Abstract
Cancer biomarkers allow diagnosis, risk assessment, monitoring disease progression, and prediction of therapeutic response in oncology. Different forms and types of cancer biomarkers exist and it covers a broad range of biochemical entities including DNA, miRNA, cirRNA, and proteins. The advances in high-throughput technologies including genomics, transcriptomics, proteomics, and metabolomics have generated great opportunities for the discovery of new and effective cancer biomarkers. Due to the heterogeneous nature of cancer cells, it is difficult to identify the clinically useful precise cancer biomarkers. Multiomics data analyses using a systems approach play a vital role in the discovery of cancer biomarkers. In this chapter, a brief classification system for cancer biomarkers has been provided according to their biochemical nature and based on clinical utility along with the application of recent high-throughput approaches used in cancer biomarker discovery. Several databases and bioinformatics tools applied in cancer biomarker discovery have been mentioned. In summary, different cancer biomarker types, omics approaches used for cancer biomarker discovery, and dedicated cancer-related databases and tools have been discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
3.1 Introduction
Cancer, a heterogeneous group of diseases, represents the second leading cause of death globally and is responsible for an estimated ten million deaths worldwide in 2020 (Sung et al. 2021). For advancement and better treatment in cancer, patient’s early diagnosis and accurate detection of cancer are the crucial steps. In the past few decades, the biomarker field along with the improved quality of medical services and technologies has transformed the ability of cancer researchers to easily diagnose and classify cancer at the molecular level and resulted in improved drug development and clinical trial design (Hu and Dignam 2019; Parker et al. 2021; Goossens et al. 2015; Goyal et al. 2021). The definition of biomarker has been given differently by different groups. Some of the definitions limit the scope of biomarkers up to biological molecule or biochemical features; on the other hand, a broader definition of biomarker provided by the Biomarker Consortium (Foundation of National Institute of Health) increases the probability of discovering new biomarkers in the ever-changing era of research biology (Califf 2018; Wu and Qu 2015; Strimbu and Tavel 2010). According to United Nations, World Health Organization (WHO) biomarkers can be defined as any measurable substance, structure, or process or its product that can predict the incidence of disease outcome (World Health Organization 2001). In simple words, any measurable indicator of disease condition and treatment response indicator can be considered as a biomarker (Zare Jeddi et al. 2021; Lassere 2008). In a study in 1848, light chain of immunoglobulin was identified as the first-ever cancer biomarker in the myeloma patient urine sample (Solomon 1980). Since then, numerous cancer biomarkers have been identified such as alfa-fetoprotein, carcinoembryonic antigen (CEA), and prostate-specific antigen (PSA). Every era of cancer biomarker discovery has been closely associated with the new and powerful technology (Tatarinov 1964; Xu et al. 2021; Campos-da-Paz et al. 2018; Cózar et al. 2021; Wang et al. 1979).
In the past few decades, high-throughput technologies such as genomics, metagenomics, transcriptomics, proteomics, and metabolomics have generated a significant amount of data for cancer biology. For example, a proteomics-based study identified DNA-dependent protein kinase catalytic subunit (DNA-PKcs) as a potential diagnostic biomarker for clinical outcomes in breast cancer (Asleh et al. 2021). A combined study of transcriptomics with metabolomics revealed a panel of important serum biomarkers such as 2-hydroxybutyric acid and 4-hydroxybutyric acid as candidate diagnostic biomarkers for lung cancer proliferation through the Ca2+ signaling pathway (Zheng et al. 2021). In addition, miRNA-194 was identified as a favorable prognostic biomarker for gastric cancer (Wang et al. 2021a). Moreover, the machine learning-based computational algorithm helps to classify the complex pattern of cancer research outcomes generated by a plethora of high-throughput experiments (Echle et al. 2021). As a result, these omics approaches when coupled with bioinformatics and computational methods provide great opportunities for biomarker discovery and facilitate therapeutic development for cancer (Menyhárt and Győrffy 2021; Yan et al. 2016).
As with time, the information about cancer biomarkers has expanded, and so also the complexities of tumor biology evolve adding challenges to the development of efficient biomarkers. More powerful tools, cross-validation techniques, and system biology approaches should be deployed in future to increase the yield of biomarkers in cancer therapeutics (Ileana Dumbrava et al. 2018; Sheng et al. 2020; Louie et al. 2021). In this chapter, we have highlighted three aspects of cancer biomarkers. The first part deals with the cancer biomarker classification according to the molecular types and the potential role they play in clinical oncology. The second part relates to the omics approaches that are nowadays considered as a powerful strategy to untangle the complex biological behavior of cancer cells. These high-throughput technologies help to discover molecular biomarkers with prognostic, diagnostic, and targeted therapeutic values. The third part discusses some important bioinformatics tools and software related to cancer biomarker discovery. The multiomics data integration and molecular regulatory network-based analysis tool can revolutionize the biomarker field for the effective treatment of cancer.
3.2 Categorization of Cancer Biomarkers
In biomedical science, cancer biomarker identification is one of the major multidisciplinary areas and their categorization should be considered contextual. Further, due to the enhancement in recent advanced technologies, enormous cancer biomarkers have been reported thereby it is difficult to categorize cancer biomarkers by considering only one aspect (Henry and Hayes 2012). However, according to contemporary findings, several attempts have been applied to classify cancer biomarkers (Nguyen et al. 2020). A basic schematic representation for the classification of cancer biomarkers is shown in Fig. 3.1.
3.2.1 Classification of Cancer Biomarkers Based on Biomolecules
3.2.1.1 DNA Cancer Biomarkers
Genetic alteration, gene rearrangement, and point mutation are responsible for cancer progression (Housman et al. 2014; Torgovnick and Schumacher 2015; Li et al. 2021a). The most common cancer DNA biomarker includes single nucleotide polymorphisms (SNPs). In a recent study, SNP–SNP interactions have been identified as a potential indicator of prostate cancer aggressiveness (Lin et al. 2021). Another study demonstrated m6A-associated functional SNPs (PLEKHA8, SMUG1, CDC123, RMI2, ACSM5) as major functional variants for thyroid cancer (Ruan et al. 2021). Moreover, researchers have defined circulating tumor DNA in localized nonsmall cell lung cancer as a prognostic biomarker and predicted the survival rate (Peng et al. 2020). Epigenetic CpG methylation also provides a broad range for early cancer detection and can be used as a cancer biomarker (Locke et al. 2019).
3.2.1.2 RNA Cancer Biomarkers
Quantitative Reverse Transcription Polymerase Chain Reaction (RT-qPCR), microarray, Serial Analysis of Gene Expression (SAGE), differential display, microfluid card, and bead-based methods are commonly used to detect RNA or miRNA cancer biomarkers (Xi et al. 2017; Yamashita et al. 2008). Various identified mRNA and miRNA act as effective diagnostic and prognostic cancer biomarkers (Paramasivam 2021; Wang et al. 2021b; Bautista-Sánchez et al. 2020). In blood-based liquid biopsies, platelet RNA has been detected as an early diagnostic biomarker for cancer (Wurdinger et al. 2020). A recent article has indicated that miRNA plays an important role in identification of drug and decision-making of drug delivery in cancer therapeutics (Paramasivam 2021). Clinical researches reported that circRNAs expression indicates the cancer prognosis in different types of cancer, for example in a recent study circRNA expression in peripheral blood has been found to be correlated with cancer size (Li and Han 2019; Pan et al. 2019; Chen et al. 2017).
3.2.1.3 Protein Cancer Biomarkers
The most commonly used techniques used to identify protein-based biomarkers include Polyacrylamide Gel Electrophoresis (PAGE) and Two-Dimensional fluorescence Difference Gel Electrophoresis (2D-DIGE) (Issaq and Veenstra 2008). The high-throughput method includes proteomics study based on Mass Spectroscopy (MS), Surface-Enhanced Laser Absorption Desorption Ionization Time of Flight (SELDI-TOF), and Matrix-Associated Laser Absorption Desorption Ionization Time of Flight (MALDI-TOF) and discovered various protein and peptide biomarkers for ovarian and breast cancer (Zeidan et al. 2009; Liu 2011; Swiatly et al. 2017; Lv et al. 2019). Quantitative proteomics has also been applied to identify potential cancer protein biomarkers in different types of cancers (Kwon et al. 2021). To discover secreted protein-mediated interaction between the cancer cell and nonmalignant stroma, Stable Isotope Labeling with Amino Acids in Cell culture (SILAC) was performed and further in pancreatic cancer, Wntless homolog protein (WLS) and Myristoylated Alanine-rich C-Kinase Substrate (MARCKS) found to be associated with oxaliplatin resistance (Kim et al. 2021; Wang et al. 2018). Isobaric Tags for Relative and Absolute Quantitation (iTRAQ), a quantitative proteomics approach was applied to find differentially expressed protein in metformin-treated cervical cancer cells and reported that metformin increases tumor suppressor gene expression IGFBP7 (Xia et al. 2020). Liquid Chromatography-Mass Spectrometry/Mass Spectrometry (LC-MS/MS) and antibody arrays are used in lung breast and colon cancer to find a panel of potential protein biomarkers (Wang et al. 2016a; Huang and Zhu 2017). Protein-based biomarkers are considered as a more valuable biomarker as compared to DNA- and RNA-based biomarkers as they are involved in functional molecular pathways and determine the disease initiation and progression state (Zhang et al. 2019). In a recent study, PINK1 protein has been reported as a prognostic biomarker for cancer (Zhu et al. 2020).
3.2.1.4 Carbohydrate Cancer Biomarkers
Cancer progression is often associated with changes in the expression of surface carbohydrates such as N-linked and O-linked glycans (Leney et al. 2017). The glycobiomarkers [glycoprotein, glycolipid, and proteoglycan] serve as candidate epidemiological cancer biomarkers (Lan et al. 2016; Daniotti et al. 2013). Mass spectrometry such as MALDI-TOF and Electrospray Ionization (ESI) are generally used for profiling of N- and O-linked glycosylation at serine and threonine residue of candidate protein molecules in human sera and cancer cell lines (Dube and Bertozzi 2005; An et al. 2006, 2010; Drake et al. 2017). Glycan microarray analysis has been found a good biomarker identification method for the diagnosis of breast cancer (Wang et al. 2008). Further in hepatocellular carcinoma, cancer-associated carbohydrate antigens (DSGG, fucosyl GM1, and Gb2 of CACAs) have been reported as potential biomarkers for early detection of cancer (Wu et al. 2012).
3.2.2 Classification of Cancer Biomarkers Based on Clinical Utility
Based on the putative application, cancer biomarkers can be classified under the following categories. Although some biomarkers are overlapping in nature, for example, the grading and staging cancer biomarker is also used as a prediction and screening biomarker (Ludwig and Weinstein 2005).
3.2.2.1 Prediction Cancer Biomarker
Predictive biomarkers predict the response and efficacy of the treatment and also help to determine the optimal dose of the drug at the initial treatment stages (Alves Martins et al. 2019; Bai et al. 2020). As cancer is a heterogeneous disease and the same cancer type responds differently to a drug thereby these types of biomarkers help in selecting a successful treatment process and minimizing the drug toxicity. A common predictive biomarker is overexpression of HER2, which predicts breast cancer’s response to drugs like trastuzumab (Jørgensen and Hersom 2016). A high level of circulating IFN-γ predicts the response of immunotherapies to immune checkpoint blockade in melanoma and nonsmall cell lung cancer patients (Karachaliou et al. 2018). A more recent study predicts that overexpression of excision repairs cross-complementation group 1 (ERCC1) increases DNA excision repair and imparts resistance to platinum-based drugs (Chung 2021). Additionally, in colorectal cancer, the mutation in MAPK pathway genes serves as a predictive biomarker for EGFR therapy and indicates resistance to cetuximab drug (Boussios et al. 2019).
3.2.2.2 Detection/Diagnostic Cancer Biomarker
The screening or detection of cancer biomarkers is the real indicator of the presence of cancer. These biomarkers help to identify benign cancer before metastasis. The tumor cells produce several immune factors, serum proteins, and circulating free DNA and RNA and these molecules can serve as cancer detection biomarkers (Parker et al. 2018). Diagnostic biomarkers play an important role in classifying patients into subtypes and also detecting the presence of the disease. Prostate-specific antigen (PSA) is the best-known cancer biomarker for prostate cancer early detection (Welch and Albertsen 2009). Cancer Antigen 19-9 (CA-19-9) is a diagnostic serum biomarker for pancreatic ductal carcinoma (Poruk et al. 2013). Further, another cancer antigen CA 125 is also used as a classical biomarker for the detection of ovarian cancer (Felder et al. 2014). The utility of cytokines as diagnostic biomarkers is increasing rapidly although further validation is required. IL-6 and VEGF serve as possible diagnostic biomarkers for ovarian and gastric cancer (Liang et al. 2015; Monastero and Pentyala 2017). Diagnostic biomarkers are often used in conjunction with other specific biomarkers to increase the specificity and diagnosis in the general population (Califf 2018).
3.2.2.3 Prognostic Cancer Biomarkers
Prognostic biomarkers allow to monitor the disease status, detect the recurrence rate, and provide an idea about the overall patient survival rate, independent of therapy (Sechidis et al. 2018; Ruberg and Shen 2015). It allows estimating the risk of the disease. In colon, lung, and breast cancer, patient’s carcinoembryonic antigen (CEA) signifies a poor survival rate (Boonpipattanapong and Chewatanakornkul 2006; Su et al. 2012). Some diagnostic biomarkers such as Cancer antigen 19-9 (CA-19-9) and cancer antigen (CA 125) have also prognostic values and their presence can predict the survival rate in pancreatic ductal carcinoma and ovarian cancer, respectively (Poruk et al. 2013; Felder et al. 2014). Other prognostic biomarkers include miR-155 which suggests a poor clinical prognosis in hepatocellular carcinoma (Nalejska et al. 2014). A recent study reported five signature miRNAs having prognostic values in colon cancer (Lv et al. 2020). Further for breast cancer, circulating tumor cells serve as a prognostic indicator in nonmetastatic breast cancer as their presence is correlated with metastasis (Lucci et al. 2012).
3.2.2.4 Pharmacodynamics Cancer Biomarkers
This type of biomarkers is the new classification-based biomarker that determines the degree of the drug response (Sarker and Workman 2007). Pharmacodynamic biomarkers provide an idea about the interaction between a drug and its suspected target and whether the drug exerted a cellular response or not, and thereby guide treatment decision-making plans in real-time (Jackson 2012). For example, in nonsmall cell, lung cancer patient measurement of Mitogen-Activated Protein Kinase (MAPK) pathway inhibition receiving BRAF inhibitors can suggest a direct interaction between drug and target genes (Gainor et al. 2014). Further, Ki67 acts as a biomarker for cell proliferation, its expression after treatment with endocrine therapy serves as a pharmacodynamic response, and indicates target drug effects (Kelloff et al. 2005). Another example of pharmacodynamic biomarker example is monitoring the activity of PARP enzyme in white blood cells for the development of anticancer drug Olaparib (Dick et al. 2021).
3.2.3 Classification of Cancer Biomarkers Based on Other Criteria
3.2.3.1 Imaging Cancer Biomarkers
X-ray, Positron Emission Tomography (PET), Computed Tomography (CT), ultrasound, radionuclide imaging, and Magnetic Resonance Imaging (MRI) are the imaging techniques that are routinely used in clinical oncology for diagnosis, screening, and staging of cancer (Dregely et al. 2018; O’Connor et al. 2017). In oncology, imaging biomarkers are cost-effective noninvasive tools that easily allow identifying the disease state including assessment of the drug response. Several attempts have been made to do a regular assessment to reduce the risk of cancer development. For example, colonoscopy and mammography have been found to reduce the risk of developing colon cancer and breast cancer, respectively (Bischoff 2014).
3.2.3.2 Pathological Cancer Biomarkers
Various types of infectious agents such as viruses and bacteria constitute 15–20% of all human cancers (Srivastava et al. 2005; McLaughlin-Drubin and Munger 2008). The presence of pathogenic agents within the tumor cell makes them attractive pathogenic cancer biomarkers. The presence of HPV is associated with cervical cancer (Burd 2003). Further, Epstein Bair Virus (EBV) is closely associated with lymphoma and nasopharyngeal carcinoma (Pagano 1999). Helicobacter pylori is an established biomarker for gastric cancer (Wroblewski et al. 2010). Besides, various cancer pathogen detection methods have been revolutionized recently for rapid biomarker detection in the complex biological sample including Bioluminescence Resonance Energy Transfer (BRET), Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-based biosensors, and ELISA (Wu and Qu 2015).
3.3 Omics Approaches in Cancer Biomarker Research
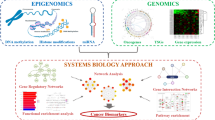
“Omics” studies have been characterized by high-throughput technologies which help to investigate the genome, transcriptome, epigenome, proteome, and metabolome of cancer cells. These omics approaches facilitate the understanding of carcinogenesis at the molecular level. Figure 3.2 depicts a schematic representation of omics approaches that are commonly used to study the cellular behavior of cancer cells and some important biomarkers identified by using these omics approaches.
3.3.1 Genomics for Cancer Biomarkers
In oncology, numerous technologies such as Next-Generation Sequencing (NGS), Whole-Genome Sequencing (WGS), Comparative Genome Hybridization (CGH), and Fluorescence in situ Hybridization (FISH) have been widely used to analyze cancer-specific mutational changes (Hu et al. 2018; Zhao et al. 2019). Additionally, genomic studies primarily focus on the analysis of copy number variation and identification of chromosomal abnormality to characterize cancer cells at the molecular endpoint (Nogrady 2020). Further, the advances in sequencing technology helped the decision-making for personalized treatment strategy instead of based on cancer type. For example, FDA-approved biomarker EGFR mutation signifies the effectiveness of EGFR inhibitors like gefitinib (Tsimberidou et al. 2020). BRCA1 and BRCA2 mutations have been identified as hereditary markers for breast and ovarian cancer syndrome (Narod and Salmena 2011; Savanevich et al. 2021). Moreover, genomic profiling of pancreatic cancer identified centromere protein F as a novel therapeutic target and proved to be responsible for cancer progression (Chen et al. 2021).
3.3.2 Transcriptomics for Cancer Biomarkers
Transcriptomics studies are engaged in quantification, detection, and identification of altered mRNA, miRNA, and lncRNA in cancer cell populations (Chakraborty et al. 2018). Tools like RNA-seq, and microarray are commonly used to study the transcriptome. Expression Quantitative Trait Loci (eQTL) is a new approach for the analysis of functional variation sequence that leads to changes in gene expression (Hong et al. 2020; Geeleher et al. 2018). Various prognostic and predictive gene signatures have been identified in lung, breast, colon, and other tumor types (Vishnubalaji et al. 2019; Xiong et al. 2020; Sheng et al. 2019; Fang et al. 2021; Li et al. 2019). The transcriptomic study-based microarray technology has also been applied in precision oncology trials and aids in the clinical classification of breast cancer, colon cancer, and gastric cancer (Salem et al. 2017; Guinney et al. 2015; Lin et al. 2015). In breast cancer, transcriptomic data with bioinformatics study reveal BRIP1 as a noteworthy prognostic biomarker and its expression found to be correlated with various clinical features of breast cancer (Khan and Khan 2021). Furthermore, from RNA-seq data, a differential gene expression pattern has been revealed for different cancer tissue and their normal counterpart which will uncover the complex molecular pattern of cancer cells (Li et al. 2017). Different types of RNA serve as independent cancer biomarkers for instance in nonsmall cell lung cancer (NSCLC), the expression profiling of snoRNAs serves as an early diagnostic cancer biomarker (Liao et al. 2010). For renal cancer, hepatic cancer and glioblastoma piRNAs may serve as diagnostic and prognostic biomarkers (Busch et al. 2015; Liu et al. 2019; Rizzo et al. 2016). In addition, lncRNAs, such as XIST reported as a potential candidate biomarker for gastric cancer through transcriptomic study (Lu et al. 2017).
3.3.3 Proteomics for Cancer Biomarkers
Proteomic-based studies are considered as one of the innovative and dynamic high-throughput technology for determining the cellular function and location of main mediators of proteins (Olivier et al. 2019). The proteome of a defined entity be it a cell, an organelle, a tissue allows for better biomarker identification and to better understand the cancer surveillance mechanisms (Sallam 2015). Several studies reported that the proteomics approach can be used to identify the drug resistance nature of cancer cells and treatment resistance biomarkers; for example through mass spectrometry, PYCR1 and ALDH18A1 expressions have been identified to be significantly associated with drug resistance in breast cancer (Shenoy et al. 2020). The drug resistance of cancer is associated with stemness and by applying the proteomics approach, new specific cancer biomarkers and therapeutic targets have been identified in the breast cancer stem cell population (Koh et al. 2020). Protein profiling of patients receiving immunotherapy is necessary to monitor the therapeutic response and thereby proteomic study can help to discover the potential prognostic biomarkers for cancer therapeutics (Chae et al. 2020; Harel et al. 2019).
3.3.4 Metabolomics for Cancer Biomarkers
Cancer affects intracellular metabolism and results in the inappropriate proliferation of cells (Vander Heiden and DeBerardinis 2017; Pavlova and Thompson 2016). Metabolomics is the study of altered metabolites that are produced by cellular processes mediated by proteins and thus it is a direct assessment of phenotype. Plasma or serum samples from patients are the major focus for metabolomic analysis of cancer cells. The methodologies that are used for metabolomic studies for biomarker detection include mass spectrometry and Nuclear Magnetic Resonance (NMR)-based imaging techniques, such as Magnetic Resonance Spectroscopic Imaging (MRSI) which can use both tissue/cell or biopsies samples for detection (Schmidt et al. 2021). Some putative metabolite biomarkers are altered carbohydrates in acute myeloid leukemia and unsaturated free fatty acids in colon cancer (Chaturvedi et al. 2013; Zhang et al. 2016). Other metabolite biomarkers include changes in citric acid, branched-chain amino acid for prostate cancer and pancreatic cancer (Giskeødegård et al. 2013; Mayers et al. 2014). Bladder cancer biomarkers detected from urinary metabolic profiling and 27 differentially metabolites have been detected (Li et al. 2021b).
3.3.5 Epigenomics for Cancer Biomarker
Epigenomics can be defined as the study of genome-wide chemical modification such as acetylation and methylation of DNA. The epigenetic modifications play an important role in uncovering the important genetic marker as these modifications regulate cellular interactions (Piunti and Shilatifard 2016). ChIP seq and Whole-Genome Bisulfite Sequencing (WGBS) are the two important powerful techniques for the identification of DNA-binding sites of transcription factors and to detect the methylated part in the sequences respectively (Chakraborty et al. 2018; Raj et al. 2017). MBD-isolated Genome Sequencing (MiGS) is another novel technique that allows the analysis of whole-genome sequencing patterns (Serre et al. 2010). For predicting the risk of head and neck cancer, DNA methylation in saliva has been found to be a potential epigenetic biomarker (Rapado-González et al. 2021). Chromatin immunoprecipitation (ChIP) studies demonstrated RUNX1T1 as an epigenetic regulator of Small Cell Lung Cancer (SCLC) (He et al. 2021). A recent finding suggests that PD-L1 methylation in CpG loci can be considered as a valuable diagnostic biomarker for gastric cancer (Amini et al. 2021).
3.4 Bioinformatics Analytical Tools for Cancer Biomarker Discovery
The emerging high-throughput technologies result in the exponential growth of cancer biomarker data set from various resources. Thereby biologists face difficulty in extracting useful information from the available repositories as these contain various types of cancer-related information. The Cancer Genome Atlas (TCGA), a resource of multiomics cancer data platform that integrates genomics, epigenomics, and transcriptomics data of more than 30 human tumor types (Wang et al. 2016b). This aims to provide publicly available comprehensive atlas for molecular alteration in cancer cell. Pan-Cancer initiative is the new version of TCGA atlas and it is dedicated for comparison and analysis of molecular alteration found in different tumor types (Cancer Genome Atlas Research Network 2013). Recently published bioinformatics cancer-related database MarkerDB provides molecular cancer biomarker information along with the clinical significance such as diagnostic marker or prognostic marker (Wishart et al. 2021). Another database, OncoMX is a knowledge base; it integrates data for cancer mutation gene signatures, differential expression genes for cancer (Dingerdissen et al. 2020). Similarly, CIViCmine is another recently published database that provides list of curative clinically relevant cancer biomarkers information (Lever et al. 2019). Different machine learning and statistical approaches allow to the identification of biomolecules of interest from the large dataset with quantitative measurements. BioPlat is a software package for cancer biomarker discovery that allows high-throughput data filtering, gene expression calculation in silico (Butti et al. 2014). Another recently developed software Q omics that enables the analysis of patient survival, gene expression, and mutation of cancer-driven data set (Lee et al. 2021). There is an R-based tool available for cancer data analysis named CAncer bioMarker Prediction Pipeline (CAMPP), a standardized framework for the analysis of quantitative biological data (Terkelsen et al. 2020). Overall, several dedicated databases and tools are available for the storage and discovery of cancer biomarkers (Table 3.1).
3.5 Future Challenges
The future of biomarkers in oncology is potentially associated with the diagnostic, predictive, and prognostic cancer biomarkers. Despite the explosion of new technologies, a number of hurdles are associated with the identification of potential cancer biomarkers to be considered in clinical trials. The major challenges for cancer biomarker discovery can be considered at three levels (Henry and Hayes 2012; Teutsch et al. 2009). First is analytic validity which can be described as pre- and postanalytical evaluation of biomarker detection assay. It determines the specificity and sensitivity of the technical aspects (Hayes 2015). The second is clinical validity, it determines the diagnostic accuracy of biomarkers by dividing the population of interest into two groups such as patient and reference group (Bossuyt 2010). The third is clinical utility, which relates to making a clinical decision with a high level of evidence to improve cancer treatment outcomes (Hayes 2021). The aim of cancer research is earlier cancer diagnosis and get better clinical outcomes for precision oncology. Nevertheless, multiomics approaches offer great advantages for translational cancer research over monogenic markers. The rapid development of omics biomarkers increases the specificity of targeted therapeutic approach and leads to enhance predictive, preventive, and personalized medicine (PPPM) practice in clinical oncology (Lu and Zhan 2018).
References
Alves Martins BA et al (2019) Biomarkers in colorectal cancer: the role of translational proteomics research. Front Oncol 9:1284
Amini M, Hejazi M, Ghorban K, Mokhtarzadeh A, Baradaran B (2021) Identification of functional methylated CpG loci in PD-L1 promoter as the novel epigenetic biomarkers for primary gastric cancer. Gene 772:145376
An HJ et al (2006) Profiling of glycans in serum for the discovery of potential biomarkers for ovarian cancer. J Proteome Res 5:1626–1635
An Z et al (2010) Integrated ionization approach for RRLC-MS/MS-based metabonomics: finding potential biomarkers for lung cancer. J Proteome Res 9:4071–4081
Asleh K et al (2021) Proteomics-derived basal biomarker DNA-PKcs is associated with intrinsic subtype and long-term clinical outcomes in breast cancer. NPJ Breast Cancer 7:114
Bai R, Lv Z, Xu D, Cui J (2020) Predictive biomarkers for cancer immunotherapy with immune checkpoint inhibitors. Biomark Res 8:34
Bautista-Sánchez D et al (2020) The promising role of miR-21 as a cancer biomarker and its importance in RNA-based therapeutics. Mol Ther Nucleic Acids 20:409–420
Bischoff M (2014) Tumor markers, mammography and colon cancer screening. Prevention in oncology—what makes sense, what does not? MMW Fortschr Med 156:27
Boonpipattanapong T, Chewatanakornkul S (2006) Preoperative carcinoembryonic antigen and albumin in predicting survival in patients with colon and rectal carcinomas. J Clin Gastroenterol 40:592–595
Bossuyt PMM (2010) Clinical validity: defining biomarker performance. Scand J Clin Lab Investig 70:46–52
Boussios S et al (2019) The developing story of predictive biomarkers in colorectal cancer. J Pers Med 9:E12
Burd EM (2003) Human papillomavirus and cervical cancer. Clin Microbiol Rev 16:1–17
Busch J et al (2015) Piwi-interacting RNAs as novel prognostic markers in clear cell renal cell carcinomas. J Exp Clin Cancer Res 34:61
Butti MD et al (2014) BioPlat: a software for human cancer biomarker discovery. Bioinformatics 30:1782–1784
Califf RM (2018) Biomarker definitions and their applications. Exp Biol Med (Maywood) 243:213–221
Campos-da-Paz M, Dórea JG, Galdino AS, Lacava ZGM, de Fatima Menezes Almeida Santos M (2018) Carcinoembryonic antigen (CEA) and hepatic metastasis in colorectal cancer: update on biomarker for clinical and biotechnological approaches. Recent Pat Biotechnol 12:269–279
Cancer Genome Atlas Research Network et al (2013) The cancer genome atlas pan-cancer analysis project. Nat Genet 45:1113–1120
Chae YK et al (2020) Mass spectrometry-based serum proteomic signature as a potential biomarker for survival in patients with non-small cell lung cancer receiving immunotherapy. Transl Lung Cancer Res 9:1015–1028
Chakraborty S, Hosen MI, Ahmed M, Shekhar HU (2018) Onco-multi-OMICS approach: a new frontier in cancer research. Biomed Res Int 2018:9836256
Chaturvedi A et al (2013) Mutant IDH1 promotes leukemogenesis in vivo and can be specifically targeted in human AML. Blood 122:2877–2887
Chen J et al (2017) Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett 388:208–219
Chen H et al (2021) Centromere protein F is identified as a novel therapeutic target by genomics profile and contributing to the progression of pancreatic cancer. Genomics 113:1087–1095
Chung C (2021) Predictive and prognostic biomarkers with therapeutic targets in colorectal cancer: a 2021 update on current development, evidence, and recommendation. J Oncol Pharm Pract 28(4):850–869. https://doi.org/10.1177/10781552211005525
Cózar JM, Hernández C, Miñana B, Morote J, Alvarez-Cubero MJ (2021) The role of prostate-specific antigen in light of new scientific evidence: an update in 2020. Actas Urol Esp (Engl Ed) 45:21–29
Daniotti JL, Vilcaes AA, Torres Demichelis V, Ruggiero FM, Rodriguez-Walker M (2013) Glycosylation of glycolipids in cancer: basis for development of novel therapeutic approaches. Front Oncol 3:306
Dick P, Jos H, B. (2021) Analytical validation of quantitative pharmacodynamic methods used in clinical cancer studies. Int Arch Clin Pharmacol 7:26
Dingerdissen HM et al (2020) OncoMX: a knowledgebase for exploring cancer biomarkers in the context of related cancer and healthy data. JCO Clin Cancer Inform 4:210–220
Drake RR et al (2017) MALDI mass spectrometry imaging of N-linked glycans in cancer tissues. Adv Cancer Res 134:85–116
Dregely I et al (2018) Imaging biomarkers in oncology: basics and application to MRI. J Magn Reson Imaging 48:13–26
Dube DH, Bertozzi CR (2005) Glycans in cancer and inflammation—potential for therapeutics and diagnostics. Nat Rev Drug Discov 4:477–488
Echle A et al (2021) Deep learning in cancer pathology: a new generation of clinical biomarkers. Br J Cancer 124:686–696
Fang Z, Xu S, Xie Y, Yan W (2021) Identification of a prognostic gene signature of colon cancer using integrated bioinformatics analysis. World J Surg Oncol 19:13
Felder M et al (2014) MUC16 (CA125): tumor biomarker to cancer therapy, a work in progress. Mol Cancer 13:129
Gainor JF, Longo DL, Chabner BA (2014) Pharmacodynamic biomarkers: falling short of the mark? Clin Cancer Res 20:2587–2594
Geeleher P et al (2018) Cancer expression quantitative trait loci (eQTLs) can be determined from heterogeneous tumor gene expression data by modeling variation in tumor purity. Genome Biol 19:130
Giskeødegård GF et al (2013) Spermine and citrate as metabolic biomarkers for assessing prostate cancer aggressiveness. PLoS One 8:e62375
Goossens N, Nakagawa S, Sun X, Hoshida Y (2015) Cancer biomarker discovery and validation. Transl Cancer Res 4:256–269
Goyal B et al (2021) Diagnostic, prognostic, and therapeutic significance of long non-coding RNA MALAT1 in cancer. Biochim Biophys Acta Rev Cancer 1875:188502
Guinney J et al (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21:1350–1356
Harel M et al (2019) Proteomics of melanoma response to immunotherapy reveals mitochondrial dependence. Cell 179:236–250.e18
Hayes DF (2015) Biomarker validation and testing. Mol Oncol 9:960–966
Hayes DF (2021) Defining clinical utility of tumor biomarker tests: a clinician’s viewpoint. JCO 39:238–248
He T et al (2021) Identification of RUNX1T1 as a potential epigenetic modifier in small-cell lung cancer. Mol Oncol 15:195–209
Henry NL, Hayes DF (2012) Cancer biomarkers. Mol Oncol 6:140–146
Hong M et al (2020) RNA sequencing: new technologies and applications in cancer research. J Hematol Oncol 13:166
Housman G et al (2014) Drug resistance in cancer: an overview. Cancers (Basel) 6:1769–1792
Hu C, Dignam JJ (2019) Biomarker-driven oncology clinical trials: key design elements, types, features, and practical considerations. JCO Precis Oncol 3:1–12. https://doi.org/10.1200/PO.19.00086
Hu T et al (2018) Forward and reverse mutations in stages of cancer development. Hum Genom 12:40
Huang Y, Zhu H (2017) Protein array-based approaches for biomarker discovery in cancer. Genom Proteom Bioinform 15:73–81
Ileana Dumbrava E, Meric-Bernstam F, Yap TA (2018) Challenges with biomarkers in cancer drug discovery and development. Expert Opin Drug Discov 13:685–690
Issaq H, Veenstra T (2008) Two-dimensional polyacrylamide gel electrophoresis (2D-PAGE): advances and perspectives. BioTechniques 44:697–698
Jackson RC (2012) Pharmacodynamic modelling of biomarker data in oncology. ISRN Pharmacol 2012:590626
Jørgensen JT, Hersom M (2016) Companion diagnostics-a tool to improve pharmacotherapy. Ann Transl Med 4:482
Karachaliou N et al (2018) Interferon gamma, an important marker of response to immune checkpoint blockade in non-small cell lung cancer and melanoma patients. Ther Adv Med Oncol 10:1758834017749748
Kelloff GJ et al (2005) Progress and promise of FDG-PET imaging for cancer patient management and oncologic drug development. Clin Cancer Res 11:2785–2808
Khan U, Khan MS (2021) Prognostic value estimation of BRIP1 in breast cancer by exploiting transcriptomics data through bioinformatics approaches. Bioinform Biol Insights 15:11779322211055892
Kim YE et al (2021) SILAC-based quantitative proteomic analysis of oxaliplatin-resistant pancreatic cancer cells. Cancers 13:724
Koh E-Y, You J-E, Jung S-H, Kim P-H (2020) Biological functions and identification of novel biomarker expressed on the surface of breast cancer-derived cancer stem cells via proteomic analysis. Mol Cells 43:384–396
Kwon YW et al (2021) Application of proteomics in cancer: recent trends and approaches for biomarkers discovery. Front Med (Lausanne) 8:747333
Lan Y et al (2016) Serum glycoprotein-derived N- and O-linked glycans as cancer biomarkers. Am J Cancer Res 6:2390–2415
Lassere MN (2008) The biomarker-surrogacy evaluation schema: a review of the biomarker-surrogate literature and a proposal for a criterion-based, quantitative, multidimensional hierarchical levels of evidence schema for evaluating the status of biomarkers as surrogate endpoints. Stat Methods Med Res 17:303–340
Lee J et al (2021) Q-omics: smart software for assisting oncology and cancer research. Mol Cells 44:843–850
Leney AC, El Atmioui D, Wu W, Ovaa H, Heck AJR (2017) Elucidating crosstalk mechanisms between phosphorylation and O-GlcNAcylation. Proc Natl Acad Sci USA 114:E7255–E7261
Lever J et al (2019) Text-mining clinically relevant cancer biomarkers for curation into the CIViC database. Genome Med 11:78
Li S, Han L (2019) Circular RNAs as promising biomarkers in cancer: detection, function, and beyond. Genome Med 11:15
Li M, Sun Q, Wang X (2017) Transcriptional landscape of human cancers. Oncotarget 8:34534–34551
Li H et al (2019) Transcriptomic analysis and identification of prognostic biomarkers in cholangiocarcinoma. Oncol Rep 42(5):1833–1842. https://doi.org/10.3892/or.2019.7318
Li L, Guan Y, Chen X, Yang J, Cheng Y (2021a) DNA repair pathways in cancer therapy and resistance. Front Pharmacol 11:629266
Li J et al (2021b) Bladder cancer biomarker screening based on non-targeted urine metabolomics. Int Urol Nephrol 54(1):23–29. https://doi.org/10.1007/s11255-021-03080-6
Liang B et al (2015) Circulating VEGF as a biomarker for diagnosis of ovarian cancer: a systematic review and a meta-analysis. Onco Targets Ther 8:1075–1082
Liao J et al (2010) Small nucleolar RNA signatures as biomarkers for non-small-cell lung cancer. Mol Cancer 9:198
Lin X, Zhao Y, Song W, Zhang B (2015) Molecular classification and prediction in gastric cancer. Comput Struct Biotechnol J 13:448–458
Lin H-Y et al (2021) KLK3 SNP–SNP interactions for prediction of prostate cancer aggressiveness. Sci Rep 11:9264
Liu C (2011) The application of SELDI-TOF-MS in clinical diagnosis of cancers. J Biomed Biotechnol 2011:1–6
Liu Y et al (2019) The emerging role of the piRNA/piwi complex in cancer. Mol Cancer 18:123
Locke WJ et al (2019) DNA methylation cancer biomarkers: translation to the clinic. Front Genet 10:1150
Louie AD, Huntington K, Carlsen L, Zhou L, El-Deiry WS (2021) Integrating molecular biomarker inputs into development and use of clinical cancer therapeutics. Front Pharmacol 12:747194
Lu M, Zhan X (2018) The crucial role of multiomic approach in cancer research and clinically relevant outcomes. EPMA J 9:77–102
Lu Q et al (2017) Potential lncRNA diagnostic biomarkers for early gastric cancer. Mol Med Rep 16:9545–9552
Lucci A et al (2012) Circulating tumour cells in non-metastatic breast cancer: a prospective study. Lancet Oncol 13:688–695
Ludwig JA, Weinstein JN (2005) Biomarkers in cancer staging, prognosis and treatment selection. Nat Rev Cancer 5:845–856
Lv P et al (2019) Exploratory study on application of MALDI-TOF-MS to detect serum and urine peptides related to small cell lung carcinoma. Mol Med Rep 21(1):51–60. https://doi.org/10.3892/mmr.2019.10794
Lv Y, Duanmu J, Fu X, Li T, Jiang Q (2020) Identifying a new microRNA signature as a prognostic biomarker in colon cancer. PLoS One 15:e0228575
Mayers JR et al (2014) Elevation of circulating branched-chain amino acids is an early event in human pancreatic adenocarcinoma development. Nat Med 20:1193–1198
McLaughlin-Drubin ME, Munger K (2008) Viruses associated with human cancer. Biochim Biophys Acta 1782:127–150
Menyhárt O, Győrffy B (2021) Multi-omics approaches in cancer research with applications in tumor subtyping, prognosis, and diagnosis. Comput Struct Biotechnol J 19:949–960
Monastero RN, Pentyala S (2017) Cytokines as biomarkers and their respective clinical cutoff levels. Int J Inflamm 2017:4309485
Nalejska E, Mączyńska E, Lewandowska MA (2014) Prognostic and predictive biomarkers: tools in personalized oncology. Mol Diagn Ther 18:273–284
Narod SA, Salmena L (2011) BRCA1 and BRCA2 mutations and breast cancer. Discov Med 12:445–453
Nguyen TTH et al (2020) Salivary biomarkers in oral squamous cell carcinoma. JKAOMS 46:301–312
Nogrady B (2020) How cancer genomics is transforming diagnosis and treatment. Nature 579:S10–S11
O’Connor JPB et al (2017) Imaging biomarker roadmap for cancer studies. Nat Rev Clin Oncol 14:169–186
Olivier M, Asmis R, Hawkins GA, Howard TD, Cox LA (2019) The need for multi-omics biomarker signatures in precision medicine. Int J Mol Sci 20:E4781
Pagano JS (1999) Epstein-Barr virus: the first human tumor virus and its role in cancer. Proc Assoc Am Physicians 111:573–580
Pan B et al (2019) Identification of serum exosomal hsa-circ-0004771 as a novel diagnostic biomarker of colorectal cancer. Front Genet 10:1096
Paramasivam G (2021) Micro-RNA (miRNA): a biomarker to identify novel compounds in drug discovery and delivery for cancer therapy. Curr Drug Discov Technol 18:e130921188092
Parker LA et al (2018) Diagnostic biomarkers: are we moving from discovery to clinical application? Clin Chem 64:1657–1667
Parker JL et al (2021) Does biomarker use in oncology improve clinical trial failure risk? A large-scale analysis. Cancer Med 10:1955–1963
Pavlova NN, Thompson CB (2016) The emerging hallmarks of cancer metabolism. Cell Metab 23:27–47
Peng M et al (2020) Circulating tumor DNA as a prognostic biomarker in localized non-small cell lung cancer. Front Oncol 10:561598
Piunti A, Shilatifard A (2016) Epigenetic balance of gene expression by Polycomb and COMPASS families. Science 352:aad9780
Poruk KE et al (2013) The clinical utility of CA 19-9 in pancreatic adenocarcinoma: diagnostic and prognostic updates. Curr Mol Med 13:340–351
Raj U, Aier I, Semwal R, Varadwaj PK (2017) Identification of novel dysregulated key genes in breast cancer through high-throughput ChIP-Seq data analysis. Sci Rep 7:3229
Rapado-González Ó et al (2021) Salivary DNA methylation as an epigenetic biomarker for head and neck cancer. Part II: a cancer risk meta-analysis. J Pers Med 11:606
Rizzo F et al (2016) Specific patterns of PIWI-interacting small noncoding RNA expression in dysplastic liver nodules and hepatocellular carcinoma. Oncotarget 7:54650–54661
Ruan X et al (2021) Genome-wide identification of m6A-associated functional SNPs as potential functional variants for thyroid cancer. Am J Cancer Res 11:5402–5414
Ruberg SJ, Shen L (2015) Personalized medicine: four perspectives of tailored medicine. Stat Biopharm Res 7:214–229
Salem H, Attiya G, El-Fishawy N (2017) Classification of human cancer diseases by gene expression profiles. Appl Soft Comput 50:124–134
Sallam RM (2015) Proteomics in cancer biomarkers discovery: challenges and applications. Dis Mark 2015:1–12
Sarker D, Workman P (2007) Pharmacodynamic biomarkers for molecular cancer therapeutics. Adv Cancer Res 96:213–268
Savanevich A, Ashuryk O, Cybulski C, Lubiński J, Gronwald J (2021) BRCA1 and BRCA2 mutations in ovarian cancer patients from Belarus: update. Hered Cancer Clin Pract 19:13
Schmidt DR et al (2021) Metabolomics in cancer research and emerging applications in clinical oncology. CA Cancer J Clin 71:333–358
Sechidis K et al (2018) Distinguishing prognostic and predictive biomarkers: an information theoretic approach. Bioinformatics 34:3365–3376
Serre D, Lee BH, Ting AH (2010) MBD-isolated genome sequencing provides a high-throughput and comprehensive survey of DNA methylation in the human genome. Nucleic Acids Res 38:391–399
Sheng M, Xie X, Wang J, Gu W (2019) A pathway-based strategy to identify biomarkers for lung cancer diagnosis and prognosis. Evol Bioinform 15:117693431983849
Sheng KL et al (2020) An integrated approach to biomarker discovery reveals gene signatures highly predictive of cancer progression. Sci Rep 10:21246
Shenoy A et al (2020) Proteomic patterns associated with response to breast cancer neoadjuvant treatment. Mol Syst Biol 16:e9443
Solomon A (1980) Monoclonal immunoglobulins as biomarkers of cancer. In: Sell S (ed) Cancer markers. Humana Press, Totowa, pp 57–87. https://doi.org/10.1007/978-1-4612-6117-9_3
Srivastava S, Verma M, Gopal-Srivastava R (2005) Proteomic maps of the cancer-associated infectious agents. J Proteome Res 4:1171–1180
Strimbu K, Tavel JA (2010) What are biomarkers? Curr Opin HIV AIDS 5:463–466
Su C-H et al (2012) The carcinoembryonic antigen as a potential prognostic marker for neuroendocrine carcinoma of the breast. Anticancer Res 32:183–188
Sung H et al (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J Clin 71:209–249
Swiatly A et al (2017) MALDI-TOF-MS analysis in discovery and identification of serum proteomic patterns of ovarian cancer. BMC Cancer 17:472
Tatarinov IS (1964) Detection of embryo-specific ALPHA-globulin in the blood serum of a patient with primary liver cancer. Vopr Med Khim 10:90–91
Terkelsen T, Krogh A, Papaleo E (2020) CAncer bioMarker prediction pipeline (CAMPP)—a standardized framework for the analysis of quantitative biological data. PLoS Comput Biol 16:e1007665
Teutsch SM et al (2009) The evaluation of genomic applications in practice and prevention (EGAPP) initiative: methods of the EGAPP working group. Genet Med 11:3–14
Torgovnick A, Schumacher B (2015) DNA repair mechanisms in cancer development and therapy. Front Genet 6:157
Tsimberidou AM, Fountzilas E, Nikanjam M, Kurzrock R (2020) Review of precision cancer medicine: evolution of the treatment paradigm. Cancer Treat Rev 86:102019
Vander Heiden MG, DeBerardinis RJ (2017) Understanding the intersections between metabolism and cancer biology. Cell 168:657–669
Vishnubalaji R, Sasidharan Nair V, Ouararhni K, Elkord E, Alajez NM (2019) Integrated transcriptome and pathway analyses revealed multiple activated pathways in breast cancer. Front Oncol 9:910
Wang MC, Valenzuela LA, Murphy GP, Chu TM (1979) Purification of a human prostate specific antigen. Investig Urol 17:159–163
Wang C-C et al (2008) Glycan microarray of Globo H and related structures for quantitative analysis of breast cancer. Proc Natl Acad Sci USA 105:11661–11666
Wang H et al (2016a) The clinical impact of recent advances in LC-MS for cancer biomarker discovery and verification. Expert Rev Proteom 13:99–114
Wang Z, Jensen MA, Zenklusen JC (2016b) A practical guide to the cancer genome atlas (TCGA). Methods Mol Biol 1418:111–141
Wang X et al (2018) SILAC-based quantitative MS approach for real-time recording protein-mediated cell-cell interactions. Sci Rep 8:8441
Wang J et al (2021a) miRNA-194 predicts favorable prognosis in gastric cancer and inhibits gastric cancer cell growth by targeting CCND1. FEBS Open Bio 11:1814–1826
Wang H et al (2021b) Circular RNA TMEM87A promotes cell proliferation and metastasis of gastric cancer by elevating ULK1 via sponging miR-142-5p. J Gastroenterol 56:125–138
Welch HG, Albertsen PC (2009) Prostate cancer diagnosis and treatment after the introduction of prostate-specific antigen screening: 1986–2005. J Natl Cancer Inst 101:1325–1329
Wishart DS et al (2021) MarkerDB: an online database of molecular biomarkers. Nucleic Acids Res 49:D1259–D1267
World Health Organization (2001) Biomarkers in risk assessment: validity and validation. World Health Organization, Geneva
Wroblewski LE, Peek RM, Wilson KT (2010) Helicobacter pylori and gastric cancer: factors that modulate disease risk. Clin Microbiol Rev 23:713–739
Wu L, Qu X (2015) Cancer biomarker detection: recent achievements and challenges. Chem Soc Rev 44:2963–2997
Wu C-S et al (2012) Cancer-associated carbohydrate antigens as potential biomarkers for hepatocellular carcinoma. PLoS One 7:e39466
Wurdinger T, In’t Veld SGJG, Best MG (2020) Platelet RNA as pan-tumor biomarker for cancer detection. Cancer Res 80:1371–1373
Xi X et al (2017) RNA biomarkers: frontier of precision medicine for cancer. Noncoding RNA 3:E9
Xia C, Yang F, He Z, Cai Y (2020) iTRAQ-based quantitative proteomic analysis of the inhibition of cervical cancer cell invasion and migration by metformin. Biomed Pharmacother 123:109762
Xiong Y, Feng Y, Qiao T, Han Y (2020) Identifying prognostic biomarkers of non-small cell lung cancer by transcriptome analysis. Cancer Biomark 27:243–250
Xu Y, Guo Q, Wei L (2021) The emerging influences of alpha-fetoprotein in the tumorigenesis and progression of hepatocellular carcinoma. Cancers (Basel) 13:5096
Yamashita T, Honda M, Kaneko S (2008) Application of serial analysis of gene expression in cancer research. Curr Pharm Biotechnol 9:375–382
Yan W, Xue W, Chen J, Hu G (2016) Biological networks for cancer candidate biomarkers discovery. Cancer Inform 15:1–7
Zare Jeddi M et al (2021) Towards a systematic use of effect biomarkers in population and occupational biomonitoring. Environ Int 146:106257
Zeidan BA et al (2009) SELDI-TOF MS proteomics in breast cancer. Clin Proteom 5:133–147
Zhang Y et al (2016) Serum unsaturated free fatty acids: a potential biomarker panel for early-stage detection of colorectal cancer. J Cancer 7:477–483
Zhang X, Sun X-F, Shen B, Zhang H (2019) Potential applications of DNA, RNA and protein biomarkers in diagnosis, therapy and prognosis for colorectal cancer: a study from databases to AI-assisted verification. Cancers (Basel) 11:E172
Zhao EY, Jones M, Jones SJM (2019) Whole-genome sequencing in cancer. Cold Spring Harb Perspect Med 9:a034579
Zheng Y et al (2021) Combined metabolomics with transcriptomics reveals important serum biomarkers correlated with lung cancer proliferation through a calcium signaling pathway. J Proteome Res 20:3444–3454
Zhu L et al (2020) Pan-cancer analysis of the mitophagy-related protein PINK1 as a biomarker for the immunological and prognostic role. Front Oncol 10:569887
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Firdous, S., Srivastava, S.K., Saha, S. (2022). Cancer Biomarkers in the Era of Systems Biology. In: Singh, S. (eds) Systems Biomedicine Approaches in Cancer Research. Springer, Singapore. https://doi.org/10.1007/978-981-19-1953-4_3
Download citation
DOI: https://doi.org/10.1007/978-981-19-1953-4_3
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-1952-7
Online ISBN: 978-981-19-1953-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)