Abstract
The ability to produce volatile compounds is widely distributed among bacteria. A comprehensive summary of volatiles reported to be emitted by different fungal and bacterial species can be found in the mVOC database (http://bioinformatics.charite.de/mvoc). This chapter aims to summarize different features of bacterial phyla, classes, families, and genera present in the mVOC database and review the different habitats where volatile producers have been isolated from. As a result, bacteria belonging to the phylum of Proteobacteria were the most studied mVOC producers. Moreover, soil, marine environments, and the human body turned out to be the main isolation sources of the microbes compiled in the mVOC database. Additionally, general biosynthetic routes from the primary as well as secondary metabolism are presented which ultimately can result in the production of microbial volatile metabolites.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1.1 Phylogeny and Diversity of Bacteria Comprised in the mVOC Database
Life on Earth can be categorized into three different domains: (1) Archaea, (2) Bacteria, and (3) Eukarya. In 1990, Woese et al. were the first to introduce this new taxonomic rank which was found to be superior to the rank of kingdom. According to this model, all life derived from the last universal common ancestor (LUCA) which therefore represents the origin of life. Whereas bacteria and Archaea are closer related to LUCA, Eukarya are thought to have evolved later in the history of the earth (Fig. 1.1).
The three domains of life. LUCA last universal common ancestor. Modified after Madigan et al. (2019)
This chapter will only focus on the domain of bacteria. More specifically, main emphasis will be laid on phyla and genera present in the mVOC database (Lemfack et al. 2018). This database represents the first collection of all microbial volatile organic compounds investigated so far and comprises ca. 2000 compounds produced by roughly 1000 different microorganisms consisting of 600 bacterial and 400 fungal species (Lemfack et al. 2018). Bacterial VOC producers analyzed so far are distributed in 6 phyla (Table 1.1), 15 classes, 34 orders, 74 families, and 125 genera (Fig. 1.2). Since 1012 microbial species are expected to exist on Earth (Larsen et al. 2017; Pedrós-Alió and Manrubia 2016), the wealth of mVOCs still to be explored becomes obvious.
We will first summarize the main characteristics of bacterial phyla, classes, and genera present in the mVOC database, describe their distribution in nature and, furthermore, give a brief overview about the bacterial metabolism focusing on volatile end products.
The research field of bacterial phylogeny undergoes constant reconstructions which is why numbers mentioned in this chapter have to be treated with caution. According to the List of prokaryotic names with standing in nomenclature (LPSN, Parte 2018), the kingdom of bacteria currently comprises 34 different phyla which are listed in Table 1.1.
1.1.1 Proteobacteria
The Proteobacteria represent the largest and phenotypically most diverse division among prokaryotes (Gupta 2000). To date, more than 40 % of all validly published prokaryotic genera belong to this phylum (Kersters et al. 2006). They comprise the majority of Gram-negative bacteria and formerly have been referred to as “purple bacteria and relatives” although only a small fraction possesses purple coloration. This is why the phylum was renamed after the Greek god Proteus who could take on different shapes which reflects the diversity inside this phylum (Murray et al. 1990; Stackebrandt et al. 1988). They consist of more than 200 different genera subdivided into eight classes based on 16S and 23S rRNA sequences: (1) Alphaproteobacteria, (2) Betaproteobacteria, (3) Gammaproteobacteria, (4) Deltaproteobacteria, (5) Epsilonproteobacteria, (6) Acidithiobacillia, (7) Hydrogenophilalia, and (8) Oligoflexia (Boden et al. 2017; Garrity et al. 2005; Nakai et al. 2014; Williams and Kelly 2013). Nevertheless, this classification does not reflect specific morphological or physiological traits that members of the same class have in common (Rizzatti et al. 2017). In contrast, Proteobacteria are very diverse concerning their appearance. They can form rods or cocci and curved, spiral, ring-shaped but also filamentous and sheathed cells. Most of them are motile and possess polar or peritrichous flagella. A special exception is found in the myxobacteria (Deltaproteobacteria) which possess a gliding motility (Kersters et al. 2006). The proteobacterial energy metabolism is also highly diverse including phototrophic genera (e.g., Chromatium, Rhodospirillum) as well as chemoorganotrophs (e.g., Escherichia coli) and chemolithotrophs (e.g., Nitrobacter) (Kersters et al. 2006).
Proteobacteria can be found ubiquitously in nature and are of high biological relevance since they include a vast number of human, animal, and plant pathogens (e.g., Neisseria, Pseudomonas, Shigella, Salmonella, Yersinia, Escherichia, Helicobacter spp.) (Bergey and Holt 2000; Collier et al. 1998; Dworkin et al. 2006; de Vos et al. 2009). Moreover, they represent one of the most abundant phyla in the gut microbiota (Rizzatti et al. 2017). Their cells mostly appear as free-living organisms with some exceptions. The most famous example is the genus Rhizobium which lives in symbiosis with leguminous plants. This Alphaproteobacterium is capable to reduce atmospheric nitrogen to ammonia which the host plants can readily assimilate. In turn, the plant provides the bacteria with photosynthetic products as a nutrient source. But Proteobacteria also appear as intracellular endosymbionts of protozoa and invertebrates (e.g., mussels, insects, and nematodes) (Kersters et al. 2006). Furthermore, it is anticipated that the origin of mitochondria can be traced back to the Alphaproteobacteria (Kersters et al. 2006).
When analyzing Proteobacteria present in mVOC database in detail, it appears that ca. 250 bacterial strains are described as VOC producers and are distributed in all five classes (Fig. 1.2). They are largely dominated by Gammaproteobacteria (150 strains) compared to Beta-(49), Alpha-(29), Delta-(29), and Epsilonproteobacteria (5) (Fig. 1.3a), while there are no data for Acidithiobacillia, Hydrogenophilalia, and Oligoflexia. Nevertheless, these last classes are composed of only few bacterial genera with validly published names. The most investigated genus concerning its VOC emission is Pseudomonas, accounting for one-third of all Gammaproteobacteria listed in the mVOC database (Fig. 1.3d) which is not surprising due to its high clinical relevance (summarized in Lyczak et al. 2000) raising the need for new, minimally invasive diagnostic tools. Other representatives of the Gammaproteobacteria which have been studied regarding their volatile emission are the genera Serratia and Escherichia, belonging to the family of Enterobacteriaceae. Both genera have been described to be good VOC producers (Bunge et al. 2008; Kai et al. 2010; Kai and Piechulla 2010; Umber et al. 2013; Weise et al. 2014; Yu et al. 2000; Zhu and Hill 2013) and important plant growth promoting microorganisms (Devi et al. 2016; Nautiyal et al. 2010; Zaheer et al. 2016) which is why it is even more surprising to see that they are considerably less represented in the mVOC database. This also applies to the important plant symbiont Rhizobium having only one entry in the database.
Distribution of bacterial strains present in mVOC database. The data represent the number of bacterial strains (at least 10) present in each class (a), order (b), family (c), and genus (d) compiled in mVOC database so far. All classes, orders, families, and genera with less than 10 bacterial strains are not shown
1.1.2 Firmicutes
The Firmicutes represent the second largest phylum in the mVOC database. Ca. 170 bacterial strains have been investigated concerning their volatile emission, so far.
Firmicutes mostly consist of low G/C-containing (<50 %), Gram-positive bacteria with rigid or semi-rigid cell walls containing peptidoglycan (Gibbons and Murray 1978). Nevertheless, this phylum also contains single Gram-negative members, e.g., Veillonellaceae and Syntrophomonadaceae. But just like the Proteobacteria, Firmicutes are phenotypically very diverse reflected by high fluctuations in the composition of this phylum (Seong et al. 2018). Diversity is displayed in terms of (1) cell appearance (spherical, straight, curved, helical rods or filaments, flagellated or not, spore-forming as well as non-sporulating forms), (2) metabolism (chemoorganotrophic or anoxygenic photoheterotrophs), and (3) way of life (aerobic, facultative or strict anaerobes, meso-, thermo- and halophiles, growth at neutral pH but also some alkali- and acidophiles) (de Vos et al. 2009). Bacteria belonging to the Firmicutes mainly divide by binary fission and are highly abundant in soil and aquatic environments (Baik et al. 2008; Schleifer 2009). They play a central role in the decomposition of organic matter which is why they are also often part of the normal intestinal flora of mammals (Lee et al. 2009). Nevertheless, they can also lead to the development of severe diseases in humans and animals as well as in plants (Nguyen and Götz 2016).
This large phylum is separated into three different classes: (1) “Bacilli,” (2) “Clostridia,” and (3) Erysipelotrichia (de Vos et al. 2009). Originally, the class of Mollicutes was also included, but due to low support using alternative markers and unique phenotypic properties, it was removed from the Firmicutes later on to form a phylum on its own (Ludwig and Schleifer 2005). According to the mVOC database, the class of “Bacilli” is the most investigated among Firmicutes (137 bacterial strains). “Bacilli” contain only Gram-positive members and can be divided into two orders, Bacillales and Lactobacillales. The most important genera of the Bacillales are Staphylococcus and Bacillus which are also the most investigated Firmicutes concerning their VOC emission with 28 and 27 bacterial strains, respectively.
Bacillus spp. are aerobic or facultative anaerobic, motile, endospore-forming bacteria. They can form rod-shaped, straight, or slightly curved cells and are highly heterogenous concerning their physiology, ecology, and genetics (Logan and de Vos 2009; Slepecky and Hemphill 2006). Bacillus is a very well investigated organism which has been known to mankind for long time, since its life cycle was first described already in 1872 by Cohn.
According to Lory (2014), the genus Staphylococcus currently contains 54 species. Staphylococcus spp. are widely distributed and are frequently found in association with human and animal hosts. Their appearance resembles grape-like clusters resulting from perpendicular division planes during cell division of single cocci (Lory 2014). They are facultative anaerobes which prefer to use respiratory pathways for energy generation rather than fermentation (Lory 2014). A prominent representative of the Staphylococcus genus is Staphylococcus aureus which can cause major infections in humans reaching from abscesses and food poisoning to endocarditis, toxic shock syndrome, and pneumonia (Gordon and Lowy 2008; Kravitz et al. 2005). Especially the methicillin-resistant S. aureus (MRSA) poses big challenges to medical research, due to increased fitness of the pathogen, improved evasion mechanisms of the hosts’ immune system, and unique toxin production (Gordon and Lowy 2008).
From the order Lactobacillales, 25 Lactobacillus and 22 Leuconostoc species have been investigated for VOC emission (Fig. 1.3d). Both genera are Gram-positive, catalase negative, belong to the lactic acid bacteria, and thus are chemoorganotrophs (Schleifer and Ludwig 1995). They grow anaerobically but can also thrive well under microaerophilic conditions and prefer slightly acidic growth conditions (pH 5.5–6.5) (Schleifer and Ludwig 1995). Whereas the genus Lactobacillus contains at least 13 motile species (Neville et al. 2012; Puertas et al. 2014), Leuconostocaceae were characterized as non-motile and are often found in nutrient-rich environments, e.g., green vegetation, roots, and food (Nieminen et al. 2014).
The second major class of Firmicutes found in the mVOC database is the “Clostridia.” In total, only 29 bacterial strains of this class have been investigated regarding their volatile emission so far. The data further indicate that the Clostridiaceae and more specifically the genus Clostridium account for 23 of the aforementioned 29 “Clostridia”-entries in mVOC. Members of the Clostridiaceae contain more than 30 genera that can be found ubiquitously in nature which is not surprising since they can form endospores protecting them from different kinds of stresses. They are Gram-positive, rod-shaped cells that live anaerobically and contain meso-diaminopimelic acid in their peptidoglycan which is used as a diagnostic marker (Stackebrandt 2014).
As already mentioned above, several Firmicutes are known pathogens and it appears in general that only a very low number of pathogenic bacteria were analyzed for mVOC production. Genera like Streptococcus, responsible for oral malodor, are almost not represented (Fig. 1.3d).
1.1.3 Actinobacteria
The Actinobacteria are one of the largest taxonomic units among the major lineages in the domain of bacteria composed of Gram-positive bacteria with high G/C content and typical filamentous morphology (Barka et al. 2015; Dhakal et al. 2017; Ludwig et al. 2012a). They contain mostly free-living organisms which can be found in terrestrial as well as in aquatic and marine ecosystems (Macagnan et al. 2006). In soil, Streptomyces spp. are predominantly found, whereas Salinispora spp. are a prominent example for Actinobacteria living in aquatic environments. Moreover, bacteria of this phylum also live in close symbiosis to plants, like Frankia spp., or can also have pathogenic effects on plants as well as animals (e.g., Corynebacterium, Mycobacterium, and Nocardia spp.) (Barka et al. 2015; Macagnan et al. 2006). Actinobacteria, more specifically members of the genus Bifidobacterium, also affect us humans, since they are among the first bacteria to colonize our intestinal tract and are believed to exert positive health effects although the molecular mechanisms are still not understood (Cronin et al. 2011; O’Callaghan and van Sinderen 2016). Most Actinobacteria live aerobically under mesophilic conditions (optimal temperature between 25 and 30 °C) at a pH ranging from 6 to 9 with an optimum at neutral pH values (Barka et al. 2015).
Phylogenetically, this phylum is divided into six classes: (1) Actinobacteria, (2) Acidimicrobiia, (3) Coriobacteriia, (4) Nitriliruptoria, (5) Rubrobacteria, and (6) Thermoleophilia. The class of Actinobacteria comprises 16 orders including Actinopolysporales, Actinomycetales, Bifidobacteriales, Catenulisporales, Corynebacteriales, Frankiales, Glycomycetales, Jiangellales, Kineosporiales, Micrococcales, Micromonosporales, Propionibacteriales, Pseudonocardiales, Streptomycetales, Streptosporangiales, and Incertae sedis (Ludwig et al. 2012b). Due to its large size, it is not surprising that members of this phylum are highly diverse in terms of morphology, physiology, and metabolic capabilities. Concerning morphology, most Actinobacteria form substrate mycelia in liquid and solid cultures which during growth on solid medium can transform into aerial mycelia to produce asexual exospores (Flärdh and Buttner 2009; van Dissel et al. 2014). Nevertheless, cell shapes can range from coccoid (Micrococcus), rod-coccoid (Arthrobacter spp.), and fragmented (Nocardia) to permanent and highly differentiated, branched mycelia (Streptomyces, Frankia) (Atlas 1997). Often, Actinobacteria are discriminated according to their mycelium color since most strains are capable to produce melanoid pigments which can improve survival and competitiveness of the producers. Additionally, the appearance and structure of exospores can be used to differentiate actinobacterial species (Barka et al. 2015). The most common soil-dwelling genera found are the saprophytic Streptomyces, already accounting for 70%, Nocardia and Micromonospora (Yokota 1997) which play important roles in the decomposition of organic matter, e.g., cellulose and chitin. As a result, they are crucial for the carbon fluxes in nature (Tarkka and Hampp 2008). Moreover, Streptomyces species are used extensively for the discovery of new bioactive secondary metabolites and produce 70–80% of all substances applied in pharmacy or agrochemistry, e.g., antibiotics like actinomycin or streptomycin (Bérdy 2005; Hopwood 2007; Manteca et al. 2008). Secondary metabolite production is most efficient during morphological differentiation of the cells, i.e., during transition from vegetative to aerial growth (Bibb 2005; Granozzi et al. 1990). So far, more than 500 species have been described in the genus Streptomyces (Tarkka and Hampp 2008) of which only 51 were also investigated regarding their volatile emission. Summarizing everything, although Actinobacteria are ubiquitous and a rich source of natural products, they are less investigated concerning volatile emission compared to other phyla or classes with Streptomyces being the most investigated genus, while other genera like Corynebacterium or Actinomyces are significantly less represented in the data compiled so far (Figs. 1.2 and 1.3).
1.1.4 Bacteroidetes
The phylum of Bacteroidetes is composed of Gram-negative, rod-shaped, chemoorganotrophic bacteria which form no spores and are mostly non-motile or possess a gliding motility (Hahnke et al. 2016; McBride and Zhu 2013; Paster et al. 1994; Woese 1987). It includes ca. 7000 species (Thomas et al. 2011) which are, according to the Bergey’s Manual of Systematic Bacteriology, distributed into four different classes: (1) Bacteroidia, (2) Cytophagia, (3) Flavobacteriia, and (4) Sphingobacteriia (Krieg et al. 2011a). Bacteroidetes are widespread in nature and can be found in soil, decaying plant material, freshwater, marine environments, algae, and dairy products (Bernardet and Nakagawa 2006; Reichenbach 2006). Nevertheless, different classes of Bacteroidetes are also distributed differentially. Accordingly, strains of the class Cytophagia were found only in marine habitats or soil, whereas Flavobacteriia colonize more diverse ecosystems, like soil, sediment, freshwater, brackish water, or seawater including some pathogenic species for humans, mammals, and fish (Bernardet 2015; Krieg et al. 2011b). Just like the Actinobacteria, environmental Bacteroidetes are thought to play a role in the decomposition of complex organic matter, e.g., of polysaccharides and proteins (Bowman 2006; Church 2008; Fernández-Gómez et al. 2013; Kirchman 2002). Even more importantly, Bacteroidetes are found in the gastrointestinal (GI) tract of animals as well as humans. Here, they can account for up to 50% of the microbial gut flora which makes them the dominating phylum together with the Firmicutes (Eckburg et al. 2005; Ley et al. 2008; Smith et al. 2006; Thomas et al. 2011). Moreover, recent studies indicated that the ratio between Firmicutes and Bacteroidetes in the gut microbiota raised with increasing body mass index (BMI) suggesting a role of Bacteroidetes in obesity (Barlow et al. 2015; Koliada et al. 2017; Sweeney and Morton 2013).
In terms of volatile emission, Bacteroidetes are rather unexplored so far. Only 48 entries (<10%) can be found in the mVOC database (Fig. 1.2), most of which belong to the class of Bacteroidia (43 bacterial strains) with the main genera being Bacteroides (20 strains) and Prevotella (17 strains) (Fig. 1.3a, d). Both, Bacteroides spp. and Prevotella spp. contain solely anaerobic bacteria, in contrast to the other three Bacteroidetes classes which comprise only aerobic or facultative anaerobic species. Species of the genus Bacteroides are generally considered as friendly commensals, although the term mutualism is much more fitting since both partners are benefitting from each other (Bäckhed et al. 2005). Moreover, this genus alone accounts for ca. 25% of the total human gut microbiota (Wexler 2007). Bacteroides spp. become a part of the human microbiome already very early in life and can be passed from mother to child during vaginal birth making a crucial role for the human body apparent (Reid 2004). Specifically, it has been shown that Bacteroides spp. produce volatile fatty acids which can be reabsorbed from the host through the large intestine and, in turn, be used as an energy source (Hooper et al. 2002). Nevertheless, when Bacteroides escape their native niche, for example, through rupture of the GI tract or surgical interventions, they can cause severe pathologies outside the gut, like formation of abscesses or bacteremia (summarized in Wexler 2007). The genus Prevotella basically fulfils equal functions in the human host. The only difference is that Prevotella spp. are associated with a plant-rich diet as well as with chronic inflammatory conditions which is why gut microbial communities contain either Bacteroides or Prevotella, but not both at the same time (Brook 1998, 2004; Ley 2016; Nagy 2010).
1.1.5 Cyanobacteria
Cyanobacteria are thought to have emerged ca. 3 Ga which was when the transition to oxygenic conditions through the evolution of photosynthesis took place (Schirrmeister et al. 2011). They belong to the most diverse and widely distributed Prokaryotes and can be found in freshwater, marine, terrestrial, planktonic, and benthic habitats as well as in extreme environments, i.e., frozen lakes, hot springs, or salt works (Walter et al. 2017; Whitton 1992). Cyanobacteria are usually Gram-negative but appear much larger than most bacteria ranging from 1 μm for unicellular forms to more than 30 μm for filamentous forms (Singh and Montgomery 2011). In general, species of this phylum are oxygenic phototrophs which makes them independent from specific carbon sources like other bacteria. Nevertheless, also species growing heterotrophically on organic compounds have been reported (Halm et al. 2012; Walter et al. 2017). Furthermore, some secondary metabolites produced by cyanobacteria are potentially toxic and can lead to harmful effects on health during algal blooms (Percival and Williams 2014). Morphologically Cyanobacteria are very diverse. They exist either as unicellular or filamentous forms whereby also the unicells can be bound together by mucilaginous secretions resulting in complex formations (Broady and Merican 2012; Singh and Montgomery 2011). These complexes can also become very large which form mats, crusts, or gelatinous masses on rocks, sediments, soil, and vegetation getting visible with the naked eye (Broady and Merican 2012). Moreover, single cells can differentiate into specialized compartments like heterocysts or akinetes. Heterocysts are rounder, thick-walled cells that are capable to fix atmospheric nitrogen and produce ammonia which can further be used for amino acid biosynthesis making it an available N-source for the surrounding vegetative cells and thus promote growth during nitrogen starvation (Kumar et al. 2010; Meeks and Elhai 2002). Akinetes represent thick-walled resting spores which secure survival of the population under worst conditions (Walter et al. 2017). Moreover, some cyanobacterial species can produce gas vacuoles which allow them to float in aquatic environments or exhibit a gliding motility (Hoiczyk 2000; Lyra et al. 2005; Percival and Williams 2014).
In total, the phylum Cyanobacteria includes ca. 150 genera containing about 2000 species (Percival and Williams 2014). They can be divided into five different orders: (1) Chroococcales, (2) Pleurocapsales, (3) Oscillatoriales, (4) Nostocales, and (5) Stigonematales (Waterbury 2006). In mVOC only a total of 17 validated cyanobacterial strains have been investigated for their volatile emission profiles. Most of them belong to the orders Nostocales (8 strains) and Oscillatoriales (8 strains) and one representative of the Chroococcales. Nostocales include filamentous cyanobacteria that are capable to differentiate into the aforementioned heterocysts and akinetes which makes them truly multicellular organisms since differentiation is irreversible and functional specialization takes place (Komárek and Johansen 2015; Waterbury 2006). They divide by binary fission in one plane at right angles to the long axis of the trichomes (Waterbury 2006). Species belonging to the order of Nostocales listed in mVOC are of the genera Anabaena (1), Calothrix (3), Rivularia (2), and Tolypothrix (2). Oscillatoriales also contain filamentous cyanobacteria which are, in contrast to Nostocales, not differentiated and divide by binary fission in a single plane. Genera found in mVOC are Lyngbya (1), Phormidium (1), Plectonema (3), and Oscillatoria (3). As already mentioned, there is only one representative of the order Chroococcales to be found in mVOC database, Spirulina platensis. Generally, Chroococcales entail only unicellular cyanobacteria that reproduce by binary fission or budding occurring in one to three planes at right angles to one another or in irregular planes. Cell shapes range from single cocci and rods to cell aggregates held together by sheath material, amorphous slime, or capsular material (Waterbury 2006).
1.1.6 Fusobacteria
The Fusobacteria are a rather unexplored bacterial phylum. They consist of Gram-negative, anaerobic, non-sporulating, non-motile bacilli with tapered rod-shaped cells (Brennan and Garrett 2019). Naturally, Fusobacteria species can be found as free-living organisms in marine environments (Ilyobacter spp.) but also in the human oral cavity (Fusobacterium spp.), intestinal and urogenital tracts (Leptotrichia and Sneathia spp.) and in the intestines of fishes and whales (Cetobacterium spp.) (Brennan and Garrett 2019). Interestingly, single Fusobacteria can harbor unique metabolic capabilities, e.g., Psychrilyobacter atlanticus which was shown to break down nitramine explosives (Zhao et al. 2009). Members of this phylum are differentiated in two families: (1) Leptotrichiaceae and (2) Fusobacteriaceae. In mVOC only five representatives of the latter are found including three strains of the species Fusobacterium nucleatum besides one F. necrophorum and F. simiae species each.
Fusobacterium spp. are generally found in the mouth and other mucosal sites inside humans or animals. Their presence in healthy tissues suggest that they are natural constituents of the microbiota (Brennan and Garrett 2019). Especially the genus F. nucleatum caught scientific interest. It is most abundant in the oral cavity and was shown to play a central role in biofilms that contribute to periodontal health and disease. It mediates biofilm formation by serving as a bridge organism between primary (e.g., Streptococcus spp.) and secondary colonizers (e.g., Porphyromonas gingivalis) (Brennan and Garrett 2019; Kolenbrander et al. 2010). Nevertheless, F. nucleatum is also considered as an opportunistic pathogen since it was found to play a role during periodontitis by shaping host responses and increasing its infectivity by other pathogens (Binder Gallimidi et al. 2015). Furthermore, this species was also implicated to be involved in other diseases such as appendicitis (Swidsinski et al. 2011), brain abscesses (Han et al. 2003), osteomyelitis (Gregory et al. 2015), pericarditis (Truant et al. 1983), and chorioamnionitis (Altshuler and Hyde 1988) although the definite role of F. nucleatum in these cases remains unclear. Still, this species is known to promote inflammatory responses by binding/invading diverse cell types in the human body strengthening the notion of F. nucleatum as an opportunistic pathogen (Han et al. 2000; Ikegami et al. 2009; Strauss et al. 2011).
1.2 Natural Habitat of Bacteria in mVOC Database
Among the 600 bacterial strains registered in mVOC database so far, the isolation sites of ca. 500 of them were compiled, while there were no data existing in the literature for the 100 other strains. It turned out that most of the bacteria present in the database were isolated from the rhizosphere (17%), soil (15%), aquatic environment (15%), clinical samples (15%), humans (13.5%), food products (10%), animals (5%), plants (5%), plant waste (3%), and fungi (1.5%) (Fig. 1.4). Although the volatile profiles of microbes are independent of their isolation site, the number of bacteria isolated from animals and investigated for VOC emission remains very low. Likewise, only the volatile profiles of plant-associated bacteria like Pseudomonas, Bacillus, and Burkholderia are commonly studied (Piechulla et al. 2017). Most of these microorganisms are isolated from the rhizosphere/soil, while the upper parts of the plants as well as animal-associated bacteria (e.g., insects) still represent an important unexplored potential.
Altogether, these analyses of the mVOC database reveal that among the huge number of bacterial species existing on Earth (1012; Larsen et al. 2017; Pedrós-Alió and Manrubia 2016) only a very small proportion, i.e., 0.00000006%, can presently be found in the database. Furthermore, it became obvious that from 26 known bacterial phyla (most of them listed in Fig. 1.1) only six have been investigated concerning their volatile emission, so far. Therefore, the plethora of bacteria which remain to be investigated represents a huge potential in the discovery of further information regarding their ecological role(s).
1.3 Bacterial Volatiles Derived from the Primary Metabolism
The emission of volatiles by bacteria is known for decades. These volatiles are mainly generated during the catabolic activity of these microorganisms. In general, these substances are considered either as primary or as secondary metabolites, depending on whether they are produced during the exponential growth phase or during the transition to or in the stationary growth phase, respectively. The vast majority of volatiles is certainly formed during secondary metabolism. Despite their limited number, also primary metabolism volatiles such as CO2 should not be neglected, e.g., when the interaction between organisms is determined (Piechulla 2017). In the following section, the formation of volatiles by bacteria during primary metabolism is described.
The diversity of bacteria is primarily not marked by a large variability in morphology but by a vast number of different metabolic, especially catabolic, pathways. Bacteria can make a living by generation of energy (ATP) for biosynthetic activity using either light (phototrophy) or the oxidation of chemical compounds (chemotrophy) as energy source. Besides organic material (chemoorganotrophy), a number of inorganic molecules can serve as electron donors for energy generation (chemolithotrophy). The carbon source of bacteria is either organic material or CO2. In Table 1.2, the different types of chemotrophic metabolism and the main volatile products are depicted.
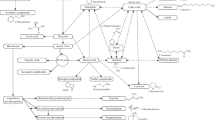
In general, during primary metabolism the ultimate ambition of the bacteria is to generate as much energy (ATP) for growth as possible. In addition, the amount of energy gained during chemotrophic metabolism (oxidation of the substrate) depends on the electron acceptor available or utilizable by a specific bacterium. Some bacteria are very flexible with this respect. Enteric bacteria such as E. coli are a good example for this. They can oxidize glucose, e.g., completely to the inorganic volatile CO2 in the presence of oxygen. If no oxygen but nitrate is on hand, it carries out anaerobic respiration and reduces nitrate to NH3, which also is released into the atmosphere. If no external electron acceptor is present, E. coli switches to fermentation and transfers the electrons to internal acceptors, which finally leads to the excretion of acetate, ethanol, and formate which are altogether organic volatiles (Fig. 1.5). Thus, with respect to the production of volatiles by bacteria the incomplete oxidation of an organic substrate is of special interest, since a greater variety of volatiles might be generated compared to the main volatile CO2 as end product of complete oxidation. In addition to the main alternative electron acceptors nitrate and sulfate listed in Table 1.2, several other compounds can serve as electron sink during growth of certain bacteria, e.g., dimethyl sulfoxide (DMSO) is reduced to dimethyl sulfide and trimethyl amine-N-oxide (TMAO) to trimethyl amine (Fuchs et al. 2007; Madigan et al. 2017). A special metabolic type is the incomplete oxidation of the substrate despite the fact that oxygen is present. Here, a complete oxidation is not possible, since either, an enzyme of the tricarboxylic acid cycle is missing (acidic acid bacteria) or is repressed (bacilli) (Gottschalk 1986). Examples of an incomplete oxidation of glucose and ethanol and the formation of corresponding volatiles are shown in Fig. 1.6. Bacterial fermentation processes lead to the emission of a number of organic volatiles (alcohols, fatty acids, ketones) in addition to the gases CO2 and H2. Figure 1.7 demonstrates in more detail the diversity of volatiles produced from glucose by different fermentative bacteria. Especially clostridia carrying out butyric acid or an acetone–butanol fermentation can emit a rich bouquet of volatiles.
Volatile production caused by incomplete oxidation of the substrate in the presence of oxygen. (a) incomplete oxidation of ethanol by acidic acid bacteria; (b) incomplete oxidation of glucose by acidic acid bacteria; (c) incomplete oxidation of glucose by Bacillus subtilis. Volatiles are highlighted by a red box
In summary, which volatiles (and other products) bacteria produce as primary metabolites depends on the substrate they grow on and on the growth conditions.
1.4 Bacterial Volatiles Derived from the Secondary Metabolism
Many organisms produce metabolites which are not essential for the central processes of growth and development (= primary metabolism). These compounds are referred to as secondary or specialized metabolites, or natural products. They are often unique to individual species or groups of species and mediate interactions with other organisms (defense and attraction arsenal). Typical secondary metabolite classes found in plants are terpenes, cyanogenic glycosides glucosinolates, phenylpropanoids, alkaloids, fatty acid derivatives, S- and N-containing compounds. Chemical convergence of some biosynthetic pathways between plants and insects was documented recently (Beran et al. 2019), while chemical convergences between the microbial and plant and/or animal kingdoms are less well studied, despite the fact that microorganisms release a wealth of secondary metabolites.
One prominent ecological role of specialized compounds released by bacteria is to structure the microbial community and populations living in the same habitat. Many of these compounds are well-known antibiotics that are produced to inhibit the growth of different (microbial) species and are therefore often used in human health care. Most secondary metabolites have unusual structures and their biosyntheses are catalyzed by enzymes that are normally clustered on the chromosome and infrequently on plasmids (e.g., cyt P450 enzymes, glucosyltransferases). Despite the huge variety of chemical structures, the sequence of reactions by which they are made can be grouped into three polymerization reactions:
-
1.
Condensation of acetate-malonate units (polyketide biosynthesis).
-
2.
Condensation of amino acids to oligopeptides (non-ribosomal peptide biosynthesis).
-
3.
Condensation of carbohydrate units (often amino sugars).
The polyketide biosynthetic pathway among prokaryotes is prominent in actinomycetes, but some polyketide compounds are also produced by myxobacteria, cyanobacteria, Bacillus sp., and pseudomonads. Polyketides produced by microorganisms show an extraordinary diversity (Helfrich et al. 2014; Jenke-Kodama and Dittmann 2009), despite its core biosynthesis based on repeated cycles of decarboxylative Claisen-like condensations of simple acyl-CoA building blocks which resemble fatty acid biosynthesis. A multienzyme complex (type II fatty acid biosynthesis) is present in bacteria and plants, while type I single multifunctional FAS are present in invertebrates. Polyketides are typically synthesized by type II PKS. Acetate and malonate or alternatively propionate and methylmalonate form chains in which the keto groups and methylene groups alternate. When methylmalonate is used instead of malonate, the chain becomes branched with methyl groups. The biosynthesis starts with acetyl CoA and malonyl CoA, both bound to the synthase as thioesters (Fig. 1.8a). Acetate (= initiator) binds to the condensing enzyme domain and malonate binds to acyl carrier protein (ACP). Acetate is condensed with the methylene carbon of malonate, while at the same time the carboxyl group of malonate eliminates carbon dioxide resulting in an acetoacetate bound to ACP. In fatty acid biosynthesis three reactions reduce the keto group of acetoacetate to a methylene group. These steps are partially or totally omitted during secondary metabolite biosynthesis. Consequently, the chain can bear keto or hydroxyl groups or double bonds are formed adjacent to methylene groups. The extended chain is then transferred to the condensing enzyme and another malonate-ACP is used for chain elongation.
Polyketide biosynthesis (PKS). (a) First condensation step of the classical fatty acid biosynthesis. (b) A single round of elongation in a type I cis-acyltransferase (AT) PKS module. Examples of modifications and derivatizations are introduced via various domains (red X). In trans-AT PKS, the AT and ER domains are usually missing. (c) Modification reactions, DH dehydratase, ER enoyl reductase, KR ketoreductase, MT methyltransferase, Ox: oxygenase. Modified based on Nguyen et al. (2008) and Meoded et al. (2018)
The complexity and diversification of polyketides is large and depends on (1) the selection of building blocks, (2) the facultative enzymatic modifications, and (3) additional activity of a variety of auxiliary enzymes during or after chain elongation (e.g., dehydratase, enoylreductase, ketoreductase, methyltransferase, oxygenase, Fig. 1.8c, Meoded et al. 2018; Nguyen et al. 2008). Depending on the nature of the enzymes involved as well as altered initiator and extender molecules, the chain can be converted by the aldol reaction into aromatic rings and either linear molecules or macrocyclic rings are formed, subsequently many different structures are produced, e.g., erythromycin, tetracycline, rifamycin, and monensin A. Polyketide synthases (PKS) also encompass different types regarding their enzyme architecture (non-modular, mono-modular, multi-modular) and mode of operations (iterative, non-iterative, cis- and trans-AT). They can also form hybrid enzymes containing components of different PKS classes and/or non-ribosomal peptide synthetases (NRPS) (Helfrich et al. 2014). Very recently, several putative Diels-Alderases (cycloaddition) have been characterized in PKS/NRPS pathways which act in tailoring events (summarized in Scott and Piel 2019).
The vast majority of peptide antibiotics are synthesized by the thiotemplate mechanism (non-ribosomal protein biosynthesis). This process starts with the activation of the amino acids as adenylates, followed by the condensation of the carboxyl group of the amino acid to thiol groups of the non-ribosomal peptide synthases (NRPSs) to form thioesters and stepwise polymerization. The polymerization initiates with the formation of a peptide bond between the carboxyl group of the first amino acid and the amino group of the second amino acid (Fig. 1.9a). These condensation reactions are repeated until the chain is completed. A thioesterase releases the peptide. The NRPSs that catalyze this process can comprise up to four multifunctional enzymes, containing domains which catalyze the activation of an amino acid, its esterification to the thiol group of a pantetheine moiety and the formation of the peptide bond. The enzymes may also encompass domains that catalyze isomerization from l- to d-isomers or methylation of the nitrogen of a newly formed amide, resulting in, e.g., gramicidin biosynthesis (Fig. 1.9). This principle of this process resembles that of polyketide biosynthesis.
Non-ribosomal peptide synthesis (NRPS). (a) Amino acids are linked via thioesters to mono- or multifunctional enzymes. (b) Amino group and carboxyl group of amino acid 2 and 1, respectively, form peptide bonds until the peptide is released by a thioesterase from the enzyme. The mono- as well as multifunctional enzymes may encompass domains which modify the amino acids, e.g., isomerization from l- to d isomer (*)
Antibiotics often contain sugar residues. The biosynthesis of oligosaccharides is identical with the polysaccharide biosynthesis of bacterial cell walls, they are formed by the assembly of monomers, activated as nucleoside diphosphates at the anomeric carbon. Unusual oligosaccharides, often present in antibiotics, either are first assembled and then modified, or sugars are first modified and then stepwise condensed to the precursor.
In contrast to the well-known and established biosynthesis pathways for high molecular weight compounds in microorganisms/bacteria, the biosyntheses of some small molecular compounds of microorganisms, such as terpenes and pyrazines, are less well studied.
Presently, ca. 2000 VOCs released from microorganisms are known (mVOC database, Lemfack et al. 2018) which are categorized into fatty acid derivatives, aromatic compounds, nitrogen-containing compounds, volatiles sulfur compounds, terpenoids and others such as halogenated compounds, and metalloid compounds (summarized in Schulz and Dickschat 2007).
Schenkel et al. (2015) used the mVOC database to quantify and compare compound classes released from microbes and plant roots. It is interesting to note that the same compound classes are found in both headspaces, while the quantitative distribution is different in these organisms. It is presently not studied in detail whether the appearance of identical or similar compounds in both kingdoms are due to evolutionary convergence (in analogy as described by Beran et al. 2019) or due to horizontal gene transfer (Jia et al. 2019). Fatty acids and respective derivatives such as alkanes, alkenes, aldehydes, ketones, alcohols, as well as ethers and esters are most likely products of incomplete oxidations of the primary metabolism. However, typical secondary metabolites are found in the groups of terpenes, aromatic compounds, furans, and S- and N-containing compounds.
Terpene Biosynthesis
Terpenoids are the most diverse class of natural products, 80,000 compounds are estimated to be biosynthesized (Christianson 2017). Monoterpene (C10) and sesquiterpene (C15) compounds are most relevant as volatile organic compounds. Terpene synthases catalyze the most complex chemical reactions in biology since the carbon atoms of the substrates undergo complicated changes in bonding and hybridization during single enzyme catalyzed cyclization reactions. The classical substrates of terpene synthases are geranyl pyrophosphate (GPP), farnesyl pyrophosphate (FPP), and geranylgeranyl pyrophosphate (GGPP) which are synthesized from C5 building blocks (isopentenyl pyrophosphate IPP and dimethylallyl pyrophosphate DMAPP). The latter derive either from the mevalonate or MEP pathway present in the different bacterial species (examples given in Fig. 1.10). Many terpene synthases are very specific and accept only one substrate (single substrate enzymes), while multisubstrate enzymes react with more than one prenyl pyrophosphate. However, the most outstanding and common feature of terpene synthases is their ability to produce multiple products from one substrate (multiproduct enzymes). Often the products are released in defined ratios indicating that precise pathways of biosynthesis are underlying.
Beside the canonical substrates of terpene synthases GPP and FPP and their respective isomers (E,E; Z,Z; E,Z), NPP (neryl pyrophosphate) was described as a substrate (Jia et al. 2018; Sun et al. 2016), and it was shown that methyl-GPP was the substrate for methylisoborneol biosynthesis in Streptomyces coelicolor (Komatsu et al. 2008; Wang and Cane 2008). The latter opened a new route of structural diversity due to the fact that C11 compounds are also substrates for terpene synthases (Kschowak et al. 2018). Such methylation reactions also occur with IPP in Streptomyces species resulting in C6 substrates (Drummond et al. 2019) and FPP as shown in Serratia plymuthica 4Rx13 (Fig. 1.11a; von Reuss et al. 2018). The latter is particularly interesting because the FPP-methyltransferase (FPPMT) not only methylates the C15 FPP to a C16 compound but also performs a cyclisation reaction which is unique for methyltransferases. The product of the FPPMT is presodorifen pyrophosphate and expands the repertoire of non-canonical substrates of terpene synthases uniquely. So far, the methylation and cyclization reactions of IPP, GPP, and FPP (Fig. 1.11b) were only found in the bacterial metabolism.
(a) Biosynthesis of the extraordinary biosynthesis of sodorifen by S. plymuthica (after von Reuss et al. 2018). In a first step a methyltransferase methylates and forms a 5-carbon ring from the canonical substrate FPP; presodorifen is subsequently rearranged to sodorifen by a terpene synthase. (b) Schematic presentation of the canonical and non-canonical terpene biosynthesis to reveal methylated terpene products
Ca. 100 volatile monoterpenes and sesquiterpenes of bacterial origin were summarized by Schulz and Dickschat (2007). Several genome mining approaches were performed (Cane and Ikeda 2012; Yamada et al. 2012), however, up to now only 63 bacterial terpene synthases, primarily sesquiterpene synthases from Streptomyces species, have been isolated (Dickschat 2016). The architecture of bacterial terpene synthases is distinct compared to respective plant enzymes which typically are built of alpha, beta, and gamma domains, while respective bacterial enzymes are comprised of either single or double alpha domains or beta-gamma domains. The “alpha-only” type is most prevalent in bacteria. Furthermore, the characteristic aspartate-rich motive of plant terpene synthases (DDxxD) is slightly altered to DDxxxD in bacteria (Jia et al. 2018, 2019).
Aromatic Compounds
The basal biosynthetic pathway for aromatic secondary metabolites in plants and bacteria is the shikimate pathway whose primary products are the aromatic amino acids tyrosine, phenylalanine, and tryptophane. While the phenylpropane biosynthesis is very widespread and common in plants, the aromatic compound biosynthesis is (to date) not universally observed and well-studied in bacteria. However, compounds like 2-phenylethanol, phenol, benzyl alcohol, methyl benzoate, benzaldehyde, acetophenone, and closely related compounds were shown to be released from several bacteria (summarized by Schulz and Dickschat 2007, mVOC database Lemfack et al. 2018). Two alternative pathways (phenylalanine lyase pathway and phenylpyruvate–phenylacetate–phenylglycolate pathway, Figs. 1.12 and 1.13, respectively) are known to be involved in the biosynthesis of the above-mentioned compounds. However, it is difficult to distinguish these two pathways by feeding experiments with isotope labelled intermediates, and in many cases the bacteria developed individually altered enzymatic reactions supporting and expanding these general pathways. While benzaldehyde can be biosynthesized via both pathways, it seems straightforward that many bacteria synthesize β-phenylethanol via phenylacetaldehyde.
Pyrazines are a special class of volatile heterocyclic compounds that are often produced by many bacteria during fermentation processes (mVOC database Lemfack et al. 2018).
They exhibit strong odor properties and are therefore used as flavoring compounds from the aroma industry. Particularly widespread are pyrazines with one to four methyl or ethyl groups (simple alkylated pyrazines). Caution has to be taken because these pyrazines may also originate from cultivation media or are formed during heating or autoclaving. Pyrazines with longer side chains are less often found as bacterial volatiles. To clarify the biosynthesis of pyrazines, more research is needed. At present, one non-enzymatic pathway via amination of acyloins is postulated for the biosynthesis of simple pyrazines. This biosynthesis results in the formation of aminocarbonyl compounds. Condensation of two aminocarbonyl compounds leads to unstable dihydropyrazines (Fig. 1.14) which are easily oxidized to pyrazines. Higher alkylpyrazines require enzymatic reactions and derive from amino acids (Fig. 1.15). The methoxy methyl-group originates from S-adenosyl methionine.
Indole is a very prominent volatile biosynthesized by E. coli, other Enterobacteriaceae such as Klebsiella and Enterobacter, and other bacteria as well (e.g., Loktanella). It derives from a one-step reaction of the enzyme tryptophanase of the aromatic amino acid tryptophane (Fig. 1.16). The malodourous skatole also derives from tryptophane; it is released from Calothrix and biosynthesized via indole acetic acid (IAA) which is a well-known phytohormone.
N-Compounds
Ammonia is an inorganic, highly volatile compound and produced by many bacteria, including Enterobacter, Serratia, Klebsiella, Staphylococcus, Micrococcus, and Bacillus by degradation of amino acids, by nitrite ammonification, by urease-mediated hydrolytic degradation of urea, and by decarboxylation of amino acids (summarized in Piechulla et al. 2017). Biogenic amines such as trimethylamine, 2-methylpropylamine, 2-methylbutylamine, 3-methylbutylamine, cyclohexylamine, and phenylethylamine are also often found in the headspace of bacteria.
Biogenic amines can be converted by aminooxidases to respective aldehydes which furthermore can react with biogenic amines to produce imines.
S-Compounds
Dimethyl disulfide (DMDS) and dimethyl trisulfide (DMTS) are the most prominent volatile sulfur organic compounds released from bacteria (mVOC database, Lemfack et al. 2018). Three major pathways are known for their biosynthesis (Fig. 1.17). Marine bacteria (Alpha-, Beta-, Gamma-, Deltaproteobacteria) mainly use the dimethylsulfoniopropionate (DMSP) which is produced by algae in high amounts from l-methionine and is therefore prevalent in oceans (Fig. 1.17a). Depending on the bacterial species and its genetic repertoire, DMSP is converted to acrylate and DMS via CoA ester and acyl-CoA transferase, lytically cleaved or degraded by an unknown mechanism of DMSP lyase. Alternatively, DMSP can also be degraded on the demethylation pathway to 3-(methylmercapto) propionic acid by the DMSP demethylase and further to methanethiol and acrylate by an unknown enzyme (Dickschat et al. 2010). In freshwater habitats, bacteria produce methanethiol and DMS through methylation of inorganic sulfide (Fig. 1.17b). In a first step, sulfate is reduced to sulfite by sulfite reductase, which is an evolutionary old reaction/enzyme also present in archaea.
Three biosynthetic pathways for volatile sulfur compounds in bacteria. (a) Dimethylsulfoniopropionate (DMSP) from bacteria of marine or estuarine habitats is converted into acrylate and dimethyl sulfide (DMS). (b) Bacteria in freshwater mainly produce methanethiol from sulfate. (c) l-Methionine is degraded by methionine lyase to ammonia, 2-oxobutyrate, and methanethiol or transaminated to ketomethylthiobutyric acid (KMBA). The latter may be converted by a proposed demethiolase to 2-oxobutyrate and methanethiol. KMBA can be chemically (Mn2+, O2) degraded to methyl mercapto acetaldehyde and subsequently converted to methanethiol and acetaldehyde, or to 3-(methylsulfanyl) propanal and several derivatives. (d) Methanethiol is a precursor for DMS, dimethyl disulfide (DMDS), dimethyl trisulfide (DMTS)
Methanethiol and DMS are then either produced by a SAM dependent methyltransferase or via methoxylated aromatic compounds (e.g. in, Halophaga foetida, Sporobacter termitidis, Sporobacterium olearium; and Parasporobacetrium paucivorans, respectively). Finally, L-methionine is the major source for volatile sulfur compounds of dairy product producing bacteria such as Brevibacterium, Corynebacterium, Staphylococcus, Lactococcus, and Lactobacillus. These bacteria produce a wide array of sulfur compounds biosynthesized via two methionine degradation pathways, (1) direct cleavage of methionine and (2) transamination to ketomethylthiobutyric acid (KMBA) and subsequent reductive demethiolation or decarboxylation (Fig. 1.17c). In the direct cleavage pathway methionine lyase produces ammonia, 2-oxobutyrate and methanethiol. The second pathway is initiated by a transaminase reaction and demethiolase to reveal 2-oxobutyrate and methanethiol, or alternatively KMBA is converted to 3-(methylsulfanyl) propanal, which is the precursor for subsequent reduction and oxidation reactions forming sulfur compound derivatives. Methanethiol can be converted by rapid autooxidation to DMDS, or by reaction with H2S to DMTS. Alternatively, two molecules of DMDS can be transformed by disproportionation to DMS and DMTS (Fig. 1.17d).
Taken together, investigating more bacterial strains, even those from less representative phyla, classes, families, or genera, could help to discover new interesting and bioactive natural products, especially those derived from the secondary metabolism. These could, in turn, be used for new applications since mVOCs are seen as new frontier in bioprospecting and can be applied as eco-friendly alternatives to synthetic compounds for biotechnological applications.
References
Altshuler G, Hyde S (1988) Clinicopathologic considerations of fusobacteria chorioamnionitis. Acta Obstet Gynecol Scand 67:513–517. https://doi.org/10.3109/00016348809029862
Atlas RM (1997) Principles of microbiology, 2nd edn. WCB Brown, Dubuque, IA
Bäckhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI (2005) Host-bacterial mutualism in the human intestine. Science 307:1915–1920. https://doi.org/10.1126/science.1104816
Baik KS, Park SC, Kim EM, Bae KS, Ahn J-H, Ka J-O, Chun J, Seong CN (2008) Diversity of bacterial community in freshwater of Woopo wetland. J Microbiol 46:647–655. https://doi.org/10.1007/s12275-008-0135-x
Barka EA, Vatsa P, Sanchez L, Gaveau-Vaillant N, Jacquard C, Klenk H-P, Clément C, Ouhdouch Y, van Wezel GP, Meier-Kolthoff JP (2015) Taxonomy, physiology, and natural products of Actinobacteria taxonomy, physiology, and natural products of Actinobacteria. Microbiol Mol Biol Rev 80:1–43
Barlow GM, Yu A, Mathur R (2015) Role of the gut microbiome in obesity and diabetes mellitus. Nutr Clin Pract 30:787–797. https://doi.org/10.1177/0884533615609896
Beran F, Köllner TG, Gershenzon J, Tholl D (2019) Chemical convergence between plants and insects: biosynthetic origins and functions of common secondary metabolites. New Phytol 223:52–67. https://doi.org/10.1111/nph.15718
Bérdy J (2005) Bioactive microbial metabolites. J Antibiot 58:1–26. https://doi.org/10.1038/ja.2005.1
Bergey DH, Holt JG (eds) (2000) Bergey’s manual of determinative bacteriology, 9. [Nachdr.]. Lippincott Williams & Wilkins, Philadelphia
Bernardet J-F (2015) Flavobacteriaceae. In: Whitman WB (ed) Bergey’s manual of systematics of archaea and bacteria. Wiley, Hoboken, NJ, pp 1–18
Bernardet JF, Nakagawa Y (2006) An introduction to the family Flavobacteriaceae. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 7. Springer, New York
Bibb MJ (2005) Regulation of secondary metabolism in streptomycetes. Curr Opin Microbiol 8:208–215. https://doi.org/10.1016/j.mib.2005.02.016
Binder Gallimidi A, Fischman S, Revach B, Bulvik R, Maliutina A, Rubinstein AM, Nussbaum G, Elkin M (2015) Periodontal pathogens Porphyromonas gingivalis and Fusobacterium nucleatum promote tumor progression in an oral-specific chemical carcinogenesis model. Oncotarget 6:22613–22623. https://doi.org/10.18632/oncotarget.4209
Boden R, Hutt LP, Rae AW (2017) Reclassification of Thiobacillus aquaesulis (Wood & Kelly, 1995) as Annwoodia aquaesulis gen. nov., comb. nov., transfer of Thiobacillus (Beijerinck, 1904) from the Hydrogenophilales to the Nitrosomonadales, proposal of Hydrogenophilalia class. nov. within the ‘Proteobacteria’, and four new families within the orders Nitrosomonadales and Rhodocyclales. Int J Syst Evol Microbiol 67:1191–1205. https://doi.org/10.1099/ijsem.0.001927
Bowman JP (2006) The marine clade of the family Flavobacteriaceae: the genera Aequorivita, Arenibacter, Cellulophaga, Croceibacter, Formosa, Gelidibacter, Gillisia, Maribacter, Mesonia, Muricauda, Polaribacter, Psychroflexus, Psychroserpens, Robiginitalea, Salegentibacter, Tenacibaculum, Ulvibacter, Vitellibacter and Zobellia. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 7. Springer, New York, pp 677–694
Brennan CA, Garrett WS (2019) Fusobacterium nucleatum – symbiont, opportunist and oncobacterium. Nat Rev Microbiol 17:156–166. https://doi.org/10.1038/s41579-018-0129-6
Broady PA, Merican F (2012) Phylum cyanobacteria: blue-green bacteria, blue-green algae. In: Gordon DP (ed) New Zealand inventory of biodiversity. Canterbury University Press, Christchurch, pp 50–69
Brook I (1998) Microbiology of common infections in the upper respiratory tract. Prim Care 25:633–648
Brook I (2004) Anaerobic pulmonary infections in children. Pediatr Emerg Care 20:636–640
Bunge M, Araghipour N, Mikoviny T, Dunkl J, Schnitzhofer R, Hansel A, Schinner F, Wisthaler A, Margesin R, Märk TD (2008) On-line monitoring of microbial volatile metabolites by proton transfer reaction-mass spectrometry. Appl Environ Microbiol 74:2179–2186. https://doi.org/10.1128/AEM.02069-07
Cane DE, Ikeda H (2012) Exploration and mining of the bacterial terpenome. Acc Chem Res 45:463–472. https://doi.org/10.1021/ar200198d
Christianson DW (2017) Structural and chemical biology of terpenoid cyclases. Chem Rev 117(17):11570–11648
Church MJ (2008) Resource control of bacterial dynamics in the sea. In: Kirchman DL (ed) Microbial ecology of the oceans, 2nd edn. Wiley-Blackwell, Hoboken, pp 335–382
Cohn F (1872) Untersuchungen über Bacterien. Beiträge zur Biologie der Pflanzen 1:127–224
Collier L, Balows A, Sussman M (eds) (1998) Topley & Wilson’s microbiology and microbial infections: volume 2: systematic bacteriology, 9th edn. London, Arnold
Cronin M, Ventura M, Fitzgerald GF, van Sinderen D (2011) Progress in genomics, metabolism and biotechnology of bifidobacteria. Int J Food Microbiol 149:4–18. https://doi.org/10.1016/j.ijfoodmicro.2011.01.019
de Vos P, Garrity GM, Jones D, Bergey DH (eds) (2009) Bergey’s manual of systematic bacteriology: volume 3: the Firmicutes, 2nd edn. Dordrecht, London, Springer
Devi KA, Pandey P, Sharma GD (2016) Plant growth-promoting endophyte Serratia marcescens AL2-16 enhances the growth of Achyranthes aspera L., a medicinal plant. Hayati J Biosci 23:173–180. https://doi.org/10.1016/j.hjb.2016.12.006
Dhakal D, Pokhrel AR, Shrestha B, Sohng JK (2017) Marine rare actinobacteria: isolation, characterization, and strategies for harnessing bioactive compounds. Front Microbiol 8:1106. https://doi.org/10.3389/fmicb.2017.01106
Dickschat JS (2016) Bacterial terpene cyclases. Nat Prod Rep 33:87–110. https://doi.org/10.1039/C5NP00102A
Dickschat JS, Zell C, Brock NL (2010) Pathways and substrate specificity of DMSP catabolism in marine bacteria of the Roseobacter clade. Chembiochem 11:417–425. https://doi.org/10.1002/cbic.200900668
Drummond L, Kschowak MJ, Breitenbach J, Wolff H, Shi Y-M, Schrader J, Bode HB, Sandmann G, Buchhaupt M (2019) Expanding the isoprenoid building block repertoire with an IPP methyltransferase from Streptomyces monomycini. ACS Synth Biol 8:1303–1313. https://doi.org/10.1021/acssynbio.8b00525
Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) (2006) The prokaryotes. Springer, New York
Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Gill SR, Nelson KE, Relman DA (2005) Diversity of the human intestinal microbial flora. Science 308:1635–1638. https://doi.org/10.1126/science.1110591
Fernández-Gómez B, Richter M, Schüler M, Pinhassi J, Acinas SG, González JM, Pedrós-Alió C (2013) Ecology of marine Bacteroidetes: a comparative genomics approach. ISME J 7:1026–1037. https://doi.org/10.1038/ismej.2012.169
Flärdh K, Buttner MJ (2009) Streptomyces morphogenetics: dissecting differentiation in a filamentous bacterium. Nat Rev Microbiol 7:36–49. https://doi.org/10.1038/nrmicro1968
Fuchs G, Schlegel HG, Eitinger T (2007) Allgemeine Mikrobiologie: 53 Tabellen, 8., vollst. überarb. und erw. Auflage Thieme, Stuttgart
Garrity GM, Brenner DJ, Krieg NR, Staley JT, Bergey DH (2005) Bergey’s manual of systematic bacteriology: volume 2: the Proteobacteria, 2nd edn. Springer, New York
Gibbons NE, Murray RGE (1978) Proposals concerning the higher taxa of Bacteria. Int J Syst Bacteriol 28:1–6
Gordon RJ, Lowy FD (2008) Pathogenesis of methicillin-resistant Staphylococcus aureus infection. Clin Infect Dis 46:S350–S359. https://doi.org/10.1086/533591
Gottschalk G (1986) Bacterial metabolism, Springer series in microbiology, 2nd edn. Springer, New York
Granozzi C, Billetta R, Passantino R, Sollazzo M, Puglia AM (1990) A breakdown in macromolecular synthesis preceding differentiation in Streptomyces coelicolor A3(2). J Gen Microbiol 136:713–716. https://doi.org/10.1099/00221287-136-4-713
Gregory SW, Boyce TG, Larson AN, Patel R, Jackson MA (2015) Fusobacterium nucleatum osteomyelitis in 3 previously healthy children: a case series and review of the literature. J Pediatric Infect Dis Soc 4:e155-9. https://doi.org/10.1093/jpids/piv052
Gupta RS (2000) The phylogeny of proteobacteria: relationships to other eubacterial phyla and eukaryotes. FEMS Microbiol Rev 24:367–402.https://doi.org/10.1111/j.1574-6976.2000.tb00547.x
Hahnke RL, Meier-Kolthoff JP, García-López M, Mukherjee S, Huntemann M, Ivanova NN, Woyke T, Kyrpides NC, Klenk H-P, Göker M (2016) Genome-based taxonomic classification of Bacteroidetes. Front Microbiol:7. https://doi.org/10.3389/fmicb.2016.02003
Halm H, Lam P, Ferdelman TG, Lavik G, Dittmar T, LaRoche J, D'Hondt S, Kuypers MMM (2012) Heterotrophic organisms dominate nitrogen fixation in the South Pacific Gyre. ISME J 6:1238–1249. https://doi.org/10.1038/ismej.2011.182
Han YW, Shi W, Huang GT, Kinder Haake S, Park NH, Kuramitsu H, Genco RJ (2000) Interactions between periodontal bacteria and human oral epithelial cells: Fusobacterium nucleatum adheres to and invades epithelial cells. Infect Immun 68:3140–3146. https://doi.org/10.1128/iai.68.6.3140-3146.2000
Han XY, Weinberg JS, Prabhu SS, Hassenbusch SJ, Fuller GN, Tarrand JJ, Kontoyiannis DP (2003) Fusobacterial brain abscess: a review of five cases and an analysis of possible pathogenesis. J Neurosurg 99:693–700. https://doi.org/10.3171/jns.2003.99.4.0693
Helfrich EJN, Reiter S, Piel J (2014) Recent advances in genome-based polyketide discovery. Curr Opin Biotechnol 29:107–115. https://doi.org/10.1016/j.copbio.2014.03.004
Hoiczyk E (2000) Gliding motility in cyanobacterial: observations and possible explanations. Arch Microbiol 174:11–17
Hooper LV, Midtvedt T, Gordon JI (2002) How host-microbial interactions shape the nutrient environment of the mammalian intestine. Annu Rev Nutr 22:283–307. https://doi.org/10.1146/annurev.nutr.22.011602.092259
Hopwood DA (2007) Streptomyces in nature and medicine: the antibiotic makers. Oxford University Press, Oxford, New York
Ikegami A, Chung P, Han YW (2009) Complementation of the fadA mutation in Fusobacterium nucleatum demonstrates that the surface-exposed adhesin promotes cellular invasion and placental colonization. Infect Immun 77:3075–3079. https://doi.org/10.1128/IAI.00209-09
Jenke-Kodama H, Dittmann E (2009) Bioinformatic perspectives on NRPS/PKS megasynthases: advances and challenges. Nat Prod Rep 26(7):874
Jia Q, Kollner TG, Gershenzon J, Chen F (2018) MTPSLs: new Terpene synthases in nonseed plants. Trends Plant Sci 23:121–128. https://doi.org/10.1016/j.tplants.2017.09.014
Jia Q, Chen X, Köllner TG, Rinkel J, Fu J, Labbé J, Xiong W, Dickschat JS, Gershenzon J, Chen F (2019) Terpene synthase genes originated from bacteria through horizontal gene transfer contribute to terpenoid diversity in fungi. Sci Rep 9. https://doi.org/10.1038/s41598-019-45532-1
Kai M, Piechulla B (2010) Impact of volatiles of the rhizobacteria Serratia odorifera on the moss Physcomitrella patens. Plant Signal Behav 5:444–446. https://doi.org/10.4161/psb.5.4.11340
Kai M, Crespo E, Cristescu SM, Harren FJM, Francke W, Piechulla B (2010) Serratia odorifera: analysis of volatile emission and biological impact of volatile compounds on Arabidopsis thaliana. Appl Microbiol Biotechnol 88:965–976. https://doi.org/10.1007/s00253-010-2810-1
Kersters K, de Vos P, Gillis M, Swings J, Vandamme P, Stackebrandt E (2006) Introduction to the Proteobacteria. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 5. Springer, New York
Kirchman DL (2002) The ecology of Cytophaga-Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39:91–100. https://doi.org/10.1111/j.1574-6941.2002.tb00910.x
Kolenbrander PE, Palmer RJ, Periasamy S, Jakubovics NS (2010) Oral multispecies biofilm development and the key role of cell-cell distance. Nat Rev Microbiol 8:471–480. https://doi.org/10.1038/nrmicro2381
Koliada A, Syzenko G, Moseiko V, Budovska L, Puchkov K, Perederiy V, Gavalko Y, Dorofeyev A, Romanenko M, Tkach S, Sineok L, Lushchak O, Vaiserman A (2017) Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol 17. https://doi.org/10.1186/s12866-017-1027-1
Komárek J, Johansen JR (2015) Chapter 4 – filamentous cyanobacteria. In: Wehr JD, Sheath RG, Kociolek JP (eds) Freshwater algae of North America: ecology and classification, 2nd edn. Elsevier, Academic Press, Amsterdam, pp 135–235
Komatsu M, Tsuda M, Omura S, Oikawa H, Ikeda H (2008) Identification and functional analysis of genes controlling biosynthesis of 2-methylisoborneol. Proc Natl Acad Sci U S A 105:7422–7427. https://doi.org/10.1073/pnas.0802312105
Kravitz GR, Dries DJ, Peterson ML, Schlievert PM (2005) Purpura fulminans due to Staphylococcus aureus. Clin Infect Dis 40:941–947. https://doi.org/10.1086/428573
Krieg NR, Whitman WB, Bergey DH (eds) (2011a) Bergey’s manual of systematic bacteriology: volume 4: the Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes, vol 4, 2nd edn. Springer, New York
Krieg NR, Ludwig W, Euzéby J, Whitman WB (2011b) Phylum XIV. Bacteroidetes phyl. nov. In: Krieg NR, Whitman WB, Bergey DH (eds) Bergey’s manual of systematic bacteriology: volume 4: the Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes, vol 47, 2nd edn. Springer, New York, pp 25–469
Kschowak MJ, Wortmann H, Dickschat JS, Schrader J, Buchhaupt M (2018) Heterologous expression of 2-methylisoborneol / 2 methylenebornane biosynthesis genes in Escherichia coli yields novel C11-terpenes. PLoS One 13:e0196082. https://doi.org/10.1371/journal.pone.0196082
Kumar K, Mella-Herrera RA, Golden JW (2010) Cyanobacterial heterocysts. Cold Spring Harb Perspect Biol 2:a000315. https://doi.org/10.1101/cshperspect.a000315
Larsen BB, Miller EC, Rhodes MK, Wiens JJ (2017) Inordinate fondness multiplied and redistributed: the number of species on earth and the new pie of life. Q Rev Biol 92:229–265. https://doi.org/10.1086/693564
Lee DY, Seo Y-S, Rayamajhi N, Kang ML, Lee SI, Yoo HS (2009) Isolation, characterization, and evaluation of wild isolates of Lactobacillus reuteri from pig feces. J Microbiol 47:663–672. https://doi.org/10.1007/s12275-009-0124-8
Lemfack MC, Gohlke B-O, Toguem SMT, Preissner S, Piechulla B, Preissner R (2018) mVOC 2.0: a database of microbial volatiles. Nucleic Acids Res 46:D1261–D1265. https://doi.org/10.1093/nar/gkx1016
Ley RE (2016) Gut microbiota in 2015: Prevotella in the gut: choose carefully. Nat Rev Gastroenterol Hepatol 13:69–70. https://doi.org/10.1038/nrgastro.2016.4
Ley RE, Hamady M, Lozupone C, Turnbaugh P, Ramey RR, Bircher JS, Schlegel ML, Tucker TA, Schrenzel MD, Knight R, Gordon JI (2008) Evolution of mammals and their gut microbes. Science 320:1647–1651. https://doi.org/10.1126/science.1155725
Logan NA, de Vos P (2009) Genus I. Bacillus Cohn 1872, 174AL. In: de Vos P, Garrity GM, Jones D, Bergey DH (eds) Bergey’s manual of systematic bacteriology: volume 3: the Firmicutes, 2nd edn. Springer, Dordrecht, London
Lory S (2014) The family Staphylococcaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: Firmicutes and Tenericutes, 4th edn. Springer, Berlin, Heidelberg, pp 364–366
Ludwig W, Schleifer KH (2005) Molecular phylogeny of bacteria based on comparative sequence analysis of conserved genes. In: Sapp J (ed) Microbial phylogeny and evolution: concepts and controversies. Oxford University Press, New York, pp 70–98
Ludwig W, Euzéby J, Schumann P, Busse H-J, Trujillo ME, Kämpfer P, Whitman WB (2012a) Road map of the phylum Actinobacteria. In: Goodfellow M, Kämpfer P, Busse H-J, Trujillo ME, Suzuki K-I (eds) Bergey’s manual of systematic bacteriology, vol 57. Springer, New York, pp 1–28
Ludwig W, Euzéby J, Whitman WB (2012b) Taxonomic outline of the phylum Actinobacteria. In: Goodfellow M, Kämpfer P, Busse H-J, Trujillo ME, Suzuki K-I, Ludwig W, Whitman WB (eds) Bergey’s manual® of systematic bacteriology: volume five: the Actinobacteria, part A and B, 2nd edn. Springer, New York
Lyczak JB, Cannon CL, Pier GB (2000) Establishment of Pseudomonas aeruginosa infection: lessons from a versatile opportunist1. Microbes Infect 2(9):1051–1060
Lyra C, Laamanen M, Lehtimäki JM, Surakka A, Sivonen K (2005) Benthic cyanobacteria of the genus Nodularia are non-toxic, without gas vacuoles, able to glide and genetically more diverse than planktonic Nodularia. Int J Syst Evol Microbiol 55:555–568. https://doi.org/10.1099/ijs.0.63288-0
Macagnan D, Romeiro RS, de Souza JT, Pomella AWV (2006) Isolation of actinomycetes and endospore-forming bacteria from the cacao pod surface and their antagonistic activity against the witches’ broom and black pod pathogens. Phytoparasitica 34:122–132. https://doi.org/10.1007/BF02981312
Madigan MT, Bender KS, Buckley DH, Sattley WM, Stahl DA (2017) Brock biology of microorganisms, 15th edn. Pearson, New York
Madigan MT, Bender KS, Buckley DH, Sattley WM, Stahl DA (2019) Brock biology of microorganisms, 15, global edition. Pearson, New York
Manteca A, Alvarez R, Salazar N, Yagüe P, Sanchez J (2008) Mycelium differentiation and antibiotic production in submerged cultures of Streptomyces coelicolor. Appl Environ Microbiol 74:3877–3886. https://doi.org/10.1128/AEM.02715-07
McBride MJ, Zhu Y (2013) Gliding motility and por secretion system genes are widespread among members of the phylum Bacteroidetes. J Bacteriol 195:270–278. https://doi.org/10.1128/JB.01962-12
Meeks JC, Elhai J (2002) Regulation of cellular differentiation in filamentous cyanobacteria in free-living and plant-associated symbiotic growth states. Microbiol Mol Biol Rev 66:94–121. https://doi.org/10.1128/mmbr.66.1.94-121.2002
Meoded RA, Ueoka R, Helfrich EJN, Jensen K, Magnus N, Piechulla B, Piel J (2018) A polyketide synthase component for oxygen insertion into polyketide backbones. Angew Chem Int Ed Engl 57:11644–11648. https://doi.org/10.1002/anie.201805363
Murray RGE, Brenner DJ, Colwell RR, Vos PDE, Goodfellow M, Grimont PAD, Pfennig N, Stackebrandt E, Zavarzin GA (1990) Report of the ad hoc committee on approaches to taxonomy within the Proteobacteria. Int J Syst Evol Microbiol 40:213–215. https://doi.org/10.1099/00207713-40-2-213
Nagy E (2010) Anaerobic infections: update on treatment considerations. Drugs 70:841–858. https://doi.org/10.2165/11534490-000000000-00000
Nakai R, Nishijima M, Tazato N, Handa Y, Karray F, Sayadi S, Isoda H, Naganuma T (2014) Oligoflexus tunisiensis gen. nov., sp. nov., a gram-negative, aerobic, filamentous bacterium of a novel proteobacterial lineage, and description of Oligoflexaceae fam. nov., Oligoflexales ord. nov. and Oligoflexia classis nov. Int J Syst Evol Microbiol 64:3353–3359. https://doi.org/10.1099/ijs.0.060798-0
Nautiyal CS, Rehman A, Chauhan PS (2010) Environmental Escherichia coli occur as natural plant growth-promoting soil bacterium. Arch Microbiol 192:185–193. https://doi.org/10.1007/s00203-010-0544-1
Neville BA, Forde BM, Claesson MJ, Darby T, Coghlan A, Nally K, Ross RP, O'Toole PW (2012) Characterization of pro-inflammatory flagellin proteins produced by Lactobacillus ruminis and related motile lactobacilli. PLoS One 7:e40592. https://doi.org/10.1371/journal.pone.0040592
Nguyen MT, Götz F (2016) Lipoproteins of gram-positive bacteria: key players in the immune response and virulence. Microbiol Mol Biol Rev 80:891–903. https://doi.org/10.1128/MMBR.00028-16
Nguyen T, Ishida K, Jenke-Kodama H, Dittmann E, Gurgui C, Hochmuth T, Taudien S, Platzer M, Hertweck C, Piel J (2008) Exploiting the mosaic structure of trans-acyltransferase polyketide synthases for natural product discovery and pathway dissection. Nat Biotechnol 26:225–233. https://doi.org/10.1038/nbt1379
Nieminen TT, Säde E, Endo A, Johansson P, Björkroth J (2014) The family leuconostocaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: firmicutes and tenericutes, 4th edn. Springer, Heidelberg, pp 215–240
O’Callaghan A, van Sinderen D (2016) Bifidobacteria and their role as members of the human gut microbiota. Front Microbiol 7. https://doi.org/10.3389/fmicb.2016.00925
Parte AC (2018) LPSN – list of prokaryotic names with standing in nomenclature (bacterio.Net), 20 years on. Int J Syst Evol Microbiol 68:1825–1829. https://doi.org/10.1099/ijsem.0.002786
Paster BJ, Dewhirst FE, Olsen I, Fraser GJ (1994) Phylogeny of bacteroides, prevotella, and Porphyromonas spp. and related bacteria. J Bacteriol 176:725–732. https://doi.org/10.1128/jb.176.3.725-732.1994
Pedrós-Alió C, Manrubia S (2016) The vast unknown microbial biosphere. Proc Natl Acad Sci U S A 113:6585–6587. https://doi.org/10.1073/pnas.1606105113
Percival SL, Williams DW (2014) Chapter five – cyanobacteria. In: Percival SL (ed) Microbiology of waterborne diseases: microbiological aspects and risks, 2nd edn. Academic Press, Elsevier, London, Amsterdam, Heidelberg, pp 79–88
Piechulla B (2017) Considering microbial CO2 during microbe-plant cocultivation. Plant Physiol 173:1529. https://doi.org/10.1104/pp.16.01584
Piechulla B, Lemfack MC, Kai M (2017) Effects of discrete bioactive microbial volatiles on plants and fungi. Plant Cell Environ 40:2042–2067. https://doi.org/10.1111/pce.13011
Puertas AI, Arahal DR, Ibarburu I, Elizaquível P, Aznar R, Dueñas MT (2014) Lactobacillus sicerae sp. nov., a lactic acid bacterium isolated from Spanish natural cider. Int J Syst Evol Microbiol 64:2949–2955. https://doi.org/10.1099/ijs.0.059980-0
Reichenbach H (2006) The order cytophagales. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 7. Springer, New York, pp 549–590
Reid G (2004) When microbe meets human. Clin Infect Dis 39:827–830. https://doi.org/10.1086/423387
Rizzatti G, Lopetuso LR, Gibiino G, Binda C, Gasbarrini A (2017) Proteobacteria: a common factor in human diseases. Biomed Res Int. https://doi.org/10.1155/2017/9351507
Schenkel D, Lemfack MC, Piechulla B, Splivallo R (2015) A meta-analysis approach for assessing the diversity and specificity of belowground root and microbial volatiles. Front Plant Sci 6:707. https://doi.org/10.3389/fpls.2015.00707
Schirrmeister BE, Antonelli A, Bagheri HC (2011) The origin of multicellularity in cyanobacteria. BMC Evol Biol 11:45. https://doi.org/10.1186/1471-2148-11-45
Schleifer KH (2009) Phylum XIII. Firmicute Gibbons and Murray 1978, 5 (Firmacutes [sic] Gibbons and Murray 1978, 5). In: de Vos P, Garrity GM, Jones D, Bergey DH (eds) Bergey’s manual of systematic bacteriology: volume 3: the Firmicutes, 2nd edn. Springer, Dordrecht, London, p 19
Schleifer KH, Ludwig W (1995) Phylogeny of the genus lactobacillus and related genera. Syst Appl Microbiol 18(4):461–467
Schulz S, Dickschat JS (2007) Bacterial volatiles: the smell of small organisms. Nat Prod Rep 24:814–842. https://doi.org/10.1039/b507392h
Scott TA, Piel J (2019) The hidden enzymology of bacterial natural product biosynthesis. Nat Rev Chem 3:404. https://doi.org/10.1038/s41570-019-0107-1
Seong CN, Kang JW, Lee JH, Seo SY, Woo JJ, Park C, Bae KS, Kim MS (2018) Taxonomic hierarchy of the phylum Firmicutes and novel Firmicutes species originated from various environments in Korea. J Microbiol 56:1–10. https://doi.org/10.1007/s12275-018-7318-x
Singh SP, Montgomery BL (2011) Determining cell shape: adaptive regulation of cyanobacterial cellular differentiation and morphology. Trends Microbiol 19:278–285. https://doi.org/10.1016/j.tim.2011.03.001
Slepecky RA, Hemphill HE (2006) The genus Bacillus—nonmedical. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 4. Springer, New York, pp 530–562
Smith CJ, Rocha ER, Paster BJ (2006) The medically important Bacteroides spp. in health and disease. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 7. Springer, New York, pp 381–427
Stackebrandt E (2014) The family Clostridiaceae, other genera. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E, Thompson F (eds) The prokaryotes: Firmicutes and Tenericutes, 4th edn. Springer, Berlin, Heidelberg, pp 67–73
Stackebrandt E, Murray RGE, Trüper HG (1988) Proteobacteria classis nov., a name for the phylogenetic taxon that includes the “purple Bacteria and their relatives”. Int J Syst Evol Microbiol 38:321–325. https://doi.org/10.1099/00207713-38-3-321
Strauss J, Kaplan GG, Beck PL, Rioux K, Panaccione R, Devinney R, Lynch T, Allen-Vercoe E (2011) Invasive potential of gut mucosa-derived Fusobacterium nucleatum positively correlates with IBD status of the host. Inflamm Bowel Dis 17:1971–1978. https://doi.org/10.1002/ibd.21606
Sun P, Schuurink RC, Caissard J-C, Hugueney P, Baudino S (2016) My way: noncanonical biosynthesis pathways for plant volatiles. Trends Plant Sci 21:884–894. https://doi.org/10.1016/j.tplants.2016.07.007
Sweeney TE, Morton JM (2013) The human gut microbiome: a review of the effect of obesity and surgically induced weight loss. JAMA Surg 148:563–569. https://doi.org/10.1001/jamasurg.2013.5
Swidsinski A, Dörffel Y, Loening-Baucke V, Theissig F, Rückert JC, Ismail M, Rau WA, Gaschler D, Weizenegger M, Kühn S, Schilling J, Dörffel WV (2011) Acute appendicitis is characterised by local invasion with Fusobacterium nucleatum/necrophorum. Gut 60:34–40. https://doi.org/10.1136/gut.2009.191320
Tarkka M, Hampp R (2008) Secondary metabolites of soil streptomycetes in biotic interactions. In: Karlovsky P (ed) Secondary metabolites in soil ecology. Springer-Verlag, Berlin, Heidelberg, pp 107–126
Thomas F, Hehemann J-H, Rebuffet E, Czjzek M, Michel G (2011) Environmental and gut Bacteroidetes: the food connection. Front Microbiol 2. https://doi.org/10.3389/fmicb.2011.00093
Truant AL, Menge S, Milliorn K, Lairscey R, Kelly MT (1983) Fusobacterium nucleatum pericarditis. J Clin Microbiol 17:349–351
Umber BJ, Shin H-W, Meinardi S, Leu S-Y, Zaldivar F, Cooper DM, Blake DR (2013) Gas signatures from Escherichia coli and Escherichia coli-inoculated human whole blood. Clin Transl Med 2:13. https://doi.org/10.1186/2001-1326-2-13
van Dissel D, Claessen D, van Wezel GP (2014) Morphogenesis of Streptomyces in submerged cultures. Adv Appl Microbiol 89:1–45. https://doi.org/10.1016/B978-0-12-800259-9.00001-9
von Reuss SH, Domik D, Lemfack MC, Magnus N, Kai M, Weise T, Piechulla B (2018) Sodorifen biosynthesis in the Rhizobacterium Serratia plymuthica involves methylation and cyclization of MEP-derived Farnesyl pyrophosphate by a SAM-dependent C-Methyltransferase. J Am Chem Soc 140:11855–11862. https://doi.org/10.1021/jacs.8b08510
Walter JM, Coutinho FH, Dutilh BE, Swings J, Thompson FL, Thompson CC (2017) Ecogenomics and taxonomy of cyanobacteria phylum. Front Microbiol 8. https://doi.org/10.3389/fmicb.2017.02132
Wang C-M, Cane DE (2008) Biochemistry and molecular genetics of the biosynthesis of the earthy odorant methylisoborneol in Streptomyces coelicolor. J Am Chem Soc 130:8908–8909. https://doi.org/10.1021/ja803639g
Waterbury JB (2006) The cyanobacteria – isolation, purification and identification. In: Dworkin M, Falkow S, Rosenberg E, Schleifer K-H, Stackebrandt E (eds) The prokaryotes, vol 4. Springer, New York, pp 1053–1073
Weise T, Thürmer A, Brady S, Kai M, Daniel R, Gottschalk G, Piechulla B (2014) VOC emission of various Serratia species and isolates and genome analysis of Serratia plymuthica 4Rx13. FEMS Microbiol Lett 352:45–53. https://doi.org/10.1111/1574-6968.12359
Wexler HM (2007) Bacteroides: the good, the bad, and the Nitty-gritty. Clin Microbiol Rev 20:593–621. https://doi.org/10.1128/CMR.00008-07
Whitton BA (1992) Diversity, ecology, and taxonomy of the cyanobacteria. In: Mann NH, Carr NG (eds) Photosynthetic prokaryotes, vol 9. Springer, Boston, MA, pp 1–51
Williams KP, Kelly DP (2013) Proposal for a new class within the phylum Proteobacteria, Acidithiobacillia classis nov., with the type order Acidithiobacillales, and emended description of the class Gammaproteobacteria. Int J Syst Evol Microbiol 63:2901–2906. https://doi.org/10.1099/ijs.0.049270-0
Woese CR (1987) Bacterial evolution. Microbiol Rev 51:221–271
Woese CR, Kandler O, Wheelis ML (1990) Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci U S A 87:4576–4579. https://doi.org/10.1073/pnas.87.12.4576
Yamada Y, Cane DE, Ikeda H (2012) Diversity and analysis of bacterial terpene synthases. Methods Enzymol 515:123–162. https://doi.org/10.1016/B978-0-12-394290-6.00007-0
Yokota A (1997) Phylogenetic relationship of actinomycetes. In: Miyadoh S (ed) Atlas of actinomycetes. Asakura Publ, Tokyo
Yu K, Hamilton-Kemp TR, Archbold DD, Collins RW, Newman MC (2000) Volatile compounds from Escherichia coli O157:H7 and their absorption by strawberry fruit. J Agric Food Chem 48:413–417. https://doi.org/10.1021/jf990576b
Zaheer A, Mirza BS, Mclean JE, Yasmin S, Shah TM, Malik KA, Mirza MS (2016) Association of plant growth-promoting Serratia spp. with the root nodules of chickpea. Res Microbiol 167:510–520. https://doi.org/10.1016/j.resmic.2016.04.001
Zhao J-S, Manno D, Hawari J (2009) Psychrilyobacter atlanticus gen. nov., sp. nov., a marine member of the phylum Fusobacteria that produces H2 and degrades nitramine explosives under low temperature conditions. Int J Syst Evol Microbiol 59:491–497. https://doi.org/10.1099/ijs.0.65263-0
Zhu J, Hill JE (2013) Detection of Escherichia coli via VOC profiling using secondary electrospray ionization-mass spectrometry (SESI-MS). Food Microbiol 34:412–417. https://doi.org/10.1016/j.fm.2012.12.008
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Lemfack, MC., Bahl, H., Piechulla, B., Magnus, N. (2020). The Domain of Bacteria and Their Volatile Metabolic Potential. In: Ryu, CM., Weisskopf, L., Piechulla, B. (eds) Bacterial Volatile Compounds as Mediators of Airborne Interactions. Springer, Singapore. https://doi.org/10.1007/978-981-15-7293-7_1
Download citation
DOI: https://doi.org/10.1007/978-981-15-7293-7_1
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-7292-0
Online ISBN: 978-981-15-7293-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)