Abstract
Although the fragment molecular orbital (FMO) method enables electronic structure calculations with near-linear scaling behavior with respect to system size, the computational cost of ab initio methods typically employed in conjunction with FMO is still prohibitive for routine calculations of very large systems or long timescale molecular dynamics simulations. We, therefore, combined the FMO and density-functional tight-binding (DFTB) method, which is one of the emerging semi-empirical quantum chemical methods, and have demonstrated that FMO-DFTB is capable of performing geometry optimizations for systems containing up to one million atoms using limited computational resources. In this chapter, we will review the basics of the DFTB method first before introducing FMO-DFTB, focusing on the relationship with density functional theory and other FMO methodologies. We also demonstrate the latest scalings of FMO-DFTB2 and DFTB3 using three-dimensional water clusters, showing that the most favorable scaling is \(\mathscr {O} \left( N^{1.16} \right) \). Applications of FMO-DFTB to various systems are briefly summarized, and an outlook to future applications is provided.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Density-functional tight-binding
- Approximate electronic structure theory
- Molecular dynamics
- Near-linear scaling
1 Introduction
In performing quantum chemical calculations, finding a reasonable trade-off between accuracy and computational cost is always inevitable. If an infinitely fast computer would be available, the choice should always be the full configuration interaction method, which allows to compute the exact nonrelativistic solution of the Schrödinger equation within the Born–Oppenheimer approximation. Such a computer, anticipated by some as the ultimate goal of current efforts in quantum computing, is obviously still far from realization. With the currently available limited computational resources, we have to choose a method which (hopefully) yields the highest accuracy with the lowest computational cost, carefully considering the expected accuracy and cost. In principle, accurate methods exhibit unfortunate, steep scaling with respect to system size, such as polynomial or, as in the case of full configuration interaction, factorial. The scaling of all standard quantum chemical calculations is at least equal or greater than quadratic. Quadratic scaling in electronic structure calculations can be achieved by using integral prescreening [1] and density fitting [2] techniques. If the bottleneck of integral calculation can be overcome, for instance, by efficient parallelization or parameterization, matrix diagonalization and other linear algebra related to Hamiltonian and density matrices become dominant, and these operations are commonly associated with cubic scaling. Thus, one always suffers from rapid increase of the computational cost as far as we treat dense matrices. This problem may be actually less severe; we can simply wait for long calculations to finish or, if the code parallelizes well, we can use more and more CPU cores. However, the memory requirement is another, and possibly more severe, problem. If we cannot store a very large matrix in memory, we cannot even execute the calculation. For instance, storing a 100,000 \(\times \) 100,000 matrix in memory requires 74.5 GB in double precision, so some computers with a medium-sized memory space may not handle this large calculation. In any case, calculations for large systems always suffer from these problems.
In this context, a number of linear-scaling methods have been developed, and one of them is the fragment molecular orbital (FMO) method [3], the main theme of this book. After the first idea by Kitaura et al., it has been combined with various ab initio methods, and numerous methodological simplifications and advances have been reported. Various FMO approaches have been successfully applied to a great many systems. In spite of the usefulness of the approach itself, computational costs of the ab initio methods themselves are still high, even when they are combined with FMO. One may find that treating the whole system in a straightforward fashion with semi-empirical quantum mechanics (QM) methods is sometimes faster than FMO combined with ab initio methods, if the system consists of less than a few thousand atoms.
The density-functional tight-binding (DFTB) method [4,5,6,7,8] is one of the semi-empirical QM methods, and it has been applied to a number of nanomaterials [9], chemical [10] and biosystems [11]. The method is known as an approximation of density functional theory (DFT), utilizing tight-binding approximations such as minimal Slater-type basis set and two-center approximations for Hamiltonian matrix elements, and diatomic repulsive potentials. The earliest DFTB development perhaps began with the seminal works by Porezag et al. [12] and Seifert et al. [13]. This option is known as nonself-consistent-charge DFTB (NCC-DFTB) or DFTB1, and it has been applied to systems in which charge polarization is not significant. As the name implies, NCC-DFTB requires only a single diagonalization of a model Hamiltonian employing a reference electron density. It is conceptually similar to extended Hückel-type methods, although the potentials are more sophisticated. The second generation of DFTB is the self-consistent-charge (SCC) formalism [14], also recently called DFTB2. This extension is based on the fundamental ideas of Foulkes and Haydock [15]. In the SCC formalism, interactions between atomic charges are explicitly taken into account, and the electronic structure is determined as a minimum of electronic energy with respect to the change of variational parameters, improving the description of charge–charge interactions. It requires self-consistent treatment of charge–charge interactions, as is evident from the name. The third generation is the DFTB3 formalism introduced in 2011 by Gaus et al. [16], which includes a third-order term related to the charge-dependence of the Hubbard parameter introduced in DFTB2. More details on DFTB methods are reviewed later.
In terms of the computational cost, even though the pre-factor of DFTB is relatively small, all the DFTB generations require matrix multiplications and diagonalization of the Hamiltonian (roughly equivalently, Fock matrix in Hartree–Fock) matrix more than once, so the computational scaling of these steps formally scales as cubic, and applications to large systems (more than 1000 atoms) are, therefore, not straightforward. For instance, a DFTB energy and gradient calculation for a water cluster containing 4000 molecules take about 6 d on a single CPU [17]. To circumvent this problem, various linear-scaling approaches have been combined with DFTB. One realization is a rather direct method; large systems are treated as they are, i.e., without fragmentation. Focusing only on the approaches combined with DFTB, this category may include the use of an OpenMP sparse matrix solver [18], shift-and-inverse parallel spectral transformations (SIPs) [19], and graph-based Fermi operator expansion [20]. A key of these methods is the exploitation of the sparsity of Hamiltonian and overlap matrices. Another realization divides large systems into smaller spatial segments or fragments: modified divide-and-conquer (mDC) [21], fragment molecular orbital (FMO) [17] and divide-and-conquer (DC) [22, 23]. Although it would be certainly interesting to compare the performance and the applicability of these linear scaling methods, our focus in this chapter is the combination of FMO and DFTB, so we are not going to discuss the DC methods further.
Since 2014, we have been developing an ultra-fast semi-empirical quantum mechanical method, FMO-DFTB, which combines FMO and DFTB. The scaling of the computational cost was close to linear, \(\mathscr {O} \left( N^{1.2} \right) \), even for three-dimensional water clusters. The performance of the method has been reported several times since its initial conception, and the largest system it was applied to consisted of 1,180,800 atoms for which each single point gradient calculation took only 22 min, using a single computer node with 24 CPU cores. In this chapter, we review our FMO-DFTB method after briefly introducing the DFTB method itself. We will discuss previous applications of DFTB and FMO-DFTB and finally conclude with a short summary and future prospect.
2 Basics of the DFTB Method
The basics of the DFTB method are briefly reviewed in this section. The formalism introduced in this subsection is limited for the purpose of this review chapter, and we point the reader to more detailed introductions, for instance, in Refs. [4,5,6,7,8] Furthermore, semi-empirical quantum mechanical methods have been compared in detail in Ref. [10].
As mentioned, the DFTB method is derived from the DFT method itself. Here, given the (perturbed) total density \(\rho \), the total Kohn–Sham DFT energy can be written as
where \(f_i\) is the occupation number of ith orbital \(\psi _i\), \(V^\mathrm {ext}\) is the external potential, \(E^\mathrm {xc} [\rho ]\) is the exchange–correlation energy, and \(E^\mathrm {nuc}\) is the nuclear repulsion. Additionally, the following abbreviations were used: \(\int = \int \mathrm {d} \mathbf {r}\), \(\int ^\prime = \int \mathrm {d} \mathbf {r}^\prime \), \(\rho = \rho \left( \mathbf {r} \right) \), and \(\rho ^\prime = \rho \left( \mathbf {r}^\prime \right) \). For convenience, two more abbreviations are introduced later in the text: \(\int ^{\prime \prime } = \int \mathrm {d} \mathbf {r}^{\prime \prime }\) and \(\rho ^{\prime \prime } = \rho \left( \mathbf {r}^{\prime \prime } \right) \).
A series of approximations can then be introduced. The perturbed total density \(\rho \) can be expressed as the sum (superposition) of a reference electron density \(\rho ^0\) and a density perturbation \(\delta \rho \): \(\rho = \rho ^0 + \delta \rho \). The reference density \(\rho ^0\) usually corresponds to the density of a free atom in vacuum. Free atoms in a bound molecular system or solid affect each other, perturbing their density, and this density fluctuation can be expressed as \(\delta \rho \). Using the density partitioning \(\rho = \rho ^0 + \delta \rho \), the exchange–correlation energy can be expanded in a Taylor series around the reference density in terms of the density perturbation:
With this expansion, the DFT total energy in Eq. (1) can be expanded as
where \(V^\mathrm {xc}\) is the exchange–correlation potential. Each term in Eq. (4) corresponds to one line in Eq. (3) and is related to the truncation order of the Taylor expansion in Eq. (2). The last equation is a simplified expression of the DFTB energy:
where the truncation of terms distinguishes the level of DFTB: DFTB1, DFTB2, and DFTB3.
The repulsion energy above is defined by
Clearly, the first term takes into account the nuclear repulsion, whereas the other terms avoid double-counting Coulomb interactions and the exchange–correlation energy of the reference density. \(E^\mathrm {1st}\) is sometimes referred to as the band structure or electronic energy, defined by
This term is further simplified to yield
where \(\hat{H} [\rho ^{0}]\) is the Hamiltonian, which is only dependent on the reference density.
The second-order contribution
addresses Coulomb interactions and exchange–correlation contributions of the perturbed electron density. The third-order contribution
originates in the third-order Taylor expansion of the exchange–correlation energy. Although higher order DFTB models (with fourth- and higher order Taylor expansion) may also be formulated, no efforts for including them have been devoted. The formulations employed during computation are briefly reviewed in the following subsections. As usual, the linear combination of atomic orbitals (AOs) is applied:
where \(\phi _\mu \) represents \(\mu \)th AO and \(C_{\mu i}\) is the expansion coefficient.
2.1 DFTB1
The NCC-DFTB method is sometimes referred to as DFTB1, indicating both its expansion order, and indicating that it is the first generation of the DFTB hierarchy of methods. The total energy of DFTB1 can be given by the sum of the repulsion and one-electron contributions:
Here, the density matrix \(D_{\mu \nu }\) is defined by
and \(H_{\mu \nu }^0\) is sometimes referred to as the non-perturbed Hamiltonian. After neglecting several contributions, the matrix elements of \(H_{\mu \nu }^0\) are written as
where A and B represent the index of atoms and \(\mu \in A\) and \(\nu \in B\). The orbital energy of \(\mu \)th AO is \(\varepsilon _\mu ^\mathrm {neutral~free~atom}\), and \(V_0^A\) is the potential generated by the atom A at a neutral state. The terms in the right-hand side are obtained from reference DFT calculations. In most cases, the Perdew–Burke–Ernzerhof (PBE) [24, 25] exchange–correlation functional is employed in the DFT calculation of atoms and diatomic molecules in vacuum. For \(A \ne B\), the term is determined so that the DFTB calculations with parametrized \(H_{\mu \nu }^0\) reproduce the band structure obtained with DFT. In actual DFTB calculations, tabulated (discretized) values in the Slater–Koster files are normally interpolated using cubic or fifth-order polynomials, so no integrals are explicitly computed. DFTB was originally developed using Slater-type AOs, although this is only relevant during parametrization, and other choices including numerical AOs and Gaussian AOs have been used as well.
Applying the standard variational method, the optimum MO coefficients can be obtained by solving the generalized eigenvalue problem
where \(S_{\mu \nu }\) is the (non-orthogonal) overlap matrix and \(\varepsilon _i\) are the eigenvalues of the ith vector (molecular orbital). The overlap matrix elements are also parametrized, tabulated, and computed by interpolations, as in the case of \(H_{\mu \nu }^0\). The Hamiltonian, here corresponding to \(H_{\mu \nu }^0\), is directly constructed with Slater–Koster tables, and is thus not dependent on the electronic structure. Therefore, a single diagonalization suffices to determine the electronic structure and to compute the total energy within DFTB1; no self-consistent field (SCF) cycles are required. Note that the DFTB community usually uses SCC to refer to SCF in Hartree–Fock. However, when combined with FMO, SCC formally means monomer iterations to determine electrostatic potential (ESP). To avoid unnecessary confusion, “SCF” cycles are exclusively used here to determine MO coefficients.
The repulsive potential \(E_{AB}^\mathrm {rep}\) is empirically determined to minimize the deviations of geometrical parameters from reference geometries optimized with DFT. It is usually obtained after having optimized the AOs used to construct the diagonal and off-diagonal matrix elements of the Hamiltonian. We note in passing that parameterization of both electronic parameters as well as repulsive potentials simultaneously may lead to instabilities in the optimization procedure.
2.2 DFTB2
DFTB1 has been successfully applied to some homogeneous systems, such as carbon-based nanomaterials [9] and some bulk systems [14]. Nevertheless, DFTB1 truncates the Taylor series at the first order and does not include charge–charge interactions in the formalism, limiting the accuracy for systems with heteroatoms, and is thus not anymore widely used. Elstner et al. proposed the self-consistent version of DFTB as a next-generation model in 1998 [14], DFTB2; the total energy within DFTB2 can be written by
The last term represents the second-order contributions, the third line of Eq. (3). The \(\gamma _{AB}\) function is dependent on the distance between atoms A and B and on the Hubbard values of them. The Hubbard values are related to the atomic ionization potentials and electron affinities or to chemical hardness. At a long distance, \(\gamma _{AB}\) closely follows the inverse of the distance between atoms A and B, \(R_{AB}\) and thus corresponds to the pure Coulomb interaction between two charges. On the other hand, at the \(R_{AB} \rightarrow 0\) limit, \(\gamma _{AB}\) will be the Hubbard value \(U_{A}\), leading to charge self-interaction on a give site (atom) [26]. The Mulliken charge \(\varDelta Q_A\) is the difference between the Mulliken population \(Q_A\) and the reference density \(Q_A^0\) of atom A: \(\varDelta Q_A = Q_A - Q_A^0\). The Mulliken population is calculated by
Only valence electrons are considered in DFTB, e.g., the \(Q_A^0\) of oxygen is six, not eight. In DFTB2, the Hamiltonian matrix \(H_{\mu \nu }\), conceptually corresponding to the Fock matrix in Hartree–Fock, is
for \(A \in \mu \) and \(B \in \nu \). Since \(\varDelta Q_C\) is dependent on the density matrix, the Hamiltonian matrix is dependent on the electronic structure. Therefore, the variational parameters within DFTB2 are determined iteratively by solving the eigenvalue problem self-consistently. Consequently, DFTB2 is 5–15 times slower than DFTB1.
2.3 DFTB3
In 2011, Gaus et al. [16] introduced two formal advancements to the DFTB2 as described in 2006 [11]. First, the third-order Taylor expansion term of the exchange–correlation energy in Eq. (2) was introduced. Intuitively, the chemical hardness of an atom should be dependent on its charge state. However, chemical hardness, which is relevant to Hubbard value, is constant within DFTB2; this has severely limited applying DFTB to highly charged systems, such as an anion and cation system. The third-order term introduced the charge-dependence of the \(\gamma _{AB}\) function in the DFTB formalism. Omitting the detailed derivation, the final energy within DFTB3 was expressed as
The last term is derived from the third-order expansion term in Eq. (3). The newly introduced function \(\varGamma _{AB}\) is dependent on the distance between atoms A and B, the Hubbard values of them, and the derivative of the Hubbard values with respect to charge fluctuation:
Analytic expressions of \(\varGamma _{AB}\) and \(\gamma _{AB}\) are derived in the Supporting Information Ref. [16]. Historically, the DFTB3 was used with diagonal terms only; \(\varGamma = 0\) for \(A \ne B\). Nowadays, this diagonal approximation is not usually employed. The Hamiltonian matrix in DFTB3 is
Thus, DFTB3 also requires iterative treatment. The convergence of SCF with DFTB3 is usually similar to that with DFTB2, but one or two more cycles may be needed.
The second advancement was the modification of the gamma function introduced in DFTB2: \(\gamma _{AB} \rightarrow \gamma _{AB}^\mathrm {h}\). The necessity of this modification arose from the difference of chemical hardness between isolated and covalently bonded hydrogen atoms. Thus, this modification improved the calculation of the binding energy in hydrogen bonding. Here, \(\gamma _{AB}\) and \(\gamma _{AB}^\mathrm {h}\) are not explicitly distinguished because the difference only affects the electronic structure indirectly. \(\varGamma _{AB}\) is a function of \(\gamma _{AB}\) and is, therefore, also affected.
2.4 Long-Range Corrected DFTB
The most recent and important development in DFTB is the long-range corrected DFTB (LC-DFTB). The first two practical implementation and benchmark studies were reported in 2015 by Humeniuk et al. [27] and Lutsker et al. [28], although the initial formulation was outlined in 2012 by a coauthor of the latter study [29]. The central idea of LC-DFTB is to add Hartree–Fock-like exchange contributions to the DFTB formalism. As the DFTB total energy was derived from the DFT energy with a GGA exchange–correlation functional, the addition of Hartree–Fock-like exchange terms is not completely straightforward [28]. In its current implementation, the reference Hamiltonian matrix elements are obtained using the BNL range-separated functional[30], and the Hartree–Fock-like exchange terms for the density perturbation are obtained using difference density matrices of the one-particle density.
The total energy of LC-DFTB [28] can be written as
where
and \(\varDelta D_{\mu \nu }\) is the difference between \(D_{\mu \nu }\) and the reference density matrix, \(D_{\mu \nu }^0\): \(\varDelta D_{\mu \nu } = D_{\mu \nu } - D_{\mu \nu }^0\). The reference density matrix is a diagonal matrix with the number of electrons of free neutral atoms in each AO. The last term in Eq. (22) is the newly added Hartree–Fock-like exchange term. The long-range gamma function \(\gamma _{AB}^\mathrm {lr}\) is similar to \(\gamma _{AB}\) in DFTB2 and DFTB3, but with an additional dependence on the range-separated parameter \(\omega \). In LC-DFT literature, the parameter may be written as \(\mu \). This formalism is based on DFTB2; as there is no term with \(\varGamma _{AB}\), there are no reports for LC-DFTB3 developments at the moment.
LC-DFTB was shown to be roughly ten times more expensive than conventional DFTB, although DFT calculations are 50 times slower than LC-DFTB [28]. Nevertheless, LC-DFTB decreases the self-interaction error, and applications to excited-state calculations [27, 31] have demonstrated that HOMO–LUMO gaps and charge transfer excitation energies are much improved.
2.5 Parameters
In tight binding calculations, in general, it is essential to prepare Slater–Koster parameters prior to simulation, and DFTB is no exception, as already mentioned above. These parameters are defined for each pair of elements; for water, four combinations are needed: O–O, O–H, H–O, and H–H. The two mixed parameters are required because of possible sign changes in integrals involving orbitals with non-zero angular momentum. Various parameters are employed in DFTB and a number of these parameters are available on the DFTB website [32]. To perform DFTB1 calculations, users have to supply \(H_{\mu \nu }^0\), \(S_{\mu \nu }^0\), and \(E_{AB}^\mathrm {rep}\) parameters. These are already summarized in Slater–Koster tables, so users simply have to specify the file locations. Users also need Hubbard values for DFTB2, and these have also been tabulated. For DFTB3, users need the derivative of the Hubbard values with respect to charge fluctuation, which has to be given separately. Because many parameters are still missing, not all elements can be treated with DFTB.
3 Basics of the FMO-DFTB Method
A brief history of the methodological development of FMO-DFTB is first presented. The first FMO-DFTB study reported the geometry optimization of a one-million-atom system and was published in 2014 [17]. At that time, only two-body interactions within FMO (FMO2) were considered and the DFTB model employed was only DFTB2: FMO2-DFTB2. The implemented gradient was not fully analytic and response terms coming from the use of ESP that are determined in the monomer cycle were neglected. In the next year, extensions to FMO2-DFTB3 [33] and fully analytic gradient [34] were implemented. Since then, all FMO-DFTB developments have included a fully analytic gradient and an extension to DFTB3. In 2016, FMO-DFTB was combined with the polarizable continuum model (PCM) [35], and an approximate Hessian with FMO-DFTB was implemented by Nakata [36]. In 2017, three-body interactions were included, namely FMO3-DFTB [37]. In 2018, FMO-DFTB was combined with an alternative fragmentation approach for cutting covalent bonds, termed as the adaptive frozen orbital (AFO) method [38]. At that time, FMO-DFTB was the only method using an analytic gradient within AFO because of the complexity of determining the electronic structure [39].
In the following subsections, only the FMO2-DFTB3 formalism is employed. DFTB2 formalism can be obtained by setting \(\varGamma _{AB} = 0\). As the extension to FMO3 is straightforward and there is a large overlap between DFTB and other ab initio methods, three-body terms are not explicitly introduced here.
All of these features presented in this chapter are publicly available through the official version of GAMESS-US [40, 41]. DFTB and FMO-DFTB in GAMESS-US were first officially released in 2014. Major updates were executed in 2016 and 2018, and the latter release includes almost all developments presented in this chapter.
3.1 Formalism of FMO-DFTB: Energy
Again, the total energy within FMO2 is given by
where N is the number of fragments and \(E_I^\prime \) is the internal energy of fragment I, defined by
This expression is almost identical to Eq. (19) but includes the fragment index. For convenience, the hybrid orbital projection operator \(P_{\mu \nu }^X\) is included in the non-perturbed Hamiltonian \(H_{\mu \nu }^{0,X}\). \(P_{\mu \nu }^X\) for FMO-DFTB is computed here similarly as in other methods. The energy of the charge transfer between fragments I and J in embedding ESP can be written as
In the case of DFTB, the electrostatic interaction between the atoms in fragment X and the atoms in the total system excluding fragment Y is given by
The first term in Eq. (27) originates from the second-order term, corresponding to the last term in Eq. (16), which originates in the sum of Coulomb interactions and the second-order expansion of the exchange–correlation energy (see the third line of Eq. (3)). The second term in Eq. (27) comes from the third-order term describing an expansion of the exchange–correlation functional, thus physically corresponding to exchange–correlation contributions to the ESP.
The Hamiltonian matrix within FMO-DFTB3 (with hybrid orbital projection) is written as
for \(\mu \in A\) and \(\nu \in B\), where the second and third terms come from the internal (atoms in fragment X) and external (atoms outside the fragment X) embedding, respectively. By defining for atoms \(A, B \in X\) as
the external embedding potential can be written as
Again, the second and third terms in Eq. (29) come from exchange–correlation-like contributions. Since the ESP is already expressed with Mulliken charges, the point charge approximation is always employed within DFTB.
It is important to analyze the \(E^\mathrm {2nd}\) term for \(A = B\):
Because \(U_A\) is non-zero, \(E^\mathrm {2nd}\) for \(A = B\) is also non-zero. This is a typical symptom of the self-interaction error. The residual value comes from the second-order term of the Taylor expansion (the term proportional to the inverse of the distance is canceled [14]), so the origin of this self-charge interaction seemingly inherits from the self-interaction error of the exchange–correlation functional in DFT. Special care thus has to be paid when evaluating ESP within FMO-DFTB if the system under consideration requires fragmentation across covalent bonds. ESP contributions from the same atom belonging to different fragments cannot be ignored. This also applies to third-order terms.
The introductions of two-electron integral-like terms in DFTB3 for \(\mu \in A\), \(\nu \in B\), \(\rho \in C\), and \(\sigma \in D\):
allows the ESP to be written as
This expression has some connections with FMO-HF and FMO-DFT, although FMO-DFTB benefits from a number of simplifications, resulting in expressions with Mulliken charges (Eqs. 29 and 30).
Efforts are also being made to combine FMO with the recently developed LC-DFTB [28]. In terms of the formalism, the main difference is the addition of exchange contributions to the two-electron integral-like term (Eq. 32):
Only intra-fragment exchange contributions are considered, as they are not considered in other FMO-related methods.
3.2 Formalism of FMO-DFTB: Gradient
As the total energy of FMO is not fully variationally determined, differentiation of the energy gives rise to response terms of the electronic structure. In FMO, such response contributions can be efficiently computed by solving self-consistent Z-vector (SCZV) equations [42]. This approach has been applied to ab initio methods. DFTB is not an exception; the response terms are computed to obtain accurate first-order derivatives in a similar manner.
As reported by Ref. [34], the error of the gradient without response contributions is on the order of \(5.0 \times 10^{-4}\) Hartree/Bohr, is greater than the default convergence criterion of GAMESS-US, and is particularly severe when combined with the AFO approach. With FMO2-DFTB3/AFO, the maximum error has been reported as large as \(10^{-3}\) Hartree/Bohr. The analytic gradient is 100 times more accurate, and the remaining error likely stems from the limited accuracy of numerical derivatives. Approximate gradients are rather accurate with FMO3, but the error is still on the order of \(10^{-4}\) Hartree/Bohr.
After derivation, first-order derivatives with FMO-DFTB (with hybrid orbital projection) may be written as
where \(E_X^{\prime a}\) and \(\varDelta E_{IJ}^{V,a}\) are the terms that come from the derivative of the integrals for \(E_X^{\prime }\) and \(\varDelta E_{IJ}^V\), respectively. \(\mathfrak {R}^a\) represents the response and can be written by
where the Lagrangian is
and the unitary transformation matrix \(U_{ij}^{a,I}\) is relevant to the derivative of the MO coefficient
In FMO-DFTB, the coupled-perturbed (CP) DFTB equation for fragment X can be written as
where
and
\(F_{ij}^{(a),X}\) and \(S_{ij}^{a,X}\) contains the derivative of integrals, and the former also contains the renormalization terms. After solving the Z-vector equation
for all fragments K (\(k \in \mathrm {virt}\), \(l \in \mathrm {occ}\)) self-consistently, the response contribution can be computed by
These expressions are similar to those given for FMO-HF.
Employing AFO further complicates the algorithm. The first complication is due to the frozen and projected orbital terms in the Hamiltonian matrix. This necessitates solving Z-vector equations in dimers and trimers in addition to SCZV equations. The second complication comes from the use of localized orbitals determined in model systems. Consequently, two types of Z-vector equations must be solved, derived from CP localization and the standard CP equations. In total, four types of Z-vector equations are solved, as outlined in Ref. [39], which contains 111 equations.
3.3 Implementation Notes
As all implementations of FMO-DFTB are parallelized with the generalized distributed data interface (GDDI) [43], FMO-DFTB is parallelized to utilize multiple computer nodes. In GDDI, all of the CPU cores are divided into groups and independent jobs (e.g., fragment calculations) are performed in each group. In principle, each group may consist of more than one CPU core and can do jobs in parallel. At present, however, FMO-DFTB is not efficiently parallelized within each group. Therefore, although it is possible to perform hybrid parallelization (GDDI/DDI; similar to MPI/OpenMPI), this is not efficient. Thus, the developers of FMO-DFTB always assign one CPU core per group.
Dispersion corrections are also implemented for FMO-DFTB. Universal force-field (UFF) [44, 45], Slater–Kirkwood [46], and Grimme’s dispersion [47,48,49] models can be combined. Since these dispersion models are not dependent on the electronic structure of the system, the value computed with FMO-DFTB exactly reproduces that of full DFTB (i.e., without fragmentation).
In the most recent implementation, results demonstrated the possibility to significantly eliminate the ESP evaluation for dimers (and trimers). As ESP evaluation scales as quadratic, improving this step is of great importance. ESP for dimers may be written and transformed as
\(\varOmega _{AB}^I\) and \(\varOmega _{AB}^{J}\) are the ESP for fragment I and J, respectively. As they are not dependent on the combination of I and J, they can be constructed before dimer calculations begin. The computation of \(\varOmega _{AB}^{IJ}\) is then reduced to subtracting the \(\varOmega _{AB}^{I(J)}\) and \(\varOmega _{AB}^{J(I)}\) terms. They are calculated in the dimer IJ and thus can be evaluated quickly. A native implementation requires the evaluation of ESP \(N_\mathrm {dimer}\) times, where \(N_\mathrm {dimer}\) is the number of dimers, but the transformation above indicates that the number of the evaluation is decreased to only one. This simplification is only applicable to FMO-DFTB. In addition, if boundary atoms are bond-detached atoms, double counting has to be avoided.
3.4 Computational Efficiency of FMO-DFTB
In 2014, FMO-DFTB scaling for water clusters was reported as \(\mathscr {O} \left( N^{1.21} \right) \) with up to 18,432 atoms. After a number of improvements, new scaling values must be reported. A comparison between the time required to perform a single point gradient calculation at the level of full DFTB3 and FMO2-DFTB3 for the same water clusters is reported in Fig. 1a. The time of full DFTB3 for \(N_\mathrm {at} > 9216\) is an extrapolated estimate. The observed scaling with full DFTB3 is \(\mathscr {O} \left( N^{2.97} \right) \), very close to the theoretical scaling (cubic). The DSYEVD driver, which is likely the fastest LAPACK diagonalization, was employed. FMO2-DFTB3 took only 129.0 s. The observed scaling was \(\mathscr {O} \left( N^{1.45} \right) \), higher than the previously reported scaling. This is because the most time-consuming step in FMO-DFTB became the evaluation of ESP during monomer SCC iteration, which scales as purely quadratic. Monomer SCC and dimer calculations took 48.1 and 20.3 s, respectively. In addition, evaluating the gradients require solving SCZV equations, which are conceptually similar to monomer SCC, so this step also involves the quadratic scaling step. Nevertheless, FMO-DFTB is more than one thousand times faster than full DFTB for the largest system.
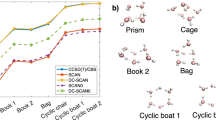
It is also interesting to compare the performance of FMO-DFTB2 and -DFTB3 (Fig. 1b). Overall, FMO-DFTB3 is roughly twice as expensive as FMO-DFTB2, because the evaluation of ESP, which is the most time-consuming step, involves more terms (\(\varGamma \), see Eq. 29). The scaling of FMO-DFTB2 energy is \(\mathscr {O} \left( N^{1.16} \right) \), which is to be compared with the previous scaling \(\mathscr {O} \left( N^{1.21} \right) \) [17]. As mentioned, solving a set of SCZV equations requires a similar computational effort to solving a monomer SCC. Therefore, gradient evaluation requires almost twice the computational effort.
The scaling of the computation of the Hessian with FMO-DFTB was \(\mathscr {O} \left( N^{2.00} \right) \) and \(\mathscr {O} \left( N^{1.98} \right) \) with one and six CPU cores, respectively [36]. With six CPU cores (Xeon E5-1650 v3), the second-order derivative calculation for 10,041 atoms (water) finished in only 17.2 min, whereas normal mode analysis required 262.4 min for computation. The bottleneck of the analysis was the diagonalization of the Hessian matrix. Considering the performance of FMO-DFTB, it could easily be applied to larger systems; however, as the memory requirement scales as quadratic, this will be a severe problem.
4 Selected Applications
4.1 Molecular Dynamics Simulations
Molecular dynamics (MD) simulations with FMO-DFTB have been applied to halogen halides (HF, HCl, HBr, and HI) [34], solvated sodium cation [37], and boron nitride nano-rings [39]. In all cases, the gradient was computed analytically, as gradient inaccuracy introduces artificial effects such as an energy drift in the energy-conservative ensemble. In the first example [34], one million MD steps were performed for systems consisting of 2,000 atoms and the first peak of halogen–halogen radial pair distribution functions obtained experimentally and with FMO-DFTB were compared. Agreement between experimental and computational functions was rather dependent on the chosen DFTB and dispersion models. After many test calculations, FMO-DFTB2 with the UFF-type dispersion model was selected for use. The second example demonstrated a comparative performance of FMO2- and FMO3-DFTB3 [37]. A sodium cation was placed in the center of 473 water molecules; 100 ps MD simulations were performed (100,000 steps). Because the charge in the central sodium cation is prone to delocalize over the surrounding atoms, the inclusion of three-body effects is essential. The coordination numbers obtained with FMO2 and FMO3 were 8.9 and 6.5, respectively; FMO3 gave a value closer to other simulations (5–6), whereas FMO2 significantly overestimated it. This is likely because FMO2 tends to overestimate the binding between fragments in the confined relaxation space. In this simulation, the time step was 1 fs with the help of RATTLE [50] constraints.
The last case employed boron nitride nano-rings [39] and demonstrated that FMO-DFTB may be applicable to MD simulations for one-million-atom systems. Five hundred MD steps over the course of 181 h were performed for the system, which contained 1,180,800 atoms. Unlike other linear-scaling methods, FMO-DFTB is uniquely suited to performing on laboratory-scale computers; the calculation was performed with only 24 CPU cores on a single computer node. Although this simulation was rather short, it demonstrates that, provided more computer resources are available, FMO-DFTB may be applicable to longer and larger simulations and will be a useful tool for understanding the dynamics of large systems.
4.2 Proteins
Due to self-interaction, DFTB cannot accurately describe orbital energies; applying FMO-DFTB to proteins had thus been problematic until it was combined with PCM. Specifically, deriving DFTB with DFT causes the electronic structure parameters to be fitted to results obtained with the PBE exchange–correlation functional. Additionally, DFTB also underestimates the gap between the highest occupied MO (HOMO) and the lowest unoccupied MO (LUMO). This effect is particularly prominent when fragments have a net charge. The use of DFTB3 improved charge–charge interactions, but not necessarily orbital energies.
A typical consequence of self-interaction is that the orbital levels of fragments with positive and negative charges are akin to be low and high, respectively, with FMO-DFTB. This problem does not directly affect calculations during monomer SCC because orbital levels of fragments do not couple directly. However, in the dimer calculation, I and J are merged and this causes a problem. For instance, let’s assume that fragments I and J have a positive and negative charge (both have four electrons), respectively, and that the LUMO of fragment I is lower than the HOMO of fragment J when constructing the dimer IJ. Next, an initial guess can be constructed by placing electrons obeying the Aufbau principle. Now, the atoms that belong to fragments I and J get six and two electrons, respectively, not four and four, resulting in a charge transfer state. Consequently, a charge-transferred electronic structure or a lack of SCF convergence is possible. The situation is schematically depicted in Fig. 2. To combat this, the 1L2Y protein in the original FMO-DFTB paper was neutralized [17].

Reproduced from Ref. [35] by permission of the PCCP Owner Societies
Origin of the problem of charge transfer states in dimers. a Orbital energies in monomers I (red solid lines) and J (blue dashed lines) b Initial levels in dimer IJ c Population of initial levels in dimer IJ
Once FMO-DFTB was combined with PCM, no such switch occurs. The origin of incorrect orbital energies is partially attributed to strong charge–charge interactions within charged residues and could be alleviated by adding solvent screening effects. This problem is not unique to DFTB; as discussed in Ref. [35], a similar problem occurs with GGA and some hybrid functionals. A more systematic solution may employ a long-range corrected functional. At the Hartree–Fock limit, no such problem occurs, as there is no self-interaction error.
There are four FMO2/PCM (PCM[1], PCM\(\langle 1 \rangle \), PCM[1(2)], and PCM[2]) and six (the former four and PCM[1(3)] and PCM[3]) FMO3/PCM levels, depending on the construction of ESP on the apparent surface charge (ASC) and the scaling for the solute–solvent interaction energy. FMO-DFTB can be combined with any PCM variants. The difference is summarized in Table 1 in Ref. [35]. Analytic derivatives have been developed only when combined with PCM\(\langle 1 \rangle \).
With FMO2-DFTB3-D(UFF)/PCM\(\langle 1 \rangle \), medium-sized proteins up to 3,578 atoms could be optimized. The root-mean-square deviations of the largest optimized structure (PDB: 2CGA) compared with the experimental structure was 0.720 Å. This experimental structure was obtained from X-ray crystallography with a resolution of 1.8 Å. Calculations combined with PCM are somewhat slower than those in a vacuum because linear equations must be solved to obtain ASCs. Nevertheless, the observed scaling (FMO2-DFTB3-D/PCM\(\langle 1 \rangle \) energy + gradient) was \(\mathscr {O} \left( N^{1.39} \right) \), and the performance is still useful for medium to large systems.
Although adding solvent effects circumvented the problem, a more promising and fundamental solution is to apply long-range corrections to DFTB. Ref. [35] showed that the HOMO–LUMO gap problem disappeared when long-range corrected functionals were employed in DFT calculations. Ref. [28] demonstrated that LC-DFTB allows gas phase calculations of the proteins for which the conventional DFTB failed because of the underestimation of HOMO–LUMO gap, which originates from the self-interaction error. Preliminary results demonstrate that FMO-LC-DFTB also does not suffer from this problem.
4.3 Chemoinformatics
Apart from FMO-DFTB, DFTB has been often applied to biosystems; detailed expositions may be found in published reviews [4, 5, 10, 51]. Considering that the development of DFTB2 and DFTB3 has been motivated by describing proper interactions within charged systems, DFTB should be a promising tool for biosystems. Moreover, the fast execution of DFTB calculations should be an appealing feature in the area of chemoinformatics, as the structures and properties of thousands of molecules can quickly be calculated with DFTB. One important development in applying DFTB to biosystems is the combination of DFTB with molecular mechanics (MM), DFTB/MM. An early implementation has been realized in 2001 [52] by Cui et al. DFTB/MM was applied to ATP hydrolysis [53, 54] and cytochrom c oxidase [55]. It was also recently applied to evaluate the binding interactions between proteins and ligands, including binding interactions between drugs and H1N1 neutraminidase-1 [56] and docking simulations of zinc-bound ligands [57]. A more practical application of DFTB to chemoinformatics may be found in a study by Qu et al. [58]. Here, DFTB was used to optimize more than 900 molecules to train machine learning models. The trained data could then predict bond dissociation energies of 100 molecules.
Although DFTB is much faster than other ab initio methods, its cubic-scaling computational cost has been a major limitation in applying DFTB to larger and/or a large number of systems. Using the QM/MM approach is very useful, especially when focusing on the reaction center of the protein. However, when treating the whole system on equal footing, e.g., to investigate interactions between a protein and ligands, full DFTB or QM/MM approaches may not be the best choice. Furthermore, the combination with FMO is beneficial for the use of pair interaction energy decomposition analysis (PIEDA). Details of PIEDA are described somewhere in this book. Recently, FMO-DFTB was also combined with PIEDA [59]. In this context, in spite of the aforementioned challenge of orbital energies, FMO-DFTB combined with PCM is seemingly a promising tool to investigate interactions between proteins and ligands. FMO-DFTB was applied to the assessment of receptor-ligand interactions and total interaction energies [60]. Here, the quantities computed with FMO-DFTB were validated by comparison with experimental data and FMO-MP2 results; good correlation was found between total interaction energies computed with FMO-DFTB and FMO-MP2 (\(R^2 > 0.9\)), while the computational cost was reduced by 1000 times.
A similar correlation study was presented by Ref. [35]. Comparing the solvent screening and pair interactions obtained with FMO-DFTB and FMO-DFT (LC-BOP and M11 functionals) showed that the correlations of these parameters were very high with \(R^2\) values greater than 0.97, while the computational cost was reduced by 4,840 times for the 1IO5 protein (1,961 atoms). These pilot demonstrations indicate that FMO-DFTB is potentially very useful in the process of drug discovery in which a number of interacting energies between proteins and ligands must be evaluated with a sufficiently accurate method.
4.4 Vibrational Frequency Analysis
Vibrational frequency analysis was applied using FMO-DFTB in two studies [36, 37]. In both, non-resonance Raman spectra were also simulated. Polarizability derivatives were evaluated by the numerical differentiation of gradients under electric fields. Nakata et al. simulated infrared spectra of the epoxy amine oligomers in a system of more than 1000 atoms and compared them with experimental data [36]. Three characteristic peaks were experimentally observed: 1183, 1260, and 3450 cm\(^{-1}\) resulting from CH\(_3\) groups in isopropylidene, benzene rings, and hydroxyl groups, respectively. The simulation used one chain of epoxy amine (279 atoms) to show that FMO2-DFTB3 predicted a shift of approximately 150 cm\(^{-1}\), but the use of four chains predicted a peak at 3431 cm\(^{-1}\). Systems of this size are not easy to evaluate even with the conventional DFTB, as the scaling of Hessian is formally \(\mathscr {O} \left( N^4 \right) \).
Another study applied FMO3-DFTB to three isomers of polyalanine and solvated sodium cation [37]. The maximum frequency deviation in FMO3-DFTB3 was less than 10 cm\(^{-1}\), compared with the corresponding frequency computed with full DFTB calculations. Comparison of infrared and Raman spectra of the solvated sodium cation showed that FMO3-DFTB had a higher accuracy than FMO2 by a factor of 2.5.
4.5 Charge Transport Materials
DFTB has also been applied to describe the electron transfer mechanism of organic molecules and conductors, DNA, and peptides [61,62,63,64,65,66]. This approach, called “fragment orbital”-DFTB, is similar to the FMO method. It begins with dividing a system into fragments, obtaining HOMOs of each fragment, and then constructing the charge transfer (coupling) integrals by the standard AO–MO transformation:
where I and J are the index of fragments and \(i \in I\) and \(j \in J\) represent the index of HOMOs. On-site energies are the orbital level of the HOMO.
Conceptually, there is a large overlap between the fragment orbital and FMO approaches. FMO uses more well-defined fragments with the help of hybrid orbital projection, allowing the application of it to covalently bonded systems to be straightforward and robust. A similar analysis using the linear combination of fragment molecular orbital (LCMO) approach has been studied previously [67, 68]. Recently, Kitoh-Nishioka et al. applied the LCMO approach to FMO-DFTB and performed detailed analysis of the charge transport properties of covalent organic frameworks by combining classical MD, FMO-DFTB, and carrier dynamics simulations [69]. In estimating charge transfer integrals, the standard DFTB parameters were rather “confined” and thus not suitable for evaluation, so unconfined parameters called “8–\(\infty \)” were employed, similar to previous studies [65, 66]. The confinement of the standard parameters originates from the tight-binding DFTB characters. In spite of the use of unconfined parameters, DFTB still underestimated transfer integrals, and scaling was essential to reproduce LC-BLYP results. Nevertheless, the combination of FMO-DFTB with LCMO should be an important tool for more practical applications using larger systems.
5 Conclusion and Outlook
FMO-DFTB is currently the fastest quantum chemical method among the available FMO-electronic structure method combinations. The scaling of FMO2-DFTB3 is at present \(\mathscr {O}\left( N^{1.45} \right) \) for evaluating the energy gradient and \(\mathscr {O} \left( N^{2.00} \right) \) for evaluating the Hessian. The most favorable scaling was achieved with FMO2-DFTB2, \(\mathscr {O} \left( N^{1.16} \right) \). For a water cluster consisting of 18,432 atoms, a single point gradient calculation with FMO2-DFTB3 finished in 129.0 0 s with only one CPU core. This development has enabled geometry optimizations and short MD simulations for systems consisting of more than one million atoms and vibrational frequency calculations for those of ten thousand atoms. Typical applications are soft matter and polymers including biosystems. A recent development employing AFOs further widened the range of application into the range of materials.
In spite of these advancements, there is still large room for development. Recent developments are still restricted to closed-shell single electron structure and FMO-DFTB still cannot be applied to systems with radicals. FMO-DFTB can be potentially combined with linear-response time-dependent DFTB, which has already been developed in GAMESS-US. Extensions using periodic boundary conditions are of great importance for bulk systems.
The limitations of DFTB caused by the self-interaction inherent in DFT pose serious problems. The underestimation of HOMO–LUMO gaps is attributed to this drawback. Although this problem was circumvented by adding solvent screening effects, a more pragmatic solution lies in further development of long-range corrected DFTB (LC-DFTB). Developments of FMO around LC-DFTB in GAMESS-US are currently in progress [70]. Another limitation comes from the restriction of available parameters. However, qualified parameters are expected to be routinely generated with machine learning or artificial intelligence technologies in the near future.
References
Almlöf J, Faegri Jr, K, Korsell K (1982) Principles for a direct SCF approach to LICAO–MOab-initio calculations. J Comput Chem 3(3):385–399. https://doi.org/10.1002/jcc.540030314
Burow AM, Sierka M, Mohamed F (2009) Resolution of identity approximation for the Coulomb term in molecular and periodic systems. J Chem Phys 131(21):214101. https://doi.org/10.1063/1.3267858
Kitaura K, Ikeo E, Asada T, Nakano T, Uebayasi M (1999) Fragment molecular orbital method: an approximate computational method for large molecules. Chem Phys Lett 313:701–706
Cui Q, Elstner M (2014) Density functional tight binding: values of semi-empirical methods in an ab initio era. Phys Chem Chem Phys 16:14368–14377. https://doi.org/10.1039/C4CP00908H
Gaus M, Cui Q, Elstner M (2014) Density functional tight binding: application to organic and biological molecules. WIREs Comput Mol Sci 4(1):49–61
Gotthard S (2017) Tight-binding density functional theory: an approximate Kohn–Sham DFT scheme. J Phys Chem A 111(26):5609–5613. https://doi.org/10.1021/jp069056r
Koskinen P, Mäkinen V (2009) Density-functional tight-binding for beginners. Comput Mater Sci 47:237–253
Seifert G, Joswig JO (2012) Density-functional tight binding—an approximate density-functional theory method. WIREs Comput Mol Sci 2:456–465
Irle S, Zheng G, Wang Z, Morokuma K (2006) The C\(_{60}\) formation puzzle "solved": QM/MD simulations reveal the shrinking hot giant road of the dynamic fullerene self-assembly mechanism. J Phys Chem B 110(30):14531–14545
Christensen AS, Kubař T, Cui Q, Elstner M (2016) Semiempirical quantum mechanical methods for noncovalent interactions for chemical and biochemical applications. Chem. Rev. 116(9):5301–5337. https://doi.org/10.1021/acs.chemrev.5b00584
Elstner M (2006) The SCC-DFTB method and its application to biological systems. Theor Chem Acc 116:316–325
Porezag D, Frauenheim T, Köhler T, Seifert G, Kaschner R (1995) Construction of tight-binding-like potentials on the basis of density-functional theory: application to carbon. Phys Rev B 51(19):12947–12957
Seifert G, Porezag D, Frauenheim T (1996) Calculations of molecules, clusters, and solids with a simplified LCAO-DFT-LDA scheme. Int J Quantum Chem 58(2):185–192
Elstner M, Porezag D, Jungnickel G, Elsner J, Haugk M, Frauenheim T, Suhai S, Seifert G (1998) Self-consistent-charge density-functional tight-binding method for simulations of complex materials properties. Phys Rev B 58:7260–7268
Foulkes WMC, Haydock R (1989) Tight-binding models and density-functional theory. Phys Rev B 39:12520–12536. https://doi.org/10.1103/PhysRevB.39.12520
Gaus M, Cui Q, Elstner M (2011) DFTB3: extension of the self-consistent-charge density-functional tight-binding method (SCC-DFTB). J Chem Theory Comput 7(4):931–948
Nishimoto Y, Fedorov DG, Irle S (2014) Density-functional tight-binding combined with the fragment molecular orbital method. J Chem Theory Comput 10:4801–4812
Scemama A, Renon N, Rapacioli M (2014) A sparse self-consistent field algorithm and its parallel implementation: application to density-functional-based tight binding. J Chem Theory Comput 10(6):2344–2354. https://doi.org/10.1021/ct500115v
Murat K, Hong Z, Peter Z, Dixon DA, Wagner AF (2016) Shift-and-invert parallel spectral transformation eigensolver: massively parallel performance for density- functional based tight- binding. J Comput Chem 37(4):448–459 . https://doi.org/10.1002/jcc.24254
Niklasson AMN (2017) Next generation extended lagrangian first principles molecular dynamics. J Chem Phys 147(5):054103. https://doi.org/10.1063/1.4985893
Giese TJ, Chen H, Dissanayake T, Giambasu GM, Heldenbrand H, Huang M, Kuechler ER, Lee TS, Panteva MT, Radak BK, York DM (2013) A variational linear-scaling framework to build practical, efficient next-generation orbital-based quantum force fields. J Chem Theory Comput 9:1417–1427
Hiroaki N, Yoshifumi N, Masato K, Stephan I, Hiromi N (2016) Three pillars for achieving quantum mechanical molecular dynamics simulations of huge systems: Divide-and-conquer, density-functional tight-binding, and massively parallel computation. J Comput Chem 37(21):1983–1992. https://doi.org/10.1002/jcc.24419
Hu H, Lu Z, Elstner M, Hermans J, Yang W (2007) Simulating water with the self-consistent-charge density functional tight binding method: from molecular clusters to the liquid state. J Phys Chem A 111:5685–5691
Perdew JP (1986) Density-functional approximation for the correlation energy of the inhomogeneous electron gas. Phys Rev B 33:8822–8824. https://doi.org/10.1103/PhysRevB.33.8822
Perdew JP, Burke K, Ernzerhof M (1996) Generalized gradient approximation made simple. Phys Rev Lett 77:3865–3868. https://doi.org/10.1103/PhysRevLett.77.3865
Lundberg M, Nishimoto Y, Irle S (2012) Delocalization errors in a hubbard-like model: consequences for density-functional tight-binding calculations of molecular systems. Int J Quantum Chem 112(6):1701–1711. https://doi.org/10.1002/qua.23178
Humeniuk A, Mitrić R (2015) Long-range correction for tight-binding TD-DFT. J Chem Phys 143(13):134120. https://doi.org/10.1063/1.4931179
Lutsker V, Aradi B, Niehaus TA (2015) Implementation and benchmark of a long-range corrected functional in the density functional based tight-binding method. J Chem Phys 143(18):184107. https://doi.org/10.1063/1.4935095
Niehaus TA, Della Sala F (2012) Range separated functionals in the density functional based tight-binding method: formalism. Phys Status Solidi B 249(2):237–244. https://doi.org/10.1002/pssb.201100694
Livshits E, Baer R (2007) A well-tempered density functional theory of electrons in molecules. Phys Chem Chem Phys 9:2932–2941. https://doi.org/10.1039/B617919C
Kranz JJ, Elstner M, Aradi B, Frauenheim T, Lutsker V, Garcia AD, Niehaus TA (2017) Time-dependent extension of the long-range corrected density functional based tight-binding method. J Chem Theory Comput 13(4):1737–1747. https://doi.org/10.1021/acs.jctc.6b01243
Nishimoto Y, Fedorov DG, Irle S (2015) Third-order density-functional tight-binding combined with the fragment molecular orbital method. Chem Phys Lett 636:90–96
Nishimoto Y, Nakata H, Fedorov DG, Irle S (2015) Large-scale quantum-mechanical molecular dynamics simulations using density-functional tight-binding combined with the fragment molecular orbital method. J Phys Chem Lett 6(24):5034–5039. https://doi.org/10.1021/acs.jpclett.5b02490
Nishimoto Y, Fedorov DG (2016) The fragment molecular orbital method combined with density-functional tight-binding and the polarizable continuum model. Phys Chem Chem Phys 18:22047–22061. https://doi.org/10.1039/C6CP02186G
Nakata H, Nishimoto Y, Fedorov DG (2016) Analytic second derivative of the energy for density-functional tight-binding combined with the fragment molecular orbital method. J Chem Phys 145(4):044113. https://doi.org/10.1063/1.4959231
Nishimoto Y, Fedorov DG (2017) Three-body expansion of the fragment molecular orbital method combined with density-functional tight-binding. J Comput Chem 38(7):406–418. https://doi.org/10.1002/jcc.24693
Fedorov DG, Jensen JH, Deka RC, Kitaura K (2008) Covalent bond fragmentation suitable to describe solids in the fragment molecular orbital method. J Phys Chem A 112(46):11808–11816. https://doi.org/10.1021/jp805435n
Nishimoto Y, Fedorov DG (2018) Adaptive frozen orbital treatment for the fragment molecular orbital method combined with density-functional tight-binding. J Chem Phys 148(6):064115. https://doi.org/10.1063/1.5012935
Gordon MS, Schmidt MW (2005) Advances in electronic structure theory: GAMESS a decade later. In: Dykstra CE, Frenking G, Kim KS, Scuseria GE (eds) Theory and applications of computational chemistry, the first forty years. Elsevier, Amsterdam, pp 1167–1189
Schmidt MW, Baldridge KK, Boatz JA, Elbert ST, Gordon MS, Jensen JH, Koseki S, Matsunaga N, Nguyen KA, Su S, Windus TL, Dupuis M, Montgomery JA (1993) General atomic and molecular electronic structure system. J Comput Chem 14:1347–1363
Nagata T, Brorsen K, Fedorov DG, Kitaura K, Gordon MS (2011) Fully analytic energy gradient in the fragment molecular orbital method. J Chem Phys 134:124115
Fedorov DG, Olson RM, Kitaura K, Gordon MS, Koseki S (2004) A new hierarchical parallelization scheme: generalized distributed data interface (GDDI), and an application to the fragment molecular orbital method (FMO). J Comput Chem 25:872–880
Rappé AK, Casewit CJ, Colwell KS, Goddard WA III, Skiff WM (1992) UFF, a full periodic table force field for molecular mechanics and molecular dynamics simulations. J Am Chem Soc 114:10024–10035
Zhechkov L, Heine T, Patchkovskii S, Seifert G, Duarte HA (2005) An efficient a posteriori treatment for dispersion interaction in density-functional-based tight binding. J Chem Theory Comput 1:841–847. https://doi.org/10.1021/ct050065y
Elstner M, Hobza P, Frauenheim T, Suhai S, Kaxiras E (2001) Hydrogen bonding and stacking interactions of nucleic acid base pairs: a density-functional-theory based treatment. J Chem Phys 114(12):5149–5155
Grimme S (2006) Semiempirical GGA-type density functional constructed with a long-range dispersion correction. J Comput Chem 27(15):1787–1799. https://doi.org/10.1002/jcc.20495
Grimme S, Antony J, Ehrlich S, Krieg H (2010) A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J Chem Phys 132:154104
Grimme S, Ehrlich S, Goerigk L (2011) Effect of the damping function in dispersion corrected density functional theory. J Comput Chem 32(7):1456–1465. https://doi.org/10.1002/jcc.21759
Andersen HC (1983) Rattle: A “velocity” version of the shake algorithm for molecular dynamics calculations. J Comput Phys 52:24–34
Elstner M, Frauenheim T, Suhai S (2003) An approximate DFT method for QM/MM simulations of biological structures and processes. J Mol Struct: THEOCHEM 632(1):29–41. https://doi.org/10.1016/S0166-1280(03)00286-0
Cui Q, Elstner M, Kaxiras E, Frauenheim T, Karplus M (2001) A QM/MM implementation of the self-consistent charge density functional tight binding (SCC-DFTB) method. J Phys Chem B 105(2):569–585. https://doi.org/10.1021/jp0029109
Lu X, Ovchinnikov V, Demapan D, Roston D, Cui Q (2017) Regulation and plasticity of catalysis in enzymes: insights from analysis of mechanochemical coupling in myosin. Biochemistry 56(10):1482–1497. https://doi.org/10.1021/acs.biochem.7b00016
Yang Y, Yu H, Cui Q (2008) Extensive conformational transitions are required to turn on atp hydrolysis in myosin. J Mol Biol 381(5):1407–1420. https://doi.org/10.1016/j.jmb.2008.06.071
Ghosh N, Prat-Resina X, Gunner MR, Cui Q (2009) Microscopic pKa analysis of Glu286 in cytochrome c oxidase (rhodobacter sphaeroides): toward a calibrated molecular model. Biochemistry 48(11):2468–2485. https://doi.org/10.1021/bi8021284
Yeng- Tseng W, Yu- Ching C (2014) Insights from QM/MM modeling the 3d structure of the 2009 H1N1 influenza a virus neuraminidase and its binding interactions with antiviral drugs. Mol Inf 33(3):240–249. https://doi.org/10.1002/minf.201300117
Chaskar P, Zoete V, Röhrig UF (2014) Toward on-the-fly quantum mechanical/molecular mechanical (QM/MM) docking: development and benchmark of a scoring function. J Chem Inf Model 54(11):3137–3152. https://doi.org/10.1021/ci5004152
Qu X, Latino DA, Aires-de Sousa J (2013) A big data approach to the ultra-fast prediction of dft-calculated bond energies. J Cheminformatics 5(1):34. https://doi.org/10.1186/1758-2946-5-34
Fedorov DG, Kitaura K (2018) Pair interaction energy decomposition analysis for density functional theory and density-functional tight-binding with an evaluation of energy fluctuations in molecular dynamics. J Phys Chem A 122(6):1781–1795. https://doi.org/10.1021/acs.jpca.7b12000
Inaki M, Fedorov GD, Roger R, Michelle S, Andrea T, Bodkin JM, Alexander H (2017) Rapid and accurate assessment of GPCR–ligand interactions using the fragment molecular orbital- based density-functional tight-binding method. J Comput Chem 38(23):1987–1990. https://doi.org/10.1002/jcc.24850
Gillet N, Berstis L, Wu X, Gajdos F, Heck A, de la Lande A, Blumberger J, Elstner M (2016) Electronic coupling calculations for bridge-mediated charge transfer using constrained density functional theory (CDFT) and effective hamiltonian approaches at the density functional theory (DFT) and fragment-orbital density functional tight binding (FODFTB) level. J Chem Theory Comput 12(10):4793–4805. https://doi.org/10.1021/acs.jctc.6b00564
Heck A, Kranz JJ, Elstner M (2016) Simulation of temperature-dependent charge transport in organic semiconductors with various degrees of disorder. J Chem Theory Comput 12(7):3087–3096. https://doi.org/10.1021/acs.jctc.6b00215
Heck A, Kranz JJ, Kubař T, Elstner M (2015) Multi-scale approach to non-adiabatic charge transport in high-mobility organic semiconductors. J Chem Theory Comput 11(11):5068–5082. https://doi.org/10.1021/acs.jctc.5b00719
Kubar T, Woiczikowski PB, Cuniberti G, Elstner M (2008) Efficient calculation of charge-transfer matrix elements for hole transfer in dna. J Phys Chem B 112:7937–7947
Kubas A, Gajdos F, Heck A, Oberhofer H, Elstner M, Blumberger J (2015) Electronic couplings for molecular charge transfer: benchmarking cdft, fodft and fodftb against high-level ab initio calculations. Phys Chem Chem Phys 17:14342–14354. https://doi.org/10.1039/C4CP04749D
Kubas A, Hoffmann F, Heck A, Oberhofer H, Elstner M, Blumberger J (2014) Electronic couplings for molecular charge transfer: benchmarking CDFT, FODFT, and FODFTB against high-level ab initio calculations. J Chem Phys 140(10):104105. https://doi.org/10.1063/1.4867077
Nishioka H, Ando K (2011) Electronic coupling calculation and pathway analysis of electron transfer reaction using ab initio fragment-based method. I. FMO-LCMO approach. J Chem Phys 134(20):204109. https://doi.org/10.1063/1.3594100
Tsuneyuki S, Kobori T, Akagi K, Sodeyama K, Terakura K, Fukuyama H (2009) Molecular orbital calculation of biomolecules with fragment molecular orbitals. Chem Phys Lett 476(1):104–108. https://doi.org/10.1016/j.cplett.2009.05.069
Kitoh-Nishioka H, Welke K, Nishimoto Y, Fedorov DG, Irle S (2017) Multi-scale simulations on charge transport in covalent organic frameworks: Including dynamics of transfer integrals from FMO-DFTB/LCMO. J Phys Chem C 121:17712–17726. https://doi.org/10.1021/acs.jpcc.7b05779
Vuong VQ, Nishimoto Y, Fedorov DG, Sumpter BG, Niehaus TA, Irle S (2019) The fragment molecular orbital method based on long-range corrected density-functional tight-binding. J Chem Theor Comput 15(5): 3008–3020. https://doi.org/10.1021/acs.jctc.9b00108
Acknowledgements
The FMO-DFTB study has been supported by JSPS KAKENHI Grant Numbers 24\(\cdot \)4986, 15H06316, 16K05677, and 17K14436. S.I. acknowledges support by the Laboratory Directed Research and Development (LDRD) Program of Oak Ridge National Laboratory. ORNL is managed by UT-Battelle, LLC, for DOE under contract DE-AC05-00OR22725. We acknowledge fruitful discussions with Drs. Dmitri G. Fedorov, Hiroya Nakata, and Hirotaka Kitoh-Nishioka.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Nishimoto, Y., Irle, S. (2021). The FMO-DFTB Method. In: Mochizuki, Y., Tanaka, S., Fukuzawa, K. (eds) Recent Advances of the Fragment Molecular Orbital Method. Springer, Singapore. https://doi.org/10.1007/978-981-15-9235-5_23
Download citation
DOI: https://doi.org/10.1007/978-981-15-9235-5_23
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-9234-8
Online ISBN: 978-981-15-9235-5
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)