Abstract
The soil is a complex mixture of organic matter and minerals, supporting a discrete array of life. Severely polluted soils have been detoxified using a variety of microorganisms. Bioremediation is a process of removal of environmental contaminants utilizing microbes through a variety of enzymatic processes. In situ processing, high public acceptance, and a comparatively lower cost hasten the overall process of bioremediation. However, it is not always effective due to its relatively long time scales and the variable range of contaminants. Varying degrees of success rate have been noticed at different sites worldwide. This chapter attempts to link the traditional and cutting edge technologies such as metagenomics, metatranscriptomics, metaproteomics, and metabolomics to numerous bioremediation techniques as they play a symbolic role in the study of the regulation of numerous mineralization pathways. Extensive data are being generated using these techniques, but their application is still in the infant stage. A stepwise organization of data is needed within the instructive databases. Microbial-assisted contaminant attenuation and in-depth analysis of the organism’s metabolism will accelerate the overall process of bioremediation. Thereafter, the next decade will going to decipher the cellular mechanisms and molecular manipulations using an integrated omic tool approach.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction
The soil is the most common home of more than 1016 diverse microbes/ton due to its heterogeneity, favoring the formation of micro-niches (Olaniran et al. 2013). Soil contamination is a change in the biological and physiochemical nature of the soil which has a detrimental effect on the living organisms residing in it. The different types of soil contamination (Fig. 5.1) are agricultural waste, industrial waste, urban waste, and radioactive waste. The fertilizers, pesticides, industrial effluents, and radionuclides flow down to the nearby water bodies or any other soil location resulting in biomagnification. This creates an interruption in the biochemical pathways and leads to harmful diseases. Improper dumping of waste in landfills and public places results in erroneous disintegration of the waste and deposition of contaminants in the soil. Limitless deposition of waste results in increased bacterial growth that causes a rise in the generation of methane gas which eventually leads to global warming. Nuclear power plants and nuclear testing add wavering amount of radioactive material to the soil (Mishra et al. 2015).
Earlier, the disposal of waste was done by throwing the waste in a hole, but due to the lack of new areas, the practice was difficult. With the emerging techniques like chemical decomposition and incineration at high temperature, the disposal of waste became effective, but they came with several disadvantages like obscure methods, expensive, and others. Alternative techniques like bioremediation were hence implemented (Karigar and Rao 2011).
Bioremediation is the process of speeding up the process of natural biodegradation in the contaminated areas by the application of microbes (Calvo et al. 2009). The various strategies that are being used to remediate the soil are either removing the pollutant present in the soil or reducing its effect by stabilizing it (containment) (Cunningham and Berti 1993 ). In general, bioremediation strategies can be classified into the following three processes:
-
1.
Biodegradation: Organic compounds are fragmented down into reduced inorganic or organic compounds.
-
2.
Biotransformation: The hazardous molecules are reformed into a reduced or nonhazardous molecule.
-
3.
Biomineralization: The organic compounds are entirely degraded into inorganic compounds like carbon dioxide or H2O4.
The type of contaminant determines the remediation process that will be implemented. The soil remediation cost depends on soil properties, site conditions, volume to be remediated, and type of contaminant. The increasing urbanization and industrialization lead to contamination of soil by organic and inorganic pollutants and, hence, have led to the deterioration of the environment and the human health (Dong et al. 2013).
5.2 Traditional Technologies for Soil Remediation
Along with biological, physical, and chemical concerns, remediation strategy depends upon the legitimate and economic considerations as well. Such strategies are preferred that result in minimum adulteration to the soil.
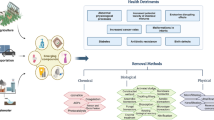
Different strategies for bioremediation of contaminated soil (Fig. 5.2) are:
5.2.1 Physical Remediation
It includes predominantly thermal desorption and soil replacement. It can be further done by replacing clean soil with contaminated soil, followed by its treatment, soil spading (the contaminated area is dug deep to spread the contaminants into deep sites and naturally degrading the pollutant), and soil importing (clean soil is added to the affected site on the surface, and mixing is done to decrease the concentration of the pollutant).
Thermal desorption is the removal of the pollutant by its volatility.
5.2.2 Chemical Remediation
-
1.
Chemical leaching: Soil washing/flushing is done with reagents, surfactants, water, and chelating agents. Soil washing is a strategy where liquids like aqueous solutions are used to separate the pollutants from the soil. The contaminants adhere to the soil particles, but they have low water solubility. To increase the solubility, additives like surfactants and chelating agents are applied along with the process (Mao et al. 2015).
-
2.
Chemical fixation: The movement of heavy metals is decreased by adding reagents. The reagents make the heavy metals insoluble in soil, thereby decreasing its toxicity (Yao et al. 2012).
-
3.
Electrokinetic remediation: In this strategy, voltage is applied at the two sides of the soil to create an electric field gradient. This technique provides minimum disturbance to the topsoil and treats the lower surface contaminants (in situ) (Gan et al. 2009).
-
4.
Vitrify remediation: The organic matter present in the soil is heated at 1400–2000 °C, and it is volatized. The end products (after pyrolysis and steam) are collected and cooled to form a rock-type substance that creates hindrance in the movement of the heavy metals. This technique can be applied in situ as well as ex situ (Yao et al. 2012).
-
5.
Ball mill process: Soil sample along with grinding media is added to the reactor (i.e., mill pot). In the absence of any chemical agents, the grinding process removes the contaminants and maintains the soil property as well (Shin et al. 2016).
-
6.
Subcritical water extraction process: Instead of organic solvent, superheated water is used as a solvent. The water is heated at a pressure less than 22.1 MPa and temperature range of 100–374 °C. It follows the principle of pressurized liquid extraction (PLE) (Islam et al. 2013).
5.2.3 Biological Remediation
-
1.
Phytoremediation is containment or removal of the contaminants by the use of green plants. Phytoremediation generally includes three processes: phytostabilization (adsorption, reduction, and precipitation of the pollutants at the roots of the plants), phytovolatilization (converting pollutant to a gaseous state), and phytoextraction (tolerant and accumulating plants are used) (Yao et al. 2012).
-
2.
Bio-augmentation is the introduction of microorganisms at the contaminated site. The microbes are generally added to such areas where the microbes that can degrade the pollutants are in a low amount (autochthonous microorganisms) or the population present at the site does not have the catabolic pathway to degrade the pollutants (Yao et al. 2012).
-
3.
Biosorption: Biomass (containing inactive, dead microbes) can bind to heavy metals and concentrate them. First, physical adsorption at the cell surface occurs, and then the metal ions gain access to the cytoplasm via the cell membrane. The type of microorganisms used defines the type and quantity of metal binding on them (Yao et al. 2012).
5.3 Traditional Tools of Omics
Specific genes (often 16s rRNA gene) were cloned in early environmental gene sequencing to produce a microbial diversity profile unlike microbial genome sequencing and traditional microbiology which depends upon cultivated cultures. The inherent soil microbial functions are nutrient recycling along with essential elements, the formation of organic matter, and decomposition aiding the natural process of soil bioremediation (Garbeva et al. 2004). The microbial world primarily constitutes of the organisms that are already cultured consisting of 1% of the overall soil microbial community. Using culture-based techniques, Bacteroidetes, Proteobacteria, Actinobacteria, and Firmicutes are the most governing phyla isolated from soil (Schloss and Handelsman 2004). The viable and nonculturable microorganisms inherently propagate in habitual environments, but then they are dormant in the laboratory or artificial surroundings. Standard culture-based approaches cannot culture these organisms, but they embrace a copious standing in the ecosystem. Hence lipids, nucleic acids, or proteins were used from the soil samples for direct assessment of their function to overcome this problem. Familiar genes such as ITS, 18S rRNA, and 16S rRNA are used as a biomarker for identification of microbial community population in the culture-independent techniques. Hence for an enhanced phylogenetic and functional categorization of the microbial community in the soil, amalgamation of specific molecular tools such as genetic fingerprinting, quantitative PCR, fluorescence in situ hybridization technique (FISH), microbial lipid analysis, stable isotope probing, microradiography, clone library method, and DNA microarray has been developed to understand the interaction of the microorganisms with various natural factors in the soil microenvironment.
Genetic fingerprinting techniques perform the direct analysis of specific molecular biomarker genes using their amplified PCR products. The relationship between diverse communities of microbes is studied using cluster-assisted analysis which compares fingerprints from various samples using software such as GelCompar. Temperature gradient gel electrophoresis and denaturing gradient gel electrophoresis (TGGE/DGGE), single-stranded conformation polymorphism (SSCP), random amplified polymeric DNA (RAPD), terminal restriction fragment length polymorphism (T-RFLP), ribosomal intergenic spacer analysis (RISA), amplified ribosomal DNA restriction analysis (ARDRA), and length heterogeneity PCR (LH-PCR) are the most prominent techniques used in genetic fingerprinting. Multiple samples are evaluated at a glance through a generated community fingerprint based on sequence polymorphism.
Amplified ribosomal DNA fragments get separated using DGGE and TGGE. Identical length DNA fragments get separated based on their variable and nucleotide composition. At the 5′ end, a GC-rich primer prevents the thorough alienation of the PCR fragments. The illustration of a solo species by several bands is the constraint associated with this method (Dowd et al. 2008). The solicitation of traditional omic tools along with their sampling source is presented in Table 5.1. Using this technique, analysis of rhizospheric bacterial populations and assessment of the microbial soil community have been done in paddy agricultural soils in recent times (Srivastava et al. 2016; Schloter et al. 2018). RAPD, due to its high speed and ease of use, is considered a simpler technique for the assessment of inherently allied bacterial species, functional and structural interpretation of the microbial communities in the soil, and genetic fingerprinting. Synthetic oligonucleotide primers having random nucleotide sequences are annealed at multiple locations on the genomes at low temperature. Assessment of laboratory-scale biodegradation of fuel oil-contaminated soil has been studied using this technique (Piñón-Castillo et al. 2017). SSCP uses the principle of separation of a single-stranded DNA by electrophoresis. T-RFLP uses fluorescently labeled 5′ primers. Advanced throughput analysis and evaluation of numerous assorted samples at a lone time is the major advantage of this technique. Using bacterial 16S rRNA or 18S rRNA gene amplicon and fungal communities to breed restriction fragment contours, ARDRA serves as the most crucial tool in the process of unique clone identification obtained from the environment. Analysis of nitrogen-fixing bacterial communities in peak rice-growing season has been done using this technique (Chakraborty and Islam 2018). RISA analyzes the phylogenetic diversity of the microbes built on the intergenic length adjustment in the transcribed spacer region within the 23S and 16S genes for prokaryotes and 23S and 18S encoding rRNA genes for eukaryotes (Fuhrman et al. 2008). Quantitative PCR determines the manifestation and plethora of operative and taxonomic gene markers in the exploration of soil microbial communities (Bustin et al. 2005). SYBR green fluorescent dyes or fluorescent probes measure the accumulated amplicons in each cycle of PCR. Quantification of Fusarium species in root rot complex in field pea soil has been studied using quantitative PCR (Zitnick-Anderson et al. 2018). The conjugation of a fluorochrome with an oligonucleotide probe is the basic principle behind FISH. Due to its immense genetic stability and high copy number, probes of 16S rRNA are conventionally exploited in this technique. On hybridizing the homologous sequence to its fluorescent probe, the fluorescent intensity is measured using a fluorescent microscope. Lipids as opposed to nucleic acids are used in microbial lipid analysis for the assessment of soil microbial communities (Banowetz et al. 2006). The biomass of a cell has a constant amount of fatty acids which is stable in nature and gives a clear differentiating picture between the different taxonomic populations of microbes. Extraction of fatty acids is done using saponification and derivatization, generating FAMEs which are further analyzed through gas chromatography. The radiolabelled substrate is used in microradiography by metabolic active cells. Combining microradiography with FISH identifies the phylogenetic active cells which are radioactive in nature and consume the substrate (Rogers et al. 2007). The individual gene fragments are sequenced following cloning in the clone library method. The PCR-obtained sequences are compared to green gene, ribosomal data projects, and gene banks (DeSantis et al. 2007). Characterization of the microorganisms in environmental samples is also done using DNA microarrays. DNA from the environmental samples generates fluorescently labeled amplified PCR products which are directly hybridized to the microarrays having known sequence molecular probes (Gentry et al. 2006). The overall evaluation process is enhanced due to the rapid replication by the DNA microarray, aiding as a significant advantage. The intensity of the signal on the microarray is directly related to the ampleness of the target organism in the sample. Besides having enhanced responses, these conventional techniques also have their specific limitations. However, with the integration of advanced omic tools, the process of microbial community analysis has been undergoing an enhanced revolution with improved understanding and application to decipher the role of microbial communities in soil bioremediation.
5.4 Advanced Omic Tools
The DNA-based molecular techniques do not provide detailed information about gene expressions under in situ conditions. Hence sequences within the metagenomic databases from uncultured microbes provide fruitful insight into the functional microbial diversity. Therefore, post-genomic methodologies such as metagenomics, metatranscriptomics, proteogenomics, and metaproteomics provide a potential connection among the genetic and functional resemblances between numerous communities of microbes, hastening the process of soil bioremediation.
The direct collection of microbial genomes from ecological samples is known as metagenomics, community genomics, or environmental genomics (Riesenfeld et al. 2004). The communication of uncultured organisms with altered environmental factors and their biochemical role can hence be studied using metagenomics. Using functional metagenomic libraries, various functional molecules such as microbial enzymes (lipase, amylase, cellulase) and antibiotics, by companies such as Terragen are derived (Rondon et al. 2000). In the aerobic conditions, the acid sulfate soil microbial community was characterized to study the structural and functional genes, using the tools of metagenomics. Both the topsoil and parent materials underwent significant changes on incubating in aerobic conditions. The archaeal community significantly decreased, whereas the sulfur-cycling genes enhanced in the parent material (Su et al. 2017). However, at the genetic level, the relationship between community composition and taxonomic diversity remains to be determined.
Metatranscriptomics or environmental transcriptomics is the study of the variations in the microbial expression of genes under specific conditions by capturing the total mRNA (Moran 2009). Recently, the overall phylogenetic pool of the functionally and taxonomically appropriate microbial communities has been analyzed using the double-RNA method or both the rRNA and mRNA, noticing a considerable diversity among the microbial communities. Recently, through direct species electron transfer, the interaction between Methanothrix and Geobacter was studied in paddy soils under methanogenic terrestrial environments. Methanothrix is the prominent microbial contributor in the global methane production, but very little is known about its physiology and ecology. The transformation of methane from acetate serves as an important contribution by Methanothrix in terrestrial ecosystems (Holmes et al. 2017). Nitrogen-transforming reductive gene transcripts were identified through metatranscriptomics in waterlogged paddy soils. Reductive nitrogen transformation was actively induced due to the presence of anoxic environments (Masuda et al. 2017). Using rice straw as a source of carbon, the severity of seawater salinity on paddy fields was studied to observe the significant changes in mRNA expression throughout the whole community (Peng et al. 2017). The diversity of microorganisms is crucially analyzed using metatranscriptomics, elucidating the community composition and deciphering their potential in soil bioremediation.
Metaproteomics or environmental proteomics is the qualitative and quantitative study of proteins on a large scale of diverse microbial species (Wilmes and Bond 2006). Under stressful conditions, proteofingerprints are generated to indicate the functional status of microbial societies (Keller and Hettich 2009). In recent times, metaproteomics has been exercised in various environments such as sediments, soil, freshwater, and marine systems. However incorrect metagenomic information, flush variety of microbial organisms, and soil heterogeneity have a negative influence on the process described (Wang et al. 2016). The bacterial metabolic functions correlated with a plant in serpentine soil contaminated with nickel, cobalt, and chromium were also interpreted using metaproteomic approach. (Mattarozzi et al. 2017). A bacterial protein database was constructed through the genera identified using 16S DNA profiling. A continuum of bacteria is revealed by the proteins involved in response to stimulus and transportation of nutrients. The bacterial biocatalysts play a vital part in the evolution of a post-petroleum bio-based economy, but the difficulty in analyzing the genetic information limits the biocatalyst’s capability (Sukul et al. 2017). The bioactivity from the proteome of an environmental sample can hence be efficiently analyzed using functional metaproteomics. Hence biomass quantification in metaproteomics serves as an important tool to interpret the correlation between various soil microbial communities in a sustainable environment.
Metabolomics characterizes the response of soil microbial communities to specific biological factors, abiotic pressures, and its immediate environment. Organic, low molecular weight biomolecules occurring naturally in a cell, tissue, or biofluid are known as metabolites. A comprehensive study of metabolomics is the production of a range of metabolites or metabolomes in communication with a natural environmental stimulation (Miller 2007). Recently, metabolomics has had numerous prominent applications in the field of environmental sciences such as the development of biomarkers, feedback to altered levels of environmental stress, assessment of risks on toxicant exposure, and disease monitoring and diagnosis (Viant 2009). Prominent deviations were observed in the metabolic pathways of S. meliloti. Simultaneously this could be related to other bacteria experiencing a series of abiotic pressure. Some of the noticeable modification changes were the presence of intermediates in myoinositol degradation and changes in the biosynthesis of exopolysaccharides and pentose phosphate pathway (PPP). In accordance to metabolic adaptation, enhanced acid tolerance phenotypes and improved competitiveness in nodulation are associated with the same in rhizobia (Draghi et al. 2017). Osmotic adjustment and osmoprotection were calculated in the classical species of cowpea (Goufo et al. 2017). The organic solutes in plants uphold an optimal turgor by enacting as osmolytes or as radical scavengers to protect the metabolic functions and hence survive the extreme drought conditions. By analyzing the salt-tolerant mechanisms and the metabolic profile of plants, enhanced sustainable crop production can be achieved worldwide, leading to the development and protection of natural plant reserves and hence aiding the bioremediation potential of the soil microbial community.
5.5 Application of Omic Tools in Bioremediation
Microorganism utilizes organic compounds as a sole carbon source and to manage their biomass and assemble suitable enzymes and cofactors for their oxidation/reduction. Hence, the organic compounds should be nontoxic or less damaging to microbial growth. The microorganisms participating in the metabolic degradation of organic compounds are heterotrophic. Molecular methods like cloning, fingerprinting, ARISA, RFLP, etc. are used to study microbial diversity (Fig. 5.3). These techniques yield information on how environmental factors change the microbial community structure. More advanced techniques like Illumina and 454 sequencing are also being used to study the microbial diversity of the polluted areas. Different approaches are used to remediate contaminated soils (Yergeau et al. 2014). The present scenario includes the implementation of various omic tools (Table 5.2) to study the microbial diversity of the contaminated soil with that of uncontaminated soil, thus providing better insight for the development of the new remediation technique or improving the already existing methods.
The uptake of heavy metals like mercury can lead to biomagnification. The heavy metals interrupt with the energy metabolism of the plants. Transcriptomics helps in the early detection at molecular levels. The changes in the genes in the presence of a low and high concentration of metals can also be studied (Villiers et al. 2012; Beauvais-Flück et al. 2017). The tools of genomics like DGGE (denaturing gradient gel electrophoresis) of 16S rRNA enhance the study of several communities of microbes in non-polluted and polluted soils and, therefore, help in the isolation of the heavy metal-resistant bacterial strains (Altimira et al. 2012; Utturkar et al. 2013). The adaptation of any organisms to the surroundings is reflected in their biological activities which can be calculated by doing transcriptomics, proteomics, and metabolomics analysis. Thus, the techniques of the bioremediation can be enhanced, and the scope of remediation technique can be improved (Hediji et al. 2010; Tripathi et al. 2012). The bacterial soil community affects the uptake of metals by plants by either stimulating the plant growth or by metabolizing the heavy metals. Pyrosequencing, a next-generation tool, of 16S rRNA provides a better picture of plant-metal-microbe interaction in the soil (Muehe et al. 2015).
Phytoremediation is one of the cost-effective remediation techniques in use for years now. The purpose of plants to attenuate the xenobiotics makes them more feasible method than the physical and chemical processes. The transgenic plants result in either degradation of the xenobiotics or increased resistance of the plant to the pollutant (Abhilash et al. 2009). The industrial effluents are estimated to be 5% saline and hypersaline. Microbial diversity is less as compared to non-extreme environments. Thus degradation of the pollutant becomes a significant problem in such regions. The halophilic microorganisms are proposed to be a favorable applicant for the remediation of the hypersaline environments. Though the metabolization of hydrocarbon reduces at high salt concentration, a lengthy exposition period has shown a significant amount of degradation and metabolized hydrocarbons (Le Borgne et al. 2008). The genomic sequence analysis discloses the genes that might be involved in the degradation. Further, proteomic analysis of the microbe in the presence of different concentrations of hydrocarbons affirms the genes involved in the degradation (Yun et al. 2014; Wei et al. 2017). Application of techniques like RT-qPCR quantifies the expression of various hydrocarbon-degrading genes and thus provides an insight into the shift in the microbial communities (Yergeau et al. 2012).
5.6 Future Prospects
There have been unprecedented changes in the field of microbial ecology with the augmentation and advancement of numerous molecular-based omic tools. The inherent functional and taxonomical diversity of the innate communities of microbes present in the soil has been investigated using diverse post-genomic approaches, revealing the superficial knowledge about the metabolic and genetic heterogeneity present in the most copious organisms in the planet, known as the prokaryotes. Implicit questions, for instance, “How the physical, chemical, and biological factors regulate the microbial communities?” and “How many bacterial species are currently present in the planet?”, and the broad knowledge about the metabolic diversity in the elementally present microorganisms still remain to be understood. As most of the classified genes have no autologous sequences in the present databases, deciphering the utilitarian roles of uncultured organisms has become an appalling task. Numerous technical challenges still remain to be overcome although there has been immense progress in the field of identification and classification of the intrinsic microbial communities in the soil by the application of proteogenomic, transcriptomic, and metagenomic approaches. New insights into the soil microbiology can be provided using interdisciplinary omic system technologies, revealing the interaction between proteins, genes, and environmental factors. Hence the upcoming years will see a prioritized area of research in the development of consequential multi-omics approaches.
References
Abdallah RZ, Wegner CE, Liesack W (2019) Community transcriptomics reveals drainage effects on paddy soil microbiome across all three domains of life. Soil Biol Biochem 132:131–142
Abhilash PC, Jamil S, Singh N (2009) Transgenic plants for enhanced biodegradation and phytoremediation of organic xenobiotics. Biotechnol Adv 27:474–488
Ahn JH, Kim MS, Kim MC, Lim JS, Lee GT, Yun JK, Kim TS, Kim TS, Ka JO (2006) Analysis of bacterial diversity and community structure in forest soils contaminated with fuel hydrocarbon. J Microbiol Biotechnol 16:704–715
Ai C, Liang G, Sun J, Wang X, He P, Zhou W (2013) Different roles of rhizosphere effect and long-term fertilization in the activity and community structure of ammonia oxidizers in a calcareous fluvo-aquic soil. Soil Biol Biochem 57:30–42
Altimira F, Yáñez C, Bravo G, González M, Rojas LA, Seeger M (2012) Characterization of copper-resistant bacteria and bacterial communities from copper-polluted agricultural soils of Central Chile. BMC Microbiol 12:193
Angel R, Panhölzl C, Gabriel R, Herbold C, Wanek W, Richter A, Eichorst SA, Woebken D (2018) Application of stable-isotope labelling techniques for the detection of active diazotrophs. Environ Microbiol 20:44–61
Awasthi AK, Li J, Pandey AK, Khan J (2019) An overview of the potential of bioremediation for contaminated soil from municipal solid waste site. In: Emerging and eco-friendly approaches for waste management. Springer, Singapore, pp 59–68
Badin AL, Mustafa T, Bertrand C, Monier A, Delolme C, Geremia RA, Bedell JP (2012) Microbial communities of urban stormwater sediments: the phylogenetic structure of bacterial communities varies with porosity. FEMS Microbiol Ecol 81:324–338
Baldan E, Basaglia M, Fontana F, Shapleigh JP, Casella S (2015) Development, assessment and evaluation of a biopile for hydrocarbons soil remediation. Int Biodeterior Biodegradation 98:66–72
Banowetz GM, Whittaker GW, Dierksen KP, Azevedo MD, Kennedy AC, Griffith SM, Steiner JJ (2006) Fatty acid methyl ester analysis to identify sources of soil in surface water. J Environ Qual 35:133–140
Bargiela R, Mapelli F, Rojo D, Chouaia B, Tornés J, Borin S, Richter M, Del Pozo MV, Cappello S, Gertler C, Genovese M (2015) Bacterial population and biodegradation potential in chronically crude oil-contaminated marine sediments are strongly linked to temperature. Sci Rep 5:11651
Bastida F, Nicolás C, Moreno JL, Hernández T, Garcia C (2010) Tracing changes in the microbial community of a hydrocarbon-polluted soil by culture-dependent proteomics. Pedosphere 20:479–485
Baudoin E, Nazaret S, Mougel C, Ranjard L, Moënne-Loccoz Y (2009) Impact of inoculation with the phytostimulatory PGPR Azospirillum lipoferum CRT1 on the genetic structure of the rhizobacterial community of field-grown maize. Soil Biol Biochem 41:409–413
Beauvais-Flück R, Slaveykova VI, Cosio C (2017) Cellular toxicity pathways of inorganic and methyl mercury in the green microalga Chlamydomonas reinhardtii. Sci Rep 7:8034
Bell TH, Yergeau E, Martineau C, Juck D, Whyte LG, Greer CW (2011) Identification of nitrogen-incorporating bacteria in petroleum-contaminated arctic soils by using [15N] DNA-based stable isotope probing and pyrosequencing. Appl Environ Microbiol 77:4163–4171
Benndorf D, Balcke GU, Harms H, Von Bergen M (2007) Functional metaproteome analysis of protein extracts from contaminated soil and groundwater. ISME J 1:224
Bevivino A, Dalmastri C (2017) Impact of agricultural land management on soil bacterial community: a case study in the Mediterranean area. In: Soil biological communities and ecosystem resilience. Springer, Cham, pp 77–95
Bhaduri AM, Fulekar MH (2015) Biochemical and RAPD analysis of Hibiscus rosa sinensis induced by heavy metals. Soil Sediment Contam 24:411–422
Bharagava RN, Purchase D, Saxena G, Mulla SI (2019) Applications of metagenomics in microbial bioremediation of pollutants: from genomics to environmental cleanup. In: Microbial diversity in the genomic era. Academic, Oxford, pp 459–477
Bustin SA, Benes V, Nolan T, Pfaffl MW (2005) Quantitative real-time RT-PCR–a perspective. J Mol Endocrinol 34:597–601
Buyer JS, Baligar VC, He Z, Arévalo-Gardini E (2017) Soil microbial communities under cacao agroforestry and cover crop systems in Peru. Appl Soil Ecol 120:273–280
Calvo C, Manzanera M, Silva-Castro GA, Uad I, González-López J (2009) Application of bioemulsifiers in soil oil bioremediation processes. Sci Total Environ 407:3634–3640
Camacho-Montealegre CM, Rodrigues EM, Tótola MR (2019) Microbial diversity and bioremediation of rhizospheric soils from Trindade Island-Brazil. J Environ Manag 236:358–364
Carini P, Marsden PJ, Leff JW, Morgan EE, Strickland MS, Fierer N (2017) Relic DNA is abundant in soil and obscures estimates of soil microbial diversity. Nat Microbiol 2:16242
Chakraborty A, Islam E (2018) Temporal dynamics of total and free-living nitrogen-fixing bacterial community abundance and structure in soil with and without history of arsenic contamination during a rice growing season. Environ Sci Pollut Res 25:4951–4962
Couger MB, Youssef NH, Struchtemeyer CG, Liggenstoffer AS, Elshahed MS (2015) Transcriptomic analysis of lignocellulosic biomass degradation by the anaerobic fungal isolate Orpinomyces sp. strain C1A. Biotechnol Biofuels 8:208
Cunningham SD, Berti WR (1993) Remediation of contaminated soils with green plants: an overview. In Vitro Cell Dev Plant 29:207–212
De Cárcer DA, Martín M, Mackova M, Macek T, Karlson U, Rivilla R (2007) The introduction of genetically modified microorganisms designed for rhizoremediation induces changes on native bacteria in the rhizosphere but not in the surrounding soil. ISME J 1:215
DeSantis TZ, Brodie EL, Moberg JP, Zubieta IX, Piceno YM, Andersen GL (2007) High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb Ecol 53:371–383
Dong ZY, Huang WH, Xing DF, Zhang HF (2013) Remediation of soil co-contaminated with petroleum and heavy metals by the integration of electrokinetics and biostimulation. J Hazard Mater 260:399–408
Dowd SE, Sun Y, Secor PR, Rhoads DD, Wolcott BM, James GA, Wolcott RD (2008) Survey of bacterial diversity in chronic wounds using pyrosequencing, DGGE, and full ribosome shotgun sequencing. BMC Microbiol 8:43
Draghi WO, Del Papa MF, Barsch A, Albicoro FJ, Lozano MJ, Pühler A, Niehaus K, Lagares A (2017) A metabolomic approach to characterize the acid-tolerance response in Sinorhizobium meliloti. Metabolomics 13:71
Dunbar J, Takala S, Barns SM, Davis JA, Kuske CR (1999) Levels of bacterial community diversity in four arid soils compared by cultivation and 16S rRNA gene cloning. Appl Environ Microbiol 65:1662–1669
Espínola F, Dionisi HM, Borglin S, Brislawn CJ, Jansson JK, Mac Cormack WP, Carroll J, Sjöling S, Lozada M (2018) Metagenomic analysis of subtidal sediments from polar and subpolar coastal environments highlights the relevance of anaerobic hydrocarbon degradation processes. Microb Ecol 75:123–139
Farinati S, DalCorso G, Panigati M, Furini A (2011) Interaction between selected bacterial strains and Arabidopsis halleri modulates shoot proteome and cadmium and zinc accumulation. J Exp Bot 62:3433–3447
Festa S, Coppotelli BM, Madueño L, Loviso CL, Macchi M, Tauil RM, Valacco MP, Morelli IS (2017) Assigning ecological roles to the populations belonging to a phenanthrene-degrading bacterial consortium using omic approaches. PLoS One 12:e0184505
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007
Foti M, Sorokin DY, Lomans B, Mussman M, Zacharova EE, Pimenov NV, Kuenen JG, Muyzer G (2007) Diversity, activity, and abundance of sulfate-reducing bacteria in saline and hypersaline soda lakes. Appl Environ Microbiol 73:2093–2100
Frerichs J, Oppermann BI, Gwosdz S, Möller I, Herrmann M, Krüger M (2013) Microbial community changes at a terrestrial volcanic CO2 vent induced by soil acidification and anaerobic microhabitats within the soil column. FEMS Microbiol Ecol 84:60–74
Fu B, Conrad R, Blaser M (2018) Potential contribution of acetogenesis to anaerobic degradation in methanogenic rice field soils. Soil Biol Biochem 119:1–10
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL, Brown JH (2008) A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci 105:7774–7778
Gan S, Lau EV, Ng HK (2009) Remediation of soils contaminated with polycyclic aromatic hydrocarbons (PAHs). J Hazard Mater 172:532–549
Garbeva PV, Van Veen JA, Van Elsas JD (2004) Microbial diversity in soil: selection of microbial populations by plant and soil type and implications for disease suppressiveness. Annu Rev Phytopathol 42:243–270
Gasser I, Müller H, Berg G (2009) Ecology and characterization of polyhydroxyalkanoate-producing microorganisms on and in plants. FEMS Microbiol Ecol 70:142–150
Gentry TJ, Wickham GS, Schadt CW, He Z, Zhou J (2006) Microarray applications in microbial ecology research. Microb Ecol 52:159–175
Gieg LM, Toth CR (2018) Anaerobic biodegradation of hydrocarbons: metagenomics and metabolomics. In: Consequences of microbial interactions with hydrocarbons, oils, and lipids: biodegradation and bioremediation. Springer, Cham, pp 1–42
Gielnik A, Pechaud Y, Huguenot D, Cébron A, Riom JM, Guibaud G, Esposito G, van Hullebusch ED (2019) Effect of digestate application on microbial respiration and bacterial communities’ diversity during bioremediation of weathered petroleum hydrocarbons contaminated soils. Sci Total Environ 670:271–281
Goufo P, Moutinho-Pereira JM, Jorge TF, Correia CM, Oliveira MR, Rosa EA, António C, Trindade H (2017) Cowpea (Vigna unguiculata L. Walp.) metabolomics: osmoprotection as a physiological strategy for drought stress resistance and improved yield. Front Plant Sci 8:586
Grube M, Cernava T, Soh J, Fuchs S, Aschenbrenner I, Lassek C, Wegner U, Becher D, Riedel K, Sensen CW, Berg G (2015) Exploring functional contexts of symbiotic sustain within lichen-associated bacteria by comparative omics. ISME J 9:412
Guerrero-Molina MF, Winik BC, Pedraza RO (2012) More than rhizosphere colonization of strawberry plants by Azospirillum brasilense. Appl Soil Ecol 61:205–212
Guida M, Cannavacciuolo PL, Cesarano M, Borra M, Biffali E, D’Alessandro R, De Felice B (2014) Microbial diversity of landslide soils assessed by RFLP and SSCP fingerprints. J Appl Genet 55:403–415
Habtom H, Demanèche S, Dawson L, Azulay C, Matan O, Robe P, Gafny R, Simonet P, Jurkevitch E, Pasternak Z (2017) Soil characterisation by bacterial community analysis for forensic applications: a quantitative comparison of environmental technologies. Forensic Sci Int Genet 26:21–29
Hamamura N, Olson SH, Ward DM, Inskeep WP (2006) Microbial population dynamics associated with crude-oil biodegradation in diverse soils. Appl Environ Microbiol 72:6316–6324
Hassan SE, Boon E, Ma S-A, Hijri M (2011) Molecular biodiversity of arbuscular mycorrhizal fungi in trace metal-polluted soils. Mol Ecol 20:3469–3483
Hediji H, Djebali W, Cabasson C, Maucourt M, Baldet P, Bertrand A, Zoghlami LB, Deborde C, Moing A, Brouquisse R, Chaïbi W (2010) Effects of long-term cadmium exposure on growth and metabolomic profile of tomato plants. Ecotoxicol Environ Saf 73:1965–1974
Hilton S, Bennett AJ, Keane G, Bending GD, Chandler D, Stobart R, Mills P (2013) Impact of shortened crop rotation of oilseed rape on soil and rhizosphere microbial diversity in relation to yield decline. PLoS One 8:e59859
Holmes DE, Shrestha PM, Walker DJ, Dang Y, Nevin KP, Woodard TL, Lovley DR (2017) Metatranscriptomic evidence for direct interspecies electron transfer between Geobacter and Methanothrix species in methanogenic rice paddy soils. Appl Environ Microbiol 83:e00223–e00217
Hoppe T, Schnittler M (2015) Characterization of myxomycetes in two different soils by TRFLP-analysis of partial 18S rRNA gene sequences. Mycosphere 6:216–227
Hou L, Wu Q, Gu Q, Zhou Q, Zhang J (2018) Community structure analysis and biodegradation potential of aniline-degrading bacteria in biofilters. Curr Microbiol 75:918–924
Huang WE, Stoecker K, Griffiths R, Newbold L, Daims H, Whiteley AS, Wagner M (2007) Raman-FISH: combining stable-isotope Raman spectroscopy and fluorescence in situ hybridization for the single cell analysis of identity and function. Environ Microbiol 9:1878–1889
Inui H, Hirota M, Goto J, Yoshihara R, Kodama N, Matsui T, Yamazaki K, Eun H (2015) Zinc finger protein genes from Cucurbita pepo are promising tools for conferring non-Cucurbitaceae plants with ability to accumulate persistent organic pollutants. Chemosphere 123:48–54
Islam MN, Jo YT, Jung SK, Park JH (2013) Evaluation of subcritical water extraction process for remediation of pesticide-contaminated soil. Water Air Soil Pollut 224:1652
Ivanov P, Eickhorst T, Wehrer M, Georgiadis A, Rennert T, Eusterhues K, Totsche KU (2017) Natural attenuation of aged tar-oil in soils: a case study from a former gas production site. InEGU General Assembly Conference Abstracts 19:8586
Jayaraman S, Thangaiyan S, Mani K, Nagarajan K, Muthukalingan K (2019) Identification of a novel gene through the metagenomic approach to degrade the targeted pollutant. Microb Biodegrad Xeno Comp 30:204
Jiang Y, Qian H, Wang X, Chen L, Liu M, Li H, Sun B (2018) Nematodes and microbial community affect the sizes and turnover rates of organic carbon pools in soil aggregates. Soil Biol Biochem 119:22–31
Karigar CS, Rao SS (2011) Role of microbial enzymes in the bioremediation of pollutants: a review. Enzyme Res. https://doi.org/10.4061/2011/805187
Keenan SW, Schaeffer SM, Jin VL, DeBruyn JM (2018) Mortality hotspots: nitrogen cycling in forest soils during vertebrate decomposition. Soil Biol Biochem 121:165–176
Keller M, Hettich R (2009) Environmental proteomics: a paradigm shift in characterizing microbial activities at the molecular level. Microbiol Mol Biol Rev 73:62–70
Khudur LS, Shahsavari E, Miranda AF, Morrison PD, Nugegoda D, Ball AS (2015) Evaluating the efficacy of bioremediating a diesel-contaminated soil using ecotoxicological and bacterial community indices. Environ Sci Pollut Res 22:14809–14819
Kim S, Krajmalnik-Brown R, Kim JO, Chung J (2014) Remediation of petroleum hydrocarbon-contaminated sites by DNA diagnosis-based bioslurping technology. Sci Total Environ 497:250–259
Kolb S, Knief C, Stubner S, Conrad R (2003) Quantitative detection of methanotrophs in soil by novel pmoA-targeted real-time PCR assays. Appl Environ Microbiol 69:2423–2429
Koshlaf E, Shahsavari E, Haleyur N, Osborn AM, Ball AS (2019) Effect of biostimulation on the distribution and composition of the microbial community of a polycyclic aromatic hydrocarbon-contaminated landfill soil during bioremediation. Geoderma 338:216–225
Kotoky R, Rajkumari J, Pandey P (2018) The rhizosphere microbiome: significance in rhizoremediation of polyaromatic hydrocarbon contaminated soil. J Environ Manag 217:858–870
Krolicka A, Boccadoro C, Nilsen MM, Baussant T (2017) Capturing early changes in the marine bacterial community as a result of crude oil pollution in a Mesocosm experiment. Microbes Environ 2:358–366
Labbé D, Margesin R, Schinner F, Whyte LG, Greer CW (2007) Comparative phylogenetic analysis of microbial communities in pristine and hydrocarbon-contaminated alpine soils. FEMS Microbiol Ecol 59:466–475
Le Borgne S, Paniagua D, Vazquez-Duhalt R (2008) Biodegradation of organic pollutants by halophilic bacteria and archaea. J Mol Microbiol Biotechnol 5:74–92
Lee DW, Lee H, Lee AH, Kwon BO, Khim JS, Yim UH, Kim BS, Kim JJ (2018) Microbial community composition and PAHs removal potential of indigenous bacteria in oil contaminated sediment of Taean coast, Korea. Environ Pollut 234:503–512
Lee MS, Do JO, Park MS, Jung S, Lee KH, Bae KS, Park SJ, Kim SB (2006) Dominance of Lysobacter sp. in the rhizosphere of two coastal sand dune plant species, Calystegia soldanella and Elymus mollis. Anton Leeuw Int J G 90:19–27
Li Y, Ying Y, Zhao D, Jin S, Ding W (2010) Genetic diversity analysis on rhizosphere soil microbial population of Panax ginseng and Panax quinquefolium by RAPD. Zhongcaoyao = Chin Tradit Herb Drug 41:1871–1875
Liang X, Zhuang J, Löffler FE, Zhang Y, DeBruyn JM, Wilhelm SW, Schaeffer SM, Radosevich M (2019) Viral and bacterial community responses to stimulated Fe (III)—bioreduction during simulated subsurface bioremediation. Environ Microbiol 21:2043–2055
Lu Y, Abraham WR, Conrad R (2007) Spatial variation of active microbiota in the rice rhizosphere revealed by in situ stable isotope probing of phospholipid fatty acids. Environ Microbiol 9:474–481
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, Del Rio TG, Edgar RC (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86
Luque-Almagro VM, Moreno-Vivián C, Roldán MD (2016) Biodegradation of cyanide wastes from mining and jewellery industries. Curr Opin Biotechnol 38:9–13
Lynn TM, Ge T, Yuan H, Wei X, Wu X, Xiao K, Kumaresan D, Wu J, Whiteley AS (2017) Soil carbon-fixation rates and associated bacterial diversity and abundance in three natural ecosystems. Microb Ecol 73:645–657
Mao X, Jiang R, Xiao W, Yu J (2015) Use of surfactants for the remediation of contaminated soils: a review. J Hazard Mater 285:419–435
Maqbool F, Wang Z, Xu Y, Zhao J, Gao D, Zhao YG, Bhatti ZA, Xing B (2012) Rhizodegradation of petroleum hydrocarbons by Sesbania cannabina in bioaugmented soil with free and immobilized consortium. J Hazard Mater 237:262–269
Masuda Y, Itoh H, Shiratori Y, Isobe K, Otsuka S, Senoo K (2017) Predominant but previously-overlooked prokaryotic drivers of reductive nitrogen transformation in paddy soils, revealed by metatranscriptomics. Microbes Environ 32:180–183
Mathew RP, Feng Y, Githinji L, Ankumah R, Balkcom KS (2012) Impact of no-tillage and conventional tillage systems on soil microbial communities. Appl Environ Soil Sci. https://doi.org/10.1155/2012/548620
Mattarozzi M, Manfredi M, Montanini B, Gosetti F, Sanangelantoni AM, Marengo E, Careri M, Visioli G (2017) A metaproteomic approach dissecting major bacterial functions in the rhizosphere of plants living in serpentine soil. Anal Bioanal Chem 409:2327–2339
Meneghine AK, Nielsen S, Varani AM, Thomas T, Alves LM (2017) Metagenomic analysis of soil and freshwater from zoo agricultural area with organic fertilization. PLoS One 12:e0190178
Miller MG (2007) Environmental metabolomics: a SWOT analysis (strengths, weaknesses, opportunities, and threats). J Proteome Res 6:540–545
Mishra RK, Mohammad N, Roychoudhury N (2015) Soil pollution: causes, effects and control. Tropical Forest Research Institute 3:20–30
Moran MA (2009) Metatranscriptomics: eavesdropping on complex microbial communities-large-scale sequencing of mRNAs retrieved from natural communities provides insights into microbial activities and how they are regulated. Microbe 4:329
Moter A, Göbel UB (2000) Fluorescence in situ hybridization (FISH) for direct visualization of microorganisms. J Microbiol Methods 41:85–112
Muehe EM, Weigold P, Adaktylou IJ, Planer-Friedrich B, Kraemer U, Kappler A, Behrens S (2015) Rhizosphere microbial community composition affects cadmium and zinc uptake by the metal-hyperaccumulating plant Arabidopsis halleri. Appl Environ Microbiol 81:2173–2181
Mukherjee A, Yadav R, Marmeisse R, Fraissinet-Tachet L, Reddy MS (2019) Heavy metal hypertolerant eukaryotic aldehyde dehydrogenase isolated from metal contaminated soil by metatranscriptomics approach. Biochimie 160:183–192
Navarrete AA, Kuramae EE, de Hollander M, Pijl AS, van Veen JA, Tsai SM (2013) Acidobacterial community responses to agricultural management of soybean in Amazon forest soils. FEMS Microbiol Ecol 83:607–621
Navarro E, Fabrègue O, Scorretti R, Reboulet J, Simonet P, Dawson L, Demanèche S (2015) RisaAligner software for aligning fluorescence data between Agilent 2100 Bioanalyzer chips: application to soil microbial community analysis. BioTechniques 59:347–349
Oates LG, Read HW, Gutknecht JL, Duncan DS, Balser TB, Jackson RD (2017) A lipid extraction and analysis method for characterizing soil microbes in experiments with many samples. J Vis Exp 16:e55310
Okabe S, Kindaichi T, Ito T (2004) MAR-FISH—An ecophysiological approach to link phylogenetic affiliation and in situ metabolic activity of microorganisms at a single-cell resolution. Microbes Environ 19:83–98
Olaniran AO, Balgobind A, Pillay B (2013) Bioavailability of heavy metals in soil: impact on microbial biodegradation of organic compounds and possible improvement strategies. Int J Mol Sci 14:10197–101228
Oshoma CE, Igbeta B, Omonigho SE (2017) Analysis of microbiological and physiochemical properties of top soil from municipal dumpsites in Benin City. J Appl Sci Environ Manage 21:985–990
Pal S, Roy A, Kazy SK (2019) Exploring microbial diversity and function in petroleum hydrocarbon associated environments through Omics approaches. In: Microbial diversity in the genomic era. Academic Press, Cambridge, pp 171–194
Panigrahi S, Velraj P, Rao TS (2019) Functional microbial diversity in contaminated environment and application in bioremediation. In: Microbial diversity in the genomic era. Academic Press, pp 359–385
Panosyan H, Hakobyan A, Birkeland NK, Trchounian A (2018) Bacilli community of saline-alkaline soils from the Ararat plain (Armenia) assessed by molecular and culture-based methods. Syst Appl Microbiol 41:232–240
Peng J, Wegner CE, Liesack W (2017) Short-term exposure of paddy soil microbial communities to salt stress triggers different transcriptional responses of key taxonomic groups. Front Microbiol 8:400
Petrucco-Toffolo E, Basso A, Kerdelhué C, Ipekdal K, Mendel Z, Simonato M, Battisti A (2018) Evidence of potential hybridization in the Thaumetopoea pityocampa-wilkinsoni complex. Agric For Entomol 20:9–17
Phillips LA, Greer CW, Germida JJ (2006) Culture-based and culture-independent assessment of the impact of mixed and single plant treatments on rhizosphere microbial communities in hydrocarbon contaminated flare-pit soil. Soil Biol Biochem 38:2823–2833
Piñón-Castillo HA, Gutiérrez DL, Hernández-Castillo D, Muñoz-Castellanos LN, Rivera-Chavira BE, Nevárez-Moorillón GV (2017) Laboratory-scale biodegradation of fuel oil no. 6 in contaminated soils by Autochthonous bacteria. In: Advances in bioremediation and phytoremediation. IntechOpen
Rapp JZ, Bienhold C, Offre P, Boetius A (2016) Polysaccharide degradation potential of bacterial communities in Arctic deep-sea sediments (1200–5500 m water depth). In: 16th international symposium on microbial ecology, Montréal, Canada, 21 August 2016–26 August
Rastogi G, Stetler LD, Peyton BM, Sani RK (2009) Molecular analysis of prokaryotic diversity in the deep subsurface of the former Homestake gold mine, South Dakota, USA. J Microbiol 47:371–384
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities. Annu Rev Genet 38:525–552
Ritchie NJ, Schutter ME, Dick RP, Myrold DD (2000) Use of length heterogeneity PCR and fatty acid methyl ester profiles to characterize microbial communities in soil. Appl Environ Microbiol 66:1668–1675
Rogers SW, Moorman TB, Ong SK (2007) Fluorescent in situ hybridization and micro-autoradiography applied to ecophysiology in soil. Soil Sci Soc Am J 71:620–631
Rondon MR, August PR, Bettermann AD, Brady SF, Grossman TH, Liles MR, Loiacono KA, Lynch BA, MacNeil IA, Minor C, Tiong CL (2000) Cloning the soil metagenome: a strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl Environ Microbiol 66:2541–2547
Schloss PD, Handelsman J (2004) Status of the microbial census. Microbiol Mol Biol Rev 68:686–691
Schloter M, Nannipieri P, Sørensen SJ, van Elsas JD (2018) Microbial indicators for soil quality. Biol Fertil Soils 54:1–10
Shahsavari E, Schwarz A, Aburto-Medina A, Ball AS (2019) Biological degradation of polycyclic aromatic compounds (PAHs) in soil: a current perspective. Curr Pollut Rep 5:1–9
Shanmugam BK, Easwaran SN, Lakra R, Deepa PR, Mahadevan S (2017) Metabolic pathway and role of individual species in the bacterial consortium for biodegradation of azo dye: a biocalorimetric investigation. Chemosphere 188:81–89
Shekhar SK, Godheja J, Modi DR (2020) Molecular Technologies for Assessment of bioremediation and characterization of microbial communities at pollutant-contaminated sites. In: Bioremediation of industrial waste for environmental safety. Springer, Singapore, pp 437–474
Shibulal B, Al-Bahry SN, Al-Wahaibi YM, Elshafie AE, Al-Bemani AS, Joshi SJ (2018) Analysis of bacterial diversity in different heavy oil Wells of a reservoir in South Oman with alkaline pH. Scientifica 2018:10
Shin YJ, Park SM, Yoo JC, Jeon CS, Lee SW, Baek K (2016) A new approach for remediation of as-contaminated soil: ball mill-based technique. Environ Sci Pollut Res 23:3963–3970
Singh SK, Rai MK, Sahoo L (2012) An improved and efficient micropropagation of Eclipta alba through transverse thin cell layer culture and assessment of clonal fidelity using RAPD analysis. Ind Crop Prod 37:328–333
Slade EM, Roslin T, Santalahti M, Bell T (2016) Disentangling the ‘brown world’ faecal–detritus interaction web: dung beetle effects on soil microbial properties. Oikos 125:629–635
Srivastava M, Kaushik MS, Mishra AK (2016) Linking the physicochemical properties with the abundance and diversity of Rhizospheric bacterial population inhabiting Paddy soil based on a concerted multivariate analysis of PCR-DGGE and RISA. Geomicrobiol J 33:894–905
Srivastava M, Mishra AK (2018) Comparative responses of diazotrophic abundance and community structure to the chemical composition of paddy soil. Environ Sci Pollut Res 25:399–412
Stackebrandt E, Liesack W, Goebel BM (1993) Bacterial diversity in a soil sample from a subtropical Australian environment as determined by 16S rDNA analysis. FASEB J 7:232–236
Stefanis C, Alexopoulos A, Voidarou C, Vavias S, Bezirtzoglou E (2013) Principal methods for isolation and identification of soil microbial communities. Folia Microbiol 58:61–68
Su JQ, Xia Y, Yao HY, Li YY, An XL, Singh BK, Zhang T, Zhu YG (2017) Metagenomic assembly unravel microbial response to redox fluctuation in acid sulfate soil. Soil Biol Biochem 105:244–252
Sukul P, Schäkermann S, Bandow JE, Kusnezowa A, Nowrousian M, Leichert LI (2017) Simple discovery of bacterial biocatalysts from environmental samples through functional metaproteomics. Microbiome 5:28
Sun X, Zhang J, Zhang H, Ni Y, Zhang Q, Chen J, Guan Y (2010) The responses of Arabidopsis thaliana to cadmium exposure explored via metabolite profiling. Chemosphere 78:840–845
Sutton NB, Maphosa F, Morillo JA, Al-Soud WA, Langenhoff AA, Grotenhuis T, Rijnaarts HH, Smidt H (2012) Impact of long term diesel contamination on soil microbial community structure. Appl Environ Microbiol 79:619–630
Szewczyk R, Soboń A, Słaba M, Długoński J (2015) Mechanism study of alachlor biodegradation by Paecilomyces marquandii with proteomic and metabolomic methods. J Hazard Mater 291:52–64
Toljander JF, Lindahl BD, Paul LR, Elfstrand M, Finlay RD (2007) Influence of arbuscular mycorrhizal mycelial exudates on soil bacterial growth and community structure. FEMS Microbiol Ecol 61:295–304
Toplin JA, Norris TB, Lehr CR, McDermott TR, Castenholz RW (2008) Biogeographic and phylogenetic diversity of thermoacidophilic cyanidiales in Yellowstone National Park, Japan, and New Zealand. Appl Environ Microbiol 74:2822–2833
Tribelli PM, Rossi L, Ricardi MM, Gomez-Lozano M, Molin S, Iustman LJ, Lopez NI (2018) Microaerophilic alkane degradation in pseudomonas extremaustralis: a transcriptomic and physiological approach. J Ind Microbiol Biotechnol 45:15–23
Tripathi RD, Tripathi P, Dwivedi S, Dubey S, Chakrabarty D (2012) Arsenomics: omics of arsenic metabolism in plants. Front Physiol 3:275
Utturkar SM, Bollmann A, Brzoska RM, Klingeman DM, Epstein SE, Palumbo AV, Brown SD (2013) Draft genome sequence for Caulobacter sp. strain OR37, a bacterium tolerant to heavy metals. Genome Announc 1:e00322–e00313
Utturkar SM, Cude WN, Robeson MS, Yang ZK, Klingeman DM, Land ML, Allman SL, Lu TY, Brown SD, Schadt CW, Podar M (2016) Enrichment of root endophytic bacteria from populus deltoides and single-cell-genomics analysis. Appl Environ Microbiol 82:5698–5708
Viant MR (2009) Applications of metabolomics to the environmental sciences. Metabolomics 5:1–2
Villiers F, Hugouvieux V, Leonhardt N, Vavasseur A, Junot C, Vandenbrouck Y, Bourguignon J (2012) Exploring the plant response to cadmium exposure by transcriptomic, proteomic and metabolomic approaches: potentiality of high-throughput methods, promises of integrative biology. In: Metal toxicity in plants: perception, signaling and remediation. Springer, Berlin, pp 119–142
Visioli G, Vincenzi S, Marmiroli M, Marmiroli N (2012) Correlation between phenotype and proteome in the Ni hyperaccumulator Noccaea caerulescens subsp. caerulescens. Environ Exp Bot 77:156–164
Vivas A, Moreno B, del Val C, Macci C, Masciandaro G, Benitez E (2008) Metabolic and bacterial diversity in soils historically contaminated by heavy metals and hydrocarbons. J Environ Monit 10:1287–1296
Wanapaisan P, Laothamteep N, Vejarano F, Chakraborty J, Shintani M, Muangchinda C, Morita T, Suzuki-Minakuchi C, Inoue K, Nojiri H, Pinyakong O (2018) Synergistic degradation of pyrene by five culturable bacteria in a mangrove sediment-derived bacterial consortium. J Hazard Mater 342:561–570
Wang DZ, Kong LF, Li YY, Xie ZX (2016) Environmental microbial community proteomics: status, challenges and perspectives. Int J Mol Sci 17:1275
Wang X, Sharp CE, Jones GM, Grasby SE, Brady AL, Dunfield PF (2015) Stable-isotope probing identifies uncultured Planctomycetes as primary degraders of a complex heteropolysaccharide in soil. Appl Environ Microbiol 81:4607–4615
Wang Y, Tian H, Huang F, Long W, Zhang Q, Wang J, Zhu Y, Wu X, Chen G, Zhao L, Bakken LR (2017) Time-resolved analysis of a denitrifying bacterial community revealed a core microbiome responsible for the anaerobic degradation of quinoline. Sci Rep 7:14778
Wei K, Yin H, Peng H, Liu Z, Lu G, Dang Z (2017) Characteristics and proteomic analysis of pyrene degradation by Brevibacillus brevis in liquid medium. Chemosphere 178:80–87
Wilmes P, Bond PL (2006) Metaproteomics: studying functional gene expression in microbial ecosystems. Trends Microbiol 14:92–97
Wu T, Chellemi DO, Graham JH, Rosskopf EN (2008) Assessment of fungal communities in soil and tomato roots subjected to diverse land and crop management systems. Soil Biol Biochem 40:1967–1970
Yang MM, Xu LP, Xue QY, Yang JH, Xu Q, Liu HX, Guo JH (2012a) Screening potential bacterial biocontrol agents towards Phytophthora capsici in pepper. Eur J Plant Pathol 134:811–820
Yang S, Wen X, Jin H, Wu Q (2012b) Pyrosequencing investigation into the bacterial community in permafrost soils along the China-Russia crude oil pipeline (CRCOP). PLoS One 7:e52730
Yao Z, Li J, Xie H, Yu C (2012) Review on remediation technologies of soil contaminated by heavy metals. Procedia Environ Sci 16:722–729
Yergeau E, Sanschagrin S, Beaumier D, Greer CW (2012) Metagenomic analysis of the bioremediation of diesel-contaminated Canadian high arctic soils. PLoS One 7:e30058
Yergeau E, Sanschagrin S, Maynard C, St-Arnaud M, Greer CW (2014) Microbial expression profiles in the rhizosphere of willows depend on soil contamination. ISME J 8:344
Yun SH, Choi CW, Lee SY, Lee YG, Kwon J, Leem SH, Chung YH, Kahng HY, Kim SJ, Kwon KK, Kim SI (2014) Proteomic characterization of plasmid pLA1 for biodegradation of polycyclic aromatic hydrocarbons in the marine bacterium, Novosphingobium pentaromativorans US6-1. PLoS One 9:e90812
Zachow C, Berg C, Müller H, Meincke R, Komon-Zelazowska M, Druzhinina IS, Kubicek CP, Berg G (2009) Fungal diversity in the rhizosphere of endemic plant species of Tenerife (Canary Islands): relationship to vegetation zones and environmental factors. ISME J 3:79
Zampolli J, Zeaiter Z, Di Canito A, Di Gennaro P (2019) Genome analysis and-omics approaches provide new insights into the biodegradation potential of Rhodococcus. Appl Environ Microbiol 103:1069–1080
Zeng XW, Qiu RL, Ying RR, Tang YT, Tang L, Fang XH (2011) The differentially-expressed proteome in Zn/cd hyperaccumulator Arabis paniculata Franch. In response to Zn and cd. Chemosphere 82:321–328
Zhao L, Zhang H, White JC, Chen X, Li H, Qu X, Ji R (2019) Metabolomics reveals that engineered nanomaterial exposure in soil alters both soil rhizosphere metabolite profiles and maize metabolic pathways. Environ Sci Nano 6:1716–1727
Zhou X, Wu F (2012) P-Coumaric acid influenced cucumber rhizosphere soil microbial communities and the growth of Fusarium oxysporum f. sp. cucumerinum Owen. PLoS One 7:e48288
Zitnick-Anderson K, Simons K, Pasche JS (2018) Detection and qPCR quantification of seven Fusarium species associated with the root rot complex in field pea. Can J Plant Pathol 40:261–271
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Gupta, K., Biswas, R., Sarkar, A. (2020). Advancement of Omics: Prospects for Bioremediation of Contaminated Soils. In: Shah, M. (eds) Microbial Bioremediation & Biodegradation. Springer, Singapore. https://doi.org/10.1007/978-981-15-1812-6_5
Download citation
DOI: https://doi.org/10.1007/978-981-15-1812-6_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-1811-9
Online ISBN: 978-981-15-1812-6
eBook Packages: Earth and Environmental ScienceEarth and Environmental Science (R0)