Abstract
The use of Magnetic Resonance Images (MRI) is a frequently used tool in disease detection. The use of healthcare professionals to examine MRI images and to identify diseases are among traditional methods. Therefore, one way to improve clinical health care is to present and analyze medical images more efficiently and intelligently. Brain tumors can be of different types, and accordingly, they can cause serious health problems in adults and children. Such bulks can occur anywhere in the brain in different sizes and densities. This is not a standardized situation due to its nature. The diagnoses are revealed by the experts by analyzing the tumor images manually. In the proposed model, it is aimed at automating the process and reducing human errors in the system. The model is based on the deep learning technique, which is a probabilistic neural network to identify unwanted masses in the brain. In this study, a model has been created with VGG and CNN (Convolutional Neural Network) architectures, which are among the deep learning techniques. The performance values of the model outputs, accuracy, error rates, and specificity separators are discussed comparatively.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Healthcare professionals rely on medical images for disease detection and treatment when visual inspection is not possible. For this reason, medical images should be examined more efficiently in order to develop this method, which is frequently preferred in health services [1]. On the other hand, the devices and the created images are developed to obtain better quality results. Thus, the analysis of images with smart systems in order to facilitate their use in healthcare services has become a research topic. Physicians and radiologists can obtain more effective results in image analysis with computer-aided systems. Computing and data-based repetitive processes can be automated with computers and have become an important reference source for healthcare professionals [2].
In classical machine learning techniques, the feature extraction process is revealed manually, unlike deep learning methods, and the obtained images can be defined numerically. The manual feature extraction method is not an approach that can be possible in every field and on the other hand the necessary competence in the medical field may not be achieved. In addition, manually extracted features cannot produce effective results in all cases and successful results cannot be obtained in complex scenarios. One possible solution is to learn research-related features directly from medical data. Such a data-driven approach can naturally learn domain information from data without manual feature engineering. However, constraints such as optimization difficulty and hardware limitation are encountered. Since such architectures also lead to the use of shallow architectures, healthier results can be obtained with deep learning models [3].
El Kaitoun et al. have used an improved Markov method and a U-net-based deep learning method for brain tumor detection [4]. Ramirez et al. have presented a new variational model for MRI image segmentation [5]. Sobhaninia et al. have obtained more effective results by using multiple scales and the utilization of two cascade networks as an image segmentation method [6]. Wu et al. have used a 3D U-net-based deep learning model for tumor image segmentation [7]. Chetty et al. have presented a new approach model for classifying brain MRI images based on the 3D U-Net deep learning architecture [17].
The development of computer hardware and artificial learning techniques in recent years has enabled deep learning models on problems. Thus, classification with deep learning models, object perception, and computer vision problems became research topics. In this context, calculations that have transformed from traditional machine learning methods to deep learning methods have gained great momentum. Although deep learning in medical fields has become widespread, there is still much field for improvement. In this study, a model has been developed for the classification problem on brain tumor images with CNN and VGG deep learning methods, which are among the computer vision methods, is presented.
2 Materials and Methods
In this study, a classification model has been created using MR images frequently used by healthcare professionals. With this model, brain tumor symptoms in patients can be detected by computer vision methods. As a data source, publicly available data on Kaggle has been used [8]. The dataset includes 155 tumor patients images separated according to patient data and 98 healthy MR images. Figure 1 contains sample data from the tumor and healthy brain images in the dataset.
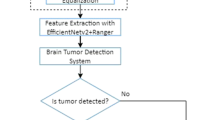
Performance results of classification models created with CNN and VGG architectures, among deep learning techniques, have been compared. Images are rearranged to a fixed pixel size of 224 × 224. The created model is designed with the steps as in Fig. 2.
2.1 CNN and VGG Architecture
A convolutional neural network consists of layers in Fig. 3. The image is processed with the Convolutional Layer, Pooling Layer, Fully Connected Layer, and Output Layer steps, respectively, as indicated in Fig. 4 [9,10,11,12,13,14,15,16].
Each step in which the image is processed consists of the following processes.
Convolutional Layer and Activation Function: It takes place to reveal the features and the non-linearity is introduced to the model through the activation function.
Pooling Layer: This layer is added after convolutional layer and helps to reduce the number of weights in the model. Thus, by reducing the number of parameters in the network, it allows a reduction of computational complexity.
Flattening Layer: Prepares the classical neural network data by making the matrix that consists of convolutional layer and pooling layer steps into one-dimensional array.
Fully Connected Layer: The standard neural network method used in the classification process is applied to the one-dimensional array taken from the pooling layer.
Relu Activation Function: This layer comes into effect after convolutional layers. The main task of this layer is to convert negative values from input data to zero. The model that comes from the operations before this layer has a linear structure and is required to turn it into a nonlinear structure. Thus, the model learns more efficiently [10, 11].
Softmax Activation Function: Softmax is mainly used in the output layer. Calculates the probability that the input belongs to a particular class. This is calculated by generating values between 0 and 1 with a probabilistic interpretation. It is mathematically expressed as in Eq. (1). The input vector x is a real number and consequently, a probability result p is produced [12, 13].
2.2 Model Implementation and Performance Analysis
The model has been implemented with the Python programming language version 3.7 and using the Keras deep learning library [15]. The dataset that used in the study has been divided into two groups as train and test datasets. The rate is 75–25%. First of all, the images are adjusted to a fixed size. Images have been rearranged in 224 × 224 size and thus it has aimed to achieve more effective results. Two different models proposed that based on CNN and VGG architectures have been designed. Layer information and the number of parameters for presented models have been created as shown in Table 1–2.
Model performances have been measured after training with two different models. According to the performance results, since the success rate in model 2 is higher and the loss function is closer to the value of zero, a better classification result has been obtained compared to model 1. As a result, a classification performance result of 92% in model 2 and 85% in model 1 has been obtained. It has been observed that model 2 with a result closer to 0 is more effective. As the performance values obtained from these models, accuracy and loss function graphs have obtained for 50 epochs as in Table 3. Only the accuracy criterion does not give sufficient results in cases where there are unbalanced data sets [14], thus accuracy, precision, and recall values have also been observed as other performance metrics in Table 4.
TP (True positive): Correct detection of the tumor image.
FP (False positive): Incorrect detection of the healthy image.
TN (True negative): Correct detection of the healthy image.
FN (False negative): False detection of the tumor image.
Precision rate (P) = TP / (TP + FP).
Recall rate (R) = TP / (TP + FN).
3 Conclusion
In this study, the performance performances of CNN- and VGG-based proposed models for tumor detection through MRI images, which are frequently used in health care, in classification have been evaluated. The number of convolutional layers, dataset quality, number of epochs can be among the main criteria that can affect the success of the model during training. The number of patient samples may be limited in the training set and thus this situation may lead to over-learning. It has been observed that better results can be obtained with a limited number of images with the VGG-based model, which has a pre-trained architecture for such cases. As a result, it has been observed that pre-trained VGG-based models have high applicability in the health field where data acquisition is limited even if they are trained with objects with different characteristics. It has been observed that the learned features during training can be transferred with high accuracy on different models and it makes such models a viable option for classification problems in the healthcare field.
References
Sangeetha R, Mohanarathinam A, Aravindh G, Jayachitra S, Bhuvaneswari M (2020) Automatic detection of brain tumor using deep learning algorithms. In: Proceedings of 2020 4th International Conference on Electronics, Communication and Aerospace Technology (ICECA), Coimbatore, India, pp 1–4. https://doi.org/10.1109/ICECA49313.2020.9297536
Santos J, Santos dos HDP, Vieira R (2020) Fall detection in clinical notes using language models and token classifier. In: Proceedings of 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS) Rochester, MN, USA, pp 283–288. https://doi.org/10.1109/CBMS49503.2020.00060
Sufri NAJ, Rahmad NA, Ghazali NF, Shahar N, As’ari MA (2019) Vision based system for banknote recognition using different machine learning and deep learning approach. In: Proceedings of 2019 IEEE 10th Control and System Graduate Research Colloquium (ICSGRC) Shah Alam, Malaysia, pp 5–8. https://doi.org/10.1109/ICSGRC.2019.8837068
El kaitouni SEI, Tairi H (2020) Segmentation of medical images for the extraction of brain tumors: a comparative study between the Hidden Markov and deep learning approaches. In: Proceedings of 2020 International Conference on Intelligent Systems and Computer Vision (ISCV), Fez, Morocco, pp 1–5. https://doi.org/10.1109/ISCV49265.2020.9204319
Ramírez I, Martín A, Schiavi E (2018) Optimization of a variational model using deep learning: an application to brain tumor segmentation. 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018) Washington, DC, USA, pp 631–634. https://doi.org/10.1109/ISBI.2018.8363654
Sobhaninia Z, Rezaei S, Karimi N, Emami A, Samavi S (2020) Brain tumor segmentation by cascaded deep neural networks using multiple image scales. In Proceedings of 2020 28th Iranian Conference on Electrical Engineering (ICEE), Tabriz, Iran, pp 1–4. https://doi.org/10.1109/ICEE50131.2020.9260876
Wu P, Chang Q (2020) Brain tumor segmentation on multimodal 3D-MRI using deep learning method. In: Proceedings of 2020 13th International Congress on Image and Signal Processing, BioMedical Engineering and Informatics (CISP-BMEI) Chengdu, China, pp 635–639. https://doi.org/10.1109/CISP-BMEI51763.2020.9263614
Chakrabarty N (2019) Brain mri images for brain tumor detection. Retrieved April 10, 2021, from https://www.kaggle.com/navoneel/brain-mri-images-for-brain-tumor-detection
Basaveswara SK (2019) CNN Architectures, a Deep-dive - towards data science. Medium. https://towardsdatascience.com/cnn-architectures-a-deep-dive-a99441d18049
Stursa D, Dolezel P (2019) Comparison of ReLU and linear saturated activation functions in neural network for universal approximation. In: Proceedings of 2019 22nd International Conference on Process Control (PC19), Strbske Pleso, Slovakia, pp 146–151. https://doi.org/10.1109/PC.2019.8815057
Kirana KC, Wibawanto S, Hidayah N, Cahyono GP, Asfani K (2019) Improved neural network using Integral-RELU based prevention activation for face detection. In: Proceedings of 2019 International Conference on Electrical, Electronics and Information Engineering (ICEEIE), Denpasar, Indonesia, pp 260–263. https://doi.org/10.1109/ICEEIE47180.2019.8981443
Galindo O, Ayub C, Ceberio M, Kreinovich V (2019) Faster quantum alternative to softmax selection in deep learning and deep reinforcement learning. 2019 IEEE Symposium Series on Computational Intelligence (SSCI) Xiamen, China, pp 815–818. https://doi.org/10.1109/SSCI44817.2019.9003167
Alabassy B, Safar M, El-Kharashi MW (2020) A high-accuracy implementation for softmax layer in deep neural networks. 2020 15th Design & Technology of Integrated Systems in Nanoscale Era (DTIS) Marrakech, Morocco, pp 1–6. https://doi.org/10.1109/DTIS48698.2020.9081313
Steiniger Y, Stoppe J, Meisen T, Kraus D (2020) Dealing with highly unbalanced sidescan sonar image datasets for deep learning classification tasks. Global Oceans, 2020 Singapore–U.S Gulf Coast, Biloxi, MS, USA, pp 1–7. https://doi.org/10.1109/IEEECONF38699.2020.9389373
Team K (n.d.) Simple. Flexible. Powerful. Retrieved from https://keras.io/
Allibhai E (2019) Building a Convolutional Neural Network (CNN) in Keras. Medium. https://towardsdatascience.com/building-a-convolutional-neural-network-cnn-in-keras-329fbbadc5f5
Chetty G, Singh M, White M (2019) Automatic brain image analysis based on multimodal deep learning scheme. In: Proceedings of International Conference on Machine Learning and Data Engineering (iCMLDE) 2019, pp 97–100. https://doi.org/10.1109/iCMLDE49015.2019.00028
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Kırelli, Y., Arslankaya, S., Alcan, P. (2022). MRI Image Analysis with Deep Learning Methods in Brain Tumor Diagnosis. In: Sen, Z., Oztemel, E., Erden, C. (eds) Recent Advances in Intelligent Manufacturing and Service Systems. Lecture Notes in Mechanical Engineering. Springer, Singapore. https://doi.org/10.1007/978-981-16-7164-7_4
Download citation
DOI: https://doi.org/10.1007/978-981-16-7164-7_4
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-7163-0
Online ISBN: 978-981-16-7164-7
eBook Packages: EngineeringEngineering (R0)