics 34(21):3666–3674. https://doi.org/10.1093/bioinformatics/bty374
Stokes JM, Yang K, Swanson K et al (2020) A deep learning approach to antibiotic discovery. Cell 181(2):475–483. https://doi.org/10.1016/j.cell.2020.04.001
Wong CH, Siah KW, Lo AW (2019) Estimation of clinical trial success rates and related parameters. Biostatistics 20(2):273–286. https://doi.org/10.1093/biostatistics/kxx069
Yadav V, Tonk RK, Khatri R (2020) Molecular docking, 3D-QSAR, fingerprint-based 2D-QSAR, analysis of pyrimidine, and analogs of ALK (anaplastic lymphoma kinase) inhibitors as an anticancer agent. Lett Drug Des Discov 18(5):509–521. https://doi.org/10.2174/1570180817999201123163617
Yadav V, Reang J, Vinita TRK (2022) Ligand-based drug design (LBDD). In: Rudrapal M, Egbuna CBT-CADD (CADD): FL-BM to S-BA (eds) . Elsevier, Drug discovery update, pp 57–99
Zhavoronkov A, Vanhaelen Q, Oprea TI (2020) Will artificial intelligence for drug discovery impact clinical pharmacology? Clin Pharmacol Ther 107(4):780–785. https://doi.org/10.1002/cpt.1795
Zhu Z, Zhang F, Hu H, Bakshi A, Robinson MR et al (2016) Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 48(5):481–487. https://doi.org/10.1038/ng.3538
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Yadav, V., Reang, J., Vinita, Tonk, R.K. (2023). Artificial Intelligence and Machine Learning in Drug Discovery. In: Rudrapal, M., Khan, J. (eds) CADD and Informatics in Drug Discovery. Interdisciplinary Biotechnological Advances. Springer, Singapore. https://doi.org/10.1007/978-981-99-1316-9_9
Download citation
DOI: https://doi.org/10.1007/978-981-99-1316-9_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-99-1315-2
Online ISBN: 978-981-99-1316-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)