Abstract
Phenolics are one of the most abundant groups of secondary metabolites found throughout the plant kingdom. Phenolic compounds are widely distributed in plant tissues, particularly contributing to color, flavor, taste, and astringency to fruits. Phenolics are aromatic benzene ring compounds with one or more hydroxyl groups attached to it. Phenolics constitute most likely the largest group of plant secondary metabolites, varying in size from a simple structure with an aromatic ring to complex ones such as lignins. Most phenolic compounds are thought to be by-products of the metabolism of the aromatic amino acid phenylalanine. Phenolics are considered to play a vital role in the evolution of plants by providing an adaptive advantage to cope up with the changing environments. Furthermore, they are believed to be involved in leaf and petal senescence. Change in the leaf or petal color during senescence is also due to anthocyanin pigments, a kind of plant phenolics.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
18.1 Introduction
The plant phenolics play an important role in the survival of plants by providing specific adaptation to the changing environmental conditions. Phenolics are defined as the molecule that is not directly involved in the internal economy of the plant that produces it. They are essential for plant survival and defense mechanisms and act like an immune system of the plants. Phenolics are diverse compounds that differ from species to species and from tissue to tissue. They are produced by plants and compartmented in plant vacuole. They occur ubiquitously in plants in a variety of tissues. The characteristic feature of all phenolics is the presence of one or more benzene rings with one or more hydroxyl groups that may be variously elaborated with methyl, methoxyl, amino, or glycosyl groups. Phenolics size is ranging from the simple phenol (C6H6O) of 94.1 g/mol to large complex molecules such as tannins having more than 1000 g/mol (Hagerman et al. 1998). Based on their chemical composition, their function in plants is important in plant development, growth, and defense mechanisms (Coley et al. 1985). They provide an adaptive advantage to the plants itself that produce it, e.g., wade off competition, antigerminative or toxic for other plants (allelopathic), better adapted to harsh environments, and able to combat feeding animals such as insects or cattle (antifeedant) (Delgoda and Murray 2017). Several conceivable and clearly functional roles for some of these compounds have been proposed and documented (Beckman 2000). They include the following:
-
(1)
Protection from injurious UV radiation
-
(2)
Coloration of flowers that attract pollinating animals
-
(3)
Deterrence to grazing animals and feeding insects because of their astringent, toxic nature
-
(4)
Providing a tanglefoot mechanism for trapping feeding insects
-
(5)
Resistance to pathogens
-
(6)
Volatiles that affect the growth of other neighboring plants
Thus, the primary established roles are clearly ecological in nature, some having dual or even multiple functions (Beckman 2000). In addition to their role in defense, plant growth, and development, plant phenolics also have taxonomic and evolutionary importance. The metabolic pathway that produces phenolics and their link with taxonomic distribution displayed that phenolics play a crucial role in gradual diversification of plants to suit varying lifestyles during plant evolution (Delgoda and Murray 2017). Indeed, when the first plants moved from aquatic life to terrestrial life form, they faced many different stressful conditions, such as ultraviolet (UV) radiation which is involved in damaging of the biomolecules (Caretto et al. 2015). To cope up with this condition, land plants started synthesizing the phenolic compounds such as phenylpropanoids include flavonoids, monolignols, phenolic acids, stilbenes, and caumarins which act as sunscreens for the plants (Caretto et al. 2015).
The sequential action of different biosynthetic pathways in plants leads to the production of phenolics. The glycolysis and pentose phosphate pathways provide precursors (phosphoenolpyruvate and erythrose 4-phosphate, respectively) to the shikimate pathway (Caretto et al. 2015). Phenylalanine, produced by the shikimate pathway, is the precursor of phenylpropanoid metabolism which, in turn, feeds the diverse specific flavonoid pathways (Caretto et al. 2015). The biosynthesis of phenolics is an energetically expensive process even though plants produce it due to their advantageous nature (Delgoda and Murray 2017). To make this process more favorable to plants, the genes involved in the production of these molecules reside next to each other including the resistance genes that demonstrate the natural selection of it (Delgoda and Murray 2017). They are not merely artifacts of isolation procedures and do in fact exist in nature (Delgoda and Murray 2017).
18.2 Biosynthesis of Phenolics
Plants have developed the ability to produce large number of secondary metabolites that are not required in the primary growth and development but are of vital importance for their interaction with the environment (Cheynier et al. 2013). This “chemodiversity” is mainly developed in land plants, which have to encounter numerous environmental challenges, and in particular in vascular plants that also need to maintain efficient transport of the metabolites, structural rigidity, and a fine regulation of homeostasis (Caputi et al. 2012). Several classes of phenolics have been categorized on the basis of their basic skeletons like C6 (phenol), C6-C1 (phenolic acid and aldehyde), C6-C2 (phenylacetic acid), C6-C3 (hydroxycinnamic acids), C6-C1-C6 (xanthones), C6-C2-C6 (stilbenes), and C6-C3-C6 (flavonoids) (Cheynier et al. 2013). Phenolics are formed by three different biosynthetic pathways: (i) the shikimate/chorismate or succinylbenzoate pathway, which produces the phenylpropanoid derivatives (C6-C3); (ii) the acetate/malonate or polyketide pathway, which produces the side-chain-elongated phenylpropanoids, including the large group of flavonoids (C6-C3-C6) and some quinones; and (iii) the acetate/mevalonate pathway, which produces the aromatic terpenoids, mostly monoterpenes, by dehydrogenation reactions (Bhattacharya et al. 2010).
Phenylpropanoid-derived metabolites are a huge class of secondary metabolites produced from primary metabolites, phenylalanine or tyrosine, through a series of enzymatic reactions. Based on chemical structures, phenylpropanoids can be divided into five groups, including flavonoids, monolignols, phenolic acids, stilbenes, and coumarins (Noel et al. 2005; Vogt 2010; Liu et al. 2015). Among them, flavonoids, monolignols, and phenolic acids are three most common groups which can be found in almost all land plants. In the past decades, phenylpropanoids have stirred a wide concern in the fields of plant biology and medicines for their essential roles in plant survival and broad biological activities in the human body. Many studies in the recent years have shown that some chronic diseases such as cardiovascular, diabetic, and some cancer can be reduced by taking phenylpropanoid-pathway-derived metabolites (Roupe et al. 2006; Ojala et al. 2000; Kumar and Pandey 2013; Venugopala et al. 2013;Yang et al. 2016; Xia et al. 2017).
The initial steps of the phenylpropanoid pathway are collectively referred to as the general phenylpropanoid pathway (GPP) (Winkel 2004; Ferrer et al. 2008; Vogt 2010). Derivation of phenylpropanoids from phenylalanine undergoes a series of enzymatic reactions to be converted to other metabolites. In the first enzymatic step of phenylpropanoid pathway, phenylalanine is deaminated by phenylalanine ammonia lyase (PAL) to yield cinnamic acid, which is then hydroxylated and transformed to p-coumaric acid by cinnamate-4-hydroxylase (C4H) in the second enzymatic step. On the contrary, in some monocots, fungi and bacterial species, tyrosine ammonia lyase (TAL), or bifunctional ammonia lyase (PTAL) can directly convert tyrosine into p-coumaric acid, which bypasses the C4H intermediate (Watts et al. 2004; Jendresen et al. 2015; Barros et al. 2016). Following this, 4-coumaroyl-CoA ligase (4CL) catalyzes the conversion of p-coumaric acid into p-coumaroyl-CoA, which is an important branch point leading to generate various phenylpropanoid compounds (Vogt 2010; Liu et al. 2015). One of the well-understood downstream branches following the GPP is the flavonoid pathway. In Arabidopsis thaliana and Vitis vinifera, the majority of enzymatic steps have been described and well understood (Appelhagen et al. 2014; Ichino et al. 2014; Petrussa et al. 2013).
Biosynthesis of flavonoids starts with one p-coumaroyl-CoA and three malonyl-CoA molecules by committed enzymatic step and chalcone synthase (CHS) resulting in the production of naringenin chalcone (Falcone Ferreyra et al. 2012). In the second enzymatic step, chalcone isomerase (CHI) catalyzes the stereospecific cyclization of naringenin chalcone or isoliquiritigenin chalcone to flavanone (naringenin flavanone or isoliquiritigenin flavanone) (Ngaki et al. 2012). Naringenin flavanone is a common intermediate of numerous flavonoid subgroups, including flavones, flavanonols, flavonols, anthocyanins, proanthocyanidins (PA), and isoflavones. However, isoliquiritigenin flavanone can be only converted to legume-specific isoflavonoids (Ngaki et al. 2012).
Dihydroflavonols, also called flavanonols or 3-hydroxy-flavanone, possess a hydroxyl group at C3 position, but there is no C2-C3 double bond. Dihydrokaempferol possesses a hydroxyl group at 4′ position of the B-ring. Dihydroquercetin has two hydroxyl groups, which are located at 3′ and 4′ positions, respectively. Dihydromyricetin has three hydroxyl groups located at 3′, 4′, and 5′ positions, respectively. Dihydrokaempferol is synthesized by hydroxylation at C3 position of naringenin flavanone under the catalysis of a 2-ODD superfamily member, flavanone 3-hydroxylase (F3H) (Turnbull et al. 2004). Dihydrokaempferol can be subsequently hydroxylated into dihydroquercetin or dihydromyricetin by P450 superfamily members flavonoid 3′-hydroxylase (F3’H) or flavonoid 3′5’-hydroxylase (F3’5’H), respectively. F3’H catalyzes the hydroxylation at C3’ position of B-ring, whereas F3’5’H catalyzes the hydroxylation at both C3’ and C5’ positions. Except for dihydrokaempferol, naringenin flavanone can also be direct substrate of F3’H and F3’5’H, which convert it into eriodictyol and pentahydroxyflavone, respectively. The products are subsequently hydroxylated to dihydroquercetin or dihydromyricetin by F3H (Petrussa et al. 2013). Interestingly, dihydroflavonols are main branch-point intermediates of the flavonoid pathway. One branch leads to the production of flavonols, and another one flows into the biosynthesis of anthocyanins and proanthocyanidins.
Flavonols are the most abundant flavonoids in plants. The 3-hydroxyflavone backbone of flavonols is characterized by a hydroxyl group at C3 position. Flavonol synthase (FLS) is a key enzyme in the flavonol branch, and under the catalysis of FLS, different dihydroflavonols (dihydrokaempferol, dihydroquercetin, and dihydromyricetin) can be oxidized into the corresponding flavonols (kaempferol, quercetin, and myricetin) (Petrussa, et al. 2013; Saito et al. 2013).
Anthocyanins are one of the classes of flavonoids accountable for the color of grape berries. Anthocyanins are the glycosylated products of aglycone, known as anthocyanidins. Six different types of anthocyanidins are found in nature, and they include cyanidin (Cy), pelargonidin (Pg), delphinidin (Dp), malvidin (Mv), peonidin (Pn), and petunidin (Pt). The first committed step directed into the biosynthesis of Cy, Pg, and Dp is catalyzed by dihydroflavonol reductase (DFR) with its cofactor, NAPDH (Petrussa et al. 2013; Shi and Xie 2014). DFR competes with FLS on the substrate (dihydroflavonols) and catalyzes the reduction of the 4-keto group of dihydroflavonol to leucoanthocyanidins (flavan-3,4-diols), such as leucocyanidin, leucopelargonidin, and leucodelphinidin. The second step in this branch is catalyzed by anthocyanidin synthase (ANS), which is also known as leucoanthocyanidin dioxygenase, LDOX. ANS/LDOX is one of the four 2-ODD enzymes in the flavonoid pathway (Turnbull et al. 2004; Cheng et al. 2014). The other three 2-ODDs are F3H, FLS, and FNSI. Using 2-oxoglutarate and oxygen as co-substrates, ANS/LDOX oxidizes leucoanthocyanidins into the corresponding anthocyanidins (Cy, Pg, and Dp). UDP glucose/flavonoid-3-O-glucosyltransferase (UFGT) is critical for conversion of anthocyanidin to anthocyanin biosynthesis (Zheng et al. 2013).
Proanthocyanidins (PA), also known as condensed tannins (CTs), are named because they release anthocyanidins on acid hydrolysis. They are usually found in seed coat but also exist in leaves, flowers, and stems (Dixon et al. 2013). Based on the cis- or trans-stereochemistry at C2 and C3 positions of the C-ring, flavan-3-ols can be divided into 2,3-trans-flavanols and 2,3-cis-flavanols, which are represented by (+)-catechin and (−)-epicatechin, respectively. 2,3-trans-flavanols and 2,3-cis-flavanols share the same upstream biosynthetic pathway with anthocyanins but have different downstream branches catalyzed by distinct enzymes. NADPH-dependent leucoanthocyanidin reductase (LAR) competes with ANS/LDOX on the leucoanthocyanidin (flavan-3,4-diols) substrate and catalyzes the reduction reaction to produce 2,3-trans-flavanols, such as (+)-catechin, (+)-afzelechin, and (+)-gallocatechin. In contrast, 2,3-cis-flavan-3-ols are synthesized from a distinct branch with different substrates (Tanner et al. 2003; Dixon et al. 2013; Zhou et al. 2015). NADPH-dependent anthocyanidin reductase (ANR) converts anthocyanidins (pelargonidin, cyanidin, or delphinidin) to the corresponding 2,3-cis-flavan-3-ols (e.g., (−)-epiafzelechin, (−)-epicatechin, or (−)-epigallocatechin) (Xie et al. 2003; Zhou et al. 2015). Subsequently, flavan-3-ol units are condensed into PAs. Polymerization of PA is probably catalyzed by laccase-like polyphenol oxidases, but the real enzyme and the specific clustering mechanism remain in study.
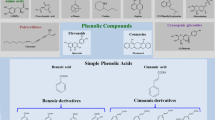
Furthermore, methylation catalyzed by O-methyltransferases and acylation and glycosylation by acyltransferases (AT) and glycosyltransferases (GTs) respectively of secondary metabolites, including phenylpropanoids and various other derived phenolic compounds, are the fundamental chemical modifications which generate enormous diversity of secondary metabolites (Tanaka et al. 2008). The complete biosynthetic pathway of phenolic acids is depicted in Fig. 18.1.
Schematic representation of biosynthetic pathways of phenolic acids (Deng and Lu 2017). PAL, phenylalanine ammonia lyase; C4H, cinnamate 4-hydroxylase; 4CL, 4-coumaroyl-CoA ligase
18.3 Regulation of Phenolics
The control of the synthesis of polyphenols involves a cascade of overlapping regulatory signals. These include developmental signals, such as during lignification of new growth or the production of anthocyanins during fruit and flower development and ripening, and environmental signals for protection against abiotic and biotic stresses. For some polyphenols, such as the flavonoids and lignin, there is now an excellent understanding on the regulation of biosynthetic genes.
Studies across a range of systems, including berry ripening, flower development, and stress-induced vegetative pigmentation, have found that the regulation of flavonoid production occurs principally through changes in the level of transcription of the biosynthetic genes. This is accomplished by the transcription factors (TFs). MYBs are the largest transcription factor (TF) families involved in activation or inhibition of gene transcription in the plant kingdom. Among them, R2R3MYBs are the largest MYB subfamily characterized in plants.
R2R3MYBs not only contribute in cell fate and identity, plant developmental processes, and responses to biotic and abiotic stresses but also play regulatory roles in primary and secondary metabolism, particularly in the biosynthesis of phenylpropanoids, such as flavonoids and monolignols (Jin and Martin 1999; Dubos et al. 2010). MYB-involved regulation of anthocyanin, proanthocyanidin, flavonol, and other flavonoid biosynthesis is largely conserved in plants (Hichri et al. 2011; Liu et al. 2015; Xu et al. 2015; Zhou et al. 2015). In general, MYBs involved in flavonoid biosynthesis can be divided into two types, independent MYB TFs and transcriptional complexes (MYB-bHLH-WDR complex (MBW) or MYB-bHLH complex). AtFLS, AtCHS, AtCHI, AtF3H, and AtF3’H are activated by independent MYB TFs, such as AtMYB12/PFG1 (production of flavonol glycoside 1), AtMYB11/PFG2, and AtMYB111/PFG3 that belong to subgroup 7 (Stracke et al. 2007, 2010). AtCHS and AtF3H are also activated by MYB-bHLH heterodimers consisting of AtMYB12 or AtMYB111 and a plant-specific bHLH TF, AtTCP3 (Li and Zachgo 2013). It has been reported that regulation of late biosynthetic genes such as production of anthocyanins and PAs required MBW complex (Dubos et al. 2010) which is composed of specific members of the R2R3MYB and basic helix-loop-helix (bHLH) TF families in conjunction with a WD-repeat (WDR; tryptophan-aspartic acid (W-D) dipeptide repeat) protein. Variant MBW complexes can form from different MYB and bHLH components, and these can have different target genes and vary in their activation or repression actions. The complex can activate the expression of late anthocyanin biosynthetic genes, such as AtDFR, AtANS/LDOX, and UF3GT (Gonzalez et al. 2008; Shi and Xie 2014; Xu et al. 2015, Kuhn et al. 2013). In grapes, there are positive transcriptional regulators involved in the general flavonoid pathway (VvMYB5a and VvMYB5b; Deluc et al. 2006, 2008) and flavonols (VvMYBF1; Czemmel et al. 2009), anthocyanins (VlMYBA1 and VlMYBA2; Kobayashi et al. 2004), and synthesis of tannins (VvMYBPA1; Bogs et al. 2007). In tomato fruit, anthocyanin synthesis is absent because of the bottleneck of CHI expression. Recently, Butelli et al. (2008) have selected two different snapdragon TF Delia (Del) which is bHLH and Rossea 1 (Ros1) R2R2MYB-type TF and overexpressed in tomato (cv. Micro-Tom) using fruit-specific E8 promoter. Whereas transgenic plants show normal vegetative growth, transgenic fruit started to synthesize anthocyanin at the end of green stage and continued to accumulate these pigment during subsequent ripening ultimately reaching an intense, uniform purple coloration both in peel and the skin (Butelli et al. 2008).
The PA-related R2R3MYBs were first described for A. thaliana, with the R2R3MYB TT2 interacting with TT8 (bHLH) and TTG1 (WDR) to form the MBW complex (Feller et al. 2011; Hichri et al. 2011). Subsequently, the genes involved in regulation of PA biosynthesis have been characterized for other species such as grape, in which several PA-related R2R3MYBs have been identified (Bogs et al. 2007; Deluc et al. 2008; Terrier et al. 2009) and various legumes (Dixon et al. 2013). A significant recent breakthrough is the identification of an R2R3MYB from rabbit’s foot clover (Trifolium arvense L.) that when overexpressed in Medicago sativa L. or Trifolium repens L. induces foliar PA accumulation at up to 1.8% dry weight (Hancock et al. 2012). In grape, PAs are major influences on the sensory qualities of the resultant wine. Persimmon is a very interesting study system, as in persimmon PAs can accumulate to such levels that they render the fruit highly astringent (Akagi et al. 2012).
Except anthocyanins, PA, and flavonols, there are only a few polyphenol biosynthetic pathways for which TFs have been characterized. Most notably, the regulation of lignin biosynthesis has been extensively studied in A. thaliana and to a lesser extent the grasses (Gray et al. 2012; Zhao and Dixon 2011). It includes AtMYB4, AtMYB26, AtMYB32, AtMYB43, AtMYB46, AtMYB52, AtMYB54, AtMYB58, AtMYB61, AtMYB63, AtMYB75/PAP1, AtMYB83, AtMYB85, and AtMYB103 (Newman et al. 2004; Zhong et al. 2008; McCarthy et al. 2009; Kim et al. 2012; Romano et al. 2012; Zhong and Ye 2014; Ko et al. 2014; Yoon et al. 2015; Nakano et al. 2015). These MYBs work together with other TFs, such as the secondary wall NAC (SWN). They form a complicated transcriptional network to regulate the transcription of lignin biosynthetic genes (Zhao and Dixon 2011; Ko et al. 2014; Yoon et al. 2015). Overexpression of AtMYB85 results in specific lignin ectopic deposition in the epidermal and cortical cells of stems (Zhong et al. 2008).
The knowledge of MYBs regulating other phenylpropanoids is limited and fragmented, and they may act as activators or repressors in the biosynthetic pathway of phenolic acids. One such example is AtMYB4 which acts as an active repressor controlling sinapate ester biosynthesis in a UV-dependent manner (Jin et al. 2000) and Petunia hybrida (Hook.) Vilm. MYB4 represses formation of phenylpropanoid scent compounds in the petals (Colquhoun et al. 2011). Moreover in Salvia miltiorrhiza, SmMYB39 represses phenolic acid biosynthesis through negatively regulating the expression of SmC4H and tyrosine aminotransferase gene (SmTAT) (Zhang et al. 2013). There has been recent progress on understanding the regulation of another branch of the phenylpropanoid pathway that is producing the isoflavonoids. Changes in TF gene expression associated with isoflavonoid production were characterized for the Lotus japonicus L. and a group of R2R3MYBs (subgroup 2) identified as candidate activators for the necessary biosynthetic genes (Shelton et al. 2012).
Several TFs with a repressive effect on flavonoid biosynthesis are now known. The first repressor TF to be identified from plants was the R2R3MYB mentioned earlier, AtMYB4. AtMYB4 production is downregulated by exposure to UV-B irradiation, which in turn releases the cinnamate-4-hydroxylase gene from AtMYB4-based suppression, thus stimulating the production of sinapate esters for UV-B protection (Jin et al. 2000). A potential repressor for anthocyanin production was identified soon after AtMYB4, FaMYB1 from strawberry. Expression of FaMYB1 in tobacco (Aharoni et al., 2001) or Lotus corniculatus L. (Paolocci et al. 2011) represses anthocyanin or PA biosynthesis, respectively. Both AtMYB4 and FaMYB1 possess residues required for R2R3MYB interaction with the bHLH partners and also have a domain in the C-terminal that mediates active transcriptional repression, called the ethylene-responsive transcription factor (ERF)-associated amphiphilic repression (EAR) domain. This suggests that they can form MBW complexes that repress, rather than activate, target genes.
While many of the recent advances have come from the model species, such as A. thaliana, Petunia, and Antirrhinum majus L., there have also been significant advances in the depth of understanding of regulation of phenylpropanoid biosynthesis in important horticultural species, in particular grape and apple (Hichri et al. 2011; Allan et al. 2008). The completion of the genome sequences of these two species has enabled more extensive analysis of the phenylpropanoid-related TF families. The R2R3MYBs regulating anthocyanin production in grape have been extensively studied over the last decade (Hichri et al. 2011), but more recently, those for the PA (Bogs et al. 2007; Deluc et al. 2008; Terrier et al. 2009) and flavonol (Czemmel et al. 2009) pathway branches have been identified, as well as the phenylpropanoid-related bHLHs and WDR (Hichri et al. 2011; Matus et al. 2010). Lin-Wang et al. (2010) characterized a large number of anthocyanin-related R2R3MYB activators from Rosaceae species, with a focus on apple, and also identified a range of apple R2R3MYBs containing the EAR domain that were able to inhibit anthocyanin gene activation in a transient assay system (Lin-Wang et al. 2011).
The use of TFs as transgenes for modification of the biosynthesis of polyphenolics is now well proven, with many different genes shown to be effective and a large number of species successfully targeted (Hichri et al. 2011, Dixon et al. 2013). However, there are still issues to be resolved in using TFs for pathway regulation. In particular, great variation is observed in transgenic phenotype depending on the specific transgene/host species combination. For example, in a given species, it may vary whether the R2R3MYB or bHLH of the flavonoid-related MBW complex is more effective, or two R2R3MYBs from the same gene family may generate dramatically different phenotypes. These variations likely reflect differences in the activation strength and specific target DNA motifs of the individual TF proteins and/or hierarchies of regulation among the TFs (Feller et al. 2011). Some TFs may activate a wide range of target genes when overexpressed, including in pathways for which they are not normally key regulatory factors. For example, AtMYB12 can induce production of both flavonols and caffeoylquinic acids when overexpressed in tomato (Luo et al. 2008). For some target pathways, the correct combination of activator and repressor TFs may be required if the desired level of phenolic production is to be achieved (Shelton et al. 2012).
The understanding of the regulation of polyphenol production is well advanced compared to the state of knowledge for other plant secondary metabolite pathways. However, there are still many areas in which progress is limited. In particular, there is little depth of knowledge outside of the phenylpropanoid pathway, with information lacking for groups of compounds as significant as the coumarins, anthraquinones, naphthoquinones, xanthones, and hydrolyzable tannins.
18.4 Evolutionary Significance of Plant Phenolics
Phenolic compounds have been produced in plants to provide adaptive advantage and metabolic plasticity to survive under different environmental challenges throughout the course of evolution. Plant phenolics played a significant role in the colonization of land plant from aquatic life. This transition faced three major challenges: large and rapid temperature changes, desiccation, and direct harsh ultraviolet light (Delgoda and Murray 2017). Additional challenges included the need for anchorage on soil and rock, the combat of new forms of microorganic occupancies in the new soil environment, and eventually competition by other plants for space and resources as well as grazing predators (Delgoda and Murray 2017). It appears that the production of phenolics has played a crucial role in adaption to new environment and overcoming these abiotic and biotic hurdles.
The biosynthesis of the phenolics, phenylpropanoids, flavonoids, and sporopollenins, present in land plants, provided protection against UV radiation, which enable them to survive in the harsh terrestrial landscape with direct exposure to sunlight (Delgoda and Murray 2017). These metabolites have played a critical role in helping the early algal forms to transition into the mosses and liverworts found in terrestrial environments (Delgoda and Murray 2017). Cuticle development in bryophyte also helps in the avoidance of some level of desiccation, although their approach was a rather rudimentary on and off method for metabolism, for wet and dry times, respectively (Delgoda and Murray 2017).
Nearly 40 million years ago, the production of lignin was first observed in tracheophytes to overcome the problems of desiccation (Delgoda and Murray 2017). In addition to protection against desiccation, lignin, a phenolic, complex polymer, provided physical strength to the plant and enables them to adopt erect and upright form against soil and also rigidity required for vasculature. It also made the cell wall rigid and tough required for vasculature and internal irrigation through xylem and phloem cell formations (Delgoda and Murray 2017).
The formation of true leaves having trichomes (in addition to mechanical defenses such as thorns, hairs, and crystals) occurs in late Paleozoic Era (Levin 1973). Trichome provided defense against insect feeder. Different types of trichomes were also storage place for exuding terpenes, phenolics, alkaloids, and others with repellent properties (Levin 1973).
By the late Devonian Era, a well-develop root system in plants was observed which helps the plant for its establishment in the soil and taking water from the soil. Phenolics like isoflavonoids used to be produced by many plant roots against soilborne microorganism during its establishment in the soil (Delgoda and Murray 2017).
During the Mesozoic Era, tannin biosynthesis in the seed coat was observed with the appearance of seed plants in gymnosperms (Delgoda and Murray 2017). Tannin biosynthesis acts as antifeedant and protects the seed. With the evolution of the angiosperms around 145 million years ago, many compounds like anthocyanins and carotenoids (terpenoids) appeared to provide color and fragrance to the flower which help them for pollination and reproductive success of the flowering plants (Delgoda and Murray 2017).
18.5 Role of Plant Phenolics in Senescence
Plant senescence is a degenerative and highly regulated process which occurs at last stage of development and required for fitness and survival of the plant. It involves the recycling and reallocation of nutrients to actively growing parts of plant. The senescence is governed by external as well as internal stimuli in a coordinated manner. The expression of many different genes and transcription factors (TF) was also observed during the courses of senescence at molecular level. The total process of senescence can be divided into three phases: initiation, reorganization, and termination (Bresson et al. 2017). The initiation phase involves the integration of different external as well as internal signals at cellular, tissue, and organ levels. During the reorganization phase, breakdown of macromolecules and remobilization of nutrients occur in the cell (Bresson et al. 2017). Many different proteases and their regulator get activated and are involved in breakdown of proteins. During this process, large macromolecules were converted into transportable and useful by-products. However, macromolecule degradation also resulted in the generation of many toxic intermediates and by-products (Bresson et al. 2017). The removal of these toxic intermediates is essential to maintain nucleus and mitochondria remain functional in order to make a transcription control and provide sufficient energy till the end of senescence. Therefore, antioxidative enzymes get activated to detoxify these toxic intermediate compounds (Bresson et al. 2017). Anthocyanin, a natural antioxidant, was also used to protect against oxidative stress-induced damage and increases during senescence (He and Giusti 2010). During the termination phase, disruption of vacuole resulted in gradual degradation of the cytoplasm due to release of nuclease and protease (Bresson et al. 2017). Finally, Cell death occurs (Bresson et al. 2017).
Terpene compounds also play an important role in plant senescence. They are synthesized by two pathways in the cytosol, endoplasmic reticulum, peroxisomes, and plastids and stored in glandular cells of leaves and resin ducts of needles (Korankye et al. 2017). Terpene synthesis and plant senescence have been tied to photosynthesis of a plant since photosynthesis is reported to serve as a carbon source in initiating terpene biosynthesis (Korankye et al. 2017). However, there have been speculations of alternative carbon sources such as xylem and chloroplast in studies where plants showed reduced rates of photosynthesis but increased in terpene biosynthesis (Korankye et al. 2017). Role of terpenes in plant senescence was reported in Arabidopsis thaliana exposed to citral, peppermint, β-pinene, α-pinene, and camphene (Korankye et al. 2017). Postharvest studies have also shown that after trees such as balsam fir are cut, excessive synthesis and/or emission of terpene compounds such as β-pinene, β-terpinene, camphene, and 3-carene is induced prior to needle abscission (Korankye et al. 2017).
18.5.1 Leaf Senescence
In plant leaves, the common physical indicators of senescence most often occur in a much later stage than the actual onset but are usually characterized when the mesophyll tissue begins to lose its greenness and turn to yellow or red (Korankye et al. 2017). The color change is due to both preferential degradation of chlorophyll compared to carotenoids and synthesis of new compounds, such as anthocyanins and phenolics (Matile 1980), and then the ultimate consequence of senescence, which may or may not result in organ abscission (Korankye et al. 2017). Plants, being a sessile in nature, have to respond rapidly to the adverse environmental conditions. In general, they respond by shedding off the affected part of plants and saving the rest of the plant. For instance, a leaf affected with disease undergoes senescence and finally drops off the plant, thereby preventing the spread of disease and allowing the rest of the plant to continue in its development. Leaf senescence occurs in response to the many factors like disease, drought, and nutrient deficiency. A study on green leaves and red leaves of Prunus indicated that the anthocyanins help the red leaves to perform better during early stage of senescence and also in photoprotection (Lo Piccolo et al. 2018). Metabolomics study of natural early tobacco leaf senescence indicates decreases in membrane lipids, free sterols, and acylated sterol glucosides along with the accumulation of sterol esters and alkaloids. The amino acid levels were significantly decreased, particularly those of N-rich amino acids (glutamine and asparagine), thus reflecting N translocation. Sugar alcohols and polyphenols accumulated when the lower leaves turned yellow (Li et al. 2016).
18.5.2 Autumn Senescence
Deciduous tree commonly shed their leaves during autumn, and this process is known as autumn senescence. This process is important for winter storage of plants in which remobilization of nutrient occurs from leaves. Autumn senescence is also a kind of leaf senescence which is characterized by chlorophyll degradation and flavonoid synthesis (Mattila et al. 2018). The spectacular autumn colors are partly caused by exposure of carotenoids due to faster degradation of chlorophyll (Mattila et al. 2018). In addition, some species also synthesize specific flavonoids, e.g., anthocyanins (Falcone Ferreyra et al. 2012) which are largely responsible for the red colors in senescing leaves (Mattila et al. 2018). Anthocyanins are suggested to act as antioxidants or metal ion chelators, in blocking ultraviolet (UV) or visible radiation or playing roles in defense and signalling (Archetti 2009; Landi et al. 2015). The amounts of other flavonoid compounds, flavonols or flavonol glycosides, have been reported to increase during age-related senescence (Torras-Claveria et al. 2012), but their roles are poorly understood. In contrast to anthocyanins, flavonols do not absorb visible light. They are found in the cuticle and sometimes inside the cells, and they block UV radiation (Solovchenko and Merzlyak 2008) and may act as ROS scavengers or signal molecules (Pollastri and Tattini 2011). Flavonol glycosides have also been suggested to be involved in supercooling of xylem parenchyma cells of Cercidiphyllum japonicum (Kasuga et al. 2008).
Keskitalo et al. (2005) described the cellular time table of autumn senescence in Populus tremula. Similarly, Mattila et al. (2018) studied the autumn senescence in Sorbus aucuparia, Acer platanoides, Betula pendula, and Prunus padus and measured the chlorophyll and flavonol contents every morning and evening during the whole autumn with a nondestructive method from individual leaves. Increase in flavonols is commonly observed during senescence which is accompanied with the rapid degradation of chlorophyll. Moy et al. (2015) recorded increase in anthocyanin content during late autumn in maple tree. Autumn senescence is considered to be coordinated and synchronized at the organism level; neither all leaves of a tree nor all cells of a single leaf senesce simultaneously (Keskitalo et al. 2005).
18.5.3 Petal Senescence
Petal senescence is a rapid and synchronous process that represents the final stage of petal development. Prior to senescence, petals play an important role in pollination of the plants by producing scent and color that attract pollinators. Anthocyanins, betalains, and carotenoids are the major flower pigment and provide color to the petals and other organs of the plants. Color change, rolling, wilting, etc. are the visible signs of petal senescence, while collapse of mesophyll cells occurs before the appearance of visible morphological changes (Ma et al. 2018). Petals are evolutionarily derived from leaves; therefore, many events are common in both senescence; these includes disintegration of intracellular structures, degradation of macromolecules and membrane systems, and recycling of substances (Ma et al. 2018). However, petal and leaf senescence are also distinct from each other in a number of ways (Ma et al. 2018).
First, leaf senescence can be reversible in some species, and re-greening is also possible, before a point of “no return,” while the process of petal senescence can only be delayed, but not arrested or reversed (Ma et al. 2018). Petal senescence is greatly activated or accelerated by pollination and is thus tightly regulated by developmental signals, rather than environmental signals, which contrasts with leaf senescence (Ma et al. 2018). Secondly, petals are decorative organs that contain relatively lower levels of nutrients and are considered to be “sink” organs, while leaves are “source” organs and the primary site of photosynthesis, so nutrient remobilization is a critical event during leaf senescence (Ma et al. 2018). Nutrient remobilization from petals is therefore less critical during senescence, although nutrients can be transferred from petals to the ovary during seed development (Ma et al. 2018). Thirdly, petal senescence is relatively rapid, and petal abscission can occur even while the petal is still turgid in some plant species (Ma et al. 2018). In contrast, leaf senescence involves gradual yellowing and wilting, and the leaf abscission occurs after nutrient recycling has been completed (Ma et al. 2018).
The genus Oenothera, evening primrose, is known to undergo a flower color change during senescence. The flowers of this genus bloom in the evening and fade in the morning (Teppabut et al. 2018). When fully opened, the petals of O. tetraptera are white, and then, they become pink in the morning (Teppabut et al. 2018). Those of O. laciniata as well as O. stricta are yellow, and then, they turn orange as they fade. These phenomena strongly indicate that an anthocyanin is biosynthesized during senescence (Teppabut et al. 2018).
18.6 Conclusions
Phenolics are ubiquitously present throughout the plant kingdom and play a significant role in plant development and defense by acting as an immune system of plant. In recent past, many researchers have focus on identification and extraction of different plant phenolics due to their antioxidant properties and remarkable effect in preventing many diseases including cancer. Plant phenolics also played a critical role in diversification of plant during evolution. In addition to this, their role in different types of plant senescence indicates their importance for plant life cycle. However, more findings are required to understand their involvement during senescence.
References
Aharoni A, De Vos CR, Wein M, Sun Z, Greco R, Kroon A, Mol JN, O'connell AP (2001) The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. Plant J 28(3):319–332
Akagi T, Katayama-Ikegami A, Kobayashi S, Sato A, Kono A, Yonemori K (2012) Seasonal abscisic acid signal and a basic leucine zipper transcription factor, DkbZIP5, regulate proanthocyanidin biosynthesis in persimmon fruit. Plant Physiol 158(2):1089–1102
Allan AC, Hellens RP, Laing WA (2008) MYB transcription factors that colour our fruit. Trends Plant Sci 13(3):99–102
Appelhagen I, Thiedig K, Nordholt N, Schmidt N, Huep G, Sagasser M, Weisshaar B (2014) Update on transparent testa mutants from Arabidopsis thaliana: characterisation of new alleles from an isogenic collection. Planta 240(5):955–970
Archetti M (2009) Classification of hypotheses on the evolution of autumn colours. Oikos 118:328–333
Barros J, Serrani-Yarce JC, Chen F, Baxter D, Venables BJ, Dixon RA (2016) Role of bifunctional ammonia-lyase in grass cell wall biosynthesis. Nature Plants 2(6):16050
Beckman CH (2000) Phenolic-storing cells: keys to programmed cell death and periderm formation in wilt disease resistance and in general defence responses in plants? Physiol Mol Plant Pathol 57:101–110
Bhattacharya A, Sood P, Citovsky V (2010) The roles of plant phenolics in defence and communication during agrobacterium and rhizobium infection. Mol Plant Pathol 11(5):705–719
Bogs J, Jaffé FW, Takos AM, Walker AR, Robinson SP (2007) The grapevine transcription factor VvMYBPA1 regulates proanthocyanidin synthesis during fruit development. Plant Physiol 143(3):1347–1361
Bresson J, Bieker S, Riester L, Doll J, Zentgraf U (2017) A guideline for leaf senescence analyses: from quantification to physiological and molecular investigations. J Exp Bot 69:769–786
Butelli E, Titta L, Giorgio M, Mock HP, Matros A, Peterek S, Schijlen EG, Hall RD, Bovy AG, Luo J, Martin C (2008) Enrichment of tomato fruit with health-promoting anthocyanins by expression of select transcription factors. Nat Biotechnol 26(11):1301
Caputi L, Malnoy M, Goremykin V, Nikiforova S, Martens S (2012) A genome-wide phylogenetic reconstruction of family 1 UDP-glycosyltransferases revealed the expansion of the family during the adaptation of plants to life on land. Plant J 69(6):1030–1042
Caretto S, Linsalata V, Colella G, Mita G, Lattanzio V (2015) Carbon fluxes between primary metabolism and phenolic pathway in plant tissues under stress. Int J Mol Sci 16:26378–26394
Cheng A-X, Han X-J, Wu Y-F, Lou H-X (2014) The function and catalysis of 2-oxoglutarate-dependent oxygenases involved in plant flavonoid biosynthesis. Int J Mol Sci 15(1):1080–1095
Cheynier V, Comte G, Davies KM, Lattanzio V, Martens S (2013) Plant phenolics: recent advances on their biosynthesis, genetics, and ecophysiology. Plant Physiol Biochem 72:1–20
Coley PD, Bryant JP, Chapin FS (1985) Resource availability and plant antiherbivore defense. Science 230:895–899
Colquhoun TA, Kim JY, Wedde AE, Levin LA, Schmitt KC, Schuurink RC, Clark DG (2011) PhMYB4 fine-tunes the floral volatile signature of Petunia× hybrida through PhC4H. J Exp Bot 62(3):1133–1143
Czemmel S, Stracke R, Weisshaar B, Cordon N, Harris NN, Walker AR, Robinson SP, Bogs J (2009) The grapevine R2R3-MYB transcription factor VvMYBF1 regulates flavonol synthesis in developing grape berries. Plant Physiol 151(3):1513–1530
Delgoda R, Murray J (2017) Evolutionary perspectives on the role of plant secondary metabolites, Pharmacognosy. Elsevier, New York, pp 93–100
Deluc L, Barrieu F, Marchive C, Lauvergeat V, Decendit A, Richard T, Carde JP, Mérillon JM, Hamdi S (2006) Characterization of a grapevine R2R3-MYB transcription factor that regulates the phenylpropanoid pathway. Plant Physiol 140(2):499–511
Deluc L, Bogs J, Walker AR, Ferrier T, Decendit A, Merillon JM, Robinson SP, Barrieu F (2008) The transcription factor VvMYB5b contributes to the regulation of anthocyanin and proanthocyanidin biosynthesis in developing grape berries. Plant Physiol 147(4):2041–2053
Deng Y, Lu S (2017) Biosynthesis and regulation of phenylpropanoids in plants. Crit Rev Plant Sci 36:257–290
Dixon RA, Liu C, Jun JH (2013) Metabolic engineering of anthocyanins and condensed tannins in plants. Curr Opin Biotechnol 24(2):329–335
Dubos C, Stracke R, Grotewold E, Weisshaar B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581
Falcone Ferreyra ML, Rius S, Casati P (2012) Flavonoids: biosynthesis, biological functions, and biotechnological applications. Front Plant Sci 3:222
Feller A, Machemer K, Braun EL, Grotewold E (2011) Evolutionary and comparative analysis of MYB and bHLH plant transcription factors. Plant J 66(1):94–116
Ferrer JL, Austin MB, Stewart C Jr, Noel JP (2008) Structure and function of enzymes involved in the biosynthesis of phenylpropanoids. Plant Physiol Biochem 46(3):356–370
Gonzalez A, Zhao M, Leavitt JM, Lloyd AM (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53(5):814–827
Gray J, Caparrós-Ruiz D, Grotewold E (2012) Grass phenylpropanoids: regulate before using! Plant Sci 184:112–120
Hagerman AE, Riedl KM, Jones GA, Sovik KN, Ritchard NT, Hartzfeld PW, Riechel TL (1998) High molecular weight plant polyphenolics (tannins) as biological antioxidants. J Agric Food Chem 46:1887–1892
Hancock KR, Collette V, Fraser K, Greig M, Xue H, Richardson K, Jones C, Rasmussen S (2012) Expression of the R2R3-MYB transcription factor TaMYB14 from Trifolium arvense activates proanthocyanidin biosynthesis in the legumes Trifolium repens and Medicago sativa. Plant Physiol 159(3):1204–1220
He J, Giusti MM (2010) Anthocyanins: natural colorants with health-promoting properties. Annu Rev Food Sci Technol 1:163–187
Hichri I, Barrieu F, Bogs J, Kappel C, Delrot S, Lauvergeat V (2011) Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J Exp Bot 62(8):2465–2483
Ichino T, Fuji K, Ueda H, Takahashi H, Koumoto Y, Takagi J, Tamura K, Sasaki R, Aoki K, Shimada T, Hara-Nishimura I (2014) GFS9/TT9 contributes to intracellular membrane trafficking and flavonoid accumulation in Arabidopsis thaliana. Plant J 80(3):410–423
Jendresen CB, Stahlhut SG, Li M, Gaspar P, Siedler S, Förster J, Maury J, Borodina I, Nielsen AT (2015) Highly active and specific tyrosine ammonia-lyases from diverse origins enable enhanced production of aromatic compounds in bacteria and Saccharomyces cerevisiae. Appl Environ Microbiol 81(13):4458–4476
Jin H, Martin C (1999) Multifunctionality and diversity within the plant MYB-gene family. Plant Mol Biol 41(5):577–585
Jin H, Cominelli E, Bailey P, Parr A, Mehrtens F, Jones J, Tonelli C, Weisshaar B, Martin C (2000) Transcriptional repression by AtMYB4 controls production of UV-protecting sunscreens in Arabidopsis. EMBO J 19(22):6150–6161
Kasuga J, Hashidoko Y, Nishioka A, Yoshiba M, Arakawa K, Fujikawa S (2008) Deep supercooling xylem parenchyma cells of katsura tree (Cercidiphyllum japonicum) contain flavonol glycosides exhibiting high anti-ice nucleation activity. Plant Cell Environ 31:1335–1348
Keskitalo J, Bergquist G, Gardeström P, Jansson S (2005) A cellular timetable of autumn senescence. Plant Physiol 139:1635–1648
Kim WC, Ko JH, Han KH (2012) Identification of a cis-acting regulatory motif recognized by MYB46, a master transcriptional regulator of secondary wall biosynthesis. Plant Mol Biol 78(4–5):489–501
Ko JH, Jeon HW, Kim WC, Kim JY, Han KH (2014) The MYB46/MYB83-mediated transcriptional regulatory programme is a gatekeeper of secondary wall biosynthesis. Ann Bot 114(6):1099–1107
Kobayashi S, Goto-Yamamoto N, Hirochika H (2004) Retrotransposon-induced mutations in grape skin color. Science 304(5673):982–982
Korankye EA, Lada R, Asiedu S, Caldwell C (2017) Plant senescence: the role of volatile terpene compounds (VTCs). Am J Plant Sci 8:3120–3139
Kuhn N, Guan L, Dai ZW, Wu BH, Lauvergeat V, Gomès E, Li SH, Godoy F, Arce-Johnson P, Delrot S (2013) Berry ripening: recently heard through the grapevine. J Exp Bot 65(16):4543–4559
Kumar S, Pandey AK (2013) Chemistry and biological activities of flavonoids: an overview. The Scientific World Journal 2013:1–16
Landi M, Tattini M, Gould KS (2015) Multiple functional roles of anthocyanins in plant-environment interactions. Environ Exp Bot 119:4–17
Levin DA (1973) The role of trichomes in plant defense. Q Rev Biol 48:3–15
Li S, Zachgo S (2013) TCP3 interacts with R2R3-MYB proteins, promotes flavonoid biosynthesis and negatively regulates the auxin response in Arabidopsis thaliana. Plant J 76(6):901–913
Li L, Zhao J, Zhao Y, Lu X, Zhou Z, Zhao C, Xu G (2016) Comprehensive investigation of tobacco leaves during natural early senescence via multi-platform metabolomics analyses. Sci Rep 6:37976
Lin-Wang K, Bolitho K, Grafton K, Kortstee A, Karunairetnam S, McGhie TK, Espley RV, Hellens RP, Allan AC (2010) An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biology 10(1):50
Lin-Wang KUI, Micheletti D, Palmer J, Volz R, Lozano L, Espley R, Hellens RP, Chagne D, Rowan DD, Troggio M, Iglesias I (2011) High temperature reduces apple fruit colour via modulation of the anthocyanin regulatory complex. Plant Cell Environ 34(7):1176–1190
Liu J, Osbourn A, Ma P (2015) MYB transcription factors as regulators of phenylpropanoid metabolism in plants. Mol Plant 8(5):689–708
Lo PE, Landi M, Pellegrini E, Agati G, Giordano C, Giordani T, Lorenzini G, Malorgio F, Massai R, Nali C (2018) Multiple consequences induced by epidermally-located anthocyanins in young, mature and senescent leaves of Prunus. Front Plant Sci 9:917
Luo J, Butelli E, Hill L, Parr A, Niggeweg R, Bailey P, Weisshaar B, Martin C (2008) AtMYB12 regulates caffeoylquinic acid and flavonol synthesis in tomato: expression in fruit results in very high levels of both types of polyphenol. Plant J 56(2):316–326
Ma N, Ma C, Liu Y, Shahid MO, Wang C, Gao J (2018) Petal senescence: a hormone view. J Exp Bot 69:719–732
Matile P (1980) Catabolism of chlorophyll: involvement of peroxidase? Z Pflanzenphysiol 99:475–478
Mattila H, Valev D, Havurinne V, Khorobrykh S, Virtanen O, Antinluoma M, Mishra KB, Tyystjärvi E (2018) Degradation of chlorophyll and synthesis of flavonols during autumn senescence—the story told by individual leaves. AoB Plants 10:ply028
Matus JT, Poupin MJ, Cañón P, Bordeu E, Alcalde JA, Arce-Johnson P (2010) Isolation of WDR and bHLH genes related to flavonoid synthesis in grapevine (Vitis vinifera L.). Plant Mol Biol 72(6):607–620
McCarthy RL, Zhong R, Ye ZH (2009) MYB83 is a direct target of SND1 and acts redundantly with MYB46 in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell Physiol 50(11):1950–1964
Moy A, Le S, Verhoeven A (2015) Different strategies for photoprotection during autumn senescence in maple and oak. Physiol Plant 155:205–216
Nakano Y, Yamaguchi M, Endo H, Rejab NA, Ohtani M (2015) NAC-MYB-based transcriptional regulation of secondary cell wall biosynthesis in land plants. Front Plant Sci 6:288
Newman LJ, Perazza DE, Juda L, Campbell MM (2004) Involvement of the R2R3-MYB, AtMYB61, in the ectopic lignification and dark-photomorphogenic components of the det3 mutant phenotype. Plant J 37(2):239–250
Ngaki MN, Louie GV, Philippe RN, Manning G, Pojer F, Bowman ME, Li L, Larsen E, Wurtele ES, Noel JP (2012) Evolution of the chalcone-isomerase fold from fatty-acid binding to stereospecific catalysis. Nature 485(7399):530
Noel JP, Austin MB, Bomati EK (2005) Structure–function relationships in plant phenylpropanoid biosynthesis. Curr Opin Plant Biol 8(3):249–253
Ojala T, Remes S, Haansuu P, Vuorela H, Hiltunen R, Haahtela K, Vuorela P (2000) Antimicrobial activity of some coumarin containing herbal plants growing in Finland. J Ethnopharmacol 73(1-2):299–305
Paolocci F, Robbins MP, Passeri V, Hauck B, Morris P, Rubini A, Arcioni S, Damiani F (2011) The strawberry transcription factor FaMYB1 inhibits the biosynthesis of proanthocyanidins in Lotus corniculatus leaves. J Exp Bot 62(3):1189–1200
Petrussa E, Braidot E, Zancani M, Peresson C, Bertolini A, Patui S, Vianello A (2013) Plant flavonoids—biosynthesis, transport and involvement in stress responses. Int J Mol Sci 14(7):14950–14973
Pollastri S, Tattini M (2011) Flavonols: old compounds for old roles. Ann Bot 108:1225–1233
Romano JM, Dubos C, Prouse MB, Wilkins O, Hong H, Poole M, Kang KY, Li E, Douglas CJ, Western TL, Mansfield SD (2012) AtMYB61, an R2R3-MYB transcription factor, functions as a pleiotropic regulator via a small gene network. New Phytol 195(4):774–786
Roupe KA, Remsberg CM, Yáñez JA, Davies NM (2006) Pharmacometrics of stilbenes: seguing towards the clinic. Curr Clin Pharmacol 1(1):81–101
Saito K, Yonekura-Sakakibara K, Nakabayashi R, Higashi Y, Yamazaki M, Tohge T, Fernie AR (2013) The flavonoid biosynthetic pathway in Arabidopsis: structural and genetic diversity. Plant Physiol Biochem 72:21–34
Shelton D, Stranne M, Mikkelsen L, Pakseresht N, Welham T, Hiraka H, Tabata S, Sato S, Paquette S, Wang TL, Martin C (2012) Transcription factors of Lotus: regulation of isoflavonoid biosynthesis requires coordinated changes in transcription factor activity. Plant Physiol 159(2):531–547
Shi M-Z, Xie D-Y (2014) Biosynthesis and metabolic engineering of anthocyanins in Arabidopsis thaliana. Recent Pat Biotechnol 8(1):47–60
Solovchenko A, Merzlyak M (2008) Screening of visible and UV radiation as a photoprotective mechanism in plants. Russ J Plant Physiol 55:719
Stracke R, Ishihara H, Huep G, Barsch A, Mehrtens F, Niehaus K, Weisshaar B (2007) Differential regulation of closely related R2R3-MYB transcription factors controls flavonol accumulation in different parts of the Arabidopsis thaliana seedling. Plant J 50(4):660–677
Stracke R, Jahns O, Keck M, Tohge T, Niehaus K, Fernie AR, Weisshaar B (2010) Analysis of PRODUCTION OF FLAVONOL GLYCOSIDES-dependent flavonol glycoside accumulation in Arabidopsis thaliana plants reveals MYB11-, MYB12-and MYB111-independent flavonol glycoside accumulation. New Phytol 188(4):985–1000
Tanaka Y, Sasaki N, Ohmiya A (2008) Biosynthesis of plant pigments: anthocyanins, betalains and carotenoids. Plant J 54(4):733–749
Tanner GJ, Francki KT, Abrahams S, Watson JM, Larkin PJ, Ashton AR (2003) Proanthocyanidin biosynthesis in plants purification of legume leucoanthocyanidin reductase and molecular cloning of its cDNA. J Biol Chem 278(34):31647–31656
Teppabut Y, Oyama K-i, Kondo T, Yoshida K (2018) Change of Petals′ Color and Chemical Components in Oenothera Flowers during Senescence. Molecules 23:1698.
Terrier N, Torregrosa L, Ageorges A, Vialet S, Verries C, Cheynier V, Romieu C (2009) Ectopic expression of VvMybPA2 promotes proanthocyanidin biosynthesis in grapevine and suggests additional targets in the pathway. Plant Physiol 149(2):1028–1041
Torras-Claveria L, Jáuregui O, Codina C, Tiburcio AF, Bastida J, Viladomat F (2012) Analysis of phenolic compounds by high-performance liquid chromatography coupled to electrospray ionization tandem mass spectrometry in senescent and water-stressed tobacco. Plant Sci 182:71–78
Turnbull JJ, Nakajima J-i, Welford RWD, Yamazaki M, Saito K, Schofield CJ (2004) Mechanistic studies on three 2-oxoglutarate-dependent oxygenases of flavonoid biosynthesis anthocyanidin synthase, flavonol synthase, and flavanone 3β-hydroxylase. J Biol Chem 279(2):1206–1216
Venugopala KN, Rashmi V, Odhav B (2013) Review on natural coumarin lead compounds for their pharmacological activity. BioMed Res Int 1:1–15
Vogt T (2010) Phenylpropanoid biosynthesis. Mol Plant 3:2–20
Watts KT, Lee PC, Schmidt-Dannert C (2004) Exploring recombinant flavonoid biosynthesis in metabolically engineered Escherichia coli. Chembiochem 5(4):500–507
Winkel BS (2004) Metabolic channeling in plants. Annu Rev Plant Biol 55:85–107
Xia N, Daiber A, Förstermann U, Li H (2017) Antioxidant effects of resveratrol in the cardiovascular system. Br J Pharmacol 174(12):1633–1646
Xie D-Y, Sharma SB, Paiva NL, Ferreira D, Dixon RA (2003) Role of anthocyanidin reductase, encoded by BANYULS in plant flavonoid biosynthesis. Science 299(5605):396–399
Xu W, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB–bHLH–WDR complexes. Trends Plant Sci 20(3):176–185
Yang L, Yang C, Li C, Zhao Q, Liu L, Fang X, Chen XY (2016) Recent advances in biosynthesis of bioactive compounds in traditional Chinese medicinal plants. Sci Bull 61(1):3–17
Yoon J, Choi H, An G (2015) Roles of lignin biosynthesis and regulatory genes in plant development. J Integr Plant Biol 57(11):902–912
Zhang S, Ma P, Yang D, Li W, Liang Z, Liu Y, Liu F (2013) Cloning and characterization of a putative R2R3 MYB transcriptional repressor of the rosmarinic acid biosynthetic pathway from Salvia miltiorrhiza. PloS One 8(9):e73259
Zhao Q, Dixon RA (2011) Transcriptional networks for lignin biosynthesis: more complex than we thought? Trends Plant Sci 16(4):227–233
Zheng Y, Li JH, Xin HP, Wang N, Guan L, Wu B-H, Li SH (2013) Anthocyanin profile and gene expression in berry skin of two red Vitis vinifera grape cultivars that are sunlight dependent versus sunlight independent. Aust J Grape Wine Res 19(2):238–248
Zhong R, Ye ZH (2014) Complexity of the transcriptional network controlling secondary wall biosynthesis. Plant Sci 229:193–207
Zhong R, Lee C, Zhou J, McCarthy RL, Ye ZH (2008) A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 20(10):2763–2782
Zhou M, Wei L, Sun Z, Gao L, Yu M, Tang Y, Wu Y (2015) Production and transcriptional regulation of proanthocyanidin biosynthesis in forage legumes. Appl Microbiol Biotechnol 99(9):3797–3806
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Tyagi, K., Shukla, P., Rohela, G.K., Shabnam, A.A., Gautam, R. (2020). Plant Phenolics: Their Biosynthesis, Regulation, Evolutionary Significance, and Role in Senescence. In: Lone, R., Shuab, R., Kamili, A. (eds) Plant Phenolics in Sustainable Agriculture . Springer, Singapore. https://doi.org/10.1007/978-981-15-4890-1_18
Download citation
DOI: https://doi.org/10.1007/978-981-15-4890-1_18
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-4889-5
Online ISBN: 978-981-15-4890-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)