Abstract
Autophagy is an intracellular recycling process that maintains cellular homeostasis by orchestrating immunity upon viral infection. Following viral infection, autophagy is often initiated to curtail infection by delivering viral particles for lysosomal degradation and further integrating with innate pattern recognition receptor signaling to induce interferon (IFN)-mediated viral clearance. However, some viruses have evolved anti-autophagy strategies to escape host immunity and to promote viral replication. In this chapter, we illustrate how autophagy prevents viral infection to generate an optimal anti-viral milieu, and then concentrate on how viruses subvert and hijack the autophagic process to evade immunosurveillance, thereby facilitating viral replication and pathogenesis. Understanding the interplays between autophagy and viral infection is anticipated to guide the development of effective anti-viral therapeutics to fight against infectious diseases.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction
Autophagy, an evolutionarily conserved degradative process, is required to maintain organismal homeostasis and promote the clearance of intracellular waste and invading pathogens by the host immune system [76, 77]. Triggered by various physiological processes, autophagy is a frequent by-product of infection due to the cellular stress caused by viral infection and replication [22, 93]. Autophagy is also selective for the recognition and degradation of specific cargoes tagged by ubiquitination by a group of E3 ligase family proteins such as the tripartite motif (TRIM) proteins [142]. Depending on the cargos being sorted for destruction, selective autophagy can be classified into mitophagy (damaged mitochondria), pexophagy (peroxisome), ribophagy (ribosomes), ER-phagy (ER), glycophagy (glycogen), xenophagy (pathogens), and lipophagy (lipid droplets) [119]. Specifically, the xenophagy, a type of selective autophagy specifically senses intracellular microorganisms, including viruses, and physically targets them to autophagosomes for further degradation [75]. Autophagy, programmed to dispose of cytoplasmic components, is first activated by the innate immune system to degrade and clear invading viruses [22, 34], and then facilitates antigen processing followed by the induction of adaptive immune responses at later stages of infection [22, 103, 114].
Although autophagy aims at clearances, some viruses, those persisting and adaptable ones, have evolved a variety of strategies to inhibit, escape or manipulate multiple steps during autophagy to the elemental goal of survival and propagation. Physically, these viruses settle down in the membrane-bound, the protected environment offered by the autophagosome; and metabolically, they utilize autophagy-generated energy and metabolites. In short, these viruses suppress the autophagic process to avoid being degraded or use the autophagosome as the site for replication. Lipophagy, another form of autophagy that degrades intracellular lipid droplets, can also be manipulated by viruses [53]. Lipid droplets serve as a desirable platform for virion assembly, and directly inducing lipophagy allows viruses to sustain the high ATP levels needed for viral replication. In brief, current evidence supports the notion that viruses have evolved strategies to either combat or exploit autophagy to benefit their own life cycle and survival.
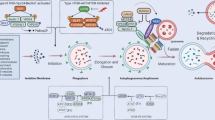
The viral proteins [19] (Fig. 5.1, Step 1) or any single step in the viral life cycle, including virus attachment and entry, membrane fusion, exposure of viral components and replication, may trigger autophagy [105]. We provide herein several representative examples to illustrate how viruses induce autophagy at multiple phases during infection (Fig. 5.1, Steps 2–5).
Viral infection induces autophagy initiation. (1) Viral protein itself is able to trigger autophagy. (2) The engagement of CD46, a ubiquitous human surface pathogen receptor for measles virus (MeV) is sufficient to induce autophagy through a CD46-Cyt1/GOPC pathway. (3) Fusion activity of the HIV-1 envelope glycoproteins gp120 and gp41 induce autophagy in uninfected CD4 T cells through binding to CD4 and CXCR4, leading to HIV entry, T cell apoptosis, and immunodeficiency. (4) The recognition process of certain viruses via TLR7 requires transport of cytosolic viral replication intermediates into the lysosome by autophagy. (5) Chikungunya virus (CHIKV)-triggered autophagy is mediated by the ER and oxidative stress pathways
The very first chance for viruses to induce autophagy is through virion binding [20]. CD46 serves as the binding and entry receptor for the measles virus (MeV) to induce autophagy by interacting with the phosphatidylinositol 3-kinase (VPS34)/Beclin-1 complex through a scaffold protein Golgi associated PDZ and coiled coil motif-containing (GOPC) (Fig. 5.1, Step 2). This pathway is only sensitive to vaccinal/attenuated strains that utilize CD46 to infect cells [59, 91, 96, 106, 112, 118]. In fact, MeV also activates autophagy by targeting an autophagy associated protein, named as immunity-associated GTPase family M (IRGM) [45, 46, 107].
Autophagy can also be activated by viral membrane fusion. Human immunodeficiency virus type 1 (HIV-1) envelope glycoproteins (gp120 and gp41, called Env) up-regulate autophagy with their fusogenic activity between HIV-infected cells and uninfected CD4 T cells, which could be prevented by HIV fusion inhibitors T20 and C34, leading to the cell death of uninfected T cells (Fig. 5.1, Step 3). There is not much known about the specific pathways and mechanisms that make up this process, although it hints that bioactive lipids involved in this fusion process and increased reactive oxygen species (ROS) production may mediate the activation of autophagy [33]. It is also important to note that the signaling activity of CD4 and CXCR4 are not associated with autophagy activated by Env [33].
Events following the fusion initiate autophagy via distinct mechanisms. For instance, the delivery of viral components into the cytosol can lead to cytosolic pattern recognition receptor (PRR)-induced autophagy. This is best illustrated by viral particles containing single-stranded RNA (ssRNA) such as vesicular stomatitis virus (VSV): the cytosolic viral replication intermediates in VSV-infected plasmacytoid dendritic cells (pDCs) can be introduced and transported by autophagy to lysosome compartment for the recognition by Toll-like receptor (TLR) 7, which leads to the activation type I interferon (IFN) signaling and the production of IFN-α. Atg5-deficient pDCs were not allowed to recognize VSV infection through TLR7, further demonstrating the critical step of VSV-induced autophagy in the host defense of viral infection (Fig. 5.1, Step 4) [71].
Viral replication offers a distinct model of deterioration in homeostasis leading to up-regulated autophagy [57]. One such example is the chikungunya virus (CHIKV) whose replication competent form has been reported to promote autophagy through endoplasmic reticulum (ER) stress and the generation of ROS [60]. ER stress is thought to be activated via the accumulation of viral polyproteins [51], which trigger the unfolded protein response (UPR) to restore homeostasis [51, 57]. In the case of CHIKV, its replication promotes the activation of inositol-requiring Ser/Thr protein kinase/endonuclease a (IRE-1a) pathway for provoking the UPR-elicited autophagy through the splicing of X-box-binding protein 1. CHIKV replication also induces increased ROS and reactive nitrogen species, which stimulates autophagy via AMPK-mediated inhibition of the mechanistic target of rapamycin complex 1 (mTORC1) (Fig. 5.1, Step 5) [60].
In summary, multiple steps during virus infection can activate autophagy by sensing of viral genomes or proteins, acting indirectly through cellular stress pathways to modulate homeostasis, and/or directly interacting with autophagy regulatory proteins.
5.2 Virus-Mediated Inhibition of Autophagy
Autophagy is a compilated but well-coordinated cellular event, which can be further divided into the processes of induction, nucleation of the phagophore, elongation, fusion, and degradation artificially [79]. Smart enough, viruses have evolved a variety of strategies to escape or manipulate these autophagic processes to benefit their own replication and propagation.
5.2.1 Inhibition of Autophagy Prior to the Initiation Phase
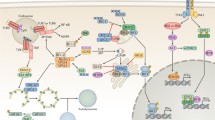
Mechanistic target of rapamycin (mTOR) works as a central homeostatic regulator of cell growth by promoting anabolic–metabolic processes like nucleotide synthesis and suppressing catabolic processes such as autophagy [64]. In light of the central role of mTORC1 in the prevention of autophagy, it is by no means out of the ordinary that some viruses have evolved tactics to boost mTORC1 activity, leading to the indirect suppression of the Beclin-1/PI3KIII complex and subsequent autophagy. HIV-1, whose envelope activates the mTOR pathway in dendritic cells (DCs), causes autophagy exhaustion [12]. Fusion-defective HIV-1 and CD4 agonist antibodies recapitulate these discoveries, underlining that HIV-1 might well suppress autophagy preceding viral entry (Fig. 5.2). Furthermore, the v-G protein-coupled receptor (v-GPCR), a Kaposi’s sarcoma-associated herpesvirus (KSHV) protein, activates the mTOR pathway to negatively regulate autophagy (Fig. 5.2) [8]. Aside from activating mTOR, v-GPCR is able to mimic the cellular homolog GPCR and down-regulates autophagy via suppressing ATG14L expression (Fig. 5.2) [152]. Interestingly, Beclin-2 may affect the v-GPCR protein level, enhancing its endolysosomal degradation [37].
Viral infection suppresses autophagy. Viruses have evolved a variety of strategies to escape or manipulate autophagic steps to benefit their own survival. HIV-1 envelope up-regulates the mTOR pathway in DCs, resulting in autophagy exhaustion which promotes cell-associated HIV-1 and transfer of HIV-1 infection to CD4 T cells. KSHV viral protein, v-GPCR which also modulates the mTOR signaling pathway. Besides activating mTOR, v-GPCR can mimic the cellular homolog GPCR and suppress autophagy by blocking the expression of ATG14L. ICP34.5 encoded by HSV-1, binds to Beclin-1 and inhibits autophagy function. And the mutant HSV-1 virus lacking the Beclin-1-binding domain of ICP34.5 cannot block autophagy in neurons. HCMV proteins, TRS1 and IRS1, suppress autophagosome biogenesis by interacting with Beclin-1. In addition, the anti-apoptotic protein, Bcl-2, interacts with the evolutionarily conserved autophagy protein, Beclin-1. The majority of members of the gamma-herpesvirus family encode and express vBcl-2, their cellular counterparts (cBCL-2), and inhibit autophagy by directly interacting with Beclin-1. Like KSHV encodes orf16 and MHV-68 produces M11. There is a checkpoint of the autophagy pathway in which cellular and KSHV FLIPs limit the Atg3-mediated step of LC3 conjugation to regulate autophagosome biogenesis. In addition, KSHV K7 protein interacts with the Rubicon autophagy protein and blocks the autophagosome maturation step by suppressing VPS34 enzymatic activity. Red line, inhibition; green line, promotion
5.2.2 Inhibition of Vesicle Nucleation
Nucleation of the phagophore is impelled by the induction of the Beclin-1/PI3KIII complex. Herpes simplex virus type 1 (HSV-1) encodes the neurovirulence factor ICP34.5 that binds to Beclin-1 and suppresses autophagy (Fig. 5.2) [100]. Human cytomegalovirus (HCMV) encodes a functional homolog of ICP34.5 called TRS1 that works against autophagy as well (Fig. 5.2) [17]. Unlike ICP34.5, the PKR binding domain of TRS1 is irrelevant to autophagy inhibition. Instead, TRS1 interacts with Beclin-1 through its N-terminal region, and this binding is indispensable to suppress autophagy. Besides TRS1, IRS1, another protein encoded by HCMV, has also been proven to inhibit autophagy by interacting with Beclin-1(Fig. 5.2) [95]. A myriad of viruses encode viral BCL-2 (vBCL-2), a protein mimicking its cellular counterpart (cBCL-2), and inhibit autophagy by directly interacting with Beclin-1 [104]. Biochemical and structural analyses showed that, compared with cBCL-2, vBCL-2 lacks the regulatory loop of cBCL-2, which is required for its phosphorylation by JUN N-terminal kinase (JNK). As a result, the association of cBCL-2 with Beclin-1 segregates Beclin-1 from the autophagy initiation complex, thereby attenuating the autophagosome formation [67, 149]. Human gamma-herpesvirus 4 (Epstein–Barr virus, EBV) encodes BHRF1 and BALF-1, two ortholog proteins of cellular Bcl-2, but their inhibitory effects on autophagy remain unclear [3]. Most members of the gamma-herpesvirus family encode and express vBcl-2 during their productive lytic infection process. For example, KSHV and murine γ-herpesvirus 68 (MHV68) use ORF16 and M11 to antagonize autophagy (Fig. 5.2) [30]. These studies have collectively illustrated that vBCL-2 has evolved to become a highly mighty autophagy suppressor.
5.2.3 Inhibition of Vesicle Elongation and Autophagosome Formation
KSHV encodes a homolog of the cellular FLICE-like inhibitor protein (FLIP; also known as ORF71), called vFLIP, that prevents ATG3 from binding to and processing LC3 in the autophagosome elongation process (Fig. 5.2) [72]. HSHV also expresses K7 that boosts the Rubicon–Beclin-1 interplay to attenuate the enzymatic activity of VPS34, thus hampering the fusion of autophagosomes with lysosomes (Fig. 5.2) [80].
5.3 Autophagy as a Mechanism of Promoting Virus Replication
Double-membrane compartments formed in autophagy serve as an excellent physical platform for viral replication, as they concentrate essential intermediates locally and prevent viral RNAs from detection with innate immune sensors and degradation. This phenomenon was first observed over three decades ago [9, 21, 40]. It is also important to note that RNA viruses are among the most frequent “hackers” of autophagy to promote their own replication.
5.3.1 (+) ssRNA Viruses
Studies have illustrated the accumulation of double-membrane vesicles (DMVs) following picornaviral infection. These small RNA viruses take advantage of autophagosomes as membrane scaffolds for their own RNA assembly and replication [2, 55, 148]. Moreover, the role of the autophagy machinery in inducing the non-lytic release of picornaviruses has emerged. Picornaviruses, a group of non-enveloped viruses, are conventionally thought to exit infected cells only through cell rupture. However, growing evidence shows that picornaviruses, including poliovirus (PV) and coxsackievirus, are able to spread in a non-lytic manner among cells via extracellular microvesicles (EMVs), including autophagosome derived EMVs (Fig. 5.3) [11, 18, 41, 113]. These viruses also acquire a defensive advantage by cloaking inside the host-derived vesicles to protect themselves against host immune assaults.
Viruses manipulate autophagy to promote their replication. Double-membrane vesicles (DMVs), following picornaviral infection, furnish the virus an excellent physical platform for viral RNA assembly and replication. And some picornaviruses, such as PV and coxsackievirus, can spread via extracellular microvesicles (EMVs) in a non-lytic manner between cells. The poliovirus triggered membranes can be specifically induced by the co-expression of two viral proteins, 2BC and 3A. HCV infection prompts the expression of Rubicon and UVRAG, which separately increases and decreases the maturation of autophagosomes. And Rubicon can be triggered by HCV NS4B protein alone. IRGM, known to contribute to autophagy, is localized at the Golgi apparatus and regulates the fragmentation of Golgi membranes in response to HCV infection, resulting in co-localization of Golgi vesicles with replicating HCV. Non-structural viral proteins 2B, 2C and 3A with LC3 and viral structural protein VP1 with Atg5, and LC3 with LAMP-1 co-localize in FMDV-infected cells. DENV stimulates and needs AMPK signaling and AMPK-independent suppression of mTORC1 activity for proviral lipophagy. ZIKV NS4A and NS4B, down-regulate cooperatively the Akt-mTOR pathway and induce cellular dysregulation. ZIKV NS3-mediated cleavage of FAM134B blocks the formation of ER and viral protein enriched autophagosomes, and the reticulophagy pathway further. As for HIV, nondegradative stages of autophagy promote its yields at early stage; HIV Gag-derived proteins bind to and interact with LC3, and autophagy enhances productive Gag processing. And when autophagy progresses to the maturation stages, HIV protein Nef plays as an anti-autophagic maturation factor by the interaction with the autophagy regulatory factor Beclin-1, which protects HIV from degradation. The dual and delicate interaction of HIV with the autophagy pathway enhances viral yields by utilizing the early stages while inhibiting the late stages. SARS-CoV and MHV, activate the formation of DMVs. And MHV utilizes the pathway of EDEMosome formation to generate the DMVs. IAV M2 interacts with LC3 and leads to LC3 re-localization. And a highly pathogenic avian H5N1 strain of IAV is able to block mTOR, activating autophagy. HPIV3 induces incomplete autophagy by blocking autophagosome-lysosome fusion, leading to increased virus production. The viral phosphoprotein binds to SNAP29 and suppresses its interaction with syntaxin17, therefore preventing these two host SNARE proteins from mediating autophagosome-lysome fusion. Matrix protein of HPIV3 shuttles to mitochondria and interacts with TUFM. The interaction between M and the LC3 protein that mediates autophagosome formation. These interactions with both TUFM and LC3 are required for the induction of mitophagy and result in inhibition of the type I interferon response. In RV-infected cells, RV NSP4 co-localized with LC3 in cap-like structures associated with viroplasms. And NSP4 enhances the release of calcium from the ER into the cytoplasm, leading to CaMKK-β signaling to trigger autophagy. HBV HBx maintains interrelationships with PI3KC3 and DAPK, and directly activates Beclin-1 to trigger autophagy. EBV LMP1 up-regulates PERK and the unfolded protein response to drive its own synthesis
The very first representative to show the benefits that viruses receive from remolding intracellular membranes is PV. Current evidence suggests that rapamycin, which induces autophagy, up-regulates poliovirus replication, while the silencing of some key genes of autophagosome formation down-regulates it [55]. PV is able to activate the formation of autophagosome-like membranes for RNA replication, virion maturation, and non-lytic viral spread [11, 55, 111]. A further study has shown that the PV protein, 2BC, alone is adequate for inducing the lipidation of LC3 but not for the construction of autophagosomes [139]. Nonetheless, the co-expression of both 2BC and 3A is able to promote the formation of DMVs containing markers of autophagosomes (Fig. 5.3) [55, 111, 131, 139]. Additionally, a recent study revealed that the ULK1 complex is non-essential for PV-induced autophagy [24].
Coxsackievirus B3 (CVB3) is in the same Picornaviridae family as PV. The mechanisms by which picornaviruses use to exploit autophagy for their benefits are still unclear. Whether picornaviral infection results in incomplete versus complete autophagy is disputable. Several studies have shown that CVB3 infection restricts the fusion of autophagosomes with lysosomes, leading to the production of giant autophagy-related vesicles during infections [63, 113, 148]. By contrast, another report suggests that CVB3 prompts complete autophagy [121]. A third recently published study showed that CVB3 infection compromises the autophagosome-lysosome/endosome fusion and, at least in part, promotes the accumulation of autophagosomes [94]. A new mechanism has been proposed: synaptosomal-associated protein 29 (SNAP29) and adaptor protein pleckstrin homology domain-containing protein family member 1 (PLEKHM1), known as regulators in autophagosome fusion, are both indispensable to the accumulation of autophagosomes. By cleaving SNAP29 and PLEKHM1 with proteinase 3C, CVB3 curtails autophagic flux and the resulting impaired versions of SNAP29/PLEKHM1 prompt viral replication [94].
Hepatitis C virus (HCV) induces autophagy by promoting the accumulation of autophagosomes and utilizing autophagosomal membranes as the spot for its RNA replication [1, 38, 122]. However, it is still controversial whether HCV is able to efficiently prompt the fusion between autophagosomes and lysosomes. Several studies lean toward the viewpoint that HCV induces autophagosome formation but obstructs the fusion to benefit viral replication and to prevent virion degradation [126, 127, 136]. For example, Sir et al. demonstrated that HCV induces the accumulation of autophagosomes without causing autophagic protein degradation in cells, and this inducement relies on UPR [126]. Dreux et al. suggested that the autophagy pathway is required for the translation of incoming HCV RNA but not for the maintenance of replication [39]. In contrast, Ke et al. found that the entire autophagic process used to complete autolysosome maturation is essential for supporting HCV RNA replication [62]. Nevertheless, during the early stage of infection, the HCV RNA-dependent RNA polymerase NS5B binds to ATG5, meaning that HCV utilizes ATG5 as a proviral factor at the onset of infection. The resultant downregulation of autophagy via ATG5 silencing obstructs HCV replication and persistence (Fig. 5.3) [47]. Two autophagy regulatory proteins, ultraviolet radiation resistance-associated gene protein (UVRAG), and Rubicon, expressed with different kinetics upon HCV infection activate and suppress the maturation of autophagosomes (Fig. 5.3). HCV is capable of temporally regulating autophagy by inducing the expression of these two proteins differentially to enhance its replication [145]. The early induction of Rubicon by HCV suppresses the fusion between autophagosomes and lysosomes, as a result of the accumulation of autophagosomes and encouragement of HCV replication [145]. Additionally, immunity-related GTPase family M protein (IRGM), an IFN-inducible GTPase, has been reported to regulate autophagy and the development of a variety of intracellular membrane compartments [46]. Upon HCV infection, IRGM interacts with Golgi apparatus-specific brefeldin A-resistance guanine nucleotide exchange factor 1 (GBF1) and facilitates AMPK-mediated GBF1 phosphorylation, thus activating GTPase ADB ribosylation factor 1 (ARF1) for Golgi apparatus fragmentation and coordinating viral replication (Fig. 5.3) [49]. Furthermore, the IRGM-mediated phosphorylation of ULK1 is triggered by HCV infection [16]. The sum of evidence points to the fact that HCV dynamically modulates autophagy to promote viral replication (Fig. 5.3).
Similarly, Foot-and-mouth disease virus (FMDV) leads to ATG5-dependent autophagosome formation as well as the redistribution of LC3 to punctate vesicles. The PI3K activity of VPS34 is non-essential for this induction and occurs very early, as ultraviolet-inactivated FMDV is still able to provoke the autophagosome formation [6]. In addition, co-localization of viral non-structural proteins 2B, 2C, and 3A with LC3 was observed and autophagosomes induced by FMDV contained VP1, the viral capsid protein, which co-localizes with p62, suggesting that autophagosome formation is activated at FMDV entry (Fig. 5.3) [97]. A recent study offered evidence that the expression of FMDV capsid protein VP2 is able to induce autophagy through the EIF2S1-ATF4-AKT-mTOR cascade. VP2 was found to interact with HSPB1 (heat shock protein beta-1) and up-regulate the EIF2S1-ATF4 signaling, leading to autophagy and enhanced FMDV replication [135].
Dengue virus (DENV) has been reported to activate the proliferation of LC3-containing membranes [73, 92]. Using 3-methyladenine or spautin-1, two autophagy inhibitors affect DENV infection [52, 89]. Lipophagy, a form of autophagy, regulates the storage of cellular lipids by lysosomal degradation [84, 124]. Within starving cells, lipophagy breaks down lipid droplets (LDs), in which the eukaryotic cells stock lipids to provide mitochondria with fatty acids, which are oxidized to create acetyl-CoA [84]. Viruses can also take advantage of lipophagy for their own benefits. The number of LDs is increased in DENV-infected cells, and in turn, the inhibition of LD formation remarkably damages DENV replication. Viral capsid proteins are contained in these LDs, which means that these DENV-induced LDs offer a platform for nucleocapsid formation as well as viral replication [117]. Moreover, lipophagy is activated in DENV-infected cells; and the stored triglycerides are depleted. β-oxidation and energy production are increased in this process, which creates a seedbed of viral replication. If exogenous free fatty acids are added into autophagy-deficient cells, DENV replication will be rescued. Etomoxir, a drug that blocks fatty acid transport into the mitochondria, will prevent it [52]. So, the quantity of free fatty acids and ATP released by lipophagy may be required for DENV replication and persistence. Moreover, DENV induces AMPK kinase activity, which prohibits mTORC1, and this modulation is crucial for virus-induced lipophagy (Fig. 5.3) [58]. Recently, AUP1, a lipid droplet-localized type-III membrane protein with dual localization marks for LDs and ER, was shown to be utilized by DENV to trigger lipophagy. Interaction of unmodified AUP1with the viral non-structural proteins NS4A and NS4B in DENV-infected cells triggers the acyltransferase activity of AUP1, generating phospholipids as the source of membrane components necessary for lipophagy formation and subsequent viral replication [151]. This mechanism seems to be a general phenomenon in flaviviruses and underlies the key role of post-translational modifications during viral infections [151].
Zika virus (ZIKV) has been found to induce the formation of LC3-containing membranes as well [81]. Moreover, the spread of ZIKV might be up-regulated by noncanonical secretory autophagy, as it is for PV and CVB3 [153]. In human neural progenitor cells ER rearrangement and the formation of vesicular clusters in ZIKV infection were thought to be the sites for viral RNA replication and virion assembly [25, 98]. In ZIKV-infected primary fibroblasts, multi-membrane structures are formed resembling autophagic vesicles [48]. In addition, increased lapidated LC3 in ZIKV-infected placentae and decreased viral titers in ATG16-deficient mouse fetuses both work in favor of the proviral role of autophagy [15]. Zika virus (ZIKV) utilizes the ER as a source of membranes to establish their viral replication, assembly and maturation. A selective form of ER degradation by autophagy, or reticulophagy has evolved in the host to restrict DENV and ZIKV, mediated by an ER-resident reticulophagy receptor FAM134B [7]. The virally encoded proteases NS3 in several flaviviruses including ZIKV, DENV, and West Nile Virus (WNV) cleaves FAM134B to suppress the formation of ER and viral protein enriched autophagosomes, as a strategy that viruses manipulate autophagy for their replication (Fig. 5.3) [74]. Furthermore, upon ZIKV infection in fetal neural stem cells AKT phosphorylation and subsequent mTOR activation will be inhibited through the viral protein NS4A and NS4B, which leads to the aberrant activation of autophagy and defective neurogenesis, thus promoting viral replication (Fig. 5.3) [81].
HIV skillfully manipulates the autophagy process by utilizing its two proteins to interact with two different autophagic factors separately. On the one hand, HIV Gag-derived proteins co-localize with and bind to LC3, and autophagy supports productive Gag processing in early and nondegradative stages of autophagy to promote HIV yields (Fig. 5.3). On the other hand, when autophagy enters its maturation stages, HIV protein Nef serves as an anti-autophagic maturation factor through interactions with Beclin-1, thus protecting HIV from degradation. Therefore, the perturbation of the early and late stage of autophagy process promotes HIV survival and replication [68]. However, during permissive infection, HIV attenuates autophagy in order to avoid proteolytic degradation. Normally, mTOR phosphorylates transcription factor EB (TFEB) and restricts its translocation by favoring its retention within the cytosol. When mTOR is suppressed, TFEB gets dephosphorylated and is allowed to transfer to the nucleus, where it can promote autophagy and lysosomal gene expression. Within infected macrophages, the interplay between HIV and TLR8 activates autophagy, which relies on the dephosphorylation and nuclear translocation of TFEB. During permissive infection, Nef interacts with Beclin-1, leading to mTOR activation, TFEB phosphorylation, and cytosolic sequestration, as well as the suppression of autophagy [14].
5.3.2 (–) ssRNA Viruses
Upon infection, Coronaviruses (CoVs) like the severe acute respiratory syndrome coronavirus (SARS-CoV) and mouse hepatitis virus (MHV) activate the formation of DMVs in host cells and target their replication and transcription complexes (RTCs) on the DMVs-limiting membranes [31, 43, 116]. However, the exact derivation of the DMV lipid bilayers, the host protein content, and the identification of the cellular factors essential for DMVs formation remains unclear [66]. The probable participation of autophagy in the conversion of host membranes into DMVs has been reported [26, 87, 134]. The precise mechanisms that explain why CoVs limits subsequent autophagosome expansion are still a mystery. The non-structural protein 6 (NSP6) has been reported to trigger the autophagic pathway and limit autophagosome expansion to favor CoVs infection [26, 27]. Atg5, according to a study, is non-essential for MHV replication [154]. Contradictory evidence showed either the presence [108, 154] or the absence [31, 129] of LC3/Atg8 on DMVs. Another theory about the origin of virus-induced DMVs suggests that these DMVs are part of a reticulovesicular network of modified ER membranes and contain dsRNA in their interior, which came from a natural and intuitive analysis of SARS-CoV and MHV-infected cells via electron tomography [66]. This idea has been supported by several findings [50, 61, 99]. NSP4, when separately expressed, was shown to localize to the ER and then translocate to the DMVs upon infection [99]. But the deficiency of ER, ER-Golgi intermediate compartment (ERGIC), or Golgi protein markers within CoV-induced DMVs might well mean that their biogenesis does not rely on the traditional pathway [99, 129, 143]. Of special interest is a study that determined MHV hijacks the pathway of EDEMosome (a vesicle involved in ER-associated degradation, ERAD) formation to generate the DMVs (Fig. 5.3). In doing so, MHV trapped two ERAD regulatory proteins into the DMVs, and therefore, exploited the ERAD pathway for viral replication [109]. In addition, this study also revealed an autophagy-independent role for nonlipidated LC3-I [109].
Influenza A virus (IAV) infection also triggers the accumulation of autophagosomes for viral replication [157]. IAV Matrix 2 (M2) ion-channel protein is credited with the manipulation of autophagy, which blocks the fusion of autophagosomes with lysosomes [42]. Further study showed that M2 hijacks autophagy with its LC3-interacting region [4]. M2 interacts with LC3 and induces LC3 re-localization to the plasma membrane, and disruption of this interaction down-regulates virion budding and stability (Fig. 5.3). Another protein, NS1 triggers autophagy by promoting the synthesis of hemagglutinin (HA) and M2 [155]. Recently, IAV M2 protein was reported to interact with MAVS and positively regulate MAVS-mediated innate immunity. Moreover, ROS production induced by M2 is pivotal for the activation of autophagy and the amplification of the MAVS signaling pathway [146]. In addition, a highly pathogenic avian H5N1 strain of IAV is able to activate autophagy by inhibiting mTOR [85].
Human parainfluenza virus type 3 (HPIV3) suppresses autophagosome maturation as well as triggers the accumulation of autophagosomes [35]. HPIV3 phosphoprotein (P) binds to the SNARE domains of SNAP29 and blocks the interaction between STX17 and SNAP29, which eventually prevents autophagosome-lysosome fusion (Fig. 5.3) [35]. In addition, the matrix protein (M) of HPIV3 interacts with TUFM and binds LC3 to trigger TUFM-mediated mitophagy (Fig. 5.3), a form of autophagy that selectively removes damaged mitochondria and suppresses the subsequent IFN response. These findings suggest that a viral protein is enough to activate mitophagy via bridging autophagosomes and mitochondria [36].
5.3.3 dsRNA Viruses
Within rotavirus (RV)-infected cells, NSP4, whose appearance relies on the intracellular calcium levels co-localizes with LC3 on viroplasms, sites of viral genome replication and immature particle assembly [5]. Further study found that NSP4 activates the release of calcium from the ER into the cytoplasm, inducing calcium/calmodulin-dependent kinase kinase-β (CaMKK-β) signaling to trigger autophagy (Fig. 5.3) [28, 29]. Besides CaMKK-β signaling, a mutually complementary mechanism about a new small RNA was found in RV-initiated autophagy. RV-vsRNA1755 encoded by the NSP4 gene targets the host cell IGF1R which is the part of the PI3K/Akt/mTOR signaling process. In the initial stage of infection RV-vsRNA1755 activates autophagy by obstructing induction of the mTOR pathway [156].
5.3.4 dsDNA Virus
Hepatitis B virus (HBV) has been shown to induce autophagy whether it is in its productive or nonproductive cycles making autophagy vital for its replication [125, 128, 140]. Hepatitis B x protein (HBx) has been linked to an extraordinarily diverse group of pathways, like ones that maintain interrelationships with PI3KC3, or the ones that induce death associated protein kinase (DAPK) in a way that involves Beclin-1 [150], or the ones that directly activate Beclin-1 expression [137] to trigger autophagy (Fig. 5.3). Another one of its encoded proteins SHBs, can induce autophagy as well [78]. As an intermediate process, the accumulation of autophagosomes mirrors the balance between the rate of their generation and conversion into autolysosomes. Tang et al. suggested the view that HBx, at the initiation stage of autophagic progression triggers autophagy in a Beclin-1-dependent fashion (Fig. 5.3) [137]. Wang et al. suggested that HBV induces autophagy at the initiation stage by the interaction of HBx and c-myc to influence miR-192-3p-XIAP, which in turn regulates Beclin-1 [144]. Meanwhile, Liu et al. revealed that in the late phase of autophagy HBx induces the formation of autophagosomes where HBx evidently damages the lysosomal degradative ability [83]. And they partly supported the conclusion of Sir et al., which stated that HBx is enough to induce autophagosomes [83].
It has also been reported that human gamma-herpesvirus 4 (Epstein–Barr virus, EBV) employ several strategies to interact with autophagic proteins and favor their own survival [23, 90, 123]. Specific autophagy inhibitors are able to encourage EBV lytic replication and might very well influence its oncogenesis [32]. The six-transmembrane spanning domains (6TM) of LMP1 up-regulate PERK, resulting in UPR-mediated autophagy (Fig. 5.3) [69, 70]. Moreover, EBNA1-fragments instead of EBNA3C and EBNA2 are presented via MHC class-II through the autophagy-lysosomal process [138]. And, the accumulation of EBNA1 in autophagosomes suppresses the lysosomal acidification, resulting in a reduction of EBNA1-antigen presentation for CD4+ T lymphocytes recognition [138]. In summary, these findings illustrated that EBV latent antigens hijack autophagy and subsequently influence B-cell lymphomagenesis.
The first study aiming at understanding the implication of autophagy on KSHV replication was performed by Wen et al., who believed that KSHV replication and transcription activator (RTA) enhances autophagy activation to facilitate KSHV lytic replication [147]. Later on, Granato et al. confirmed the function of autophagy in provoking KSHV replication triggered by RTA as well as butyrate combination (T/B), which revealed that the last autophagic steps are suppressed [44].
5.4 Autophagy-Mediated Restriction of Viral Replication
As a piece of vital machinery that responds to environmental stresses rapidly, it is not shocking that autophagy plays a pivotal role in both innate and adaptive immunity to keep cellular homeostasis [114]. But, here, we illustrate that autophagy restricts viral replication by degrading viral components, viral particles or even host factors required for viral replication rather than cooperating with innate and/or adaptive immunity. This process of autophagy targeting individual viral components for degradation is termed virophagy [102]. It’s important to note that virophagy targets neosynthesized viral components, while xenophagy targets entire viral particles [86].
Core proteins for HCV virion particles assembly and release are mainly localized within the ER [56]. Overload of HCV in infected cells induces ER stress-associated HPR and subsequent autophagy activation to promote viral replication [62]. The abilities of HCV to evade autophagic destruction and make use of autophagy for its own benefit have been extensively studied. However, a recent study highlights that an IFN-β-inducible SCOTIN (ER-resident protein, also named SHISA5) recruits HCV non-structural protein 5A (NS5A) to autophagysome for degradation, thereafter suppressing HCV replication (Fig. 5.4) [65].
Autophagy-mediated restriction of viral replication. HCV NS5A protein interacts with the IFN-β-inducible protein SHISA5, which transfers NS5A to autophagosomes for further degradation. SMURF1 is indispensable for the co-localization of the SINV capsid protein to p62, which prompts virophagy by shuttling the viral capsid to autophagosomes. FANCC also interacts with SINV capsid protein (not known to be ubiquitinated) and enhance virophagy. Poliovirus breaks the endosomal membrane and releases its genome into the cytoplasm, and Galectin-8 detects the permeated endosomes and marks them for autophagic degradation, but PLA2G16 facilitates viral genome translocation and prevents clearance. Upon HIV viral fusion, TRIM5α induces the recruitment of Atg5 to the TRIM5α–Atg16L1–HIV-1p24 capsid complex, promoting lipidation of LC3 (LC3 II) and thereby mediating autophagosome formation. HIV Vif interacts with the HD6A/APOBEC3G complex to induce its rapid degradation. Autophagy selectively degrades the HIV-1 transactivator Tat, a protein that is essential for HIV-1 transcription and virion production
Upon the Sindbis virus (SINV) infection, Beclin-1 was and Atg5 were reported to protect the host from SINV-mediated encephalitis [82, 101]. Interestingly, knockdown of p62 or other autophagy-related genes up-regulates viral capsid accumulation and progresses virus-induced cell death without influencing virus replication [101]. An E3-ubiquitin ligase, SMURF1 is indispensable for the co-localization of the SINV capsid protein with p62; this interaction advances virophagy by allowing the movement of the SINV viral capsid to autophagosomes [102]. The Fanconi anemia group C protein (FANCC) was also reported to interact with the SINV capsid protein and enhance virophagy [132, 133]. The fact that SMURF1 and FANCC target HSV-1 for virophagy as well and suggests that they often function as virophagic factors (Fig. 5.4) [133].
Picornaviruses are sensed by galectin 8 which restricts viral infection by triggering the autophagic degradation of the viral RNA genome [130]. When poliovirus pierces the endosomal membrane to dump its genome into the cytoplasm, β-galactosides are exposed and activate galectin 8 which results in the detection of punctured endosomes and marks them for further autophagic degradation. Poliovirus, in turn, utilizes the host protein HRAS-like suppressor 3 (PLA2G16) to escape this detection and help genome delivery. Coxsackievirus B3 (CVB3), another picornavirus cleaves p62 and inhibits virophagy by hijacking the viral protease 2A (Fig. 5.4) [120].
HIV-1 is subjected to autophagic degradation as well. In order to surmount innate immunity, the virion infectivity factor (Vif) induces the degradation of an HIV-1 restriction factor APOBEC3G favoring the HIV replication [88]. However, histone deacetylase 6 (HDAC6), in turn, forms a complex with APOBEC3G and provokes autophagy-dependent Vif degradation which down-regulates HIV-1 replication (Fig. 5.4) [141]. Moreover, within CD4+ T cells the transactivator Tat, a protein that promotes viral transcription was selectively degraded by autophagy [115]. In Langerhans cells, which are dendritic immune skin cells, HIV was degraded by the restriction factor tripartite motif-containing protein 5α (TRIM5α) and its ability to regulate the assembly of autophagy activating complexes (Fig. 5.4) [110].
Autophagy also possesses anti-viral capabilities independent of its role in degradation. For mouse norovirus (MNV) infection in vivo, the ATG5/ATG12/ATG16L1 complexplays a key role in autophagosome formation and is essential for IFNγ-mediated anti-viral defense [54]. In ATG16L1 hypomorphic mice, MNV infection induced a phenotype that resembled Crohn’s disease [13]. Interestingly, the initiation, fusion, and degradative activities of autophagy were indispensable; while IFNγ-inducible GTPases, which were targeted to MNV replication complexes by LC3 suppressed viral replication [10].
5.5 Conclusion
The control of viral infection by autophagy is a multi-faceted, dynamic physiological, and pathological process. On one hand, autophagy destructs viruses, regulates inflammatory responses, and provokes antigen presentation. On the other hand, viruses try all means to enhance their immune escape, replication, and release from infected cells by sabotaging or taking advantage of autophagy. Different virus finds its own strategies to survive the autophagic destruction while secures the membrane source provided by autophagy for viral replication. In summary, autophagy and viral infection are highly connected and continuing investigations on the virus autophagy interplays will be a fruitful area of scientific inquiry for many years to come.
References
Ait-Goughoulte M, Kanda T, Meyer K, Ryerse JS, Ray RB, Ray R (2008) Hepatitis C virus genotype 1a growth and induction of autophagy. J Virol 82:2241–2249
Alirezaei M, Flynn CT, Wood MR, Whitton JL (2012) Pancreatic acinar cell-specific autophagy disruption reduces coxsackievirus replication and pathogenesis in vivo. Cell Host Microbe 11:298–305
Altmann M, Hammerschmidt W (2005) Epstein-Barr virus provides a new paradigm: a requirement for the immediate inhibition of apoptosis. PLoS Biol 3:e404
Beale R, Wise H, Stuart A, Ravenhill BJ, Digard P, Randow F (2014) A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability. Cell Host Microbe 15:239–247
Berkova Z, Crawford SE, Trugnan G, Yoshimori T, Morris AP, Estes MK (2006) Rotavirus NSP4 induces a novel vesicular compartment regulated by calcium and associated with viroplasms. J Virol 80:6061–6071
Berryman S, Brooks E, Burman A, Hawes P, Roberts R, Netherton C, Monaghan P, Whelband M, Cottam E, Elazar Z et al (2012) Foot-and-mouth disease virus induces autophagosomes during cell entry via a class III phosphatidylinositol 3-kinase-independent pathway. J Virol 86:12940–12953
Bhaskara RM, Grumati P, Garcia-Pardo J, Kalayil S, Covarrubias-Pinto A, Chen W, Kudryashev M, Dikic I, Hummer G (2019) Curvature induction and membrane remodeling by FAM134B reticulon homology domain assist selective ER-phagy. Nat Commun 10:2370
Bhatt AP, Damania B (2012) AKTivation of PI3K/AKT/mTOR signaling pathway by KSHV. Front Immunol 3:401
Bienz K, Egger D, Pasamontes L (1987) Association of polioviral proteins of the P2 genomic region with the viral replication complex and virus-induced membrane synthesis as visualized by electron microscopic immunocytochemistry and autoradiography. Virology 160:220–226
Biering SB, Choi J, Halstrom RA, Brown HM, Beatty WL, Lee S, McCune BT, Dominici E, Williams LE, Orchard RC et al (2017) Viral replication complexes are targeted by LC3-guided interferon-inducible GTPases. Cell Host Microbe 22(74–85):e77
Bird SW, Maynard ND, Covert MW, Kirkegaard K (2014) Nonlytic viral spread enhanced by autophagy components. Proc Natl Acad Sci USA 111:13081–13086
Blanchet FP, Moris A, Nikolic DS, Lehmann M, Cardinaud S, Stalder R, Garcia E, Dinkins C, Leuba F, Wu L et al (2010) Human immunodeficiency virus-1 inhibition of immunoamphisomes in dendritic cells impairs early innate and adaptive immune responses. Immunity 32:654–669
Cadwell K, Patel KK, Maloney NS, Liu TC, Ng AC, Storer CE, Head RD, Xavier R, Stappenbeck TS, Virgin HW (2010) Virus-plus-susceptibility gene interaction determines Crohn’s disease gene Atg16L1 phenotypes in intestine. Cell 141:1135–1145
Campbell GR, Rawat P, Bruckman RS, Spector SA (2015) Human immunodeficiency virus Type 1 Nef Inhibits autophagy through transcription factor EB sequestration. PLoS Pathog 11:e1005018
Cao B, Parnell LA, Diamond MS, Mysorekar IU (2017) Inhibition of autophagy limits vertical transmission of Zika virus in pregnant mice. J Exp Med 214:2303–2313
Chauhan S, Mandell MA, Deretic V (2015) IRGM governs the core autophagy machinery to conduct antimicrobial defense. Mol Cell 58:507–521
Chaumorcel M, Lussignol M, Mouna L, Cavignac Y, Fahie K, Cotte-Laffitte J, Geballe A, Brune W, Beau I, Codogno P et al (2012) The human cytomegalovirus protein TRS1 inhibits autophagy via its interaction with Beclin 1. J Virol 86:2571–2584
Chen YH, Du W, Hagemeijer MC, Takvorian PM, Pau C, Cali A, Brantner CA, Stempinski ES, Connelly PS, Ma HC et al (2015) Phosphatidylserine vesicles enable efficient en bloc transmission of enteroviruses. Cell 160:619–630
Cheng CY, Chi PI, Liu HJ (2014) Commentary on the regulation of viral proteins in autophagy process. Biomed Res Int 2014:962915
Chiramel AI, Brady NR, Bartenschlager R (2013) Divergent roles of autophagy in virus infection. Cells 2:83–104
Cho MW, Teterina N, Egger D, Bienz K, Ehrenfeld E (1994) Membrane rearrangement and vesicle induction by recombinant poliovirus 2C and 2BC in human cells. Virology 202:129–145
Choi Y, Bowman JW, Jung JU (2018) Autophagy during viral infection—a double-edged sword. Nat Rev Microbiol 16:341–354
Cirone M (2018) EBV and KSHV infection dysregulates autophagy to optimize viral replication, prevent immune recognition and promote tumorigenesis. Viruses 10
Corona Velazquez A, Corona AK, Klein KA, Jackson WT (2018) Poliovirus induces autophagic signaling independent of the ULK1 complex. Autophagy 14:1201–1213
Cortese M, Goellner S, Acosta EG, Neufeldt CJ, Oleksiuk O, Lampe M, Haselmann U, Funaya C, Schieber N, Ronchi P et al (2017) Ultrastructural characterization of Zika virus replication factories. Cell Rep 18:2113–2123
Cottam EM, Maier HJ, Manifava M, Vaux LC, Chandra-Schoenfelder P, Gerner W, Britton P, Ktistakis NT, Wileman T (2011) Coronavirus nsp6 proteins generate autophagosomes from the endoplasmic reticulum via an omegasome intermediate. Autophagy 7:1335–1347
Cottam EM, Whelband MC, Wileman T (2014) Coronavirus NSP6 restricts autophagosome expansion. Autophagy 10:1426–1441
Crawford SE, Estes MK (2013) Viroporin-mediated calcium-activated autophagy. Autophagy 9:797–798
Crawford SE, Hyser JM, Utama B, Estes MK (2012) Autophagy hijacked through viroporin-activated calcium/calmodulin-dependent kinase kinase-beta signaling is required for rotavirus replication. Proc Natl Acad Sci USA 109:E3405–E3413
Cuconati A, White E (2002) Viral homologs of BCL-2: role of apoptosis in the regulation of virus infection. Genes Dev 16:2465–2478
de Haan CA, Reggiori F (2008) Are nidoviruses hijacking the autophagy machinery? Autophagy 4:276–279
De Leo A, Colavita F, Ciccosanti F, Fimia GM, Lieberman PM, Mattia E (2015) Inhibition of autophagy in EBV-positive Burkitt’s lymphoma cells enhances EBV lytic genes expression and replication. Cell Death Dis 6:e1876
Denizot M, Varbanov M, Espert L, Robert-Hebmann V, Sagnier S, Garcia E, Curriu M, Mamoun R, Blanco J, Biard-Piechaczyk M (2008) HIV-1 gp41 fusogenic function triggers autophagy in uninfected cells. Autophagy 4:998–1008
Deretic V, Saitoh T, Akira S (2013) Autophagy in infection, inflammation and immunity. Nat Rev Immunol 13:722–737
Ding B, Zhang G, Yang X, Zhang S, Chen L, Yan Q, Xu M, Banerjee AK, Chen M (2014) Phosphoprotein of human parainfluenza virus type 3 blocks autophagosome-lysosome fusion to increase virus production. Cell Host Microbe 15:564–577
Ding B, Zhang L, Li Z, Zhong Y, Tang Q, Qin Y, Chen M (2017) The matrix protein of Human Parainfluenza virus type 3 induces Mitophagy that suppresses interferon responses. Cell Host Microbe 21(538–547):e534
Dong X, Cheng A, Zou Z, Yang YS, Sumpter RM Jr, Huang CL, Bhagat G, Virgin HW, Lira SA, Levine B (2016) Endolysosomal trafficking of viral G protein-coupled receptor functions in innate immunity and control of viral oncogenesis. Proc Natl Acad Sci USA 113:2994–2999
Dreux M, Chisari FV (2009) Autophagy proteins promote hepatitis C virus replication. Autophagy 5:1224–1225
Dreux M, Gastaminza P, Wieland SF, Chisari FV (2009) The autophagy machinery is required to initiate hepatitis C virus replication. Proc Natl Acad Sci USA 106:14046–14051
Egger D, Teterina N, Ehrenfeld E, Bienz K (2000) Formation of the poliovirus replication complex requires coupled viral translation, vesicle production, and viral RNA synthesis. J Virol 74:6570–6580
Feng Z, Hensley L, McKnight KL, Hu F, Madden V, Ping L, Jeong SH, Walker C, Lanford RE, Lemon SM (2013) A pathogenic picornavirus acquires an envelope by hijacking cellular membranes. Nature 496:367–371
Gannage M, Dormann D, Albrecht R, Dengjel J, Torossi T, Ramer PC, Lee M, Strowig T, Arrey F, Conenello G et al (2009) Matrix protein 2 of influenza A virus blocks autophagosome fusion with lysosomes. Cell Host Microbe 6:367–380
Gosert R, Kanjanahaluethai A, Egger D, Bienz K, Baker SC (2002) RNA replication of mouse hepatitis virus takes place at double-membrane vesicles. J Virol 76:3697–3708
Granato M, Santarelli R, Filardi M, Gonnella R, Farina A, Torrisi MR, Faggioni A, Cirone M (2015) The activation of KSHV lytic cycle blocks autophagy in PEL cells. Autophagy 11:1978–1986
Gregoire IP, Rabourdin-Combe C, Faure M (2012) Autophagy and RNA virus interactomes reveal IRGM as a common target. Autophagy 8:1136–1137
Gregoire IP, Richetta C, Meyniel-Schicklin L, Borel S, Pradezynski F, Diaz O, Deloire A, Azocar O, Baguet J, Le Breton M et al (2011) IRGM is a common target of RNA viruses that subvert the autophagy network. PLoS Pathog 7:e1002422
Guevin C, Manna D, Belanger C, Konan KV, Mak P, Labonte P (2010) Autophagy protein ATG5 interacts transiently with the hepatitis C virus RNA polymerase (NS5B) early during infection. Virology 405:1–7
Hamel R, Dejarnac O, Wichit S, Ekchariyawat P, Neyret A, Luplertlop N, Perera-Lecoin M, Surasombatpattana P, Talignani L, Thomas F et al (2015) Biology of Zika virus infection in human skin cells. J Virol 89:8880–8896
Hansen MD, Johnsen IB, Stiberg KA, Sherstova T, Wakita T, Richard GM, Kandasamy RK, Meurs EF, Anthonsen MW (2017) Hepatitis C virus triggers Golgi fragmentation and autophagy through the immunity-related GTPase M. Proc Natl Acad Sci USA 114:E3462–E3471
Harcourt BH, Jukneliene D, Kanjanahaluethai A, Bechill J, Severson KM, Smith CM, Rota PA, Baker SC (2004) Identification of severe acute respiratory syndrome coronavirus replicase products and characterization of papain-like protease activity. J Virol 78:13600–13612
He B (2006) Viruses, endoplasmic reticulum stress, and interferon responses. Cell Death Differ 13:393–403
Heaton NS, Randall G (2010) Dengue virus-induced autophagy regulates lipid metabolism. Cell Host Microbe 8:422–432
Heaton NS, Randall G (2011) Dengue virus and autophagy. Viruses 3:1332–1341
Hwang S, Maloney NS, Bruinsma MW, Goel G, Duan E, Zhang L, Shrestha B, Diamond MS, Dani A, Sosnovtsev SV et al (2012) Nondegradative role of Atg5–Atg12/ Atg16L1 autophagy protein complex in anti-viral activity of interferon gamma. Cell Host Microbe 11:397–409
Jackson WT, Giddings TH Jr, Taylor MP, Mulinyawe S, Rabinovitch M, Kopito RR, Kirkegaard K (2005) Subversion of cellular autophagosomal machinery by RNA viruses. PLoS Biol 3:e156
Jones DM, McLauchlan J (2010) Hepatitis C virus: assembly and release of virus particles. J Biol Chem 285:22733–22739
Jordan TX, Randall G (2012) Manipulation or capitulation: virus interactions with autophagy. Microbes Infect 14:126–139
Jordan TX, Randall G (2017) Dengue virus activates the AMP kinase-mTOR axis to stimulate a proviral lipophagy. J Virol 91
Joubert PE, Meiffren G, Gregoire IP, Pontini G, Richetta C, Flacher M, Azocar O, Vidalain PO, Vidal M, Lotteau V et al (2009) Autophagy induction by the pathogen receptor CD46. Cell Host Microbe 6:354–366
Joubert PE, Werneke SW, de la Calle C, Guivel-Benhassine F, Giodini A, Peduto L, Levine B, Schwartz O, Lenschow DJ, Albert ML (2012) Chikungunya virus-induced autophagy delays caspase-dependent cell death. J Exp Med 209:1029–1047
Kanjanahaluethai A, Chen Z, Jukneliene D, Baker SC (2007) Membrane topology of murine coronavirus replicase nonstructural protein 3. Virology 361:391–401
Ke PY, Chen SS (2011) Activation of the unfolded protein response and autophagy after hepatitis C virus infection suppresses innate anti-viral immunity in vitro. J Clin Invest 121:37–56
Kemball CC, Alirezaei M, Flynn CT, Wood MR, Harkins S, Kiosses WB, Whitton JL (2010) Coxsackievirus infection induces autophagy-like vesicles and megaphagosomes in pancreatic acinar cells in vivo. J Virol 84:12110–12124
Kim J, Guan KL (2019) mTOR as a central hub of nutrient signalling and cell growth. Nat Cell Biol 21:63–71
Kim N, Kim MJ, Sung PS, Bae YC, Shin EC, Yoo JY (2016) Interferon-inducible protein SCOTIN interferes with HCV replication through the autolysosomal degradation of NS5A. Nat Commun 7:10631
Knoops K, Kikkert M, Worm SH, Zevenhoven-Dobbe JC, van der Meer Y, Koster AJ, Mommaas AM, Snijder EJ (2008) SARS-coronavirus replication is supported by a reticulovesicular network of modified endoplasmic reticulum. PLoS Biol 6:e226
Ku B, Woo JS, Liang C, Lee KH, Hong HS, Xiaofei E, Kim KS, Jung JU, Oh BH (2008) Structural and biochemical bases for the inhibition of autophagy and apoptosis by viral BCL-2 of murine gamma-herpesvirus 68. PLoS Pathog 4:e25
Kyei GB, Dinkins C, Davis AS, Roberts E, Singh SB, Dong C, Wu L, Kominami E, Ueno T, Yamamoto A et al (2009) Autophagy pathway intersects with HIV-1 biosynthesis and regulates viral yields in macrophages. J Cell Biol 186:255–268
Lee DY, Sugden B (2008) The latent membrane protein 1 oncogene modifies B-cell physiology by regulating autophagy. Oncogene 27:2833–2842
Lee DY, Sugden B (2008) The LMP1 oncogene of EBV activates PERK and the unfolded protein response to drive its own synthesis. Blood 111:2280–2289
Lee HK, Lund JM, Ramanathan B, Mizushima N, Iwasaki A (2007) Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315:1398–1401
Lee JS, Li Q, Lee JY, Lee SH, Jeong JH, Lee HR, Chang H, Zhou FC, Gao SJ, Liang C et al (2009) FLIP-mediated autophagy regulation in cell death control. Nat Cell Biol 11:1355–1362
Lee YR, Lei HY, Liu MT, Wang JR, Chen SH, Jiang-Shieh YF, Lin YS, Yeh TM, Liu CC, Liu HS (2008) Autophagic machinery activated by dengue virus enhances virus replication. Virology 374:240–248
Lennemann NJ, Coyne CB (2017) Dengue and Zika viruses subvert reticulophagy by NS2B3-mediated cleavage of FAM134B. Autophagy 13:322–332
Levine B (2005) Eating oneself and uninvited guests: autophagy-related pathways in cellular defense. Cell 120:159–162
Levine B, Kroemer G (2008) Autophagy in the pathogenesis of disease. Cell 132:27–42
Levine B, Mizushima N, Virgin HW (2011) Autophagy in immunity and inflammation. Nature 469:323–335
Li J, Liu Y, Wang Z, Liu K, Wang Y, Liu J, Ding H, Yuan Z (2011) Subversion of cellular autophagy machinery by hepatitis B virus for viral envelopment. J Virol 85:6319–6333
Liang C, Oh BH, Jung JU (2015) Novel functions of viral anti-apoptotic factors. Nat Rev Microbiol 13:7–12
Liang Q, Chang B, Brulois KF, Castro K, Min CK, Rodgers MA, Shi M, Ge J, Feng P, Oh BH et al (2013) Kaposi’s sarcoma-associated herpesvirus K7 modulates Rubicon-mediated inhibition of autophagosome maturation. J Virol 87:12499–12503
Liang Q, Luo Z, Zeng J, Chen W, Foo SS, Lee SA, Ge J, Wang S, Goldman SA, Zlokovic BV et al (2016) Zika virus NS4A and NS4B proteins deregulate Akt-mTOR Signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 19:663–671
Liang XH, Kleeman LK, Jiang HH, Gordon G, Goldman JE, Berry G, Herman B, Levine B (1998) Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J Virol 72:8586–8596
Liu B, Fang M, Hu Y, Huang B, Li N, Chang C, Huang R, Xu X, Yang Z, Chen Z et al (2014) Hepatitis B virus X protein inhibits autophagic degradation by impairing lysosomal maturation. Autophagy 10:416–430
Liu K, Czaja MJ (2013) Regulation of lipid stores and metabolism by lipophagy. Cell Death Differ 20:3–11
Ma J, Sun Q, Mi R, Zhang H (2011) Avian influenza A virus H5N1 causes autophagy-mediated cell death through suppression of mTOR signaling. J Genet Genomics 38:533–537
Ma Y, Galluzzi L, Zitvogel L, Kroemer G (2013) Autophagy and cellular immune responses. Immunity 39:211–227
Maier HJ, Britton P (2012) Involvement of autophagy in coronavirus replication. Viruses 4:3440–3451
Marin M, Rose KM, Kozak SL, Kabat D (2003) HIV-1 Vif protein binds the editing enzyme APOBEC3G and induces its degradation. Nat Med 9:1398–1403
Mateo R, Nagamine CM, Spagnolo J, Mendez E, Rahe M, Gale M Jr, Yuan J, Kirkegaard K (2013) Inhibition of cellular autophagy deranges dengue virion maturation. J Virol 87:1312–1321
McFadden K, Hafez AY, Kishton R, Messinger JE, Nikitin PA, Rathmell JC, Luftig MA (2016) Metabolic stress is a barrier to Epstein-Barr virus-mediated B-cell immortalization. Proc Natl Acad Sci USA 113:E782–E790
Meiffren G, Joubert PE, Gregoire IP, Codogno P, Rabourdin-Combe C, Faure M (2010) Pathogen recognition by the cell surface receptor CD46 induces autophagy. Autophagy 6:299–300
Miller S, Krijnse-Locker J (2008) Modification of intracellular membrane structures for virus replication. Nat Rev Microbiol 6:363–374
Mizushima N, Levine B (2010) Autophagy in mammalian development and differentiation. Nat Cell Biol 12:823–830
Mohamud Y, Shi J, Qu J, Poon T, Xue YC, Deng H, Zhang J, Luo H (2018) Enteroviral infection inhibits autophagic flux via disruption of the SNARE complex to enhance viral replication. Cell Rep 22:3292–3303
Mouna L, Hernandez E, Bonte D, Brost R, Amazit L, Delgui LR, Brune W, Geballe AP, Beau I, Esclatine A (2016) Analysis of the role of autophagy inhibition by two complementary human cytomegalovirus BECN1/Beclin 1-binding proteins. Autophagy 12:327–342
Naniche D, Varior-Krishnan G, Cervoni F, Wild TF, Rossi B, Rabourdin-Combe C, Gerlier D (1993) Human membrane cofactor protein (CD46) acts as a cellular receptor for measles virus. J Virol 67:6025–6032
O’Donnell V, Pacheco JM, LaRocco M, Burrage T, Jackson W, Rodriguez LL, Borca MV, Baxt B (2011) Foot-and-mouth disease virus utilizes an autophagic pathway during viral replication. Virology 410:142–150
Offerdahl DK, Dorward DW, Hansen BT, Bloom ME (2017) Cytoarchitecture of Zika virus infection in human neuroblastoma and Aedes albopictus cell lines. Virology 501:54–62
Oostra M, Hagemeijer MC, van Gent M, Bekker CP, te Lintelo EG, Rottier PJ, de Haan CA (2008) Topology and membrane anchoring of the coronavirus replication complex: not all hydrophobic domains of nsp3 and nsp6 are membrane spanning. J Virol 82:12392–12405
Orvedahl A, Alexander D, Talloczy Z, Sun Q, Wei Y, Zhang W, Burns D, Leib DA, Levine B (2007) HSV-1 ICP34.5 confers neurovirulence by targeting the Beclin 1 autophagy protein. Cell Host Microbe 1:23–35
Orvedahl A, MacPherson S, Sumpter R Jr, Talloczy Z, Zou Z, Levine B (2010) Autophagy protects against Sindbis virus infection of the central nervous system. Cell Host Microbe 7:115–127
Orvedahl A, Sumpter R Jr, Xiao G, Ng A, Zou Z, Tang Y, Narimatsu M, Gilpin C, Sun Q, Roth M et al (2011) Image-based genome-wide siRNA screen identifies selective autophagy factors. Nature 480:113–117
Paludan C, Schmid D, Landthaler M, Vockerodt M, Kube D, Tuschl T, Munz C (2005) Endogenous MHC class II processing of a viral nuclear antigen after autophagy. Science 307:593–596
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, Packer M, Schneider MD, Levine B (2005) Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 122:927–939
Perot BP, Ingersoll MA, Albert ML (2013) The impact of macroautophagy on CD8(+) T-cell-mediated antiviral immunity. Immunol Rev 255:40–56
Petkova DS, Verlhac P, Rozieres A, Baguet J, Claviere M, Kretz-Remy C, Mahieux R, Viret C, Faure M (2017) Distinct contributions of autophagy receptors in measles virus replication. Viruses 9
Petkova DS, Viret C, Faure M (2012) IRGM in autophagy and viral infections. Front Immunol 3:426
Prentice E, Jerome WG, Yoshimori T, Mizushima N, Denison MR (2004) Coronavirus replication complex formation utilizes components of cellular autophagy. J Biol Chem 279:10136–10141
Reggiori F, Monastyrska I, Verheije MH, Cali T, Ulasli M, Bianchi S, Bernasconi R, de Haan CA, Molinari M (2010) Coronaviruses Hijack the LC3-I-positive EDEMosomes, ER-derived vesicles exporting short-lived ERAD regulators, for replication. Cell Host Microbe 7:500–508
Ribeiro CM, Sarrami-Forooshani R, Setiawan LC, Zijlstra-Willems EM, van Hamme JL, Tigchelaar W, van der Wel NN, Kootstra NA, Gringhuis SI, Geijtenbeek TB (2016) Receptor usage dictates HIV-1 restriction by human TRIM5alpha in dendritic cell subsets. Nature 540:448–452
Richards AL, Jackson WT (2012) Intracellular vesicle acidification promotes maturation of infectious poliovirus particles. PLoS Pathog 8:e1003046
Richetta C, Gregoire IP, Verlhac P, Azocar O, Baguet J, Flacher M, Tangy F, Rabourdin-Combe C, Faure M (2013) Sustained autophagy contributes to measles virus infectivity. PLoS Pathog 9:e1003599
Robinson SM, Tsueng G, Sin J, Mangale V, Rahawi S, McIntyre LL, Williams W, Kha N, Cruz C, Hancock BM et al (2014) Coxsackievirus B exits the host cell in shed microvesicles displaying autophagosomal markers. PLoS Pathog 10:e1004045
Romao S, Gannage M, Munz C (2013) Checking the garbage bin for problems in the house, or how autophagy assists in antigen presentation to the immune system. Semin Cancer Biol 23:391–396
Sagnier S, Daussy CF, Borel S, Robert-Hebmann V, Faure M, Blanchet FP, Beaumelle B, Biard-Piechaczyk M, Espert L (2015) Autophagy restricts HIV-1 infection by selectively degrading Tat in CD4+ T lymphocytes. J Virol 89:615–625
Salonen A, Ahola T, Kaariainen L (2005) Viral RNA replication in association with cellular membranes. Curr Top Microbiol Immunol 285:139–173
Samsa MM, Mondotte JA, Iglesias NG, Assuncao-Miranda I, Barbosa-Lima G, Da Poian AT, Bozza PT, Gamarnik AV (2009) Dengue virus capsid protein usurps lipid droplets for viral particle formation. PLoS Pathog 5:e1000632
Santiago C, Celma ML, Stehle T, Casasnovas JM (2010) Structure of the measles virus hemagglutinin bound to the CD46 receptor. Nat Struct Mol Biol 17:124–129
Sharma V, Verma S, Seranova E, Sarkar S, Kumar D (2018) Selective autophagy and xenophagy in infection and disease. Front Cell Dev Biol 6:147
Shi J, Wong J, Piesik P, Fung G, Zhang J, Jagdeo J, Li X, Jan E, Luo H (2013) Cleavage of sequestosome 1/p62 by an enteroviral protease results in disrupted selective autophagy and impaired NFKB signaling. Autophagy 9:1591–1603
Shi X, Chen Z, Tang S, Wu F, Xiong S, Dong C (2016) Coxsackievirus B3 infection induces autophagic flux, and autophagosomes are critical for efficient viral replication. Arch Virol 161:2197–2205
Shrivastava S, Raychoudhuri A, Steele R, Ray R, Ray RB (2011) Knockdown of autophagy enhances the innate immune response in hepatitis C virus-infected hepatocytes. Hepatology 53:406–414
Silva LM, Jung JU (2013) Modulation of the autophagy pathway by human tumor viruses. Semin Cancer Biol 23:323–328
Singh R, Kaushik S, Wang Y, Xiang Y, Novak I, Komatsu M, Tanaka K, Cuervo AM, Czaja MJ (2009) Autophagy regulates lipid metabolism. Nature 458:1131–1135
Sir D, Ann DK, Ou JH (2010) Autophagy by hepatitis B virus and for hepatitis B virus. Autophagy 6:548–549
Sir D, Chen WL, Choi J, Wakita T, Yen TS, Ou JH (2008) Induction of incomplete autophagic response by hepatitis C virus via the unfolded protein response. Hepatology 48:1054–1061
Sir D, Liang C, Chen WL, Jung JU, Ou JH (2008) Perturbation of autophagic pathway by hepatitis C virus. Autophagy 4:830–831
Sir D, Tian Y, Chen WL, Ann DK, Yen TS, Ou JH (2010) The early autophagic pathway is activated by hepatitis B virus and required for viral DNA replication. Proc Natl Acad Sci USA 107:4383–4388
Snijder EJ, van der Meer Y, Zevenhoven-Dobbe J, Onderwater JJ, van der Meulen J, Koerten HK, Mommaas AM (2006) Ultrastructure and origin of membrane vesicles associated with the severe acute respiratory syndrome coronavirus replication complex. J Virol 80:5927–5940
Staring J, von Castelmur E, Blomen VA, van den Hengel LG, Brockmann M, Baggen J, Thibaut HJ, Nieuwenhuis J, Janssen H, van Kuppeveld FJ et al (2017) PLA2G16 represents a switch between entry and clearance of Picornaviridae. Nature 541:412–416
Suhy DA, Giddings TH Jr, Kirkegaard K (2000) Remodeling the endoplasmic reticulum by poliovirus infection and by individual viral proteins: an autophagy-like origin for virus-induced vesicles. J Virol 74:8953–8965
Sumpter R Jr, Levine B (2011) Selective autophagy and viruses. Autophagy 7:260–265
Sumpter R Jr, Sirasanagandla S, Fernandez AF, Wei Y, Dong X, Franco L, Zou Z, Marchal C, Lee MY, Clapp DW et al (2016) Fanconi anemia proteins function in mitophagy and immunity. Cell 165:867–881
Sun MX, Huang L, Wang R, Yu YL, Li C, Li PP, Hu XC, Hao HP, Ishag HA, Mao X (2012) Porcine reproductive and respiratory syndrome virus induces autophagy to promote virus replication. Autophagy 8:1434–1447
Sun P, Zhang S, Qin X, Chang X, Cui X, Li H, Zhang S, Gao H, Wang P, Zhang Z et al (2018) Foot-and-mouth disease virus capsid protein VP2 activates the cellular EIF2S1-ATF4 pathway and induces autophagy via HSPB1. Autophagy 14:336–346
Taguwa S, Kambara H, Fujita N, Noda T, Yoshimori T, Koike K, Moriishi K, Matsuura Y (2011) Dysfunction of autophagy participates in vacuole formation and cell death in cells replicating hepatitis C virus. J Virol 85:13185–13194
Tang H, Da L, Mao Y, Li Y, Li D, Xu Z, Li F, Wang Y, Tiollais P, Li T et al (2009) Hepatitis B virus X protein sensitizes cells to starvation-induced autophagy via up-regulation of beclin 1 expression. Hepatology 49:60–71
Taylor GS, Long HM, Haigh TA, Larsen M, Brooks J, Rickinson AB (2006) A role for intercellular antigen transfer in the recognition of EBV-transformed B cell lines by EBV nuclear antigen-specific CD4+ T cells. J Immunol 177:3746–3756
Taylor MP, Kirkegaard K (2007) Modification of cellular autophagy protein LC3 by poliovirus. J Virol 81:12543–12553
Tian Y, Sir D, Kuo CF, Ann DK, Ou JH (2011) Autophagy required for hepatitis B virus replication in transgenic mice. J Virol 85:13453–13456
Valera MS, de Armas-Rillo L, Barroso-Gonzalez J, Ziglio S, Batisse J, Dubois N, Marrero-Hernandez S, Borel S, Garcia-Exposito L, Biard-Piechaczyk M et al (2015) The HDAC6/APOBEC3G complex regulates HIV-1 infectiveness by inducing Vif autophagic degradation. Retrovirology 12:53
van Gent M, Sparrer KMJ, Gack MU (2018) TRIM proteins and their roles in antiviral host defenses. Annu Rev Virol 5:385–405
Verheije MH, Raaben M, Mari M, Te Lintelo EG, Reggiori F, van Kuppeveld FJ, Rottier PJ, de Haan CA (2008) Mouse hepatitis coronavirus RNA replication depends on GBF1-mediated ARF1 activation. PLoS Pathog 4:e1000088
Wang J, Chen J, Liu Y, Zeng X, Wei M, Wu S, Xiong Q, Song F, Yuan X, Xiao Y et al (2019) Hepatitis B virus induces autophagy to promote its replication by the axis of miR-192-3p-XIAP through NF kappa B signaling. Hepatology 69:974–992
Wang L, Tian Y, Ou JH (2015) HCV induces the expression of Rubicon and UVRAG to temporally regulate the maturation of autophagosomes and viral replication. PLoS Pathog 11:e1004764
Wang R, Zhu Y, Lin X, Ren C, Zhao J, Wang F, Gao X, Xiao R, Zhao L, Chen H et al (2019) Influenza M2 protein regulates MAVS-mediated signaling pathway through interacting with MAVS and increasing ROS production. Autophagy 15:1163–1181
Wen HJ, Yang Z, Zhou Y, Wood C (2010) Enhancement of autophagy during lytic replication by the Kaposi’s sarcoma-associated herpesvirus replication and transcription activator. J Virol 84:7448–7458
Wong J, Zhang J, Si X, Gao G, Mao I, McManus BM, Luo H (2008) Autophagosome supports coxsackievirus B3 replication in host cells. J Virol 82:9143–9153
Yamamoto K, Ichijo H, Korsmeyer SJ (1999) BCL-2 is phosphorylated and inactivated by an ASK1/Jun N-terminal protein kinase pathway normally activated at G(2)/M. Mol Cell Biol 19:8469–8478
Zhang HT, Chen GG, Hu BG, Zhang ZY, Yun JP, He ML, Lai PB (2014) Hepatitis B virus x protein induces autophagy via activating death-associated protein kinase. J Viral Hepat 21:642–649
Zhang J, Lan Y, Li MY, Lamers MM, Fusade-Boyer M, Klemm E, Thiele C, Ashour J, Sanyal S (2018) Flaviviruses exploit the lipid droplet protein AUP1 to trigger lipophagy and drive virus production. Cell Host Microbe 23(819–831):e815
Zhang T, Dong K, Liang W, Xu D, Xia H, Geng J, Najafov A, Liu M, Li Y, Han X et al (2015) G-protein-coupled receptors regulate autophagy by ZBTB16-mediated ubiquitination and proteasomal degradation of Atg14L. Elife 4:e06734
Zhang ZW, Li ZL, Yuan S (2016) The role of secretory autophagy in Zika virus transfer through the placental barrier. Front Cell Infect Microbiol 6:206
Zhao Z, Thackray LB, Miller BC, Lynn TM, Becker MM, Ward E, Mizushima NN, Denison MR, Virgin HW (2007). Coronavirus replication does not require the autophagy gene ATG5. Autophagy 3:581–585
Zhirnov OP, Klenk HD (2013) Influenza A virus proteins NS1 and hemagglutinin along with M2 are involved in stimulation of autophagy in infected cells. J Virol 87:13107–13114
Zhou Y, Geng P, Liu Y, Wu J, Qiao H, Xie Y, Yin N, Chen L, Lin X, Liu Y et al (2018) Rotavirus-encoded virus-like small RNA triggers autophagy by targeting IGF1R via the PI3K/Akt/mTOR pathway. Biochim Biophys Acta Mol Basis Dis 1864:60–68
Zhou Z, Jiang X, Liu D, Fan Z, Hu X, Yan J, Wang M, Gao GF (2009) Autophagy is involved in influenza A virus replication. Autophagy 5:321–328
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mao, J., Lin, E., He, L., Yu, J., Tan, P., Zhou, Y. (2019). Autophagy and Viral Infection. In: Cui, J. (eds) Autophagy Regulation of Innate Immunity. Advances in Experimental Medicine and Biology, vol 1209. Springer, Singapore. https://doi.org/10.1007/978-981-15-0606-2_5
Download citation
DOI: https://doi.org/10.1007/978-981-15-0606-2_5
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-15-0605-5
Online ISBN: 978-981-15-0606-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)