Abstract
The discovery of genome-wide transcription through high-throughput sequencing in the mammals has revealed that only ~2 % of the genome is expressed into protein-coding mRNAs and the rest ~98 % makes different types of intergenic, intronic and repeat sequence-rich, small and long regulatory noncoding RNAs with multitude of biological functions. The complexity of mammalian brain has been largely attributed to diverse region-specific transcriptomes, a major portion of which has been recently found to consist of innumerable forms of long (>200 nt.) noncoding RNAs (lncRNAs), implicated in various functions such as brain development, cell-lineage specification, learning and memory. However, their relative association with processes involved in aging and age-related disorders has not been sufficiently explored. Here, we have characterized a repeat sequence containing long intergenic noncoding RNA (lincRNA), LINC-RBE (rat brain expressed) from the rat genome, which is differentially expressed in the brain during maturation and aging. Through expression analysis, LINC-RBE was shown to express in specific cell types and neuroanatomical compartments, e.g., cortex, hippocampus and cerebellum of the rat brain in an age-dependent manner. Thus, our study showed the possible interrelationship between lincRNAs and various brain functions during aging, which may provide an alternative basis to study various age-related neurological diseases and disorders.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Multicellular diploid organisms develop either from single diploid cells by asexual methods such as vegetative growth of totipotent cells or from haploid cells by sexual method by a process of fertilization of a female gonadal cell, the mature oocyte by a male gonadal cell, the sperm. The latter mechanism has been selected for mammals during evolution. In this process, the oocyte or the mother cell contributes the essential genomic information along with the mitochondria, while the sperm or the father cell contributes the variations in the genomic information to the next generation and thereby to the gene pool of the population. Along with this, two more major genetic contributions are also made during the process of fertilization. The first is by the transposable elements (TEs) and the second is by the noncoding RNAs present in the haploid cells. These two components of the genome are now known to contribute to both genetic and epigenetic aspects of the zygotic cell, which determines certain aspects of the subsequent steps of the embryonic development. Following embryonic development, the multicellular organism undergoes functional specialization and maturation to adulthood functions. Reproduction is one of the major functions of adulthood. Soon after attainment of adulthood or sexual maturity to reproduce itself, the organism slowly starts experiencing the aging process. Aging sets in slowly but surely. Aging is manifested in all types of cells, organs and organisms, which finally expresses into progressive and cumulative decline in all physiological functions of the cells and tissues. Prominent manifestations among them are the following: decrease in production of reproductive cells ultimately leading to cessation of reproductive ability; weakening of the bones and muscles; wrinkling of the skin; greying of the hairs; decreased physical ability for alertness, movement and capacity to work; decreased mental ability often leading to loss of memory and cognition; lowered immune system; lowered metabolism in almost all the organs; decreased level of confidence and overall loss of physiological balance and coordination among various organ systems of the body. Thus old age makes a person susceptible to many diseases. Often old patients feel less cared for medical treatments. The geriatric medicine is usually either less developed or less cared for in many countries. With all these difficulties, we still notice many old persons keep themselves alive with joyful personality, effective interaction with others and substantial human contribution to their family, immediate neighbourhood and the society at large. Until they remain useful to others, they get attention for their own life. However, many other elderly, due to poor conditions of their health, often become less fortunate in the later part of life. Generally, old age is prolonged for a considerable period of years, thereby adding problems due to lack of quality of life, healthcare issues and family as well as societal responsibilities towards the elderly. Thus a biological natural process of aging of a living organism ultimately gives rise to a multitude of challenges for scientists, clinicians, policy makers and organizations as well as the government. Many individuals and organizations are actively thinking to convert these challenges in the field of aging into opportunities for research on aging, drug-development for elderly and old age-associated diseases, making use of traditional medicine system for healthy aging, innovations in pharmaceutical industry, advancements in clinical practices in geriatric medicine and patient care in hospitals as well as winning the good will and blessings of the elderly populations of the world for the policy makers, organizations, government and society. As biologists, we need to understand the basic mechanism(s) of the aging process in mammals at cellular, biochemical and molecular levels. The present research article is an example of a piece of basic or fundamental research on one of the aspects of the aging process in the rat brain as an experimental model system.
In the mid-20th century, with the discovery of DNA as the genetic material (Watson and Crick 1953), one of the greatest challenges of biology had emerged as how to explain the decoding of information in the form of nucleotide sequence in DNA, a double helical nucleic acid macromolecule, to the stretch of amino acid sequences in proteins, presumed to perform a diversity of cellular functions in all organisms. This was answered following comprehensive experiments and collective insights about the role of a special class of ribonucleic acid molecule, called as ‘messenger RNA’ (mRNA). It is synthesized from DNA (transcription) and carries information to ribosomes for protein synthesis (translation). This proposed the concept of “central dogma of molecular biology” (DNA makes RNA makes protein) (Amiel 1965; Geiduschek and Haselkorn 1969). Subsequently, it was realized that a rather small fraction of mammalian genome was assigned for all proteins, the rest of the DNA especially the repetitive (~40 %), intergenic (~40 %) and intronic (~15 %) DNA was not represented in proteins. They were inadvertently called as “Junk DNA” with no function. With the sequencing of mammalian genomes during the last two decades (Ananda et al. 2014; Gibbs et al. 2004; Venter et al. 2001; Waterston et al. 2002) together with the advancements in the high-throughput genomic screening techniques such as microarrays for global gene expression analysis and next generation sequencing (NGS) (Capobianco 2014; Kawaji et al. 2014) and data acquired from ENCODE (Encyclopedia of DNA elements) (Birney et al. 2007; Siggens and Ekwall 2014) and FANTOM (Functional annotation of mammalian cDNAs) consortia (Carninci et al. 2005; Liang et al. 2015; Lizio et al. 2015), a whole new revelation was made. The ~98 % of the non-protein coding DNA or “dark matter” of the genome is now known to pervasively transcribe into multifunctional forms of non-protein coding RNA molecules, called as noncoding RNAs (ncRNAs), and only ~1.5–1.8 % consists of protein-coding sequences (Denas et al. 2015; Guttman et al. 2009; Iyer et al. 2015; Quek et al. 2015). Thus, instead of having only protein-coding genes with multiple transcription start sites, alternative promoter and enhancer elements, and variable 5ʹ and 3ʹ-upstream regions, maximum portion of mammalian genome coding for myriad of complex ncRNAs of intergenic and intronic origins define the complexity and flexibility of the genome with respect to its evolution (Liang et al. 2015; Marques and Ponting 2014). Until now, many of these ncRNAs have been functionally associated with almost all biological processes from development, lineage specification, differentiation (Ballarino et al. 2015; Cajigas et al. 2015; Gong et al. 2015; Shahryari et al. 2015), cellular reprogramming (Flynn and Chang 2014; Kim et al. 2015; Loewer et al. 2010) to cell proliferation, growth and cell death. Repetitive sequences, besides being prevalently associated with heterochromatin and telomeric organization (Arning et al. 2015; Chow et al. 2010; Lee 2011; Redon et al. 2013; Saksouk et al. 2014), X-chromosome inactivation (dosage compensation), translational regulation, neocentromere formation and generation of small regulatory RNAs (Faulkner et al. 2009; Hoffman et al. 2014; Kapusta et al. 2013), have also emerged as an important source of noncoding RNAs in the mammalian genome (Singh and Rath 2012), however, still their functional relevance in the genome is largely unknown (Faulkner et al. 2009). The revised “central dogma of molecular biology” now includes the ncRNAs as regulatory molecules (Fig. 1) (Wahlestedt 2013).
Schematic representation of the revised “central dogma of molecular biology” depicting the flow of genetic information in eukaryotes and various roles of lncRNAs along with proteins in regulating the steps. The generalized view of the flow of genetic information includes: DNA is the source of all genetic information and from DNA the information is decoded in the form of messenger RNA (mRNA) and then to protein through the ribosomal machinery. The earlier assumption was that proteins perform all functions in a cell for sustenance of life. This concept has been revised with the discovery of various regulatory noncoding RNAs including microRNAs (miRNAs) acting on the mRNAs and long noncoding RNAs (lncRNAs). LncRNAs together with various proteins are involved in maintenance and epigenetic modification of chromatin, transcriptional and post-transcriptional gene regulation, translational initiation, protein metabolism and transport and many others. This gives a whole new concept of exploring the genome and cellular functions in a new dimension (Mattick and Rinn 2015)
Noncoding RNAs, based on their expression and function, are divided into two major categories: (a) housekeeping or structural ncRNAs, that include well-characterized infrastructural RNAs such as ribosomal RNAs, transfer RNAs, and small nuclear/spliceosomal RNAs; and (b) regulatory ncRNAs, which comprise of various types of small noncoding RNAs (sncRNAs) such as microRNAs or miRNAs (~19–22 nt.), PIWI-interacting RNAs or piRNAs (~30–32 nt.), small nucleolar RNAs or snoRNAs, promoter-associated small RNAs or PASRs (Hu et al. 2012) and transcription initiation RNAs or tiRNAs (Zaramela et al. 2014), among others, and a wide range of thousands of long noncoding RNAs (lncRNAs). SncRNAs comprise of substantially characterized and highly conserved fraction of transcripts, primarily implicated in functions such as post-transcriptional modulation of mRNA stability and degradation (Olive et al. 2015; Philippe et al. 2015), dosage compensation of X chromosome (Song et al. 2009) and chromatin organization (Chen et al. 2014). Besides this, recently many of them have been studied for their involvement in aging and age-related diseases in C. elegans (lin-14, mir-71, mi-238 and mi-246) (Boehm and Slack 2005; de Lencastre et al. 2010) and animal (mouse, human) models (miR-21, miR-130a and miR-494) (Serna et al. 2012) (Table 1). LncRNAs, on the other hand, are ≥200 to several thousand nt. in size (e.g., XACT, ~250 kb; Xist, ~17 kb) (Mattick and Rinn 2015; Ponting et al. 2009; St Laurent et al. 2015). Also, some recently discovered class of heterogeneous, polyadenylated and 5ʹ-methyl-capped, regulatory RNAs that are spatio-temporally expressed in sense, antisense, overlapping and bi-directional manners from the intronic, intergenic, repeat sequence-rich regions of mammalian genome (Guttman et al. 2009; Porro et al. 2014; Kurokawa 2011; Milligan and Lipovich 2014; Mortimer et al. 2014; Pnueli et al. 2015; St Laurent et al. 2015; Vucicevic et al. 2015). Although most of the lncRNAs are pervasively transcribed in the genome, they show poor sequence conservation and are highly unstable or present in few copies per cell. However, the dynamics, developmental and tissue/cell type-specific expression patterns of lncRNAs are regulated by transcriptional factors (e.g., Oct-4, Nanog, p53 etc.) and epigenetic modifications of histones and DNA methylation, this together with conservation of their promoter sequences strongly propose for their biological function and evolutionary significance (Ponting et al. 2009; Wang and Chang 2011). Their intrinsic tendency to form various secondary structures for functional aspects, is a prerequisite for RNA-protein/RNA-chromatin interactions, this provides them with sensory, guiding, scaffolding and allosteric capacities (Flintoft 2013; Gonzalez-Buendia et al. 2015; Mortimer et al. 2014; Somarowthu et al. 2015). Their biological functions include sex-dependent dosage compensation, gene imprinting (McHugh et al. 2015; Mercer and Mattick 2013; Rinn 2014), chromatin modification and heterochromatinization (Bao et al. 2015; Khalil et al. 2009; Marchese and Huarte 2014; Mercer and Mattick 2013; Wang et al. 2015), epigenetic regulation of gene expression, post-transcriptional modulation of gene expression, via regulation of alternative splicing, RNA editing, RNA transport (Singh 2012), maintenance of nuclear structure, translational regulation by modulating RNA stability and degradation (Legnini et al. 2014) (Huarte et al. 2010), nuclear-cytoplasmic protein trafficking (Hu et al. 2014; Willingham et al. 2005) and many others (Fig. 1). The altered expression of lncRNAs, therefore, has been linked to the progression and prognosis of various cardiovascular and neurological diseases and disorders along with development of various types of cancers (Akula et al. 2014; Sigdel et al. 2015; Xue et al. 2015; Yang et al. 2014). Recently, many lncRNAs have been found to regulate the onset of cellular senescence or various genes/gene networks/processes directly associated with the progression of aging and age-related disorders (Table 2), however, direct implication of repeat sequence containing lncRNA in the aging process has not yet been fully explored. We have recently reviewed the possible link between various lncRNAs, associated with chromatin modulation, telomeric maintenance, p53-mediated cell cycle regulation, with the onset and pathophysiology of aging and age-related neurological, cardiovascular and immunological diseases/disorders (Kour and Rath 2016c).
In mammals, the higher order sensory-regulatory, cognitive and behavioral functions are performed by the complex, dynamic and intricate networks of neurons and glial cells in the brain or central nervous system (CNS). Brain is structurally and functionally heterogeneous in nature but with high levels of coordination among its different functional regions. This has been attributed to various cell types and developmental stage-specific gene expression patterns and their epigenetic regulatory mechanisms such as chromatin remodelling via differential histone modifications and DNA-methylation marks that drive, control and coordinate the gene expression patterns (Graff and Mansuy 2008; Weichenhan and Plass 2013). In a recent study, many cell/tissue/region-specific and spatio-temporally expressed long noncoding RNAs, which bind and recruit epigenetic regulatory enzyme complexes and other transcriptional factors, have been characterized to be essential for plethora of brain functions (Goff et al. 2015; Guennewig and Cooper 2014; Khalil et al. 2009; Lasalle et al. 2013; Mercer et al. 2008) such as brain development (Feng et al. 2006; Lin et al. 2014a; Lv et al. 2013), differentiation (Lin et al. 2014a; Mercer et al. 2010; Ramos et al. 2015), myelination (Lin et al. 2014b), synaptic transmission, strength and plasticity (Bernard et al. 2010), learning and memory, neurogenesis (Aprea et al. 2013; Ng et al. 2013). They are also dysregulated in many neurological diseases (Clark and Blackshaw 2014; Johnson 2012; Roberts et al. 2014b; Ziats and Rennert 2013). However, involvement of lncRNAs in brain aging and age-associated neurological diseases needs further investigation.
In our laboratory, a novel 1339 bp long, repeat sequence containing cDNA, named as LINC-RBE (long intergenic noncoding RNA-rat brain expressed; Accession no. GQ463152) has been isolated by screening of a rat testis λgt11 cDNA library by using a 227 bp rat genomic simple repeat DNA (Accession No. X97459) as a probe (Bajaj 2002; Dey 2000; Dey and Rath 2005). Bioinformatically, LINC-RBE (cDNA) was characterized as a trans-spliced non-protein coding transcript from the rat chromosome 5 and 3, with ~11.7 % of different types of repeat sequences (GA, CA and SINE B2/B4) (Mishra 2009). Through expression analysis by RT-PCR and Northern blotting, LINC-RBE was found to be strongly expressed in multiple rat tissues and when used as a probe, detected various large cellular RNAs in the size range of 10–0.2 kb as well as small RNAs of 20–30 nt., which is in accordance with the in silico based sequence homology of LINC-RBE (cDNA) with eight rat-specific piRNAs, therefore, suggesting their possible processing from the larger transcript(s) and playing role as precursor transcripts in the generation of the small regulatory RNAs. Thus, LINC-RBE represented a class of repeat sequence containing lncRNAs from the rat genome with unknown function. In the present study, analysis of LINC-RBE expression in different neurobiologically distinct compartments of the brain and their age-dependent functional significance was studied in 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats. Through RT-PCR and RNA in situ hybridization, LINC-RBE was reported to be expressed distinctly and differentially as a sense transcript in the cortex, hippocampus and cerebellum regions of the rat brain. LINC-RBE expression increased from young to adult and decreased from adult to old. Therefore, it suggests that repeat-rich lincRNAs could be crucially linked to various complex functions of the brain during maturation from young to adult phases of life, and they are also profoundly altered in the brain during aging from adulthood to old age (Kour 2015).
2 Materials and Methods
2.1 Bioinformatic Analyses
The genomic organization of LINC-RBE cDNA, its chromosomal location, sequence conservation among different vertebrates and homology with other RNA sequences was analyzed by using ESEMBL, UCSC genome browser (http://genome.ucsc.edu/cgibin/hg Blat) and BLAST (http://www.Ncbi.nlm.nih.gov/BLAST) search engines. Its coding potential was found through ORF finder (www.ncbi.nlm.nih.gov/gorf/gorf.htm) and Codon Potential Calculator (CPU) (http://cpc.cbi.pku.edu.cn/docs/terms.jsp), respectively. The functional regulatory elements (putative promoter region, transcriptional factor binding sites, polyA signal, presence of untranslated region or UTR element and possible miRNA target site) in the 2000 bp up/downstream genomic region of LINC-RBE was scanned by using PromoScan (www.bimas.cit.nih.gov/molbio/proscan), GENSCAN (http://genes.mit.edu/GENSCAN.html), RegRNA2.0 (http://regrna2.mbc.nctu.edu.tw/), and Promo3 (http://alggen.lsi.upc.es/cgi-bin/promo_v3), respectively.
2.2 Semi-quantitative Reverse Transcription Polymerase Chain Reaction (RT-PCR)
Most of the molecular biology methods were performed as per the instructions given in the Sambrook’s molecular cloning laboratory manual (Sambrook 2001). The 4 weeks (young), 16 weeks (adult) and 70 weeks (old) male rats were anesthetized with 80 mg/kg Ketamine and 20 mg/kg Xylazine injection (as approved by the Institutional Animal Ethics Committee) and the whole brains were collected. Total RNA was isolated from the brains of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) Wistar rats (Rattus norvegicus) by following the Trizol-method as recommended by the manufacturer (Sigma-Aldrich). Prior to cDNA synthesis, 10 μg of total RNA was treated with RNase-free DNase I (Sigma-Aldrich) to remove the DNA contamination and then purified by phenol-chloroform extraction method. The cDNA was synthesized by reverse transcription from 1.0 μg of DNase-treated RNA in a 25 μl reaction mixture containing 500 ng of oligo-(dT) primer, 0.5 mM dNTPs, 1× M-MLV RT-reaction buffer, 20 U RNasin and 100 U of M-MLV Reverse Transcriptase at 37 °C for 1 h. For the amplification of LINC-RBE and GAPDH (a positive internal control) cDNAs, PCR was carried out by using 2.5–5.0 μl of cDNA in a 25 µl PCR-reaction mixture containing 1× Taq-buffer with 2 mM MgCl2, dNTPs (0.2 mM), Taq DNA polymerase (1 U) and 25 pmol of primer pair specific for LINC-RBE (RBE Fwd: 5ʹCCCAAAATGAGCAAGTAAGGAA3ʹ and RBE Rev: 5ʹTGTCAACAGAAGCCCTTTTTCA3ʹ) and GAPDH mRNA (GAPDHFwd: 5′ACCACAGTCCATGCCATCAC3′; GAPDH Rev: 5′ TCCACCACCCTGTTGCT GTA3ʹ). The amplification conditions used were as follows: initial denaturation at 95 °C for 4 min; followed by 35 cycles of denaturation at 95 °C for 45 s, annealing at 54 °C (LINC-RBE) or 60 °C (GAPDH) for 45 s and extension at 72 °C for 45 s and a final extension at 72 °C for 10 min.
2.3 Strand-Specific Expression by RT-PCR
For strand-specific RT-PCR, cDNA was synthesized from 1 µg DNase I-treated total RNA at 50 °C for 1 h by using 0.7–0.9 µM of either sense- or antisense strand-specific primer in a 25 µl reaction mixture containing dNTPs (1 mM), RNase-inhibitor (40 U), 1× M-MLV RT-buffer, M-MLV reverse transcriptase (200 U). The synthesized cDNA was PCR-amplified by using LINC-RBE specific primer pairs for total of 35 cycles under conditions as follows: denaturation at 94 °C for 45 s, primer annealing at 54 °C for 45 s and extension at 72 °C for 45 s. The amplification from the oligo-(dT) primer synthesized cDNA was used as a positive control.
2.4 RNA In Situ Hybridization in Rat Brain Tissue
2.4.1 Gelatin-Coated Slide Preparation
The glass slides, pre-cleaned with 0.2 % HCl and DEPC-treated H2O, were incubated with gelatin solution (0.5 % gelatin and 0.05 % chromium potassium sulphate) in a coupling jar for 1–2 min. The excess gelatin was drained out and the slides were dried at 42 °C for 2–3 h and stored at RT for future use.
2.4.2 Tissue Perfusion, Paraffin Wax Embedding and Tissue Sectioning
The brain tissues from 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats were incubated in 4 % formaldehyde solution for overnight for further fixation and later on dehydrated with a series of increasing ethanol concentrations: 30 % for 30 min., 50 % for 30 min, 70 % for overnight, 95 % for 1 h and absolute ethanol for 1 h at 4 °C. The dehydrated tissue was incubated in xylene and paraffin wax (pre-melted at 60 °C) solution (1:1) at 60 °C for 30 min and subsequently immersed in pre-melted paraffin wax for 3 h at 60 °C. Then ~6–10 µm thick sagittal sections of the wax-moulded tissue were cut by using microtome and the sections were mounted on 0.5 % gelatine-coated slides.
2.4.3 Preparation of RNA Probe by In Vitro Transcription
The cDNA for the noncoding RNA was subcloned with either T7 or T3 RNA polymerase promoter region flanking at the 5ʹ-end or 3ʹ-end. The pBluescript KS(+) construct (1 ng) containing such DNA template was PCR-amplified in a 25 µl PCR-reaction consisting of 1 × Taq buffer, MgCl2 (2 mM), dNTPs (0.2 mM), Taq DNA polymerase (1U) and 25 pmol of strand-specific primer pair for LINC-RBE (sense primer pair: RBE-T7Fwd: 5ʹTAATACGACTCACTATAGGCGGCCCAAAATGAG3ʹ; RBE-T7Rev: 5ʹATGCAATTCT TTGTGTT 3ʹ) and antisense primer pair (RBE-T3Fwd: 5ʹAATTAACCCTCACTAAAGGA TGCAATTCTTTGTGTT3ʹ; RBE-T3Rev: 5ʹCGGCCCAAAATGAG3ʹ), with forward primer containing either T7 or T3 promoter sequence at their 5ʹ-end, to amplify the cDNA from both orientations. The PCR conditions used were: 30 cycles of denaturation at 94 °C for 45 s, annealing at 55 °C for 45 s and extension at 72 °C for 1 min, 30 s. Finally, 500 ng of purified PCR-amplified DNA was used for in vitro transcription of the RNA probes at 37 °C for 1 h in a reaction mixture (50 µl) containing T3/T7 transcription buffer (1×), DTT (2 mM), rNTP mixture (1 mM each of ATP, GTP, CTP; 0.65 mM of UTP and 0.35 mM of UTP-Digoxygenin (dig), RNase inhibitor (40 U) and 40 U of T7/T3 RNA polymerase. The integrity of the RNA product was checked by electrophoresis in a 1.2 % TBE-agarose gel.
2.4.4 Hydrolysis of RNA Probe
The optimal length (~500 bp) of the dig-labelled RNA probes was obtained by hydrolysis method as given by Cox et al. (1984) with few modifications (Cox et al. 1984). Briefly, the in vitro synthesized RNA probe was incubated with two volumes of carbonate buffer (pH 10.2) at 65 °C for optimum time period ‘t’, calculated using the formula, t = (Lo − Lf)/k (Lo)(Lf), where t = time in minutes; Lf = final probe size; Lo = initial probe size and k = 0.11. The hydrolysis reaction was stopped by adding equal volume of neutralizing solution (sodium acetate, pH 5.0), the RNA probe was precipitated with 2.5 volumes of ethanol and one tenth volume of 3 M sodium acetate (pH 5.2) and diluted into a 50 % deionized formamide and 4× SSC solution.
2.4.5 RNA In Situ Hybridization
For RNA in situ hybridization, paraffin-embedded tissue sections were dewaxed, hydrated and permeabilized with 5 μg/ml RNase-free Proteinase K solution in 1× PBS-Triton X-100 for 30 min. at 37 °C. After stopping the reaction with 100 mM glycine solution, the sections were post-fixed in 4 % paraformaldehyde for 10 min. at 4 °C, rinsed twice with DEPC-treated 1× PBS and acetylated twice with 0.1 M tri-ethanolamine (TEA) buffer (pH 8.0) containing 0.25 % (v/v) acetic anhydride for 5 min each. The sections were pre-hybridized in a buffer containing 50 % deionized paraformaldehyde, 4× SSC, 1× Denhardt’s solution at 50 °C and after 2 h, the solution was replaced with hybridization buffer (50 % deionized paraformaldehyde, 4× SSC, 1× Denhardt’s solution, 1 mg/ml denatured herring sperm DNA, 10 % dextran sulphate) containing 30 ng of either sense or antisense digoxygenin-labelled RNA probes and 1 mg/ml tRNA, which was pre-incubated at 65 °C for 5 min. The sections were incubated at 50 °C overnight in a moist chamber. After hybridization, the sections were successively washed in 2× SSC, 1× SSC and 0.1× SSC, treated with 20 µg/ml RNase A to remove any single stranded RNA probe and incubated with blocking solution (100 mM Tris.HCl, pH 7.5, 100 mM NaCl, 2 % BSA and 0.1 % Triton X-100) for 30 min at RT. The dig-labelled hybridized probe was detected by incubating the sections in a blocking solution containing 1:1000 dilution of sheep anti-digoxigenin-alkaline phosphatase-Fab fragment antibody (Roche) for 2 h in a humid chamber followed by development of color due to the alkaline phosphatase activity with the NBT/BCIP substrate solution. The sections were stored at RT for overnight and images were captured under bright field microscope (Nikon-TiS) at 50× and 600× optical magnifications.
2.5 Quantification and Statistical Analysis
For comparison of the RNA expression patterns in the brains of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats, the number of in situ hybridization (ISH)-positive cells were counted and normalized with the number of haematoxylin and eosin-stained cells counted from total of nine different areas of the cortex, three from cerebellum and one each from CA1, CA2, CA3 and dentate gyrus of hippocampus. Similarly, by keeping the base value of black and white to 0 and 1, the mean intensity of ISH-positive cells were quantified by using the Nikon NIS-AR software and the percentage mean intensity of expression relative to its expression in the adult was calculated. The significance of RNA expression was determined by using ANOVA at 5 % significance level and the comparison of the significance between different age-groups was carried out with Turkey test. Each experiment was repeated three times.
3 Results
In order to explore biological function of repetitive DNA present in mammalian genome, we had earlier isolated a novel genomic simple repeat DNA sequence (Accession No. X97459) from the Wistar rat (Rattus norvegicus) genome, which showed triplex (H-DNA) structure in vitro; RNA-homology with many eukaryotic mRNAs mostly in the 5′-untranslated region (5′-UTR) and 3′-UTR due to presence of simple repeat sequences in the mRNAs and expression of endogenous, large-sized, complementary RNA in the rat brain suggesting constitutive expression of long noncoding RNAs from the repetitive DNA of the rat genome under in vivo conditions (Dey 2000; Dey and Rath 2005). This was one of the initial examples of long noncoding RNA expression in mammalian tissue. In order to clone RNAs with repeat sequences, the X97459 genomic simple repeat DNA was used as a probe to screen cDNA library from the rat and human testis and several repeat sequence containing cDNAs were isolated (Bajaj 2002; Dey 2000). Upon DNA sequencing, bioinformatic analyses and RNA expression studies, these cDNAs represented some long noncoding RNAs from the rat genome (Mishra 2009). LINC-RBE is one such repeat sequence containing lncRNA and was originally deposited in the NCBI Genbank as pRT5.5 (Accession no. GQ463152). This was later renamed as LINC-RBE. Here we describe sequence analysis and differential expression patterns of LINC-RBE in the rat brain by RT-PCR and in different parts of the rat brain by in situ RNA hybridization in the young (4 weeks), adult (16 weeks) and old (70 weeks) rats, which has been recently reported (Kour and Rath 2015). The results are discussed in the light of an example of the lincRNAs, differentially expressed in the cortex, hippocampus (subregions: CA1, CA2, CA3 and dentate gyrus) and cerebellum regions of the mammalian brain in an age-dependent manner and its possible functional significance during maturation and aging of the brain. This example argues in favour of the possible role of lncRNAs in maturation and aging of mammalian brain.
3.1 Sequence Characterization of LINC-RBE in the Rat Genome
The sequence analysis of any given transcript in relation to its genomic location, homology to other functional RNAs, evolution and conservation over time among different species, presence of putative regulatory elements in its up/downstream genomic region provide sufficient basic information necessary for analyzing its biological function in an organism. Similarly, we tried to find out the functional sequence characteristics of LINC-RBE in rat genome.
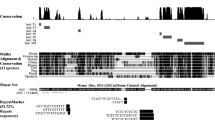
By using rat Mar. 2012 (RGSC 5.0/rn5) assembly on UCSC Genome browser, we found that the 5ʹ1–1213 nt. (~92 %) region of LINC-RBE (cDNA) sequence is homologous to the intergenic region of rat chromosome 5 (Chr.5 q33) and the remaining 1213–1334 3ʹ nt. (8 %) region is homologous to the intergenic region of rat chromosome 3 (Chr.3 q36), therefore, suggesting it to be a trans-spliced intergenic transcript. Also, Chr.5 q33 region, upstream of LINC-RBE (genomic DNA), contains transcription sites for various piRNAs (Girard et al. 2006; Lau et al. 2006), eight of which (DQ73550, DQ628056, DQ752533, DQ747205, DQ619691, DQ751276, DQ748367 and DQ613867) were found to be generated from within the LINC-RBE (cDNA) sequence (Fig. 2a; Table 3). Although the sequence comparison study of 5ʹ1–1213 nt. region of LINC-RBE (cDNA) in UCSC genome browser showed LINC-RBE to be less evolutionarily constrained and moderately conserved among 13 different vertebrate species (Chodroff et al. 2010; Johnsson et al. 2014; Ponjavic and Ponting 2007), the presence of various tandem conserved regions in its upstream region at chromosome 5 and its highly conserved syntenic similarity with the piRNA clusters in the mouse chromosome 4 intergenic region argues for its possible function as a piRNA-precursor long noncoding RNA (Fig. 2b) (Kour and Rath 2015). One of these highly conserved regions, lod = 19, showed homology to four mouse (Mus musculus) piRNAs (DQ687281, DQ725370, DQ699388 and DQ726299) and one rat (Rattus norvegicus) (DQ618908) piRNA (Fig. 2a).
Sequence analysis of LINC-RBE (cDNA). a Sequence homology of 1339 bp LINC-RBE (cDNA) (5′-1217 bp region) to the rat chromosome 5 (q33.3) and various rat and mouse piRNAs (already described in the rat genome database) transcribed from this region. b Conservation studies using UCSC Genome browser showed LINC-RBE (cDNA) to be moderately conserved among vertebrates with one highly conserved syntenic region in the intergenic region of mouse chromosome 4. c The coding potential of LINC-RBE (cDNA), analyzed by using ORF Finder tool, showed putative ORF of less than 300 codons but the predicted peptides did not show any homology with known peptides in the protein database. d Similarly, the coding potential of LINC-RBE (cDNA) by using codon potential calculator showed very low coding score and ORF coverage. Thus, LINC-RBE is a long noncoding transcript [from Fig. 1a and 1b of Kour and Rath (2015) with permission]
Furthermore, the scanning of 2000 bp up/downstream sequence of LINC-RBE (cDNA) on the chromosome 5 by using RegRNA2.0 and Genomatix Genome analyzer found binding sites for many important transcriptional factors involved in cell growth, proliferation and development, such as Activator Protein-1 (AP-1), SRY (sex determining region Y)-box 9 (Sox9), Retinoic Acid Receptor (RAR-alpha: RXR-alpha), Glucocorticoid Receptor (GR) and many others (Fig. 3b); 14 binding sites of the RNA binding protein, Musashi, which is a characteristic translational activator of certain temporally expressed mRNAs in Xenopus oocyte and a known context-dependent translational regulator in proliferating mammalian cells through its modulation of cytoplasmic polyadenylation (Rutledge et al. 2014; Sutherland et al. 2013) (Fig. 3c); two polyadenylation signals (Fig. 3b), one within (5ʹ AAATAAATCCAAACTCCAAATTGCCTTT3ʹ) and one ~903 bp downstream of LINC-RBE (genomic DNA) sequence (5ʹ AATAATCATCTGATGGTTTCATGTTACCTTTGT TT TC3ʹ); and one target site of the miRNA, rno-mir466-b within LINC-RBE (genomic DNA) (Fig. 3d). Thus, it suggests the multi-factorial regulation of LINC-RBE expression in the rat genome. Besides, a putative promoter region (250 bp) containing the regulatory binding sites for the transcription factors: Nuclear Factor kappa B (NF-κB), Serum Response Factor (SRF), Octamer-binding transcription factor (Oct factors) and Transcription Factor IID (TFIID) at 1250 bp downstream of LINC-RBE (genomic DNA) and −26 to −276 bp from the putative transcriptional start site (TSS) (+1) at chromosome 5 was also found by using PromoterScan (Fig. 3a). However, through Codon Potential Calculator (CPC) (Grote et al. 2005) and NCBI Open Reading Frame (ORF) finder tools, we found that LINC-RBE has a characteristic feature of a non-protein coding transcript with a low coding potential score of −1.11689 (lod score 31.92) and ORF coverage of 21.73 % (lod score value = 31.92) (Fig. 2c). The predicted small ORFs (128, 292, 132, 138 codon size) showed considerably small sequence homology with any known peptides in the protein database, thus, falling short of the criteria for a polypeptide to be considered as a functional protein (Fig. 2d). Overall, bioinformatic studies have confirmed that LINC-RBE is an intergenic lncRNA, which despite being less evolutionarily conserved, could possibly be regulated by various factors and might act as a precursor RNA for various piRNAs in rat cells and tissues.
Presence of various regulatory elements in the up/downstream genomic region of LINC-RBE. a Schematic representation showing presence of the putative promoter region, consisting of binding sites for the transcription factors: NF-κB, Oct-2, NF-S, TFIID, at 1250 bp downstream of LINC-RBE (genomic DNA) in the rat chromosome 5. b–d Schematic representation of presence of various regulatory elements such as polyadenylation signals (b), transcriptional factor binding sites (b), UTR-elements (c), and possible miRNA target sites (d) in the 2000 bp up/downstream region of LINC-RBE (genomic DNA) in the rat chromosome 5 were analyzed by using RegRNA 2.0, Genomatix Genome analyzer and Promo 3.0, respectively [from Fig. 1c of Kour and Rath (2015) with permission]
3.2 Transcriptional Strand-Specificity of LINC-RBE in Adult Rat Brain
The different modes of transcription of various lncRNAs, such as sense, antisense, overlapping and bidirectional, in the mammalian genome could provide information about their biological role. Therefore, we assessed the transcriptional strand-specificity of LINC-RBE in the adult rat brain to gain insight into their possible role by using strand-specific sense or antisense primer during the reverse transcription step of RT-PCR (Fig. 4a). We found that in case of LINC-RBE, an amplicon of 545 bp was obtained from the reaction containing cDNA synthesized by using the antisense LINC-RBE specific-primer (RBE-Fwd), whereas no amplification was obtained in case of the sense-specific primer (RBE-Rev) (Kour and Rath 2015). Therefore, based on the complementarity of RBE-Fwd primer to the reverse (minus) strand of chromosome 5, we inferred that LINC-RBE is transcribed as a sense-transcript from the intergenic region of rat chromosome 5. The amplification from the oligo-(dT) primer synthesized cDNA suggested that LINC-RBE is a polyadenylated lincRNA.
Age-dependant expression of LINC-RBE in the rat brain by RT-PCR. a Strand-specificity of transcription of LINC-RBE in the adult (16 weeks) rat brain was determined by using strand-specific sense (RBE-Fwd) or antisense (RBE-Rev) primer during cDNA synthesis step of RT-PCR. b Expression of LINC-RBE in the brain of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats was studied by RT-PCR. GAPDH mRNA was used as an internal control. c, d LINC-RBE expression was normalized to that of GAPDH mRNA. −RT Negative control; M 100 bp DNA-ladder; LC loading control (545 bp DNA-amplicon); *** p < 0.001 (n = 4). [from Fig. 2 of Kour and Rath (2015) with permission]
3.3 Analysis of LINC-RBE Expression by RT-PCR in Rat Brain During Aging
In mammals, through extensive high-throughput RNA sequence analysis, it has been shown that the brain has the highest transcriptome complexity in comparison to all other organs (Soumillon et al. 2013). Diverse groups of lncRNAs with different neurobiological functions such as brain development, differentiation, myelination etc. have been described in literature, however, association of lncRNAs with the process of aging and age-related brain diseases and disorders such as Alzheimer, Parkinson etc. have recently become evident. Since LINC-RBE was strongly expressed in the adult rat brain, we, therefore, investigated its likely involvement during maturation and aging of the brain by studying its expression pattern in the brains of 4 weeks (young), 16 weeks (adult), and 70 weeks (old) rats by RT-PCR (Fig. 4b, c). The expression of GAPDH mRNA was taken as an internal control and used for normalization of LINC-RBE expression. We found that the expression of LINC-RBE, measured as integrated density value (IDV), significantly (~2× fold) increased from young to adult (p = 0.0075) and then decreased (~1.7× fold) with aging from adult to old (p = 0.017) in the rat brain (Fig. 4c) (Kour and Rath 2015). Thus, the age-dependent differential expression patterns of LINC-RBE in the rat brain suggest its possible involvement in functions related to maturation and aging of the brain.
3.4 Expression and Localization of LINC-RBE by RNA In Situ Hybridization in Rat Brain During Aging
Recently, numerous studies have shown that many functionally characterized lncRNAs follow distinct cell type-, tissue- and developmental stage-specific expression patterns in mammalian brain, which define and relate to their biological roles. The age-dependent expression of LINC-RBE in rat brain has, therefore, led us to further elaborate on its possible function based on its cell type and sub-cellular expression pattern in different functionally specialized neuroanatomical regions of rat brain during aging. This was performed by RNA in situ hybridization in paraffin-embedded brain sections of immature (4 weeks), adult (16 weeks) and old (70 weeks) rats by using digoxygenin-labelled sense- and antisense LINC-RBE-specific riboprobes. The LINC-RBE was found to be differentially expressed with respect to cell types, cell number and intensity of expression, specifically in the pyramidal and granule cells of the forebrain (cortex and hippocampus) and granule and Purkinje cells of cerebellum regions of the brain in an age-dependent manner (Figs. 5, 6 and 7).
Age-dependent expression of LINC-RBE in the cortex of the rat brain by RNA in situ hybridization. a Differential expression pattern of LINC-RBE in paraffin-embedded sections (10 µm thick) of the cortex of the brain of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats by in situ RNA hybridization by using digoxygenin-labelled sense- and antisense strand-specific RNA probes as well as no probe negative control. Age-dependent differential expression pattern of LINC-RBE in the pyramidal and granule cells of different cortical layers of the rat brain with respect to the three age-groups are shown. Bar Scale, 100 and 5 µm. b Number of cells positive for LINC-RBE expression normalized to the number of cells stained by Haematoxylin/Eosin in the cortex of the brain. c Alterations in the intensity of expression of LINC-RBE in the cortex of the brain of young and old relative to that of the adult rats. *** p < 0.001; ** p < 0.01 (n = 3). [from Fig. 3 of Kour and Rath (2015) with permission]
Age-dependent expression of LINC-RBE in the hippocampus of the rat brain by RNA in situ hybridization. a Differential expression pattern of LINC-RBE in paraffin-embedded sections (10 µm thick) of the hippocampus of the brain of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats by in situ RNA hybridization by using digoxygenin-labelled sense- and antisense strand-specific RNA probes as well as no probe negative control. Bar Scale, 100 µm. b–e Expression of LINC-RBE in the granule cells of the dentate gyrus (b), and pyramidal cells of CA1 (c), CA2 (d), CA3 (e) sub-regions of the hippocampus. Bar Scale, 5 µm. (f) Number of cells positive for LINC-RBE expression normalized to the number of cells stained by Haematoxylin/Eosin in the hippocampal sub-regions of the brain. (g) Alterations in the intensity of LINC-RBE expression in the hippocampal sub-regions of the brain of young and old relative to that of the adult rats. *** p < 0.001; ** p < 0.01; p < 0.05 (n = 3). [from Fig. 4 of Kour and Rath (2015) with permission]
Age-dependent expression of LINC-RBE in the cerebellum of the rat brain by RNA in situ hybridization. a Differential expression pattern of LINC-RBE in the granule (G) and Purkinje (P) cells of the cerebellum in paraffin-embedded sections (10 µm thick) of the brain of 4 weeks (young), 16 weeks (adult) and 70 weeks (old) rats by in situ RNA hybridization by using digoxigenin-labelled sense- and antisense strand-specific RNA probes as well as no probe negative control. Bar Scale, 100 and 5 µm. b Number of cells positive for LINC-RBE expression normalized to the number of cells stained by Haematoxylin/Eosin in the cerebellum of the brain. c Alterations in the expression of LINC-RBE in the cerebellum of the young and old relative to the adult rats. *** p < 0.001 (n = 3). [from Fig. 5 of Kour and Rath (2015) with permission]
In brain cortex, LINC-RBE showed varied cell type-specific expression profile with a punctate expression in the granule cells of young and granule/pyramidal cells of old rats, but, a distinct and strong expression pattern was observed in adult rats, which was confined mainly to outer pyramidal and granule molecular cortical cell layers. The relative percentage mean intensity of expression of LINC-RBE, calculated with respect to adult, showed a significant decrease from adult to young and old (F = 93.844, p < 0.001) (Fig. 5a, b), however, this decrease did not correlate with any possible decrease in the number of cells expressing LINC-RBE with increasing age (Fig. 5c) (Kour and Rath 2015). Similarly, in the hippocampus, expression and localization of LINC-RBE showed a high variability in relative intensity in pyramidal cells of CA1, CA2, CA3 regions and granule cells of dentate gyrus among young, adult and old rats (Fig. 6). During brain aging, LINC-RBE expression in hippocampus showed a significant increase of 24 % and 13.1 % in pyramidal cells of CA2 (F = 38.105, p < 0.001) (Fig. 6d, f) and CA3 (F = 7.318, p = 0.025) (Fig. 6e, f) regions, and 18.2 % in granule cells of supra-pyramidal blade of dentate gyrus (F = 28.462, p < 0.001) (Fig. 6b, f), respectively from young to adult, and a decrease of 19.41 % in CA2 (p = 0.001); 11.82 % in CA3 (p = 0.046) and 14.5 % in the dentate gyrus (p = 0.003) regions from adult to old rats. The CA1 region showed no profound change in LINC-RBE expression during brain aging (Fig. 6c, f). Besides this, the number of LINC-RBE positive cells showed a significant variation in CA2 and dentate gyrus, whereas CA1 and CA3 regions showed no change with increasing age (Fig. 6g). The LINC-RBE positive cell population was ~30–35 % in the hippocampal CA2 regions of young and old relative to ~45 % of cells in the adult (F = 8.173, p = 0.019). On the contrary, in suprapyramidal blade of dentate gyrus, LINC-RBE expressing cells showed ~threefold reduction from young and adult to old brains, thus, suggesting the dynamic expression of LINC-RBE with respect to maturation and aging in the rat hippocampal sub-regions (Fig. 6g). Furthermore, LINC-RBE expression was also observed in the Purkinje and granule cells of the cerebellum in young, adult and old rats. Similar to cortical region, the expression of LINC-RBE initially showed a significant increase of ~21 % from young to adult and then a decrease of ~8 % from adult to old in granule cells (F = 158.151, p < 0.001) (Fig. 7a, b). There were ~45 % of LINC-RBE positive cells in adult cerebellum as compared to ~35 % in young and old rats during aging (F = 55.787, p < 0.001) (Fig. 7a, c) (Kour and Rath 2015). Thus, the differential cell type and region-specific expression pattern of LINC-RBE in the three transcriptomically and functionally complex regions of the rat brain, i.e., cortex, hippocampus and cerebellum, with increasing age might suggest its potential functional significance in age-related cognitive processes and neurological diseases.
4 Discussion
Evolutionary conservation of newly discovered protein-coding genes are generally based on their nucleotide or amino acid sequences, which provide invaluable means to understand and evaluate their relatedness, functional significance and probable lineage-specific phenotypes. However, although lncRNAs are abundantly expressed and functionally diverse, these criteria do not explain their selection process during evolution and functional relevance among various species in depth (Chodroff et al. 2010; Guo et al. 2014; Johnsson et al. 2014). Refraining from the conventional concept of linking conservation of sequence to function, there are examples of many functionally characterized mammalian lncRNAs such as Myocardial Infarction Associated Transcript (MIAT), HOX transcript antisense RNA (HOTAIR), X inactive-specific transcript (XIST), Embryonic ventral forebrain-2 (Evf-2) and Antisense Igf2r RNA (Air) with poor sequence conservation among species (Diederichs 2014; Johnsson et al. 2014; Roberts et al. 2014a; Wood et al. 2013). Further, despite showing rapid evolution with accumulating mutation and lacking sequence ortholouge, many lncRNAs show syntenic locus conservation among different species with highly conserved regulatory regions and show specific sub-cellular, cell/tissue and developmental expression patterns in mammals, which as a whole, argues for their function (Diederichs 2014; Mercer et al. 2008; Ponting et al. 2009).
Similarly, by homology studies we found modest sequence conservation of LINC-RBE among 13 different vertebrates with the presence of a highly conserved syntenic locus in mouse chromosomes 4. The presence of various small conserved regions in the up/downstream region of LINC-RBE in chromosome 5 are the sites for synthesis for different piRNAs in the rat and mouse genome, further asserting its function as a precursor lncRNA that may be processed by RNA processing mechanisms to produce many small noncoding regulatory RNAs (piRNAs). In accordance with presence of up/downstream conserved spots, through sequence analysis, we found presence of binding sites for various cell growth/differentiation-specific regulatory transcriptional factors such as RAR-alpha:RXR-alpha, Oct-1, PARP, GR and AP-1, NF-κB and SOX9 in the 2000 bp up/downstream region of LINC-RBE (genomic DNA) in rat chromosome 5. Taken together, our results may suggest the multifactorial regulation of expression and functional relevance of LINC-RBE in the rat genome. Involvement of LINC-RBE in regulation of aging process or its related diseases is emphasized due to the presence of binding sites for major age-related transcriptional factors, GR and NF-κB, (Adler et al. 2007; Rapicavoli et al. 2013), in the up/downstream sequence of LINC-RBE (genomic DNA).
Mammalian genome expresses a huge number of different lncRNAs that are transcribed in sense/antisense orientation with respect to protein-coding genes (Wood et al. 2013). One such example is the long intergenic noncoding RNA repeat-rich sense-antisense transcript (LINC-RSAS) from the chromosome 17, which is expressed in both sense and antisense orientations in the rat brain (Kour and Rath 2016a). Similarly, transcriptional orientation studied for LINC-RBE from the rat chromosome 5 showed its expression in sense orientation in the brain. In mammals, many lncRNAs with variable biological roles showed distinct tissue/organ and developmental stage-specific expression, which specified their tissue-specific function. LINC-RBE was found to be expressed in multiple rat tissues (Mishra 2009) and its expression was high in the adult rat brain. We, therefore, studied its expression in the rat brain during aging.
Aging is defined as a global intrinsic biological phenomenon of continuous and cumulative deterioration of neurological, immunological and physiological as well as cellular and molecular functions in an organism with increasing age (Robert et al. 2010). Alteration of many biological pathways/processes/mechanisms as well as cellular protein levels are proposed to be involved in its onset and progression. Aging is considered to be a major risk factor in the patho-physiology of various diseases and disorders (Robert et al. 2010; Sinha et al. 2014). Recently, through advanced RNA sequencing techniques, vast number of functionally diverse regulatory RNAs, sncRNAs and lncRNAs, have been reported, which could provide another way to explore and discourse the complexity of aging process and its related diseases (Kour and Rath 2016c).
Involvement of small noncoding RNAs in aging and age-related diseases such as cardiac malfunctions (mir-18, mir-19, mir-241, mir-214, mir-217, mir-146) (van Almen et al. 2011; van Balkom et al. 2013), neurological disorders (miR-34a, mir-29, mir-144, mir-107) have been studied extensively. However, function and age-related expression of lncRNAs are much less explored. Till date, only a few reports of modulation of long noncoding RNAs during cellular senescence are known, e.g., senescence-associated known long noncoding RNAs such as Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1), MIAT, Taurine up-regulated gene 1 (TUG1) and novel lncRNAs such as XLOC025918, XLOC025931, XLOC023166 mediate proliferation defects and induce senescence phenotype in cultured human fibroblasts upon their down-regulation (Abdelmohsen et al. 2013). Many lncRNA such as H19, Antisense noncoding RNA in the INK4 locus (ANRIL), Air, Telomeric repeat-containing RNAs (TERRA), HOTAIR, Long intergenic non coding RN-p21 (lincRNA-p21) could possibly be implicated in aging process based on their regulatory role in heterochromatin and telomere maintenance, p53-mediated cell proliferation and apoptosis, a paramount factor during progression of aging and its related diseases (Kour and Rath 2016c). Similarly, LINC-RSAS, a repeat-rich 942 bp intergenic lncRNA has been found to be differentially expressed in sense-antisense orientations and age-dependent manner in the rat brain (Kour and Rath 2016a). However, still much is unclear regarding their direct implication and mechanism through which they regulate aging process. In our study, by using RT-PCR and in situ RNA hybridization, we found strong and differential expression of a novel lincRNA, LINC-RBE in the rat brain during aging. Its expression in the brain showed a marked increase from the young (4 weeks) to adult (16 weeks) followed by a subsequent and significant decrease from the adult to old (70 weeks) rats. This suggested the possible involvement of LINC-RBE in functions related to maturation and aging of the brain.
Mammalian brain is a heterogeneous, complex functional system, primarily consisting of various arrangements of interconnected neurons and glial cells in transcriptomically diverse compartments but with elaborate networks. The cortex has many functional areas associated with visual, auditory, motor functions and cognitive functions such as memory, language, emotions, creativity and judgement. Cortex contains neurons of various shapes, sizes, density and different arrangements of neural fibres organized into multiple molecular layers with variable functions. The layer V and VI consisting mainly of pyramidal neurons are highly developed in the cortical regions involved in motor functions and interconnect cortex to subcortical regions by giving rise to efferent cortical projections to brain stem, spinal cord, basal ganglia and thalamus. The hippocampus consists of varied arrangements of extremely larger pyramidal neurons into CA1, CA2 and CA3 regions, and smaller granule neurons in the dentate gyrus region. The changes in dendritic complexity, synapse number, transmission, plasticity and formation of new neurons or synapses via neurogenesis in the hippocampus throughout life plays an important role in learning, generation and storage of memory processes. The cerebellum accounts for 50 % of total neurons in the brain. Anatomically, cerebellar cortex comprises of innermost granular layer, middle Purkinje cell layer and outermost molecular layer consisting of axons of granule cells and the dendrites of Purkinje cells. Cerebellum is primarily involved in motor functions (voluntary movements, balance and posture) and in cognitive functions such as language. The cortex, hippocampus and cerebellum regions of brain function together in controlling various cognitive functions such as spatial learning and generation of episodic memory.
High-throughput sequencing and microarray techniques to evaluate organ level transcriptome along with inputs from Allen Brain Atlas have reported the cell type- and developmental stage-specific dynamic expression of vast number of sense and antisense transcripts from specific regions/compartments of mammalian brain, which via their innumerable associations with various proteins/complexes are involved in complex brain functions such as learning, memory formation, synaptic variations and plasticity, myelination, neuron development and differentiation (Carrieri et al. 2015; Goff et al. 2015; Kadakkuzha et al. 2015; Mercer et al. 2008). Of late, analysis of RNAseq dataset of mouse brain has reported region-specific expression of 2759 lncRNAs in the hippocampus and 2561 lncRNAs in the prefrontal cortex region, the two main regions involved in various cognitive functions and neuropsychiatric disorders, and together they account for 70 % of the total annotated lncRNAs in the mouse genome (Kadakkuzha et al. 2015).
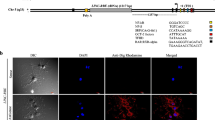
Similarly, through RNA in situ hybridization based expression studies in brain of young, adult and old rats, we found dynamic, age-dependent expression profile of LINC-RBE in different neuroanatomical regions of the brain such as cortex; CA1, CA2, CA3, and dentate gyrus subregions of hippocampus and cerebellum, involved primarily in generation of episodic and spatial learning and memory. The strong, dynamic expression profile of LINC-RBE, in terms of both cell number and intensity, in adult compared to young and old rat brains in the complex brain regions involved in cognitive functions (learning and memory), i.e., cortex, hippocampus and cerebellum, might suggest the possible involvement of LINC-RBE in modulation of various brain functions including neurogenesis during maturation and lack of it during aging. Furthermore, progression of many neurological disorders and diseases with impaired cognitive functions (memory and learning) such as Alzheimer’s disease, autism, Huntington disease have been associated with onset of brain aging (Carrieri et al. 2015). Therefore, further evaluation of LINC-RBE function(s) in the hippocampal and cortical regions of the brain during aging or onset and prognosis of many age-related neurological diseases/disorders would highlight a way to study its relevance in various brain functions and neuro-pathological conditions. Since the varied transcriptome based functional compartmentalization of mammalian brain is the result of differential regional- and cell-specific, epigenetically chromatin-modulated gene expression patterns, transcription of LINC-RBE in the cortex, hippocampus and cerebellum regions might suggest either its chromatin association or chromatin-based regulation of gene expression in the brain during maturation and aging. It may also be involved in various RNA processing pathways in the brain during maturation and aging. In another study, the relevance of LINC-RBE in the function of hippocampus was investigated in primary hippocampal neurons from the adult rat in the presence of a vitamin A derivative, all-trans retinoic acid (atRA), a known regulator of brain development, adult neurogenesis, synaptic plasticity and memory formation. AtRA was found to significantly upregulate expression of LINC-RBE in the nucleus and cytoplasm of the neurons along with the dendrites in a time- and dose-dependent manner (Kour and Rath 2016b). The atRA-mediated induction of LINC-RBE expression was found to be inhibited by actinomycin D, hence, it was regulated at transcriptional level. The possible binding of atRA along with the retinoic acid receptor (RAR: RXR) to the two binding sites present in the putative promoter of LINC-RBE was proposed. Altogether, the decrease in LINC-RBE expression in the hippocampus during brain aging and its transcriptional induction by atRA in the primary hippocampal neurons, could possibly argue for its significance in cognitive functions such as synaptic regulation, learning and memory formation as well as their decline during aging.
5 Conclusion
Transcription, gene expression and RNA processing are complex and heterogeneous in the brain. LncRNAs have emerged as major regulators of these processes. This study is a conclusive representation of possible role of lincRNAs in mammalian brain during maturation from young to adulthood and aging from adulthood to old age. Since the onset of many neurological diseases such as dementia, Alzheimer, Parkinson, Huntington occur late in life, study of molecular aspects of brain functions such as impairment of cognitive functions like short and long term memory formation, thinking capability, decision making etc. with increasing age would provide an important basis for understanding details of such age-related diseases and, therefore, would pave new avenues for development of successful therapeutics and treatments. LncRNAs in coordination with various factor(s)/complex(es) have been found to modulate almost each and every process of the flow of genetic information as depicted in the revised “central dogma of molecular biology”, i.e., chromatin organization, gene expression, RNA processing, translation and RNA/protein trafficking, thus, regulating nearly every aspect of biological processes required for life. The immense transcription and functional implication of lncRNAs in the mammalian brain, thus, would provide a foremost way to explore the biological process of aging and its related diseases extensively. The age-dependent and differential expression of LINC-RBE in specific cell types/regions of the brain, e.g., cortex, hippocampus and cerebellum, the three interconnected regions involved in memory processing and other cognitive functions, may further contribute to more detailed study of brain maturation, aging and age-related diseases at molecular level. LINC-RBE may be used as a biomarker for brain aging.
References
Abdelmohsen K, Panda A, Kang MJ, Xu J, Selimyan R, Yoon JH, Martindale JL, De S, Wood WH 3rd, Becker KG et al (2013) Senescence-associated lncRNAs: senescence-associated long noncoding RNAs. Aging Cell 12:890–900
Adler AS, Sinha S, Kawahara TL, Zhang JY, Segal E, Chang HY (2007) Motif module map reveals enforcement of aging by continual NF-kappaB activity. Genes Dev 21:3244–3257
Akula N, Barb J, Jiang X, Wendland JR, Choi KH, Sen SK, Hou L, Chen DT, Laje G, Johnson K et al (2014) RNA-sequencing of the brain transcriptome implicates dysregulation of neuroplasticity, circadian rhythms and GTPase binding in bipolar disorder. Mol. Psychiatry 19:1179–1185
Amiel JL (1965) The Nobel prize for Medicine 1965 is awarded to professors Lwoff, Monod and Jacob. Rev Fr Etud Clin Biol 10:1015–1017
Ananda G, Takemon Y, Hinerfeld D, Korstanje R (2014). Whole-genome sequence of the C57L/J mouse inbred strain. G3 (Bethesda) 4:1689–1692
Aprea J, Prenninger S, Dori M, Ghosh T, Monasor LS, Wessendorf E, Zocher S, Massalini S, Alexopoulou D, Lesche M et al (2013) Transcriptome sequencing during mouse brain development identifies long non-coding RNAs functionally involved in neurogenic commitment. EMBO J 32:3145–3160
Arning L, Ocklenburg S, Schulz S, Ness V, Gerding WM, Hengstler JG, Falkenstein M, Epplen JT, Gunturkun O, Beste C (2015) Handedness and the X chromosome: the role of androgen receptor CAG-repeat length. Sci Rep 5:8325. doi:10.1038/srep08325
Bajaj G (2002) Molecular cloning and characterization of human cDNAs by a simple repeat DNA probe, identification of novel candidate genes, Ph.D thesis. Jawaharlal Nehru University, New Delhi
Ballarino M, Cazzella V, D’Andrea D, Grassi L, Bisceglie L, Cipriano A, Santini T, Pinnaro C, Morlando M, Tramontano A et al (2015) Novel long noncoding RNAs (lncRNAs) in myogenesis: a miR-31 overlapping lncRNA transcript controls myoblast differentiation. Mol Cell Biol 35:728–736
Bao X, Wu H, Zhu X, Guo X, Hutchins AP, Luo Z, Song H, Chen Y, Lai K, Yin M et al (2015) The p53-induced lincRNA-p21 derails somatic cell reprogramming by sustaining H3K9me3 and CpG methylation at pluripotency gene promoters. Cell Res 25:80–92
Bates DJ, Li N, Liang R, Sarojini H, An J, Masternak MM, Bartke A, Wang E (2010) MicroRNA regulation in Ames dwarf mouse liver may contribute to delayed aging. Aging Cell 9:1–18
Bernard D, Prasanth KV, Tripathi V, Colasse S, Nakamura T, Xuan Z, Zhang MQ, Sedel F, Jourdren L, Coulpier F et al (2010) A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J 29:3082–3093
Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE et al (2007) Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447:799–816
Boehm M, Slack F (2005) A developmental timing microRNA and its target regulate life span in C. elegans. Science 310:1954–1957
Boon RA, Iekushi K, Lechner S, Seeger T, Fischer A, Heydt S, Kaluza D, Tréguer K, Carmona G, Bonauer A et al (2013) MicroRNA-34a regulates cardiac ageing and function. Nature 495:107–110
Cajigas I, Leib DE, Cochrane J, Luo H, Swyter K, Chen S, Clark BS, Thompson J, Yates JR 3rd, Kingston RE et al (2015) Evf2 lncRNA/BRG1/DLX1 interactions reveal RNA-dependent chromatin remodeling inhibition. Development 142:2641–2652
Capobianco E (2014) RNA-Seq data: a complexity journey. Comput Struct Biotechnol J 11:123–130
Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C et al (2005) The transcriptional landscape of the mammalian genome. Science 309:1559–1563
Carrieri C, Forrest AR, Santoro C, Persichetti F, Carninci P, Zucchelli S, Gustincich S (2015) Expression analysis of the long non-coding RNA antisense to Uchl1 (AS Uchl1) during dopaminergic cells’ differentiation in vitro and in neurochemical models of Parkinson’s disease. Front Cell Neurosci 9:114. doi:10.3389/fncel.2015.00114
Chassin C, Hempel C, Stockinger S, Dupont A, Kübler JF, Wedemeyer J, Vandewalle A, Hornef MW (2012) MicroRNA-146a-mediated downregulation of IRAK1 protects mouse and human small intestine against ischemia/ reperfusion injury. EMBO Mol Med 4:1308–1319
Chen D, Fu LY, Zhang Z, Li G, Zhang H, Jiang L, Harrison AP, Shanahan HP, Klukas C, Zhang HY et al (2014) Dissecting the chromatin interactome of microRNA genes. Nucleic Acids Res 42:3028–3043
Chiyomaru T, Fukuhara S, Saini S, Majid S, Deng G, Shahryari V, Chang I, Tanaka Y, Enokida H, Nakagawa M et al (2014). Long non-coding RNA HOTAIR is targeted and regulated by miR-141 in human cancer cells. J Biol Chem 289:12550–12565
Chodroff RA, Goodstadt L, Sirey TM, Oliver PL, Davies KE, Green ED, Molnar Z, Ponting CP (2010) Long noncoding RNA genes: conservation of sequence and brain expression among diverse amniotes. Genome Biol 11:R72. doi:10.1186/gb-2010-11-7-r72
Chow JC, Ciaudo C, Fazzari MJ, Mise N, Servant N, Glass JL, Attreed M, Avner P, Wutz A, Barillot E et al (2010) LINE-1 activity in facultative heterochromatin formation during X chromosome inactivation. Cell 141:956–969
Clark BS, Blackshaw S (2014) Long non-coding RNA-dependent transcriptional regulation in neuronal development and disease. Front Genet 5:164. doi:10.3389/fgene.2014.00164
Cox KH, DeLeon DV, Angerer LM, Angerer RC (1984) Detection of mrnas in sea urchin embryos by in situ hybridization using asymmetric RNA probes. Dev Biol 101:485–502
Cushing L, Costinean S, Xu W, Jiang Z, Madden L, Kuang P, Huang J, Weisman A, Hata A, Croce CM (2015) Disruption of miR-29 Leads to Aberrant Differentiation of Smooth Muscle Cells Selectively Associated with Distal Lung Vasculature. PLoS Genet 11:e1005238
de Lencastre A, Pincus Z, Zhou K, Kato M, Lee SS, Slack FJ (2010) MicroRNAs both promote and antagonize longevity in C. elegans. Curr Biol 20:2159–2168
Dellago H, Preschitz-Kammerhofer B, Terlecki-Zaniewicz L, Schreiner C, Fortschegger K, Chang MW, Hackl M, Monteforte R, Kühnel H, Schosserer M et al (2013) High levels of oncomiR-21 contribute to the senescence-induced growth arrest in normal human cells and its knock-down increases the replicative lifespan. Aging Cell 12:446–458
Denas O, Sandstrom R, Cheng Y, Beal K, Herrero J, Hardison RC, Taylor J (2015) Genome-wide comparative analysis reveals human-mouse regulatory landscape and evolution. BMC Genom 16:87. doi:10.1186/s12864-015-1245-6
Deng S, Zhu S, Wang B, Li X, Liu Y, Qin Q, Gong Q, Niu Y, Xiang C, Chen J et al (2014) Chronic pancreatitis and pancreatic cancer demonstrate active epithelial-mesenchymal transition profile, regulated by miR-217-SIRT1 pathway. Cancer Lett 355:184–191
Dey I (2000) A study of chromatin structure and transcription using LINE DNA and simple repeat DNA probes. PhD thesis. School of Life Sciences, Jawaharlal Nehru University, New Delhi
Dey I, Rath PC (2005) A novel rat genomic simple repeat DNA with RNA-homology shows triplex (H-DNA)-like structure and tissue-specific RNA expression. Biochem Biophys Res Commun 327:276–286
Diederichs S (2014) The four dimensions of noncoding RNA conservation. Trends Genet 30:121–123
Dimmeler S, Nicotera P (2013) MicroRNAs in age-related diseases. EMBO Mol Med 5:180–190
Drummond MJ, McCarthy JJ, Sinha M, Spratt HM, Volpi E, Esser KA, Rasmussen BB (2011) Aging and microRNA expression in human skeletal muscle: a microarray and bioinformatics analysis. Physiol Genomics 43:595–603
Faulkner GJ, Kimura Y, Daub CO, Wani S, Plessy C, Irvine KM, Schroder K, Cloonan N, Steptoe AL, Lassmann T et al (2009) The regulated retrotransposon transcriptome of mammalian cells. Nat Genet 41:563–571
Feng J, Bi C, Clark BS, Mady R, Shah P, Kohtz JD (2006) The Evf-2 noncoding RNA is transcribed from the Dlx-5/6 ultraconserved region and functions as a Dlx-2 transcriptional coactivator. Genes Dev 20:1470–1484
Flintoft L (2013) Non-coding RNA: structure and function for lncRNAs. Nat Rev Genet 14:598. doi:10.1038/nrg3561
Flynn RA, Chang HY (2014) Long noncoding RNAs in cell-fate programming and reprogramming. Cell Stem Cell 14:752–761
Gao M, Wang X, Zhang X, Ha T, Ma H, Liu L, Kalbfleisch JH, Gao X, Kao RL, Williams DL (2015) Attenuation of Cardiac Dysfunction in Polymicrobial Sepsis by MicroRNA-146a Is Mediated via Targeting of IRAK1 and TRAF6 Expression. J Immunol 195:672–682
Geiduschek EP, Haselkorn R (1969) Messenger RNA. Annu Rev Biochem 38:647–676
Gibbs RA, Weinstock GM, Metzker ML, Muzny DM, Sodergren EJ, Scherer S, Scott G, Steffen D, Worley KC, Burch PE et al (2004) Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428:493–521
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442:199–202
Goff LA, Groff AF, Sauvageau M, Trayes-Gibson Z, Sanchez-Gomez DB, Morse M, Martin RD, Elcavage LE, Liapis SC, Gonzalez-Celeiro M et al (2015) Spatiotemporal expression and transcriptional perturbations by long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. U. S. A. 112:6855–6862
Gong C, Li Z, Ramanujan K, Clay I, Zhang Y, Lemire-Brachat S, Glass DJ (2015) A long non-coding RNA, LncMyoD, regulates skeletal muscle differentiation by blocking IMP2-mediated mRNA translation. Dev Cell 34:181–191
Gonzalez-Buendia E, Saldana-Meyer R, Meier K, Recillas-Targa F (2015) Transcriptome-wide identification of in vivo interactions between RNAs and RNA-binding proteins by RIP and PAR-CLIP assays. Methods Mol Biol 1288:413–428
Graff J, Mansuy IM (2008) Epigenetic codes in cognition and behaviour. Behav Brain Res 192:70–87
Grillari J, Hackl M, Grillari-Voglauer R (2010) miR-17-92 cluster: ups and downs in cancer and aging. Biogerontology 11:501–506
Grote A, Hiller K, Scheer M, Munch R, Nortemann B, Hempel DC, Jahn D (2005) JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res 33:W526–W531
Guennewig B, Cooper AA (2014) The central role of noncoding RNA in the brain. Int Rev Neurobiol 116:153–194
Guo L, Zhao Y, Yang S, Zhang H, Wu Q, Chen F (2014) An integrated evolutionary analysis of miRNA-lncRNA in mammals. Mol Biol Rep 41:201–207
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP et al (2009) Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458:223–227
Hackl M, Brunner S, Fortschegger K, Schreiner C, Micutkova L, Mück C, Laschober GT, Lepperdinger G, Sampson N, Berger P (2010) miR-17, miR-19b, miR-20a, and miR-106a are down-regulated in human aging. Aging Cell 9:291–296
Hamrick MW, Herberg S, Arounleut P, He HZ, Shiver A, Qi RQ, Zhou L, Isales CM, Mi QS (2010) The adipokine leptin increases skeletal muscle mass and significantly alters skeletal muscle miRNA expression profile in aged mice. Biochem Biophys Res Commun 400:379–383
Hoffman Y, Pilpel Y, Oren M (2014) microRNAs and Alu elements in the p53-Mdm2-Mdm4 regulatory network. J Mol Cell Biol 6:192–197
Hu G, Lou Z, Gupta M (2014) The long non-coding RNA GAS5 cooperates with the eukaryotic translation initiation factor 4E to regulate c-Myc translation. PLoS ONE 9:e107016
Hu J, Chen Z, Xia D, Wu J, Xu H, Ye ZQ (2012) Promoter-associated small double-stranded RNA interacts with heterogeneous nuclear ribonucleoprotein A2/B1 to induce transcriptional activation. Biochem J 447:407–416
Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D, Khalil AM, Zuk O, Amit I, Rabani M et al (2010) A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142:409–419
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, Barrette TR, Prensner JR, Evans JR, Zhao S et al (2015) The landscape of long noncoding RNAs in the human transcriptome. Nat Genet 47:199–208
Jazbutyte V, Fiedler J, Kneitz S, Galuppo P, Just A, Holzmann A, Bauersachs J, Thum T (2013) MicroRNA-22 increases senescence and activates cardiac fibroblasts in the aging heart. Age 35:747–762
Jiang L, Wang C, Lei F, Zhang L, Zhang X, Liu A, Wu G, Zhu J, Song L (2015) miR-93 promotes cell proliferation in gliomas through activation of PI3K/Akt signalling pathway. Oncotarget 6:8286–8299
Johnson R (2012) Long non-coding RNAs in Huntington’s disease neurodegeneration. Neurobiol Dis 46:245–254
Johnsson P, Lipovich L, Grander D, Morris KV (2014) Evolutionary conservation of long non-coding RNAs; sequence, structure, function. Biochim Biophys Acta 1840:1063–1071
Kadakkuzha BM, Liu XA, McCrate J, Shankar G, Rizzo V, Afinogenova A, Young B, Fallahi M, Carvalloza AC, Raveendra B et al (2015) Transcriptome analyses of adult mouse brain reveal enrichment of lncRNAs in specific brain regions and neuronal populations. Front Cell Neurosci 9:63. doi:10.3389/fncel.2015.00063
Kapusta A, Kronenberg Z, Lynch VJ, Zhuo X, Ramsay L, Bourque G, Yandell M, Feschotte C (2013) Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. PLoS Genet 9:e1003470
Karp X, Hammell M, Ow MC, Ambros V (2011) Effect of life history on microRNAs expression during C. elegans development. RNA 17:639–651
Kato M, Chen X, Inukai S, Zhao H, Slack FJ (2011) Age-associated changes in expression of small, noncoding RNAs, including microRNAs, in C. elegans. RNA 17:1804–1820
Kawaji H, Lizio M, Itoh M, Kanamori-Katayama M, Kaiho A, Nishiyori-Sueki H, Shin JW, Kojima-Ishiyama M, Kawano M, Murata M et al (2014) Comparison of CAGE and RNA-seq transcriptome profiling using clonally amplified and single-molecule next-generation sequencing. Genome Res 24:708–717
Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A et al (2009) Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 106:11667–11672
Khanna A, Muthusamy S, Liang R, Sarojini H, Wang E (2011) Gain of survival signalling by down-regulation of three key miRNAs in brain of calorie-restricted mice. Aging 3:223–236
Kim K, Vinayagam A, Perrimon N (2014) A rapid genome-wide microRNA screen identifies miR-14 as a modulator of Hedgehog signaling. Cell Rep 7:2066–2077
Kim DH, Marinov GK, Pepke S, Singer ZS, He P, Williams B, Schroth GP, Elowitz MB, Wold BJ (2015) Single-cell transcriptome analysis reveals dynamic changes in lncRNA expression during reprogramming. Cell Stem Cell 16:88–101
Kour S (2015) Functional characterization of the noncoding RNAs from the repetitive DNA of mammalian genome. Jawaharlal Nehru University, New Delhi, PhD
Kour S, Rath PC (2015) Age-dependent differential expression profile of a novel intergenic long noncoding RNA in rat brain. Int J Dev Neurosci 47:286–297
Kour S, Rath PC (2016a) Age-related expression of a repeat-rich intergenic long noncoding RNA in the rat brain. Mol Neurobiol [Epub ahead of print]
Kour S, Rath PC (2016b) All-trans retinoic acid induces expression of a novel intergenic long noncoding RNA in adult rat primary hippocampal neurons. J Mol Neurosci 58:266–276
Kour S, Rath PC (2016c) Long noncoding RNAs in aging and age-related diseases. Ageing Res Rev 26:1–21
Kurokawa R (2011) Promoter-associated long noncoding RNAs repress transcription through a RNA binding protein TLS. Adv Exp Med Biol 722:196–208
Hyun S, Lee JH, Jin H, Nam J, Namkoong B, Lee G, Chung J, Kim VN (2009) Conserved MicroRNA miR-8/miR-200 and its target USH/FOG2 control growth by regulating PI3K. Cell 139:1096–1108
Lasalle JM, Powell WT, Yasui DH (2013) Epigenetic layers and players underlying neurodevelopment. Trends Neurosci 36:460–470
Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP, Kingston RE (2006) Characterization of the piRNA complex from rat testes. Science 313:363–367
Lee JT (2011) Gracefully ageing at 50, X-chromosome inactivation becomes a paradigm for RNA and chromatin control. Nat Rev Mol Cell Biol 12:815–826
Legnini I, Morlando M, Mangiavacchi A, Fatica A, Bozzoni I (2014) A feedforward regulatory loop between HuR and the long noncoding RNA linc-MD1 controls early phases of myogenesis. Mol Cell 53:506–514
Li N, Muthusamy S, Liang R, Sarojini H, Wang E (2011a) Increased expression of miR-34a and miR-93 in rat liver during aging, and their impact on the expression of Mgst1 and Sirt1. Mech Ageing Dev 132:75–85
Li X, Khanna A, Li N, Wang E (2011b) Circulatory miR34a as an RNA based, noninvasive biomarker for brain aging. Aging 3:985–1002
Li N, Bates DJ, An J, Terry DA, Wang E (2011c) Up-regulation of key microRNAs, and inverse down-regulation of their predicted oxidative phosphorylation target genes, during aging in mouse brain. Neurobiol Aging 32:944–955
Liang R, Khanna A, Muthusamy S, Li N, Sarojini H, Kopchick JJ, Masternak MM, Bartke A, Wang E (2011) Post-transcriptional regulation of IGF1R by key microRNAs in long-lived mutant mice. Aging Cell 10:1080–1088
Liang C, Forrest AR, Wagner GP (2015) The statistical geometry of transcriptome divergence in cell-type evolution and cancer. Nat Commun 6:6066. doi:10.1038/ncomms7066
Lin N, Chang KY, Li Z, Gates K, Rana ZA, Dang J, Zhang D, Han T, Yang CS, Cunningham TJ et al (2014a) An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Mol Cell 53:1005–1019
Lin ST, Heng MY, Ptacek LJ, Fu YH (2014b) Regulation of Myelination in the Central Nervous System by Nuclear Lamin B1 and Non-coding RNAs. Transl Neurodegener 3:4. doi:10.1186/2047-9158-3-4
Liu N, Landreh M, Cao K, Abe M, Hendriks GJ, Kennerdell JR, Zhu Y, Wang LS, Bonini NM (2012) The microRNA miR-34 modulates ageing and neurodegeneration in Drosophila. Nature 482:519–523
Lizio M, Harshbarger J, Shimoji H, Severin J, Kasukawa T, Sahin S, Abugessaisa I, Fukuda S, Hori F, Ishikawa-Kato S et al (2015) Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol 16:22. doi:10.1186/s13059-014-0560-6
Loewer S, Cabili MN, Guttman M, Loh YH, Thomas K, Park IH, Garber M, Curran M, Onder T, Agarwal S et al (2010) Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet 42:1113–1117
Lucanic M, Graham J, Scott G, Bhaumik D, Benz CC, Hubbard A, Lithgow GJ, Melov S (2013) Age-related micro-RNA abundance in individual C. elegans. Aging 5:394–411
Lv J, Liu H, Huang Z, Su J, He H, Xiu Y, Zhang Y, Wu Q (2013) Long non-coding RNA identification over mouse brain development by integrative modeling of chromatin and genomic features. Nucleic Acids Res 41:10044–10061
Maes OC, An J, Sarojini H, Wang E (2008) Murine microRNAs implicated in liver functions and aging process. Mech Ageing Dev 129:534–541
Marasa BS, Srikantan S, Masuda K, Abdelmohsen K, Kuwano Y, Yang X, Martindale JL, Rinker-Schaeffer CW, Gorospe M (2009). Increased MKK4 abundance with replicative senescence is linked to the joint reduction of multiple microRNAs. Sci Signal 2:ra69
Marchese FP, Huarte M (2014) Long non-coding RNAs and chromatin modifiers: their place in the epigenetic code. Epigenetics 9:21–26
Mariño G, Ugalde AP, Fernández AF, Osorio FG, Fueyo A, Freije JM, López-Otín C (2010) Insulin-like growth factor 1 treatment extends longevity in a mouse model of human premature aging by restoring somatotroph axis function. Proc Natl Acad Sci USA 107:16268–16273
Martinez I, Cazalla D, Almstead LL, Steitz JA, DiMaio D (2011) miR-29 and miR-30 regulate B-Myb expression during cellular senescence. Proc Natl Acad Sci USA 108:522–527
Marques AC, Ponting CP (2014) Intergenic lncRNAs and the evolution of gene expression. Curr Opin Genet Dev 27:48–53
Mattick JS, Rinn JL (2015) Discovery and annotation of long noncoding RNAs. Nat Struct Mol Biol 22:5–7
McHugh CA, Chen CK, Chow A, Surka CF, Tran C, McDonel P, Pandya-Jones A, Blanco M, Burghard C, Moradian A et al (2015) The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature. doi:10.1038/nature14443
Menghini R, Casagrande V, Cardellini M, Martelli E, Terrinoni A, Amati F, Vasa-Nicotera M, Ippoliti A, Novelli G, Melino G et al (2009) MicroRNA 217 modulates endothelial cell senescence via silent information regulator 1. Circulation 120:1524–1532
Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS (2008) Specific expression of long noncoding RNAs in the mouse brain. Proc Natl Acad Sci USA 105:716–721
Mercer TR, Mattick JS (2013) Structure and function of long noncoding RNAs in epigenetic regulation. Nat Struct Mol Biol 20:300–307
Mercer TR, Qureshi IA, Gokhan S, Dinger ME, Li G, Mattick JS, Mehler MF (2010) Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci 11:14. doi:10.1186/1471-2202-11-14
Meza-Sosa KF, Pedraza-Alva G, Pérez-Martínez L (2014) MicroRNAs: key triggers of neuronal cell fate. Front Cell Neurosci 8:175
Milligan MJ, Lipovich L (2014) Pseudogene-derived lncRNAs: emerging regulators of gene expression. Front Genet 5:476. doi:10.3389/fgene.2014.00476
Mishra RR (2009) Expression and functional characterization of candidate cDNAs isolated by a simple repeat DNA probe. Ph.D thesis. School of Life Sciences, Jawaharlal Nehru University, New Delhi
Mogilyansky E, Rigoutsos I (2013) The miR-17/92 cluster: a comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ 20:1603–1614
Mortimer SA, Kidwell MA, Doudna JA (2014) Insights into RNA structure and function from genome-wide studies. Nat Rev Genet 15:469–479
Nelson C, Ambros V, Baehrecke EH (2014) miR-14 regulates autophagy during developmental cell death by targeting ip3-kinase 2. Mol Cell 56:376–388
Ng SY, Bogu GK, Soh BS, Stanton LW (2013) The long noncoding RNA RMST interacts with SOX2 to regulate neurogenesis. Mol Cell 51:349–359
Noren Hooten N, Abdelmohsen K, Gorospe M, Ejiogu N, Zonderman AB, Evans MK (2010) MicroRNA expression patterns reveal differential expression of target genes with age. PLoS One 5:e10724
Olive V, Minella AC, He L (2015) Outside the coding genome, mammalian microRNAs confer structural and functional complexity. Sci Signal 8, re2. doi:10.1126/scisignal.2005813
Persengiev S, Kondova I, Otting N, Koeppen AH, Bontrop RE (2011) Genome-wide analysis of miRNA expression reveals a potential role for miR-144 in brain aging and spinocerebellar ataxia pathogenesis. Neurobiol Aging 32:2316.e17–27
Philippe JV, Ayadi L, Branlant C, Behm-Ansmant I (2015) Probing small non-coding RNAs structures. Methods Mol Biol 1296:119–136
Pincus Z, Smith-Vikos T, Slack FJ (2011) MicroRNA predictors of longevity in Caenorhabditis elegans. PLoS Genet 7(9):e1002306
Pnueli L, Rudnizky S, Yosefzon Y, Melamed P (2015) RNA transcribed from a distal enhancer is required for activating the chromatin at the promoter of the gonadotropin alpha-subunit gene. Proc Natl Acad Sci USA 112:4369–4374
Ponjavic J, Ponting CP (2007) The long and the short of RNA maps. BioEssays 29:1077–1080
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641
Porro A, Feuerhahn S, Lingner J (2014) TERRA-reinforced association of LSD1 with MRE11 promotes processing of uncapped telomeres. Cell Rep 6:765–776
Quek XC, Thomson DW, Maag JL, Bartonicek N, Signal B, Clark MB, Gloss BS, Dinger ME (2015) lncRNAdb v2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res 43:D168–D173
Ramos AD, Andersen RE, Liu SJ, Nowakowski TJ, Hong SJ, Gertz CC, Salinas RD, Zarabi H, Kriegstein AR, Lim DA (2015) The long noncoding RNA Pnky regulates neuronal differentiation of embryonic and postnatal neural stem cells. Cell Stem Cell 16:439–447
Rapicavoli NA, Qu K, Zhang J, Mikhail M, Laberge RM, Chang HY (2013) A mammalian pseudogene lncRNA at the interface of inflammation and anti-inflammatory therapeutics. Elife 2:e00762
Redon S, Zemp I, Lingner J (2013) A three-state model for the regulation of telomerase by TERRA and hnRNPA1. Nucleic Acids Res 41:9117–9128
Rinn JL (2014) lncRNAs: linking RNA to chromatin. Cold Spring Harb Perspect Biol 6. doi:10.1101/cshperspect.a018614
Robert L, Labat-Robert J, Robert AM (2010) Genetic, epigenetic and posttranslational mechanisms of aging. Biogerontology 11:387–399
Roberts TC, Morris KV, Weinberg MS (2014a) Perspectives on the mechanism of transcriptional regulation by long non-coding RNAs. Epigenetics 9:13–20
Roberts TC, Morris KV, Wood MJ (2014b) The role of long non-coding RNAs in neurodevelopment, brain function and neurological disease. Philos Trans R Soc Lond B Biol Sci 369. doi:10.1098/rstb.2013.0507
Rokavec M, Li H, Jiang L, Hermeking H (2014) The p53/miR-34 axis in development and disease. J Mol Cell Biol 6:214–230
Rutledge CE, Lau HT, Mangan H, Hardy LL, Sunnotel O, Guo F, MacNicol AM, Walsh CP, Lees-Murdock DJ (2014) Efficient translation of Dnmt1 requires cytoplasmic polyadenylation and Musashi binding elements. PLoS ONE 9:e88385
Saksouk N, Barth TK, Ziegler-Birling C, Olova N, Nowak A, Rey E, Mateos-Langerak J, Urbach S, Reik W, Torres-Padilla ME et al (2014) Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Mol Cell 56:580–594
Sambrook JRD (2001) Molecular Cloning. Cold Spring Harbour Press, New York, A laboratory manual
Serna E, Gambini J, Borras C, Abdelaziz KM, Belenguer A, Sanchis P, Avellana JA, Rodriguez-Manas L, Vina J (2012) Centenarians, but not octogenarians, up-regulate the expression of microRNAs. Sci Rep 2:961. doi:10.1038/srep00961
Shahryari A, Jazi MS, Samaei NM, Mowla SJ (2015) Long non-coding RNA SOX2OT: expression signature, splicing patterns, and emerging roles in pluripotency and tumorigenesis. Front Genet 6:196. doi:10.3389/fgene.2015.00196
Shi Z, Hayes G, Ruvkun G (2013) Dual regulation of the lin-14 target mRNA by the lin-4 miRNA. PLoS One 8:e75475
Sigdel KR, Cheng A, Wang Y, Duan L, Zhang Y (2015) The emerging functions of long noncoding RNA in immune cells: autoimmune diseases. J Immunol Res 2015:848790. doi:10.1155/2015/848790
Siggens L, Ekwall K (2014) Epigenetics, chromatin and genome organization: recent advances from the ENCODE project. J Intern Med 276:201–214
Singh DK, Rath PC (2012) Long interspersed nuclear elements (LINEs) show tissue-specific, mosaic genome and methylation-unrestricted, widespread expression of noncoding RNAs in somatic tissues of the rat. RNA Biol 9:1380–1396
Singh M (2012) Dysregulated A to I RNA editing and non-coding RNAs in neurodegeneration. Front Genet 3:326. doi:10.3389/fgene.2012.00326
Sinha JK, Ghosh S, Swain U, Giridharan NV, Raghunath M (2014) Increased macromolecular damage due to oxidative stress in the neocortex and hippocampus of WNIN/Ob, a novel rat model of premature aging. Neuroscience 269:256–264
Smith-Vikos T, Slack FJ (2012) MicroRNAs and their roles in aging. J Cell Sci 125:7–17
Somarowthu S, Legiewicz M, Chillon I, Marcia M, Liu F, Pyle AM (2015) HOTAIR forms an intricate and modular secondary structure. Mol Cell 58:353–361
Song R, Ro S, Michaels JD, Park C, McCarrey JR, Yan W (2009) Many X-linked microRNAs escape meiotic sex chromosome inactivation. Nat Genet 41:488–493
Soumillon M, Necsulea A, Weier M, Brawand D, Zhang X, Gu H, Barthes P, Kokkinaki M, Nef S, Gnirke A et al (2013) Cellular source and mechanisms of high transcriptome complexity in the mammalian testis. Cell Rep 3:2179–2190
St Laurent G, Wahlestedt C, Kapranov P (2015) The Landscape of long noncoding RNA classification. Trends Genet 31:239–251
Sutherland JM, McLaughlin EA, Hime GR, Siddall NA (2013) The Musashi family of RNA binding proteins: master regulators of multiple stem cell populations. Adv Exp Med Biol 786:233–245
Takahashi M, Eda A, Fukushima T, Hohjoh H (2012) Reduction of type IV collagen by upregulated miR-29 in normal elderly mouse and klotho-deficient, senescence-model mouse. PLoS One 711:e48974
Trümbach D, Prakash N (2015) The conserved miR-8/miR-200 microRNA family and their role in invertebrate and vertebrate neurogenesis. Cell Tissue Res 359:161–177
Toledano H, D'Alterio C, Czech B, Levine E, Jones DL (2012) The let-7-Imp axis regulates ageing of the Drosophila testis stem-cell niche. Nature 485:605–610
Turner MJ, Jiao AL, Slack FJ (2014) Autoregulation of lin-4 microRNA transcription by RNA activation (RNAa) in C. elegans. Cell Cycle 13:772–781
Ugalde AP, Ramsay AJ, de la Rosa J, Varela I, Mariño G, Cadiñanos J, Lu J, Freije JM, López-Otín C (2011) Aging and chronic DNA damage response activate a regulatory pathway involving miR-29 and p53. EMBO J 30:2219–2232
van Almen GC, Verhesen W, van Leeuwen RE, van de Vrie M, Eurlings C, Schellings MW, Swinnen M, Cleutjens JP, van Zandvoort MA, Heymans S et al (2011) MicroRNA-18 and microRNA-19 regulate CTGF and TSP-1 expression in age-related heart failure. Aging Cell 10:769–779
van Balkom BW, de Jong OG, Smits M, Brummelman J, den Ouden K, de Bree PM, van Eijndhoven MA, Pegtel DM, Stoorvogel W, Wurdinger T, et al (2013) Endothelial cells require miR-214 to secrete exosomes that suppress senescence and induce angiogenesis in human and mouse endothelial cells. Blood 121:3997–4006, S3991–3915
Vasa-Nicotera M, Chen H, Tucci P, Yang AL, Saintigny G, Menghini R, Mahè C, Agostini M, Knight RA, Melino G et al (2011) miR-146a is modulated in human endothelial cell with aging. Atherosclerosis 217:326–330
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG, Smith HO, Yandell M, Evans CA, Holt RA et al (2001) The sequence of the human genome. Science 291:1304–1351
Vucicevic D, Corradin O, Ntini E, Scacheri PC, Orom UA (2015) Long ncRNA expression associates with tissue-specific enhancers. Cell Cycle 14:253–260
Wahlestedt C (2013) Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat Rev Drug Discov 12:433–446
Wan DY, Zhang Z, Yang HH (2015) Cardioprotective effect of miR-214 in myocardial ischemic postconditioning by down-regulation of hypoxia inducible factor 1, alpha subunit inhibitor. Cell Mol Biol 61:1–6
Wang WX, Wilfred BR, Madathil SK, Tang G, Hu Y, Dimayuga J, Stromberg AJ, Huang Q, Saatman KE, Nelson PT (2010) miR-107 regulates granulin/progranulin with Implications for traumatic brain injury and neurodegenerative disease. Am J Pathol 177:334–345
Wang KC, Chang HY (2011) Molecular mechanisms of long noncoding RNAs. Mol Cell 43:904–914
Wang L, Zhao Y, Bao X, Zhu X, Kwok YK, Sun K, Chen X, Huang Y, Jauch R, Esteban MA et al (2015) LncRNA Dum interacts with Dnmts to regulate Dppa2 expression during myogenic differentiation and muscle regeneration. Cell Res 25:335–350
Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P et al (2002) Initial sequencing and comparative analysis of the mouse genome. Nature 420:520–562
Watson JD, Crick FH (1953) Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature 171:737–738
Weichenhan D, Plass C (2013) The Evolving Epigenome. Hum Mol Genet 22:R1–R6
Willingham AT, Orth AP, Batalov S, Peters EC, Wen BG, Aza-Blanc P, Hogenesch JB, Schultz PG (2005) A strategy for probing the function of noncoding RNAs finds a repressor of NFAT. Science 309:1570–1573
Wood EJ, Chin-Inmanu K, Jia H, Lipovich L (2013) Sense-antisense gene pairs: sequence, transcription, and structure are not conserved between human and mouse. Front Genet 4:183. doi:10.3389/fgene.2013.00183
Xu D, Takeshita F, Hino Y, Fukunaga S, Kudo Y, Tamaki A, Matsunaga J, Takahashi RU, Takata T, Shimamoto A et al (2011) miR-22 represses cancer progression by inducing cellular senescence. J Cell Biol 193:409–424
Xue Y, Gu D, Ma G, Zhu L, Hua Q, Chu H, Tong N, Chen J, Zhang Z, Wang M (2015) Genetic variants in lncRNA HOTAIR are associated with risk of colorectal cancer. Mutagenesis 30:303–310
Yang G, Lu X, Yuan L (2014) LncRNA: a link between RNA and cancer. Biochim Biophys Acta 1839:1097–1109
Yu X, Zhang L, Wen G, Zhao H, Luong LA, Chen Q, Huang Y, Zhu J, Ye S, Xu Q, Wang W, Xiao Q (2015) Upregulated sirtuin 1 by miRNA-34a is required for smooth muscle cell differentiation from pluripotent stem cells. Cell Death Differ 22:1170–1180
Zaramela LS, Vencio RZ, ten-Caten F, Baliga NS, Koide T, 2014. Transcription start site associated RNAs (TSSaRNAs) are ubiquitous in all domains of life. PLoS One 9, e107680
Zhang X, Zabinsky R, Teng Y, Cui M, Han M (2011) MicroRNAs play critical roles in the survival and recovery of Caenorhabditis elegans from starvation-induced L1 diapause. Proc Natl Acad Sci USA 108:17997–18002
Zhao H, Wen G, Huang Y, Yu X, Chen Q, Afzal TA, Luong le A, Zhu J, Ye S, Zhang L et al (2015) MicroRNA-22 regulates smooth muscle cell differentiation from stem cells by targeting methyl CpG-binding protein 2. Arterioscler Thromb Vasc Biol 35:918–929
Zheng Y, Xu Z (2014) MicroRNA-22 induces endothelial progenitor cell senescence by targeting AKT3. Cell Physiol Biochem 34:1547–1555
Ziats MN, Rennert OM (2013) Aberrant expression of long noncoding RNAs in autistic brain. J Mol Neurosci 49:589–593
Acknowledgments
We thank financial support from the University Grants Commission (UGC)-Research Networking Resource Centre, Department of Science and Technology (DST)-FIST & PURSE to the School of Life Sciences (SLS), Jawaharlal Nehru University (JNU) & PCR and the UGC-Junior & Senior Research Fellowship to SK for Ph.D.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media Singapore
About this chapter
Cite this chapter
Kour, S., Rath, P.C. (2017). Differential Expression of Long Noncoding RNA in the Rat Brain During Aging. In: Rath, P., Sharma, R., Prasad, S. (eds) Topics in Biomedical Gerontology. Springer, Singapore. https://doi.org/10.1007/978-981-10-2155-8_10
Download citation
DOI: https://doi.org/10.1007/978-981-10-2155-8_10
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-2154-1
Online ISBN: 978-981-10-2155-8
eBook Packages: MedicineMedicine (R0)