Abstract
Sleep disorder diseases have one of the major health issues across the world. To handle this issue, the primary step taken by most of the sleep experts is the sleep staging classification. In this paper, we proposed an automated deep one-dimensional convolution neural network (1D-CNN) for multi-class sleep stages through polysomnographic signals. The proposed 1D-CNN model comprises eleven layers with learnable parameters: nine convolution layers and two-fully connected layers. The main objective of designing such a 1D-CNN model is to achieve higher classification accuracy for multiple sleep stage classifications with reduced learnable parameters. The proposed network architecture is tested on two different subgroups subject sleep recordings of ISRUC-Sleep datasets, namely ISRUC-Sleep subgroup-I (SG-I) and ISRUC-Sleep subgroup-III (SG-III). The proposed deep 1D-CNN model achieved the highest classification accuracy of 98.44, 99.03, 99.50, and 99.03% using the ISRUC-Sleep SG-I dataset and 98.51, 98.88, 98.76, and 98.67% using SG-III dataset for two to five sleep stage classification, respectively, with single channel of EEG signals. It has been observed that the obtained results from the proposed 1D-CNN model give the best classification accuracy performance on multiple sleep stage classifications incomparable to the existing literature works. The developed 1D-CNN deep learning architecture is ready for clinical usage with high PSG data.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Nowadays, it has been observed that the neurocognitive system directly decides the mental and cognitive performance in a particular task [1]. It’s very difficult to determine the subjects neurocognitive performance (NCP) very accurately either numerically or any standard evaluation procedures. Currently, NCP is an open challenge in the medical domain concerning different diseases such as neurology disorder, rehabilitation, and psychology-related disorders. It’s also very difficult to assess with a scenario of changes NCP with a known predictable manner. These types of diseases are more challenging concerning analysis and to get proper diagnosis solutions. Different sleep-related diseases are considered under these changes NCP cases since the brain-behavior changes according to different stages of sleep [1]. Sleep is one of the important ingredients for good human health and also responsible for maintaining the fitness and functioning of the different core systems of our body. It also puts an impact on our proper functioning of mental and cognitive systems.
For human life, a total of one-third of its duration is constituted of the sleep cycle. It has been observed from several studies that sleep deficiency causes so many consequences like inability to solve the problem, not able to make proper decisions, not controlling the emotions, and reflected several changes in people [2, 3]. Sometimes the improper quality of sleep influenced different types of sleep-related disorders such as sleep apnea, insomnia, depression, narcolepsy, hypersomnia, breathing-related disorders, and circadian rhythm disorders [4]. Sometimes it has been seen that sleep deprivation is considered as stress-related disorders or sleep pathology, which causes high risk in performing some common cognitive risks such as workplace incidents, road accidents happened [5]. According to a report of the National Highway Traffic Administration of USA, due to drowsiness, around one lakh car accidents happened, as consequences more than 1500 death cases resulted and injuries cases reported around 71,000 annually [6]. In this scenario, proper analysis of sleep stages is very important for identifying the sleep-related irregularities. So that it is very essential to analyze the sleep stage's behavior and accurate scoring of sleep states is a very crucial segment of the sleep staging process.
The polysomnography test is the primary step for any types of sleep-related disorders. It is a combination of a different physiological signal which is useful during analyzing the sleep patterns of an individual subject. Several polysomnographic recordings are included such as the electroencephalogram (EEG) signal, the electrooculogram (EOG) signal, and electromyogram (EMG) signals have tracked the changes in behavior muscle tone. The entire sleep staging process is generally conducted through visualizing the sleep patterns of the subject during sleep periods by well-trained sleep domain technicians according to the Rechtschaffen and Kales (R and K) [7] and the AASM sleep standards [8].
This traditional way of monitoring sleep stages methods has so many disadvantages such as requires more sleep experts to monitor the sleep recordings, time-consuming, and erroneous [9]. Due to more human interpretations during recording, it may not report good classification accuracy in the diagnosis of sleep stage classification [10]. Based on the above-mentioned drawbacks, automated classification of sleep stages is introduced, which ultimately gives benefits for quick diagnosis and also reported with increases of high classification accuracy [11].
Sleep staging analysis and its scoring is a complicated procedure because of changes in sleep characteristics related to different sleep stages and also its non-stationary nature of the signal information.
2 Related Work
Most of the authors are proposed an automatic sleep stage classification system for identifying the sleep patterns and diagnosis of several types of sleep-related disorders [12,13,14,15]. In general sleep, staging procedures are conducted mainly on two strategies, one with single-channel input recording, and the other is multi-channel input recordings. In the first approach, only one channel is considered for extracting the informative features about the sleep characteristics of the subjects. There is a common approach taken by the different researchers during sleep staging as follows: (1) acquisitions of the signals, (2) signal pre-processing, (3) extraction of the properties, (4) feature reduction, and (5) classification [16]. The (3) feature extraction step is used for extracting the different characteristics parameter from preprocessed signal stage (2). These feature values can be extracted in frequency, time, time–frequency and nonlinear domains [17]. It has been seen that some of the ASSC system, one additional step used by authors that is feature reduction or dimensionality reduction stage. It is very helpful in screening the relevant features for the classification model. Currently, research on sleep staging plays an important role in NCP and human–machine interaction (HMI). There are several studies related to automated sleep staging using various physiological datasets and multimedia data, such as EEG, EMG, EOG, ECG, and audio, etc. One of the most popular contributions of sleep stage classification is the study of sleep behavior through human brain–computer interaction (BCI). We now look upon some of the recent contributions presented by different authors related to sleep staging using deep learning concepts.
2.1 Polysomnography (PSG) Based Sleep Staging Using Deep Learning Approaches
Nowadays, the researchers are majorly focused on deep learning techniques for sleep staging because of its robustness, scalability, and adaptability with related to handle large amounts of signal recordings and it’s processing. Another important advantage related to deep learning models is no need to require any explicit features for discriminating the subject's sleep behavior. In [18], the author used a deep convolution neural network for automated sleep staging with the input of single-channel EEG. The model achieved an overall accuracy of 74%. Sors et al. [19] presented automatic sleep stages scoring for five-sleep states based on one-channel of EEG using the CNN model, and the results reported for the proposed model are 87%. Chambon et al. [20] introduced a deep learning model with the concept of multi-variate signal analysis such as EEG, EOG, and EMG using KNN. The proposed model reached an overall accuracy of 80% with combinations of EEG + EOG + EMG. In [21] the authors obtained five-layer convolution layers for classifying the sleep stages based on two-channels of EEG and EOG signal and one-channel of EMG signal and achieved result for the model is 83%. Tripathy et al. [22] introduced a novel approach of sleep scoring based on coupling features of EEG data and RR time-series information using deep neural networks. The model resulted in an average accuracy of 95.71, 94.03, and 85.51% for the classification in between NREM versus REM, deep sleep versus light sleep, and sleep vs wake respectively. Cui et al. [23] proposed a sleep scoring system with input of 30 s multi-channel signal information based on CNN and fine-grained properties, the model reported an average accuracy of 92.2% with the ISRUC-Sleep public dataset. Supra et al. [24] designed a system of sleep scoring through extracted time-invariant information using CNN and find sleep stages transition information from the bidirectional LSTM network. The reported classification accuracy performance reached to 86.2%. According to the existing contribution to sleep scoring, major challenges found that choosing the correct features which helps to distinguish the sleep stages. It has found that the maximum researchers extracted the time, frequency, and time–frequency features, then after finalizing the relevant features either manually or applied some conventional feature selection algorithm. In some cases, this selection algorithm increases the complexity factor and consumes more time. Another challenge related to feature selection is that some features are well fitted for some of the subjects but the same may not apply for another one. Sometimes this imbalance of sleep information may produce biased results with conventional machine learning algorithms.
Another limitation regarding selected features, some of the features well suitable for classification for some of the subject cases may be the same many not applicable for other categories of subjects. This may create a problem to achieve higher classification accuracy.
2.2 Contribution
The main contributions of our proposed research works are explained below:
-
1.
We propose a 1D-CNN architecture for classifying multiple sleep classes based on multivariate signals using two different categories of subjects sleep recordings.
-
2.
The complete sleep staging process was analyzed with the input of single-channel EEG.
-
3.
The proposed methodology uses fewer parameters for train the model and extracting the prominent features from the input signal data automatically, which sup-ports achieving the high classification accuracy incomparable to the earlier contributions.
The remainder of the paper is organized as follows: Sect. 2 describes detailed on the proposed methodology including experimental data preparation, data preprocessing, feature extraction, and feature screening. Section 4 discusses the experimental results of the proposed methodology results. Section 5 presents the brief discussion about the proposed research work and makes result analysis with the state-of-the-art methods. Section 6 ends with concluding remarks with future work description.
3 Methodology
3.1 Experimental Data
In this paper, we used two different subgroups of sleep recordings, which were recorded during the sleep hours of the subjects under the direct supervision of the sleep experts of the Hospital of Coimbra University, Portugal. This dataset is called as ISRUC-Sleep dataset [25]. In the present work, two different categories (ISRUC-Sleep subgroup-I (SG-I) and ISRUC-Sleep subgroup-III (SG-III)) of the subjects considered for the purpose of experimental works, under SG-I, all the subjects were affected with the sleep diseases and SG-III, all the subjects were completely healthy category. The sleep epoch distributions among the different sleep stages are presented in Table 1.
3.2 Preprocessing
It has been better to remove the artifacts like muscle twitching, muscle movements, and eye blinks information for the better analysis the sleep behavior of the subjects. To remove these irrelevant portions from the raw signal, we have applied a Butterworth bandpass filter with order 10 at the frequency ranges of 0.1–60 Hz.
3.3 CNN Model Architecture
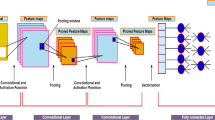
The proposed 1D-CNN model comprises 20 layers: nine 1-D convolution layers (CONV-1 to CONV-9), nine max-pooling layers (Polling-1 to Polling-9), and one fully connected layer. Apart from these layers, the proposed model considered nine batch normalization and ReLU layers. The proposed 1D-CNN architecture is shown in Fig. 1.
The proposed network extracted hierarchical feature information automatically using a set of hidden layers. As we have mentioned earlier, the use of more FC layers may increase the computational overhead. Therefore in this research work, we have only considered one FC layer, and finally, the model resulted in a vector of size five corresponding to five different classes of sleep stages, and at last, a softmax activation function is applied in the FC layer to determine the final class label.
The main objective of designing such a custom model is to increase the classification accuracy compared to the preexisting trained model. The first convolution layer of the proposed model takes an input of preprocessed polysomnography signals of size 3000 × 1 sample points , and the first layer model convolves it with 16 × 8 filters and with four stride ratios to produce the resulting feature map of size 750 × 16. The second layer of the proposed CNN model is the pooling layer; in this work, we have considered max-pooling techniques with filter size 2 × 2 and a stride 1 to produce a lower dimension size of output volume of size 375 × 16. The BN layer and ReLU activation are applied over the output of each max-pooling layer of the models. Next, to the pooling layer, we have again applied the second convolution layer and it carried the previous convolution layer output information and convolves it with 32 × 3 kernel of size with two stride ratios; it provides the output volume of size 352 × 32.
Finally, the output result of the convolution layer is fully connected to five neurons of a fully connected layer and applied softmax function to determine the probability distribution over each class label and finally, the output decided upon the neurons having the maximum probability score.
In this study, for model designing only three PSG signals and five sleep stages of the ISRUC-Sleep subgroup-I and subgroup-III dataset are used. Besides, both the subgroups datasets are split into 70% training data and 30% testing data.
4 Experiments and Results
The whole experiment was executed on the most popular and widely used ISRUC-Sleep subgroup-I and subgroup-III datasets. In this study, we have considered both categories of subjects for sleep staging, (1) subjects affected with the sleep disorder and (2) healthy controlled subjects.
4.1 Experimental Setup
In this research study, we have considered EEG signals, and the collected EEG signals are segmented into a 30 s time frame. All the experiments of this proposed study executed using MATLAB software with system configuration Intel Xeon 2.4 Ghz CPU processor, 16 GB RAM, and used 4 GB GPU from NVIDIA Quadro K2200 platform running under Windows 10 operating system. The entire dataset was partitioned into training and testing parts. For training, we selected 70% recordings from the entire dataset, and the remaining 30% data were used for testing purposes.
The proposed 1D-CNN model recognizes the sleep behavior through learns high-level properties using several cyclic executions and to achieve consistent performance from the model, a set of hyperparameters are used during training and the detailed hyperparameter settings for this research work are described in Table 2. To evaluate the performance of the proposed 1D-CNN architecture, we considered confusion matrix outcome with subject to sleep scoring using multivariate signals. The confusion matrix four terms True Positive (TP), False Positive (FP), True Negative (TN), and False Negative (FN) are used for measuring the sensitivity [26], precision [27], F1 Score [28] and classification accuracy [29].
4.2 Results with the Input of ISRUC-Sleep Subgroup-I Dataset Using EEG Signal
In this experiment, we considered with single-channel C3-A2 of EEG signal recordings of five sleep-disordered subjects. For this experiment also, the ratio for training and testing is 70:30 with the same number of features are used. Table 3 presents the confusion-matrix generated from the training set and testing set for five-state sleep staging. The reported performance metrics results for the proposed methodology is presented in Table 4. The overall accuracy performance for two-five sleep states classification problems are presented in Table 5. The training and testing accuracy performance for 100 epoch iterations is shown in Figure 2.
The highest classification accuracy performance as 98.88% with training samples and 99.50% as testing samples.
The highest accuracy resulted for training set data as 98.88% and for testing set data as 99.50% for four sleep state classifications. Similarly, the performances of sensitivity, precision, and F1-score reported more than 96% for all the five sleep stages.
Results with the SG-III dataset
The experiment was conducted with the SG-III with same channel and the same training and testing samples. The same model layer parameters used with the previous dataset (SG-I dataset) were also applied for this dataset also (ISRUC-Sleep subgroup-III).The graphical representation of resulted accuracy from both training and testing data using single-channel EEG is shown in Fig. 3. The detailed accuracy results for two-five sleep states using SG-III dataset are given in Table 6.
4.3 Summary of Experimental Results
In the current research work, we have conducted sleep staging studies considering two subgroups of sleep recordings (Subgroup-I/Subgroup-III) of the Sleep-ISRUC dataset. For all the individual experiments, we used only the proposed 1D-CNN model and with the same training and testing dataset size (70:30). The proposed model is effective with subjects to sleep scoring without any manual feature extraction or feature screening. The reported summary results obtained from both the datasets with the input of single-channel EEG using the proposed 1D-CNN model presents in Table 7.
5 Discussion
Many similar research works have been conducted by different researchers on multiple sleep staging using different methodologies through machine learning techniques. Till now very few of the studies were conducted based on the 1D-CNN model with deep learning techniques. The proposed 1D-CNN architecture can automatically be learning high-level features from the input signals directly. The obtained results indicate that the proposed scheme achieved improved sleep stages classification accuracy compared to earlier similar contributed works using the deep learning (DL) approach. Table 8 presents the performance comparison results between the proposed methodology and the existing similar multiple sleep staging studies using different deep learning techniques. It has been found that the proposed framework achieved higher classification accuracies in comparison to the other state art of the contributions. The proposed 1D-CNN model results indicated that the model is performed excellently concerning two-five sleep stages classifications incomparable to the existing literature works. It has been seen that several similar works are completely based on handcrafted features and shallow classifiers. From this point, the proposed model completely differs from the rest of the work and provides good potential for sleep staging analysis. Despite the improved performance on classification accuracy, the proposed model reported some more advantages related to the other works: (i) The proposed 1D-CNN model eliminates the traditional techniques for classification using multi-stage pipeline architecture, which creates a lot of complexity during execution and arises a lot of errors. (ii) The proposed model learned the features automatically from the polysomnography signals during the model training; therefore, it does not require any types of handcrafted features; (iii) the proposed model used less number of learnable parameters for training model incomparable to the existing some of the pre-trained DL models, (iv) with the same hyperparameter values, the sleep staging performance is significantly improved for two-five sleep states classification using both the subgroups of a dataset and (v) the proposed architecture obtains higher classification accuracy performance in comparisons with the existing state-of-the-art works.
6 Conclusion
In this paper, we proposed a 1D-CNN model for automated sleep stage classification using polysomnography signals. The proposed architecture contains nine learnable layers, which helps to learn the features automatically from the single-channel EEG signals. The main objective of designing such architecture is to improve the classification accuracy results with better learnable parameters compared to the traditional shallow learning models. The proposed 1D-CNN architecture achieved the highest classification accuracy of 98.44, 99.07, 99.50, and 99.03% using the SG-I dataset; similarly, the same model reported accuracy using the SG-III dataset of 98.51, 98.88, 98.76, and 98.67% with single-channel EEG signals for two-five sleep stages classification. The proposed model not required any types of handcrafted features and shallow learning classification models for the classification of polysomnography signals, and it can assist clinicians during the sleep scoring. In future, we will collect the biomedical signals data from different medical institutes and test the efficiency of the model. Further, we also focus on several tasks which ranging from classification task to biomedical signal analysis. Furthermore, the proposed work to be extended for different types of sleep related diseases. We will also include the data augmentation techniques to overcome the data imbalance issues. Finally, it has been observed that the proposed framework improves the existing state of the art and achieves better classification results for the two-five sleep classification tasks. The proposed fully automated sleep staging classification systems could replace the traditional error-prone classification models.
References
Heyat MBB, Lai D, Khan FI, Zhang Y (2019) Sleep bruxism detection using decision tree method by the combination of C4-P4 and C4-A1 channels of scalp EEG. IEEE Access 7:102542–102553. https://doi.org/10.1109/ACCESS.2019.2928020
Chung MH, Kuo TB, Hsu N, Chu H, Chou KR, Yang CC (2009) Sleep and autonomic nervous system changes—enhanced cardiac sympathetic modulations during sleep in permanent night shift nurses. Scand J Work Environ Health 35(3):180–187. https://doi.org/10.5271/sjweh.1324
Aboalayon K, Faezipour M, Almuhammadi W, Moslehpour S (2016) Sleep stage classification using EEG signal analysis: a comprehensive survey and new investigation. Entropy 18(9):272. https://doi.org/10.3390/e18090272
Reynolds CF, O’Hara R (2013) DSM-5 sleep-wake disorders classification: overview for use in clinical practice. Am J Psychiatry 170(10):1099–1101. https://doi.org/10.1176/appi.ajp.2013.13010058
Goel N, Rao H, Durmer J, Dinges D (2009) Neurocognitive consequences of sleep deprivation. Semin Neurol 29(04):320–339. https://doi.org/10.1055/s-0029-1237117
Garcés Correa A, Orosco L, Laciar E (2014) Automatic detection of drowsiness in EEG records based on multimodal analysis. Med Eng Phys 36(2):244–249. https://doi.org/10.1016/j.medengphy.2013.07.011
Kogure T, Shirakawa S, Shimokawa M, Hosokawa Y (2011) Automatic sleep/wake scoring from body motion in bed: validation of a newly developed sensor placed under a mattress. J Physiol Anthropol 30(3):103–109. https://doi.org/10.2114/jpa2.30.103
Rosenberg RS, Van Hout S (2013) The american academy of sleep medicine inter-scorer reliability program: sleep stage scoring. J Clin Sleep Med 9(1):81–87. https://doi.org/10.5664/jcsm.2350
Boashash B, Ouelha S (2016) Automatic signal abnormality detection using time-frequency features and machine learning: a newborn EEG seizure case study. Knowl-Based Syst 106:38–50. https://doi.org/10.1016/j.knosys.2016.05.027
Penzel T, Conradt R (2000) Computer based sleep recording and analysis. Sleep Med Rev 4(2):131–148. https://doi.org/10.1053/smrv.1999.0087 PMID: 12531163
Li Y, Luo M-L, Li K (2016) A multiwavelet-based time-varying model identification approach for time–frequency analysis of EEG signals. Neurocomputing 193:106–114. https://doi.org/10.1016/j.neucom.2016.01.062
Holland JV, Dement WC, Raynal DM (1974) Polysomnography: a response to a need for improved communication. Presented at the 14th Annual Meeting Association Psychophysiology Study Sleep. [Online]
Acharya UR, Bhat S, Faust O, Adeli H, Chua EC-P, Lim WJE, Koh JEW (2015) Nonlinear dynamics measures for automated EEG-based sleep stage detection. Eur Neurol 74(5–6):268–287. https://doi.org/10.1159/000441975
Acharya UR, Oh SL, Hagiwara Y, Tan JH, Adeli H (2018) Deep convolutional neural network for the automated detection and diagnosis of seizure using EEG signals. Comput Biol Med 100:270–278. https://doi.org/10.1016/j.compbiomed.2017.09.017
Acharya UR, Oh SL, Hagiwara Y, Tan JH, Adeli H, Subha DP (2018) Automated EEG-based screening of depression using deep convolutional neural network. Comput Methods Prog Biomed 161:103–113. https://doi.org/10.1016/j.cmpb.2018.04.012
Acharya UR, Vinitha Sree S, Swapna G, Martis RJ, Suri JS (2013) Automated EEG analysis of epilepsy: a review. Knowl Based Syst 45:147–165. https://doi.org/10.1016/j.knosys.2013.02.014
Ahmadlou M, Adeli H, Adeli A (2011) Fractality and a wavelet-chaos-methodology for EEG-based diagnosis of Alzheimer disease. Alzheimer Dis Assoc Disord 25(1):85–92. https://doi.org/10.1097/WAD.0b013e3181ed1160
Tsinalis O, Matthews PM, Guo Y, Zafeiriou S (2016) Automatic sleep stage scoring with single-channel eeg using convolutional neural networks; arXiv preprint arXiv:1610.01683
Sors A, Bonnet S, Mirek S, Vercueil L, Payen J-F (2018) A convolutional neural network for sleep stage scoring from raw single-channel eeg. Biomed Signal Process Control 42:107–114. https://doi.org/10.1016/j.bspc.2017.12.001
Chambon S, Galtier MN, Arnal PJ, Wainrib G, Gramfort A (2018) A deep learning architecture for temporal sleep stage classification using multivariate and multimodal time series. IEEE Trans Neural Syst Rehabil Eng 26(4):758–769. https://doi.org/10.1109/TNSRE.2018.2813138
Fernández-Varela I, Hernández-Pereira E, Moret-Bonillo V (2018) A convolutional network for the classification of sleep stages. Proceedings 2(18):1174. https://doi.org/10.3390/proceedings2181174
Tripathy RK, Rajendra Acharya U (2018) Use of features from RR-time series and EEG signals for automated classification of sleep stages in deep neural network framework. Biocybern Biomed Eng. https://doi.org/10.1016/j.bbe.2018.05.005
Cui Z, Zheng X, Shao X, Cui L (2018) Automatic sleep stage classification based on convolutional neural network and finegrained segments. Hindawi Complex 9248410. https://doi.org/10.1155/2018/9248410
Supratak A, Dong H, Wu C, Guo Y (2017) DeepSleepNet: a model for automatic sleep stage scoring based on raw single-channel EEG. IEEE Trans Neural Syst Rehabil Eng 25(11):1998–2008. https://doi.org/10.1109/TNSRE.2017.2721116
Khalighi S, Sousa T, Santos JM, Nunes U (2016) (2016) ISRUC-Sleep: a comprehensive public dataset for sleep researchers. Comput Methods Programs Biomed 124:180–192. https://doi.org/10.1016/j.cmpb.2015.10.013
Bajaj V, Pachori RB (2013) Automatic classification of sleep stages based on the time-frequency image of EEG signals. Comput Methods Programs Biomed 112(3):320–328. https://doi.org/10.1016/j.cmpb.2013.07.006
Yıldız A, Akin M, Poyraz M, Kirbas G (2009) Application of adaptive neuro-fuzzy inference system for vigilance level estimation by using wavelet-entropy feature extraction. Expert Syst Appl 36:7390–7399. https://doi.org/10.1016/j.eswa.2008.09.003
Sanders TH, McCurry M, Clements MA (2014) Sleep stage classification with cross frequency coupling. Annu Int Conf IEEE Eng Med Biol Soc 2014:4579–4582. https://doi.org/10.1109/EMBC.2014.6944643
Powers D, Ailab (2011) Evaluation: from precision, recall and F-measure to ROC, informedness, markedness and correlation. J Mach Learn Technol 2:2229–3981. https://doi.org/10.9735/2229-3981
Yildirim O, Baloglu U, Acharya U (2019) A deep learning model for automated sleep stages classification using PSG signals. Int J Environ Res Public Health 16(4):599. https://doi.org/10.3390/ijerph16040599
Fernandez-Blanco E, Rivero D, Pazos A (2019) Convolutional neural networks for sleep stage scoring on a two-channel EEG signal. Soft Comput. https://doi.org/10.1007/s00500-019-04174-1
Ieracitano C, Mammone N, Bramanti A, Hussain A, Morabito F (2018) A convolutional neural network approach for classification of Dementia stages based on 2D-spectral representation of EEG recordings. Neurocomputing 323. https://doi.org/10.1016/j.neucom.2018.09.071
Nagabushanam P, Thomas George S, Radha S (2019) EEG signal classification using LSTM and improved neural network algorithms. Soft Comput. https://doi.org/10.1007/s00500-019-04515-0
Michielli N, Acharya UR, Molinari F (2019) Cascaded LSTM recurrent neural network for automated sleep stage classification using single-channel EEG signals. Comput Biol Med 106:71–81. https://doi.org/10.1016/j.compbiomed.2019.01.013
Li X, La R, Wang Y, Niu J, Zeng S, Sun S, Zhu J (2019) EEG-based mild depression recognition using convolutional neural network. Med Biol Eng Comput. https://doi.org/10.1007/s11517-019-01959-2
Banluesombatkul N, Ouppaphan P, Leelaarporn P, Lakhan P, Chaitusaney B, Jaimchariyatam N, Chuangsuwanich E, Chen W, Phan H, Dilokthanakul N, Wilaiprasitporn T (2020) MetaSleepLearner: a pilot study on fast adaptation of bio-signals-based sleep stage classifier to new individual subject using meta-learning
Mousavi Z, Yousefi Rezaii T, Sheykhivand S, Farzamnia A, Razavi SN (2019) Deep convolutional neural network for classification of sleep stages from single-channel EEG signals. J Neurosci Methods 108312. https://doi.org/10.1016/j.jneumeth.2019.108312
Zhang X, Xu M, Li Y, Su M, Xu Z, Wang C, et al, (2020) Automated multi-model deep neural network for sleep stage scoring with unfiltered clinical data. Sleep Breath. https://doi.org/10.1007/s11325-019-02008-w
Nakamura T, Adjei T, Alqurashi Y, Looney D, Morrell MJ, Mandic DP (2017) Complexity science for sleep stage classification from EEG. In: 2017 international joint conference on neural networks (IJCNN). https://doi.org/10.1109/ijcnn.2017.796641
Hassan AR, Bhuiyan MIH (2017) Automated identification of sleep states from EEG signals by means of ensemble empirical mode decomposition and random under sampling boosting. Comput Methods Programs Biomed 140:201–210. https://doi.org/10.1016/j.cmpb.2016.12.015
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this paper
Cite this paper
Satapathy, S.K., Sharathkumar, S., Loganathan, D. (2021). Automated Sleep Staging Using Convolution Neural Network Based on Single-Channel EEG Signal. In: Sharma, H., Gupta, M.K., Tomar, G.S., Lipo, W. (eds) Communication and Intelligent Systems. Lecture Notes in Networks and Systems, vol 204. Springer, Singapore. https://doi.org/10.1007/978-981-16-1089-9_51
Download citation
DOI: https://doi.org/10.1007/978-981-16-1089-9_51
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-1088-2
Online ISBN: 978-981-16-1089-9
eBook Packages: EngineeringEngineering (R0)