Abstract
Increasing evidence suggests that non-coding RNAs have multiple important roles in transcriptional regulation, and also contribute to the expansion of genome complexity. Here, we attempted to perform an integrative analysis of miRNA–lncRNA in mammals to reveal their potential evolutionary relationships in transcriptional regulation network. The lncRNA of H19 gene includes an internal small non-coding RNA, miR-675. The H19 gene was conserved as well as the neighbor Igf2 coding gene and internal miR-675 gene, although they were involved in various location distributions and phylogenetic relationships. Both miR-675-5p/3p have been reported as potential negative regulatory molecules, but the canonical miRNA was more conserved than the passenger strand. These results implicated that the imprinted coding and non-coding gene cluster indicated similar evolutionary patterns. The lncRNA and internal miRNAs may have versatile roles in multiple biological processes, including in tumorigenesis as potential ncRNA regulatory molecules. The phylogenetical conservation of the imprinted cluster may be also derived from the functional implications and evolutionary pressures. The study will enrich the study of potential crosstalk of miRNA–lncRNA and mRNA–ncRNA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The entire eukaryotic genome is pervasively transcribed and yields to a myriad of non-protein-coding RNA species with complex expression and regulation patterns [1]. These non-coding RNAs (ncRNAs) contribute to evolutionary processes of the genome complexity with increasing amounts of information [2–5]. They have multiple roles in biological processes, including occupy critical edges and nodes in physiological networks, frequently involved in feedback loops [6]. The function of them is more difficult and complexity than we thought, but they will contribute to complex genomic pervasive transcription on cell biology and evolution [7].
As non-protein-coding sequences, most ncRNAs tends to be only weakly constrained than mRNAs through evolutionary processes. The potential function of ncRNA is still enigmatic, although functional ncRNAs are identified as regulatory molecules and play roles in multiple biological processes, such as a class of small (~22-nts) ncRNAs, microRNAs (miRNAs). miRNAs have been identified as crucial regulators of gene expression in multiple physiological and pathological processes [8–12]. Recently, long non-coding RNAs (lncRNAs), a large proportion of non-coding transcripts (longer than 200-nts), have been studied as another class of ncRNAs. The lncRNAs may also play a role in tumorigenesis, such as H19 [13–15]. Indeed, dysregulation of many ncRNAs contribute to pathological processes via versatile functional roles. Different ncRNAs may have close relationships according to their location distributions and sequence similarity. For example, H19 is a host gene of miR-675 and imprinted and endogenously expressed, and H19 can function as pri-miRNA of miR-675 [16, 17].
LncRNAs and miRNAs are widely studied because of their crucial regulatory roles in biological processes and evolutionary players, especially for the well-conserved small miRNAs. Although they are characterized as ncRNAs and negative regulatory molecules, they are thought to have different evolutionary patterns. Here, to further understand evolutionary and crosstalk of miRNA–lncRNA, we performed an integrated analysis of miR-675 and Hg19 genes across different mammalians. Simultaneously, the neighboring Igf2 gene of H19 was also involved in the integrative analysis. The study will enrich the ncRNA study, especially for the potential crosstalk of miRNA–lncRNA and mRNA–ncRNA, which provides potential interaction in the coding-non-coding RNA regulatory network.
Results
The long non-coding RNA, H19 gene, was located on different chromosomes with various length distributions across mammalians (Table 1). The lncRNA included the small non-coding RNA, mir-675 gene. Although both of H19 and mir-675 were identified as ncRNAs, they showed inconsistent nucleotide divergence and evolutionary patterns. Compared to small miRNAs, H19 was involved in higher levels of nucleotide transition/transversion, insertion/deletion across different animal species (Table 2). The hypothesized pri-miRNA sequences showed similar nucleotide divergence pattern with H19, while both pre-miRNA and mature miRNAs were well-conserved phylogenetically as expected. Indeed, compared to other broadly conserved miRNA gene families (such as let-7 gene family), the miR-675 was identified as a poorly conserved and mammalian-specific gene family.
Nucleotide compositions indicated that guanine (G) was the dominant nucleotide in the lncRNA, while cytosine (C) was the most dominant nucleotide in pri-miRNA (Fig. 1a, b). Interestingly, distributions of nucleotides in mouse and rat were inconsistent with other animals based on H19 and hypothesized pri-mir-675, but the diverse distributions were moderated in pre-mir-675. The mature small RNAs showed inconsistent nucleotide compositions with other longer sequences, although miR-675-5p/3p were the part sequences of them (Fig. 1). The two mature miRNAs showed inverse nucleotide compositions, which ensured the stem-loop structure of pre-miRNA.
Frequency distributions of nucleotide composition. Frequency distributions of nucleotides (A, T(U), C and G across different animal species; distributions of average nucleotide pair frequency; percentage of average nucleotide pair frequency) of H19 (a), hypothesized pri-mir-675 (b), pre-mir-675 (c), miR-675-5p (d) and miR-675-3p (e) sequences. H19 and pri-mir-675 show consistent distribution patterns, and Rodentia (mmu and rno) restricts divergence of nucleotide composition than other mammals
Reconstructed phylogenetic trees showed that pre-mir-675 showed inconsistent phylogenetic relationships between H19 and pri-mir-675 sequences (Fig. 2, Fig. S1). No significant divergence could be detected between trees of H19 and pri-mir-675. The evolutionary analysis of H19 and pri-mir-675 indicated that mouse (mmu) and rat (rno) had larger genetic distances with other animal species, but the rodentians were clustered together with primates in evolutionary relationships of pre-mir-675. All the precursor miRNAs showed stable stem-loop structures with various minimum folding free energy (MFE) and loop regions (Table S1). Although both the two arms could yield potential negative regulatory molecules, miR-675-3p was involved in higher level of nucleotide divergence than miR-675-5p (Fig. 3). miR-675-5p was identified as the original annotated mature miRNA, and had higher or dominant expression level than its partner [16]. Phylogenetic networks indicated that miR-675-3p had more complex evolutionary pattern, and was involved in more median vectors (hypothesized sequence types in reconstructed network) (Fig. 3b).
Phylogenetic relationships based on different sequences. a NJ tree of H19 sequence. b NJ tree of hypothesized primary mir-675 sequences (pri-675). c NJ tree of precursor miR-675 sequences (pre-675). Phylogenetic tree of pre-675 sequences show inconsistent distributions with H19 and pri-675 sequences
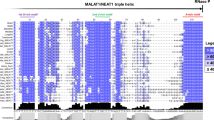
Sequence diversity and phylogenetic networks of mir-675. a Multiple sequence alignment of mir-675 and the two mature miRNAs (miR-675-5p and miR-675-3p). Compared to other well-conserved miRNAs, miR-675 sequences are poorly conserved across different animal species. They are always involved in larger genetic distances via nucleotide divergences. b The two products show inconsistent evolutionary patterns
Further, according to predicted target mRNAs of miR-675-5p/3p, dog (cfa) and cow (bta) showed more private or specific target mRNAs, although other species shared more common targets (Table S2). Strikingly, the ever thought as passenger strand, miR-675-3p, indicated more predicted potential target mRNAs than miR-675-5p. Functional enrichment analysis suggested that some target mRNAs were involved in important biological processes, such as lysine degradation, taurine and hypotaurine metabolism, valine, leucine and isoleucine biosynthesis in human, and etc. (Table S2). These results indicated that miR-675 played a versatile biological role via negatively regulating multiple target mRNAs.
Moreover, to understand the evolutionary pattern of host gene of lncRNA, we simultaneously analyzed the Igf2 gene. The Igf2 gene might be located in the upstream region or downstream of the H19 gene across mammalians (Tables 1, 3). Compared to related ncRNAs, including lncRNA and miRNA, protein-coding genes were also involved in higher level of nucleotide substitutions, insertions/deletions (Fig. S2; Table S3). The Igf2 gene showed similar phylogenetic relationships with pre-miRNAs and mature miRNAs (Figs. S1, S2).
Discussion
Non-protein-coding RNAs (ncRNAs), including miRNAs and lncRNAs, are major players that increase organismal complexity as robust regulatory molecules. These regulatory RNA species are involved in complex overlapped expression and regulation patterns [1], which contributes to coding-non-coding RNA regulatory network. Deregulated expressed ncRNAs may also play a role in physiological and pathological processes, even in tumorigenesis.
The longer (lncRNA) and small (miRNA) ncRNAs may be involved in potential relationships. For example, some lncRNAs have been identified as potential pri-miRNAs to yield mature miRNAs, such as H19 gene [16, 17]. The lncRNA is imprinted with maternal expression and plays a role in tumorigenesis [13–15], and simultaneously it is also identified as a primary transcript of miR-675 [16, 17]. Although distributions of average nucleotide pair frequency in H19, hypothesized pri-mir-675, pre-miRNA and mature miR-675 are quite similar, while frequency distributions of nucleotides are various (Fig. 1). The diversity indicates the bias of nucleotide compositions with different sequence lengths, which may contribute to functional need as well as nucleotide compositions in miRNAs. H19 contains miR-675, which suggests close relationships between miRNA and lncRNA. The phenomenon is not random, and other imprinted ncRNAs might also give rise to miRNAs [16]. The interesting relationships may increase cross-talk between different ncRNAs, and further enrich coding-non-coding RNA regulatory network via their versatile biological roles.
Although the miRNA is located in the H19 gene, phylogenetic relationships based on H19, hypothesized pri-miRNA and pre-miRNA show various distributions, especially for between pre-miRNA and pri-miRNA/H19 (Figs. 1, 2 and Fig. S1). The longer sequences are always prone to be involved in more nucleotide substitutions, insertion/deletion (Table 2), although some special regions are well-conserved (Fig. S3). We found that internal miRNA do not show special conserved region than other regions, except for special nucleotide compositions (Fig. 1). Indeed, the miRNA is a mammalian-specific species according to the annotated miRNAs in the miRBase database (Release 19.0), and it has been identified as a poorly conserved gene family (Fig. 3). Both miR-675-5p and miR-675-3p can be detected nucleotide divergence across mammalians, even though in the nucleotides 2–8 (the typical “seed sequences” of miRNA) (Fig. 3a). The inconsistent “seed sequences” lead to various target mRNAs, which simultaneously enriches the more potential versatile biological roles of the miRNA family. The two mature miRNAs also show specific evolutionary networks, and miR-675-3p is involved in more nucleotide substitutions and median vectors (Table; Fig. 3b). These results implicates that the two miRNAs are differentially evolved that is mainly derived from the functional pressures. As the reported canonical miRNA sequence, miR-675-5p is dominantly expressed as a crucial regulatory molecule, but its partner is always rarely expressed although it is also potential active regulatory molecule. The two miRNAs may have important roles in multiple biological processes via interacting with their potential target mRNAs (Table S2).
Imprinted genes in mammalian are always clustered with lncRNAs [18]. The imprinting mechanism of Igf2-H19 is conserved in therians, but the distributions can be flexible across different animal species (Tables 1, 3). Between the coding and non-coding genes, Igf2 and H19 show similar evolutionary patterns, although lncRNAs are always thought to be poorly conserved (Table 2; Figs. 1, 2, Figs. S1, S2; Table S3). The main reasons may be derived from the imprinting mechanism and functional implication and evolutionary pressure. Compared to the two longer coding and non-coding sequences, the internal miRNA, is also well-conserved. However, compared to other broadly conserved miRNAs, the mammalian-specific miRNA species is identified as a poorly conserved gene family. On contrast, the H19 lncRNA is conserved in evolutionary process, although the lncRNAs are always thought as poorly conserved RNA molecules. The H19 and mir-675 show inconsistent distributions of the MFE structures, despite both of them show mountain plots (Fig. S4). Specially, although both of the two arms of mir-675 can yield mature miRNAs (miR-675-5p/3p), the two regulatory molecules show inconsistent evolutionary patterns (Fig. 3). The identified as canonical miRNA (miR-675-5p) is more conserved than ever thought as passenger strand of miR-675-3p. Accumulating evidences have shown that miRNA passenger strands, also termed miRNA* sequences, may be potential regulatory molecules and have potential biological roles in specific developmental stages [19–23]. However, perhaps because of dynamic and temporal functional need, miRNA* sequences are always involved in rapid evolutionary processes inter- and extra-species [21]. Functional analysis based on predicted target mRNAs also suggests versatile biological roles of miR-675 (Table S2). Collectively, the evolutionary process of imprinted cluster may be slightly diverged that is mainly derived from functional implication, although all of them are well-conserved across complex evolutionary patterns inter- and extra-species.
Materials and methods
All the H19 sequences in the study were obtained from the GenBank (Table 1). The pre-miRNA of miR-675 were collected from the miRBase database (Release 19.0, http://www.mirbase.org/) [24]. Although H19 can be thought to be a pri-miRNA of miR-675, to further understand the evolutionary patterns of miRNAs, we also hypothesized the pri-miRNA sequences through extending bilaterally to 500 nt of pre-miRNAs. The miRNAs in some species (such as Bos taurus and Canis familiaris) were not annotated or reported. We hypothesized these pre-miRNAs, pre-miRNAs and mature miRNAs by homology search, because the small ncRNAs are well-conserved phylogenetically [9, 25]. The secondary structures of pre-miRNAs were estimated in MiPred web server [26]. The secondary structure of H19 and pri-miRNAs were predicted in RNAfold WebServer (http://rna.tbi.univie.ac.at/cgi-bin/RNAfold.cgi) [27].
Phylogenetic relationships of H19, pri-/pre-mir-675 and miR-675-5p/3p were reconstructed with MEGA 5.10 [28] using Neighbor-Joining method, SplitsTree 4.10 [29] using Neighbor-Net method [30], and Network 4.6.1.0 (http://www.fluxus-engineering.com/) using Median-Joining method. These sequences were aligned with Clustal X 2.0 [31]. The nucleotide composition and nucleotide pair frequency were estimated in MEGA software. Further, target mRNAs of miR-675-5p/3p were predicted by TargetScan program [32], and functional enrichment analysis was performed using CapitalBio Molecule Annotation System V4.0 (MAS, http://bioinfo.capitalbio.com/mas3/).
References
Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641
Sempere LF, Cole CN, McPeek MA, Peterson KJ (2006) The phylogenetic distribution of metazoan microRNAs: insights into evolutionary complexity and constraint. J Exp Zool B 306:575–588
Taft R, Pheasant M, Mattick J (2007) The relationship between non-protein-coding DNA and eukaryotic complexity. BioEssays 29:288–299
Mattick JS (2007) A new paradigm for developmental biology. J Exp Biol 210:1526–1547
Berezikov E (2011) Evolution of microRNA diversity and regulation in animals. Nat Rev Genet 12:846–860
Gregory CS, Georges SL, Claes W (2012) The emerging role of non-coding RNAs in drug addiction. Front Genet 3:106. doi:10.3389/fgene.2012.00106
van Bakel H, Hughes TR (2009) Establishing legitimacy and function in the new transcriptome. Brief Funct Genomic Proteomic 8:424–436
He L, Hannon GJ (2004) Micrornas: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Wu W, Sun M, Zou GM, Chen JJ (2007) MicroRNA and cancer: current status and prospective. Int J Cancer 120:953–960
Garzon R, Fabbri M, Cimmino A, Calin GA, Croce CM (2006) MicroRNA expression and function in cancer. Trends Mol Med 12:580–587
Gabory A, Ripoche MA, Yoshimizu T, Dandolo L (2006) The H19 gene: regulation and function of a non-coding RNA. Cytogenet Genome Res 113:188–193
Hao Y, Crenshaw T, Moulton T, Newcomb E, Tycko B (1993) Tumour-suppressor activity of H19 RNA. Nature 365:764–767
Matouk IJ, DeGroot N, Mezan S, Ayesh S, Abu-lail R et al (2007) The H19 non-coding RNA is essential for human tumor growth. PLoS ONE 2:e845
Cai XZ, Cullen BR (2007) The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 13:313–316
Mineno J, Okamoto S, Ando T, Sato M, Chono H et al (2006) The expression profile of microRNAs in mouse embryos. Nucleic Acids Res 34:1765–1771
Latos PA, Pauler FM, Koerner MV, Senergin HB, Hudson QJ et al (2012) Airn transcriptional overlap, but not its lncRNA products, induces imprinted Igf2r silencing. Science 338:1469–1472
Okamura K, Phillips MD, Tyler DM, Duan H, Chou YT et al (2008) The regulatory activity of microRNA star species has substantial influence on microRNA and 3′ UTR evolution. Nat Struct Mol Biol 15:354–363
Okamura K, Ishizuka A, Siomi H, Siomi MC (2004) Distinct roles for argonaute proteins in small RNA-directed RNA cleavage pathways. Genes Dev 18:1655–1666
Guo L, Lu ZH (2010) The fate of miRNA* strand through evolutionary analysis: implication for degradation as merely carrier strand or potential regulatory molecule? PLoS ONE 5:e11387
Jagadeeswaran G, Zheng Y, Sumathipala N, Jiang HB, Arrese EL et al (2010) Deep sequencing of small RNA libraries reveals dynamic regulation of conserved and novel microRNAs and microRNA-stars during silkworm development. BMC Genomics 11:52
Guo L, Sun B, Wu Q, Yang S, Chen F (2012) miRNA–miRNA interaction implicates for potential mutual regulatory pattern. Gene 511:187–194
Kozomara A, Griffiths-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39:D152–D157
Plasterk RHA (2006) Micro RNAs in animal development. Cell 124:877–881
Jiang P, Wu H, Wang W, Ma W, Sun X et al (2007) MiPred: classification of real and pseudo microRNA precursors using random forest prediction model with combined features. Nucleic Acids Res 35:W339–W344
Hofacker IL (2003) Vienna RNA secondary structure server. Nucleic Acids Res 31:3429–3431
Tamura K, Peterson D, Peterson N, Stecher G, Nei M et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Huson DH (1998) SplitsTree: analyzing and visualizing evolutionary data. Bioinformatics 14:68–73
Bryant D, Moulton V (2004) Neighbor-Net: an agglomerative method for the construction of phylogenetic networks. Mol Biol Evol 21:255–265
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA et al (2007) Clustal W and clustal X version 2.0. Bioinformatics 23:2947–2948
Lewis BP, Shih IH, Jones-Rhoades MW, Bartel DP, Burge CB (2003) Prediction of mammalian microRNA targets. Cell 115:787–798
Acknowledgments
The work was supported by the National Natural Science Foundation of China (No. 61301251, 81072389, 81373102 and 81102182), the Research Found for the Doctoral Program of Higher Education of China (No. 211323411002), the China Postdoctoral Science Foundation funded project (No. 2012M521100), the key Grant of Natural Science Foundation of the Jiangsu Higher Education Institutions of China (No. 10KJA33034), the National Natural Science Foundation of Jiangsu (No. BK20130885), the Natural Science Foundation of the Jiangsu Higher Education Institutions (No. 12KJB310003 and 13KJB330003), the Jiangsu Planned Projects for Postdoctoral Research Funds (No. 1201022B), the Science and Technology Development Fund Key Project of Nanjing Medical University (No. 2012NJMU001), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Phylogenetic relationships of different sequences. These phylogenetic networks are reconstructed based on the H19 sequences, hypothesized primary miR-675 sequences (pri-mir-675), precursor miR-675 sequences (pre-mir-675), and mature products of mir-675 (miR-675-5p and miR-675-3p). Pre-mir-675 shows inconsistent distribution with H19 and pri-mir-675. The two products (miR-675-5p and miR-675-3p) show similar evolutionary patterns, although they are involved in various genetic distances across animal species. (TIFF 444 kb)
Fig. S2
Phylogenetic relationships and nucleotide compositions of Igf2 gene. Reconstructed phylogenetic trees based on Neighbor-Joining (A) and Neighbor-Net (B) methods. Frequency of nucleotide compositions (C), distributions of nucleotide pair frequency (D) and percentage of nucleotide pair frequency (E). Avg: indicates the average the nucleotide pair frequency. (TIFF 327 kb)
Fig. S3
Various nucleotide divergence patterns in H19. (A)-(C) are selected randomly conserved regions in H19, and (D) is pre-mir-675 sequence in H19. Many regions in H19 are well-conserved. (TIFF 163 kb)
Fig. S4
MFE structures (mfe), the thermodynamic ensemble of RNA structures (pf), the centroid structure (centroid) and the positional entropy for each position. (TIFF 239 kb)
Rights and permissions
About this article
Cite this article
Guo, L., Zhao, Y., Yang, S. et al. An integrated evolutionary analysis of miRNA–lncRNA in mammals. Mol Biol Rep 41, 201–207 (2014). https://doi.org/10.1007/s11033-013-2852-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-013-2852-4