Abstract
Artificial regulation of gene expression through RNA-directed DNA methylation (RdDM)-mediated epigenome editing is one the most important and attractive next-generation technologies for plant trait improvement, often called “new plant breeding techniques” (NPBTs). RdDM can induce transcriptional gene silencing (TGS) of a target gene via modification of the cytosine methylation levels of its promoter region; thus, RdDM is useful as a method for suppression of gene expression without changing the genomic DNA sequence. Likewise, several types of strict epigenetic regulation occur at both the DNA and chromatin levels under normal growth conditions in plants. Recent studies have revealed genome-wide and organ-specific landscapes of epigenetic modifications and their close relationship to plant growth regulation. Therefore, understanding recent findings concerning epigenetic regulation in plants is very important to the future application of epigenome editing in plant breeding. In this chapter, we illustrate several aspects of theoretical and applied epigenetics in plants through discussion of recent studies.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Epigenetics
- Chromatin
- Histone
- Methylation
- New plant breeding techniques
- RNA-directed DNA methylation
- Transcriptional gene silencing
16.1 Mechanisms of Epigenetic Regulation in Plants
16.1.1 DNA Methylation
Cytosine DNA methylation (5mC) is a covalent modification of the fifth carbon residue of cytosine. 5mC is conserved in eukaryotes including mammals and plants, although absent in some organisms such as Drosophila melanogaster and Caenorhabditis elegans (Law and Jacobsen 2010). There are three main strategies to assay DNA methylation levels: (1) digestion of methylated/unmethylated DNA fragments with methylation-sensitive restriction enzymes (MSREs), (2) methylated DNA immunoprecipitation (MeDIP) using an antibody against methylated cytosine, and (3) bisulfite sequencing, in which unmethylated cytosine is converted to uracil by sodium bisulfite, which is reported as thymine in sequence reads. Each strategy can be combined with high-throughput sequencing technology and extended to genome-wide analysis (Cokus et al. 2008; Down et al. 2008; Lister et al. 2008; Maunakea et al. 2010). However, bisulfite sequencing is the most comprehensive and accurate way to quantify DNA methylation levels, so it is widely used for genome-wide analysis (Urich et al. 2015).
In plants, 5mC occurs in three distinct sequence contexts: CG and CHG, which are both symmetric, and CHH (H = C, A, or T), which is asymmetric (Law and Jacobsen 2010). Based on extensive studies in Arabidopsis thaliana, four distinct DNA methylation pathways are known. CG methylation is catalyzed by DNA METHYLTRANSFERASE 1 (MET1) and maintained in a semiconservative manner during DNA replication (Kankel et al. 2003). CHG methylation is catalyzed by CHROMOMETHYLASE3 (CMT3), which recognizes methylation at the 9th lysine residue of the H3 tail (H3K9) (Bartee et al. 2001; Lindroth et al. 2001). CMT2, a homolog of CMT3, catalyzes CHG and CHH methylation in deep heterochromatic regions (Zemach et al. 2013; Stroud et al. 2014). DOMAINS OF REARRANGED METHYLTRANSFERASE 2 (DRM2) catalyzes cytosine methylation in all three sequence contexts, in a process referred to as RNA-directed DNA methylation (RdDM) (Cao and Jacobsen 2002; Law and Jacobsen 2010; Kawashima and Berger 2014; Cuerda-Gil and Slotkin 2016). In RdDM, 21- or 24-nucleotide (nt) small interfering RNAs (siRNAs) guide DRM2 to target regions, marked with DNA methylation and H3K9me, through association with three conserved Argonaute proteins, AGO4, AGO6, and AGO9 (Gao et al. 2010; Havecker et al. 2010; McCue et al. 2015). Non-CG DNA methylation and histone methylation (see Sect. 16.1.2) form a self-reinforcing loop, in which H3K9me controls non-CG DNA methylation and non-CG DNA methylation controls H3K9me (Stroud et al. 2014). Interplays between these pathways have been implicated by comprehensive methylome analysis of components in these pathways (Stroud et al. 2013).
DNA methylation can be actively removed by DNA demethylases. In Arabidopsis, DNA glycosylases DEMETER (DME), REPRESSOR OF SILENCING 1 (ROS1)/DEMETER-LIKE 1 (DML1), DML2, and DML3, are involved in DNA demethylation (Choi et al. 2002; Gong et al. 2002; Penterman et al. 2007; Ortega-Galisteo et al. 2008). In contrast to mutants of DNA methylation components, which are usually viable, dme is embryo lethal, highlighting the importance of active demethylation (Bartee et al. 2001; Cao and Jacobsen 2002; Choi et al. 2002; Kankel et al. 2003). DNA methylation and demethylation activities are balanced by a feedback loop between RdDM and ROS1 (Lei et al. 2015; Williams et al. 2015). ROS1 expression is promoted by RdDM in the ROS1 promoter region and repressed by ROS1 activity itself.
CG methylation is broadly distributed across the genome and often resides within gene bodies (transcribed regions); this type of methylation is called gene body methylation (gbM). The distributions of gbM and histone variant H2A.Z are mutually exclusive, and gbM is associated with higher gene expression (Tran et al. 2005; Zhang et al. 2006; Zilberman et al. 2007). Therefore, gbM is thought to exclude H2A.Z and allow constitutive expression (Zilberman et al. 2008; Coleman-Derr and Zilberman 2012). However, recent studies on intra- and interspecies variations of DNA methylation indicate that gbM does not have a great effect on gene expression or affect H2A.Z distribution within genes (Bewick et al. 2016; Kawakatsu et al. 2016a). Currently, the role of gbM is unclear. Co-localization of CG and non-CG methylation is a characteristic of heterochromatin and transposable elements (TEs) and contributes to gene and TE silencing (Law and Jacobsen 2010). Population-wide methylome variations are largely associated with structural variations such as TE insertion or deletion (Kawakatsu et al. 2016a) and are enriched near signaling pathway genes or immune response genes. TE transposition has shaped the epigenome of Arabidopsis and has introduced variation in environmental responses during diversification.
DNA methylome studies are not limited to Arabidopsis, currently extending to nearly 100 species (Gent et al. 2013; Project 2013; Schmitz et al. 2013a; Stroud et al. 2013; Zhong et al. 2013; Seymour et al. 2014; Ong-Abdullah et al. 2015; Ausin et al. 2016; Niederhuth et al. 2016; Takuno et al. 2016). In addition, transgenerational, populational methylome variations, tissue- and cell-type-specific methylomes, and stress-responsive methylomes have been reported (Hsieh et al. 2009; Schmitz et al. 2011; Calarco et al. 2012; Dowen et al. 2012; Ibarra et al. 2012; Schmitz et al. 2013a, b; Garg et al. 2015; Secco et al. 2015; Hsieh et al. 2016; Kawakatsu et al. 2016a, b; Narsai et al. 2016; Park et al. 2016; Wibowo et al. 2016; Hossain et al. 2017). Recent advances in single-molecule real-time sequencing enable detection of methylated cytosines from long reads without bisulfite conversion (Rand et al. 2017; Simpson et al. 2017). These technologies potentially offer a paradigm shift in DNA methylome analysis, especially in crop species with large genomes and/or multiploidy.
16.1.2 Histone Modification
Histone proteins package genomic DNA into nucleosomes, which in turn form chromatin (Roudier et al. 2009). Histones are conserved in eukaryotes. Four major histones (H2A, H2B, H3, and H4) act as core histones and H1 acts as a linker histone (Kornberg 1974; Thoma and Koller 1977; Luger et al. 1997). The histone core is an octamer complex: two H2A-H2B dimers and an H3-H4 tetramer. Approximately 147 bp of DNA wraps around each histone core and forms a nucleosome. Several histone variants share homology with major histone proteins (Deal and Henikoff 2011a). In addition, histone tails can be posttranslationally modified through methylation (me) and acetylation (ac). These modifications are implicated in flowering, leaf development, seed maturation, flower development, circadian rhythm, and chloroplast development (Deal and Henikoff 2011a; Merini and Calonje 2015; Mozgova et al. 2015). Chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq) is widely used for analyzing the genome-wide distribution of histones, histone variants, and histone modifications (Luo and Lam 2014). Native chromatin digested by micrococcal nuclease (MNase) or cross-linked chromatin fragmented by sonication can be subjected to immunoprecipitation (N-ChIP [native ChIP] or X-ChIP [cross-linked ChIP]) using histone-, histone variant-, or histone modification-specific antibodies (Jackson 1978; O’Neill and Turner 1995; Barski et al. 2007; Schmid and Bucher 2007) (Fig. 16.1).
Comparison of ChIP-seq protocols for genome-wide chromatin mark(s) distribution. Native chromatin is digested by micrococcal nuclease (MNase) in native ChIP (N-ChIP). Cross-linked chromatin is fragmented by sonication in cross-linked ChIP (X-ChIP). Nucleosomes are subjected to immunoprecipitation with specific antibodies
Although histone modifications are conserved in eukaryotes, their distribution patterns and functions vary. For example, mono-methylation of H3 (H3K9me1) is enriched in heterochromatin in Arabidopsis but is enriched at the transcription start sites (TSS) of active genes in animals (Fransz et al. 2006; Fuchs et al. 2006). H3K9 di-methylation (H3K9me2) is also enriched in heterochromatin (Turck et al. 2007). In contrast, H3K9 tri-methylation (H3K9me3) is enriched in euchromatin and is associated with active genes (Turck et al. 2007). H3K9 methylation is catalyzed by SET- and RING-associated (SRA) domain-containing SU(VAR) HOMOLOGUE 1 (SUVH1), SUVH4–6, and SET- and WIYLD-domain-containing SU(VAR)3–9 related 4 (SUVR4) (Ebbs et al. 2005; Ebbs and Bender 2006). INCREASED BONSAI METHYLATION 1 (IBM1) demethylates H3K9 (Inagaki et al. 2010).
H3K27 methylation tends to be associated with low-expression genes and tissue-specific genes (Turck et al. 2007; Roudier et al. 2011). H3K27me1 and H3K27me2 are distributed in both euchromatin and heterochromatin, and H3K27me3 is mainly observed in euchromatin (Roudier et al. 2011). H3K27me1, me2, and me3 mark distinct sets of genes. H3K27me1 and H3K27me3 are enriched in the transcribed regions of marked genes, relative to flanking regions. However, H3K27me2 levels are uniformly higher in both the transcribed and flanking regions of marked genes than in unmarked genes. H3K27me1 is catalyzed by ARABIDOPSIS TRITHORAX-RELATED PROTEIN 5 (ATXR5) and ATXR6 (Jacob et al. 2009). H3K27m3 is catalyzed by Polycomb Repressive Complex 2 (PRC2) (Margueron and Reinberg 2011). Drosophila PRC2 consists of four subunits: (1) enhancer of zeste (E(z)), (2) suppressor of zeste 12 (Su(z)12), (3) nucleosome-remodeling factor 55 kDa subunit (NURF55), and (4) extra sex combs (ESC). Arabidopsis possesses three E(z) homologs (CURLY LEAF [CLF], MEDEA [MEA], and SWINGER [SWN]), three Su(z)12 homologs (EMBRYONIC FLOWER 2 [EMF2], VERNALIZATION 2 [VRN2], and FERTILIZATION-INDEPENDENT SEED 2 [FIS2]), five NURF55 homologs (MULTI-SUBUNIT SUPPRESSOR OF IRA 1–5 [MSI1–5]), and one ESC homolog (FERTILIZATION-INDEPENDENT ENDOSPERM [FIE]) (Ach et al. 1997; Goodrich et al. 1997; Grossniklaus et al. 1998; Kenzior and Folk 1998; Luo et al. 1999; Gendall et al. 2001; Yoshida et al. 2001; Hennig et al. 2003; Chanvivattana et al. 2004; Jullien et al. 2006; Makarevich et al. 2006; Zhang et al. 2007; Jiang et al. 2008; Kim et al. 2010; Lafos et al. 2011; Pazhouhandeh et al. 2011; Derkacheva et al. 2013). PRC2 target genes especially depend on E(z) homologs.
PRC1 is also required for transcriptional repression of H3K27me3-marked genes. PRC1 can catalyze histone H2A mono-ubiquitination (H2Aub) of target genes (de Napoles et al. 2004; Wang et al. 2004). Drosophila PRC1 consists of four subunits (Gil and O’Loghlen 2014): (1) chromodomain protein Polycomb (Pc), (2) RING-finger protein Posterior sex comb (Psc), (3) RING-finger protein Drosophila RING1 (dRING1), and (4) Polyhomeotic (Ph). In Arabidopsis, LIKE HETEROCHROMATIN PROTEIN 1 (LHP1)/TERMINAL FLOWER 2 (TFL2) plays a similar role to Pc (Turck et al. 2007). Drosophila Psc possesses a RING-finger domain, a RING-finger and WD40-associated ubiquitin-like (RAWUL) domain, and a long intrinsically disordered C-terminal region (CTR). The CTR domain of Psc is involved in inhibition of nucleosome remodeling, gene repression, and chromatin compaction; however, it is missing from Arabidopsis Psc homologs AtBMI1A, AtBMI1B, and AtBMI1C (Sanchez-Pulido et al. 2008). EMBRYONIC FLOWER 1 (EMF1) is similar to the CTR domain and acts in a similar manner (Aubert et al. 2001; Calonje et al. 2008). AtRING1A and AtRING1B correspond to dRING1 (Schoorlemmer et al. 1997; Xu and Shen 2008). No plant homolog of Ph has yet been identified.
JUMONJI (JMJ) proteins RELATIVE OF EARLY FLOWERING 6 (REF6)/JMJ12, EARLY FLOWERING 6 (ELF6)/JMJ11, JMJ30, and JMJ32 are H3K27 demethylases (Noh et al. 2004; Lu et al. 2011; Gan et al. 2014). Notably, four C2H2 zinc finger domains of REF6 recognize a CTCTGYTY motif and guide REF6 to its binding sites to modulate H3K27me3 levels (Cui et al. 2016; Li et al. 2016).
A hierarchical model for gene repression, in which PRC2 acts upstream of PRC1, has been widely accepted (Mozgova et al. 2015). In this model, PRC2 methylates H3K27 (to H3K27me3) in target genes as the first step. Second, PRC1 is guided there through H3K27me3 recognition by Pc or LHP1, and target genes are marked with H2Aub, leading to gene repression. However, the hierarchical order of PRC2 and PRC1 is more complicated than once thought (Merini and Calonje 2015). For example, PRC1 is required for H3K27me3 at many PRC2 target genes (Kim et al. 2012; Yang et al. 2013), and PRC2 is not necessarily required for H2Aub at target genes (i.e., PRC1 recruits) (Pengelly et al. 2015). H2Aub can recruit PRC2 and promote H3K27me3 in animals (Blackledge et al. 2014; Cooper et al. 2014; Kalb et al. 2014). In addition, some PRC1 and PRC2 components interact with each other (Xu and Shen 2008; Derkacheva et al. 2013; Wang et al. 2014). Therefore, the interactions between PRC1 and PRC2 may include a positive feedback loop, direct interplay, and mutually independent mechanisms.
H3K36me3 is associated with actively expressed genes (Roudier et al. 2011). H3K36me3 is most prevalent within TSSs but is distributed throughout transcribed regions. SDG8, and possibly SDG4 and SDG25/ATXR7, catalyze H3K36me2 and H3K36me3 (Zhao et al. 2005; Cartagena et al. 2008; Xu et al. 2008; Berr et al. 2009). H3K36me3 and H3K27me3 play antagonistic roles—activation and repression—and rarely co-exist on the same histone tail (Roudier et al. 2011; Yang et al. 2014).
H3K4 methylation is mostly found in genes and other euchromatin (Roudier et al. 2011). In contrast to enhancer-associated H3K4me1 in animals, H3K4me1 in Arabidopsis is distributed inside transcribed regions but is less prevalent near TSSs and transcription end sites (TESs). Both H3K4me2 and H3K4me3 are enriched around TSSs but depleted around TESs. H3K4me3 is associated with highly expressed genes, whereas H3K4me1 and H3K4me2 are associated with tissue-specific genes (Roudier et al. 2011). ARABIDOPSIS TRITHORAX 1 (ATX1)/SET DOMAIN GROUP 27 (SDG27) and Complex Proteins Associated with Set1 (COMPASS)-like complex catalyze H3K4me3, and ATX2/SDG30 catalyzes H3K4me2 (Saleh et al. 2008; Jiang et al. 2011). SDG2 catalyzes H3K4me1, H3K4me2, and H3K4me3 in vitro, but in vivo, sdg2 shows reduction only in H3K4me3 (Berr et al. 2010; Guo et al. 2010). JMJ14 and homologs of LYSINE-SPECIFIC HISTONE DEMETHYLASE 1 (LSD1), FLOWERING LOCUS D (FLD)/LSD1-like 1 (LDL1), and LDL2 are required for H3K4 demethylation (Deleris et al. 2010; Lu et al. 2010; Greenberg et al. 2013).
H3K9ac and H3K27ac are associated with active gene expression (Charron et al. 2009). Levels of both H3K9ac and H3K27ac peak near TSSs and are distributed inside gene bodies. The H3K9ac and H3K27ac target regions are largely the same but distinct from H3K27me3 target regions. H3K9ac and H3K27ac are catalyzed by histone acetyltransferase (HAT) family proteins, such as homologs of general control non-derepressible (GCN5) and TATA binding protein-associated factor 1 (TAF1) (Pandey et al. 2002; Benhamed et al. 2006). AtGCN5 also catalyzes H4K14ac. Histone deacetylase (HDAC) family proteins, such as HDA6, are responsible for histone deacetylation (Pandey et al. 2002; Earley et al. 2006; To et al. 2011; Liu et al. 2014). HATs and HDACs act as transcriptional co-activators and co-repressors, respectively.
Combinations of histone modifications are thought to be important for the precise expression state and responsiveness of a gene. Two opposing histone marks, for example, H3K27me3 (repressing) and H3K4me3 (activating), can be co-localized in the same genomic regions (Roudier et al. 2011). As in animals, bivalent chromatin regions in plants are associated with several transcription factors (TFs) that are normally expressed at low levels but are induced at specific timing and/or in specific tissues by developmental cues (Saleh et al. 2007; Jiang et al. 2008; Berr et al. 2010; Roudier et al. 2011). It is also possible that a mixture of different cell types with different chromatin modification states could be misinterpreted as co-localization of opposing histone marks. Cell-type-specific profiling would promote further understanding not only of cell-type-specific properties but also of the combinatorial functions of histone modifications. Several recent developments show considerable promise in this area. For example, low-input ChIP-seq methods and high-throughput sequencing technologies are evolving (Adli and Bernstein 2011; Brind’Amour et al. 2015; Schmidl et al. 2015). Recently developed simple but highly efficient INTACT (Isolation of Nuclei Tagged in specific Cell Types) is feasible for cell-type-specific profiling (Deal and Henikoff 2011b).
16.1.3 Chromatin Accessibility
Transcriptional activation is primarily regulated by TF binding to regulatory DNA elements, where chromatin is open or accessible. Genome-wide chromatin accessibility can be assayed directly or indirectly through a combination of nuclease digestion and high-throughput sequencing (Meyer and Liu 2014). As in N-ChIP, MNase digests bare DNA that is not protected by nucleosomes, whereas DNase I cleaves unprotected DNA. Therefore, MNase digestion followed by sequencing (MNase-seq) identifies nucleosome positioning and indirectly detects open chromatin regions (Schones et al. 2008), whereas DNase I cleavage followed by sequencing (DNase-seq) directly detects open chromatin regions (Boyle et al. 2008). FAIRE-seq (Formaldehyde-Assisted Isolation of Regulatory Elements) also directly detects open chromatin regions by isolating un-cross-linked DNA with nucleosomes (Giresi et al. 2007). Transposase Accessible Chromatin sequencing (ATAC-seq) uses Tn5 transposase to insert sequencing-ready adaptor sequences into open chromatin regions, starting with as few as 500 cells (Buenrostro et al. 2013). Genome-wide chromatin accessibility studies have been limited in plants (Zhang et al. 2012a, 2012b; Li et al. 2014; Wu et al. 2014; Zhang et al. 2015; Lu et al. 2016). Nevertheless, these studies clearly demonstrate that the identified open chromatin regions are associated with gene expression and TF binding sites. ChIP-seq has been used for assays of genome-wide TF binding sites in vivo (Song et al. 2016). However, preparing antibodies against a wide variety of TFs or transgenic plants expressing tagged TFs with native promoters is time-, cost-, and labor-consuming. DNA affinity purification sequencing (DAP-seq) is a new technology to cost-effectively identify TF binding sites in vitro (O’Malley et al. 2016). The combination of chromatin accessibility assays and DAP-seq is expected to greatly advance our knowledge of transcriptional regulatory networks.
16.2 Application Studies on Epigenetics in Plants
16.2.1 Application of Epigenome Editing to Plant Breeding
Gene manipulation (GM) techniques have been used as molecular breeding tools to develop various GM crops with excellent traits such as resistance to insect pests, plant diseases, and specific herbicides. Furthermore, “golden rice,” which accumulates provitamin A in the seed, will be practical to use in the Philippines in the near future. GM crops are at present commercially cultivated in 28 countries worldwide. The total global cultivated area of transgenic crops was estimated to have reached approximately 179.7 million hectares by 2015 (James 2016). On the other hand, conventional GM techniques are still sometimes viewed as a serious issue in numerous countries because the transgene is integrated into the genome of the target plant, and a number of people are concerned about gene flow from GM crops to the environment.
In recent years, technologies referred to as “new plant breeding techniques” (NPBTs) have been proposed as a solution for issues surrounding conventional GM crops. Genome editing (ZFN, TALEN, CRISPR/Cas9), “grafting with GM plants,” “reverse breeding,” Agrobacterium infiltration, and RdDM can all be classified as NPBTs (Lusser et al. 2012; Schaart et al. 2016). When these NPBTs are applied, it is difficult to distinguish between the newly introduced artificial mutation and natural mutations. In particular, changes to genomic DNA caused by RdDM cannot be detected by conventional molecular analysis methods such as PCR and DNA sequencing because there is no change to the DNA sequence. A key characteristic of RdDM-mediated transcriptional gene silencing (TGS) is the production of double-strand RNA (dsRNA) with homology to the promoter sequence of the target gene. The dsRNA is cleaved into 21–24 nt pieces of siRNA by DICER like (DCL) protein. In plants, these siRNAs become a cause of epigenetic modification of cytosine residues in CG, CHG, and CHH contexts into methylated cytosine (5mC). Two types of DNA-dependent RNA polymerases (Pol IV and Pol V) are necessary to advance the process of RdDM (Matzke et al. 2015). The increased methylation levels in the target gene promoter induce TGS, which is associated with changes in chromatin structure through histone modification (see Sect. 16.1.2).
Although posttranscriptional gene silencing (PTGS) has been used for functional analyses of target genes for about 20 years, TGS has an important advantage over PTGS. Artificially induced methylation and TGS via RdDM may be preserved and inherited after removal of the trigger gene cassette; thus RdDM-mediated TGS could be considered as a type of NPBT. However, there are few research papers that discuss the relationship between the preservation or loss of TGS and the presence or absence of the trigger gene (Kasai and Kanazawa 2013).
The remainder of this chapter describes the application of epigenetic modification to improvement of plant traits through various strategies of RdDM-mediated TGS. CRISPR/Cas9-mediated epigenome editing is also briefly discussed.
16.2.2 Viral Vector-Mediated TGS in Plants
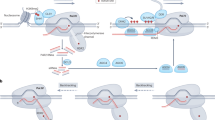
Plant viral vectors have been used to induce RdDM-mediated TGS (Fig. 16.2a). In this strategy, the plant defense response toward virus infection (called recovery) is applied to the production of dsRNA from target gene promoters. Viral vector-mediated TGS in plants has been successfully induced in both reporter genes and endogenous genes. Kanazawa and co-authors used recombinant Cucumber mosaic virus (CMV) to induce TGS of Chalcone synthase-A and LeSPL-CNR genes in petunia and tomato, respectively (Kanazawa et al. 2011). Both plants showed clear phenotypic changes in association with epigenetic modification of the target gene promoters (Kanazawa et al. 2011). Interestingly, these new traits have been observed in subsequent generations even though the viral vector was not detected in these progenies (Kanazawa et al. 2011). These authors further reported that 2b protein, one of the endogenous proteins derived from CMV, is useful for stable induction of RdDM because it increases the expression of TGS induction-related genes and decreases the expression of demethylation-related genes. The 2b protein functions as an RNA silencing suppressor that can inhibit PTGS and virus-induced gene silencing (VIGS) (Goto et al. 2007). These results suggest that the combination of dsRNA and the 2b protein leads to highly efficient induction of RdDM-mediated TGS in a viral vector system.
As another example, TGS induction of Chalcone synthase-A gene in petunia was achieved via the use of apple latent spherical virus (ALSV) as a viral vector (Kon and Yoshikawa 2014). Tobacco rattle virus (TRV) vectors have also been used for induction of TGS, but most of the target genes have been reporter genes such as GFP and GUS under the control of the CaMV 35S promoter (Jones et al. 2001). However, recently, Bond and Baulcombe reported that TGS induction of endogenous gene (FLOWERING WAGENINGEN, FWA) using TRV viral vector system in Arabidopsis and this report deeply discussed about initiation, establishment, and maintenance of TGS in endogenous gene, using FWA TGS Arabidopsis and various mutants of gene silencing-related genes (Bond and Baulcombe 2015).
16.2.3 Agrobacterium (T-DNA)-Mediated TGS in Plants
RdDM-mediated TGS in plants is generally induced by using T-DNA harboring gene cassettes to express dsRNA directed toward the promoter region of the target gene (Fig. 16.2b). Gene cassettes to express the dsRNA are introduced into the plant genome via Agrobacterium-mediated transformation. T0 plants expressing dsRNA derived from a foreign gene cassette should be treated as GM plants, whereas progenies of TGS plants after removal of T-DNA by segregation can be treated as non-GM plants in some world areas.
Although T-DNA-mediated TGS is very simple and easy to use, most research papers describe TGS of a reporter gene under the control of the CaMV 35S promoter. There are only a few reports of T-DNA-mediated TGS of endogenous genes in plants. In rice, RdDM-mediated TGS was attempted using a reporter gene (GFP under the control of the CaMV 35S promoter) and several endogenous genes. In these experiments, TGS could be easily induced for GFP; TGS of most endogenous genes could not be induced in spite of highly efficient induction of cytosine methylation of the target gene promoters (Okano et al. 2008). Although different levels of chromatin modification were observed between the CaMV 35S promoter and the endogenous gene promoter, it is not yet understood why reporter gene constructs such as CaMV 35S promoter::GFP can be silenced by TGS more easily than endogenous genes. On the other hand, successful induction of TGS of an endogenous gene by T-DNA-mediated expression of dsRNA corresponding to the target promoter region has been reported in petunia (Sijen et al. 2001), Arabidopsis (Deng et al. 2014), and potato (Kasai et al. 2016; Heilersig et al. 2006). However, it seems that a reproducible method for stable induction of TGS via T-DNA has not yet been found in plants.
As an alternative Agrobacterium-mediated strategy, T-DNA harboring a viral vector sequence is used to deliver the viral vector to plant cells. After Agrobacterium-mediated transformation or Agrobacterium infiltration, the viral vector is transferred into the nuclei of plant cells as a part of the T-DNA and can function independently as a viral vector. Viral vectors released from the T-DNA induce RdDM-mediated TGS (Ju et al. 2016).
16.2.4 Grafting-Mediated TGS in Plants
Grafting is a plant-specific strategy for inducing TGS. In vascular plants, the vascular bundle system functions to transport water, minerals, nutrients, proteins, and photosynthate from sink to source organs (or from source to sink). Some RNA molecules such as siRNA and microRNA (miRNA) are also transported by vascular bundle system. Specifically, these small RNA molecules are exclusively transported from sink to source organs through the phloem (Melnyk et al. 2011; Ham and Lucas 2017) and can move from cell to cell through the plasmodesmata (Melnyk et al. 2011; Ham and Lucas 2017). When a scion artificially expressing siRNA toward a target gene promoter (GM plant) is grafted onto a wild-type rootstock (non-GM), RdDM-mediated TGS can be induced in the rootstock via siRNA movement through the vascular bundle system and plasmodesmata. If a regenerated plantlet is obtained from the TGS rootstock, it would be transcriptionally silenced without insertion of foreign DNA into the genome. For this reason, “grafting on GM” is considered a type of NPBT (Fig. 16.2c). Bai and co-authors grafted transgenic tobacco scions expressing dsRNA directed toward parts of the CaMV 35S promoter region and under the control of the companion cell-specific Commelina yellow mottle virus (CoYMV) promoter onto transgenic tobacco rootstocks expressing GFP under the control of the CaMV 35S promoter. GFP fluorescence was drastically suppressed in lateral roots of the rootstock, indicating that TGS was epigenetically induced in the rootstock (Bai et al. 2011). This study showed that the movement of siRNA from scion to rootstock was more efficient than from rootstock to scion (Bai et al. 2011). These same authors have also produced an epigenetically improved potato by grafting with transgenic tobacco as the TGS inducer, resulting in modified amylose content through suppression of granule-bound starch synthase I (GBSSI) gene without changes in the genomic DNA sequences of the host potato (Kasai et al. 2016).
Vegetatively propagated crops such as potato and apple may have an advantage over seed-propagated crops with respect to the use of RdDM-mediated TGS because vegetatively propagated crops do not require meiosis for self-reproduction; thus, the modified methylation level may be preserved more stably than in seed-propagated crops. However, further investigation would be necessary to clarify this point.
16.2.5 CRISPR/Cas9-Mediated Epigenome Modification
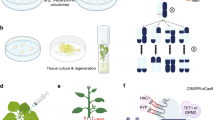
The CRISPR/Cas9 system is a convenient and powerful tool for genome editing in many organisms (Cong et al. 2013; Mali et al. 2013; Fauser et al. 2014). This system is very simple, consisting of the combination of a guide RNA and Cas9 nuclease (Fig. 16.3a). A modified CRISPR/Cas9 system can be applied to epigenome editing. A nuclease-activity-disrupted Cas9 (dCas9) fused with enzymes to modify genomic DNA or histone can be used as an epigenome editing inducer (Fig. 16.3b). For example, a fusion protein consisting of nuclease-disrupted Cas9 protein and human acetyltransferase p300 successfully catalyzed the acetylation of histone H3 at target sites in human cells, resulting in the robust transcriptional activation of target genes (Hilton et al. 2015). Further modifications of CRISPR/Cas9-mediated epigenome editing will continue to be developed in animals and plants (Johnson et al. 2014; Konermann et al. 2013).
CRISPR/Cas9-mediated epigenome editing. (a), Normal scheme of CRISPR/Cas9 system to induce double-strand break toward target genomic DNA. (b) Epigenome editing technology using fusion protein consists of effector and catalytically dead Cas9 (dCas9) lacking only endonuclease activity to induce the other modification such as demethylation or acetylation toward target genomic DNA or chromatin
16.3 Future Perspectives
This chapter describes new findings from both basic and applied studies on epigenetics in plants. Recently, the study of epigenetics has developed rapidly because of an increase in the precision of genome-wide association studies (GWAS), which have received a lot of attention in both animals and plants. At present, application of RdDM-mediated epigenome editing to plant breeding is not yet practical owing to the need to obtain stable induction of TGS toward endogenous genes and stable inheritance of the modified epigenome after removal of the trigger gene. However, we expect that many interesting findings will continue to be reported in the epigenetics field, with the result that RdDM-mediated epigenome editing will become a promising technology to produce trait-improved plants in the near future.
References
Ach RA, Taranto P, Gruissem W (1997) A conserved family of WD-40 proteins binds to the retinoblastoma protein in both plants and animals. Plant Cell 9:1595–1606
Adli M, Bernstein BE (2011) Whole-genome chromatin profiling from limited numbers of cells using nano-ChIP-seq. Nat Protoc 6:1656–1668
Aubert D, Chen L, Moon YH, Martin D, Castle LA, Yang CH, Sung ZR (2001) EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 13:1865–1875
Ausin I, Feng S, Yu C, Liu W, Kuo HY, Jacobsen EL, Zhai J, Gallego-Bartolome J, Wang L, Egertsdotter U, Street NR, Jacobsen SE, Wang H (2016) DNA methylome of the 20-gigabase Norway spruce genome. Proc Natl Acad Sci U S A 113:E8106–E8113
Bai S, Kasai A, Yamada K, Li T, Harada T (2011) A mobile signal transported over a long distance induces systemic transcriptional gene silencing in a grafted partner. J Exp Bot 62:4561–4570
Barski A, Cuddapah S, Cui K, Roh TY, Schones DE, Wang Z, Wei G, Chepelev I, Zhao K (2007) High-resolution profiling of histone methylations in the human genome. Cell 129:823–837
Bartee L, Malagnac F, Bender J (2001) Arabidopsis cmt3 chromomethylase mutations block non-CG methylation and silencing of an endogenous gene. Genes Dev 15:1753–1758
Benhamed M, Bertrand C, Servet C, Zhou DX (2006) Arabidopsis GCN5, HD1, and TAF1/HAF2 interact to regulate histone acetylation required for light-responsive gene expression. Plant Cell 18:2893–2903
Berr A, Xu L, Gao J, Cognat V, Steinmetz A, Dong A, Shen WH (2009) SET DOMAIN GROUP25 encodes a histone methyltransferase and is involved in FLOWERING LOCUS C activation and repression of flowering. Plant Physiol 151:1476–1485
Berr A, McCallum EJ, Ménard R, Meyer D, Fuchs J, Dong A, Shen WH (2010) Arabidopsis SET DOMAIN GROUP2 is required for H3K4 trimethylation and is crucial for both sporophyte and gametophyte development. Plant Cell 22:3232–3248
Bewick AJ, Ji L, Niederhuth CE, Willing EM, Hofmeister BT, Shi X, Wang L, Lu Z, Rohr NA, Hartwig B, Kiefer C, Deal RB, Schmutz J, Grimwood J, Stroud H, Jacobsen SE, Schneeberger K, Zhang X, Schmitz RJ (2016) On the origin and evolutionary consequences of gene body DNA methylation. Proc Natl Acad Sci U S A 113:9111–9116
Blackledge NP, Farcas AM, Kondo T, King HW, McGouran JF, Hanssen LL, Ito S, Cooper S, Kondo K, Koseki Y, Ishikura T, Long HK, Sheahan TW, Brockdorff N, Kessler BM, Koseki H, Klose RJ (2014) Variant PRC1 complex-dependent H2A ubiquitylation drives PRC2 recruitment and polycomb domain formation. Cell 157:1445–1459
Bond DM, Baulcombe DC (2015) Epigenetic transitions leading to heritable, RNA-mediated de novo silencing in Arabidopsis thaliana. Proc Natl Acad Sci U S A 112:917–922
Boyle AP, Davis S, Shulha HP, Meltzer P, Margulies EH, Weng Z, Furey TS, Crawford GE (2008) High-resolution mapping and characterization of open chromatin across the genome. Cell 132:311–322
Brind’Amour J, Liu S, Hudson M, Chen C, Karimi MM, Lorincz MC (2015) An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat Commun 6:6033
Buenrostro JD, Giresi PG, Zaba LC, Chang HY, Greenleaf WJ (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10:1213–1218
Calarco JP, Borges F, Donoghue MT, Van Ex F, Jullien PE, Lopes T, Gardner R, Berger F, Feijo JA, Becker JD, Martienssen RA (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151:194–205
Calonje M, Sanchez R, Chen L, Sung ZR (2008) EMBRYONIC FLOWER1 participates in polycomb group-mediated AG gene silencing in Arabidopsis. Plant Cell 20:277–291
Cao X, Jacobsen SE (2002) Role of the arabidopsis DRM methyltransferases in de novo DNA methylation and gene silencing. Curr Biol 12:1138–1144
Cartagena JA, Matsunaga S, Seki M, Kurihara D, Yokoyama M, Shinozaki K, Fujimoto S, Azumi Y, Uchiyama S, Fukui K (2008) The Arabidopsis SDG4 contributes to the regulation of pollen tube growth by methylation of histone H3 lysines 4 and 36 in mature pollen. Dev Biol 315:355–368
Chanvivattana Y, Bishopp A, Schubert D, Stock C, Moon YH, Sung ZR, Goodrich J (2004) Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 131:5263–5276
Charron JB, He H, Elling AA, Deng XW (2009) Dynamic landscapes of four histone modifications during deetiolation in Arabidopsis. Plant Cell 21:3732–3748
Choi Y, Gehring M, Johnson L, Hannon M, Harada JJ, Goldberg RB, Jacobsen SE, Fischer RL (2002) DEMETER, a DNA glycosylase domain protein, is required for endosperm gene imprinting and seed viability in Arabidopsis. Cell 110:33–42
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452:215–219
Coleman-Derr D, Zilberman D (2012) Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PLoS Genet 8:e1002988
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Cooper S, Dienstbier M, Hassan R, Schermelleh L, Sharif J, Blackledge NP, De Marco V, Elderkin S, Koseki H, Klose R, Heger A, Brockdorff N (2014) Targeting polycomb to pericentric heterochromatin in embryonic stem cells reveals a role for H2AK119u1 in PRC2 recruitment. Cell Rep 7:1456–1470
Cuerda-Gil D, Slotkin RK (2016) Non-canonical RNA-directed DNA methylation. Nat Plants 2:16163
Cui X, Lu F, Qiu Q, Zhou B, Gu L, Zhang S, Kang Y, Ma X, Yao Q, Ma J, Zhang X, Cao X (2016) REF6 recognizes a specific DNA sequence to demethylate H3K27me3 and regulate organ boundary formation in Arabidopsis. Nat Genet 48:694–699
de Napoles M, Mermoud JE, Wakao R, Tang YA, Endoh M, Appanah R, Nesterova TB, Silva J, Otte AP, Vidal M, Koseki H, Brockdorff N (2004) Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev Cell 7:663–676
Deal RB, Henikoff S (2011a) Histone variants and modifications in plant gene regulation. Curr Opin Plant Biol 14:116–122
Deal RB, Henikoff S (2011b) The INTACT method for cell type-specific gene expression and chromatin profiling in Arabidopsis thaliana. Nat Protoc 6:56–68
Deleris A, Greenberg MV, Ausin I, Law RW, Moissiard G, Schubert D, Jacobsen SE (2010) Involvement of a Jumonji-C domain-containing histone demethylase in DRM2-mediated maintenance of DNA methylation. EMBO Rep 11:950–955
Deng S, Dai H, Arenas C, Wang H, Niu HW, Chua NH (2014) Transcriptional silencing of Arabidopsis endogens by single-stranded RNAs targeting the promoter region. Plant Cell Physiol 55:823–833
Derkacheva M, Steinbach Y, Wildhaber T, Mozgová I, Mahrez W, Nanni P, Bischof S, Gruissem W, Hennig L (2013) Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J 32:2073–2085
Dowen RH, Pelizzola M, Schmitz RJ, Lister R, Dowen JM, Nery JR, Dixon JE, Ecker JR (2012) Widespread dynamic DNA methylation in response to biotic stress. Proc Natl Acad Sci U S A 109:E2183–E2191
Down TA, Rakyan VK, Turner DJ, Flicek P, Li H, Kulesha E, Gräf S, Johnson N, Herrero J, Tomazou EM, Thorne NP, Bäckdahl L, Herberth M, Howe KL, Jackson DK, Miretti MM, Marioni JC, Birney E, Hubbard TJ, Durbin R, Tavaré S, Beck S (2008) A Bayesian deconvolution strategy for immunoprecipitation-based DNA methylome analysis. Nat Biotechnol 26:779–785
Earley K, Lawrence RJ, Pontes O, Reuther R, Enciso AJ, Silva M, Neves N, Gross M, Viegas W, Pikaard CS (2006) Erasure of histone acetylation by Arabidopsis HDA6 mediates large-scale gene silencing in nucleolar dominance. Genes Dev 20:1283–1293
Ebbs ML, Bender J (2006) Locus-specific control of DNA methylation by the Arabidopsis SUVH5 histone methyltransferase. Plant Cell 18:1166–1176
Ebbs ML, Bartee L, Bender J (2005) H3 lysine 9 methylation is maintained on a transcribed inverted repeat by combined action of SUVH6 and SUVH4 methyltransferases. Mol Cell Biol 25:10507–10515
Fauser F, Schiml S, Puchta H (2014) Both CRISPR/Cas-based nucleases and nickases can be efficiently for genome engineering in Arabidopsis thaliana. Plant J 79:348–359
Fransz P, ten Hoopen R, Tessadori F (2006) Composition and formation of heterochromatin in Arabidopsis thaliana. Chromosom Res 14:71–82
Fuchs J, Demidov D, Houben A, Schubert I (2006) Chromosomal histone modification patterns – from conservation to diversity. Trends Plant Sci 11:199–208
Gan ES, Xu Y, Wong JY, Goh JG, Sun B, Wee WY, Huang J, Ito T (2014) Jumonji demethylases moderate precocious flowering at elevated temperature via regulation of FLC in Arabidopsis. Nat Commun 5:5098
Gao Z, Liu H, Daxinger L, Pontes O, He X, Qian W, Lin H, Xie M, Lorkovic Z, Zhang S, Miki D, Zhan X, Pontier D, Lagrange T, Jin H, Matzke A, Matzke M, Pikaard C, Zhu J (2010) An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation. Nature 465:106–109
Garg R, Narayana Chevala V, Shankar R, Jain M (2015) Divergent DNA methylation patterns associated with gene expression in rice cultivars with contrasting drought and salinity stress response. Sci Rep 5:14922
Gendall AR, Levy YY, Wilson A, Dean C (2001) The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 107:525–535
Gent JI, Ellis NA, Guo L, Harkess AE, Yao Y, Zhang X, Dawe RK (2013) CHH islands: de novo DNA methylation in near-gene chromatin regulation in maize. Genome Res 23:628–637
Gil J, O’Loghlen A (2014) PRC1 complex diversity: where is it taking us? Trends Cell Biol 24:632–641
Giresi PG, Kim J, McDaniell RM, Iyer VR, Lieb JD (2007) FAIRE (Formaldehyde-Assisted Isolation of Regulatory Elements) isolates active regulatory elements from human chromatin. Genome Res 17:877–885
Gong Z, Morales-Ruiz T, Ariza RR, Roldan-Arjona T, David L, Zhu JK (2002) ROS1, a repressor of transcriptional gene silencing in Arabidopsis, encodes a DNA glycosylase/lyase. Cell 111:803–814
Goodrich J, Puangsomlee P, Martin M, Long D, Meyerowitz EM, Coupland G (1997) A Polycomb-group gene regulates homeotic gene expression in Arabidopsis. Nature 386:44–51
Goto K, Kobori T, Kosaka Y, Natsuki T, Masuta C (2007) Characterization of silencing suppressor 2b of cucumber mosaic virus based of examination of its small RNA-bindng abilities. Plant Cell Physiol 48:1050–1060
Greenberg MV, Deleris A, Hale CJ, Liu A, Feng S, Jacobsen SE (2013) Interplay between active chromatin marks and RNA-directed DNA methylation in Arabidopsis thaliana. PLoS Genet 9:e1003946
Grossniklaus U, Vielle-Calzada JP, Hoeppner MA, Gagliano WB (1998) Maternal control of embryogenesis by MEDEA, a polycomb group gene in Arabidopsis. Science 280:446–450
Guo L, Yu Y, Law JA, Zhang X (2010) SET DOMAIN GROUP2 is the major histone H3 lysine [corrected] 4 trimethyltransferase in Arabidopsis. Proc Natl Acad Sci U S A 107:18557–18562
Ham BK, Lucas WJ (2017) Phloem-mobile RNAs as systemic signaling agents. Annu Rev Plant Biol 68:173–195
Havecker ER, Wallbridge LM, Hardcastle TJ, Bush MS, Kelly KA, Dunn RM, Schwach F, Doonan JH, Baulcombe DC (2010) The Arabidopsis RNA-directed DNA methylation argonautes functionally diverge based on their expression and interaction with target loci. Plant Cell 22:321–334
Heilersig BHJB, Loonen AEHM, Janssen EM, Wolters AA, Visser RGF (2006) Efficiency of transcriptional gene silencing of GBSSI in potato depends on the promoter region that is used in an inverted repeat. Mol Gen Genomics 275:437–449
Hennig L, Taranto P, Walser M, Schönrock N, Gruissem W (2003) Arabidopsis MSI1 is required for epigenetic maintenance of reproductive development. Development 130:2555–2565
Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, Gersbach CA (2015) Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol 33:510–517
Hossain MS, Kawakatsu T, Kim KD, Zhang N, Nguyen CT, Khan SM, Batek JM, Joshi T, Schmutz J, Grimwood J, Schmitz RJ, Xu D, Jackson SA, Ecker JR, Stacey G (2017) Divergent cytosine DNA methylation patterns in single-cell, soybean root hairs. New Phytol 214:808–819
Hsieh TF, Ibarra CA, Silva P, Zemach A, Eshed-Williams L, Fischer RL, Zilberman D (2009) Genome-wide demethylation of Arabidopsis endosperm. Science 324:1451–1454
Hsieh PH, He S, Buttress T, Gao H, Couchman M, Fischer RL, Zilberman D, Feng X (2016) Arabidopsis male sexual lineage exhibits more robust maintenance of CG methylation than somatic tissues. Proc Natl Acad Sci U S A 113:15132–15137
Ibarra CA, Feng X, Schoft VK, Hsieh TF, Uzawa R, Rodrigues JA, Zemach A, Chumak N, Machlicova A, Nishimura T, Rojas D, Fischer RL, Tamaru H, Zilberman D (2012) Active DNA demethylation in plant companion cells reinforces transposon methylation in gametes. Science 337:1360–1364
Inagaki S, Miura-Kamio A, Nakamura Y, Lu F, Cui X, Cao X, Kimura H, Saze H, Kakutani T (2010) Autocatalytic differentiation of epigenetic modifications within the Arabidopsis genome. EMBO J 29:3496–3506
Jackson V (1978) Studies on histone organization in the nucleosome using formaldehyde as a reversible cross-linking agent. Cell 15:945–954
Jacob Y, Feng S, LeBlanc CA, Bernatavichute YV, Stroud H, Cokus S, Johnson LM, Pellegrini M, Jacobsen SE, Michaels SD (2009) ATXR5 and ATXR6 are H3K27 monomethyltransferases required for chromatin structure and gene silencing. Nat Struct Mol Biol 16:763–768
James C (2016) 20th anniversary of the global commercialization of biotech crops (1996 to 2015) and biotech crop highlights in 2015. International Service for the Acquisition of Agri-biotech Applications (ISAAA) http://www.isaaa.org/resources/publications/briefs/51/executivesummary/pdf/B51-ExecSum-English.pdf
Jiang D, Wang Y, He Y (2008) Repression of FLOWERING LOCUS C and FLOWERING LOCUS T by the Arabidopsis Polycomb repressive complex 2 components. PLoS One 3:e3404
Jiang D, Kong NC, Gu X, Li Z, He Y (2011) Arabidopsis COMPASS-like complexes mediate histone H3 lysine-4 trimethylation to control floral transition and plant development. PLoS Genet 7:e1001330
Johnson LM, Du J, Hale CJ, Bischof S, Feng S, Chodavarapu RK, Zhong X, Marson G, Pellegrini M, Segal DJ, Patel DJ, Jacobsen SE (2014) SRA- and SET-domain-containing proteins like RNA polymerase V occupancy to DNA methylation. Nature 507:124–128
Jones L, Ratcliff F, Baulcombe DC (2001) RNA-directed transcriptional gene silencing in plants can be inherited independenly of the RNA trigger and requires Met1 for maintenance. Curr Biol 11:747–757
Ju Z, Wang L, Cao D, Zuo J, Zhu H, Fu LY, Zhu B (2016) A viral satellite DNA vector-induced transcriptional gene silencing via DNA methylation of gene promoter in Nicotiana benthamiana. Virus Res 223:99–107
Jullien PE, Katz A, Oliva M, Ohad N, Berger F (2006) Polycomb group complexes self-regulate imprinting of the Polycomb group gene MEDEA in Arabidopsis. Curr Biol 16:486–492
Kalb R, Latwiel S, Baymaz HI, Jansen PW, Müller CW, Vermeulen M, Müller J (2014) Histone H2A monoubiquitination promotes histone H3 methylation in Polycomb repression. Nat Struct Mol Biol 21:569–571
Kanazawa A, Inaba J, Shimura H, Otagaki S, Tsukahara S, Matsuzawa A, Kim BM, Goto K, Masuta C (2011) Virus-mediated efficient induction of epigenetic modifications of endogenous genes with phenotypic changes in plants. Plant J 65:156–168
Kankel MW, Ramsey DE, Stokes TL, Flowers SK, Haag JR, Jeddeloh JA, Riddle NC, Verbsky ML, Richards EJ (2003) Arabidopsis MET1 cytosine methyltransferase mutants. Genetics 163:1109–1122
Kasai M, Kanazawa A (2013) Induction of RNA-directed DNA methylation and heritable transcriptional gene silencing as a tool to engineer novel traits in plants. Plant Biotechnol 30:233–241
Kasai A, Bai S, Hojo H, Harada T (2016) Epigenome editing of potato by grafting using transgenic tobacco as siRNA donor. PLoS One 11:e0161729
Kawakatsu T, Huang SS, Jupe F, Sasaki E, Schmitz RJ, Urich MA, Castanon R, Nery JR, Barragan C, He Y, Chen H, Dubin M, Lee CR, Wang C, Bemm F, Becker C, O’Neil R, O’Malley RC, Quarless DX, Genomes C, Schork NJ, Weigel D, Nordborg M, Ecker JR (2016a) Epigenomic diversity in a global collection of Arabidopsis thaliana accessions. Cell 166:492–505
Kawakatsu T, Stuart T, Valdes M, Breakfield N, Schmitz RJ, Nery JR, Urich MA, Han X, Lister R, Benfey PN, Ecker JR (2016b) Unique cell-type-specific patterns of DNA methylation in the root meristem. Nat Plants 2:16058
Kawashima T, Berger F (2014) Epigenetic reprogramming in plant sexual reproduction. Nat Rev Genet 15:613–624
Kenzior AL, Folk WR (1998) AtMSI4 and RbAp48 WD-40 repeat proteins bind metal ions. FEBS Lett 440:425–429
Kim SY, Zhu T, Sung ZR (2010) Epigenetic regulation of gene programs by EMF1 and EMF2 in Arabidopsis. Plant Physiol 152:516–528
Kim SY, Lee J, Eshed-Williams L, Zilberman D, Sung ZR (2012) EMF1 and PRC2 cooperate to repress key regulators of Arabidopsis development. PLoS Genet 8:e1002512
Kon T, Yoshikawa N (2014) Induction and maintenance of DNA methylation in promoter sequences by apple latent spherical virus-induced transcriptional gene silencing. Front Microbiol 5:595
Konermann S, Brigham MD, Trevino A, Hsu PD, Matthias H, Cong L, Platt RJ, Scott DA, Church GM, Zhang F (2013) Optical control of mammalian endogenous transcription and epigenetic states. Nature 500:472–476
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868–871
Lafos M, Kroll P, Hohenstatt ML, Thorpe FL, Clarenz O, Schubert D (2011) Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet 7:e1002040
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220
Lei M, Zhang H, Julian R, Tang K, Xie S, Zhu JK (2015) Regulatory link between DNA methylation and active demethylation in Arabidopsis. Proc Natl Acad Sci U S A 112:3553–3557
Li G, Liu S, Wang J, He J, Huang H, Zhang Y, Xu L (2014) ISWI proteins participate in the genome-wide nucleosome distribution in Arabidopsis. Plant J 78:706–714
Li C, Gu L, Gao L, Chen C, Wei CQ, Qiu Q, Chien CW, Wang S, Jiang L, Ai LF, Chen CY, Yang S, Nguyen V, Qi Y, Snyder MP, Burlingame AL, Kohalmi SE, Huang S, Cao X, Wang ZY, Wu K, Chen X, Cui Y (2016) Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat Genet 48:687–693
Lindroth AM, Cao X, Jackson JP, Zilberman D, CM MC, Henikoff S, Jacobsen SE (2001) Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science 292:2077–2080
Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR (2008) Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133:523–536
Liu X, Yang S, Zhao M, Luo M, Yu CW, Chen CY, Tai R, Wu K (2014) Transcriptional repression by histone deacetylases in plants. Mol Plant 7:764–772
Lu F, Cui X, Zhang S, Liu C, Cao X (2010) JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res 20:387–390
Lu F, Cui X, Zhang S, Jenuwein T, Cao X (2011) Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat Genet 43:715–719
Lu Z, Hofmeister BT, Vollmers C, DuBois RM, Schmitz RJ (2016) Combining ATAC-seq with nuclei sorting for discovery of cis-regulatory regions in plant genomes. Nucleic Acids Res 45:e41
Luger K, Rechsteiner TJ, Flaus AJ, Waye MM, Richmond TJ (1997) Characterization of nucleosome core particles containing histone proteins made in bacteria. J Mol Biol 272:301–311
Luo C, Lam E (2014) Quantitatively profiling genome-wide patterns of histone modifications in Arabidopsis thaliana using ChIP-seq. Methods Mol Biol 1112:177–193
Luo M, Bilodeau P, Koltunow A, Dennis ES, Peacock WJ, Chaudhury AM (1999) Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc Natl Acad Sci U S A 96:296–301
Lusser M, Parisi C, Plan D, Rodriguez-Cerezo E (2012) Deployment of new biotechnologies in plant breeding. Nat Biotechnol 30:231–239
Makarevich G, Leroy O, Akinci U, Schubert D, Clarenz O, Goodrich J, Grossniklaus U, Köhler C (2006) Different Polycomb group complexes regulate common target genes in Arabidopsis. EMBO Rep 7:947–952
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Margueron R, Reinberg D (2011) The Polycomb complex PRC2 and its mark in life. Nature 469:343–349
Matzke MA, Kanno T, Matzke JM (2015) RNA-directed DNA methylation: the evolution of a complex epigenetic pathway in flowering plants. Annu Rev Plant Biol 66:243–267
Maunakea AK, Nagarajan RP, Bilenky M, Ballinger TJ, D’Souza C, Fouse SD, Johnson BE, Hong C, Nielsen C, Zhao Y, Turecki G, Delaney A, Varhol R, Thiessen N, Shchors K, Heine VM, Rowitch DH, Xing X, Fiore C, Schillebeeckx M, Jones SJ, Haussler D, Marra MA, Hirst M, Wang T, Costello JF (2010) Conserved role of intragenic DNA methylation in regulating alternative promoters. Nature 466:253–257
McCue AD, Panda K, Nuthikattu S, Choudury SG, Thomas EN, Slotkin RK (2015) ARGONAUTE 6 bridges transposable element mRNA-derived siRNAs to the establishment of DNA methylation. EMBO J 34:20–35
Melnyk CW, Molnar A, Baulconbe DC (2011) Intercelluar and systemic novement of RNA silencing signals. EMBO J 30:3553–3563
Merini W, Calonje M (2015) PRC1 is taking the lead in PcG repression. Plant J 83:110–120
Meyer CA, Liu XS (2014) Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat Rev Genet 15:709–721
Mozgova I, Köhler C, Hennig L (2015) Keeping the gate closed: functions of the polycomb repressive complex PRC2 in development. Plant J 83:121–132
Narsai R, Secco D, Schultz MD, Ecker JR, Lister R, Whelan J (2016) Dynamic and rapid changes in the transcriptome and epigenome during germination and in developing rice (Oryza sativa) coleoptiles under anoxia and re-oxygenation. Plant J 89:805–824
Niederhuth CE, Bewick AJ, Ji L, Alabady MS, Kim KD, Li Q, Rohr NA, Rambani A, Burke JM, Udall JA, Egesi C, Schmutz J, Grimwood J, Jackson SA, Springer NM, Schmitz RJ (2016) Widespread natural variation of DNA methylation within angiosperms. Genome Biol 17:194
Noh B, Lee SH, Kim HJ, Yi G, Shin EA, Lee M, Jung KJ, Doyle MR, Amasino RM, Noh YS (2004) Divergent roles of a pair of homologous jumonji/zinc-finger-class transcription factor proteins in the regulation of Arabidopsis flowering time. Plant Cell 16:2601–2613
O’Malley RC, Huang SC, Song L, Lewsey MG, Bartlett A, Nery JR, Galli M, Gallavotti A, Ecker JR (2016) Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165:1280–1292
O’Neill LP, Turner BM (1995) Histone H4 acetylation distinguishes coding regions of the human genome from heterochromatin in a differentiation-dependent but transcription-independent manner. EMBO J 14:3946–3957
Okano Y, Miki D, Shimamoto K (2008) Small interfering RNA (siRNA) targeting of endogenous promoters induces methylation but not necessarily gene silencing, in rice. Plant J 53:65–77
Ong-Abdullah M, Ordway JM, Jiang N, Ooi SE, Kok SY, Sarpan N, Azimi N, Hashim AT, Ishak Z, Rosli SK, Malike FA, Bakar NA, Marjuni M, Abdullah N, Yaakub Z, Amiruddin MD, Nookiah R, Singh R, Low ET, Chan KL, Azizi N, Smith SW, Bacher B, Budiman MA, Van Brunt A, Wischmeyer C, Beil M, Hogan M, Lakey N, Lim CC, Arulandoo X, Wong CK, Choo CN, Wong WC, Kwan YY, Alwee SS, Sambanthamurthi R, Martienssen RA (2015) Loss of karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525:533–537
Ortega-Galisteo AP, Morales-Ruiz T, Ariza RR, Roldan-Arjona T (2008) Arabidopsis DEMETER-LIKE proteins DML2 and DML3 are required for appropriate distribution of DNA methylation marks. Plant Mol Biol 67:671–681
Pandey R, Müller A, Napoli CA, Selinger DA, Pikaard CS, Richards EJ, Bender J, Mount DW, Jorgensen RA (2002) Analysis of histone acetyltransferase and histone deacetylase families of Arabidopsis thaliana suggests functional diversification of chromatin modification among multicellular eukaryotes. Nucleic Acids Res 30:5036–5055
Park K, Kim MY, Vickers M, Park JS, Hyun Y, Okamoto T, Zilberman D, Fischer RL, Feng X, Choi Y, Scholten S (2016) DNA demethylation is initiated in the central cells of Arabidopsis and rice. Proc Natl Acad Sci U S A 113:15138–15143
Pazhouhandeh M, Molinier J, Berr A, Genschik P (2011) MSI4/FVE interacts with CUL4-DDB1 and a PRC2-like complex to control epigenetic regulation of flowering time in Arabidopsis. Proc Natl Acad Sci U S A 108:3430–3435
Pengelly AR, Kalb R, Finkl K, Müller J (2015) Transcriptional repression by PRC1 in the absence of H2A monoubiquitylation. Genes Dev 29:1487–1492
Penterman J, Zilberman D, Huh JH, Ballinger T, Henikoff S, Fischer RL (2007) DNA demethylation in the Arabidopsis genome. Proc Natl Acad Sci U S A 104:6752–6757
Project AG (2013) The Amborella genome and the evolution of flowering plants. Science 342:1241089
Rand AC, Jain M, Eizenga JM, Musselman-Brown A, Olsen HE, Akeson M, Paten B (2017) Mapping DNA methylation with high-throughput nanopore sequencing. Nat Methods 14:411–413
Roudier F, Teixeira FK, Colot V (2009) Chromatin indexing in Arabidopsis: an epigenomic tale of tails and more. Trends Genet 25:511–517
Roudier F, Ahmed I, Bérard C, Sarazin A, Mary-Huard T, Cortijo S, Bouyer D, Caillieux E, Duvernois-Berthet E, Al-Shikhley L, Giraut L, Després B, Drevensek S, Barneche F, Dèrozier S, Brunaud V, Aubourg S, Schnittger A, Bowler C, Martin-Magniette ML, Robin S, Caboche M, Colot V (2011) Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J 30:1928–1938
Saleh A, Al-Abdallat A, Ndamukong I, Alvarez-Venegas R, Avramova Z (2007) The Arabidopsis homologs of trithorax (ATX1) and enhancer of zeste (CLF) establish ‘bivalent chromatin marks’ at the silent AGAMOUS locus. Nucleic Acids Res 35:6290–6296
Saleh A, Alvarez-Venegas R, Yilmaz M, Le O, Hou G, Sadder M, Al-Abdallat A, Xia Y, Lu G, Ladunga I, Avramova Z (2008) The highly similar Arabidopsis homologs of trithorax ATX1 and ATX2 encode proteins with divergent biochemical functions. Plant Cell 20:568–579
Sanchez-Pulido L, Devos D, Sung ZR, Calonje M (2008) RAWUL: a new ubiquitin-like domain in PRC1 ring finger proteins that unveils putative plant and worm PRC1 orthologs. BMC Genomics 9:308
Schaart JG, Wiel CCM, Lotz LAP, Smulders JM (2016) Opportunities for products of new plant breeding techniques. Trends Plant Sci 21:438–449
Schmid CD, Bucher P (2007) ChIP-Seq data reveal nucleosome architecture of human promoters. Cell 131:831–832. author reply 832–833
Schmidl C, Rendeiro AF, Sheffield NC, Bock C (2015) ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat Methods 12:963–965
Schmitz RJ, Schultz MD, Lewsey MG, O’Malley RC, Urich MA, Libiger O, Schork NJ, Ecker JR (2011) Transgenerational epigenetic instability is a source of novel methylation variants. Science 334:369–373
Schmitz RJ, He Y, Valdes-Lopez O, Khan SM, Joshi T, Urich MA, Nery JR, Diers B, Xu D, Stacey G, Ecker JR (2013a) Epigenome-wide inheritance of cytosine methylation variants in a recombinant inbred population. Genome Res 23:1663–1674
Schmitz RJ, Schultz MD, Urich MA, Nery JR, Pelizzola M, Libiger O, Alix A, McCosh RB, Chen H, Schork NJ, Ecker JR (2013b) Patterns of population epigenomic diversity. Nature 495:193–198
Schones DE, Cui K, Cuddapah S, Roh TY, Barski A, Wang Z, Wei G, Zhao K (2008) Dynamic regulation of nucleosome positioning in the human genome. Cell 132:887–898
Schoorlemmer J, Marcos-Gutiérrez C, Were F, Martínez R, García E, Satijn DP, Otte AP, Vidal M (1997) Ring1A is a transcriptional repressor that interacts with the Polycomb-M33 protein and is expressed at rhombomere boundaries in the mouse hindbrain. EMBO J 16:5930–5942
Secco D, Wang C, Shou H, Schultz MD, Chiarenza S, Nussaume L, Ecker JR, Whelan J, Lister R (2015) Stress induced gene expression drives transient DNA methylation changes at adjacent repetitive elements. elife 4:e09343
Seymour DK, Koenig D, Hagmann J, Becker C, Weigel D (2014) Evolution of DNA methylation patterns in the Brassicaceae is driven by differences in genome organization. PLoS Genet 10:e1004785
Sijen T, Vijn I, Rebocho A, van Blokland R, Roelofs D, Mol JN, Kooter JM (2001) Transcriptional and posttranscriptional gene silencing are mechanistically related. Curr Biol 11:436–440
Simpson JT, Workman RE, Zuzarte PC, David M, Dursi LJ, Timp W (2017) Detecting DNA cytosine methylation using nanopore sequencing. Nat Methods 14:407–410
Song L, Huang SC, Wise A, Castanon R, Nery JR, Chen H, Watanabe M, Thomas J, Bar-Joseph Z, Ecker JR (2016) A transcription factor hierarchy defines an environmental stress response network. Science 354:aag1550
Stroud H, Ding B, Simon SA, Feng S, Bellizzi M, Pellegrini M, Wang GL, Meyers BC, Jacobsen SE (2013) Plants regenerated from tissue culture contain stable epigenome changes in rice. elife 2:e00354
Stroud H, Do T, Du J, Zhong X, Feng S, Johnson L, Patel DJ, Jacobsen SE (2014) Non-CG methylation patterns shape the epigenetic landscape in Arabidopsis. Nat Struct Mol Biol 21:64–72
Takuno S, Ran J-H, Gaut BS (2016) Evolutionary patterns of genic DNA methylation vary across land plants. Nat Plants 2:15222
Thoma F, Koller T (1977) Influence of histone H1 on chromatin structure. Cell 12:101–107
To TK, Kim JM, Matsui A, Kurihara Y, Morosawa T, Ishida J, Tanaka M, Endo T, Kakutani T, Toyoda T, Kimura H, Yokoyama S, Shinozaki K, Seki M (2011) Arabidopsis HDA6 regulates locus-directed heterochromatin silencing in cooperation with MET1. PLoS Genet 7:e1002055
Tran RK, Henikoff JG, Zilberman D, Ditt RF, Jacobsen SE, Henikoff S (2005) DNA methylation profiling identifies CG methylation clusters in Arabidopsis genes. Curr Biol 15:154–159
Turck F, Roudier F, Farrona S, Martin-Magniette ML, Guillaume E, Buisine N, Gagnot S, Martienssen RA, Coupland G, Colot V (2007) Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet 3:e86
Urich MA, Nery JR, Lister R, Schmitz RJ, Ecker JR (2015) MethylC-seq library preparation for base-resolution whole-genome bisulfite sequencing. Nat Protoc 10:475–483
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, Zhang Y (2004) Role of histone H2A ubiquitination in Polycomb silencing. Nature 431:873–878
Wang Y, Gu X, Yuan W, Schmitz RJ, He Y (2014) Photoperiodic control of the floral transition through a distinct polycomb repressive complex. Dev Cell 28:727–736
Wibowo A, Becker C, Marconi G, Durr J, Price J, Hagmann J, Papareddy R, Putra H, Kageyama J, Becker J, Weigel D, Gutierrez-Marcos J (2016) Hyperosmotic stress memory in Arabidopsis is mediated by distinct epigenetically labile sites in the genome and is restricted in the male germline by DNA glycosylase activity. elife 5:e13456
Williams BP, Pignatta D, Henikoff S, Gehring M (2015) Methylation-sensitive expression of a DNA demethylase gene serves as an epigenetic rheostat. PLoS Genet 11:e1005142
Wu Y, Zhang W, Jiang J (2014) Genome-wide nucleosome positioning is orchestrated by genomic regions associated with DNase I hypersensitivity in rice. PLoS Genet 10:e1004378
Xu L, Shen WH (2008) Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr Biol 18:1966–1971
Xu L, Zhao Z, Dong A, Soubigou-Taconnat L, Renou JP, Steinmetz A, Shen WH (2008) Di- and tri- but not monomethylation on histone H3 lysine 36 marks active transcription of genes involved in flowering time regulation and other processes in Arabidopsis thaliana. Mol Cell Biol 28:1348–1360
Yang C, Bratzel F, Hohmann N, Koch M, Turck F, Calonje M (2013) VAL- and AtBMI1-mediated H2Aub initiate the switch from embryonic to postgerminative growth in Arabidopsis. Curr Biol 23:1324–1329
Yang H, Howard M, Dean C (2014) Antagonistic roles for H3K36me3 and H3K27me3 in the cold-induced epigenetic switch at Arabidopsis FLC. Curr Biol 24:1793–1797
Yoshida N, Yanai Y, Chen L, Kato Y, Hiratsuka J, Miwa T, Sung ZR, Takahashi S (2001) EMBRYONIC FLOWER2, a novel polycomb group protein homolog, mediates shoot development and flowering in Arabidopsis. Plant Cell 13:2471–2481
Zemach A, Kim MY, Hsieh PH, Coleman-Derr D, Eshed-Williams L, Thao K, Harmer SL, Zilberman D (2013) The Arabidopsis nucleosome remodeler DDM1 allows DNA methyltransferases to access H1-containing heterochromatin. Cell 153:193–205
Zhang X, Yazaki J, Sundaresan A, Cokus S, Chan SW, Chen H, Henderson IR, Shinn P, Pellegrini M, Jacobsen SE, Ecker JR (2006) Genome-wide high-resolution mapping and functional analysis of DNA methylation in arabidopsis. Cell 126:1189–1201
Zhang X, Clarenz O, Cokus S, Bernatavichute YV, Pellegrini M, Goodrich J, Jacobsen SE (2007) Whole-genome analysis of histone H3 lysine 27 trimethylation in Arabidopsis. PLoS Biol 5:e129
Zhang W, Wu Y, Schnable JC, Zeng Z, Freeling M, Crawford GE, Jiang J (2012a) High-resolution mapping of open chromatin in the rice genome. Genome Res 22:151–162
Zhang W, Zhang T, Wu Y, Jiang J (2012b) Genome-wide identification of regulatory DNA elements and protein-binding footprints using signatures of open chromatin in Arabidopsis. Plant Cell 24:2719–2731
Zhang T, Zhang W, Jiang J (2015) Genome-wide nucleosome occupancy and positioning and their impact on gene expression and evolution in plants. Plant Physiol 168:1406–1416
Zhao Z, Yu Y, Meyer D, Wu C, Shen WH (2005) Prevention of early flowering by expression of FLOWERING LOCUS C requires methylation of histone H3 K36. Nat Cell Biol 7:1256–1260
Zhong S, Fei Z, Chen YR, Zheng Y, Huang M, Vrebalov J, McQuinn R, Gapper N, Liu B, Xiang J, Shao Y, Giovannoni JJ (2013) Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat Biotechnol 31:154–159
Zilberman D, Gehring M, Tran RK, Ballinger T, Henikoff S (2007) Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat Genet 39:61–69
Zilberman D, Coleman-Derr D, Ballinger T, Henikoff S (2008) Histone H2A.Z and DNA methylation are mutually antagonistic chromatin marks. Nature 456:125–129
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Wakasa, Y., Kawakatsu, T., Takaiwa, F. (2018). Theoretical and Applied Epigenetics in Plants. In: Masuda, S., Izawa, S. (eds) Applied RNA Bioscience. Springer, Singapore. https://doi.org/10.1007/978-981-10-8372-3_16
Download citation
DOI: https://doi.org/10.1007/978-981-10-8372-3_16
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-8371-6
Online ISBN: 978-981-10-8372-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)