Abstract
Structural studies on integrins have recently made great strides in recent years. Crystal structures of the complete extracellular fragments of three integrins in open and closed conformations, 6 α-I domains in complex with ligands, and at least 20 intracellular proteins in complex with cytosolic tails have been obtained; and several transmembrane and cytosolic complexes have been determined by NMR. High resolution EM studies complement these atomic resolution techniques by studying the integrin in different activation states. Although we still have only a few experimental examples among integrin family members, the high level of sequence homology between integrins means that reliable models can be built for the other members of the integrin family. These structures make sense of a lot of preceding biochemical, biophysical and mutagenesis studies, and generate many new testable hypotheses of integrin function. This chapter emphasizes new structural insights applicable to all integrins, with an emphasis on those integrins that contain an α-I domain. The structural data reinforce the notion of the integrin as a molecule in dynamic equilibrium at the cell surface, regulated by binding both to extracellular and intracellular ligands.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
8.1 Overall Structure
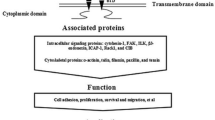
Integrins are αβ heterodimers, consisting of a head domain from which emerge two legs, one from each subunit, ending in a pair of single-pass transmembrane helices and short cytoplasmic tails, except for α6β4 (Fig. 8.1). The integrin “head” comprises a seven-bladed propeller from the α-subunit that makes an intimate contact with the β-I domain. Nine α-subunits (α1, α2, αl0, α11, αD, αE, αL, αM and αX) contain an additional domain, the α-I domain, that is inserted between two loops on the upper surface of the propeller, where it plays a central role in ligand binding [27, 41, 58]. The α-I domain contains an invariant ligand binding site called MIDAS, for Metal Ion-Dependent Adhesion Site [34], in which a metal ion is coordinated by three loops from the I domain, and a glutamic or aspartic acid from the ligand completes an octahedral coordination sphere around the metal. In those integrins that lack an α-I domain, the β-I domain and propeller form the major ligand recognition sites; in the α-I domain integrins, the β-I domain plays a regulatory role.
Cartoon of the αXβ2 structure derived by crystallography and EM studies. At left, the bent, low affinity integrin stabilized by bonds between the head, legs and cytoplasmic tails. At center, an unknown trigger causes the integrin to “stand up”, while maintaining most of its low affinity bonds. At right, binding of activated talin and/or binding of an extracellular ligand, trigger an open, high affinity form of the integrin, with TM helices separated
In the absence of ligand, bonds between the legs, tails and head are believed to hold the head in an “inactive” or “resting” conformation that has low affinity for ligand [20, 56, 61]. Recent structural data suggest that integrins possess three global conformations (see Fig. 8.1): a bent conformation in which the head adopts a “closed”, low affinity conformation and the cytoplasmic tails form an inhibitory complex; an extended conformation of the head that retains its low ligand affinity; and a high affinity form in which the legs and tails separate, and the “hybrid” domain, which is part of the head, swings away from the β-I domain, propeller and α-I domain, promoting conformational changes that create high affinity binding sites on both the head and tail [54]. During “outside-in” signaling, the head binds to ECM proteins or counter-receptors on other cells, triggering conformational changes that propagate down the “legs” and through the plasma membrane, leading to a reorganization of the C-terminal tails that allows them to bind intracellular proteins [46]. During “inside-out” signaling, cytosolic proteins bind and sequester one or both of the cytoplasmic tails, triggering conformational changes in the head that promote a high affinity “active” integrin [19, 60], in which the integrin “stands up”.
8.2 The α-I Domain
The first crystal structure of an α-I domain revealed a compact domain comprising a central mostly parallel β-sheet surrounded on both sides by amphipathic α-helices [34] (Fig. 8.2). Subsequent crystal structures of recombinant αL, α1 and α2 I domains display the same three-dimensional fold, as expected given their reasonable sequence similarity [15, 43, 45]. The MIDAS motif lies at the C-terminal end of the central β-sheet, with three loops contributing sidechains that coordinate the metal ion (Fig. 8.2 Lower panel). The metal-coordinating MIDAS residues are invariant among α-I domains, and mutagenesis of any of these residues abrogates ligand binding. Surface-exposed sidechains surrounding the MIDAS motif are more variable; they provide additional ligand contact points and hence ligand specificity [26, 41, 52].
Two conformations of the α-I domain. Upper panel conformational changes in the α2-I domain on binding collagen. Lower panel Conformational changes in the α-MIDAS motif upon binding ligand. At left, the “closed” conformation observed in the absence of ligand; at right, the “open” conformation seen when ligand is bound. These changes are mechanically linked to the tertiary changes in the domain. It is very likely that all α-I domains undergo the same switch
The structures of 6 ligand-bound α-I domains have now been determined. The first was the α2-I domain bound to a collagen-like triple helix [17]. More recently, the structures of the αL-I domain in complex with homologous fragments of ICAM-1 [51], ICAM-3 [53] and ICAM-5 [67] have been determined. Recently, the first authentic complex of an αM-I domain bound to ligand (the C3d domain of complement C3) [2] validates the earlier structure of the αM I domain bound to a “ligand-mimetic” crystal contact [33, 34]. They all demonstrate that ligand binding triggers a profound conformational switch in the α-I domain that underlies affinity regulation and signal transduction. The conformational switch is essentially identical in all these examples, strongly suggesting that all α-I domains will undergo the same switch.
8.3 Structural Determinants of Collagen Binding
Recombinant α2-I domain was crystallized as a complex with a homotrimer of a 21-mer peptide containing a critical GFOGER (where O is hydroxyproline) motif [17, 30]. The peptide closely resembles the structure of uncomplexed collagen-like peptides [16], and has the properties of a folded protein domain (i.e., stable secondary and tertiary structure). Three loops on the upper surface of the α2-I domain that comprise the MIDAS motif also engage the collagen, with a collagen glutamate completing the coordination sphere of the metal (Fig. 8.3). The critical roles of both the MIDAS and surrounding residues have been confirmed by mutagenesis [52].
Collagen binding to the α2-I domain. a Surface model of the α2-I domain colored by surface charge (red = negative, blue = positive) with a triple helical fragment of collagen bound. b Space-filling model of the complex (rotated about a horizontal axis compared with a), showing residues (in red) that are invariant in the collagen-binding integrins, α1β1, α2β1 and α10β1. The strong conservation of the binding surface suggests that these integrins will engage collagen in the same fashion. c Stereo close-up image of the α2-I:collagen complex
The buried surface area on complex formation (~1,200 A2) is at the lower limit of known protein-protein interfaces (the value is almost identical for the αL-I:ICAM complex), especially given the fact that some of the binding energy must be expended in switching the conformation of the I domain from closed to open. The quite reasonable affinity of the interaction (Kd = 35–90 nM) [24] reflects the unusually strong bonds formed by the glutamate-metal-I domain bridge, which has been estimated to contribute ~5 kcal/mol. This bridge is indeed critical, since the conservative substitution of collagen Glu to Asp in the GFOGER motif eliminates binding [31], presumably because the aspartic acid is too short to reach down from the rigid collagen triple helix to bind to the partly buried metal ion. However, in the case of the αM-C3d interaction, Asp is the preferred residue, perhaps because it lies on a flexible loop at the end of a helical segment [2].
The MIDAS motif and much of the collagen-binding surface are strictly invariant among the collagen-binding I domain integrins (αl, α2, α10 and α11), suggesting that these integrins will all engage collagen in a similar fashion, with a strict requirement for glutamate in the collagen motif. The periphery of the binding surface is more variable, however (Fig. 8.3b), which would explain their collagen type preferences. The recent structure of the α1-I domain bound to collagen containing the closely related motif, GLOGEN, confirms this [11]. Ιn addition, a gain-of-function point mutation in the α2-I domain (i.e. one that favors the open conformation) [10] displays relaxed specificity and alternate binding modes to the GFOGER motif. Given the special nature of collagen (see Chap. 3 by Zutter and Santoro), this observation may point to profound consequences for collagen recognition by activated cells.
8.4 The Integrin α-I Domain and the von Willebrand Factor (vWF) A Domain: A Caveat
The integrin α-I domain is generally categorized as a member of the vWF-A domain superfamily, based on sequence similarity and a highly conserved overall 3-dimensional structure. However, since MIDAS- and non-MIDAS containing vWFA-domains have distinct ligand-binding and allosteric properties, this author believes that much confusion could be avoided if the family was reclassified into two sub-families: Two examples illustrate my point. First, the eponymous vWF-A1 and vWF-A3 domains lack at least one of the acidic residues of authentic MIDAS motifs, and so do not bind metal; moreover, they bind ligands via distinct surfaces, and conformational changes are not induced [21]. In fact, vWF-A3, like integrin α2β1, binds triple-helical collagen, but it utilizes a different surface (one side of its β-sheet) [7]. Second, the “vWF-A domain” of Factor B does contain a functional MIDAS motif, and binds an acidic moiety in its ligand, complement iC3b, in the canonical integrin fashion; in this case, the metal ion engages the C-terminal carboxylate iC3b, triggering integrin-like conformational changes [18], suggesting that it should be classed as an I domain. Indeed, a genome-wide collection of vWF-A domains has been compiled [62] and have been these subdivided into I-like and A-like domains based on the conservation of key MIDAS residues.
8.5 Conformational Changes in the α-I Domain
Ligand binding alters the conformation of α-I domains in the same way in the three cases studied thus far (α2, αM and αL), as well as in a subset of “vWF-A domain” (as noted above) and the matrix receptor, TEM8 (in complex with pathogen; see below). Binding of an acidic residue to the α-MIDAS causes a switch in in Mg2+ coordination in which a direct bond to a MIDAS threonine is gained while a direct bond to an aspartic acid is lost (Fig. 8.2b, c). These subtle changes in metal coordination are linked to extensive secondary and tertiary changes that create a complementary surface for binding ligand and generate a 10 Å downward movement of the C-terminal helix, α7. The helix movement links the change in the upper ligand-binding surface to the lower surface of the domain. The shift of the helix α7 is highly significant in the context of the whole integrin, since the helix is packed against the propeller and β-I domain (see Sect. 8.8).
The close similarity between the structural changes seen in the three α-I domains and a subset of A domains suggests that there are just two principal conformations for I domains, “open” and “closed” (Fig. 8.2). The “open” conformation is seen in the presence of ligand or ligand mimetic, while the “closed” conformation is seen in the absence of ligand. It therefore appears to be the formation of a strong ligand-metal bond, requiring a change in metal coordination, that triggers the conformational switch. Springer’s group has engineered disulfide-linked αL-I domains with intermediate affinity and packing of the C-terminal helix, and suggested the existence of an intermediate state [51]. However, in these structures the MIDAS motif exists in only two conformations, and it remains to be seen whether the intermediate conformation has biological relevance or is an artifact of the engineered disulfide. It should be noted that it is not necessary to invoke an intermediate tertiary conformation in order to explain an intermediate affinity. In principle, a shift in the position of the equilibrium between two states is sufficient [38].
Various studies have now been published in support of the hypothesis that the open and closed conformations of the α-I domain equate with high and low affinity states. Thus, mutants of the αM-I domain that are predicted to destabilize the closed conformation and favor the open conformation increase the affinity for the ligand iC3b [41]. The epitope for an antibody that binds only to the high affinity form of the αMβ2 integrin maps to a region that undergoes extensive conformational changes between the closed and open forms [35].
Disulfide engineering studies on recombinant I domains and full-length integrins, which lock the domain either into the open or closed state, also support the hypothesis [39, 49, 50]. So does the structure of the αL-I domain in complex with the inhibitor lovastatin [25], which reveals allosteric inhibition by binding between the β-sheet and the C-terminal helix, preventing the helical shift.
It should also be appreciated that pathogens often utilize integrin α-I domains for cell entry, and there is evidence that many bind across the MIDAS motif. However, in general they bind preferentially to the (default) closed conformation, sometimes involving direct bonds to the MIDAS, but they do not induce conformational changes [4]. One counter-example is anthrax toxin, which utilizes a glutamate to engage the bona fide MIDAS motif of the “vWF A domain” of the collagen receptor, TEM8, in its open conformation [6]. There is also one clear example of gene transfer, in which the Gram-positive pathogen, Streptococcus pneumoniae, has an α-I domain inserted into the tip of its pilus, perhaps to act as a shear stress-activated adhesin for attachment to host cells [23, 36].
8.6 The β-I Domain and the Integrin Headpiece
The existence of a β-I domain was initially predicted based on the conserved and critical MIDAS-like sequence, DxSxS, and hydropathy plot comparisons with the α-I domain [34]; and later from more sophisticated sequence analysis [59]. The structure of the β3-I domain, contained within the αVβ3 crystal structure [64], confirmed that the basic fold and topology are very similar to the α-I domain, albeit with many large insertion/loops between β-strands, which had confounded conventional sequence alignment algorithms.
In contrast to α-I, the β-I domain is not folded independently, but packs rigidly against the α-subunit propeller, with the major ligand recognition elements lying at the interface (see Fig. 8.4) [42, 64]. The β-MIDAS is similar to the α-MIDAS, except that the α-MIDAS threonine is replaced by glutamate. This difference likely explains the different cation specificities—in α-MIDAS, the smaller Mg2+ ion favors ligands lacking a formal charge; while in β-MIDAS, the larger Ca2+ ion favors multiple acidic ligands. The structure of αVβ3 in complex with an RGD-style peptide shows that the Asp sidechain completes the coordination sphere of the MIDAS metal ion [65], as predicted. There are also further metal-binding sites adjacent to the β-MIDAS (the “ADMIDAS” and “SyMBS”) that play important structural and possibly regulatory roles in ligand-binding and regulation [68].
Tertiary and quaternary changes triggered by ligand binding in integrins that lack an α-I domain. Ligand binding to the β-MIDAS motif (M) causes a shift of helix α1, which generates a rotation of α7 helix (black arrow within region circled in black) and a loosening of the contacts between the β-hybrid domain and the propeller. The β-hybrid domain is then free to swing by as much as 60° away from the α-propeller
Although Xiong et al. initially proposed the opposite, the conformation of the β-I domain in their unliganded crystal structure [64] corresponds to the closed conformation of the α-I domain. Soaking of RGD ligand into preformed crystals induced small changes within the β-I domain, but these were not propagated to the rest of the headpiece; i.e., they were frustrated by the constraints of the closed quaternary structure [37]. This situation is typical in crystallography: either the ligand binds and induces small changes constrained by the lattice, or it induces large changes that destroy the lattice.
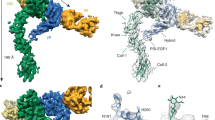
However, Springer’s group has recently discovered a rare exception to this rule, and report a crystal form of the αIIbβ3 headpiece with large solvent channels in which the lattice tolerates (and/or adjusts to) a switch from the closed to the open conformation, involving an outward swing of the hybrid domain by ~40°. Preformed crystals were simply soaked with different concentrations/durations of an RGD ligand mimetic and different Ca2+/Mg2+ ratios [69] (Fig. 8.4). This remarkable observations settles many questions with regard to head-opening, although the crystal structure of the headpiece in complex with a non-peptidic ligand is still lacking.
8.7 Quaternary Regulation in Integrins Lacking an α-I Domain
Takagi et al. [57] showed that the inactive (resting) form of the integrin αVβ3, observed in physiological concentrations of Ca2+ and Mg2+, is largely bent, and closely resembles the crystal structure, in which the C-termini of both chains are closely apposed. Based on the one case studied of an α-I domain integrin, the αXβ2 ectodomain, it also adopts a similar (although distinct) bent default conformation [63]. Other integrins tested had a lower propensity to adopt the bent conformation; however, the experiments were performed with extracellular heterodimers truncated near the plasma membrane, so that they lacked the transmembrane helices and cytoplasmic tails that are known to contribute critically to the stability of the inactive conformation. By engineering a disulfide link between the α-subunit propeller and the EGF4 domain of the β-subunit (which are 4 Å apart in the bent (crystal) structure, but would be very far apart in the “standing-up” conformation), Takagi et al. further showed that integrin expressed on the cell surface was in a low affinity state and could only be activated under reducing conditions.
The current model for integrins invokes a minimum of three distinct states: (i) bent, low affinity; (ii) standing-up, legs together, low affinity; and (iii) standing-up, legs apart, high affinity (see Fig. 8.1). The position of equilibrium depends on the concentrations and activation status of extracellular and intracellular ligands, as well as divalent cations. At the heart of the switch is an outward swing of the β-hybrid domain with respect to the β-I domain, by as much as 60° (Fig. 8.4). In αIIbβ3, the primary response to extracellular ligand binding is a concerted reorganization of the N-terminal helix (attached directly to the β-MIDAS) and the adjacent C-terminal helix. Rather than translate downward (as in the case of the α-I domain), the principle motion of α7 is a rotation about an axis close to the β-MIDAS, which is linked to the rotation of the β-hybrid domain. Thus, although some details may differ, the data support the prediction that the trigger for the integrin switch is similar in integrins that contain or lack and α-I domain: i.e., a subtle change in metal coordination at the MIDAS motif is linked to a reorganization of the I domain architecture that leads to quaternary changes toward an open, high-affinity state [33].
As noted above, these experiments were performed with truncated integrins and small peptide ligands. The nature of the trigger in the integrin head seems secure, but it remains to be seen how the quaternary changes are promulgated across the plasma membrane. Recent studies have shown that full-length integrin can be reconstituted into lipid nanodiscs and visualized by high resolution Electron Microscopy [12], so we should soon have an answer.
8.8 Quaternary Changes in Integrins Containing an α-I Domain
As noted above, in integrins that lack an α-I domain, the β-I domain and α-subunit propeller are the major recognition elements [42]. However, in integrins that contain an α-I domain, the β-I and α-propeller do not play direct roles in ligand recognition; instead they play important regulatory roles. This concept initially caused some confusion: thus, mutation of the β-MIDAS motif led to loss of iC3b binding to αMβ2 [3] which was initially interpreted as evidence for a direct role for the β-I domain in ligand; it now seems clear, however, that the mutation works allosterically, by preventing conformational changes in the α-I domain.
How does the quaternary organization of the integrin regulate the affinity of the α-I domain? We know that regulation occurs allosterically (rather than by steric masking of the binding site), since the α-I domain is a major antibody epitope. Hypotheses focused on the loss-of-function effect of mutating a Glu residue within a conserved ΦEGT motif (where Φ is any hydrophobic residue) at the end of the α-I domain C-terminal (α7) helix [1, 22, 66]; and it was suggested that the Glu could act as an intradimer ligand by completing the coordination sphere of the β-MIDAS motif. The first crystal structures of the αXβ2 headpiece (from Xie et al. [63]) were inconclusive: they showed the α-I domain in the closed conformation, but rather loosely attached to the rest of the headpiece. However, a recent structure of the αXβ2 ectodomain displays an activated α-I domain by virtue of a fortuitous crystal contact [47]. Although the rest of the αβ headpiece is in the closed conformation, the predicted “internal ligand”, Glu318, is observed coordinating the β-MIDAS motif with only minor compensatory movements in the β-I domain (Fig. 8.5). By contrast, the α-I domain adopts a fully “open” conformation, with the MIDAS threonine directly coordinating the metal and (what appears to be) a chloride ion completing the coordination sphere. The first half of the α-I domain α7 helix has shifted by ~10 Å, as expected, but the remainder is unwound, thereby switching the orientation with respect to the headpiece. Thus, the crystalline environment seems to have created a hybrid molecule with a fully active α-I domain in the context of an inactive headpiece. It is possible that such a hybrid state could exist in vivo, providing long-range, flexible and rapid (non-equilibrium) responses to the presence of ligand and/or mechanical stress; with a slow switch to the overall open (equilibrium) conformation occurring if the signal persisted.
Close-up comparison of the α-I domain-containing integrin head of αXβ2. In the open conformation, Glu318 acts as an internal ligand to the β-MIDAS that generates a 10 Å shift in the first half of helix α7 of the αI domain, while the second half of the helix unwinds, leading to a 30–40° rotation of the α-I domain about the propeller and β-I domain. Τhe open α-I domain is stabilized by a crystal contact, and the the β-I domain remains principally in the closed conformation
8.9 Transmembrane (TM) and Tail Interactions
There are abundant biochemical and genetic data supporting the notion that interactions between integrin α- and β-TM helices and cytosolic tails help to hold the resting integrin in a low affinity conformation. In a classic study by Ginsberg and colleagues, a salt bridge between αIIb Arg995 and β3 Asp723 was shown to be necessary and sufficient to hold the integrin in its resting state [20]. A definitive structure of the αIIbβ3 tails in bicelles [32] reveals a remarkably stable conformation, in which the two helices pack closely together at the extracellular end; at the intracellular end they diverge, but the void is effectively filled by a highly conserved aromatic triplet that breaks the α-subunit helix and turns inward (Fig. 8.6). Isolated α and β subunit helices studies in bicelles show a remarkably well-conserved structure: the α-subunit is always orthogonal to the membrane while the β-subunit helix is always tilted. Recently, Ginsberg has shown that Lys 716β, whose Cα is buried in the membrane, can “snorkel” to the hydrophilic headgroups by extension of the Lys sidechain, and moreover that this residue is essential for maintaining the tilted helix and TM signaling [28, 55]. Recent studies on α-I domain integrins have yielded consistent results for isolated TM regions, and structures of αβ complexes are in progress [13]. The switch to the “open” conformation may entail a simple separation of the tails, which maintain their structural integrity and reassemble rapidly when the integrin returns to the low affinity state.
Integrin-tail interactions. Upper panel Atomic interactions between the αllb and β3 tails as revealed by NMR, melded to the complex of Talin2 and β1D tail. Lower panel Aligned sequences of α and β TM segments. Important residues in αIIbβ3 (and conserved in α-I domain integrins) are circled. See text for details
The penultimate question is how cytosolic proteins interact with the cytoplasmic tails of integrins, and the number of structural examples of protein domains bound to tail peptides (mostly β, but some α) has grown rapidly in recent years (see Table 8.1). It is clear that some proteins bind strongly enough to the β-tail to promote integrin activation. Talin was the first such molecule to be thus characterized, and remains the central player [8], although the number of additional contributors, such as kindlin and filamin [9], is growing fast. A model of talin activation is presented in Fig. 8.6. Talin sequesters the β-tail, breaking the critical R995α-D723β bond. It is also clear that phosphorylation of the β-tails provides a rapid means of switching between binding partners, and thus between cell migration and adhesion [14, 40, 44].
The final question is how the cytosolic activators generate force across the membrane. In the case of talin, recent work by Ginsberg’s group suggests that dissociation of the tails, which have flexible linkages to the extracellular domains, is sufficient [29]. Key to this process is talin’s ability to bind simultaneously to the integrin β-tail and membrane (Fig. 8.6), the latter providing a pivot point to force the two helices apart.
8.10 Perspectives
Structural and structure-function studies have revealed many of the major paradigms of integrin allostery underlying affinity regulation and bi-directional signal transduction. Notably lacking is the structure of an intact integrin bound to a physiological ligand in a membrane environment that would reveal the true “active” conformation of the molecule. EM studies show the greatest promise here, most likely using nanodisks. We are beginning to understand the structures of the TM helices and their cytosolic extensions, but the biophysics of inside-out signaling in particular requires further study. The role of mechanical force, whether of intracellular (actomyosin motors) or extracellular (shear flow in the vasculature) origin has not been discussed here, but its interplay with the chemical forces that attract cognate molecules is a fascinating field for current and future study [48]. Finally, this chapter has addressed the structural basis of affinity changes within individual integrins. Lateral association (clustering) of integrins in the plasma membrane at sites of ECM contact also plays a major role in integrin signaling, and we still know little about its structural basis and regulation [5].
References
Alonso JL, Essafi M, Xiong JP, Stehle T, Arnaout MA (2002) Does the integrin αA domain act as a ligand for its βA domain? Curr Biol 12:R340–R342
Bajic G, Yatime L, Sim RB, Vorup-Jensen T, Andersen GR (2013) Structural insight on the recognition of surface-bound opsonins by the integrin I domain of complement receptor 3. Proc Natl Acad Sci USA 110:16426–16431
Bajt ML, Loftus JC, Gawaz MP, Ginsberg MH (1992) Characterization of a gain of function mutation of integrin αIIb β3 (platelet glycoprotein IIb-IIIa). J Biol Chem 267:22211–22216
Bergelson JM, Chan BM, Finberg RW, Hemler ME (1993) The integrin VLA-2 binds echovirus 1 and extracellular matrix ligands by different mechanisms. J Clin Invest 92:232–239
Boettiger D (2012) Mechanical control of integrin-mediated adhesion and signaling. Curr Opin Cell Biol 24:592–599
Bradley KA, Mogridge J, Jonah G, Rainey A, Batty S, Young JA (2003) Binding of anthrax toxin to its receptor is similar to α integrin-ligand interactions. J Biol Chem 278:49342–49347
Brondijk TH, Bihan D, Farndale RW, Huizinga EG (2012) Implications for collagen I chain registry from the structure of the collagen von Willebrand factor A3 domain complex. Proc Natl Acad Sci USA 109:5253–5258
Calderwood DA (2004) Talin controls integrin activation. Biochem Soc Trans 32:434–437
Calderwood DA, Campbell ID, Critchley DR (2013) Talins and kindlins: partners in integrin-mediated adhesion. Nat Rev Mol Cell Biol 14:503–517
Carafoli F, Hamaia SW, Bihan D, Hohenester E, Farndale RW (2013) An activating mutation reveals a second binding mode of the integrin α2 I domain to the GFOGER motif in collagens. PLoS ONE 8:e69833
Chin YK, Headey SJ, Mohanty B, Patil R, McEwan PA, Swarbrick JD et al (2013) The structure of integrin α1I domain in complex with a collagen-mimetic peptide. J Biol Chem 288:36796–36809
Choi WS, Rice WJ, Stokes DL, Coller BS (2013) Three-dimensional reconstruction of intact human integrin αIIbβ3: new implications for activation-dependent ligand binding. Blood 122:4165–4171
Chua GL, Tang XY, Patra AT, Tan SM, Bhattacharjya S (2012) Structure and binding interface of the cytosolic tails of αXβ2 integrin. PLoS ONE 7:e41924
Deshmukh L, Meller N, Alder N, Byzova T, Vinogradova O (2011) Tyrosine phosphorylation as a conformational switch: a case study of integrin β3 cytoplasmic tail. J Biol Chem 286:40943–40953
Emsley J, King SL, Bergelson JM, Liddington RC (1997) Crystal structure of the I domain from integrin α2β1. J Biol Chem 272:28512–28517
Emsley J, Knight CG, Farndale RW, Barnes MJ (2004) Structure of the integrin α2β1-binding collagen peptide. J Mol Biol 335:1019–1028
Emsley J, Knight CG, Farndale RW, Barnes MJ, Liddington RC (2000) Structural basis of collagen recognition by integrin α2β1. Cell 101:47–56
Forneris F, Ricklin D, Wu J, Tzekou A, Wallace RS, Lambris JD et al (2010) Structures of C3b in complex with factors B and D give insight into complement convertase formation. Science 330:1816–1820
Garcia-Alvarez B, de Pereda JM, Calderwood DA, Ulmer TS, Critchley D, Campbell ID et al (2003) Structural determinants of integrin recognition by talin. Mol Cell 11:49–58
Hughes PE, Diaz-Gonzalez F, Leong L, Wu C, McDonald JA, Shattil SJ et al (1996) Breaking the integrin hinge. A defined structural constraint regulates integrin signaling. J Biol Chem 271:6571–6574
Huizinga EG, Tsuji S, Romijn RA, Schiphorst ME, de Groot PG, Sixma JJ et al (2002) Structures of glycoprotein Ibα and its complex with von Willebrand factor A1 domain. Science 297:1176–1179
Huth JR, Olejniczak ET, Mendoza R, Liang H, Harris EA, Lupher ML Jr et al (2000) NMR and mutagenesis evidence for an I domain allosteric site that regulates lymphocyte function-associated antigen 1 ligand binding. Proc Natl Acad Sci USA 97:5231–5236
Izore T, Contreras-Martel C, El Mortaji L, Manzano C, Terrasse R, Vernet T et al (2010) Structural basis of host cell recognition by the pilus adhesin from Streptococcus pneumoniae. Structure 18:106–115
Jung SM, Moroi M (1998) Platelets interact with soluble and insoluble collagens through characteristically different reactions. J Biol Chem 273:14827–14837
Kallen J, Welzenbach K, Ramage P, Geyl D, Kriwacki R, Legge G et al (1999) Structural basis for LFA-1 inhibition upon lovastatin binding to the CD11a I-domain. J Mol Biol 292:1–9
Kamata T, Puzon W, Takada Y (1994) Identification of putative ligand binding sites within I domain of integrin α2β1 (VLA-2, CD49b/CD29). J Biol Chem 269:9659–9663
Kamata T, Takada Y (1994) Direct binding of collagen to the I domain of integrin α2β1 (VLA-2, CD49b/CD29) in a divalent cation-independent manner. J Biol Chem 269:26006–26010
Kim C, Schmidt T, Cho EG, Ye F, Ulmer TS, Ginsberg MH (2012) Basic amino-acid side chains regulate transmembrane integrin signalling. Nature 481:209–213
Kim C, Ye F, Hu X, Ginsberg MH (2012) Talin activates integrins by altering the topology of the β transmembrane domain. J Cell Biol 197:605–611
Knight CG, Morton LF, Onley DJ, Peachey AR, Messent AJ, Smethurst PA et al (1998) Identification in collagen type I of an integrin α2β1-binding site containing an essential GER sequence. J Biol Chem 273:33287–33294
Knight CG, Morton LF, Peachey AR, Tuckwell DS, Farndale RW, Barnes MJ (2000) The collagen-binding A-domains of integrins α1β1 and α2β1 recognize the same specific amino acid sequence, GFOGER, in native (triple-helical) collagens. J Biol Chem 275:35–40
Lau TL, Kim C, Ginsberg MH, Ulmer TS (2009) The structure of the integrin αIIbβ3 transmembrane complex explains integrin transmembrane signalling. EMBO J 28:1351–1361
Lee JO, Bankston LA, Arnaout MA, Liddington RC (1995) Two conformations of the integrin A-domain (I-domain): a pathway for activation? Structure 3:1333–1340
Lee JO, Rieu P, Arnaout MA, Liddington R (1995) Crystal structure of the A domain from the α subunit of integrin CR3 (CD11b/CD18). Cell 80:631–638
Li R, Rieu P, Griffith DL, Scott D, Arnaout MA (1998) Two functional states of the CD11b A-domain: correlations with key features of two Mn2+-complexed crystal structures. J Cell Biol 143:1523–1534
Liddington R (2010) Catching pneumonia. Structure 18:6–8
Liddington RC (2002) Will the real integrin please stand up? Structure 10:605–607
Loftus JC, Liddington RC (1997) Cell adhesion in vascular biology. New insights into integrin-ligand interaction. J Clin Invest 99:2302–2306
McCleverty CJ, Liddington RC (2003) Engineered allosteric mutants of the integrin αMβ2 I domain: structural and functional studies. Biochem J 372:121–127
McCleverty CJ, Lin DC, Liddington RC (2007) Structure of the PTB domain of tensin1 and a model for its recruitment to fibrillar adhesions. Protein Sci 16:1223–1229
Michishita M, Videm V, Arnaout MA (1993) A novel divalent cation-binding site in the A domain of the β2 integrin CR3 (CD11b/CD18) is essential for ligand binding. Cell 72:857–867
Mould AP, Askari JA, Aota S, Yamada KM, Irie A, Takada Y et al (1997) Defining the topology of integrin α5β1-fibronectin interactions using inhibitory anti-α5 and anti-β1 monoclonal antibodies. Evidence that the synergy sequence of fibronectin is recognized by the amino-terminal repeats of the α5 subunit. J Biol Chem 272:17283–17292
Nolte M, Pepinsky RB, Venyaminov S, Koteliansky V, Gotwals PJ, Karpusas M (1999) Crystal structure of the α1β1 integrin I-domain: insights into integrin I-domain function. FEBS Lett 452:379–385
Oxley CL, Anthis NJ, Lowe ED, Vakonakis I, Campbell ID, Wegener KL (2008) An integrin phosphorylation switch: the effect of β3 integrin tail phosphorylation on Dok1 and talin binding. J Biol Chem 283:5420–5426
Qu A, Leahy DJ (1995) Crystal structure of the I-domain from the CD11a/CD18 (LFA-1, αLβ2) integrin. Proc Natl Acad Sci USA 92:10277–10281
Schwartz MA, Ginsberg MH (2002) Networks and crosstalk: integrin signalling spreads. Nat Cell Biol 4:E65–E68
Sen M, Yuki K, Springer TA (2013) An internal ligand-bound, metastable state of a leukocyte integrin, αXβ2. J Cell Biol 203:629–642
Seong J, Tajik A, Sun J, Guan JL, Humphries MJ, Craig SE et al (2013) Distinct biophysical mechanisms of focal adhesion kinase mechanoactivation by different extracellular matrix proteins. Proc Natl Acad Sci USA 110:19372–19377
Shimaoka M, Shifman JM, Jing H, Takagi J, Mayo SL, Springer TA (2000) Computational design of an integrin I domain stabilized in the open high affinity conformation. Nat Struct Biol 7:674–678
Shimaoka M, Takagi J, Springer TA (2002) Conformational regulation of integrin structure and function. Annu Rev Biophys Biomol Struct 31:485–516
Shimaoka M, Xiao T, Liu JH, Yang Y, Dong Y, Jun CD et al (2003) Structures of the αL I domain and its complex with ICAM-1 reveal a shape-shifting pathway for integrin regulation. Cell 112:99–111
Smith C, Estavillo D, Emsley J, Bankston LA, Liddington RC, Cruz MA (2000) Mapping the collagen-binding site in the I domain of the glycoprotein Ia/IIa (integrin α2β1). J Biol Chem 275:4205–4209
Song G, Yang Y, Liu JH, Casasnovas JM, Shimaoka M, Springer TA et al (2005) An atomic resolution view of ICAM recognition in a complex between the binding domains of ICAM-3 and integrin αLβ2. Proc Natl Acad Sci USA 102:3366–3371
Springer TA, Dustin ML (2012) Integrin inside-out signaling and the immunological synapse. Curr Opin Cell Biol 24:107–115
Surya W, Li Y, Millet O, Diercks T, Torres J (2013) Transmembrane and juxtamembrane structure of αL integrin in bicelles. PLoS ONE 8:e74281
Takagi J, Erickson HP, Springer TA (2001) C-terminal opening mimics ‘inside-out’ activation of integrin α5β1. Nat Struct Biol 8:412–416
Takagi J, Petre BM, Walz T, Springer TA (2002) Global conformational rearrangements in integrin extracellular domains in outside-in and inside-out signaling. Cell 110:599–611
Tuckwell D, Calderwood DA, Green LJ, Humphries MJ (1995) Integrin α2 I-domain is a binding site for collagens. J Cell Sci 108:1629–1637
Tuckwell DS, Humphries MJ (1997) A structure prediction for the ligand-binding region of the integrin ß subunit: evidence for the presence of a von Willebrand factor A domain. FEBS Lett 400:297–303
Vinogradova O, Haas T, Plow EF, Qin J (2000) A structural basis for integrin activation by the cytoplasmic tail of the αIIb-subunit. Proc Natl Acad Sci USA 97:1450–1455
Vinogradova O, Velyvis A, Velyviene A, Hu B, Haas T, Plow E et al (2002) A structural mechanism of integrin αIIbβ3 “inside-out” activation as regulated by its cytoplasmic face. Cell 110:587–597
Whittaker CA, Hynes RO (2002) Distribution and evolution of von Willebrand/integrin A domains: widely dispersed domains with roles in cell adhesion and elsewhere. Mol Biol Cell 13:3369–3387
Xie C, Zhu J, Chen X, Mi L, Nishida N, Springer TA (2010) Structure of an integrin with an αI domain, complement receptor type 4. EMBO J 29:666–679
Xiong JP, Stehle T, Diefenbach B, Zhang R, Dunker R, Scott DL et al (2001) Crystal structure of the extracellular segment of integrin αVβ3. Science 294:339–345
Xiong JP, Stehle T, Zhang R, Joachimiak A, Frech M, Goodman SL et al (2002) Crystal structure of the extracellular segment of integrin αVβ3 in complex with an Arg-Gly-Asp ligand. Science 296:151–155
Yang W, Shimaoka M, Salas A, Takagi J, Springer TA (2004) Intersubunit signal transmission in integrins by a receptor-like interaction with a pull spring. Proc Natl Acad Sci USA 101:2906–2911
Zhang H, Casasnovas JM, Jin M, Liu JH, Gahmberg CG, Springer TA et al (2008) An unusual allosteric mobility of the C-terminal helix of a high-affinity αL integrin I domain variant bound to ICAM-5. Mol Cell 31:432–437
Zhang K, Chen J (2012) The regulation of integrin function by divalent cations. Cell Adh Migr 6:20–29
Zhu J, Zhu J, Springer TA (2013) Complete integrin headpiece opening in eight steps. J Cell Biol 201:1053–1068
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Liddington, R.C. (2014). Structural Aspects of Integrins. In: Gullberg, D. (eds) I Domain Integrins. Advances in Experimental Medicine and Biology, vol 819. Springer, Dordrecht. https://doi.org/10.1007/978-94-017-9153-3_8
Download citation
DOI: https://doi.org/10.1007/978-94-017-9153-3_8
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-017-9152-6
Online ISBN: 978-94-017-9153-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)