Abstract
Gp120 is a critical viral proteins required for HIV-1 entry and infection. It facilitates HIV-1 binding to target cells, human-to-human transmission, relocation of virus from mucosa to lymph nodes, cell–cell infection and syncytium formation, and the bystander effect that kills uninfected CD4+ T-cells and other human cells. Molecules that bind to gp120 can inhibit its function by stabilizing conformations of the protein, leading to the inability to infect cells, and resulting in non-permissive. Small molecule-mediated stabilization of certain conformations of gp120 may also enhance recognition of HIV-1 infected cells by neutralizing antibodies and make the virus more susceptible to effector functions such as ADCC, which could potentially be part of future cure regimens. Additionally, HIV attachment inhibitors can complex with free gp120 and potentially repress both cytopathic effects from membrane-bound or soluble gp120. Fostemsavir (RukobiaTM), a phosphate prodrug of an HIV-1 attachment inhibitor that was recently approved for use in highly treatment experienced (HTE) patients with multidrug resistant HIV-1 is a first-in-class drug with a favorable safety profile that provides an additional treatment option for treatment in this population of patients with a high medical need.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
The latest new chemical entity for the treatment of HIV-1 is fostemsavir (RukobiaTM) which received FDA approval in the U.S. in July 2020. The discovery program and early development activities were carried out at Bristol-Myers Squibb company and as a result of a business agreement, the successful late stage development work and subsequent marketing of the compound for patients was conducted by ViiV HealthCare Inc.
The Phase 3 BRIGHTE study demonstrated the benefits of fostemsavir for multidrug resistant HIV subjects. Two cohorts were evaluated by the addition of fostemsavir. The randomized cohort consisted of subjects who had at least one but no more than two remaining active approved HIV agents. These subjects received 600 mg bid fostemsavir or a placebo, while the non-randomized cohort (subjects with no remaining fully active options) all received 600 mg fostemsavir bid. In the randomized cohort, those who received fostemsavir had a significantly greater decrease in the HIV RNA level than those who received placebo during the first 8 days and this result has been sustained through 48 weeks and longer (Kozal et al. 2020). Efficacy was also observed in the non-randomized cohort.
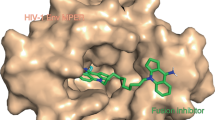
This drug is the first example of a mechanistically unique class of HIV entry inhibitor called an attachment inhibitor (HIV AI). HIV AIs bind to HIV gp120 and block the initial contact with human CD4+ T-cell, essentially interrupting the HIV life cycle at the earliest stage (Fig. 4.1) (Lin et al. 2003).
Infected CD4+ T-cells express gp120 on their membrane which is packaged into newly formed progeny infectious virus particles after the budding process. These viruses can then infect new CD4+ T-cells. However, cell-bound gp120s are also capable of mediating cell-to-cell transfection (Fig. 4.2), a second route for HIV to deliver its genetic material to uninfected CD4+. HIV AIs can suppress both virus–cell and cell–cell HIV infection.
4.2 Discovery and SAR of Fostemsavir-Related Series (BMS Series)
The original attachment inhibitor lead compound (BMS-‘216) was discovered from a whole cell, antiviral screen of a portion (~100K compounds) of Bristol-Myers Squibb’s chemical inventory (Fig. 4.3). It was the sole hit from the screen and exhibited a novel mechanism of action. BMS-‘216 exhibited an EC50 value of 153 nM in a pseudotype assay using envelopes from the CCR5-dependent JRFL strain and CXCR4-dependent LAI strain of HIV-1, and a CC50 value of 339 μM in uninfected cells (Wang et al. 2019). A subsequent combinatory effort that included a library of single position changes revealed a F atom at position 4 of the indole (BMS-‘705, EC50 2.6 nM) that enhanced activity by 58-fold (Meanwell et al. 2009a). However, BMS-‘705’s solubility in water, and especially in common excipients, was low, and unsuccessful efforts to identify an oral formulation in the context of other properties precluded advancement. A nitrogen atom was then introduced into the indole ring to improve the solubility and metabolic stability (Wang et al. 2009a). As expected, 4-/5-/6-/7-azaindole analogues of BMS-‘216-Me all displayed better aqueous solubility (>26-fold improvement) and metabolic stability (>2.3-fold improvement) (Wang et al. 2009a).
The addition of a MeO– group in the 7-azadole promptly produced BMS-‘806 (Lin et al. 2003; Wang et al. 2003), which was advanced to first-in-man (FIM) clinical studies. Unfortunately, the targeted exposure was not achieved in the Phase I trial, presumably due to the moderate metabolic stability (t1/2 47 min in HLM) and permeability (Caco-2 Pc 51 nm/s) measured for the compound in preclinical studies. A closely related 6-azaindole molecule, BMS-‘043, was evaluated soon after and proved to be a compound with improved preclinical characteristics (t1/2 > 100 min in HLM, Caco-2 Pc 178 nm/s) (Wang et al. 2009a). BMS-‘043 was advanced to Phase II clinical trials and the resulting virology data established proof-of-concept (POC) for the new and novel mechanism (Ho et al. 2006; Hanna et al. 2011). Several issues which precluded further advancement of BMS-‘043 were that the targeted viral load drop was observed in only ~68% of patients and that this result was achieved at the relatively high dose requirement of 1.8 g of compound co-administered with a high-fat meal. Further rounds of optimization focused on addressing the abovementioned shortcomings via further modification of position 7 of the 6-azaindole. Installation of amides, aromatic rings, C-linked heteroaryl and ultimately N-linked heteroaryl substituents led to the discovery of BMS-‘529 (temsavir, EC50 0.1 nM) (Wang et al. 2018). Temsavir (Nowicka-Sans et al. 2012) possessed a broader spectrum of anti-HIV activity than its predecessors. The preclinical profile achieved all of the necessary safety and physicochemical properties with the exception of aqueous solubility, which appeared to be dictated by the requirements of the target. The less than desired water solubility of 0.022 mg/mL at pH = 7.4 resulted in the lower-than-dose-proportional-increases in exposure in dose-escalation PK studies. The solubility/dissolution limited absorption issue was resolved preclinically by the use of a phosphonooxymethyl prodrug strategy which resulted in the discovery of BMS-‘068 (fostemsavir) (Wang et al. 2018; Nettles et al. 2012). In a subsequent Phase 3 clinical study (the BRIGHTE study), fostemsavir demonstrated a favorable safety and tolerability profile, and showed effective in this difficult to treat HTE population with a high medical need, leading to its approval as a first-in-class HIV AI (Kozal et al. 2020; Markham 2020; Lataillade et al. 2020).
The main focus of the BMS team’s work was the optimization of the physicochemical properties of the (aza)indole benzamide key series described above and shown in Fig. 4.3. In addition, efforts were also spent on surveying and establishing structure activity relationships (SAR) in five regions: (A) (aza)indole core, (B) glyoxamide linker, (C) piperazine spacer, (D) benzamide moiety, and (E) a region which encompasses alterations that concomitantly change both region C and region D that could not be easily described using C or D alone (Fig. 4.4). Additional SAR has also been reported by researchers outside of Bristol-Myers Squibb.
The description of the SAR is based on published data and partially based on that contained in published patent applications and granted patents where for some compounds antiviral activity is reported in categories (e.g., Category A: EC50 < 1 μM, etc.) and some compounds have specific EC50 values disclosed. In Figs. 4.5, 4.6, 4.7, 4.8, 4.9, 4.10, 4.11, and 4.12, the representative examples were selected to highlight their unique structures or substructures.
In Region A, replacements of the (aza)indole with aromatic or heteroaromatic rings consistently maintained some level of anti-HIV-1 potency. The aromatic(hetero) rings explored were bicyclic, tricyclic, and mono-cyclic (Wang et al. 2013, 2019; Lu et al. 2009; Bender et al. 2013). In general, bicyclic heteroaromatic surrogates offered the most interesting results. While monocyclic aromatics(hetero) displayed weak potency in their unsubstituted form, inclusion of a second aromatic(hetero) functionality could rescue the potency to near that of the starting indoles and azaindoles (Wang et al. 2019; Lu et al. 2009). The most impressive (aza)indole surrogates are shown in Fig. 4.5, as antiviral potencies for (5a EC50 2.4 nM, 5b EC50 0.7 nM, 5c EC50 4.6 nM) were superior to or competitive with BMS-‘216-Me’s EC50 of 4 nM (Fig. 4.3) (Wang et al. 2019). Similar to the (aza)indole series, an aromatic ring at position 6 in 5d was able to increase activity >100-fold, with the EC50 to 0.03 nM (Wang et al. 2019). These novel series remain less studied due to the successful progression of the (aza)indole series.
A bivalent compound 5e was prepared by dimerizing two molecules of BMS-‘806. In an assay using the MT-2 cell line infected with HIV-1 IIIB virus, compound 5e exhibited an IC50 value of 4.8 nM, half of that of BMS-‘806 (2.3 nM) (Wang et al. 2005). The result initially seemed remarkable in that initial SAR obtained during the optimization of BMS-‘216 seemed to show that the best potency was observed when the C-4 position of the 7-azaindole and indole contained small functionalities and larger groups were deleterious. In the work that identified dimer 5e, the same expected trends for reduced potency at C-4 with increasing size were observed except for when two BMS-806 molecules were dimerized via a long water-soluble linker. The data suggested that for dimer 5e, only one of two BMS-‘806 units was involved in gp120 binding, while the second BMS-‘806 unit simply floated around in the extracellular medium (Wang et al. 2005). The tolerance for a bulky extension at C-4 with appropriate linkage was also independently observed in the BMS-‘806 derived conjugates described by others (Sect. 4.4.2).
The modifications of Region B were reported by two independent teams. Replacement of the glyoxamide linkage with a 2-methyl-2-oxo-carboxamide linkage appeared to provide comparable anti-HIV-1 potency to the original glyoxamide series. Example 6a in Fig. 4.6 exhibited an IC50 value of 0.04 nM against HIV-1 fusion in a gp160 induced cell–cell fusion assay that used a Hela P4 cell line and a CHO-15 Tat10 cell line (Fenwick et al. 2005).
Another team made analogs in which the glyoxamide linker was replaced with a sulfonamide. In general, the sulfonamide substitution appeared to exhibit modestly decreased potency vs. the BMS glyoxamide series. The most potent sulfonamide derivative was 6b (Fig. 4.6), with an IC50 of 7 nM in an M33 pseudotype assay, while the benchmark glyoxamide derivative’s IC50 was <5 nM in the same testing conditions (Lu et al. 2007).
In Region C, replacement of the piperazine linker was explored with a variety of diamines, including fused bicyclic diamine (Wang et al. 2014c), spiral bicyclic diamine (Wang et al. 2014d), and pyrollidine (Kadow et al. 2009) shown in Fig. 4.7. Spiral diamine derivative 7a (Wang et al. 2014c), fused diamine derivatives 7b (Wang et al. 2014d) and 7c (Wang et al. 2014d) displayed anti-HIV EC50 values of 5.65 nM, 1.41 nM and 0.68 nM, respectively, in the pseudo type assay using LAI Env. Additionally, pyrrolidine compound 7d fell into the activity category A for which the EC50 value was less than 1 μM (Kadow et al. 2009).
Interestingly, another independent group also published biological data for 7c. Against JR-CSF and B41 pseudotyped HIV-1, IC50s of 0.6 nM and 2 nM, respectively, were obtained, while the benchmark BMS-‘529 showed 0.06 nM and 0.05 nM in the same experiments (Meuser et al. 1940).
Region D was explored extensively. Some of the strategies explored were: (1) replacement of the benzene ring by other aromatic heteroaromatic rings or non-aromatic elements, (2) carbonyl isosteres of the benzamide’s carbonyl group, (3) the benzamide’s carbonyl group was replaced with an aromatic or heteroaromatic ring, (4) the benzamide’s carbonyl group was fused to form a bicyclic ring with either the aryl/heteroaryl ring or alternatively with the piperazine, and (5) the benzamide’s amide bond was replaced with an alkene.
In Approach 1, the phenyl ring was replaced either by heteroaromatic rings such as the pyridine contained in analog 8a (Fig. 4.8) (Meanwell et al. 2009b), or by non-aromatic cyclic rings such as the cyclohexane in 8b and the cycloheptane in 8c (Wang et al. 2014a). The resulting compounds furnished comparable potency (EC50: 8a 4.6 nM, 8b 0.11 nM, 8c 0.14 nM) in the initial pseudotype assay compared to their corresponding benzamides BMS-‘705 (EC50 2.6 nM) and BMS-‘529 (EC50 0.1 nM) (Meanwell et al. 2009b; Wang et al. 2014a). Nitrogen containing saturated heterocyclic rings were also explored which provided ureas. The morpholine urea 8d’s EC50 was reported as Category A (EC50 < 1 μM) (Regueiro-Ren et al. 2004).
Approach 2 explored Region D via exchange of the benzamide’s carbonyl group with its bio-isosteres such as thione (Meanwell et al. 2014), oxime (Meanwell et al. 2014), enamine (Wang et al. 2009b), amidine (Wang et al. 2008a), and sulfone (Regueiro-Ren et al. 2005). As in Fig. 4.9, thione 9a and oxime 9b showed superior potency to BMS-‘529 (EC50: 0.027 nM and 0.019 nM vs. 0.1 nM) (Meanwell et al. 2014). Furthermore, examples of an enamine 9c (EC50 < 1 μM), amidines 9d (EC50 < 0.5 μM) and 9e (EC50 < 0.5 μM), and sulfone 9f (EC50 < 1 μM) all belonged to the most active Category A. Of note, a lengthy substitution was tolerated on the amidine’s nitrogen in 9e.
Approach 3 in Region D provided potentially useful SAR. As shown in Fig. 4.10, all the examples were in the most active category A (EC50 < 0.5 μM). The carbonyl group of the benzamide was able to be changed to a benzene in 10a, a mono-heterocycle in 10b, a bicyclic aromatic ring in 10c, and a saturated cycloalkane in 10d. In 10e, the distal nitrogen atom of the piperazine was transposed into the attached cyclopentane resulting in a piperidine ring rather than piperazine as the spacer. In 10f, an additional large group was attached onto the pyrazole heteroaromatic ring, implying the protein binding interface around the moiety was either flexible or there was some space in this region (Wang et al. 2010).
Retention of potency did not require the phenyl group to be attached exclusively to the 2-position of the ring appended to the piperazine. For example, potency was retained when the phenyl ring was attached to the 3-position of the middle benzene ring, as in 10g and the 6-position of a diazabenzothiophene, as in 10h (Wang et al. 2010).
Moreover, further studies showed the carbonyl surrogate at the 2-position could be varied. The terminal benzene was exchanged with a range of functionalities such as a cycloalkane in 10i, a ketone in 10j, an amide in 10k, and a simple ether in 10l (Wang et al. 2010).
Approach 4 explored what was effectively merging and modifying the benzamide’s carbonyl group to form bicyclic rings, either via attachment to the piperazine or with the distal phenyl ring. The majority of the published anti-HIV-1 data for those compounds was expressed as potency categories in the related patents, but these in combination with some published numerical EC50s shed light on the SAR (Wang et al. 2011, 2014b; Swidorski et al. 2016). Forming a new five- or six-membered ring with the piperazine linker in place of the carbonyl group, provided data which clearly showed the preference of aromatic character for retention of potency. This was exemplified by comparison of 11a (Wang et al. 2011) and 11c (Swidorski et al. 2016) with 11b (Wang et al. 2011) in Fig. 4.11. Both fused-imidazole 11a and fused-benzene 11c were more potent than BMS-‘529 in the initial pseudotype assay (EC50: 0.06 nM and 0.02 nM vs. 0.1 nM). When one of the C=C bonds in the imidazole was saturated, the resulting compound 11b was significantly weaker, with an EC50 of 13 nM (Wang et al. 2011).
A similar trend was observed in 11d and 11e, in which the carbonyl element was infused into the distal benzene or pyridine. Both compounds were active, with the aromatic fusion in 11e having a 10-fold advantage in potency over the non-aromatic fusion in 11d (EC50: 0.07 nM vs. 0.87 nM) (Wang et al. 2014b).
An extra benzene ring was added to the fused moiety to provide the tricyclic amidine 11f. Its potency was more than tenfold less than the benchmark BMS-‘529, but it was still potent (EC50 1.83 nM). In addition, the bulkiness of 11f’s tricyclic amidine suggested there was room for structural modification in the region (Wang et al. 2015a).
In Approach 5, the polarized double bond character of the benzamide’s amide inspired its transformation into an alkene group, which lacks the amide bond’s rotatory freedom and results in a rigid sub-structure. A collection of these piperidine alkene derivatives was described in a patent application (Wang et al. 2008b). Two compounds contained in that application, BMS-‘793 (12a, DS003) (Schader et al. 2012; Ketas et al. 2007; Frank and Robbiani 2011) and BMS-‘251 (12b) (Marie Pancera et al. 2017; Lai et al. 2019), have been extensively studied, suggesting the significance of the piperidine alkene chemotype. Both 12a and 12b (Fig. 4.12) are discussed in more detail in Sects. 4.4.1 and 4.5. It is noteworthy that 12c had a bulky benzyl pyrazole on the alkene, along with the distal benzene, which again implied there was sufficient space for optimization around the region (Wang et al. 2008b).
Compound 12d explored a cyclic hydrazine isostere of the piperidine alkene, which possessed one less carbon atom and one additional nitrogen atom in the ring. It possessed an EC50 of 0.29 nM but was not pursued further (Wang et al. 2015b). Alternatively, two distal phenyl groups on C=C bond were joined to give tricyclic alkene 12e which exhibited similar potency (EC50 0.28 nM) (Wang et al. 2017a).
An array of piperidine alkene fragments were also incorporated into the 2-methyl-2-oxo-carboxamide chemotype and the example 12f displayed an EC50 < 1 μM (Category A) (Wang et al. 2009c).
Region E covers all the variations which span over the piperazine benzamide moiety, but which could not be described by the definitions of Region A–D. The structures in this classification were very diverse, as shown in Fig. 4.13. Compound 13a was made from the reduction of C=C bond of the corresponding alkene derivative, whilst 13b was synthesized via the cyclopropanation of its alkene counterpart (Wang et al. 2016). Analogue 13c was the regioisomer of compound 11b, with a phenyl ring appended at the 3-position (Wang et al. 2017b). 13a, 13b and 13c all maintained sub-nM potency (EC50: 0.2 nM, 0.75 nM and 0.92 nM) respectively, even though 13c was 45-fold weaker than 11b. Compound 13d contains a homo-piperidine linker rather than a piperazine or piperidine and retained potency with a double-digit pM EC50 (0.09 nM) (Wang et al. 2017b).
Another team reported data for the phenyl ketone analog 13e, which was devoid of the nitrogen of the benzamide. 13e was tested against JR-CSF, B41, and HxBc2 pseudo typed HIV-1, showing IC50s of 3 nM, 7 nM, and 9 nM, respectively (Tuyishime et al. 2016). The significance of 13e was that it had multiple freely rotatable C–C bonds and enjoyed much greater entropy than the benzamide analogs. Due to this result, the investigators advocated that there was opportunity for the next-level innovations beyond modifications of the glyoxamide moiety.
4.3 Alternate Chemotypes for Targeting gp120 Unrelated to Keto Amides
Several betulinic acid derivatives with broad subtype coverage were reported to target gp120 (Lai et al. 2008). Structure 14a (Fig. 4.13) inhibited 25 HIV-1 clinical isolates of clades A, B, and C with an average IC50 of 506 nM. 14a was proposed to bind to the V3 loop of gp120.
Another report described the gp120 antagonist 14b, which displayed IC50 values of 63–170 nM against a panel of 56 HIV-1 pseudoviruses, covering clades A, A/D, A2/D, CRF01_AE, CRF02_AG, CRF13_cpx, B, C, D, and D/A. 14b appeared to possess pan-subtype coverage (Fig. 4.14) (Curreli et al. 2018, 2020).
The CD4 mimics (CD4mcs), exemplified by JP-III-48 (14c) and BNM-III-170 (14d), exhibited somewhat complex virology data. They interacted with gp120 trimers with biphasic dose-response curves, showing dose-dependent activation and inactivation of HIV-1 infection. The proposed mechanism of action of these compounds was that gp120 was activated and could bind to receptor when one CD4mc was bound to one gp120 monomer within the trimer, but was inactivated when three CD4mcs occupied three cavities of one Env trimer. In addition to those two scenarios, when two CD4mcs complexed with one Env trimer, the infection of CD4+ CCR5+ T-cells was inhibited, but the infection of CD4− CCR5+ T-cells was enhanced (Madani et al. 2017). Nevertheless, in the single-round pseudotype assay against YU, JRFL, AD8, and AMLV HIV-1 envelopes, 14c and 14d, respectively, exhibited IC50 values of 1.3/1.9 μM, 30.6/13.3 μM, 6.7/6.4 μM and >100/>100 μM. Recently, (S)-MCG-IV-210 (14e) was uncovered as a new class of CD4mc with a profile distinct from the 14c/14d series (Ding et al. 2019; Grenier et al. 2020).
4.4 Application of HIV gp120 Inhibitors
4.4.1 As Microbicides
The best possible course against any viral infection is prevention, fending off the virus’s invasion at the earliest possible step. General types of prevention include vaccines, microbicides, or drug treatments (Pre-exposure Prophylaxis, or PrEP). An effective HIV-1 vaccine has not been realized and remains a strategy of great interest. PrEP, uses HIV drugs to prevent HIV infection. Daily oral pill regimens are approved for PrEP and recently, monthly intramuscular injections of cabotegravir have shown superiority over daily pills in two clinical studies, HPTN 083 and HPTN 084 (https://www.hptn.org/news-and-events/press-releases/hptn-083-study-demonstrates-superiority-cabotegravir-prevention-hiv, https://www.hptn.org/news-and-events/press-releases/hptn-084-study-demonstrates-superiority-of-cab-la-to-oral-tdfftc-for). HIV Microbicide is another effective approach to preventing HIV infection. It is known that HIV-1 crosses the human body surface barriers and that uptake occurs by the Langerhans cells in the epithelium and by dendritic cells in the stroma. The interactions between the host cells and virus involve HIV-1 gp120. It was shown HIV AIs’ ability to block HIV-1’s uptake by skin cells (Nuttall et al. 2007), and, recent in vitro models using on an HIV AI from the BMS series, BMS-‘793 (12a), illustrated the potential of this class of molecules to be used in PrEP. (Herrera et al. 2021).
Initial PrEP studies used BMS-‘806, which was evaluated for its ability to protect macaques from vaginal SHIV infection, alone or with C52L (a gp41 inhibitor) and/or CMPD167 (a CCR5 antagonist), which were two other mechanistically distinct types of HIV entry inhibitors. BMS-‘806 demonstrated moderate efficacy when used alone, 6 out 8 animals avoided infection when BMS-‘806 was dosed at 5.5 mM. The efficacy was dramatically improved in combinations, with 100% protection (three animals) when 5.5 mM of BMS-806 was mixed with CMPD167 (5.0 mM) and C52L (1.5 mM) (Veazey et al. 2005).
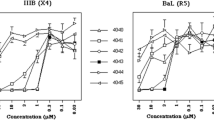
Results of studies using BMS-‘793 (Schader et al. 2012), showed it to be a much more potent analogue of BMS-‘806 as a potential microbicide, either alone or in combination with other types of HIV entry inhibitors (C52L, CMPD167 and CXCR4 ligand AMD3465), against an array of subtypes A-G clinical isolates (25 primary R5, 12 X4 and 7 R5X4 strains). Replication in human donor lymphocytes was inhibited 84% with R5 isolates, 58% with X4 isolates and 14% with R5X4 isolates, by BMS-‘793 at a concentration of 1 μM. When BMS-‘793 was used in combinations with C52L or CMPD167 or AMD3465, 100% inhibition was consistently achieved (Ketas et al. 2007). Additional studies reported inhibition of dendritic cell-driven HIV-1 infection, the main mechanism for HIV-1 mucosal transmission, using BMS-‘793, T1249 (a gp41 inhibitor) and CMPD167. Under all the experimental conditions, BMS-‘793 appeared to be the strongest inhibitor. In particular, BMS-‘793 was significantly more potent than CMPD167 in preventing infection of DCs (Frank and Robbiani 2011). More recent studies in tissue explant and trans-infection models in combination with a fusion inhibitor confirmed it’s potential as an agent for PrEP (Herrera et al. 2021). Currently, BMS-793 is in development by the International Partnership for Microbicide (IPM) as a microbicide candidate in the form of a vaginal tablet. Its Phase I study (IPM 042), a randomized, double-blinded, dose-escalating trial of HIV-negative healthy female participants, to evaluate safety, pharmacokinetics (PK) and pharmacodynamics (PD) was completed.
A molecule of another class of HIV AIs, CD4mc derivative JP-III-48, was evaluated for its ability to prevent HIV-1 vaginal transmission in bone marrow-liver-thymus (BLT) humanized mice. JP-III-48 was capable of inhibiting 100% of HIV-1 infections in experiments where 300 μM of JP-III-48 and JR-CSF virus were applied together. At a higher concentration of 3000 μM, 100% inhibition was achieved when JP-III-48 was administered 30 min prior to the HIV-1 challenge (Princiotto et al. 2018).
The same team also challenged monkeys with a simian-human immunodeficiency virus (SHIV) via the intrarectal route to assess protection by BNM-III-170, another CD4mc. BNM-III-170 had been shown to bind to Env trimer and induced conformational changes which exposed epitopes recognized by non-bNAbs. The immuno-protection in the cohort of gp120 (as immunogen) plus BNM-III-170 was 100% at week 14 and 75% at week 18, while the placebo group of gp120 plus DMSO provided 0% protection at week 9 (Madani et al. 2019).
4.4.2 As Target Recognition Element in Conjugates
The concept of using small molecule warheads to direct antibody recruiting molecules (ARM) to gp160-expressing cells has been explored to assess the potential to treat HIV-1 infection. HIV AIs were tethered to antibody recruiting elements such as a dinitrophenyl (DNP) group. Anti-DNP antibodies, which naturally occur at preexisting concentrations in the blood stream, were directed to gp160-bound viral particles and cells. Compound 15a was able to disrupt the interaction between HIV gp120 and a CD4+ T-cell, and to kill gp120-expressing cells via complement-dependent cytotoxicity (CDC). At 30 μM, compound 15a (Fig. 4.15) was capable of killing ~20% of HIV-Env-expressing CHO cells (Parker et al. 2009). Later, the second generation ARM 15b eliminated ~30% and ~70% of HIV-1 Env (JRFL)-expressing HEK293T cells at a concentration of 30 μM and 60 μM, respectively (Parker et al. 2014). It appears a more robust immune response will be needed to make this approach viable.
An alternative approach to recruiting antibodies used covalently linked HIV AIs (e.g., BMS-‘806 and BMS-‘043) and mAb 38C2 Fabs to generate multi-valent gp120 binders. Exploration of linking strategies determined that a 7-position linker strategy such as that used in the BMS-‘043 derived programmed antibody 15c performed best, with an IC50 value of 128.6 nM in a single-round neutralization assay using U87.CD4.CCR5 cells and the HIV-1 JRFL strain. The research demonstrated the ability of HIV AIs to be used to create bivalent binders. The authors stated that further assessment of 15c was warranted since the approach was also designed to use the small molecule for localization but have the conjugate provide improved properties such as pharmacokinetics and half-life since it was anticipated those properties would be defined by the antibody. Furthermore, Fc engineering of the antibodies could optimize the conjugates ability to potentially elicit antibody-dependent cellular cytotoxicity (ADCC), antibody-dependent cellular phagocytosis (ADCP) or complement dependent cytotoxicity (CDC), to enhance specific destruction of infected cells (Sato et al. 2013).
Another approach to selectively kill HIV infected cells was to link cell killing or cytotoxic agents to HIV AIs. HIV AIs were used to direct the toxic agent to gp160-expressing cells, which was expected to provide selective killing of HIV infected cells (Spiegel and Parker 2017). One example 15d demonstrated a consistent ~10% gp160-selective toxicity in an assay using a single addition of drug, while multiple additions increased gp160-selective toxicity to ~35%. Given the excellent safety of current HIV regimens, considerable progress will need to be made if cell killing agent delivery approaches are to become viable.
4.4.3 As Immuno-Protectants or Immuno-Stimulators
It is known that not all HIV-1 viral particles are infectious. The infectious viral particles (VPs) carry out their replication by invading human CD4+ T-cells and producing off-spring. The defective, non-infectious viral like particles (VLPs) that constitute the majority of gp120 particles also interact with uninfected CD4+ T-cells, CD8+ T-cell and NK cells. The cytotoxicity of gp120 in these interactions manifests itself in multiple ways. For example, binding of VLPs to some cells can induce apoptosis or initiate ADCC effects against these immune cells. This killing of what appear to be bystander, non-HIV infected CD4+ T-cells contributes to the diminished CD4 counts found in HIV-1 patients.
Early work showed VLPs induced the loss of CD4+ T-cells. VLPs with Env initiated CD4+ T-cell apoptosis in a dose-dependent manner, while VLPs without Env had no effect on apoptosis. Because HIV AIs bind to gp120, regardless of whether it is on VPs or VLPs, AIs have the capacity to inhibit bystander killing of uninfected CD4+ T-cells. In one study, the potency against the NL4-3 strain and the anti-apoptotic potency against VLPs packaged with NL4-3 Env of HIV were evaluated with entry inhibitors BMS-‘806, BMS-‘043, soluble CD4 (sCD4), AMD-3100 (a competitive inhibitor of gp120 binding to the co-receptor CXCR4) and T-20 (a peptide anti-HIV fusion inhibitor) (Alexander et al. 2009). BMS-‘806 and BMS’-043 demonstrated similar EC50 values in both the antiviral (2.9 nM and 17.4 nM) and anti-apoptotic (5.8 nM and 11.7 nM) assays. In comparison, sCD4 had an eightfold difference (12.4 nM vs. 98.1 nM) and T-20 showed a 24-fold divergence (22.4 nM vs. 548 nM), whilst AMD-3100 displayed equal potencies (10.8 nM vs. 8.8 nM). The conclusions were that HIV AIs are potent inhibitors of T-cell deletion, and secondly that the IV AI’s antiviral and anti-apoptotic effects may share a common mechanism, presumably by preventing binding of the gp120 protein to its cell surface CD4 receptor.
Immunologically, gp120 can also deregulate an immune response, as for example by blocking T-Cell activation and contributing to viral escape. One of the main mechanisms of HIV’s immune evasion is to acquire mutations in gp120 that lead to heterogenic and interconverting conformations not easily recognized by antibodies. It has also been postulated that mutations in gp120 that reduce susceptibility to HIV attachment inhibitors may stabilize epitopes and enhance antibody neutralization. This hypothesis was explored in relatively early work on AIs and showed mutations selected by HIV AI treatments could increase susceptibility of certain HIV variants to broadly neutralizing antibodies (Zhou et al. 2010). Three escape mutants of LAI virus selected by an early HIV AI contained one of the following mutations: F423Y, an amino acid located in gp120-CD4 binding pocket, and I595F or K655E, both of which are found in gp41 ectodomain. As anticipated, BMS-‘043 displayed a loss in potency against the three mutants with EC50 values of 10.9 nM (F423Y), 19.0 nM (I595F), and 6.06 nM (K655E), compared to 0.98 nM for the wild-type virus. The potency of two bNAbs antibodies, 2F5 and 4E10, were increased by >9-fold against these mutant viruses. The EC50 values for inhibition by 2F5 were 1.88 μg/mL, 0.06 μg/mL, 0.22 μg/mL and 0.02 μg/mL against the wild-type, mutant F423Y, mutant I595F and mutant K655E, respectively. The EC50 values for inhibition by 4E10 were 9.29 μg/mL, 0.72 μg/mL, 0.27 μg/mL and 0.22 μg/mL against the wild-type, F423Y, I595F and K655E, respectively. As controls, HIV entry inhibitors AMD-3100 and Enfuvirtide (T-20) did not show any differential inhibition against the variant viruses.
Recent studies by another team investigated the effects of combining temsavir and anti-HIV-1 antibodies (1:1 molar ratio) against a HIV ADA pseudotyped virus (Zhang et al. 2019). Synergy was observed between temsavir and 5 out of 7 CD4 binding site (CD4bs)-targeting bNAbs. The CD4bs-targeting bNAbs combinations were able to maintain IC50 values under 0.45 nM. Moreover, all the bNAbs, including those which targeted gp120’s CD4 binding site, gp120’s V1V2 segment, gp120’s V3 loop, and gp41-gp120 interface, were potent vs. virus containing temsavir’s key escape mutants, such as M426L, S375M, M426L/M434I, M426L/M475I and S375M/M434I. For example, the S375M/M434I mutations decreased temsavir’s IC50 from 0.14 nM to 29.58 nM. All eleven bNAbs demonstrated IC50 values against the S375M/M434I variant with values of 0.001–0.23 nM.
Mechanistically, HIV AIs are now believed to stabilize the closed conformation of gp120, which prevents access to gp120 conformations permissive for binding to the CD4 receptor. The closed conformation is also thought to be the most efficient conformation for induction and binding of bNAbs. The open conformation of gp120 trimer, which is permissible for binding to CD4 receptor on cells, also is more susceptibility to the binding of non-neutralizing Abs. The binding of non-neutralizing Abs could compete with bNAb binding, effectively inhibiting neutralization. Recent studies show how temsavir constrains the conformation change of the gp120 trimer when presented either in a soluble form or a membrane-bound mode. The stabilized Env trimers remained constrained in the pre-CD4 binding closed form and the immunogenicity of select Env immunogens were improved (Alam et al. 2020).
In contrast, the CD4mc inhibitors bound to gp120 induce the Env to convert to the open form and uncover conserved regions for non-neutralizing Abs to bind. These non-neutralizing antibodies trigger ADCC-mediated killing of HIV-1 infected cells (Richard et al. 2015, 2017). For instance, JP-III-48 (14c) increased the percentage of ADCC-mediated killing of wild-type HIV-1 infected primary CD4+ T-cells from ~2 to 8% (Richard et al. 2015). In a FACS-based ADCC assay of killing infected p24+ cells, at 50 mM concentration, BNM-III-170 (14d) and (S)-MCG-IV-210 (14e) enhanced the ADCC effect from 5% to 14% and 10%, respectively (Ding et al. 2019; Grenier et al. 2020). The key questions to be addressed in this area of research is whether enhanced levels of ADCC-mediated cell killing can be attained and will they be sufficient to provide a physiologically relevant impact.
4.4.4 As Agents Against Cytopathic and Non-cytopathic Effects of gp120
The cytopathic effects of gp120 are not specific towards CD4+ T-cell and other immune cells, as the effects are more widespread (Levy 2007). Neuronal cells, tubular epithelial cells, cardiac myocytes and endothelial cells, among others, can be damaged in the presence of bystander effects elicited by gp120 binding to CD4 present on many different cell types.
There are also numerous deleterious effects reported within cells that express gp160. Among the effects attributed to its expression are the release of calcium on mucosal surfaces, compromising integrity and enhancing monocyte migration across the blood-brain barrier, induction of pro-inflammatory cytokine release and blocking the endogenous GHRH receptor. Pathophysiological effects attributed to HIV include dementia, nephropathy, cardiomyopathy, enteropathy encephalitis, pain, and wasting, all of which may be observed in HIV-1 infected patients and may be exacerbated by the expression of gp160 (Levy 2007).
The ability of HIV AIs to bind to accessible gp160 expressed on envelope or surfaces provides considerable potential for drugs with this mechanism to improve the quality of life. For instance, the protection of neuronal cell apoptosis induced by HIV VLP was reported. In the study, two neuronal cell lines, SH-SY5Y and BE(2)-M17, were CD4−, CCR5+ and CXCR4+, which suggested that virus could not infect these cells due to lack of CD4. BMS-‘806, BMS-‘043, sCD4, TAK779 (a maraviroc-like molecule that binds to CCR5) and T-20, all HIV entry inhibitors, were assessed against the impact of HIV-1 VLP’s cell killing in the SH-SY5Y and BE(2)-M17 cells. EC50s of BMS-‘806 and BMS-‘043 were 14.7 nM and 11.3 nM against SH-SY5Y killing and 17.5 nM and 10.6 nM against BE(2)-M17 killing, respectively. sCD4 had a similar protecting capacity, whilst TAK779 and T-20 showed no effect. It was concluded that HIV gp120 was capable of interacting with and eliminating neuronal cells in a CD4-independent manner. The mechanism is believed to be distinct from that involved in the viral entry process. Thus, it was postulated that HIV AIs may protect neuronal cells from gp120’s cytopathic effects (Zhang et al. 2010). The actual benefits of fostemsavir in reducing the effects caused by VLPs or noninfective HIV particles, if clinically significant, should become more evident with expanded clinical experience and use.
4.5 Future Directions
Fostemsavir is the culmination of considerable preclinical optimization and sustained development efforts. An extended-release formulation of this prodrug enabled the twice daily dosing schedule (BID), which was ultimately pursued to maximize the chances of success in development for HTE patients, despite the fact that early clinical trials demonstrated the potential feasibility of daily (QD) dosing. Critical for its continued development has been the excellent safety profile displayed by fostemsavir and the optimization of the process chemistry and manufacturing processes. Substantive efforts were made to identify a HIV attachment inhibitor with even better properties but at this point in time, none have been disclosed to be in active development (Brown et al. 2013).
A hypothetical next generation AI could offer expanded utility to patients if ever pursued and realized. Once daily (QD) oral dosing or long-acting injectable (LAI) molecules could offer decreased frequency of dosing. Increasing the intrinsic t1/2 or potency beyond that of temsavir would help achieve these goals as might additional application of slow release technologies.
In addition, the selected dose of AIs is dependent on covering the minimum effective concentration of as broad a range of HIV-1 virus clades and subtypes as feasible. Since gp120 envelope possesses sequence variability in a number of regions, pre-existing heterogeneity has been reported to result in a wide range of susceptability (Soulie et al. 2013; Charpentier et al. 2012; Fofana et al. 2015; Alessandri-Gradt et al. 2018; Lepore et al. 2020; Bouba et al. 2020). However, in a recently published study using a Phenosense assay, 1337 individual envelopes encompassing 20 different HIV-1 subtypes were examined for their susceptibility to temsavir (Gartland et al. 2021). While there was variability in susceptibility observed within all subtypes, against the majority of tested viruses, temsavir was highly potent, with most viruses exhibiting IC50s < 10 nM. An exception was CRF01_AE viruses, where all five isolates examined exhibited IC50s > 100 nM. The spectrum and potency of temsavir is broad but because it was originally optimized vs. the HIV-1 B subtype, it does not cover every subtype such as, noted above, genotype CRF01_AE. Gp120 is heavily and nonuniformly glycosylated and is highly flexible and conformationally mobile. It is possible that a recently emerging better understanding of the exact mode of binding to gp120 and its ligands will help in further refining potential future generations of gp120 binders. For example, X-ray co-crystal structures of a number of AI analogs including gp120 + BMS-‘806, gp120 + temsavir and gp120 + BMS-‘251 have recently been reported, which may result in new insights for rational design improvements (Marie Pancera et al. 2017; Lai et al. 2019). Moreover, one of the analogs, BMS-‘251 and temsavir were screened against 208 HIV-1 strains in a cellular assay, with BMS-251’s potency enhanced versus some less susceptible HIV-1 viruses from clades A, CRF01_AE, CRF02_AG, B, CRF07_BC, C, and G (Lai et al. 2019). The IC50 geometric means were >40 nM for temsavir and <2 nM for BMS-‘251. Specifically for the CRF01_AE subtype, BMS-‘251’s IC50s against four isolates were 730 nM, 89 nM, 471 nM and 32 nM, in comparison with temsavir’s >20,000 nM, 6210 nM, > 20,000 nM and 5840 nM, respectively (Lai et al. 2019). The study demonstrates that improving the pan-subtype coverage is achievable in vitro if it could be incorporated into a molecule with an excellent drug-like profile. HIV AIs with an expanded pan-subtype coverage would potentially enable a better microbicide for HIV prevention in the absence of vaccines but realistically only if the synthesis was much shorter and less complex than that of fostemsavir. If a long-acting injectable formulation of an AI were discovered, it is possible it could find use in PrEP regimens.
Considering the recent approval of fostemsavir, there has not been time yet for definitive studies looking to see if fostemsavir will have at clinically meaningful positive additive or synergistic effects on the actions of therapeutic antibodies or bNAbs. Positive data in this arena may suggest a potential role in regimens moving closer to realizing a functional cure. The potential for using AIs for targeting and eradicating HIV infected cells has been demonstrated so if further optimization of the conjugation strategies for delivery of immunological or other specific mediators of killing those cells are realized, they could also potentially be useful in future cure regimens (Sloan et al. 2015).
4.6 Conclusions
The discovery of agents that target the HIV viral envelope gp120 has been fruitful and there is still potential for further treatment advances. The recent approval of fostemsavir (RUKOBIA™) has provided an important new option in the treatment of HTE HIV patients and clinical experience and expanded usage should further inform about potential other benefits in the future. In addition to AIs, CD4 mimics, agents that target other regions of gp120, and conjugates of these small molecules designed to elicit specific killing of HIV selected cells all offer the potential for utility in regimen’s that deplete reservoir or move closer to realizing HIV cure.
References
Alam SM, Cronin K, Parks R, Anasti K, Ding H, Go EP, Desaire H, Eaton A, Montefiori D, Sodroski J, Kappes J, Haynes BF, Saunders KO (2020) J Virol 94:e00958
Alessandri-Gradt E, Charpentier C, Leoz M, Mourez T, Descamps D, Plantier J-C (2018) J Antimicrob Chemother 73:2716
Alexander L, Zhang S, McAuliffe B, Connors D, Zhou N, Wang T, Agler M, Kadow JF, Lin P-F (2009) Antimicrob Agents Chemother 53:4726
Bender JA, Yang Z, Eggers B, Gong Y-F, Lin P-F, Parker DD, Rahematpura S, Zheng M, Meanwell NA, Kadow JF (2013) Bioorg Med Chem Lett 23:218
Bouba Y, Berno G, Fabeni L, Carioti L, Salpini R, Aquaro S, Svicher V, Federico Perno CF, Ceccherini-Silberstein F, Santoro MM (2020) J Antimicrob Chemother 75:1778
Brown J, Chien C, Timmins P, Dennis A, Doll W, Sandefer E, Page R, Nettles RE, Zhu L, Grasela D (2013) J Pharm Sci 102:1742
Charpentier C, Larrouy L, Visseaux B, Roland Landman R, Marine Levittas M, Storto A, Damond F, Yazdanpanah Y, Yeni P, Brun-Vezinet F, Descamps D (2012) J Antimicrob Chemother 67:1459
Curreli F, Belov DS, Kwon YD, Ramesh R, Furimsky AM, O’Loughlin K, Byrge PC, Iyer LV, Mirsalis JC, Kurkin AV, Altieri A, Debnath AK (2018) Eur J Med Chem 154:367
Curreli F, Ahmed S, Victor SMB, Iusupov IR, Belov DS, Markov PO, Kurkin AV, Altieri A, Debnath AK (2020) J Med Chem 63:1724
Ding S, Grenier MC, Tolbert WD, Vezina D, Sherburn R, Richard J, Prevost J, Chapleau J-P, Gendron-Lepage G, Medjehed H, Abrams C, Sodroski J, Pazgier M, Smith AB III, Fenzi A (2019) J Viol 93:e01325–e01319
Fenwick DR, Middleton DR, Stephenson PT, Tran T-D, Williams DH (2005) WO-2015111094, 22 Dec 2005
Fofana DB, Charpentier C, Maıga AI, Lambert-Niclot S, Sayon S, Desire N, Simon A, Yazdanpanah Y, Katlama C, Descamps D, Calvez V, Marcelin AG, Soulie C (2015) J Antimicrob Chemother 70:130
Frank I, Robbiani M (2011) J Acquir Immune Defic Syndr 56:204
Gartland M, Zhou N, Stewart E, Pierce A, Clark A, Ackerman P, Llamoso C, Lataillade M, Krystal M (2021) J Antimicrob Chemother 76:648
Grenier MC, Ding S, Vezina D, Chapleau J-P, Tolbert WD, Sherburn R, Schon A, Somisetti S, Abrams C, Pazgier M, Fenzi A, Smith AB III (2020) ACS Med Chem Lett 11:371
Hanna GJ, Lalezari J, Hellinger JA, Wohl DA, Nettles R, Persson A, Krystal M, Lin P-F, Colonno R, Grasela DM (2011) Antimicrob Agents Chemother 55:722
Herrera C, Harman S, Aldon Y, Rogers P, Armanasco N, Ziprin P, Stieh D, Nuttall J, Shattock R (2021) J AIDS. https://doi.org/10.1097/QAD.0000000000002974. Ahead-of-Print
Ho H-T, Fan L, Nowicka-Sans B, McAuliffe B, Li C-B, Yamanaka GA, Zhou N, Fang H, Dicker I, Dalterio R, Gong Y-F, Wang T, Yin Z, Ueda Y, Matiskella JD, Kadow JF, Clapham P, Robinson J, Colonno RJ, Lin P-FJ (2006) Virology 80:4017
Kadow JF, Xue QM, Wang T, Zhang Z, Meanwell NA (2009) U.S. 7,531,552, 12 May 2009
Ketas TJ, Schader SM, Zurita J, Teo E, Polonis V, Lu M, Klasse PJ, Moore JP (2007) Virology 364:431
Kozal M, Aberg J, Pialoux G, Cahn P, Thompson M, Molina J-M, Grinsztejn B, Diaz R, Castagna A, Kumar P, Latiff G, DeJesus E, Gummel M, Gartland M, Pierce A, Ackerman P, Llamoso C, Lataillade M (2020) N Engl J Med 382:1232
Lai W, Huang L, Ho P, Li Z, Montefiori D, Chin-Ho Chen C-H (2008) Antimicrob Agents Chemother 52:128
Lai Y-T, Wang T, Druz A, O’Dell S, Cheung CS, Gwo-Yu Chuang G-Y, Dongjun Peng D, Yang Y, Zhang B, Herschhorn A, Sodroski JG, Doria-Rose NA, Mascola JR, Langley DR, Kwong PD (2019) Nat Commun 10:47
Lataillade M, Lalezari JP, Kozal M, Aberg JA, Pialoux G, Cahn P, Thompson M, Molina J-M, Moreno S, Grinsztejn B, Diaz RS, Castagna A, Kumar PN, Latiff GH, Jesus ED, Wang M, Chabria S, Garland M, Pierce A, Ackerman P, Llamoso C (2020) Lancet HIV 7:e740
Lepore L, Fabrizio C, Bavaro DF, Milano E, Volpe A, Lagioia A, Angarano G, Saracino A, Monno L (2020) J Antimicrob Chemother 75:1580
Levy JA (2007) HIV and the pathogenesis of AIDs. ASM Press, Washington, DC
Lin P-F, Blair WS, Wang T, Spicer TP, Guo Q, Zhou N, Gong Y-F, Wang HH, Rose R, Yamanaka GA, Robinson BS, Li C-B, Kimel BJ, Fang H, Demers G, Deminie C, Fridell RA, Yang Z, Zadjura LM, Meanwell NA, Colonno RJ (2003) Proc Natl Acad Sci U S A 100:11013
Lu R-J, Tucker JA, Zinevitch T, Kirichenko O, Konoplev V, Kuznetsova S, Sviridov S, Pickens J, Tandel S, Brahmachary E, Yang Y, Wang J, Freel S, Fisher S, Sullivan A, Zhou J, Stanfield-Oakley S, Greenberg M, Bolognesi D, Bray B, Koszalka B, Jeffs P, Khasanov A, Ma Y-A, Jeffries C, Liu C, Proskurina T, Zhu T, Chucholowski A, Li R, Sexton C (2007) J Med Chem 50:6535
Lu R-J, Tucker JA, Pickens J, Ma Y-A, Zinevitch T, Kirichenko O, Konoplev V, Kuznetsova S, Sviridov S, Brahmachary E, Khasanov A, Mikel C, Yang Y, Liu C, Wang J, Freel S, Fisher S, Sullivan A, Zhou J, Stanfield-Oakley S, Baker B, Sailstad J, Greenberg M, Bolognesi D, Bray B, Koszalka B, Jeffs P, Jeffries C, Chucholowski A, Sexton C (2009) J Med Chem 52:4481
Madani N, Princiotto AM, Zhao C, Jahanbakhshsefidi F, Mertens M, Herschhorn A, Melillo B, Smith AB III, Sodroski J (2017) J Viol 91:e01880–e01816
Madani N, Princiotto AM, Mach L, Ding S, Prevost J, Richard J, Hora B, Sutherland L, Zhao CA, Conn BP, Bradley T, Moody MA, Melillo B, Finzi A, Haynes BF, Smith AB III, Santra A, Sodroski J (2019) Nat Commun 10:47
Marie Pancera M, Lai Y-T, Bylund T, Aliaksandr Druz A, Narpala S, O’Dell S, Arne Schön A, Bailer RT, Chuang G-Y, Geng H, Louder MK, Rawi R, Soumana DI, Finzi A, Herschhorn A, Madani N, Sodroski J, Freire E, Langley DR, Mascola JR, McDermott AB, Kwong PD (2017) Nat Chem Biol 13:1115
Markham A (2020) Drugs 80:1485
Meanwell NA, Wallace OB, Wang H, Deshpande M, Pearce BC, Trehan A, Yeung K-P, Qiu Z, Wright JJK, Robinson BA, Gong Y-F, Wang H-GH, Blair WS, Shi P-Y, Lin P-F (2009a) Bioorg Med Chem Lett 19:5136
Meanwell NA, Wallace OB, Wang H, Milind Deshpande M, Pearce BC, Trehan A, Yeung K-S, Qiu Z, Wright JJK, Robinson BA, Gong Y-F, Wang H-GH, Blair WS, Shi P-Y, Pin-Fang Lin P-F (2009b) Bioorg Med Chem Lett 19:5136
Meanwell NA, Wang T, Zhang Z, Yin Z, Hamann LG, Kadow JF (2014) U.S. 8,685,982, 1 April 2014
Meuser ME, Murphy MB, Rashad AA, Cocklin S (1940) Molecules 2018:23
Nettles RE, Schurmann D, Zhu L, Stonier M, Huang S-P, Chang I, Chien C, Krystal M, Wind-Rotolo M, Ray N, Hanna GJ, Bertz R, Grasela D (2012) J Infect Dis 206:1002
Nowicka-Sans B, Gong Y-F, McAuliffe B, Dicker I, Ho H-T, Zhou N, Eggers B, Lin P-F, Yamanaka GA, Ray N, Wind-Rotolo M, Zhu L, Majumdar A, Stock D, Lataillade M, Hanna GJ, Matiskella JD, Ueda Y, Wang T, Kadow JF, Meanwell NA, Krystal M (2012) Antimicrob Agents Chemother 56:3498
Nuttall J, Romano J, Douville K, Galbreath C, Nel A, Heyward W, Mitchnick M, Walker S, Rosenberg Z (2007) Infect Dis Clin N Am 21:219
Parker CG, Domaoal RA, Anderson KS, Spiegel DA (2009) J Am Chem Soc 131:16392
Parker CG, Dahlgren MK, Tao RN, Li DT, Douglass EF Jr, Shoda T, Jawanda N, Spasov KA, Lee S, Zhou N, Domaoal RA, Sutton RE, Anderson KS, Krystal M, Jorgensena WL, Spiegel DA (2014) Chem Sci 5:2311
Princiotto AM, Vrbanac VD, Melillo B, Park J, Tager AM, Smith AB III, Sodroski J, Madani N (2018) J Infect Dis 218:471
Regueiro-Ren A, Xue QM, Kadow JF, Taylor M (2004) WO 2004/011425, 5 Feb 2004
Regueiro-Ren A, Xue QM, Kadow JF, Taylor M (2005) U.S. 6,900,206, 31 May 2005
Richard J, Veillette M, Brassard N, Lyer SS, Roger M, Martin L, Pazgier M, Schon A, Freire E, Routy J-P, Smith AB III, Park J, Jones DM, Courter JR, Melillo BN, Kaufmann DE, Hahn BH, Permar SR, Haynes BF, Madani N, Sodroski J, Finzi A (2015) Proc Natl Acad Sci U S A 112:E2687
Richard J, Ding S, Finzi A (2017) AIDS Res Ther 14:42
Sato S, Inokuma T, Otsubo N, Burton DR, Barbas KF III (2013) ACS Med Chem Lett 4:460
Schader SM, Colby-Germinario SP, Quashie PK, Oliveira M, Ibanescu R-I, Moisi D, Mesplede T, Wainbeig MA (2012) Antimicrob Agents Chemother 56:4257
Sloan DD, Lam C-YK, Irrinki A, Liu L, Tsai A, Pace CS, Kaur J, Murry JP, Balakrishnan M, Moore PA, Johnson S, Nordstrom JL, Cihlar T, Koenig S (2015) PLoS Pathog 11:1
Soulie C, Lambert-Niclot S, Bocar Fofana DB, Fourati S, Ait-Arkoub Z, Sayon S, Simon A, Christine Katlama C, Calvez V, Marcelin A-G (2013) J Antimicrob Chemother 68:1243
Spiegel DA, Parker C (2017) U.S. 9,745,334, 29 Aug 2017
Swidorski JJ, Liu Z, Yin Z, Wang T, Carini DJ, Rahematpura S, Zheng M, Johnson K, Zhang S, Lin P-F, Parker DD, Li W, Meanwell NA, Hamann LG, Regueiro-Ren A (2016) Bioorg Med Chem Lett 26:160
Tuyishime M, Rae Lawrence R, Cocklin S (2016) Bioorg Med Chem Lett 26:228
Veazey RS, Klasse PJ, Schader SM, Hu Q, Ketas TJ, Lu M, Marx PA, Dufour J, Colonno RJ, Shattock RJ, Springer MS, Moore JP (2005) Nature 438:99
Wang T, Zhang Z, Wallace OB, Deshpande M, Fang H, Yang Z, Zadjura LM, Tweedie DL, Huang S, Zhao F, Ranadive S, Robinson BS, Gong Y-F, Ricarrdi K, Spicer TP, Deminie C, Rose R, Blair WS, Lin P-F, Colonno RJ, Meanwell NA (2003) J Med Chem 46:4236
Wang J, Le N, Heredia A, Song H, Redfield R, Wang LX (2005) Org Biomol Chem 3:1781
Wang T, Kadow JF, Meanwell NA, Zhang Z, Yin Z, James CA, Ruediger EH, Pearce BC (2008a) U.S. 7,396,830, 8 July 2008
Wang T, Kadow JF, Meanwell NA, Zhang Z, Yin Z, Yeung K-S, Qiu Z, Deon DH, James CA, Ruedinger EH, Bachand C (2008b) U.S. 7,348,337, 25 March 2008
Wang T, Yin Z, Zhang Z, Bender JA, Yang Z, Johnson G, Yang Z, Zadjura LM, D’Arienzo CJ, DiGiugno Parker D, Gesenberg C, Yamanaka GA, Gong Y-F, Ho H-T, Fang H, Zhou N, McAuliffe BV, Eggers BJ, Fan L, Nowicka-Sans B, Dicker IB, Gao Q, Colonno RJ, Lin P-F, Meanwell NA, Kadow J (2009a) F. J Med Chem 52:7778
Wang T, Kadow JF, Zhang Z, Yin Z, Meanwell NA (2009b) U.S. 7,504,399m 17 March 2009
Wang T, Han Y, Kadow JF, Zhang Z, Meanwell NA (2009c) U.S. 7,572,810, 11 Aug 2009
Wang T, Kadow JF, Zhang Z, Yin Z, Meanwell NA, Regueiro-Ren A, Swidorski JJ, Han Y, Carani DJ (2010) US 7,807,671, 5 Oct 2010
Wang T, Ueda Y, Hamann LG, Zhang Z, Yin Z, Regueiro-Ren A, Carini DJ, Swidorski JJ, Liu Z, Johnson BL, Meanwell NA, Kadow JF (2011) U.S. 7,960,406, 14 June 2011
Wang T, Yang Z, Zhang Z, Yamanaka GA, Lin P-F, Parker DD, Rahematpura S, Mattew M, Zhang M, Meanwell NA, Kadow JF, Bender JA (2013) Bioorg Med Chem Lett 23:213
Wang T, Zhang Z, Yin Z, Kadow JF, Meanwell NA (2014a) U.S. 8,912,195, 16 Dec 2014
Wang T, Yin Z, Zhang Z, Johnson BL, Kadow JF, Meanwell NA (2014b) WO 2014/160689, 2 Oct 2014
Wang T, Zhang Z, Langley DR, Kadow JF, Meanwell NA (2014c) U.S. 8,664,213, 4 March 2014
Wang T, Zhang Z, Kadow JF, Meanwell NA (2014d) U.S. 8,835,454, 16 Sept 2014
Wang T, Zhang Z, Yin Z, Kadow JF, Meanwell NA (2015a) U.S. 9,156,828, 13 Oct 2015
Wang T, Kadow JF, Meanwell NA, Hamann LG (2015b) U.S. 9,193,725, 24 Nov, 2015
Wang T, Zhang Z, Kadow JF, Meanwell NA, Ruediger EH, James CA, Deon DH, Carini DJ, Johnson BL (2016) U.S. 9,505,752, 29 Nov 2016
Wang T, Kadow JF, Meanwell NA, Zhang Z, Yin Z, Ruediger EH, James CA, Deon DH (2017a) U.S. 9,655,888, 23 May 2017
Wang T, Yin Z, Zhang Z, Bender JA, Johnson BL, Kadow JF, Meanwell NA (2017b) U.S. 9,586,957, 7 March 2017
Wang T, Ueda Y, Zhang Z, Yin Z, Matiskella JD, Pearce BC, Bender JA, Yang Z, Yang Z, Zheng M, Parker DD, Yamanaka GA, Gong Y-F, Ho H-T, Gao Q, Colonno RJ, Lin P-F, Meanwell NA, Kadow JF (2018) J Med Chem 61:6308
Wang T, Wallace OB, Zhang Z, Fang H, Yang Z, Robinson BA, Spicer T-P, Gong Y-F, Blair WS, Shi PY, Lin P-F, Deshpande M, Meanwell NA, Kadow JF (2019) Bioorg Med Chem Lett 29:1423
Zhang S, Alexander L, Wang T, Agler M, Zhou N, Fang H, Kadow JF, Clapham P, Lin P-F (2010) Arch Virol 155:777
Zhang Y, Chapman JH, Ulcay A, Sutton RE (2019) J Virol 93:e01446
Zhou N, Fan L, Ho H-T, Nowicka-Sans B, Sun Y, Zhu Y, Hu Y, McAuliffe B, Rose B, Fang H, Wang T, Kadow JF, Krystal M, Alexander L, Colonno RJ, Lin P-F (2010) Virology 402:256
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wang, T., Kadow, J.F., Meanwell, N.A., Krystal, M. (2022). The Genesis and Future Prospects of Small Molecule HIV-1 Attachment Inhibitors. In: Jiang, S., Lu, L. (eds) Virus Entry Inhibitors. Advances in Experimental Medicine and Biology, vol 1366. Springer, Singapore. https://doi.org/10.1007/978-981-16-8702-0_4
Download citation
DOI: https://doi.org/10.1007/978-981-16-8702-0_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-8701-3
Online ISBN: 978-981-16-8702-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)