Abstract
Peptide-based drugs/antibiotics are increasingly becoming preferable due to greater incidence of multidrug resistance towards non-peptidic drugs. In addition, several non-peptidic compounds also elicit unfavorable or toxic side effects. Consequently, besides bacteria, plants and animal-based sources, fungi have also been explored very much in the search for novel therapeutic peptides that can be more efficacious and safer. Based on the biosynthetic pathways, these peptides can be classified as (i) ribosomal peptides and (ii) non-ribosomal peptides (NRPs). Several NRPs from diverse fungal sources (soil, marine, and endophytes) have been identified and comprehensively characterized, whereas studies on ribosomal peptides of fungal origin are limited. In terms of molecular architecture, fungal peptides can be classified into four major categories: (i) linear peptides with modified N- and C-termini (e.g., peptaibiotics and peptaibols); (ii) head-to-tail (backbone) cyclized peptides; (iii) cyclic depsipeptides; and (iv) sidechain cyclized peptides. In this chapter, salient features of some major fungal peptides that have been found to elicit various kinds of biological activities are presented. A few aspects on some fungal peptide-based products of pharmaceutical or medical relevance, e.g., caspofungin (brand name: Cancidas) and cyclosporin A, which are already available in the market, are also briefly delineated.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
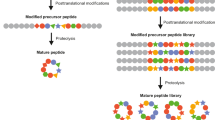
Right from the time of serendipitous finding of penicillin till now, fungi have always been sought after for discovering several drug compounds. Fungi also have been useful as biocontrol agents, particularly for agricultural applications [1]. Among various types of bioactive molecules that have been identified from different fungal species, peptides constitute an important class. Fungal peptides not only elicit very interesting in vitro biological functions, but some have also been sold as drugs, e.g., caspofungin (brand name: Cancidas; Drug Bank Accession No. DB00520) and micafungin (brand name: Mycamine, Drug Bank Accession No. DB01141), while some have been found to be useful in the preclinical stages of drug discovery, e.g., NZ2114, a derivative of a defensin plectasin [2]. The fungal peptides encompass both ribosomal and non-ribosomal classes, depending on the biosynthetic pathways. The peptides synthesized by the conventional ribosomal machinery are called as RiPPs, which means “Ribosomally synthesized and posttranslationally modified peptides” [3,4,5]. When compared to the non-ribosomal peptides from fungi, only a few studies on fungal RiPPs are available in the literature, and those are relatively recent [5]. Amatoxins and borosins are some examples of fungal RiPPs. The biosynthesis of non-ribosomal peptides involves very large multi-modular enzymes, called non-ribosomal peptide synthetases (NRPSs) [6, 7]. The NRPSs utilize both proteinogenic and non-proteinogenic amino acids in the biosynthesis of non-ribosomal peptides (NRPs). α-Amino isobutyric acid (Aib), isovaline (Iva), γ-amino butyric acid (GABA), and anthranilic acid (ATA) are a few examples of non-proteinogenic amino acids that are found in several NRPs. Additionally, several fungal NRPSs also engage “hydroxy acids” during biosynthesis, thereby producing a different class of NRPs called “depsipeptides” [8]. Because of the incorporation of the hydroxy acid, depsipeptides contain at least “an ester bond” in their backbone, in addition to the usual presence of the peptide/amide bonds.
Based on the molecular structure/architecture, fungal peptides can be classified into four major categories: (i) linear peptides with modified N- and C-termini; (ii) head-to-tail (backbone) cyclized peptides; (iii) cyclic depsipeptides; and (iv) sidechain cyclized peptides. Cyclic depsipeptides would encompass both backbone and sidechain cyclized peptides. The category, sidechain cyclized peptides, mostly pertains to disulfide-bonded peptides. The differences in the molecular architecture between these four categories can be understood from Fig. 6.1.
Fungal peptides have been identified from diverse species that inhabit various ecosystems such as soil, marine, and plants (endophytes) and also those that thrive in insect hosts, e.g., entomopathogens [9, 10]. Since several fungal NRPs have modified/blocked N- and C-termini including the cyclic peptides, sequence elucidation of such peptides has been usually carried out by spectroscopic methods rather than by Edman’s degradation method (N-terminal sequencing). Nuclear magnetic resonance (NMR) spectroscopy involving homo- and hetero-nuclear multidimensional methods and mass spectrometry (MS)-based methods have proven to be successful in deducing the molecular structure and the sequence of several fungal NRPs. Marfey’s method has been widely applied for determining stereochemical configuration of the constituting amino acid residues. High-performance liquid chromatography (HPLC), particularly reverse phase (RP)-HPLC, has been instrumental in advancing as well as accelerating the process of discovering many NRPs of fungal origin, especially to isolate pure peptidic components for the purpose of carrying out biological assays. Further, combining chromatography with MS too has been immensely fruitful not only for sequencing various fungal NRPs but also to identify several variants, which possess microheterogeneous sequences. In the earlier years, gas chromatography (GC)-MS was utilized for sequencing [11,12,13,14], and of late, especially during the last three decades or so, RP-HPLC and MS have been increasingly applied for identifying and sequencing fungal NRPs [15,16,17]. Online LC-MS also has been attempted to identify and characterize fungal peptides [18,19,20,21,22]. Salient features including molecular structural properties and biological function of both non-ribosomal and ribosomal fungal peptides are presented in this chapter.
2 Non-ribosomal Fungal Peptides
2.1 Peptaibiotics: Peptaibols (Linear NRPs)
Many fungi produce linear peptides having a very high content of Aib (c.f. vide supra) called “peptaibiotics” [23]. A major class within peptaibiotics is constituted by “peptaibols,” which possess acetylated N-termini and C-terminus alcohol [24]. The name “peptaibol” was proposed/coined considering the prevalence of “Aib” in the sequences and the presence of the C-terminus alcohol group in these peptides, viz., “pept + aib + ol” [25]. In addition to Aib, several peptaibols are also known to have “isovaline” (ethylalanine), which is another α,α-dialkyl α-amino acid similar to Aib [26, 27]. Thus far, more than 700 peptaibols have been identified, and their lengths are in the range 4–22 including the C-terminus alcohol residue [22, 28,29,30,31,32].
With regard to the source, relatively a larger number of peptaibols have been identified from Trichoderma species [22, 25, 30, 32]. In addition to the soil fungi, peptaibols have been discovered from marine fungi as well, e.g., short-length trichobrachins A and B containing 11 residues and long-chain peptaibols composed of 20 residues have been reported from marine-derived strain of Trichoderma longibrachiatum [33, 34]. Recently, five new 15-residue-long pentadecaibins I–V were reported from marine-derived Trichoderma sp. of the Harzianum clade [35]. Further, various peptaibols have been reported from Emericellopsis sp., Cephalosporium pimprina Thirum., and Acremonium sp. also [12,13,14].
Peptaibols have the ability to insert into the lipid bilayer architecture of the cellular membranes and form transmembrane ion channels or pores [36, 37]. So, peptaibols are also called as “channel-forming peptides (CFPs),” which can serve as models to understand the behavior of ion-channeling activity that is usually carried out by transmembrane (receptor) proteins [36, 38]. Most of the channels formed by the peptaibols get activated upon application of electric field, and hence, they are also referred to as voltage-dependent ion channels [38]. The first peptaibol that was subjected to voltage-dependent studies was alamethicin [39]. Incidentally, alamethicin is also the first peptaibol, whose three-dimensional (3D) molecular structure was elucidated by X-ray diffraction, which was reported by Fox and Richards in 1982 (vide infra) [40]. This 3D molecular structure could provide insights into mechanisms of ion-channel formation aiding in the translocation of metal ions across the membrane. As a representative example, the molecular structure of 20-residue-long alamethicin F-30 is shown in Fig. 6.2a. Alamethicin F-30 has eight Aib residues and C-terminus phenylalaninol. 3D molecular structures of several other natural peptaibols and synthetic analogues were also determined by X-ray diffraction, NMR spectroscopy, and circular dichroism spectroscopy [41,42,43,44,45,46,47,48,49,50].
Most of these structures have been found to adopt largely helical conformations [51, 52]. Though Aib is known to predominantly nucleate helical conformations, it gets accommodated in other types of conformations, e.g., polyproline II and extended conformations also, as reviewed by Aravinda et al. (2008) [51].
In terms of primary structure, peptaibols are often produced as mixtures of very closely related peptides that have microheterogeneous sequences (i.e., isoforms), which may be attributed to their biosynthetic route that involve NRPSs [27]. The microheterogeneity can be due to exchanges or replacements of amino acids such as Gly ↔ Ala, Ala ↔ Aib, Aib ↔ D-Iva, and Gln ↔ Glu, only at certain specific positions in their sequences, which causes differences of 1 Dalton (Da) or 14 Da or 15 Da between their molecular masses. Often such replacements do not even alter their molecular masses, but only lead to some minor variations in their sequences, viz., isobaric analogues. Another striking observation is that the occurrence of the dipeptide fragments Aib-Aib and Aib-Pro is preponderant in several peptaibol sequences [53]. Based on amino acid sequence identities, peptaibols are classified into nine superfamilies. Position-specific preferences of amino acids and the lengths of the peptides distinguish the members between the superfamilies [52].
Lipopeptaibols and Other Peptaibiotics
Other than acetylated N-terminus, peptides possessing N-terminus acyl modification, e.g., octanoyl, dodecanoyl, etc., also have been identified from some fungi. These are called “lipopeptaibols,” e.g., trichogin A IV from Trichoderma longibrachiatum [41], trikoningins KB I and II from Trichoderma koningii Oudem. (collected in Uruguay) [54], halovirs from the marine fungus of the genus Scytalidium [55], and lipovelutibols from the Himalayan Trichoderma velutinum [17]. Further, instead of C-terminus alcohol, fungal peptaibiotics with C-terminus modified by amine or amide functional group too have been discovered, for instance, leucinostatins, efrapeptins (Fig. 6.2b) and trichopolyns [26, 56]. The C-termini of efrapeptins contain an unusual moiety: 1,5-diazabicyclo [4:3:0] nonene (DBN) [57,58,59] (Fig. 6.2b). Presence of “pipecolic acid” and “β-alanine” residues are some notable features in the sequences of efrapeptins, leucinostatins, and gichigamins [26, 28, 57, 58]. Fig. 6.2b shows a representative molecular structure of an efrapeptin.
Sequence Elucidation of Peptaibiotics
In the 1970s, Rinehart’s group began to elucidate the sequences of alamethicins, emerimicins, and antiamoebin by using gas chromatography-mass spectrometry (GC-MS), involving field desorption (FD) as the mode of ionization [12,13,14]. Subsequently, in the 1980s, fast atom bombardment (FAB)-MS was utilized to deduce the sequences of other peptaibols, e.g., zervamicins, trichotoxins, paracelsins, etc. [60, 61]. HPLC, particularly the reverse phase mode in conjunction with FD and FAB-MS, also was shown to be useful to identify various classes of microheterogeneous peptaibols [11]. From the late 1990s, electrospray ionization (ESI)-MS began to be applied for identifying novel peptaibol molecules and also to characterize the extent of microheterogeneity in their sequences [15, 18, 19, 21, 33, 59, 62, 63]. NMR spectroscopic methods are also applied to verify the sequences as well as to determine the 3D conformation [16, 30, 32, 35]. The stereochemical configurations of the constituting amino acids are determined by using Marfey’s method [16, 17, 30, 32, 35].
Biological and Other Functions
As aforementioned (vide supra), several peptaibols can form ion channels by inserting into the cellular membranes and aid in the translocation of cations across the membrane [36, 38]. Based on the size of the peptaibols, different models were put forward in order to understand the mechanism of formation of ion channels within the cell membrane. “Barrel stave model” has been suggested for longer peptaibols (~17–20 amino acid residues) such as alamethicin, whereas “Carpet model” was proposed for short peptaibol sequences, e.g., trichogin A IV [52]. Further, peptaibols have also been demonstrated to be “uncouplers” of mitochondrial oxidative phosphorylation [64, 65]. Besides their insect toxicity, the efrapeptins (peptaibiotics) from the entomopathogenic Tolypocladium niveum strongly inhibit mitochondrial (F1) ATPase and photophosphorylation in chloroplasts [57, 66]. The peptaibols, e.g., trichocellins A-II and B-II, have been shown to induce catecholamine secretion from bovine adrenal chromaffin cells through Ca2+ influx, in addition to their ability to form voltage-dependent ion channels in bilayer lipid membranes [67]. Interestingly, a peptaibol from a Trichoderma sp. has been found to inhibit the formation of amyloid β-peptide in cultured neuron cells of primary guinea pig cerebral cortex, at sub-μM concentrations (IC50 = 0.1 μg/ml), and it was not observed to be cytotoxic as well for concentrations <3 μg/ml [68].
Peptaibols possessing antiviral activity too have been reported. Rowley et al. discovered a few lipopeptaibols called “halovirs,” which were potent inhibitors of herpex simplex viruses (HSV) 1 and 2 in vitro, at low and sub-micromolar concentrations [55]. The halovirs were found to be more active than the free fatty acids, thereby showing the importance of the hexapeptide portion for the potent virucidal activity. Studies on peptaibols’ activity against mammalian cells are relatively less. In this regard, a 22-mer peptaibol, gichigamin A, and some of its related analogs were investigated on a pancreatic cancer cell line (MIA PaCa-2) in vitro, and it was observed that these peptaibols exhibited a wide range of antiproliferative and cytotoxic potencies [28]. Motivated from the outcomes of these in vitro experiments, gichigamin A was chosen for in vivo experiments, which showed that this peptaibol had significant and potent anti-tumor activity in a MIA PaCa-2 xenograft mouse model. Moderate in vitro cytotoxic activities against a panel of human cancer cell lines (HL-60, LS180, MDA-MB-231, and A549) were observed with two lipovelutibols that were isolated from the Himalayan cold habitat fungus Trichoderma velutinum [17]. Further, a 14-residue peptaibol from the same Himalayan psychrotrophic Trichoderma velutinum showed cytotoxic activity against the cancer cell lines, HL-60 and MDA-MB-231, with IC50 values 4 and 7 μM, respectively [16]. In the HL-60 cells, the same 14-residue peptaibol showed apoptosis in a dose-dependent manner [16]. Recently, a few peptaibols identified from a sponge-derived Acremonium sp. have also been observed to exhibit cytotoxic activity against A549 and/or HepG2 cancer cell lines [29]. Additionally, efrapeptin J (peptaibiotic) from a marine Tolypocladium sp. has been shown to be a downregulator of a molecular chaperone GRP78, which is implicated in anti-tumor activity, whereby these experiments were done in HT1080 cells and in MKN-74 human gastric cancer cells [69].
Interestingly, peptaibols have been shown to be useful in the microbial fuel cells (MFC) also. Because of their antibacterial activity, peptaibols (neoatroviridins A–D from Trichoderma viride) have been shown to aid in improving the power generation of MFCs by inhibiting methanogenic bacteria [70].
2.2 Cyclic Peptides (NRPs)
About 290 fungal cyclic peptides have been reported in the literature, which has been reviewed by Wang et al. (2017) [9], wherein they have classified these cyclic peptides according to their size, viz., based on the number of amino acid residues constituting the cyclic peptide. Starting from cyclic tripeptides (composed of three residues), Wang et al. (2017) [9] have covered cyclic peptides of sizes up to 18 amino acid residues. Many are head-to-tail (viz., fully backbone) cyclized peptides; however, considerable number of “backbone-sidechain cyclized peptides” have also been identified. In the ensuing discussions, salient features of certain classes of cyclic peptides, especially those possessing some rare amino acid residues, are described.
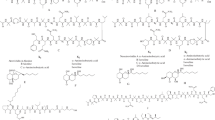
Cyclic peptides have been found from the soil fungi, marine fungi [10], entomopathogenic fungi, as well as endophytic fungi [9]. To a larger extent, fungal cyclic peptides have been identified from the genera Aspergillus, Penicillium, Fusarium, and Acremonium [9]. Cyclic peptides have been reported from psychrophilic and psychrotolerant fungi as well. In this regard, a very interesting cyclic peptide called “psychrophilin A” is of worth to be mentioned, which was identified from psychrotolerant Penicillium ribeum [71]. Psychrophilin A is the first natural cyclic peptide reported to contain a nitro group (instead of amino group). Another notable feature of psychrophilin A is, the presence of anthranilic acid (ATA), whose amino and carboxylic acid groups are part of the cyclic backbone structure, whereby the carboxylic group of the anthranilic acid forms a peptide bond with the nitrogen of the tryptophan sidechain (see Fig. 6.3a). Thus, instead of an amino group, a nitro group is actually linked to the α-carbon of the tryptophan residue. The amino group of the anthranilic acid forms a peptide bond with carboxyl group of proline. Fig. 6.3a shows this unusual molecular structure of psychrophilin A. Two more cyclic peptides, cycloaspeptides A and D, also were identified from the same psychrotolerant Penicillium ribeum [71]. These two peptides also contain ATA as part of the cyclic backbone structure.
Additionally, the cycloaspeptides A and D possess N-methylated residues: N-methyl phenylalanine and N-methyl tyrosine. Another psychrotolerant species, Penicillium algidum, was found to produce “psychrophilin D” [72], which has leucine residue instead of proline of psychrophilin A. The psychrophilin D was found to exhibit moderate activity in the P388 murine leukemia cell assay, while the cycloaspeptides A and D showed moderate activity against Plasmodium falciparum [72]. Further, four new cyclic peptides that were very closely related to psychrophilin class of peptides (as discussed above) were discovered from marine-derived fungus Aspergillus versicolor ZLN-60 [73]. From this same species of marine Aspergillus, Peng et al. (2014) also identified a hexapeptide, versicotide C that had two ATA residues and two N-methylated alanine residues. Two new cyclic peptides containing γ-Aminobutyric acid (GABA), unguisins A (Fig. 6.3b) and B, were identified from the marine-derived Emericella unguis by Malmstrom (1999), and this was the first study that reported about the occurrence of GABA as part of the ring structure of the cyclic heptapeptides [74]. An intriguing aspect is that five residues of these cycloheptapeptides have D-stereochemical configuration.
Subsequently, unguisins C and D were also reported from the same marine Emericella sp., whereby the peptide unguisin D was obtained through precursor-directed biosynthesis by adding L-leucine in the fermentation medium, as a result of which leucine got incorporated in the sequence of unguisin D, by replacing one of the valines of unguisin B [75]. Further, unguisin F was reported from an endophytic fungus Mucor irregularis, which was isolated from a medicinal plant Moringa stenopetala [76]. Aib containing cyclic heptapeptides, scytalidamides A and B, have also been reported from a marine fungus Scytalidium, wherein the scytalidamide B possessed 3-methyl proline residue, which is a rare modification [77]. Both these peptides showed in vitro cytotoxicity towards human colon carcinoma tumor cell line HCT-116 and NCI-60 cell line panel, with the IC50 values in the range 2–11 μM [77].
Sclerotides A and B from a marine-derived halotolerant Aspergillus sclerotiorum PT06-1 are interesting examples of cyclic hexapeptides in that they possess an unusual amino acid, dehydrotryptophan (Δ-Trp) [78]. Sclerotides A and B also contain ATA (Fig. 6.3c). Another notable aspect is that the sclerotide A and sclerotide B were photointerconvertible, whereby this inter-conversion occurs at the double bond of the Δ-Trp. And recently, three novel sclerotides C–E have been reported from soft coral-derived Aspergillus sclerotiorum SCSIO 41031 [79]. The same marine halotolerant Aspergillus sclerotiorum PT06-1 produced another class of cyclic peptides, which contained “ornithine” and N-methylated residues, in a nutrient-rich medium [80]. These were cyclic tripeptides named as “sclerotiotides,” wherein the δ-amino group of ornithine was involved in the formation of backbone amide (peptide) bond with the next amino acid residue. However, the α-amino group of the ornithine residue is not free; rather, it is bonded to a fatty acid by means of amide linkage, and this fatty acid contained unsaturated hydrocarbon chain. This hydrocarbon chain also contained aldehyde or hydroxyl functional group as substituent. Thus, ornithine residue-containing sclerotiotides are 12-membered ring structures. Furthermore, another sclerotiotide having lysine in the place of ornithine was also reported. Recently, from an Antarctica-sponge-derived Aspergillus insulicola HDN151418, three new sclerotiotides M–O (cyclic tripeptides) were discovered, whereby the variations in these three peptides were due to the nature of functional groups or substituents on the unsaturated hydrocarbon chain which is linked to the α-amino group of the ornithine residue [81]. Thus, sclerotiotides are examples of backbone-sidechain cyclized peptides.
As already mentioned, cyclic peptides have been discovered from entomopathogenic fungi as well. For instance, hirsutide, a cyclic tetrapeptide of sequence, cyclo-(L-NMe-Phe-L-Phe-L-NMe-Phe-L-Val), was reported from a spider-derived entomopathogenic fungus, Hirsutella sp. in New Zealand [82]. Three new cyclic tetrapeptides whose sequences were somewhat similar to the hirsutide’s sequence were identified from a fungus, Onychocola sclerotica, which was isolated from a poultry farm soil in Indonesia [83]. These three cyclic tetrapeptides were noted to block the cardiac calcium channels (Cav1.2), but not the hERG potassium channel [83]. Novel cyclic tetrapeptides, pseudoxylallemycins A–F, were discovered from a termite-associated fungus, Pseudoxylaria sp. X802, which was on the Microtermes sp. colony collected in South Africa [84]. The pseudoxylallemycins B–D possessed the unusual allenyl modification on the aromatic moiety.
Epichlicin, a cyclic octapeptide containing a β-amino acid residue, was isolated from an endophytic fungus Epichloe typhina of timothy plant (Phleum pretense L.) [85]. Epichlicin is rather a lipocyclic peptide as it possesses long hydrocarbon sidechain of the β-amino acid residue, viz., 3-amino tetradecanoic acid. Epichlicin was found to inhibit the spore germination of Cladosporium phlei, which is a pathogenic fungus of timothy plant. Recently, Ekanayake et al. had reported the identification of three cyclic octapeptides, broomeanamides from the fungus Sphaerostilbella broomeana (TFC201724) collected from the foothills of Himalayas in India [86]. Examples of a few fungal cyclic peptides possessing unusual amino acids are listed in Table 6.1.
Cyclosporin A is a very popular fungal cyclic peptide since it has been used as an immunosuppressive drug. Cyclosporin A has been identified in various fungal genera and species, e.g., Tolypocladium sp., Fusarium sp., Aspergillus sp., Trichoderma sp., and Beauveria nivea [9]. Several variants/analogs of cyclosporins have also been identified from different fungi. In addition to the immunosuppressant activity, cyclosporins also exhibit anti-inflammatory, anti-fungal, and anti-parasitic activities. Cyclosporin A and other analogues are composed of 11 amino acid residues, among which some are N-methylated residues (see Fig. 6.4).
2.3 Cyclic Depsipeptides
Cyclic depsipeptides (CDPs) are another class of NRPs that contain backbone ester (lactone) bond(s), due to hydroxy acid residue(s) amidst the conventional amide (peptide) units. Besides the hydroxy acid residue, many CDPs also contain N-methylated amino acid residues. About 350 different CDPs have been reported mainly from the genera Acremonium, Aspergillus, Beauveria, Fusarium, Metarhizium, Alternaria, etc. [88]. CDPs of different sizes have been discovered from various fungal species, albeit cyclic hexadepsipeptides (a hydroxy acid and five amino acid residues) represent the largest class of fungal CDPs that have been identified thus far [88]. CDPs may be classified into two categories based on the nature of hydroxy acid (HA): (i) α-HA-containing CDPs (α-HA CDPs) and (ii) β-HA-containing CDPs (β-HA CDPs). Molecular structural properties of some α-HA CDPs and a few β-HA CDPS are delineated herein.
α-HA CDPs
Sansalvamide, beauvenniatins, allobeauvericins, beauvericins, enniatins, destruxins, roseotoxins, isaridins, etc. are some examples of CDPs that have α-HA in their sequences. The cyclohexadepsipeptides beauvenniatins, allobeauvericins, beauvericins, and enniatins are interesting in that they have three ester and three amide bonds which are arranged in an alternating manner in their cyclic backbone structure (see Fig. 6.5a) [88]. Another intriguing aspect is that these four classes of CDPs have been identified from different fungal genera. Beauvenniatins have been identified from Acremonium sp., many beauvericins can be found in Beauveria bassiana, allobeauvericins were reported from Paecilomyces tenuipes, and enniatins have been discovered predominantly from Fusarium sp. [88]. A few beauvericins have been reported from Paecilomyces tenuipes also [88]. Enniatins have been found in endophytic as well as from entomogenous Fusarium sp. Likewise, though destruxins, roseotoxins, and isaridins have been identified from different fungal genera, these three classes may be grouped together based on the similarities in their molecular structures, wherein these peptides contain a β-amino acid residue (β-alanine) as part of their cyclic backbone, in addition to an α-HA (see Fig. 6.5b) [20, 89,90,91]. Destruxins are produced by the entomopathogenic Metarhizium anisopliae [89], roseotoxins are produced by Trichothecium roseum [90], while isaridins can be found in Isaria sp. [20, 91]. Thus, it is interesting to note that structurally very similar CDPs can be found from different fungal genera or species. In other words, fungi of different genera or species produce structurally similar CDPs. Perhaps, similar biosynthetic pathways involving NRPSs might be operating in different fungal genera.
The cyclohexadepsipeptides hirsutatins from the insect pathogenic Hirsutella nivea BCC 2594 and trichodepsipeptides from the filamentous fungus Trichothecium sp. can also be grouped in one class, although they are found in two different genera, since these peptides have identical cyclic backbone structure [92, 93]. Both hirsutatins and trichodepsipeptides contain two α-HA residues that are bonded next to each other and, hence, have two adjacent ester bonds in their cyclic backbone, albeit there are minor variations in the sidechains of one or two residues between hirsutatins and trichodepsipeptides. The leualacins reported from the fungus Hapsidospora irregularis are cyclic pentadepsipeptides, contain two α-HA residues (α-hydroxy leucine) that are not adjacent in its sequence, and have a β-amino acid residue, β-alanine [94].
The cyclic octadepsipeptide, bassionolide, and the “series of PF1022” CDPs have identical cyclic backbone architecture, which is characterized by equal number of ester and amide bonds occurring in an alternating fashion. The bassionolide was identified from Beauveria bassiana and Verticillium lecanii [95]. It is composed of four hydroxy valine and four N-methyl leucine residues, and hence, the sequence of bassionolide can be written as cyclo-[(N-Met Leu – Hyd. Val)4]. The series of “PF1022” CDPs were identified in Mycelia sterilia, and all the peptides in this series possess four N-methyl leucine residues, whereas the α-HA residues in this series can be hydroxy alanine (Hyd. Ala) or hydroxy phenylalanine (Hyd. Phe) or hydroxy tyrosine (Hyd. Tyr) [96]. The PF1022F, cyclo-[(N-Met Leu – Hyd. Ala)4] was also identified in Trichoderma asperellum, which was an endophyte of a traditional Chinese medicinal plant [97]. Thus, the four N-Met leucine residues are conserved across the sequences of bassionolide and all the CDPs of PF1022 series.
β-HA CDPs
Beauveriolides, oryzamides, isariins, and emericellamides possess β-HA residue, which has somewhat long hydrocarbon sidechain on the β-carbon (with respect to the carboxyl group). Hence, these CDPs are also called cyclic lipopeptides. Beauverolides are tetra CDPs, while oryzamides, isariins, and emericellamides are cyclic hexadepsipeptides. Beauverolides are mainly present in Beauveria sp., especially in B. bassiana [98,99,100]. Oryzamides were identified from the sponge-derived fungus Nigrospora oryzae PF18 [101]. The CDP isariin was first reported by Vining and Taber way back in 1962 from Isaria cretacea [102]. Thereafter, Baute et al. (1981) and Deffieux et al. (1981) identified three variants of isariin (isariins B, C, and D) from Isaria felina [103, 104]. Based on LC-ESI-MS/MS and NMR spectroscopy, six more variants of isariin were identified, wherein these variants differed only at one particular amino acid residue that was bonded to the hydroxyl group of the β-HA [20]. Interestingly, a novel isariin (iso-isariin B) and a known isaridin (isaridin E) were discovered in Beauveria felina also [105]. Emericellamides were reported from Aspergillus [106] as well as from marine-derived Emericella sp. [107]. Once again it is notable that peptides having very similar molecular structures and identical cyclic backbone architecture are produced by fungi belonging to different genera, as evident in the cases of oryzamides, isariins, and emericellamides.
Halobacillin, a cyclic octadepsipeptide, was first reported from a marine Bacillus. However, it was also discovered in an endophytic fungus Trichoderma asperellum that was residing in a traditional Chinese medicinal plant [97]. Halobacillin consists of a β-HA residue having a hydrocarbon sidechain (C12H25) on the β-carbon, and hence, it is also an example of cyclic lipopeptide. Examples of a few fungal cyclic depsipeptides are listed in Table 6.2.
3 Ribosomal Fungal Peptides
3.1 Disulfide-Bonded Peptides: Fungal Defensins
Disulfide-bonded peptides are also a type of cyclic peptides, whereby the formation of cyclic or ring-like structure involves the sidechain atoms rather than the backbone atoms. In other words, disulfide-bonded peptides come under the class of sidechain-sidechain cyclized peptides. Defensins are disulfide-bonded (cysteine-rich) peptides that have been identified in vertebrates including humans, in plants, as well as in insects [108,109,110,111,112]. These polypeptides constitute an important class of antimicrobial peptides (AMPs), which has a vital role as part of the innate immune system, and thus contribute to the host defense against bacterial, fungal, and viral infections [109]. When compared to the total number of defensins known from vertebrates, plants, and insects, the number of defensins or defensin-like peptides (fDLPs) identified from fungi is relatively less [113]. Nevertheless, fDLPs are touted to be of therapeutic value particularly in the context of rise in the drug-resistant strains, e.g., a derivative of a fDLP, NZ2114, could reach the preclinical stage of drug discovery [114].
Plectasin is the first fungal defensin isolated from a saprophytic ascomycete, Pseudoplectania nigrella [115]. The in vitro activity of this peptide against Streptococcus pneumoniae was comparable to that of penicillin and vancomycin. The recombinantly produced plectasin was found to be active even against the antibiotic-resistant strains of S. pneumoniae. It exhibits antimicrobial activity by directly binding to the bacterial cell wall precursor lipid II, and the stoichiometry of this binding was observed to be equimolar [116]. Plectasin is composed of 40 amino acid residues and contains 3 disulfide bonds (UniProt KB ID: Q53I06). The sequence of plectasin showed 50–55% identity with sequences of several invertebrate defensins, whereas it did not have any significant similarity to the mammalian α- and β-defensin sequences [115]. The 3D molecular structure of plectasin contains an α-helix and two antiparallel β-strands stabilized by three disulfide bonds. By considering the sequence together with the 3D molecular structural features, it was evident that plectasin indeed belonged to the defensin family [115]. A derivative of plectasin, NZ2114 was found to be potent not only against the multiply drug-resistant bacterial strains in vitro, but it also was efficacious in the in vivo pharmacodynamic investigation carried out in a murine infection model [2]. Furthermore, NZ2114 could also exhibit strong bactericidal activity in the cerebrospinal fluid (CSF) by penetrating into the CSF in an experimental meningitis model, indicating its potential to treat infections in the central nervous system (CNS), including the penicillin-resistant pneumococcal meningitis [117].
Eurocin is another new fungal defensin isolated from another ascomycete Eurotium amstelodami [118]. It is a 42-amino acid residue-long polypeptide with three disulfide bonds. Its 3D molecular structure is made up of an α-helix and two β-strands forming an antiparallel β-sheet (c.f. PDB ID 2LT8), referred to as cysteine-stabilized αβ-fold (CSαβ) [119, 120]. Thus, the eurocin’s 3D structure was found to be highly homologous to the structures reported from other fungal and invertebrate defensins. Similar to plectasin, eurocin shows more strong effect on Gram-positive bacteria than Gram-negative bacteria. From the cell-free assay experiments, it was observed that eurocin also binds to the bacterial cell wall precursor lipid II, suggesting that both eurocin and plectasin follow a similar binding mechanism. In vivo and in vitro antimicrobial assays showed that eurocin is a fast and effective antibiotic against Streptococci even at low concentrations.
Copsin is a novel antibiotic identified in the mushroom Coprinopsis cinerea, and perhaps, it is the first defensin to be identified from the fungal phylum of Basidiomycota [121]. It is composed of 57 amino acid residues with N-terminal pyroglutamic acid and 6 disulfide bonds. The sequence of copsin showed 20–27% identity with the defensins from other invertebrates, fungi, and plants. The 3D molecular structure of eurocin also consists of CSαβ fold, similar to the structures of eurocin and plectasin (vide supra). The sequence of the secondary structural elements displayed high identity to the known defensins that possessed CSαβ structural motif; however, the length and the composition of the loop regions and the termini were found to be unique. Copsin inhibited Gram-positive bacteria such as Bacillus subtilis, Listeria sp., and Enterococcus sp. in the low μg/ml range, and it showed most potent activity against Listeria monocytogenes, which is a well-known food-borne pathogen. Copsin did not elicit antibacterial activity, when treated with reducing agent (e.g., dithiothreitol, DTT), indicating the importance of disulfide bonds for the integrity of its 3D structure and its biological function.
The defensin micasin identified from a dermatophytic fungus Microsporum canis was found to act against both Gram-positive and Gram-negative bacteria suggesting that the micasin’s mode of antimicrobial action is different from eurocin and plectasin [122]. Micasin could also kill two clinical isolates of methicillin-resistant Staphylococcus aureus and the opportunistic pathogen Pseudomonas aeruginosa at low micromolar concentrations. Very recently, a non-hemolytic fDLP, Pyronesin 4 (Py4), was reported from the basal filamentous ascomycete Pyronema confluens [113]. Py4 inhibited the bacterial cell wall biosynthesis, and it was found to be highly stable in the mammalian serum.
A brief survey into UniProt KB database looking for “fungal defensins” revealed a few more peptides. Though only a few fungal defensins have been isolated and characterized, six different families of fDLPs have been identified from the available fungal genome sequences through computational prediction, whereby the computational analysis indicated very good conservation of three different types of defensins between animals and fungi [119].
3.2 Fungal RiPPs
α-Amanitin, phallacidin, ustiloxin, phomopsin, and epichloëcyclin are a few examples of fungal RiPPs [5]. Hydroxylation, methylation, epimerization, and acetylation are some posttranslational modifications that have been observed on fungal RiPPs [3]. As reviewed by Vogt and Kunzler, the fungal RiPPs identified thus far have been classified into four different families: amatoxins/phallotoxins, borosins, dikaritins, and epichloëcyclins. α-Amanitin and phallacidin represent the first fungal RiPP family, identified from the poisonous mushrooms belonging to the genus Amanita [123]. α-Amanitin (amatoxin) is a bicyclic octapeptide, while the related phallacidin (phallotoxin) is a bicyclic heptapeptide [123]. Amatoxins inhibit RNA polymerase II, and amatoxin poisoning can affect the liver, leading to death as well; however, phallotoxins elicit toxicity by stabilizing F-actin [123]. The phallotoxins exhibit toxicity only through parenteral administration and not orally due to poor absorption [123].
The family dikaritins constitutes ustiloxins, phomposins, and asperipins. These three classes of peptides were identified from ascomycetes. A striking feature in the molecular structures of dikaritins is the presence of “ether” linkage as part of the cyclic/ring structure involving the sidechain of tyrosine [5]. Ustiloxins and phomposins inhibit tubulin polymerization and suppress mitosis, whereas the biological activity of asperipin is not yet known [5]. The borosin family consists of cyclic peptides called “omphalotins” that have N-methylation at several amide bonds [124]. Prior to the identification of omphalotins, the N-methylation was mostly thought to be prevalent in non-ribosomal peptides only. Omphalotins E–I from the basidiomycete Omphalotus olearius have been reported to have nematicidal activity [124]. The endophytic fungus belonging to the genus Epichloë synthesizes cyclic nonapeptides called epichloëcyclins, whose biological function is not reported yet [125].
4 Fungal Peptides as Drugs
Caspofungin (brand name: “Cancidas”) is a semi-synthetic lipopeptide, which is the first member in the new drug class called “echinocandins,” as named by Merck & Co., Inc. (https://go.drugbank.com/drugs/DB00520). It was originally obtained as a fermentation product from the fungus Glarea lozoyensis [126]. Many clinical trials proved “caspofungin” to be efficacious for the treatment of esophageal, oropharyngeal, and invasive candidiasis as well as for invasive aspergillosis [127, 128]. It was approved by the Food and Drug Administration (FDA) for use in the United States [127, 129]. A few other members of echinocandin class are (i) micafungin (brand name: “Micamine”; Drug Bank Accession No. DB01141) and (ii) anidulafungin (brand name: “Ecalta/Eraxis”; Drug Bank Accession No. DB00362) [129]. Thus, caspofungin, micafungin, and anidulafungin became the “first-line” antifungal agents to treat invasive candidiasis. Resistance towards the echinocandin drugs also has been observed [130]. However, it was recently reported that caspofungin elicits antifungal activity even against the multidrug-resistant (MDR) Candida, when the drug was prepared in low ionic solutions [131]. The molecular structures of these three lipopeptides have cyclic peptidic backbone comprising six amide (peptide) bonds, and hence, these three are cyclic hexapeptides [9].
The cyclic undecapeptide cyclosporin A (CsA) is popularly known for its immunosuppressive activity [132]. Due to its selective immunosuppressant activities, CsA could be successfully applied in many transplantation therapies, and thereby significant improvement was observed in the survival rates after the grafting of the solid organs [132]. CsA in combination with other suitable drug has proven to be good for treating rheumatoid arthritis as well [133, 134]. Although CsA was identified from various fungal sources such as Aspergillus sp., Beauveria nivea, Fusarium oxysporum, and Trichoderma polysporum, the fungus Tolypocladium inflatum has been extensively utilized for the production of CsA by following different fermentation techniques [9, 132]. The other biological functions of CsA include antifungal, antiparasitic, and anti-inflammatory [132]. About 25 different analogues of cyclosporin have been identified, which have also been observed to have immunosuppressant effect as well as antifungal activity [9]. Cyclosporin is sold under different brand names, e.g., Sandimmune, Cequa, Gengraf, etc. (https://go.drugbank.com/drugs/DB00091).
5 Concluding Remarks
Because of the continuous emergence of resistant strains of pathogens towards various drug compounds, the search for novel antibiotics and antimicrobial compounds has not stopped for several years from now. Particularly the peptide-based antibiotics and antimicrobial peptides have been shown to elicit potent activity even against drug-resistant microbial strains [135, 136]. Therefore, exploration of peptide-based drugs and antimicrobials have become more preferable. The discussions in the previous sections clearly indicate that fungi are indeed very valuable (re)sources for exploring and identifying novel peptide antibiotics. Decades of research investigations on various fungal sources have unearthed numerous non-ribosomal peptides leading to creation of a repertoire of knowledge on these peptides. Therefore, the non-ribosomal fungal peptides have been tried and tested a lot for developing therapeutic compounds, for example, the echinocandin and the cyclosporin classes of drugs (see Sect. 6.4) [137]. Identification of cytotoxic fungal peptaibols and their synthetic analogues, e.g., gichigamins from Tolypocladium species in Michigan and lipopeptaibols from a Himalayan Trichoderma species, provides encouraging signs for development of novel anti-cancer or anti-tumor drugs [16, 17, 28]. Despite this level of progress, several studies are still ongoing, and many investigations need to be carried out in future, in order to obtain improved and clearer understanding about their natural role(s) of non-ribosomal fungal peptides [138]. In contrast, RiPPs from fungi have been reported only in the recent past. Therefore, there is enormous scope to investigate the potential of fungal RiPPs, so as to develop them into drugs of therapeutic value and make them widely available in the market. The discovery of fungal RiPPs might stimulate the interest towards “fungal sources” for searching peptide-based therapeutics [5]. Altogether, in addition to their natural multiple vital roles in various contexts in the ecosystem, fungi are indeed valuable resources for drug discovery and for various agricultural applications. Thus, the rise of fungal-based products having diverse applications towards general human welfare seems imminent, in the near future.
References
Niu X, Thaochan N, Hu Q (2020) Diversity of linear non-ribosomal peptide in biocontrol fungi. J Fungi (Basel) 6(2):61. https://doi.org/10.3390/jof6020061
Andes D, Craig W et al (2009) In vivo pharmacodynamic characterization of a novel plectasin antibiotic, NZ2114, in a murine infection model. Antimicrob Agents Chemother 53(7):3003–3009. https://doi.org/10.1128/AAC.01584-08
Arnison PG, Bibb MJ et al (2013) Ribosomally synthesized and post-translationally modified peptide natural products: overview and recommendations for a universal nomenclature. Nat Prod Rep 30(1):108–160. https://doi.org/10.1039/c2np20085f
Dang T, Süssmuth RD (2017) Bioactive peptide natural products as lead structures for medicinal use. Acc Chem Res 50(7):1566–1576. https://doi.org/10.1021/acs.accounts.7b00159
Vogt E, Künzler M (2019) Discovery of novel fungal RiPP biosynthetic pathways and their application for the development of peptide therapeutics. Appl Microbiol Biotechnol 103(14):5567–5581. https://doi.org/10.1007/s00253-019-09893-x
Kleinkauf H, Von Döhren H (1996) A nonribosomal system of peptide biosynthesis. Eur J Biochem 236:335–351. https://doi.org/10.1111/j.1432-1033.1996.00335.x
Marahiel MA, Stachelhaus T, Mootz HD (1997) Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem Rev 97:2651–2673. https://doi.org/10.1021/cr960029e
Alonzo DA, Schmeing TM (2020) Biosynthesis of depsipeptides, or Depsi: the peptides with varied generations. Protein Sci 29(12):2316–2347. https://doi.org/10.1002/pro.3979
Wang X, Lin M et al (2017) Structural diversity and biological activities of fungal cyclic peptides, excluding cyclodipeptides. Molecules 22(12):2069. https://doi.org/10.3390/molecules22122069
Youssef FS, Ashour ML et al (2019) A comprehensive review of bioactive peptides from marine fungi and their biological significance. Mar Drugs 17(10):559. https://doi.org/10.3390/md17100559
Brückner H, Przybylski M (1984) Isolation and structural characterization of polypeptide-antibiotics of the peptaibol class by high-performance liquid chromatography with field desorption and fast atom bombardment mass spectrometry. J Chromatogr 296:263–275
Pandey RC, Cook JC Jr, Rinehart KL Jr (1977) High resolution and field desorption mass spectrometry studies and revised structures of alamethicins I and II. J Am Chem Soc 99:8469–8483. https://doi.org/10.1021/ja00468a016
Pandey RC, Cook JC Jr, Rinehart KL Jr (1977) Reptaibophol antibiotics. 2. Structures of the peptide antibiotics emerimicins III and IV. J Am Chem Soc 99:5205–5206. https://doi.org/10.1021/ja00457a064
Pandey RC, Meng H et al (1977) Structure of antiamoebin I from high resolution field desorption and gas chromatographic mass spectrometry studies. J Am Chem Soc 99:5203–5205. https://doi.org/10.1021/ja00457a063
Ruiz N, Wielgosz-Collin G et al (2007) New Trichobrachins, 11-residue peptaibols from a marine strain of Trichoderma longibrachiatum. Peptides 28:1351–1358. https://doi.org/10.1016/j.peptides.2007.05.012
Singh VP, Pathania AS et al (2020) 14-residue peptaibol velutibol A from Trichoderma velutinum: its structural and cytotoxic evaluation. RSC Adv 10:31233–31242. https://doi.org/10.1039/d0ra05780k
Singh VP, Yedukondalu N et al (2018) Lipovelutibols A–D: cytotoxic lipopeptaibols from the Himalayan cold habitat fungus Trichoderma velutinum. J Nat Prod 81:219–226. https://doi.org/10.1021/acs.jnatprod.6b00873
Jaworski A, Brückner H (1999) Detection of new sequences of peptaibol antibiotics trichotoxins A-40 by on-line liquid chromatography-electrospray ionization mass spectrometry. J Chromatogr A 862:179–189. https://doi.org/10.1016/S0021-9673(99)00931-0
Poirier L, Amiard JC et al (2007) Determination of peptaibol trace amounts in marine sediments by liquid chromatography/electrospray ionization-ion trap-mass spectrometry. J Chromatogr A 1160:106–113. https://doi.org/10.1016/j.chroma.2007.04.006
Sabareesh V, Ranganayaki RS et al (2007) Identification and characterization of a library of microheterogenous cyclohexadepsipeptides from the fungus Isaria. J Nat Prod 70(5):715–729
Stoppacher N, Reithner B et al (2007) Profiling of trichorzianines in culture samples of Trichoderma atroviride by liquid chromatography/tandem mass spectrometry. Rapid Commun Mass Spectrom 21:3963–3970
Van Bohemen AI, Zalouk-Vergnoux A et al (2016) Development and validation of LC-MS methods for peptaibol quantification in fungal extracts according to their lengths. J Chromatogr B Analyt Technol Biomed Life Sci 1009-1010:25–33. https://doi.org/10.1016/j.jchromb.2015.11.039
Toniolo C, Bruckner H (2007) Peptaibiotics. Chem Biodivers 4:1021–1022. https://doi.org/10.1002/cbdv.200790093
Krause C, Kirschbaum J, Brückner H (2006) Peptaibiomics: an advanced, rapid and selective analysis of peptaibiotics/peptaibols by SPE/LC-ES-MS. Amino Acids 30:435–443. https://doi.org/10.1007/s00726-005-0275-9
Daniel JF, Filho ER (2007) Peptaibols of trichoderma. Nat Prod Rep 24(5):1128–1141. https://doi.org/10.1039/b618086h
Degenkolb T, Berg A et al (2003) The occurrence of peptaibols and structurally related peptaibiotics in fungi and their mass spectrometric identification via diagnostic fragment ions. J Pept Sci 9:666–678. https://doi.org/10.1002/psc.497
Raap J, Erkelens K et al (2005) Fungal biosynthesis of non-ribosomal peptide antibiotics and α, α-dialkylated amino acid constituents. J Pept Sci 11:331–338. https://doi.org/10.1002/psc.621
Du L, Risinger AL et al (2017) Unique amalgamation of primary and secondary structural elements transform peptaibols into potent bioactive cell-penetrating peptides. Proc Natl Acad Sci U S A 114:E8957–E8966. https://doi.org/10.1073/PNAS.1707565114
Hao X, Li S et al (2021) Acremopeptaibols A-F, 16-residue peptaibols from the sponge-derived Acremonium sp. IMB18-086 cultivated with heat-killed Pseudomonas aeruginosa. J Nat Prod 84(11):2990–3000. https://doi.org/10.1021/acs.jnatprod.1c00834
Jiao WH, Khalil Z et al (2018) Trichodermides A-E: new peptaibols isolated from the Australian termite nest-derived fungus Trichoderma virens CMB-TN16. J Nat Prod 81(4):976–984. https://doi.org/10.1021/acs.jnatprod.7b01072
Stoppacher N, Neumann NKN et al (2013) The comprehensive peptaibiotics database. Chem Biodivers 10:734–743. https://doi.org/10.1002/cbdv.201200427
Zhang SH, Yang J et al (2021) Longibramides A-E, peptaibols isolated from a mushroom derived fungus Trichoderma longibrachiatum Rifai DMG-3-1-1. Chem Biodivers 18(5):e2100128. https://doi.org/10.1002/cbdv.202100128
Mohamed-Benkada M, Montagu M et al (2006) New short peptaibols from a marine Trichoderma strain. Rapid Commun Mass Spectrom 20:1176–1180. https://doi.org/10.1002/rcm.2430
Mohamed-Benkada M, François Pouchus Y et al (2016) Identification and biological activities of long-chain peptaibols produced by a marine-derived strain of Trichoderma longibrachiatum. Chem Biodivers 13(5):521–530. https://doi.org/10.1002/cbdv.201500159
van Bohemen AI, Ruiz N et al (2021) Pentadecaibins I-V: 15-residue peptaibols produced by a marine-derived Trichoderma sp. of the Harzianum clade. J Nat Prod 84(4):1271–1282. https://doi.org/10.1021/acs.jnatprod.0c01355
Balaram P, Krishna K et al (1992) The properties of ion channels formed by zervamicins. Eur Biophys J 21:117–128. https://doi.org/10.1007/BF00185426
Nagaraj R, Balaram P (1981) Alamethicin, a Transmembrane Channel. Acc Chem Res 14:356–362. https://doi.org/10.1021/ar00071a005
Sansom MSP (1993) Structure and function of channel-forming peptaibols. Q Rev Biophys 26:365–421
Mueller P, Rudin DO (1968) Action potentials induced in biomolecular lipid membranes. Nature 217:713–719. https://doi.org/10.1038/217713a0
Fox RO, Richards FM (1982) A voltage-gated ion channel model inferred from the crystal structure of alamethicin at 1.5-Å resolution. Nature 300:325–330. https://doi.org/10.1038/300325a0
Auvin-Guette C, Rebuffat S et al (1992) Trichogin A IV, an 11-residue lipopeptaibol from Trichoderma longibrachiatum. J Am Chem Soc 114:2170–2174. https://doi.org/10.1021/ja00032a035
Gurunath R, Balaram P (1995) A nonhelical, multiple beta-turn conformation in a glycine-rich heptapeptide fragment of Trichogin A IV containing a single central alpha-aminoisobutyric acid residue. Biopolymers 35:21–29. https://doi.org/10.1002/bip.360350104
Iqbal M, Balaram P (1981) Membrane channel forming polypeptides. 270-MHz Hydrogen-1 nuclear magnetic resonance studies on the conformation of the 11–21 fragment of suzukacillin. Biochemistry 20:4866–4871. https://doi.org/10.1021/bi00520a010
Iqbal M, Balaram P (1981) The 310 helical conformation of the amino terminal decapeptide of suzukacillin. 270MHz 1H NMR evidence for eight intramolecular hydrogen bonds. J Am Chem Soc 103:5548–5552. https://doi.org/10.1021/ja00408a045
Karle IL, Flippen-Anderson J et al (1987) Conformation of a 16-residue zervamicin IIA analog peptide containing three different structural features: 3(10)-helix, alpha-helix, and beta-bend ribbon. Proc Natl Acad Sci U S A 84:5087–5091. https://doi.org/10.1073/pnas.84.15.5087
Karle IL, Flippen-Anderson JL et al (1991) Crystal structure of [Leu1] zervamicin, a membrane ion-channel peptide: implications for gating mechanisms. Proc Natl Acad Sci U S A 88:5307–5311. https://doi.org/10.1073/pnas.88.12.5307
Karle IL, Perozzo MA et al (1998) Crystal structure of the channel-forming polypeptide antiamoebin in a membrane-mimetic environment. Proc Natl Acad Sci U S A 95:5501–5504. https://doi.org/10.1073/pnas.95.10.5501
Karle IL, Sukumar M, Balaram P (1986) Parallel packing of alpha-helices in crystals of the zervamicin IIA analog Boc-Trp-Ile-Ala-Aib-Ile-Val-Aib-Leu-Aib-Pro-OMe.2H2O. Proc Natl Acad Sci U S A 83:9284–9288. https://doi.org/10.1073/pnas.83.24.9284
Rebuffat S, Prigent Y et al (1991) Tricholongins BI and BII, 19-residue peptaibols from Trichoderma longibrachiatum. Solution structure from two-dimensional NMR spectroscopy. Eur J Biochem 201:661–674
Snook CF, Woolley GA et al (1998) The structure and function of antiamoebin I, a proline-rich membrane-active polypeptide. Structure 6:783–792. https://doi.org/10.1016/S0969-2126(98)00079-3
Aravinda S, Shamala N, Balaram P (2008) Aib residues in peptaibiotics and synthetic sequences: analysis of nonhelical conformations. Chem Biodivers 5(7):1238–1262. https://doi.org/10.1002/cbdv.200890112
Chugh JK, Wallace BA (2001) Peptaibols: models for ion channels. Biochem Soc Trans 29(4):565–570. https://doi.org/10.1042/bst0290565
Whitmore L, Wallace BA (2004) Analysis of peptaibol sequence composition: implications for in vivo synthesis and channel formation. Eur Biophys J 33(3):233–237. https://doi.org/10.1007/s00249-003-0348-1
Auvin-Guette C, Rebuffat S et al (1993) Structural elucidation of Trikoningins KA and KB, peptaibols from Trichoderma koningii. J Chem Soc Perkin Trans 1:249–255
Rowley DC, Kelly S et al (2003) Halovirs A-E, new antiviral agents from a marine-derived fungus of the genus Scytalidium. Bioorg Med Chem 11:4263–4274. https://doi.org/10.1016/S0968-0896(03)00395-X
Momose I, Onodera T et al (2019) Leucinostatin Y: a peptaibiotic produced by the entomoparasitic fungus Purpureocillium lilacinum 40-H-28. J Nat Prod 82:1120–1127. https://doi.org/10.1021/acs.jnatprod.8b00839
Gupta S, Krasnoff SB et al (1991) Structures of the efrapeptins: potent inhibitors of mitochondrial ATPase from the fungus Tolypocladium niveum. J Am Chem Soc 113:707–709. https://doi.org/10.1021/ja00002a068
Gupta S, Krasnoff SB et al (1992) Structure of efrapeptins from the fungus Tolypocladium niveum: peptide inhibitors of mitochondrial ATPase. J Org Chem 57:2306–2313. https://doi.org/10.1021/jo00034a022
Uma MV, Sudha R, Balaram P (2001) Spermidine as a potential biosynthetic precursor to the 1,5-diazabicyclo[4:3:0]nonene residue in the efrapeptins. J Pept Res 58:375–379. https://doi.org/10.1034/j.1399-3011.2001.00915.x
Przybylski M, Dietrich I et al (1984) Elucidation of structure and microheterogeneity of the polypeptide antibiotics paracelsin and trichotoxin A-50 by fast atom bombardment mass spectrometry in combination with selective in situ hydrolysis. Biomed Mass Spectrom 11:569–582
Rinehart KL Jr, Gaudioso LA et al (1981) Structures of eleven zervamicin and two emerimicin peptide antibiotics studied by fast atom bombardment mass spectrometry. J Am Chem Soc 103:6517–6520
Pócsfalvi G, Ritieni A et al (1997) Microheterogeneity characterization of a Paracelsin mixture from Trichoderma reesei using high-energy collision-induced dissociation tandem mass spectrometry. Rapid Commun Mass Spectrom 11:922–930
Sabareesh V, Balaram P (2006) Tandem electrospray mass spectrometric studies of proton and sodium ion adducts of neutral peptides with modified N- and C-termini: synthetic model peptides and microheterogeneous peptaibol antibiotics. Rapid Commun Mass Spectrom 20:618–628. https://doi.org/10.1002/rcm.2349
Das MK, Raghothama S, Balaram P (1986) Membrane channel forming polypeptides. Molecular conformation and mitochondrial uncoupling activity of antiamoebin, an alpha-aminoisobutyric acid containing peptide. Biochemistry 25:7110–7117. https://doi.org/10.1021/bi00370a053
Mathew MK, Nagaraj R, Balaram P (1981) Alamethicin and synthetic peptide fragments as uncouplers of mitochondrial oxidative phosphorylation. Effect of chain length and charge. Biochem Biophys Res Commun 98:548–555. https://doi.org/10.1016/0006-291x(81)90875-5
Cross RL, Kohlbrenner WE (1978) The mode of inhibition of oxidative phosphorylation by efrapeptin (A23871). Evidence for an alternating site mechanism for ATP synthesis. J Biol Chem 253(14):4865–4873
Wada S, Iida A et al (1997) Role of the Gln/Glu residues of trichocellins A-II/B-II in ion-channel formation in lipid membranes and catecholamine secretion from chromaffin cells. Biochim Biophys Acta 1325(2):209–214. https://doi.org/10.1016/s0005-2736(96)00260-x
Hosotani N, Kumagai K et al (2007) SPF-5506-A4, a new peptaibol inhibitor of amyloid beta-peptide formation produced by Trichoderma sp. J Antibiot (Tokyo) 60(3):184–190. https://doi.org/10.1038/ja.2007.20
Hayakawa Y, Hattori Y et al (2008) Efrapeptin J, a new down-regulator of the molecular chaperone GRP78 from a marine Tolypocladium sp. J Antibiot (Tokyo) 61(6):365–371. https://doi.org/10.1038/ja.2008.51
Ghosh R, Noori MT, Ghangrekar MM (2017) Novel application of peptaibiotics derived from Trichoderma sp. for methanogenic suppression and enhanced power generation in microbial fuel cells. RSC Adv 7:10707–10717. https://doi.org/10.1039/C6RA27763B
Dalsgaard PW, Larsen TO et al (2004) Psychrophilin A and cycloaspeptide D, novel cyclic peptides from the psychrotolerant fungus Penicillium ribeum. J Nat Prod 67:878–881. https://doi.org/10.1021/np0303714
Dalsgaard PW, Larsen TO, Christophersen C (2005) Bioactive cyclic peptides from the psychrotolerant fungus Penicillium algidum. J Antibiot (Tokyo) 58(2):141–144. https://doi.org/10.1038/ja.2005.16
Peng J, Gao H et al (2014) Psychrophilins E-H and versicotide C, cyclic peptides from the marine-derived fungus Aspergillus versicolor ZLN-60. J Nat Prod 77(10):2218–2223. https://doi.org/10.1021/np500469b
Malmstrøm J (1999) Unguisins A and B: new cyclic peptides from the marine-derived fungus Emericella unguis. J Nat Prod 62(5):787–789. https://doi.org/10.1021/np980539z
Malmstrøm J, Ryager A et al (2002) Unguisin C, a GABA-containing cyclic peptide from the fungus Emericella unguis. Phytochemistry 60(8):869–872. https://doi.org/10.1016/s0031-9422(02)00150-4
Akone SH, Daletos G et al (2016) Unguisin F, a new cyclic peptide from the endophytic fungus Mucor irregularis. Z Naturforsch C J Biosci 71(1–2):15–19. https://doi.org/10.1515/znc-2015-0137
Tan LT, Cheng XC et al (2003) Scytalidamides A and B, new cytotoxic cyclic heptapeptides from a marine fungus of the genus Scytalidium. J Org Chem 68(23):8767–8773. https://doi.org/10.1021/jo030191z
Zheng J, Zhu H et al (2009) Novel cyclic hexapeptides from marine-derived fungus, Aspergillus sclerotiorum PT06-1. Org Lett 11(22):5262–5265. https://doi.org/10.1021/ol902197z
Long J, Chen Y et al (2021) Cyclic peptides from the soft coral-derived fungus Aspergillus sclerotiorum SCSIO 41031. Mar Drugs 19(12):701. https://doi.org/10.3390/md19120701
Zheng J, Xu Z et al (2010) Cyclic tripeptides from the halotolerant fungus Aspergillus sclerotiorum PT06-1. J Nat Prod 73(6):1133–1137. https://doi.org/10.1021/np100198h
Sun C, Zhang Z et al (2020) Antibacterial cyclic tripeptides from Antarctica-sponge-derived fungus Aspergillus insulicola HDN151418. Mar Drugs 18(11):532. https://doi.org/10.3390/md18110532
Lang G, Blunt JW, Cummings NJ, Cole AL, Munro MH (2005) Hirsutide, a cyclic tetrapeptide from a spider-derived entomopathogenic fungus, Hirsutella sp. J Nat Prod 68(8):1303–1305. https://doi.org/10.1021/np0501536
Pérez-Victoria I, Martín J et al (2012) Isolation and structural elucidation of cyclic tetrapeptides from Onychocola sclerotica. J Nat Prod 75(6):1210–1214. https://doi.org/10.1021/np3000987
Guo H, Kreuzenbeck NB et al (2016) Pseudoxylallemycins A-F, cyclic tetrapeptides with rare Allenyl modifications isolated from Pseudoxylaria sp. X802: a competitor of fungus-growing termite cultivars. Org Lett 18(14):3338–3341. https://doi.org/10.1021/acs.orglett.6b01437
Seto Y, Takahashi K et al (2007) Novel cyclic peptide, epichlicin, from the endophytic fungus, Epichloe typhina. Biosci Biotechnol Biochem 71(6):1470–1475. https://doi.org/10.1271/bbb.60700
Ekanayake DI, Perlatti B et al (2021) Broomeanamides: cyclic octapeptides from an isolate of the Fungicolous ascomycete Sphaerostilbella broomeana from India. J Nat Prod 84(7):2028–2034. https://doi.org/10.1021/acs.jnatprod.1c00414
Dalsgaard PW, Blunt JW et al (2004) Psychrophilin B and C: cyclic nitropeptides from the psychrotolerant fungus Penicillium rivulum. J Nat Prod 67(11):1950–1952. https://doi.org/10.1021/np0497954
Wang X, Gong X et al (2018) Structural diversity and biological activities of cyclic depsipeptides from fungi. Molecules 23(1):169. https://doi.org/10.3390/molecules23010169
Jegorov A, Havlícek V, Sedmera P (1998) Rapid screening of destruxins by liquid chromatography/mass spectrometry. J Mass Spectrom 33(3):274–280. https://doi.org/10.1002/(SICI)1096-9888(199803)33:3<274::AID-JMS630>3.0.CO;2-R
Jegorov A, Paizs B et al (2003) Profiling of cyclic hexadepsipeptides roseotoxins synthesized in vitro and in vivo: a combined tandem mass spectrometry and quantum chemical study. Eur J Mass Spectrom (Chichester) 9(2):105–116. https://doi.org/10.1255/ejms.531
Ravindra G, Ranganayaki RS et al (2004) Two novel hexadepsipeptides containing several modified amino acid residues from the fungus Isaria. Chem Biodivers 1:489–504
Isaka M, Palasarn S et al (2005) Cyclohexadepsipeptides from the insect pathogenic fungus Hirsutella nivea BCC 2594. J Nat Prod 68(11):1680–1682. https://doi.org/10.1021/np050246n
Sy-Cordero AA, Graf TN et al (2011) Cyclodepsipeptides, sesquiterpenoids, and other cytotoxic metabolites from the filamentous fungus Trichothecium sp. (MSX 51320). J Nat Prod 74(10):2137–2142. https://doi.org/10.1021/np2004243
Zhang S, Qiu Y et al (2017) Identification of cyclic depsipeptides and their dedicated synthetase from Hapsidospora irregularis. J Nat Prod 80(2):363–370. https://doi.org/10.1021/acs.jnatprod.6b00808
Kanaoka M, Isogai A et al (1978) Bassianolide, a new insecticidal cyclodepsipeptide from Beauveria bassiana and Verticillium lecanii. Agric Biol Chem 42:629–635. https://doi.org/10.1080/00021369.1978.10863029
Ohyama M, Okada Y et al (2011) Structure-activity relationship of anthelmintic cyclooctadepsipeptides. Biosci Biotechnol Biochem 75(7):1354–1363. https://doi.org/10.1271/bbb.110129
Ding G, Chen AJ et al (2012) Sesquiterpenes and cyclopeptides from the endophytic fungus Trichoderma asperellum SAMUELS, LIECKF & NIRENBERG. Chem Biodivers 9(6):1205–1212. https://doi.org/10.1002/cbdv.201100185
Elsworth JF, Grove JF (1977) Cyclodepsipeptides from Beauveria bassiana. Part 1. Beauverolides H and I. J Chem Soc Perkin Trans 13:270–273
Elsworth JF, Grove JF (1980) Cyclodepsipeptides from Beauveria bassiana. Part 2. Beauverolides A to F and their relationship to isarolide. J Chem Soc Perkin Trans 1 8:1795–1799
Grove JF (1980) Cyclodepsipeptides from Beauveria bassiana. Part 3. The isolation of beauverolides Ba, Ca, Ja, and Ka. J Chem Soc Perkin Trans 1(12):2878–2880
Ding LJ, Yuan W et al (2016) Oryzamides A-E, Cyclodepsipeptides from the sponge-derived fungus Nigrospora oryzae PF18. J Nat Prod 79(8):2045–2052. https://doi.org/10.1021/acs.jnatprod.6b00349
Vining LC, Taber WA (1962) Isariin, A new depsipeptide from Isaria cretacea. Can J Chem 40:1579–1584. https://doi.org/10.1139/v62-239
Baute R, Deffieux G et al (1981) New insecticidal cyclodepsipeptides from the fungus Isaria felina. I. Production, isolation and insecticidal properties of isariins B, C and D. J Antibiot (Tokyo) 34(10):1261–1265. https://doi.org/10.7164/antibiotics.34.1261
Deffieux G, Merlet D et al (1981) New insecticidal cyclodepsipeptides from the fungus Isaria felina. II. Structure elucidation of isariins B, C and D. J Antibiot (Tokyo) 34(10):1266–1270. https://doi.org/10.7164/antibiotics.34.1266
Langenfeld A, Blond A et al (2011) Insecticidal cyclodepsipeptides from Beauveria felina. J Nat Prod 74(4):825–830. https://doi.org/10.1021/np100890n
Chiang YM, Szewczyk E et al (2008) Molecular genetic mining of the Aspergillus secondary metabolome: discovery of the emericellamide biosynthetic pathway. Chem Biol 15(6):527–532. https://doi.org/10.1016/j.chembiol.2008.05.010
Oh DC, Kauffman CA et al (2007) Induced production of emericellamides A and B from the marine-derived fungus Emericella sp. in competing co-culture. J Nat Prod 70(4):515–520. https://doi.org/10.1021/np060381f
Kovaleva V, Bukhteeva I et al (2020) Plant defensins from a structural perspective. Int J Mol Sci 21(15):5307. https://doi.org/10.3390/ijms21155307
Lehrer RI (2004) Primate defensins. Nat Rev Microbiol 2(9):727–738. https://doi.org/10.1038/nrmicro976
Sathoff AE, Samac DA (2019) Antibacterial activity of plant defensins. Mol Plant-Microbe Interact 32(5):507–514. https://doi.org/10.1094/MPMI-08-18-0229-CR
Schneider JJ, Unholzer A et al (2005) Human defensins. J Mol Med (Berl) 83(8):587–595. https://doi.org/10.1007/s00109-005-0657-1
Koehbach J (2017) Structure-activity relationships of insect defensins. Front Chem 5:45. https://doi.org/10.3389/fchem.2017.00045
Qi S, Gao B, Zhu S (2022) A fungal defensin inhibiting bacterial cell-wall biosynthesis with non-hemolysis and serum stability. J Fungi (Basel) 8(2):174. https://doi.org/10.3390/jof8020174
Wu J, Gao B, Zhu S (2014) The fungal defensin family enlarged. Pharmaceuticals (Basel) 7(8):866–880. https://doi.org/10.3390/ph7080866
Mygind PH, Fischer RL et al (2005) Plectasin is a peptide antibiotic with therapeutic potential from a saprophytic fungus. Nature 437(7061):975–980. https://doi.org/10.1038/nature04051
Schneider T, Kruse T et al (2010) Plectasin, a fungal defensin, targets the bacterial cell wall precursor lipid II. Science 328(5982):1168–1172. https://doi.org/10.1126/science.1185723
Ostergaard C, Sandvang D et al (2009) High cerebrospinal fluid (CSF) penetration and potent bactericidal activity in CSF of NZ2114, a novel plectasin variant, during experimental pneumococcal meningitis. Antimicrob Agents Chemother 53(4):1581–1585. https://doi.org/10.1128/AAC.01202-08
Oeemig JS, Lynggaard C et al (2012) Eurocin, a new fungal defensin: structure, lipid binding, and its mode of action. J Biol Chem 287(50):42361–42372. https://doi.org/10.1074/jbc.M112.382028
Zhu S (2008) Discovery of six families of fungal defensin-like peptides provides insights into origin and evolution of the CSαβ defensins. Mol Immunol 45(3):828–838. https://doi.org/10.1016/j.molimm.2007.06.354
Zhu S, Gao B, Tytgat J (2005) Phylogenetic distribution, functional epitopes and evolution of the CSαβ superfamily. Cell Mol Life Sci 62(19–20):2257–2269. https://doi.org/10.1007/s00018-005-5200-6
Essig A, Hofmann D et al (2014) Copsin, a novel peptide-based fungal antibiotic interfering with the peptidoglycan synthesis. J Biol Chem 289(50):34953–34964. https://doi.org/10.1074/jbc.M114.599878
Zhu S, Gao B et al (2012) Dermatophytic defensin with antiinfective potential. Proc Natl Acad Sci U S A 109(22):8495–8500. https://doi.org/10.1073/pnas.1201263109
Hallen HE, Luo H et al (2007) Gene family encoding the major toxins of lethal Amanita mushrooms. Proc Natl Acad Sci U S A 104(48):19097–19101. https://doi.org/10.1073/pnas.0707340104
Liermann JC, Opatz T et al (2009) Omphalotins E-I, five oxidatively modified nematicidal cyclopeptides from Omphalotus olearius. Eur J Org Chem:1256–1262. https://doi.org/10.1002/ejoc.200801068
Johnson RD, Lane GA et al (2015) A novel family of cyclic oligopeptides derived from ribosomal peptide synthesis of an in planta-induced gene, gigA, in Epichloë endophytes of grasses. Fungal Genet Biol 85:14–24. https://doi.org/10.1016/j.fgb.2015.10.005
Letscher-Bru V, Herbrecht R (2003) Caspofungin: the first representative of a new antifungal class. J Antimicrob Chemother 51(3):513–521. https://doi.org/10.1093/jac/dkg117
Johnson MD, Perfect JR (2003) Caspofungin: first approved agent in a new class of antifungals. Expert Opin Pharmacother 4(5):807–823. https://doi.org/10.1517/14656566.4.5.807
Stone EA, Fung HB, Kirschenbaum HL (2002) Caspofungin: an echinocandin antifungal agent. Clin Ther 24(3):351–377. https://doi.org/10.1016/s0149-2918(02)85039-1
Zhang H, Chen S (2022) Cyclic peptide drugs approved in the last two decades (2001-2021). RSC Chem Biol 3:18–31. https://doi.org/10.1039/d1cb00154j
Spampinato C, Leonardi D (2013) Candida infections, causes, targets, and resistance mechanisms: traditional and alternative antifungal agents. Biomed Res Int 2013:204237. https://doi.org/10.1155/2013/204237
Sumiyoshi M, Miyazaki T et al (2020) Novel and potent antimicrobial effects of caspofungin on drug-resistant Candida and bacteria. Sci Rep 10(1):17745. https://doi.org/10.1038/s41598-020-74749-8
Survase SA, Kagliwal LD et al (2011) Cyclosporin A—a review on fermentative production, downstream processing and pharmacological applications. Biotechnol Adv 29(4):418–435. https://doi.org/10.1016/j.biotechadv.2011.03.004
Tugwell P, Pincus T et al (1995) Combination therapy with cyclosporine and methotrexate in severe rheumatoid arthritis. The methotrexate-cyclosporine combination study group. N Engl J Med 333(3):137–141. https://doi.org/10.1056/NEJM199507203330301
Yocum DE (1996) Combination therapy with cyclosporin in rheumatoid arthritis. Rheumatology 35(suppl 2):19–23. https://doi.org/10.1093/rheumatology/35.suppl_2.19
Galgóczy L, Yap A, Marx F (2019) Cysteine-rich antifungal proteins from filamentous fungi are promising bioactive natural compounds in anti-Candida therapy. Isr J Chem 59:360–370. https://doi.org/10.1002/ijch.201800168
Magana M, Pushpanathan M et al (2020) The value of antimicrobial peptides in the age of resistance. Lancet Infect Dis 20(9):e216–e230. https://doi.org/10.1016/S1473-3099(20)30327-3
Bills G, Li Y et al (2014) New insights into the echinocandins and other fungal non-ribosomal peptides and peptaibiotics. Nat Prod Rep 31(10):1348–1375. https://doi.org/10.1039/c4np00046c
Oide S, Turgeon BG (2020) Natural roles of nonribosomal peptide metabolites in fungi. Mycoscience 61(3):101–110., ISSN 1340-3540. https://doi.org/10.1016/j.myc.2020.03.001
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Sabareesh, V., Gowri, V.S. (2023). A Bird’s-Eye View of Fungal Peptides. In: Satyanarayana, T., Deshmukh, S.K. (eds) Fungi and Fungal Products in Human Welfare and Biotechnology. Springer, Singapore. https://doi.org/10.1007/978-981-19-8853-0_6
Download citation
DOI: https://doi.org/10.1007/978-981-19-8853-0_6
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-8852-3
Online ISBN: 978-981-19-8853-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)