Abstract
Avian influenza virus (AIV) belongs to the genus Influenza A virus of the family Orthomyxoviridae. The virus can infect a variety of avian species, but the low pathogenic AIVs do not usually cause explicit symptoms in poultry. In contrast, the highly pathogenic avian influenza (HPAI) viruses continue to cause outbreaks among poultry, wild birds and occasionally humans in Asia, the Middle East, North America, and Africa. Environmental factors associated with cross-species transmission have been substantially reviewed before. However, acquiring the knowledge of a number of environmental factors with spatial structures, which usually are not randomly distributed, for timely implementation of control measures rely on accurate identification of the spatial clustering in a global or local scale. In this article, we review different approaches in identifying spatial or temporal-spatial clustering in avian influenza outbreaks. In the future perspective, we propose to develop intuitive tools for timely identify the dynamic changes of clustering and viral spreading. Such tools will assist in not just the identifying the environmental factors associated with the clustering or spreading direction, but also timely control measures to prevent further damage.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Avian influenza virus (AIV)
- Cluster analysis
- Hotspot analysis
- Scan statistics
- Space–time permutation model
- Knox test
- Standard deviational ellipse (SDE) method
- Regression modeling
1 Introduction
Avian influenza, caused by influenza A virus (IAV), is a zoonotic influenza that affects a wide variety of birds, poultry and occasionally humans. Influenza A virus is the only species of the genus Alpha-influenza virus of the family Orthomyxoviridae. The structure of influenza A virus consists of a lipid envelope and a negative-sense single-stranded ribonucleic acid (RNA) genome with eight segments [1]. Influenza A viruses can be classified into subtypes based on the combination of the spike hemagglutinin (HA) attachment protein and the neuraminidase (NA) protein. To date, 18 HA subtypes (H1 to H18) and 11 NA subtypes (N1 to N11) have been identified [2], while only 131 subtypes have been detected in nature [3]. Subtypes of IAV can be further divided into clades and subclades based on the similarity of HA genes [4, 5], and subtypes can also be subdivided into genotypes based on the combination of internal gene segments. The nomenclature of influenza viruses has been standardized, and the name of a new strain consists of a combination of antigen type, original host, geographic origin, strain name, year of isolation, and subtype (HxNy) [6].
The genome segments of IAV encode different viral proteins. The structural proteins express in the envelope containing the surface proteins, which are HA attachment proteins and NA proteins, and the membrane ion channel (M2) proteins. Internal proteins include the nuclear protein (NP), matrix protein (M1), and the polymerase complex consisting of three subunits, namely polymerase basic protein 1 (PB1), polymerase basic protein 2 (PB2), and polymerase acidic protein (PA). Nonstructural protein 1 (NS1) and nonstructural protein 2 (NS2), the nuclear export protein (NEP), are encoded by segment 8. AIVs use host proteases to cleave the HA0 molecule into HA1 and HA2 subunits, which are essential for the uncoating step of viral replication. AIVs can be defined as low pathogenic avian influenza (LPAI) viruses and highly pathogenic avian influenza (HPAI) viruses based on their virulence in chickens. Thus, if mutations result in the insertion of multiple lysine and arginine residues into the HA0 cleavage site of the virus, termed the multilocus cleavage site, which can be recognized by the ubiquitous and extensive proteases in host tissues, it becomes an HPAI virus. As a corollary, HPAI viruses may replicate throughout the host, systematically destroying tissues, leading to multiple organ failure and ultimately to host death. However, LPAI viruses have only one arginine at the cleavage site, which can only be recognized by trypsin-like proteases. Therefore, replication of LPAI virus is restricted to the respiratory and gastrointestinal tracts, where expression of this protease occurs [7,8,9,10].
Although avian influenza viruses (AIVs) replicate in wild bird reservoirs, the viruses spread out through the saliva, mucus, and feces of infected birds. Spillover AIVs can be transmitted from infected wild birds to poultry, primarily through direct contact with wild birds or indirect contact through human activities and contaminated water or other media. Most AIVs cause gastrointestinal infections in chickens, while those with no or minimal clinical signs are LPAI viruses, whose distribution in wild birds varies by subtype depending on geographic location, bird abundance, and prevalence. HPAI viruses can affect poultry as well as wild birds. Infections in chickens and turkeys induce severe disease with mortality rates as high as 90–100%. So far, only the H5 and H7 subtypes of AIVs have been recorded as causing HPAI outbreaks in poultry, but most of the H5 and H7 subtypes are LPAI viruses. HPAI viruses evolve by mutation, amino acid substitution or recombination after long-term circulation and efficient replication of LPAI viruses in poultry [11] (Fig. 9.1).
The evolution and transmission routes of avian influenza virus between and among animal species and humans. Green arrows indicate the transmission routes of low pathogenic avian influenza (LPAI), red arrows indicate the transmission routes of high pathogenic avian influenza (HPAI) and blue arrows indicate the additional spread routes of avian influenza. Solid arrows represent frequent transmission events, and dashed arrows represent sporadic or limited transmission events. Once HPAI viruses become introduced into wild bird populations, the spread and maintenance of these viruses in wild birds will be determined by different factors involved the types of host birds, the viruses, and the ecology
2 Factors Associated with Zoonotic Transmission of Avian Influenza Virus
Outbreaks of HPAI were first described as “fowl plague” in the 1880s, and subsequent outbreaks in Europe from then onwards were caused exclusively by HPAI H7N7 and H7N1 viruses until the first confirmed outbreak of HPAI H5N1 occurred among chickens in Scotland in 1959 [12]. HPAI H5Nx (N1-9) and H7Nx (N1, N3, N4, N7-N9) viruses [13, 14] have been reported to cause thousands of outbreaks in domestic poultry and wild birds in more than 60 countries, killing numerous poultry through HPAI virus attacks or mass culling strategies and causing huge economic losses. During HPAI epidemics in poultry, viruses can spill back into wild birds, where they subsequently circulate asymptomatically or cause disease and death [15, 16], even generate reassortants with LPAI or wild bird-adapted strains [17,18,19]. These viruses can also spill over into mammals, including pigs, horses, whales, seals, and humans [20]. However, it is considered that AIVs do not replicate efficiently enough in humans to sustain human-to-human transmission [21] (Fig. 9.1).
Long-distance migratory birds played an influential role in the global spread of HPAI viruses [22,23,24,25,26], while wild birds may also be involved in local HPAI virus amplification and reassortment [27]. The HPAI found in wild birds was highly associated with the geographical locations of poultry farms [20, 28, 29]. However, such association has been not significant since the emergence in 2014 of the HPAI virus Gs/Gd clade 2.3.4.4 which has dominated in outbreaks in poultry and wild birds with abundant genetic reassortments resulting in H5N1, H5N2, H5N3, H5N4, H5N5, H5N6 and H5N8 subtypes [24, 30, 31].
The most predominant natural reservoirs of HPAI H5 viruses are Anseriformes, which are responsible for the maintenance, rapid transmission, and geographic expansion of these viruses. The other prominent reservoirs, Charadriiformes, are possible reasons for the rapid global spread of HPAI viruses due to their fast-moving, long-distance migration and highly gregarious during migration period [32,33,34]. The transmission rates of HPAI H5 viruses within Anseriformes and Galliformes are high, but transmission between these orders is limited [18, 35]. Understanding the mechanisms of HPAI virus transmission and maintenance in wild birds can provide a reference for surveillance strategies.
Human infection with AIVs is a rare and sporadic event, however, AIVs subtypes H5, H6, H7, H9, and H10, have been recorded infecting humans to cause clinical disease of varying severity. Exposure histories of human cases and phylogenetic analyses of AIVs isolated from wild birds, poultry, humans, and associated environments suggested that cross-species poultry-to-human transmission of AIVs frequently occurs on poultry farms. In addition, live bird markets are active sites for interspecies dissemination, where AIVs can be transmitted from birds to humans or reassort influenza gene segments in different host species [36, 37].
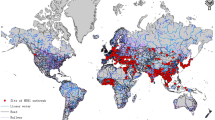
HPAI H5N1 outbreaks have occurred in a variety of ecological systems with economic, agricultural and environmental differences, which pose the threat to the poultry production sector. Factors affecting the spatial and temporal distribution of the outbreak of AIVs have been investigated in many studies previously. Marius Gilbert and Pfeiffer [38] summarized the risk factors considered for HPAI H5N1 presence in previous studies in nine categories, including(1) farming practice and local biosecurity, (2) poultry and livestock census data with longitude and latitude, (3) anthropogenic variables, (4) socio-economic variables, (5) variables indicative of the presence or abundance of wild birds, (6) variables indicative of the presence or abundance of rivers, lakes or wetlands, (7) eco-climatic variables obtained using weather station data or remote sensing, (8) land-use and cropping variables, and finally (9) topography. Among these factors, the density of domestic waterfowl, anthropogenic variables (human population density, distance to roads) and indicators of water presence were identified positively correlated with HPAI H5N1 presence across studies and regions.
3 Spatial Clustering Analysis of Avian Influenza Viruses Transmission
Commonly used statistical clustering approaches from the literatures to identify spatial distribution patterns and transmission mechanisms can provide additional information for AIV control and prevention strategies.
3.1 Cluster Analysis
The spatial distribution pattern of HPAI cases was clustered, dispersed, or randomly distributed, which can be measured by global spatial autocorrelation analysis, such as the global Moran's I statistics [39]. The null hypothesis of global Moran's I is spatial randomness. Global Moran's I index is the correlation coefficient between the eigenvalue and its surrounding values, which can be transformed into z-score and p-value to infer whether the overall spatial distribution has statistically significant clusters. A positive z-score with a statistically significant p-value indicates spatial clustering, and a negative z-score with a statistically significant p-value indicates spatial dispersion. Threshold spatial distances are analyzed by incremental spatial autocorrelation analysis for a series of increasing distances at intervals of interest over a spatial range, and spatial clustering is measured by the z-score of each distance interval. The z-score usually peaks at some distance where the spatial clustering is most salient within the specified spatial extent. The distance associated with the statistically significant peak is selected as the threshold spatial distance for a cluster.
3.2 Hotspot Analysis
However, these methods do not account for the location of clusters. The Local Indicator of Spatial Autocorrelation (LISA) with Local Moran's I statistics calculates the eigenvalues of each geographic boundary region and assesses the significance of the region's similarity to its surroundings to identify statistically significant local clusters, such as high-high hot spots and low-low cold spots, or high-low–high local spatial outliers. The high positive z-scores of the test demonstrate the statistically significant high-high cluster of hotspots [39]. Based on global and local Moran's I analyses, the distribution of H7N9 human cases in Zhejiang Province, China, showed statistically significant spatial autocorrelation in some epidemic waves and identified the statistically significantly high-high clusters and high-low outlier clusters mostly located in the northern part of this province [40]. Shan et al. [41] used global Moran's I analysis to identify that the distribution of H7N9 human cases in Mainland China showed statistically significant spatial autocorrelation during the five epidemic waves.
Liang et al. [42] evaluated environmental factors associated with clusters of outbreaks and multiple subtypes co-circulating of HPAI H5Nx viruses in Taiwan. Global Moran's I analysis was conducted to determine the grid size for covering Taiwan when measuring the clusters of H5Nx outbreak farms and found the optimal distance to be 3 km. Therefore, a 3 km square grid covering Taiwan was used for LISA and local Moran's I statistics, and the results indicated that the hotspots of H5Nx outbreak farms were located on the west coast of Taiwan from 2015 to 2017, where covered more than 75% of outbreaks farms in 2015 and 2017. Multivariate stepwise logistic regressions comparing hotspots and non-hotspots were developed to analyze four categories of variables: farm-related, farm biosecurity-related, wild bird-related, and anthropological. Notably, this study used satellite remote sensing methods to establish unregistered poultry farm data and merged it with the official poultry farm registration database to complete the poultry farm census dataset. A poultry heterogeneity index was also created in this study to describe the heterogeneity of the total number of domesticated waterfowls versus land fowl in each grid. Four risk factors consistently showed a strong association with the spatial clusters of HPAI H5N2 and H5N8 circulations during 2015 and 2017, including high poultry farm density, poultry heterogeneity index, non-registered waterfowl flock density, and a higher percentage of cropping land coverage. Using estimates from the regression models of 2015 and 2017, risk maps were generated to predict high-risk areas and further validated by using outbreaks from the first half of 2018. The results showed that the risk maps for 2015 and 2017 had a prediction rate higher than 55%.
Unlike the local Moran's I statistic, the Getis-Ord Gi* statistic calculates each feature in the context of neighboring features in the dataset to measure the degree of spatial clustering. Features in geographic boundary regions that are similar to adjacent features, and the sum of these features, including the feature itself, that differ significantly from the expected sum, will yield a statistically significant z-core and p-value result. The larger the statistically significant positive z-value, the stronger the aggregation of hot spots, and the smaller the statistically significant negative z-value, the stronger the aggregation of cold spots [43].
In order to investigate the locations of disease clusters, Shan et al. and Huang et al. [41, 44] used the Kernel density estimation to present the clustering areas of human cases caused by infection of AIVs in China in different epidemic waves. Kernel density estimation is a non-parametric statistical method to estimate the probability density function of a random variable. It converts point features into smoothly curved density surfaces by calculating the sum of a kernel function on each data point. The kernel function calculates the surface value of each point by weighting the distances of all points at each specific location in the distribution. The surface value of each point location is the highest, and it decreases with the distance from the point increases. The density of each point is the sum of all surface values of that point. If more points cluster in one location, the higher the density of that location, as the higher the probability of seeing a point at that location [45]. Based on Kernel density estimation, the Yangtze River Delta region and the Pearl River Delta region had the highest density and the intensity had gradually shifted during the epidemics [41, 46].
4 Temporal Spatial Clustering Identification of Avian Influenza Viruses Transmission
Approaches used to early and accurately characterize epidemiologic patterns of disease incidence in a temporal and spatial series are becoming increasingly important. Statistical analysis for detecting spatial–temporal clusters of health-related events is often used for epidemiological and biomedical studies. Timely identification of anomalies of disease or poisoning incidence during ongoing surveillance or an outbreak requires the use of sensitive statistical methods that recognize an incidence pattern at the time of occurrence. The following sections reviewed analytical methods commonly used to study temporal-spatial patterns.
4.1 Scan Statistics or Space–Time Permutation Model
Cluster analysis, such as scan statistics, are generally designed for retrospective detection of epidemiologic anomalies in a temporal or space–time series. Spatial scan statistics is a widely-used approach to detect spatiotemporal clustering although several conventional cluster analysis methods such as gap-statistic or K-means have been developed. The scan statistic employs a moving window, possibly with varied shapes, of predetermined radius or geographical unit with fixed population and finds the maximum number of cases revealed through the window as it slides over the entire region [47,48,49]. The scan test is structured to detect the largest cluster of incidences. The maximum number of events occurring in a window is the test statistic for the scan test. However, calibrating proper spatial and temporal windows in scan statistics is difficult, which requires a process of model tuning. Huang et al. and Dong et al. [44, 46] used the space–time permutation model to analyze the spatial–temporal clustering of H7N9 human cases. Assuming that the population changes are homogeneous, and the spatial extent of the cluster does not change during the scanning process, it only needs the spatial location and time data of the cases. Scan statistics use a varied-size cylindrical moving window with space as the base and time as the height to scan the target area in the time period of interest. Observed and expected numbers of cases were obtained from the scan of each location and size of the window, and the likelihood ratio or relative ratio statistics were used to evaluate whether there is a cluster in the cylinder. In the space–time permutation model, the spatial and temporal data of the case being studied are used to adjust multiple tests through thousands of random permutations. The cluster with the largest log-likelihood ratio is simulated for each of these permutations of the data set, and the P value for hypothesis testing is used Monte Carlo simulations [50].
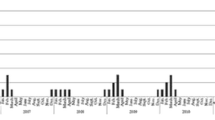
According to space–time permutation model scan statistics, the epidemic of H7N9 human cases from 2013 to 2017 showed six statistically significant clusters. In 2017, there were four clusters, with centers located in Beijing, Hubei, Sichuan and Shanghai. One cluster in Xinjiang from July to December 2014, and one cluster in Guangdong from July 2013 to March 2015 [44]. Further analysis of the first two epidemics in 2013–2014 with 5 days as the time unit, in the first and second epidemic waves, two and three statistically significant clusters were identified. In the first wave, the most likely cluster of epidemics was observed in the southeast region centered on Fujian Province from April 27 to May 11, 2013, and the second cluster of epidemics occurred in Jiangsu province and Shanghai from March 13 to April 11, 2013. In the second wave, the earlier cluster of epidemics was in Yangtze River Delta from January 12 to January 31, 2014. The second cluster of epidemics was in Pearl River delta from February 16 to March 2, 2014, and the third cluster of epidemics was in six provinces centered on Anhui Province from April 22 to May 31, 2014.
Zhang et al. [51] also analyze space–time clustering of human infection with H7N9 virus in county level in 2013–2014. The peak z-score indicates that there are obvious spatial clusters at the distance of 30 and 250 km in the incremental spatial autocorrelation analysis, and the distinct temporal clustering at the duration of 14 to 26 days in the temporal autocorrelation analysis. Based on this, 250 km and 14 days are selected as the “Threshold” of distance in space and time for the next space–time hotspot analysis. Getis-Ord Gi* z-score illustrated that there were two statistically significant space–time clustering near Shanghai and Zhejiang in March 26 to April 18, 2013, and near Guangzhou and Shenzhen from February 3 to 4, 2014. Zhang et al. also used a space–time permutation scan statistic model to investigate epidemic pattern of these human cases. The results showed that there were six statistically significant spatiotemporal clusters from 2013 to 2014. The cluster near Shanghai and Zhejiang from March 13, to April 9, 2013, and the cluster near Guangzhou and Shenzhen from February 5 to 25, 2014, were similar to the results of hotspot analysis, indicating the good consistency between these two methods.
4.2 Knox Test
Knox proposed a method that allows for statistical testing of the interaction of incidents of infectious disease in space and time that does not use an arbitrary critical value of distance or time for determining local clusters [52]. The Knox statistic is calculated by pairing all possible data points (e.g., location in space and time of the death of birds) within a clearly defined geographic area and temporal interval and testing them against assigned values of what is “close” in space and time. The number of close space–time data pairs is compared with what would be expected if there were no space–time cluster. Based on Knox settings, Barton and David [53] proposed a “intersection” approach” to obtain spatial–temporal clustering. They suggested to connect the pairs with temporal clustering by line segments to form a temporal map, and then connect the pairs with spatial clustering to form a spatial map. Combining these two maps produces a spatial–temporal clustering [54]. This is reasonable but hard to implement because in a highly dense incidence region, thousands of lines tangled together will make the discrimination among clusters difficult. In addition to the “intersection approach”, Openshaw et al. [55] considered a “geographical analysis machine” (GAM) method which draw r-radius circles for the areas with dense incidence when “r” is permitted to varied (say, r = 1, 2, or 4 km). Those corresponding dense circles visually formed bunches of circles, and is decided to be spatially clustered. See also Turnbull [56] for more discussion. In this study published in Scientific Report (2021), Wu et al. [57] showed that Knox-based approach can still display spatiotemporal clusters, in particular when the outbreaks occur in multiple places. When circling the major spatial clusters, each circle has a “diameter” within 3 km, which is the size of the control zone established once HPAI-infected farm identified in Taiwan. When an infected premises (IP) is reported, all poultry from that particular IP will be culled and all farms within 3 km radius of that infected premises will be targeted for intensive surveillance. Therefore, outside the 3 km control zone stands for the spreading of HPAI viruses requiring epidemiological investigation.
4.3 Standard Deviational Ellipse (SDE) Method
The standard deviational ellipse (SDE) method was a widely applied approach to displaying geographic distribution of occurrence of some events [58,59,60,61], including chronic diseases and infectious diseases, etc. [62,63,64]. It combines the concern of location, (two-dimensional) dispersion, and orientation (meaning direction plus shape) in a simple optimization calculation. When SDE is used repeatedly over a specified time period (say, every week during the emergent outbreak period), the mean area center is the origin of these two axes [60]. It suggests that one, two, and three standard deviation ellipses will cover approximately 68, 95, and 99% of the points [65]. The orientation of the long axis indicates the direction of the point distribution, therefore, the greater the difference between the long and short axis, the more obvious the direction trend. Connecting the centers of each ellipse offers a clue for disease transmission. This connecting line can be compared with long axes of consecutive ellipses. To sketching the spatial trends of H7N9 human cases over time in China during 2013–2017, SDE analysis was used by Huang et al. and Dong et al. [44, 46]. They analyzed the distribution of cases for each month in the epidemic wave. SDE analysis showed that the first wave of the 2013 epidemic started in the three Yangtze River Delta provinces and spread from Jiangsu to Guangdong. The second wave occurred in the southeastern coastal provinces from Jiangsu Province to Guangdong Province, and expanded the epidemic area with a coastal orientation, but spread toward the inland in the last two months. The third wave of the epidemic started from the southeast coastal area to Gansu Province, and then gradually narrowed down to the Yangtze River Delta. The fourth wave of the epidemic occurred in the southeastern coastal region and then spread to the northern coastal region. In the fifth wave, the epidemic occurred along the eastern coast and then gradually spread to most of mainland China.
4.4 Regression Modeling
The interpretation of the shape of SDE need to be cautious as it might be area-specific. While the connection between ellipses reveals a different story implying the development among sub-areas with dense emergent cases, it shows a temporary geographic pattern or latent mode of spreading of events. Using SDE method to estimate the transmission direction needs mild correction when the concerned infections have become endemic; i.e., the virus tends to be localized and existed there all year round. An alternative approach to estimate the direction of spreading is a regression model proposed in Zinszer et al. [66], hereafter called Zinszer model, which attempted to estimate local transmission directions for Ebola epidemic. It states that for an outbreak event occurred at calendar time Ti and at location (\({\mathrm{X}}_{\mathrm{i}}\), \({\mathrm{Y}}_{\mathrm{i}}\)) with corresponding explanatory variable (possibly a vector) Zi, and Ti+1 is the time of the next (Ebola) outbreak case so that the inter-outbreak “gap time” τi = Ti+1 − Ti can be modeled as:
The parameters \({\upbeta }_{1}\) and \({\upbeta }_{2}\) interpret the inverse of rate of transmission in the direction of X and Y, respectively, usually adopted as the longitude (X) and latitude (Y) of the event spot indexed by “i”. Depicting weekly SDEs and connecting consecutive centers to exhibit transmission direction employs parallel idea but roles of time and space interchange: Time interval is now not random; it is fixed to be one week. The magnitude of changes in X and Y are random, implying the velocity (speed plus direction) of transmission.
5 Future Perspectives
For infectious diseases such as avian influenza, spatial clustering of outbreaks plays a highly significant role in ecological dynamics and viral spread. However, accurate identifying the spatial cluster and predicting the direction of viral spread requires the knowledge of a number of environmental factors with spatial structures, which are not only non-randomly distributed across a country but also change through time. Instead of applying complex spatial statistics for clustering tests to detect a series of epidemiological anomalies, development of intuitive tools for timely identification of spatial–temporal clusters will assist control measures to prevent further damage. Wu et al. [57] proposed two visual approaches to identify spatial–temporal cluster with its dynamic change through two-stage methods. In the first stage, they utilized common concepts of Knox test and scan likelihood ratio statistics to determine spatiotemporal cluster. Although there is no universally feasible method to estimate the direction of transmission, the use of SDE in the second stage to visualize the geographical distribution of a series of social, biological or environmental events is still very attractive [64, 67,68,69]. Geographically, the scale wider than local infections was presented by simply connecting the centroids corresponding to each week’s ellipse. If the initial pattern was influenced by local factors, the direction connecting centroids can be exerted by a later “strength” existing among ellipses. Time-varying SDEs are applied to individual spatial clusters, defined by the Knox method, to reveal its local transmission by week. By connecting the consecutive centers of weekly SDEs, the direction of transmission can be easily visualized, which may imply the playing roles of local factors, such as wild bird movement, transportation vehicles, human activities or other meteorological factors acted within the spatial clusters [22, 38, 46, 70,71,72]. Other non-local factors, such as factors related to poultry market supply networks or the long-distance movement of certain bird species, contributing to the HPAI transmission between spatial clusters can be investigated and differentiated from the local factors [73,74,75]. Careful identification of influencing factors can help precautionary measures, public health control and prevent further outbreaks. Therefore, a Knox-based combined SDE visualization tool is suggested to identify the spatial-temporal clustering of poultry farm HPAI outbreaks in Taiwan.
On the other hand, AGC (Fig. 9.2)-based second-order aggregation maps based on scan statistics likelihood ratio as two-stage approach in a regular interval provide a quantitative risk in regional level and its dynamic change further indicates the direction of transmission [55]. The likelihood ratio statistic constructed in the first stage considers two “reference populations” to serve as the basis for statistical testing on global and local spatial clustering. A map based on drawing the AGC index, which can capture the aggregation pattern of disease clusters is very useful for displaying hotspots. That is, the aggregation of those sub-regions with higher \({\mathrm{R}}_{\mathrm{j}}\) or AGC index is called hotspots. The identified major clusters are similar in both Knox-based and AGC mapping methods. Although the AGC map inevitably depends on the choice of the critical value of the AGC index, the difference between two results is small. These major spatial clusters or hotspots could share common environmental risk factors contributing to the poultry farm outbreaks by HPAI as we published previously [42]. By monthly depicting the AGC maps, the changes in the hotspot pattern over a period of time also provide clues of the direction of HPAI viral transmission. If the AGC maps of different months remain unchanged, it means that the hotspot is very “stable” in a sense. Note that the formation of AGC map depends on the choice of the cutoff point for the number of clusters. The traditional elbow method based on minimizing the overall within-cluster variation can be applied, or the more modern gap statistics can be used in the future [76, 77].
In conclusion, various approaches to study spatial or temporal-spatial clustering in infectious diseases have been proposed. However, the knowledge of a number of environmental factors with spatial structures is necessary to accurately identify the spatial clustering in a global or local scale. Development of a visual tool in a webpage will assist in accurate identification of such clustering and predicting the direction of viral spreading.
References
Palese P, Shaw M (2007) Orthomyxoviridae: the viruses and their replication, 5th ed. Knipe DMHP, Griffin D, Lamb R, Martin M, Roizman B, Strauss S, editors. Philadelphia: Lippincott Williams and Wilkins: Fields virology
Tong S, Zhu X, Li Y, Shi M, Zhang J, Bourgeois M et al (2013) New world bats harbor diverse influenza A viruses. PLoS Pathog 9(10):e1003657
(CDC) CfDCaP (1997) Isolation of avian influenza A(H5N1) viruses from humans–Hong Kong, May–December 1997. MMWR Morb Mortal Wkly Rep 46(50):1204–7
Group WOFHNEW (2008) Toward a unified nomenclature system for highly pathogenic avian influenza virus (H5N1). Emerg Infect Dis 14(7):e1
Anderson TK, Macken CA, Lewis NS, Scheuermann RH, Van Reeth K, Brown IH et al (2016) A phylogeny-based global nomenclature system and automated annotation tool for H1 hemagglutinin genes from swine influenza A viruses. mSphere 1(6)
(WHO) WHO (1980) A revision of the system of nomenclature for influenza viruses: a WHO memorandum. Bull World Health Organ 58(4):585–91
Klenk HD, Garten W, Bosch FX, Rott R (1982) Viral glycoproteins as determinants of pathogenicity. Med Microbiol Immunol 170(3):145–153
Kawaoka Y, Webster RG (1988) Sequence requirements for cleavage activation of influenza virus hemagglutinin expressed in mammalian cells. Proc Natl Acad Sci U S A 85(2):324–328
Vey M, Orlich M, Adler S, Klenk HD, Rott R, Garten W (1992) Hemagglutinin activation of pathogenic avian influenza viruses of serotype H7 requires the protease recognition motif R-X-K/R-R. Virology 188(1):408–13
Wood GW, McCauley JW, Bashiruddin JB, Alexander DJ (1993) Deduced amino acid sequences at the haemagglutinin cleavage site of avian influenza A viruses of H5 and H7 subtypes. Arch Virol 130(1–2):209–217
Alexander DJ (2007) An overview of the epidemiology of avian influenza. Vaccine 25(30):5637–5644
Lee DH, Criado MF, Swayne DE (2021) Pathobiological origins and evolutionary history of highly pathogenic avian influenza viruses. Cold Spring Harb Perspect Med 11(2)
Naguib MM, Verhagen JH, Mostafa A, Wille M, Li R, Graaf A et al (2019) Global patterns of avian influenza A (H7): virus evolution and zoonotic threats. FEMS Microbiol Rev 43(6):608–621
Selleck PW, Arzey G, Kirkland PD, Reece RL, Gould AR, Daniels PW et al (2003) An outbreak of highly pathogenic avian influenza in Australia in 1997 caused by an H7N4 virus. Avian Dis 47(3 Suppl):806–811
Chen H, Smith GJD, Zhang SY, Qin K, Wang J, Li KS et al (2005) Avian flu: H5N1 virus outbreak in migratory waterfowl. Nature 436:191–192
Liu J, Xiao H, Lei F, Zhu Q, Qin K, Zhang XW et al (2005) Highly pathogenic H5N1 influenza virus infection in migratory birds. Science 309(5738):1206
Lycett SJ, Pohlmann A, Staubach C, Caliendo V, Woolhouse M, Beer M, Kuiken T, Global Consortium for HN, Related Influenza V (2020) Genesis and spread of multiple reassortants during the 2016/2017 H5 avian influenza epidemic in Eurasia. Proc Natl Acad Sci USA 117:20814–20825
Prosser DJ, Chen J, Ahlstrom CA, Reeves AB, Poulson RL, Sullivan JD et al (2022) Maintenance and dissemination of avianorigin influenza A virus within the northern Atlantic Flyway of North America. PLoS Pathog 18(6):e1010605
Cuia Y, Li Y, Lia M, Zhaoa L, Wang D, Tian J, Bai X, Cia Y et al (2020) Evolution and extensive reassortment of H5 influenza viruses isolated from wild birds in China over the past decade. Emerg Microbes Infect 9(1):1793–1803
Runstadler J, Hill N, Hussein IT, Puryear W, Keogh M (2013) Connecting the study of wild influenza with the potential for pandemic disease. Infect Genet Evol 17:162–187
Taubenberger JK, Morens DM (2009) Pandemic influenza–including a risk assessment of H5N1. Rev Sci Tech 28(1):187–202
Global Consortium for H5N8 and Related Influenza Viruses (2016) Role for migratory wild birds in the global spread of avian influenza H5N8. Science 354(6309):213–7
Kwon J, Youk S, Lee DH (2022) Role of wild birds in the spread of clade 2.3.4.4e H5N6 highly pathogenic avian influenza virus into South Korea and Japan. Infect Genet Evol 101:105281
Verhagen JH, Fouchier RAM, Lewis N (2021) Highly pathogenic avian influenza viruses at the wild-domestic bird interface in Europe: future directions for research and surveillance. Viruses 13(2):212
Bevins SN, Shriner SA, Cumbee JC Jr, Dilione KE, Douglass KE, Ellis JW et al (2022) Intercontinental movement of highly pathogenic avian influenza A(H5N1) clade 2.3.4.4 virus to the United States, 2021. Emerg Infect Dis 28(5):1006–1011
van der Kolk JH (2019) Role for migratory domestic poultry and/or wild birds in the global spread of avian influenza? Vet Q 39(1):161–167
Poen MJ, Bestebroer TM, Vuong O, Scheuer RD, van der Jeugd HP, Kleyheeg E et al (2018) Local amplification of highly pathogenic avian influenza H5N8 viruses in wild birds in the Netherlands, 2016 to 2017. Euro Surveill 23(4):17–00449
Kwon YK, Joh SJ, Kim MC, Lee YJ, Choi JG, Lee EK, Wee SH, Sung HW, Kwon JH, Kang MI, Kim JH (2005) Highly pathogenic avian influenza in magpies (Pica pica sericea) in South Korea. J Wildl Dis 41:618–623
Lee CW, Suarez DL, Tumpey TM, Sung HW, Kwon YK, Lee YJ, Choi JG et al (2005) Characterization of highly pathogenic H5N1 avian influenza A viruses isolated from South Korea. J Virol 79:3692–3702
Caliendo V, Leijten L, van der M Bildt, Germeraad E, Fouchier RAM, Beerens N, Kuiken T (2022) Tropism of highly pathogenic avian influenza H5 viruses from the 2020/2021 epizootic in wild ducks and geese. Viruses 14(2):280
Engelsma M, Heutink R, Harders, Germeraad EA, Beerens N (2022) Multiple introductions of reassorted highly pathogenic avian influenza H5Nx viruses clade 2.3.4.4b causing outbreaks in wild birds and poultry in The Netherlands, 2020–2021. Microbiol Spectr 10(2):e0249921
Hill NJ, Bishop MA, Trovão NS, Ineson KM, Schaefer AL, Puryear WB, Zhou K et al (2022) Ecological divergence of wild birds drives avian influenza spillover and global spread. PLoS Pathog 18(5):e1010062
Tang L, Tang W, Li X, Hu C, Wu D, Wang T, He G (2020) Avian influenza virus prevalence and subtype diversity in wild birds in Shanghai, China, 2016–2018. Viruses 12(9):1031
Runstadler J, Hill N, Hussein IT, Puryear W, Keogh M (2013) Connecting the study of wild influenza with the potential for pandemic disease. Infect Genet Evol 17:162–187
Hicks JT, Edwards K, QiuI X, Kim DK, Hixson J, Krauss S et al (2022) Host diversity and behavior determine patterns of interspecies transmission and geographic diffusion of avian influenza A subtypes among North American wild reservoir species. PLoS Pathog 18(4):e1009973
Chen Y, Liang W, Yang S, Wu N, Gao H, Sheng J et al (2013) Human infections with the emerging avian influenza A H7N9 virus from wet market poultry: clinical analysis and characterisation of viral genome. Lancet 381(9881):1916–1925
Zhang T, Bi Y, Tian H, Li X, Liu D, Wu Y et al (2014) Human infection with influenza virus A(H10N8) from live poultry markets, China, 2014. Emerg Infect Dis 20(12):2076–2079
Gilbert M, Pfeiffer DU (2012) Risk factor modelling of the spatio-temporal patterns of highly pathogenic avian influenza (HPAIV) H5N1: a review. Spat Spatiotemporal Epidemiol 3(3):173–183
Souris M, Bichaud L (2011) Statistical methods for bivariate spatial analysis in marked points. Examples in spatial epidemiology. Spat Spatiotemporal Epidemiol 2(4):227–34
Wu H, Wang X, Xue M, Wu C, Lu Q, Ding Z et al (2017) Spatial characteristics and the epidemiology of human infections with avian influenza A(H7N9) virus in five waves from 2013 to 2017 in Zhejiang Province, China. PLoS One 12(7):e0180763
Shan X, Wang Y, Song R, Wei W, Liao H, Huang H et al (2020) Spatial and temporal clusters of avian influenza a (H7N9) virus in humans across five epidemics in mainland China: an epidemiological study of laboratory-confirmed cases. BMC Infect Dis 20(1):630
Liang WS, He YC, Wu HD, Li YT, Shih TH, Kao GS et al (2020) Ecological factors associated with persistent circulation of multiple highly pathogenic avian influenza viruses among poultry farms in Taiwan during 2015–17. PLoS One 15(8):e0236581
ArcGIS Pro 2.8. How Hot Spot Analysis (Getis-Ord Gi*) works. https://pro.arcgis.com/en/pro-app/2.8/tool-reference/spatial-statistics/h-how-hot-spot-analysis-getis-ord-gi-spatial-stati.htm. Accessed 17 Jun 2022
Huang D, Dong W, Wang Q (2021) Spatial and temporal analysis of human infection with the avian influenza A (H7N9) virus in China and research on a risk assessment agent-based model. Int J Infect Dis 106:386–394
Krisp JM, Špatenková O (2010) Kernel density estimations for visual analysis of emergency response data. Geographic information and cartography for risk and crisis management, pp 395–408
Dong W, Yang K, Xu Q, Liu L, Chen J (2017) Spatio-temporal pattern analysis for evaluation of the spread of human infections with avian influenza A(H7N9) virus in China, 2013–2014. BMC Infect Dis 17(1):704
Duczmal L, Kulldorff M, Huang L (2006) Evaluation of spatial scan statistics for irregularly shaped clusters. J Comput Gr Stat 428–42
Kulldorff M, Huang L, Pickle L, Duczmal L (2006) An elliptic spatial scan statistic. Stat Med 25(22):3929–3943
Kulldorff M, Hjalmars U (1999) The Knox method and other tests for space-time interaction. Biometrics 55(2):544–552
Kulldorff M, Heffernan R, Hartman J, Assuncao R, Mostashari F (2005) A space-time permutation scan statistic for disease outbreak detection. PLoS Med 2(3):e59
Zhang Y, Shen Z, Ma C, Jiang C, Feng C, Shankar N et al (2015) Cluster of human infections with avian influenza A (H7N9) cases: a temporal and spatial analysis. Int J Environ Res Public Health 12(1):816–828
Knox EG, Bartlett MS (1964) The detection of space-time interactions. J Roy Stat Soc Ser C (Appl Stat) 13(1):25–30
Barton DE, David FN (1966) The random intersection of two graphs. Wiley, London, New York. Research papers in statistics: festschrift for J. Neyman, pp 455–9
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27(2):209–220
Openshaw S, Craft AW, Charlton M, Birch JM (1988) Investigation of leukaemia clusters by use of a Geographical Analysis Machine. Lancet 1(8580):272–273
Turnbull BW, Iwano EJ, Burnett WS, Howe HL, Clark LC (1990) Monitoring for clusters of disease: application to leukemia incidence in upstate New York. Am J Epidemiol 132(1 Suppl):S136–S143
Wu HI, Chao DY (2021) Two-stage algorithms for visually exploring spatio-temporal clustering of avian influenza virus outbreaks in poultry farms. Sci Rep 11(1):22553
Lefever DW (1926) Measuring geographic concentration by means of the standard deviational ellipse. Am J Sociol 32(1):88–94
Furfey PH (1927) A note on lefever’s “Standard Deviational Ellipse.” Am J Sociol 33(1):94–98
Yuill RS (1971) The standard deviational ellipse; an updated tool for spatial description. Geogr Annal Ser B Hum Geogr 53(1):28–39
Wang B, Shi W, Miao Z (2015) Confidence analysis of standard deviational ellipse and its extension into higher dimensional euclidean space. PLoS One 10(3):e0118537
Cressie N, Chan NH (1989) Spatial modeling of regional variables. J Am Stat Assoc 84(406):393–401
Eryando T, Susanna D, Pratiwi D, Nugraha F (2012) Standard Deviational Ellipse (SDE) models for malaria surveillance, case study: Sukabumi district-Indonesia, in 2012. Malar J 11(1):P130
Satoto TBT, Satrisno H, Lazuardi L, Diptyanusa A, Purwaningsih, Rumbiwati et al (2019) Insecticide resistance in Aedes aegypti: an impact from human urbanization? PLoS One 14(6):e0218079
Reference APT. How Directional Distribution (Standard Deviational Ellipse) works. https://pro.arcgis.com/en/pro-app/2.7/tool-reference/spatial-statistics/h-how-directional-distribution-standard-deviationa.htm. Accessed 3 Jul 2021
Zinszer K, Morrison K, Anema A, Majumder MS, Brownstein JS (2015) The velocity of Ebola spread in parts of west Africa. Lancet Infect Dis 15(9):1005–1007
Spumont F, Viti F (2018) The effect of workplace relocation on individuals’ activity travel behavior. J Trans Land Use 11(1):985–1002
Liu S, Qin Y, Xie Z, Zhang J (2020) The spatio-temporal characteristics and influencing factors of Covid-19 spread in Shenzhen, China-an analysis based on 417 cases. Int J Environ Res Public Health 17(20)
Moore TW, McGuire MP (2019) Using the standard deviational ellipse to document changes to the spatial dispersion of seasonal tornado activity in the United States. npj Clim Atmos Sci 2(1):21
Artois J, Jiang H, Wang X, Qin Y, Pearcy M, Lai S et al (2018) Changing geographic patterns and risk factors for avian influenza A(H7N9) infections in humans, China. Emerg Infect Dis 24(1):87–94
Busani L, Valsecchi MG, Rossi E, Toson M, Ferre N, Pozza MD et al (2009) Risk factors for highly pathogenic H7N1 avian influenza virus infection in poultry during the 1999–2000 epidemic in Italy. Vet J 181(2):171–177
Paul M, Tavornpanich S, Abrial D, Gasqui P, Charras-Garrido M, Thanapongtharm W et al (2010) Anthropogenic factors and the risk of highly pathogenic avian influenza H5N1: prospects from a spatial-based model. Vet Res 41(3):28
Hogerwerf L, Wallace RG, Ottaviani D, Slingenbergh J, Prosser D, Bergmann L et al (2010) Persistence of highly pathogenic avian influenza H5N1 virus defined by agro-ecological niche. EcoHealth 7(2):213–225
Lai PC, Wong CM, Hedley AJ, Lo SV, Leung PY, Kong J et al (2004) Understanding the spatial clustering of severe acute respiratory syndrome (SARS) in Hong Kong. Environ Health Perspect 112(15):1550–1556
Leibler JH, Otte J, Roland-Holst D, Pfeiffer DU, Soares Magalhaes R, Rushton J et al (2009) Industrial food animal production and global health risks: exploring the ecosystems and economics of avian influenza. EcoHealth 6(1):58–70
Everitt BS, Landau S, Leese M, Stahl D (2011) Cluster analysis, 5th ed. Wiley
Tibshirani R, Walther G, Hastie T (2001) Estimating the number of clusters in a data set via the gap statistic. J R Stat Soc B 63:411–423
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Huang, ML., Wu, HD.I., Chao, DY. (2023). Approaches for Spatial and Temporal-Spatial Clustering Analysis in Avian Influenza Outbreaks. In: Wen, TH., Chuang, TW., Tipayamongkholgul, M. (eds) Earth Data Analytics for Planetary Health. Atmosphere, Earth, Ocean & Space. Springer, Singapore. https://doi.org/10.1007/978-981-19-8765-6_9
Download citation
DOI: https://doi.org/10.1007/978-981-19-8765-6_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-8764-9
Online ISBN: 978-981-19-8765-6
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)