Summary
Environmental signals control diverse physiological processes in plant growth and development. Plants tend to adapt the structure of photosynthetic apparatus and pigment composition in response to several environmental factors. Tetrapyrroles play vital roles in various biological processes, including photosynthesis and respiration. Expression of genes encoding enzymes of tetrapyrrol biosynthesis as well as the abundances and activities of the enzymes are severely impacted by availability of water, soil salinity, low or high temperature and low or high light intensity. Plastids share many cellular metabolic pathways and alterations of plastid functions by environmental signals are known to affect various aspects of plant development. The generation of reactive oxygen species (ROS) in plants is triggered by different kinds of environmental parameters, such as high light, high or low temperature, salinity, drought and nutrient deficiency. Imbalance between production of ROS and their detoxification by enzymatic and non-enzymatic reactions causes oxidative stress. Suitable genetic manipulation of the chlorophyll (Chl) biosynthetic pathway might lead to tolerance towards environmental stresses leading to oxidative stress at the cellular level, and efficient adaptation of the photosynthetic apparatus to low and high light intensities. The present review deals with environmental modulation of Chl biosynthesis and its impact on plant productivity.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
I. Introduction
Environmental cues including light, water, temperature, soil and nutrient content have a profound impact on plant growth and development. For example, light has significant effects on morphogenesis of seedlings during the transition from heterotrophic to photoautotrophic growth. Plants tend to adapt the structure of the photosynthetic apparatus and the pigment composition to light quality and quantity and other environmental factors. Tetrapyrroles play vital roles in various biological processes, including photosynthesis and respiration (Rebeiz et al. 1994; Papenbrock and Grimm 2001; Block et al. 2007; Tanaka and Tanaka 2007; Masuda 2008; Biswal et al. 2012; Phung et al. 2012). Plants are exposed to various abiotic stresses such as low temperature, high temperature, salinity, drought, flooding, oxidative stress and heavy metal toxicity etc. either during their entire life cycle or a part thereof. Plant growth, development, photosynthesis and productivity are severely affected due to environmental stresses, particularly during early seedling growth. When seeds germinate beneath the soil, their seedlings remain in near-darkness for a while. Therefore, etiolated rice seedlings beneath the soil do not synthesize Chl and contain a special form of plastids called etioplasts or etiochloroplasts. As seedlings come out of soil, they are exposed to light and light-mediated Chl biosynthesis and other associated greening processes are initiated resulting in transformation of etioplasts to chloroplasts (See Solymosi and Aronsson, Chap. 3). During chloroplast biogenesis in the light, proplastids in meristematic tissue and etioplasts in dark-grown seedlings develop into the mature, photosynthetic chloroplast of the green leaf (Waters and Pyke 2005).
Chloroplast biogenesis and development in seedlings can be described as the differentiation process from the plastid progenitor, a proplastid, to a mature chloroplast. Plastids carry out many essential metabolic pathways and alteration of plastid functions affects various aspects of plant growth and development. Chloroplasts are responsible for the biosynthesis of carbohydrates, fatty acids, pigments, and the synthesis of amino acids and proteins from inorganic nitrogen (Staehelin and Newcomb 2000). Chloroplast development involves the biosynthesis of components of the photosynthetic apparatus involving synthesis of Chl and carotenoids, lipids and proteins which is governed in a coordinated manner by chloroplast and nuclear genomes (Leon et al. 1998; Gray et al. 2003; Nott et al. 2006). Biosynthesis of porphyrins, particularly that of Chl, during early greening stages of seedlings is elucidated in detail (Tripathy and Rebeiz 1985, 1986, 1988; Meskauskiene et al. 2001; Goslings et al. 2004; Bollivar 2006; Tanaka and Tanaka 2007; Wu et al. 2007; Wang et al. 2010; Tripathy and Pattanayak 2012).
Chl biosynthesis and chloroplast development during irradiation of dark-grown plants is impacted by external and internal factors such as light quality, temperature, nutrition, leaf age, leaf water potential, salt etc. as they influence transcription, translation and post-translational modification of proteins involved in chloroplast biogenesis (Virgin 1965; Bengtson et al. 1978; Bhardwaj and Singhal 1981; Eskins et al. 1986; Tewari and Tripathy 1998, 1999; Le Lay et al. 2000, 2001; Sood et al. 2004, 2005; Mohanty et al. 2006; Dutta et al. 2009; Mohanty and Tripathy 2011; Dalal and Tripathy 2012).
All enzymes of the Chl biosynthetic pathway are nuclear encoded and post-translationally imported into chloroplasts. Chl synthesis is synchronized with the formation of other pigments such as carotenoids and with pigment-binding proteins; Chl synthesis is also involved in the coordination between chloroplast and nucleus (Nott et al. 2006).
II. Chlorophyll Biosynthetic Enzymes and Their Modulation by Environment
A. Biosynthesis of 5-Aminolevulinic Acid
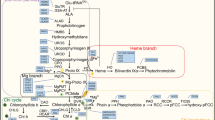
Unlike animals where one enzyme 5-aminolevulinic acid (ALA) synthase could form ALA by condensation and decarboxylation of succinyl-CoA and glycine, the synthesis of ALA in plants requires three different enzymes located in the chloroplast stroma. These are glutamyl-tRNA synthetase (GluRS) [EC 6.1.1.17], glutamyl-tRNA reductase (GluTR) [EC 1.2.1.70] and glutamate semialdehyde aminotransferase (GSA-AT) [EC 5.4.3.8] (Fig. 27.1a).
GluRS, also known as aminoacyl-tRNA synthetase, ligates glutamate to tRNAGLU (Huang et al. 1984; Kannangara et al. 1984, 1994) (Fig. 27.1a). Unlike class I aminoacyl-tRNA synthetases, GluRS avoids the aminoacyl-AMP formation in the absence of tRNA. In eukaryotic cells chloroplastic GluRS is post-translationally imported into the chloroplast where it ligates glutamate to tRNAGLU that contains the UUC anticodon (Schön et al. 1986, 1988).
GluTR, the second enzyme of the pathway, uses NADPH to reduce the activated α-carboxyl group of glutamyl-tRNA (Glu-tRNA) to synthesize glutamate 1-semialdehyde (GSA) (Hoober et al. 1988). The GluTR is a homo-pentamer of identical subunits of 54 kDa (Pontoppodian and Kannangara 1994). This enzyme is subject to feedback regulation by heme and appears to be a major control point of porphyrin biosynthesis (Kannangara et al. 1988). In A. thaliana GluTR interacts with FLU, a negative regulator of the Chl biosynthesis pathway (Meskauskiene et al. 2001; Meskauskiene and Apel 2002). FLU is a nuclear-encoded chloroplast protein, and the flu mutant has a higher level of ALA synthesis and protochlorophyllide (Pchlide) accumulation than that of wild-type plants. Probably FLU is a component of negative regulatory system for ALA synthesis when cells have high Pchlide contents. A FLU-like protein is also present in barley (Lee et al. 2003). GluTR is encoded by the HEMA gene. It has two isoforms in barley and cucumber, whereas in A. thaliana it has three isoforms.
The formation of 5-aminolevulinate/ALA from GSA is catalyzed by GSA-AT, the third and the last enzyme required for ALA biosynthesis. This enzyme is functionally an aminomutase, which transfers the amino group from carbon 2 of GSA to the neighboring carbon atom i.e., carbon 5 to form ALA (Fig. 27.1a). The enzyme is inhibited by gabaculine (Gough et al. 1992).
1. Environmental Modulation of ALA Biosynthesis
a. Light Regulation of ALA Biosynthesis
In cucumber and Arabidopsis thaliana, the HEMA1 gene is expressed in photosynthetic tissues and is induced by illumination, but no transcripts were detectable in roots (Tanaka et al. 1996; Ilag et al. 1994). Gene expression of HEMA1, and the corresponding protein abundance, increases in response to light treatment of dark-grown seedlings suggesting that an increased demand for Chl biosynthesis stimulates its expression and the gene promoter may have light-responsive elements (Mohanty et al. 2006). On the other hand, HEMA2 is preferentially expressed in non-photosynthetic tissues, and its expression is light-independent (Tanaka et al. 1996; Nagai et al. 2007). A third HEMA gene, HEMA3, has been identified in A. thaliana, but its expression is low (Matsumoto et al. 2004).
In A. thaliana light stimulates transcription of GSA (Ilag et al. 1994). The gene expression is also activated by the hormone kinetin (Yaronskaya et al. 2006). The expression of GSA and protein abundance of GSA-AT increases when etiolated seedlings are transferred to light demonstrating that it is a light-inducible gene that significantly contributes to Chl synthesis (Mohanty et al. 2006). In soybean also, the GSA is light inducible. It contains a light-regulated cis element (containing GAGA) that is found to be involved in transcriptional control (Frustaci et al. 1995). The mRNA level is high in soybean leaves (Sangwan and O’Brian 1993) whereas the mRNA is not detectable in roots (Frustaci et al. 1995).
b. Modulation of ALA Biosynthesis by Temperature
Environmental factors such as chill- or heat-stress influence gene expression, translation and post-translational modification of proteins involved in chloroplast biogenesis (Tewari and Tripathy 1998, 1999; Mohanty et al. 2006; Abdelkader et al. 2007a, b; Dutta et al. 2009). When 5-day old etiolated wheat seedlings grown at 25°C are transferred to 7°C (chill-stress), 42°C (heat-stress) or 25°C (control) and exposed to cool white fluorescent light (50 μmoles m−2 s−1) for 24 h, the Chl content gradually increases in control seedlings. In chill- and heat-stressed seedlings Chl biosynthesis is severely down-regulated. A lag period up to 12 h is observed, both in chill- and heat-stressed wheat seedlings before Chl accumulation accelerates (Fig. 27.2).
Biosynthesis of Chl (left panel (a)), Pchlide (central panel (b)), and ALA (right panel (c)) in control (25°C, squares), chill-stressed (7°C, diamonds), or heat-stressed (42°C, circles) cucumber seedlings. Each data point is the mean of three replicates; error bars represent SD. Missing error bars indicate that they are smaller than the symbols (Tewari and Tripathy 1998).
ALA synthesis in the presence of LA is almost linear up to 6 h of illumination in control and heat-stressed seedlings (Tewari and Tripathy 1998). For the first 3 h, ALA synthesis is completely inhibited in chill-stressed cucumber seedlings. As compared with the controls, the net synthesis of ALA is severely reduced in chill- and heat-stressed seedlings, respectively (Fig. 27.2) (Tewari and Tripathy 1998). Among ALA biosynthetic enzymes, the expression of HEMA is light-inducible in cucumber i.e., its expression increases in response to light in cucumber seedlings (Fig. 27.3). Its expression was down-regulated both in chill- and heat-stressed seedlings (Mohanty et al. 2006).
Modulation of gene expression (Northern blot) of Chl biosynthesis pathway enzymes by temperature stress in cucumber. Five-day old etiolated seedlings grown at 25°C were transferred to light (50 mmol m−2 s−1) or dark at 7°C, 25°C or 42°C for 24 h and Northern blotting was performed. Numbers denote temperature and D and L denote dark and light respectively (Mohanty et al. 2006).
The expression of GSA increases upon light exposure of etiolated control seedlings. However, heat-stressed etiolated seedlings display a higher GSA expression level than etiolated control seedlings. GSA expression further increases in illuminated heat-stressed seedlings and is significantly reduced in cold-treated cucumber seedlings (Fig. 27.3) (Mohanty et al. 2006).
Reduced ALA biosynthesis in cucumber at low temperature could be due to reduced gene and/or protein expression of two ALA biosynthetic enzymes GluTR and GSA-AT.
The reduced Chl synthesis was also reported in temperature-stressed maize/Pinus seedlings mostly due to down-regulation of early intermediates of Chl biosynthesis i.e., GSA and ALA (Hodgins and Van Huystee 1986; Hodgins and Oquist 2006).
c. Regulation of ALA Biosynthesis by Salinity
The Chl biosynthesis and chloroplast biogenesis are substantially regulated by salt-stress. ALA content was reduced in sunflower leaves on treatment with salt stress (Santos 2004) that may be due to reduction in the ALA precursor glutamate (Santos and Caldeira 1999; Santos et al. 2001).
d. Water-Stress and ALA Biosynthesis
In response to water-stress, Chl biosynthesis is down-regulated. The reduced Chl synthesis in water-stressed seedlings is mostly due to down-regulation of early intermediates of Chl biosynthesis i.e., GSA and ALA (Dalal and Tripathy 2012).
Reduced GSA synthesis in water-stressed rice seedlings is due to down-regulation of HEMA1 transcript abundance (Fig. 27.4). The protein/transcript abundance of GSA-AT increased (Fig. 27.4) in water-stressed rice seedlings, however the ALA contents declined suggesting that the GSA-AT, the next enzyme involved in ALA biosynthesis, may be inactivated by post-translational modification. These results show that the Chl biosynthesis pathway is down-regulated at the early steps under stress conditions to prevent the accumulation of harmful singlet oxygen generating tetrapyrroles (Dalal and Tripathy 2012).
Modulation of gene expression of chlorophyll biosynthetic enzymes due to water stress in seedlings of drought sensitive rice cultivar Pusa Basmati 1, after 24 h and 72 h of greening. Con denotes control and WS denotes water-stressed seedlings, respectively, that were treated with 50 mM PEG 6000, dissolved in nutrient solution, 16 h prior to transfer to 100 μmol m−2 s−1 white light (Dalal and Tripathy 2012).
Micromolar concentrations of Zn2+, Cu2+ and Cd2+ inhibit ALA biosynthesis in barley by impairing the activity of GluTR (Pontoppidan and Kannangara 1994).
B. Biosynthesis of Protoporphyrin IX
Protoporphyrin IX (PPIX) is synthesized from eight molecules of ALA by a series of enzymatic biochemical reactions that are largely common to plants and animals. PPIX synthesis involves several enzymes.
5-Aminolevulinic acid dehydratase (ALAD), also known as porphobilinogen (PBG) synthase is a homo-octameric metalloenzyme that catalyzes the condensation of two ALA molecules to form PBG (Fig. 27.1b) (Shemin 1976; Spencer and Jordan 1995). The ALAD of spinach is a hexamer with molecular weight of 300 kDa (Liedgens et al. 1980). The enzyme from radish cotyledons has a pH optimum of 8.0 (Shibata and Ochiai 1977) and requires Mg2+ and Mn2+ for activity. It is most active at slightly alkaline pH and shows a maximal binding of three Mg (II) per subunit (Kervinen et al. 2000).
The gene encoding ALAD is cloned from various plant sources. It has been isolated from pea, spinach, soybean and tomato.
The enzyme porphobilinogen deaminase (PBGD) is a soluble chloroplast protein (Castelfranco et al. 1988) that catalyzes the formation of the linear tetrapyrrole, hydroxymethylbilane, from four molecules of PBG (Fig. 27.1). Molecular weights from different plants range from 34 to 44 kDa. The PBGD gene has been isolated and cloned from pea (Witty et al. 1993) and A. thaliana (Lim et al. 1994). In A. thaliana, PBGD was found in both leaves and roots (Lim et al. 1994). Sequence comparison from different species shows that specific Arg and Cys residues are well conserved, and that these are implicated in catalysis and dipyrromethane cofactor binding (Witty et al. 1993). The synthesis and activity of PBGD are regulated by light and differ among cell types (Smith 1988; Shashidhara and Smith 1991; Spano and Timko 1991; He et al. 1994).
The Uroporphyrinogen III Synthase (UROS) enzyme, in concert with PBGD, catalyses formation of uroporphyrinogen III (Urogen III) from hydroxymethylbilane, a product of PBGD activity. This enzyme helps in maintaining the formation of the biologically active isomer III by inverting the ring D; in its absence, hydroxymethylbilane spontaneously cyclizes to uroporphyrinogen I (Urogen I). Inversion of ring D probably involves the production of a spiro-cyclic intermediate (Crockett et al. 1991). This enzyme has been purified from wheat germ (Higuchi and Bogorad 1975). The enzyme was found to be heat labile and the activity was enhanced by Na+ and K+. The enzyme PBGD and UROS may be present as a complex (Tsai et al. 1987). The UROS gene was isolated from A. thaliana (Tan et al. 2008). The localization of the protein in the chloroplast was confirmed by an in vitro protein import study and confocal microscopy (Tan et al. 2008). The barley uros mutant showed a necrotic phenotype in a developmental manner because of Urogen I accumulation (Ayliffe et al. 2009). The mutation in UROS also suppressed the expression of genes involved in the light reactions of photosynthesis (Ayliffe et al. 2009).
The Uroporphyrinogen III Decarboxylase (UROD) enzyme catalyzes stepwise decarboxylation of Urogen III to yield coproporphyrinogen III (Coprogen III). The enzyme catalyzes decarboxylation of all four carboxyl residues of Urogen III to yield Coprogen III. The order of Urogen III decarboxylation is substrate concentration dependent and under normal conditions enzymatic decarboxylation begins at the ring-D acetate group, in a clockwise manner (Luo and Lim 1993). Although all four isomers of Urogen are accepted by the enzyme, aromatic porphyrins are not decarboxylated (Castelfranco and Beale 1981). The discrimination between isomers, Urogen I and Urogen III for conversion into Coprogen occurs principally at the first step. Porphyrins, especially oxidation products of the substrates, have been shown to inhibit the enzyme (Smith and Francis 1981). The UROD was cloned from tobacco and barley (Mock et al. 1995). The in vitro translational product of UROD was imported into pea chloroplasts and processed into a 39 kDa product (Mock et al. 1995). Martin et al. (2001) reported the first crystal structure of a plant (tobacco) UROD.
Coproporphyrinogen oxidase (Coprox, CPOX) catalyses the oxidative decarboxylation of propionate side chains on ring A and B of Coprogen III to yield protoporphyrinogen IX (Protogen IX). In aerobic organisms, oxygen is utilized as the sole electron acceptor for enzymatic activity. The enzyme activity was found to be activated by Fe2+, Mn2+ and inhibited by EDTA and o-phenanthroline (Hsu and Miller 1970). The corresponding gene, CPOX, was isolated and characterized from soybean (Madsen et al. 1993), tobacco and barley (Kruse et al. 1995a, b), A. thaliana (Ishikawa et al. 2001) and maize (Williams et al. 2006). The CPOX mRNA is highly expressed in soybean root nodules followed by leaves, but no mRNA for CPOX was detectable in soybean roots (Madsen et al. 1993). The level of mRNA reached its maximum in developing cells and decreased drastically when cells were completely differentiated. The Coprox isoform, CPO1 fused with green fluorescent protein showed that it was localized in the plastids, whereas CPO2 appeared to localize to mitochondria (Williams et al. 2006). However, mitochondria lack CPOX activity (Smith 1988).
Protoporphyrinogen oxidase (Protox) catalyzes the oxygen-dependent aromatization of Protogen IX to protoporphyrin IX (PPIX, Proto IX). This enzyme catalyzes the six-electron oxidation of Protogen using a flavin cofactor, and molecular oxygen as terminal electron acceptor (Poulson and Polglasse 1974). Protogen is unstable and spontaneously undergoes oxidation in the presence of oxygen and its oxidation is enhanced by light (Jacobs and Jacobs 1979). Koch et al. (2004) reported on the crystal structure of mitochondrial Protox from tobacco and revealed that it contains an FAD-binding domain, a substrate-binding domain and a membrane-binding domain (Koch et al. 2004). Protox forms a loosely associated dimer that folds into an FAD-binding and substrate-binding domain. The substrate-binding domain of Protox also helps in forming a complex with the ferrochelatase enzyme. Protox has been purified from barley etioplasts (Jacobs and Jacobs 1987) and found to be localized in the envelope (stromal side) and thylakoid membranes (stromal side) of chloroplasts (Matringe et al. 1992a; Che et al. 2000). The envelope and thylakoid membranes fail to synthesize Proto IX from the substrate ALA, whereas the stromal fraction could synthesize a small amount. When however, all three components were mixed together the PPIX synthesizing capacity increased. The PPIX synthesizing capacity was reduced by oxidizing agents, and increased in the presence of reductants like dithiothreitol (DTT). ATP also increased PPIX synthesis (Manohara and Tripathy 2000).
Protox has been isolated from spinach, tobacco and A. thaliana (Narita et al. 1996; Lermontova et al. 1997; Che et al. 2000; Watanabe et al. 2001). In all these plant species, Protox was encoded by two genes, namely PPOX1 and PPOX2, and was found in both the chloroplast and mitochondria, respectively. In A. thaliana the levels of transcripts of plastid Protox were very high in leaves, whereas it was low in roots and floral buds (Narita et al. 1996). In tobacco, both transcripts were accumulated synchronously during diurnal and circadian growth (Lermontova et al. 1997). The spinach PPOX1 are preferentially localized to the stromal side of the thylakoid membrane and inner envelope membrane (Che et al. 2000). The spinach PPOX2 codes for two proteins of 59 kDa (PPOX2 L) and a 55 kDa (PPOX2 S) by using two in-frame start codons. PPOX2 L is associated with the chloroplast inner envelope membrane and PPOX2 S is associated with inner mitochondrial membranes (Watanabe et al. 2001). As it is folded into an extremely compact form, the Protox is highly resistant to proteases i.e., trypsin, endoproteinase Glu-C, or carboxypeptidases (Arnould and Camadro 1998). However, structurally bicyclic herbicides i.e., diphenyl ether-type herbicides, were shown to inhibit Protox activity in chloroplasts (Camadro et al. 1991; Matringe et al. 1992b). It has also been shown that the fungal toxin cyperin inhibits Protox activity (Dayan et al. 2008).
1. Developmental Modulation of ALAD
In cucumber and wheat ALAD expression increases upon transfer of etiolated seedlings to light (Mohanty et al. 2006). However, in pea, expression of ALAD is high in dark-grown samples as compared to light-grown samples (Li et al. 1991). In contrast, the corresponding protein level is significantly lower in dark-grown seedlings as compared to light-grown seedlings (He et al. 1994). ALAD is detectable in embryonic leaves whether the plants are grown in darkness or under continuous white-light illumination (He et al. 1994). In pea, ALAD transcript abundance is highly dependent on leaf developmental age; the transcript abundance increases with time until the leaf is fully expanded. Subsequently, its mRNA abundance decreases sharply (He et al. 1994). However, a significant amount of the protein is detected even in the matured leaves despite the mRNA abundance of ALAD being extremely low. The activity of ALAD significantly decreases during senescence (Hukmani and Tripathy 1994).
ALAD from tobacco leaves and radish cotyledons is inhibited by Zn2+ and Fe2+ (Shetty and Miller 1969; Shibata and Ochiai 1977), and arsenic inhibits its activity in maize leaves (Jain and Gadre 2004). PbCl2 and CdCl2 inhibit ALAD in Amaranthus lividus (Bhattacharjee and Mukherjee 2003).
2. Developmental Regulation of PBGD
The PBGD mRNA is slightly higher in the dark than in the light, even though the protein level is significantly lower in dark (He et al. 1994). The PBGD transcript abundance is dependent on leaf developmental age; i.e., the transcript abundance increases with increased age until the leaf is fully expanded and after that its mRNA level declines (He et al. 1994). Despite an extremely low level of PBGD mRNA, a significant amount of protein is detected even in matured leaves. Furthermore, PBGD activity rapidly declines during senescence (Hukmani and Tripathy 1994).
The enzyme is heat stable and maintains its activity at temperatures ranging from 55°C to 70°C. The PBGD enzyme from pea chloroplasts was inhibited by Fe2+, Mn2+ and Zn2+ whereas Ca2+ and Mg2+ were only weakly inhibitory at physiological concentrations (Spano and Timko 1991).
3. Environmental Regulation of UROD
The expression of the UROD gene and the corresponding protein level increase during illumination in barley (Mock et al. 1995) and cucumber (Mohanty et al. 2006).
In heat-stressed plants UROD activity, its gene and protein expression are substantially upregulated in heat-stressed seedlings whereas it is downregulated in chill-stressed plants (Tewari and Tripathy 1998; Mohanty et al. 2006) (Fig. 27.3).
The enzyme activity is inhibited by metals such as Fe2+, Co2+, Pb2+, Ni2+ and Mg2+ (Chen and Miller 1974), but stimulated by ATP (Manohara and Tripathy 2000).
4. Modulation of CPO
The CPO gene expression is not affected by light or heat stress in wheat and cucumber, however the gene expression is down-regulated by chill stress (Mohanty et al. 2006) (Fig. 27.3). In vitro protein import assays of tobacco and barley CPO protein showed that it was imported into the pea chloroplast and accumulated in the stroma. There are two isoforms of CPO (CPO1and CPO2) in maize. A. thaliana mutant defective in the lesion initiation 2 (LIN2) gene encoding CPO develops lesions on leaves, in a developmentally regulated and light-dependent manner (Ishikawa et al. 2001).
5. Environmental Modulation of Protox
The Protox activity substantially decreases in response to chill-stress (Tewari and Tripathy 1998). Its transcript and protein abundance decrease in water- stressed seedlings (Fig. 27.4) (Dalal and Tripathy 2012).
6. Environmental Regulation of Proto IX Biosynthesis
Proto IX biosynthesis is modulated by several environmental factors. In chill-stressed seedlings Proto IX synthesis from Urogen III is severely reduced whereas in heat-stressed seedlings the activity is substantially increased (Tewari and Tripathy 1998). Proto-IX synthesis from Coprogen III is reduced in chill-stressed seedlings, however in heat-stressed seedlings, Proto IX synthesis from Coprogen III is not affected (Tewari and Tripathy 1998). In heat-stressed seedlings, ALAD and PBGD were partially inhibited.
Reduced ALAD activity and gene expression were observed in water-stressed rice and chill- and heat-stressed cucumber seedlings (Figs. 27.3 and 27.4) (Mohanty et al. 2006; Dalal and Tripathy 2012). Limitation of ALA, a substrate for ALAD probably reduced its gene expression in water stress and other stress conditions. The increased or decreased availability of the substrate of the enzyme could positively or negatively regulate the gene expression of the enzyme. The enzymatic activity of PBGD that deaminates PBG to form Urogen III is reduced (Dalal and Tripathy 2012) due to down-regulation of its transcript abundance in water-stressed rice seedlings (Fig. 27.4).
UROD protein abundance decreases in water-stressed seedlings, which well correlates with the declined message abundance of UROD in response to water stress (Fig. 27.4). The UROD protein and transcript abundance also decline in chill-stressed wheat seedlings (Mohanty et al. 2006). This is in contrast to the earlier observations in cucumber and wheat where UROD activity and its transcript/protein abundance increased (Fig. 27.3) in response to heat-stress (Tewari and Tripathy 1998; Mohanty et al. 2006). The next two enzymes involved in Proto IX biosynthesis are Coprox and Protox. The enzyme activity of Coprox and Protox decreased in water-stressed rice and chill-stressed cucumber seedlings (Dalal and Tripathy 2012; Tewari and Tripathy 1998) due to down-regulation of their gene/protein abundance (Figs. 27.3 and 27.4).
C. Biosynthesis of Protochlorophyllide
The PPIX synthesis reactions are common to plants and animals. In green organisms the tetrapyrrole biosynthesis pathway branches to the formation of Fe-tetrapyrrole leading to synthesis of heme and Mg-tetrapyrrole that results in the synthesis of Chl. In green plants the Mg-branch involves insertion of Mg to PPIX by Mg-chelatase to synthesize Mg-PPIX, esterification of Mg-PPIX to Mg-protoporphyrin IX monomethylester (MPE) mediated by Mg-Protoporphyrin IX: S-adenosyl methione methyl transferase and formation of the isocyclic ring to synthesize Pchlide by MPE cyclase (Fig. 27.1c).
The insertion of Mg2+ into PPIX is catalyzed by Mg-chelatase to synthesize Mg-protoporphyrin IX (Mg-PPIX). In photosynthetic organisms, Mg-chelatase has three subunits (CHLI, CHLD and CHLH) and it catalyses the insertion of Mg2+ in two steps; an ATP-dependent activation that is followed by an ATP-dependent chelation step (Walker and Weinstein 1994; Walker and Willows 1997). The optimal ATP concentration for activation is found to be higher than that of chelation step. Out of its three subunits, CHLI is an ATPase and its ATPase activity is repressed when it forms a complex with CHLD (Jensen et al. 1999). The N-terminal halves of subunits CHLD and CHLI share high sequence similarity suggesting that the CHLD subunit is also an AAA + protein (ATPases Associated with diverse cellular Activities); however, its ATPase activity has yet not been detected (Jensen et al. 1999).
The CHLI/Chlorina9 has been cloned from soybean, barley (Jensen et al. 1996), A. thaliana (Gibson et al. 1996; Rissler et al. 2002), maize and rice (Zhang et al. 2006). This protein is localized to the stroma. In A. thaliana, most of the CHLI homozygous mutants have a pale green phenotype (Rissler et al. 2002). A second CHLI gene, CHLI-2 has been reported from A. thaliana (Rissler et al. 2002; Huang and Li 2009).
The N-terminus of CHLD shows structural similarities with the AAA domain of CHLI and therefore it is believed to contribute in complex formation and interaction with CHLH (Fodje et al. 2001). The CHLD/Chlorina1 cDNA sequence has been isolated and cloned from tobacco (Papenbrock et al. 1997) and rice (Zhang et al. 2006). The CHLD gene expression changes with respect to the diurnal changes in tobacco (Papenbrock et al. 1999). Virus-induced gene silencing of CHLH in tobacco led to the lowering of CHLD and CHLI mRNAs along with reduction in the Chl contents (Hiriart et al. 2002).
In tobacco, CHLH is strongly expressed in young leaves and less expressed in mature leaves and only traces of its transcripts were found in flowering organs (Kruse et al. 1997). CHLH expression was found to be light inducible in soybean and rice and its transcript levels were under the control of a circadian oscillation (Nakayama et al. 1998; Jung et al. 2003). CHLH protein was also found to be induced on transferring the Arabidopsis seedlings to white light from dark (Stephenson and Terry 2008). The CHLH transcripts undergo diurnal variation in A. thaliana and tobacco (Gibson et al. 1996; Papenbrock et al. 1999). Depending upon the concentration of Mg2+ in the lysis buffer, the CHLH protein migrated between stroma and the envelope membranes and was localized in the envelope membrane at very high concentrations of Mg2+ (Nakayama et al. 1998).
Mutants of CHLH have been isolated from A. thaliana (Mochizuki et al. 2001). Genome uncoupled 5 (GUN5) gene codes for CHLH subunits of Mg-chelatase. The rice CHLH mutants also showed a Chl-deficient phenotype (Jung et al. 2003; Zhang et al. 2006). In A. thaliana, the Mg-chelatase subunit CHLH regulates retrograde signaling (Mochizuki et al. 2001). Mutation in the CHLH gene results in the repressed expression of LHCB. Transgenic tobacco plants expressing antisense RNA for Mg-chelatase CHLH were Chl deficient (Papenbrock et al. 2000b). In these plants, less PPIX and heme accumulated, and a decrease in ALA synthesizing capacity was observed. A. thaliana protein GUN4 regulates Mg-chelatase activity (Larkin et al. 2003; Davison et al. 2005), and promotes the interactions between CHLH and chloroplast membranes and Chl biosynthesis (Adhikari et al. 2009, 2011).
S-adenosyl-L-methionine:Mg-PPIX methyltransferase (SAM-MgProtoMTF) catalyzes the conversion of Mg-PPIX to Mg-protoporphyrin monomethyl ester (MPE). It transfers a methyl group to the carboxyl group of the C13-propionate side chain of Mg-PPIX (Gibson et al. 1963) where SAM acts as a methyl group donor. The gene (CHLM) encoding for the SAM-MgProtoMT is cloned from A. thaliana (Block et al. 2002) and tobacco (Alawady and Grimm 2005). In tobacco, the methyltransferase physically interacts with the CHLH subunit of Mg-chelatases (Alawady et al. 2005).
The A. thaliana CHLM protein contains an N-terminal plastid transit sequence. The mature protein (without transit peptide) contains two functional regions, the N-terminal hydrophobic region that enhances the association of the protein with the envelope and thylakoid membranes and the C-terminal region that binds to Ado-met (Block et al. 2002). The A. thaliana, CHLM T-DNA insertion mutant shows albino phenotype; there is accumulation of Mg-PPIX and reduction in major Chl protein complexes in this mutant (Pontier et al. 2007). Down-regulation of the CHLM protein in antisense CHLM tobacco plants results in reduced ALA-synthesis and Mg-chelatase activities (Alawady and Grimm 2005).
Mg-protoporphyrin IX monomethylester cyclase catalyzes the formation of the isocyclic ring E of the Mg-protoporphyrins and converts MPE to Pchlide. There are two pathways for the formation of the isocyclic ring, i.e., aerobic cyclization and anaerobic cyclization. The former pathway is predominant in plants, green algae and cyanobacteria where the ketone oxygen of divinyl Pchlide (DV-Pchlide) is derived from molecular oxygen (Walker et al. 1989). The pH optimum of the cyclase activity is approximately 9.0 and the enzyme activity was found to be inhibited by CN− and N3 − (Whyte and Castelfranco 1993). The cyclase reaction with the two barley mutants xantha l and viridis K revealed that at least two plastid proteins (a membrane bound protein and a soluble protein) are required for cyclization (Walker et al. 1991; Walker and Willows 1997). Biochemical and genetic studies have demonstrated that the gene responsible for the xantha-l mutant encodes a membrane-bound cyclase subunit and it needs a soluble fraction for the cyclization reaction (Rzeznicka et al. 2005). Xantha l mutants accumulated less Chl, high MPE and had no cyclase activity (Rzeznicka et al. 2005). Three Xantha l mutants were characterized. In the leaky mutant xantha-l 35, a C-to-T point mutation resulted in an exchange of amino acid residue Ser-181 to Phe. In the non-leaky mutant xantha-l 81, a G-to-A point mutation resulted in the exchange of Gly-155 to Glu. Sequence alignment showed that Gly-155 is a highly conserved residue. In the non-leaky mutant xantha-l 82, point mutation resulted in truncation as a TGG codon corresponding to Trp-291 turned into a TGA stop codon and the truncated protein was not stable. However, the level of the protein found in the xantha-l 35 and xantha-l 81 mutants was similar as in the wild-type (Rzeznicka et al. 2005).
The gene responsible for the aerobic cyclization reaction has been isolated and characterized from different plants i.e., CHL27 from A. thaliana and xantha l from barley (Tottey et al. 2003; Rzeznicka et al. 2005). Antisense A. thaliana and tobacco plants with reduced amounts of CHL27 show chlorotic leaves with reduced abundance of all Chl-proteins and accumulate MPE (Tottey et al. 2003; Peter et al. 2010). The A. thaliana chl27 T-DNA mutant is pale green with an elevated Chl a/b ratio, and has unstacked thylakoid membranes with reduced LHCII protein. Their photosynthetic activity is reduced due to damaged Photosystem II (PS II) reaction centers (Bang et al. 2008; Hansson and Jensen 2009). In tobacco, the co-suppression of the NTZIP, which includes coding region for a di-iron motif, resulted in reduced Chl level and lower photosynthetic activity (Liu et al. 2004).
1. Environmental Regulation of Protochlorophyllide Synthesis
The accumulation of Mg-PPIX and pheophorbide inhibits Mg-chelatase activity in pea (Popperl et al. 1997). Mg-chelatase activity and the expression of the genes encoding this enzyme are up-regulated by light (Mohanty et al. 2006) (Fig. 27.3). The CHLI mRNA is induced by light (Gibson et al. 1996; Jensen et al. 1996; Nakayama et al. 1998) and constitutively expressed in matured leaves. It is also regulated by diurnal rhythm but not regulated by circadian rhythm (Matsumoto et al. 2004). CHLI could be a target for chloroplastic thioredoxin and the in vivo reduction process is light dependent (Ikegami et al. 2007).
The Mg-chelatase activity is severely down-regulated in chill- and heat- stressed cucumber seedlings (Tewari and Tripathy 1998). In wheat light treatment in control and heat-stressed seedlings leads to higher accumulation of CHLI/Chlorina9 transcripts. Its expression diminishes in cold- and heat-treated wheat seedlings. CHLI/Chlorina9 expression is also down-regulated by salt-stress in cucumber/wheat/rice seedlings.
Mg-chelatase activity is down-regulated in water-stressed rice seedlings (Dalal and Tripathy 2012) (Fig. 27.4). Moreover, the gene/protein expression of CHLI/Chlorina9 and CHLD/Chlorina1 subunits of Mg-chelatase (Zhang et al. 2006) partially declined in water-stressed seedlings. CHLI1 subunit is post-translationally regulated by chloroplastic thioredoxin (Ikegami et al. 2007) and therefore could have impaired function in altered redox environment in water-stressed seedlings. Stoichiometric imbalance among the subunits of Mg-Chelatase decreases the Mg-chelatase activity as seen in CHLI over-expressing or under-expressing transgenic Arabidopsis plants (Papenbrock et al. 2000a). Inadequate proportion of all subunits is known to hamper the correct assembly of active Mg-chelatase (Guo et al. 1998; Hansson et al. 1999; Jensen et al. 1999); therefore, nonstoichiometric abundance among its subunits may have led to decreased enzyme activity of Mg-chelatase in water stressed seedlings.
D. Phototransformation of Protochlorophyllide to Chlorophyllide
In angiosperms, Protochlorophyllide oxidoreductase (POR) has the obligate requirement of light for photo-converting Pchlide to chlorophyllide (Chlide). It catalyzes the conversion of Pchlide to Chlide in light using NADPH as reductant (Fig. 27.1c). POR converts Pchlide to Chlide, by adding two hydrogen atoms at C17 and C18 on ring D. In the POR catalytic cycle, a ternary enzyme-NADPH-Pchlide complex is formed. Light energy absorbed by the Pchlide in the complex may produce torsional strain in the molecule that provides a favorable condition for hydride/hydrogen transfer from NADPH (Begley and Young 1989). POR is a member of a large family of enzymes known as short chain dehydrogenases/reductases (SDR) (Wilks and Timko 1995), which generally catalyze NADP(H)- or NAD(H)-dependent reactions involving hydride and proton transfers. A tyrosine (Tyr) and a lysine (Lys) residues are both conserved throughout all members of the SDR family. In POR, it was also seen that Tyr and Lys residues are important for its activity (Wilks and Timko 1995; Lebedev et al. 2001). The Tyr may be deprotonated, acting as a general acid to facilitate hydride transfer to or from NAD(P)+/H (Bohren et al. 1994). The proton at the C-18 position of Pchlide is derived from Tyr and the hydride transferred to the C-17 position is derived from the pro-S face of NADPH. The close proximity of the Lys residue is thought to allow the deprotonation step to occur at physiological pH by lowering the apparent pK a of the phenolic group of the Tyr (Wilks and Timko 1995). A light-activated conformational change of the protein is necessary to activate catalysis (Heyes et al. 2008; Sytina et al. 2008). The fact that POR is light activated means that the enzyme–substrate complex can be formed in the dark. This has recently been exploited by studying Pchlide reduction at low temperatures to trap intermediates in the reaction pathway (Heyes et al. 2002, 2003; Heyes and Hunter 2004). As a result, the reaction has been shown to consist of at least three distinct steps: an initial light-driven step, followed by a series of ‘dark’ reactions. An initial photochemical step can occur below 200 K (Heyes et al. 2002), whereas two ‘dark’ steps were identified for Synechocystis sp. PCC 6803 POR, which can only occur close to or above the ‘glass transition’ temperature of proteins (Heyes et al. 2003). First, NADP+ is released from the enzyme and then replaced by NADPH, before release of the product (Chlide) and subsequent binding of Pchlide have taken place (Heyes and Hunter 2004). Monovinyl Pchlide (MV-Pchlide) and DV-Pchlide don’t influence differentially the enzyme kinetics or the steps involved in the reaction pathway (Heyes et al. 2006). The secondary structure analysis of POR reveals that it has 33% alpha helix, 19% beta-sheets, 20% turn and 28% random coil.
Mutation studies by Dahlin et al. (1999) showed that, mutation in predicted α-helical regions of the protein showed the least effect on enzyme activity, whereas mutations in the predicted β-sheet regions showed an adverse effect on enzyme function. The replacement of charged amino acids by alanine in the N- and C-terminal regions of the mature protein did not affect POR assembly, whereas mutations within the central core prevent proper attachment to the thylakoid. It is a peripheral membrane protein that accumulates to a high level in PLBs, where it forms a ternary complex with Pchlide and NADPH (Oliver and Griffiths 1982) and is present at low levels in the thylakoid membranes of developing and mature plastids. It is observed that the Cys residues of POR are crucial for its membrane association (Aronsson et al. 2001) and for NADPH and pigment binding (Townley et al. 2001; Reinbothe et al. 2006). The association of POR with Pchilde results in three different spectral forms of Pchlide based on their fluorescence emission maximum (in nm): Pchlide F631 (due to the pigment structural arrangements), Pchlide F644 (due to association of POR), and Pchlide F655 (due to localization in PLBs and/or prothylakoids) (Böddi et al. 1992, 1993). Spectroscopic studies of dark-grown bean seedlings indicated the existence of two forms of Pchlide, a main component with a red absorption band at 650 nm and a minor component absorbing at 636 nm (Shibata 1957). On the basis of flash illumination, two kinds of Pchlide can be categorized: one is transformed into Chlide and is called photoactive Pchlide, whereas the other remains unchanged and is called nonphotoactive Pchlide. The latter is assembled into various complexes with different molecular structure and spectral properties (Schoefs and Franck 2003; Masuda and Takamiya 2004). Plastids isolated from dark-grown wheat seedlings exhibit a smaller 77 K fluorescence emission peak at 632 nm due to non-phototransformable Pchlide and a larger peak at 657 nm due to phototransformable Pchlide. The non-phototransformable Pchlide emitting at 632 nm is due to a monomeric Pchlide complex or esterified Pchlide i.e., Protochlorophyll (Lindsten et al. 1988), which spontaneously dimerizes to form (POR-Pchlide-NADPH)2. The short-wavelength, monomeric Pchlide is not flash-photoactive: instead, it regenerates the long wavelength Pchlide forms (Schoefs and Franck 1993; He et al. 1994; Schoefs et al. 1994, 2000a, b). The dimer has an absorption maximum at 638 nm and an emission maximum at 645 nm (Lebedev and Timko 1999). The dimeric POR-Pchlide-NADPH complex further polymerizes to form 16-mer or larger aggregates of POR-Pchlide-NADPH complex i.e., (POR-Pchlide-NADPH)n having absorption maximum at 650 nm and emission maximum at 657 nm (Böddi et al. 1989; Wiktorsson et al. 1993) and is flash photoactive (Böddi et al. 1991). However, illumination for more than a minute usually converts non-active Pchlide to photo-active Pchlide.
Full-length cDNA clones of POR were isolated from barley (Holtorf et al. 1995), pea (Spano et al. 1992), A. thaliana (Armstrong et al. 1995; Oosawa et al. 2000), tobacco (Masuda et al. 2002), cucumber (Kuroda et al. 1995) and many other higher plants. The high degree of sequence similarity among PORs from different taxonomic groups implies a common mechanism of enzyme action.
A characteristic feature of POR accumulating in darkness is its sensitivity to illumination. The POR mRNA expression was also decreased (Santel and Apel 1981). Red and far-red light treatment also inhibit POR gene expression indicating that POR expression is controlled by phytochrome (Mosinger et al. 1985). The negative effect of light on the POR enzyme and its mRNA was observed in different dicotyledons like bean, pea, tomato and A. thaliana (Forreiter et al. 1991; Spano et al. 1992; Armstrong et al. 1995) and in the monocotyledonous plants maize and barley (Forreiter et al. 1991; Holtorf et al. 1995). However, some flowering plants have isoforms of POR. In A. thaliana, (Armstrong et al. 1995; Oosawa et al. 2000; Su et al. 2001; Pattanayak and Tripathy 2002), barley (Holtorf et al. 1995; Holtorf and Apel 1996) and tobacco (Masuda et al. 2002) there are different PORs present. The N-termini of PORA and PORB of barley etioplasts have recently been characterized (Plöscher et al. 2009). In A. thaliana there are three isoforms of POR, namely PORA, PORB and PORC. These three isoforms are differentially regulated by light. The level of PORA mRNA and protein decrease upon illumination of etiolated plants (Holtorff and Apel 1996) while that of PORC increases and was dominantly expressed in both mature and immature tissues (Oosawa et al. 2000). PORB transcript and PORB protein levels remain constant in both darkness and upon illumination (Armstrong et al. 1995; Holtorf et al. 1995; Holtorf and Apel 1996). Both PORB and PORC of A. thaliana exhibit diurnal fluctuation but only the PORB mRNA of A. thaliana exhibits circadian regulation (Su et al. 2001). PORC mRNA and PORC protein levels also increased under high light intensity (Su et al. 2001; Masuda et al. 2003). In cucumber the levels of the POR mRNA increased in etiolated cotyledons when they were illuminated with continuous light (Kuroda et al. 1995; Fusada et al. 2000). The plant hormone cytokinin regulates cucumber POR gene expression by binding to the cis-elements present at the 5′ region of the POR promoter (Fusada et al. 2005). In tobacco, two POR isoforms have been isolated, the expression of which was not negatively regulated by light, persisted in mature green tissue and showed diurnal fluctuations with a similar oscillation phase (Masuda et al. 2002).
A plant specific downstream element in the 3′ untranslated region of the PORA transcript confers PORA mRNA instability, where as it was not responsible for PORB mRNA degradation (Holtorf and Apel 1996). POR gene expression in cucumber is regulated by phytohormones, particularly by cytokinins and abscisic acid (Kuroda et al. 2001). In the lip1 mutant of pea, cytokinins restored the formation of PLB and photoactive Pchlide in the dark (Seyedi et al. 2001a), but in A. thaliana their application results in loss of PLBs (Chory et al. 1994). In lupine, POR gene expression is also regulated by cytokinins and abscisic acid (Kusnetsov et al. 1998).
POR gene expression is also organ specific. A. thaliana PORB and PORC are expressed in all photosynthetic tissues of the mature plants but not in the root (Armstrong et al. 1995; Oosawa et al. 2000). Cucumber POR gene expression is also observed in photosynthetic tissues (Kuroda et al. 1995). Plant age plays a crucial role in POR gene expression. In A. thaliana and barley PORA expression is only observed in young seedlings whereas PORB is expressed both in young and matured green tissue (Armstrong et al. 1995). In A. thaliana both PORB and PORC expression is observed in green tissue (Oosawa et al. 2000; Su et al. 2001). In the leaves of dark-grown seedlings, the highest level of expression is observed 8–10 days after germination of seedlings (Spano et al. 1992). The transcript level of pea POR did not decrease after 48 h of light exposure. Immunoblot analysis showed that there was no POR protein detectable after 48 h of light exposure. These results suggested that pchlide reductase activity in pea is primarily regulated post-transcriptionally, most likely at the level of translation initiation/elongation or protein turnover (Spano et al. 1992).
Far-red-light modulation of POR: Etiolated seedlings of A. thaliana grown under continuous far-red light are unable to green when subsequently transferred to white light; this is called far-red blocking of the greening (Buhr et al. 2008). This process involves depletion of PORA, partial depletion of PORB and the concomitant loss of PLBs resulting in photo-oxidative damage (Barnes et al. 1996; Runge et al. 1996). From these studies, PORA has been proposed to play a special role in the formation of POR ternary complexes containing photoactive Pchlide-F655, PLB assembly, and protection against photo-oxidative damage caused by non-photoactive Pchlide (Reinbothe et al. 1999). Overexpression of PORA and PORB in specific mutants overcame the photooxidative damage (Sperling et al. 1997, 1998).
Overexpression of a cyanobacterial POR protein in the A. thaliana porA mutant could restore prolamellar body formation. However, the amount of photoactive Pchlide in the etioplasts of the complementing lines was retained at a low level as in the parent PORA knockdown mutant (Masuda et al. 2009). The lip1 mutant of pea lacked PLBs but could form prolamellar bodies if treated with cytokinin (Seyedi et al. 2001a): however unlike the A. thaliana mutant, it did not undergo photooxidative damage (Seyedi et al. 2001b). The physiological function of specific POR isoforms in vivo has been well characterized in knockout mutants of A. thaliana (Frick et al. 2003; Masuda et al. 2003). Single POR mutants display no obvious phenotypes at the whole plant or chloroplast ultra-structural levels, except that etiolated PORB mutants have less extensive inner membranes. However, the PORB/PORC double mutant, which displayed a seedling-lethal xantha phenotype at the cotyledon stage, contained only a small amount of Chl a, and possessed chloroplasts with mostly unstacked thylakoid membranes (Frick et al. 2003). Masuda et al. (2003) focused on the greening process of por mutants, and showed that the etiolated PORB mutant seedlings were able to green to a similar extent as the wild type, and the greening of the PORC mutant was repressed under high light conditions.
From a molecular evolutionary perspective, the light-dependent POR (LPOR) enzymes are extraordinarily highly conserved. Comparative analysis of complete plastid genome sequences indicate that LPOR genes were lost from the plastid at some point during early evolution (Martin et al. 2002), and analysis of LPOR proteins in species of conifer shows evidence for loss of enzyme activity (Kusumi et al. 2006). The discovery of genes for LPOR in the plastid genomes of diverse cryptophyte algae suggests that these genes have been lost relatively recently.
In all photosynthetic organisms Pchlide and Chlide are originally formed as 3,8-divinyl derivates. The 8-vinyl reductase reduces the 8-vinyl group on the tetrapyrrole to an ethyl group using NADPH as the reductant. This enzymatic activity has been detected in isolated chloroplasts of barley (Tripathy and Rebeiz 1988), plastid membranes from cucumber (Parham and Rebeiz 1995), and also in solubilized crude extracts derived from etiolated barley leaves (Kolossov and Rebeiz 2001). It has been demonstrated in vitro that the monovinyl (MV) and divinyl (DV) Chl biosynthesis reactions may operate in parallel (Tripathy and Rebeiz 1986). However, the mutant of maize (Zea mays) that accumulated only DV-Chl instead of MV-Chl and capable of photosynthetic growth with DV-Chl suggests that a single gene product is responsible for the reduction of the vinyl group of Chlide (Bazzaz 1981). Nagata et al. (2005) followed by Nakanishi et al. (2005) isolated a mutant of A. thaliana which accumulates DV- Chl. By map-based cloning they detected that the gene is 8-vinyl reductase. The recombinant protein was successfully tested for the conversion of the C8-vinyl group of Chlide to an ethyl group on ring B. The 3,8-divinyl-chlide a is the major substrate of divinyl reductase (DVR) (Nagata et al. 2007). Starch granules were not found in the mutant chloroplasts, suggesting the reduction of photosynthetic activity in the mutant (Nakanishi et al. 2005). The transcript level of DVR expression is high in leaves, stems and flower buds, and low in roots.
The mutant is pale green and the Chl a/b ratio varies in between 6 and 10 depending on the developmental stage and growth conditions. This mutant is capable of photosynthesizing and growing under low-light conditions (70–90 μmol photons m−2 s−1); but it rapidly dies under high light conditions (1,000 μmoles photons m−2 s−1) (Nagata et al. 2005). The thylakoid membranes were in a disorderly fashion having no distinct grana stacks in the mutant and no significant differences in the size and the number of chloroplasts between the wild type and the mutants were observed.
1. Role of POR in Combating Oxidative Stress
Light absorbed by colored intermediates of Chl biosynthesis is not utilized in photosynthesis. Instead, it is transferred to molecular oxygen, generating singlet oxygen (1O2) (Chakraborty and Tripathy 1992). As there is no enzymatic detoxification mechanism available in plants to destroy 1O2, its generation should be minimized. The concentration of a major Chl biosynthetic intermediate, i.e., Pchlide in Arabidopsis was manipulated by overexpressing the light-inducible PORC that effectively phototransforms endogenous Pchlide to Chlide leading to minimal accumulation of the photosensitizer Pchlide in light-grown plants (Pattanayak and Tripathy 2011). In PORC overexpressing (PORCx) plants exposed to high-light, the 1O2 generation and consequent malonedialdehyde production was minimal and the maximum quantum efficiency of photosystem II remained unaffected (Fig. 27.5) demonstrating that their photosynthetic apparatus and cellular organization were intact.
Morphological and physiological responses of WT and PORCx (T-13) plants to light stress. Both WT and T-13 plants were grown in moderate light (100 μmoles photons m−2 s−1) for 22–24 days and subsequently transferred to low-light (LL) (50 μmoles photons m−2 s−1, 16 h light/8 h dark) or high-light (HL) (330 μmoles photons m−2 s−1, 16 h light/8 h dark) regimes for 6-7 d as described in experimental procedures. (a) Photosynthetic efficiency (Fv/Fm) of leaves of LL- and HL-exposed plants was monitored by PAM 2100 fluorometer. Values are mean ± SD (n = 20). (b) Anthocyanin contents of WT and T-13 plants grown under HL. (c) The gene expression study of CHS in HL-grown WT and T-13 plants was done by RT-PCR as described in experimental procedures. AtACT1 was used as an internal control. (d) Pchlide contents of HL-treated WT and T-13 plants measured 10 min after the end of dark period. (e) Singlet oxygen (1O2) contents in WT and T-13 plants. Thylakoid membranes were isolated in complete darkness from HL-exposed plants and the 1O2 production were determined in terms of RNO bleaching using histidine as a trap. (f) Malondialdehyde (MDA) production in HL-treated WT and T-13 plants. Each data point represented in all the above experiments is the average of six replicates. The error bar represents SD. (Adopted from Pattanayak and Tripathy 2011).
Further, PORCx plants treated with 5-aminolevulinicacid, when exposed to light, photo-converted over-accumulated Pchlide to Chlide, reduced the generation of 1O2 and malonedialdehyde production and reduced plasma membrane damage (Fig. 27.6). So PORCx plants survived and bolted whereas, the ALA-treated wild type plants perished. Thus, overexpression of PORC could be biotechnologically exploited in crop plants for tolerance to 1O2-induced oxidative stress, paving the use of ALA as a selective commercial light-activated biodegradable herbicide (Pattanayak and Tripathy 2011).
Contents of chlorophyll biosynthetic pathway intermediates and singlet oxygen production in ALA-treated WT and PORCx (T-13) plants. WT and T-13 plants grown for 28–32 days at 220C ± 20C under 14 h L/10 h D photoperiod (100 μmoles photons m−2 s−1) were sprayed with ALA (3 mM), dark incubated for 14 h and exposed to light (100 μmoles photons m−2 s−1) for 10 min. Leaves were harvested both from dark incubated and light exposed plants, homogenized and their tetrapyrrole contents were monitored spectrofluorometrically. (a) Pchlide, Proto IX and MP(E) contents of ALA-treated (3 mM) and 14 h-dark-incubated WT and T-13 plants. (b) After dark incubation both WT and T-13 plants were exposed to light (10 min) and their Pchlide, Proto IX and MP(E) were determined. (c) 1O2 contents in ALA-treated (+ALA) and untreated (−ALA) WT and T-13 plants. The experiments were repeated five times and each data point is the average of five replicates. The bars represents ± SD (Pattanayak and Tripathy 2011).
Reduced Pchlide content in PORCx plants released the Pchlide-mediated feed-back inhibition of ALA biosynthesis that resulted in higher ALA production. Increase of ALA synthesis up-regulated gene expression and protein level of several downstream Chl biosynthetic enzymes elucidating a regulatory net work of expression of genes involved in ALA and tetrapyrrole biosynthesis (Pattanayak and Tripathy 2011).
2. Environmental Modulation of Shibata Shift
In etiolated wheat seedlings, the phototransformable Pchlide peak (F657) is substantially higher than the non-phototransformable peak (F632) (Fig. 27.7a) demonstrating the presence of large aggregates of POR-Pchlide-NADPH ternary complexes. Although chilling arrested de novo synthesis of the Chl biosynthetic intermediate Pchlide (Tiwari and Tripathy 1998), it did not affect the ratio of non-phototransformable to phototransformable Pchlide suggesting that low temperature did not affect formation of large aggregates of POR-Pchlide-NADPH ternary complexes (Fig. 27.7a) (Mohanty and Tripathy 2011). In contrast, although heat-stress partially arrested Pchlide synthesis (Fig. 27.7a) in etiolated seedlings (Tewari and Tripathy 1998), it substantially affected the aggregation state of POR-Pchlide-NADPH complex as indicated by near absence of 657 nm peak in etiolated heat- stressed seedlings. Within hours of exposure of etiolated seedlings grown at 25°C to heat-stress, a progressive decline of the 657 nm phototransformable peak was observed suggesting the disaggregation of existing large aggregates of POR-Pchlide-NADPH polymeric complexes present in prolamellar bodies to monomeric or dimeric forms. The rate of degradation of polymeric complexes must exceed their formation, if any, in heat-stressed samples. Unlike heat-stress, in vivo application of salt-stress to excised wheat leaves reduced the peak of non-photo-transformable Pchlide and did not affect the longer wavelength phototransformable forms suggesting that prothylakoids rather than prolamellar bodies were affected by salinity (Abdelkader et al. 2007a).
(a) Low temperature (77 K) fluorescence emission spectra (E440) of plastids isolated from 6-day old etiolated control, (upper panel) chill-stressed (middle panel) and heat-stressed (lower panel) wheat seedlings showing Shibata-shift. Five-day old seedlings grown at 25°C were transferred to 7°C and 42°C in dark for 24 h. Low temperature fluorescence emission spectra were recorded before flash, immediately after flash (0.2 s) and after 1 min and 15 min post-flash incubation (Mohanty and Tripathy 2011). (b) Low temperature (77 K) fluorescence emission spectra (E440) of leaves from 6-d old etiolated control (upper panel) and water-stressed (lower panel) rice (PB1) seedlings, showing Shibata-shift. For water-stress, seedlings were treated with 50 mM PEG 6000, dissolved in nutrient solution, 16 h prior to taking spectra. Low temperature fluorescence emission spectra were recorded before the flash, immediately after a flash of 0.2 s and after 1 min and 15 min post-flash incubation (Dalal and Tripathy 2012).
In addition to changes in the aggregation status of polymeric POR-Pchlide-NADPH complexes, the flash-induced photo-transformation and the Shibata shift (Shibata 1957) leading to chloroplast biogenesis is substantially affected in temperature-stressed samples. Upon flash illumination (0.2 s) of etioplasts isolated from control seedlings the phototransformable Pchlide peak at 657 nm emanating from large aggregates of polymeric POR-Pchlide-NADPH complexes almost disappeared due to photo-reduction of Pchlide to Chlide. Transformation of Pchlide655 into Chlide692 was observed by exposing the leaf primordia of common ash (Fraxinus excelsior L.) and Hungarian ash (Fraxinus angustifolia Vahl.) (Solymosi and Böddi 2006) and that of Horse chestnut (Aesculus hippocastanum) (Solymosi et al. 2006) to a white light flash of 10 s. After 1 and 10 min of illumination, the peak at 692 nm slowly blue shifted to 676 nm (Shibata shift) (Fig. 27.7b). In chill-stressed and heat-stressed seedlings the Shibata shift was significantly arrested (Fig. 27.7a) (Mohanty and Tripathy 2011) probably due to the disaggregation of the PLB membrane particles or of the POR units as well as by their conformational changes (Böddi et al. 1990; Smeller et al. 2003).
Heating of excised etiolated barley leaves resulted in decreased accumulation of Pchlide(650), and a flash could trigger the formation of Chlide(672) instead of the formation of Chlide(684) (Eullaffroy and Popovic 1997). Similar to heat-stress, water-stress affects the Shibata shift, although the phototransformation of Pchlide to Chlide was not impaired by water deficit (Fig. 27.7b) (Le Lay et al. 2001). Both chill-stress and heat-stress affected the Shibata shift (Fig. 27.7a). This is variance to in vitro heating (40°C) of excised barley leaves where the Shibata shift was not affected (Eullaffroy et al. 1995).
Upon 16 h of water-stress treatment, the etiolated seedlings displayed an emission fluorescence peak (77 K) at 632 nm due to non-phototransformable Pchlide and a peak at 657 nm due to phototransformable Pchlide (Fig. 27.7b). However, as compared to the control, the ratio of non-photo-transformable/photo-transformable Pchlide (F632/F657) increased from 0.10 to 0.15 in stressed seedlings suggesting an impairment of aggregation of monomeric POR-Pchlide-NADPH to 16-mer or larger aggregates of POR-Pchlide-NADPH complex, i.e. (POR-Pchlide-NADPH)n. This may be due to reduced assembly or due to degradation of polymeric complexes in the stressed environment (Dalal and Tripathy 2012).
The flash-induced photo-transformation and Shibata shift leading to chloroplast biogenesis was substantially affected in 16 h-water stressed samples. Upon red light flash illumination (0.2 s) of control leaves the phototransformable Pchlide peak at 657 nm emanating from large aggregates of polymeric (POR-Pchlide-NADPH)n complexes almost disappeared due to photo-reduction of Pchlide to Chlide, and a new peak appeared at 691 nm due to formation of Chlide-LPOR-NADP+ complexes (El Hamouri et al. 1981; Oliver and Griffiths 1982; Franck 1993; Wiktorsson et al. 1993; Franck et al. 1999). Transformation of Pchlide 658 into Chlide692 was observed by exposing the leaf primordia of common ash (Fraxinus excelsior L.) and Hungarian ash, Fraxinus angustifolia Vahl. (Solymosi et al. 2006), wheat (Franck et al. 1999) and that of Horse chestnut (Aesculus hippocastanum) (Solymosi et al. 2006) to light flash. One min after flash, 691 nm-peak shifted to 694 nm (Fig. 27.7b) in control leaves due to the formation of Chlide-LPOR-NADPH complexes (El Hamouri et al. 1981; Oliver and Griffiths 1982; Franck et al. 1999). Subsequently, this peak blue-shifted to 680 nm after 15 min post-flash incubation of control leaves due to the release of Chlide from the active site of LPOR and disaggregation of multimeric complexes, a process called Shibata shift (Shibata 1957; Böddi et al. 1990; Franck 1993).
In water-stressed leaves, upon red light flash illumination of etiolated leaves the phototransformable pchlide peak at 657 nm disappeared and a new peak appeared at 692 nm due to formation of Chlide-LPOR-NADP+ complexes (El Hamouri et al. 1981; Oliver and Griffiths 1982) demonstrating that phototransformation of Pchlide to Chlide could still take place in 16 h-water-stressed samples (Dalal and Tripathy 2012). After 1 min post-flash incubation this peak shifted to 694 nm due to the formation of Chlide-LPOR-NADPH complexes (Fig. 27.7b). In water stressed leaves the shift to lower wavelengths was substantially delayed. A shoulder appeared at 680 nm after 15 min of dark incubation, in contrast to complete shift to 680 nm in control seedlings, suggesting a slow release of Chlide from the active site of LPOR (Shibata 1957; Böddi et al. 1990). In a non-physiological environment i.e. after desiccation of detached barley leaves a slowdown of Shibata was earlier reported (Le Lay et al. 2000, 2001). Upon 15 min of dark incubation after flash illumination, a good amount of phototransformable Pchlide (F657) was regenerated in control seedlings (Fig. 27.7b) and substantially less in water-stressed seedlings (Fig. 27.7b) demonstrating the down-regulation of synthesis of Pchlide and its conversion to photo-transformable form.
E. Synthesis of Chlorophyll a and Chlorophyll b
Chlorophyllide a oxygenase (CAO) catalyzes the oxidation of Chlide a to Chlide b (Fig. 27.1d). During conversion of Chlide a to Chlide b the electron is transferred from the Rieske center to the mononuclear iron with subsequent activation of molecular oxygen for oxygenation of the Chlide a methyl group (Beale and Weinstein 1990; Porra et al. 1993). Chlide b is synthesized by oxidation/conversion of the methyl group on the D ring of the porphyrin molecule to a formyl group at that position. The CAO enzyme contains domains for a [2Fe-2S] Rieske center and for a mononuclear nonheme iron-binding site and has a tyrosine radical (Eggink et al. 2004). The conserved Rieske center and non-heme-iron binding motifs of CAO are likely to be involved in electron transport from ferredoxin to molecular oxygen. The recombinant CAO protein catalyzes chlide a to chlide b in the presence of NADPH and reduced ferredoxin (Oster et al. 2000). However, Pchlide a is not a substrate for the CAO enzyme (Oster et al. 2000).
The CAO was first cloned by Tanaka et al. (1998) from Chlamydomonas and also has been cloned from A. thaliana (Espineda et al. 1999) and rice (Lee et al. 2005). Both transcript and protein level of CAO increased when A. thaliana plants were transferred from moderate to shade light (Harper et al. 2004). Rice has two CAO isoforms namely OsCAO1, OsCAO2 that are differentially regulated in light and dark. The level of the OsCAO1 transcript is less in the dark and is higher in the light whereas the OsCAO2 mRNA levels are higher in dark conditions and are reduced by exposure to light (Lee et al. 2005).
Overexpression of the CAO gene in A. thaliana led to an increase in the Chl b levels leading to reduction of the Chl a/b ratio from 2.85 to 2.65 in full green rosette leaves and at the same time there was 10–20% increase in antenna size (Tanaka et al. 2001). Overexpression of A. thaliana CAO in Synechosystis sp. PCC 6803 resulted in production of Chl b up to about 10% of total Chl content and the resulting Chl b pigments were efficiently incorporated into the Photosystem I Chl-protein complex (Satoh et al. 2001). Simultaneous overexpression of both CAO and LHC II genes in Synechosystis sp. PCC 6803 resulted in an increase in Chl b content up to 80% of total Chl (Xu et al. 2001). High light grown transgenic A. thaliana plants also showed decreased Chl a/b ratio under high light (Tanaka and Tanaka 2005). When the CAO gene of Prochlorothrix hollandica was overexpressed in A. thaliana, it was observed that approximately 40% of Chl a of the core antenna complexes was replaced by Chl b in both photosystems (Hirashima et al. 2006). The CAO sequence has been divided into four parts, the N-terminal sequence predicted to be a transit peptide, the subsequent conserved sequence unique in land plants (A-domain), a less-conserved sequence (B-domain) and the C-terminal conserved sequence common in chlorophytes and prochlorophytes (C-domain) (Nagata et al. 2004). The C-domain is sufficient for catalytic activity and the N-terminal ‘A’ domain confers protein instability by sensing the presence of Chl b and regulates the accumulation of the CAO protein (Yamasato et al. 2005). Chloroplast Clp protease is involved in regulating Chl b biosynthesis through the destabilization of CAO in response to the accumulation of Chl b (Nakagawara et al. 2007). The B domain alone is not involved in the regulation of CAO protein levels (Sakuraba et al. 2007). Further work on domain analysis also indicated that transgenic A. thaliana plants overexpressing CAO from which the A-domain had been deleted, accumulated an excess amount of Chl b during greening and the etiolated transgenic plants either died or were retarded when exposed to continuous light immediately after etiolation (Yamasato et al. 2008). This was most likely due to deregulated Chl b synthesis that reduced the energy transfer rate between photosynthetic pigments (Sakuraba et al. 2010).
Chl synthetase encoded by CHLG catalyzes the esterification of Chlide a and Chlide b to Chl (Fig. 27.1d) (Rüdiger et al. 1980). Pchlide is not the substrate for this enzyme, which indicates that reduction of the 17, 18 double bond on ring D is essential for esterification (Benz and Rüdiger 1981b). Compounds which have the 13(2)-carbomethoxy group at the same side of the macrocycle as the propionic side chain of ring D are neither substrates nor competitive inhibitors (Helfrich et al. 1994). Only compounds having the 13(2)-carbomethoxy group at the opposite site are substrates for the enzyme. Esterification of Chlide is a rapid phase, leading to esterification of 15% of total Chlide within 15–30 s, followed by a lag-phase of nearly 2 min and a subsequent main phase (Schmid et al. 2002). It has been shown that the conversion of Chlide to Chl is a four-step process including three intermediates, i.e., Chlide geranylgeraniol, Chlide dihydrogeranylgeraniol and Chlide tetrahydrogeranylgeraniol before the formation of Chlide phytol or Chl (Schoefs et al. 2000a, b).
In etioplasts, geranyl-geranyl pyrophosphate (GGPP) is used as a substrate (Rüdiger et al. 1980), while in chloroplasts the preferential substrate is phytyl diphosphate (PhPP) (Soll et al. 1983). Chl synthetase in chloroplast thylakoid membranes incorporates phytol in the presence of ATP and a stromal kinase (Benz and Rüdiger 1981a). The enzyme was not affected by the developmental stage of the plastids. In etiolated wheat, the enzyme was found in latent form in PLBs (Lindstein et al. 1990).
The CHLG gene was isolated from A. thaliana, Avena sativa, rice and tobacco (Gaubier et al. 1995; Schmid et al. 2001; Wu et al. 2007; Shalygo et al. 2009). In A. thaliana, the CHLG transcript has been detected in green or greening tissues (Gaubier et al. 1995), whereas in A. sativa, the gene is expressed equally both in dark- and light-grown seedlings (Schmid et al. 2001). Sequence analysis of cDNAs from rice yielded a putative Chl synthase homolog (Scolnik and Bartley 1996); however, the biochemical properties and physiological functions remained unknown until Wu et al. (2007) characterized a rice mutant with inactivated CHLG. The young rice Chl synthase mutant plants have yellow-green leaves with decreased Chl synthesis (Wu et al. 2007). In the mutated plants, there is accumulation of tetrapyrrole intermediates, reduced expression of LHCB1 and delayed chloroplast development.
1. Modulation of Chlorophyll b Synthesis Confers Tolerance to Low Light and High Light
Overexpression of CAO in tobacco plants resulted in a decreased Chl a/b ratio i.e., from 3.38 in wild-type plants to 2.33 in transgenic plants when grown in high light and from 2.8 to 2.4 in low light-grown plants (Pattanayak et al. 2005). The overexpression of full length CAO in tobacco (Nicotiana tabacum) resulted in an increased Chl synthesis and a decreased Chl a/b ratio in low-light-grown (LL) as well as in high-light-grown (HL) tobacco plants; this effect was more pronounced in HL-plants. The potential of Chl biosynthesis and the POR activity increased compensating for the usual loss of Chl when plants were grown in high light. Increased Chl b synthesis in CAOx plants was accompanied by an increased abundance of light-harvesting chlorophyll-proteins (LHCPs) and other proteins of electron transport chain that led to an increase in capture of light, as well as enhanced (40–80%) electron transport rates of Photosystem I and Photosystem II at both limiting and saturating light intensities. However, the increase in the whole chain electron transport was somewhat lower (20–50%). The light-saturated photosynthetic carbon assimilation, starch content and the dry matter accumulation increased in CAOx plants grown in both low and high-light regimes (Figs. 27.8 and 27.9).
Photosynthesis (net CO2 assimilation rate) light response curves and quantum yield of leaves from attached WT and CAOx plants grown in LL and HL intensities. (a) Net CO2 assimilation rates of attached leaves of WT and CAOx plants were monitored by IRGA (Licor 6400-XT portable photosynthetic system) in ambient CO2 at different light intensities. Light response curves were measured up to 1,800 μmol of photons m−2 s−1 at 28°C. (b) Relative quantum yield of CO2 fixation by leaves from WT and CAOx plants grown in LL or HL regimes. Quantum yield was measured from the above photosynthetic rate after the chamber reached to a steady-state. Light intensity curves at LL-intensities upto 80 μmol of photons m−2 s−1; the slopes of these curves provide relative quantum yield of CO2 fixation by leaves. Leaves were pre-exposed for 15 min at 700 μmol photons m−2 s−1 and 200 μmol photons m−2 s−1 for LL and HL grown plants respectively prior to CO2 assimilation measurement. These experiments were done three times with similar results. Each data point is the average of five replicates and the error bar represents SE. Asterisks indicate significant differences determined by ANOVA followed by Tukey’s test (*P < 0.05) (Biswal et al. 2012).
Diurnal starch content and dry weight measurement in WT and CAOx plants. (a) Starch content was measured from mature leaves of WT and CAOx plants grown under LL and HL at various times over a diurnal cycle as described in methods. Note the diurnal starch accumulation was maximum between 3 and 6 PM and CAOx-HL plants showed maximum starch accumulation. (b) Dry weight of WT and CAOx plants was measured after aerial parts of the plant were dried at 70°C for 5 days. HL-grown WT and CAOx plants showed significant increase in dry matter accumulation in comparison to WT-LL and CAOx-LL plants. Asterisks indicate significant differences determined by ANOVA followed by Tukey’s test (*P < 0.05; **P < 0.001). These experiments were done three times with similar results. Each data point is the average of four replicates in (a) and 15 replicate in (b) and the error bars represent SD (Biswal et al. 2012).
These results from the laboratory of the author (Biswal et al. 2012) demonstrate that controlled up-regulation of Chl b biosynthesis co-modulates the expression of chloroplast proteins that increase the antenna size and electron transport rates and enhances CO2 assimilation, starch contents and dry matter accumulation.
Chl b reductase catalyzes the conversion of Chl b to Chl a. It reduces the formyl group of Chl b to a hydroxymethyl group. It was observed that barley etioplast had Chlide b reductase activity and the enzyme needs NADPH and reduced ferredoxin for its activity (Scheumann et al. 1996, 1999). The gene encoding Chl b reductase was isolated from rice and it belongs to a family of short-chain dehydrogenase/reductases (Kusaba et al. 2007). It encodes a protein of 504 amino acids and contains a dinucleotide binding motif (TGXXXGXG) and a catalytic site (YXXXK) and uses NADPH as a cofactor. Interestingly, two genes for Chl b reductase were found in the genomes of A. thaliana and rice (Kusaba et al. 2007; Sato et al. 2009). It was also observed that disruption of the genes encoding Chl b reductase in A. thaliana resulted in non-degradation of Chl b and LHC II (Horie et al. 2009). In the presence of recombinant CAO enzyme, the Chlide a gets converted to Chlide b using NADPH, molecular oxygen and ferredoxin (Oster et al. 2000). In this in vitro assay, a small amount of 7-hydroxymehtyl Chlide a was also formed. When the 7-hydroxymethyl Chlide a was used as a substrate for the in vitro enzymatic assay, the recombinant enzyme also efficiently converted 7-hydroxymehtyl Chlide a to Chlide b (Oster et al. 2000). Then, Chl synthase converts Chlide b into Chl b. Chl b is further converted to hydroxymethyl Chl a by the enzyme Chl b reductase (Kusaba et al. 2007). This enzyme converts the formyl group of Chl b to a hydroxymethyl group using NADPH as a reductant.
Geranyl-geranyl reductase mediates the reduction of geranylgeranyl diphosphate to phytyl diphosphate. The cDNA encoding a pre-geranyl-geranyl reductase from A. thaliana has been isolated and characterized (Keller et al. 1998). The recombinant protein catalyzes the reduction of geranyl-geranyl-Chl a into phytyl-Chl a, as well as the reduction of free geranyl-geranyl diphosphate to phytyl diphosphate, suggesting this is a multifunctional enzyme.
2. Modulation of Phytol Synthesis and Its Impact on Plant Development, Photosynthesis, Tocopherol Contents and Oxidative Stress
a. Modulation by CHLP
Antisense expression of CHLP coding for geranyl-geranyl reductase affects the Chl and tocopherol contents in tobacco (Tanaka et al. 1999). The reduced tocopherol and Chl contents in CHLP antisense plants resulted in the reduction of electron transport chains and PS II activity. There are also more lipid peroxidation products in CHLP antisense plants. Havaux et al. (2003) found accumulation of xanthophylls cycle pigments in CHLP antisense plants which could be a compensatory mechanism for tochopherol deficiency. The CHLP transcript levels in peach were abundant in Chl-containing tissues and flower organs however barely detected in roots and mesocarp of the ripening fruits (Giannino et al. 2004). Its transcript level is up-regulated during etioplast to chloroplast and chloroplast to chromoplast development (Keller et al. 1998).
The responses of Chl biosynthetic enzymes to various environmental stresses are examined and summarized in Table 27.1. These stresses broadly downregulate most of the enzymes of Chl biosynthesis pathway. However, gene/protein expression of a certain enzyme i.e., GSA-AT is upregulated in most stresses, i.e., heat, water, salt etc. The expression of UroD is upregulated in high-temperature. As GSA-AT is a crucial enzyme involved in the last step of synthesis of ALA, plants most likely upregulate its expression to compensate for the reduced expression of earlier enzymes of the ALA biosynthesis.
III. Future Prospects
Plant tetrapyrroles play an important role in plant development, growth, productivity and modulation of their biosynthesis and protect plants from environmental stresses. Therefore, genetic manipulation of tetrapyrrole biosynthesis either via molecular marker assisted breeding programs or transgenic approaches will have a potential to protect crop plants from environmental stresses and increase yield.
Abbreviations
- ALA:
-
5-Aminolevulinic acid;
- ALAD:
-
5-Aminolevulinic acid dehydratase;
- CAO:
-
Chlorophyllide A oxygenase;
- Chl:
-
Chlorophyll;
- Chlide:
-
Chlorophyllide;
- Coprox:
-
Coproporphyrinogen oxidase;
- CPO:
-
Coproporphyrinogen oxidase;
- DV:
-
Divinyl;
- DV-Pchlide:
-
Divinyl protochlorophyllide;
- DVR:
-
Divinyl reductase;
- FLU:
-
Fluorescence;
- GGPP:
-
Geranyl geranyl pyrophosphate;
- GluRS:
-
Glutamyl-tRNA synthetase;
- GluTR:
-
Glutamyl-tRNA reductase;
- GSA:
-
Glutamate 1-semialdehyde;
- GSA-AT:
-
Glutamate 1-semialdehyde aminotransferase;
- GUN:
-
Genome uncoupled;
- LHC II:
-
Light-harvesting complex II;
- Lin2:
-
Lesion initiation 2;
- lip1:
-
Light-independent photomorphogenesis 1;
- MPE:
-
Mg-protoporphyrin IX monomethylester;
- MTF:
-
Mg-Protoporphyrin IX methyltransferase;
- MV:
-
Monovinyl;
- MV-Pchlide:
-
Monovinyl protochlorophyllide;
- PBG:
-
Porphobilinogen;
- PBGD:
-
Porphobilinogen deaminase;
- Pchlide:
-
Protochlorophyllide;
- PhPP:
-
Phytyl diphosphate;
- PLBs:
-
Prolamellar bodies;
- POR:
-
Protochlorophyllide oxido-reductase;
- PPIX:
-
Protoporphyrin IX;
- Protogen IX:
-
Protoporphyrinogen IX;
- Protox:
-
Protoporphyrinogen oxidase;
- ROS:
-
Reactive oxygen species;
- SAM:
-
S-adenosyl-methionine;
- SDR:
-
Short chain dehydrogenases/reductases;
- tRNAglu :
-
Glutamate conjugated tRNA;
- Urogen III:
-
Uroporphyrinogen III;
- UROS:
-
Uroporphyrinogen III synthase
References
Abdelkader AF, Aronsson H, Sundqvist C (2007a) High salt-stress in wheat leaves (Triticum aestivum) causes retardation of chlorophyll accumulation due to a limited rate of protochlorophyllide formation. Physiol Plant 130:157–166
Abdelkader AF, Aronsson H, Solymosi K, Böddi B, Sundqvist C (2007b) High salt stress induces swollen prothylakoids in dark-grown wheat and alters both prolamellar body transformation and reformation after irradiation. J Exp Bot 58:2553–2564
Adhikari ND, Orler R, Chory J, Froehlich JE, Larkin RM (2009) Porphyrins promote the association of genomes uncoupled 4 and a MG-chelatase subunit with chloroplast membranes. J Biol Chem 284:24783–24796
Adhikari ND, Froehlich JE, Strand DD, Buck SM, Kramer DM, Larkin RM (2011) GUN4-porphyrin complexes bind the ChlH/GUN5 subunit of Mg-chelatase and promote chlorophyll biosynthesis in Arabidopsis. Plant Cell 23:1449–1467
Alawady AE, Grimm B (2005) Tobacco Mg protoporphyrin IX methyltransferase is involved in inverse activation of Mg porphyrin and proto heme synthesis. Plant J 41:282–290
Alawady AE, Reski R, Yaronskaya E, Grimm B (2005) Cloning and expression of the tobacco CHLM sequence encoding Mg protoporphyrin IX methyltransferase and its interaction with Mg chelatase. Plant Mol Biol 57:679–691
Armstrong GA, Runge S, Frick G, Sperling U, Apel K (1995) Identification of NADPH: protochlorophyllide oxidoreductase A and B: a branched pathway for light-dependent chlorophyll biosynthesis in Arabidopsis thaliana. Plant Physiol 108:1505–1517
Arnould S, Camadro JM (1998) The domain structure of protoporphyrinogen oxidase, the molecular target of diphenyl ether-type herbicides. Proc Natl Acad Sci USA 95:10553–10558
Aronsson H, Sundqvist C, Timko MP, Dahlin C (2001) The importance of the C-terminal region and Cys residues for the membrane association of the NADPH: protochlorophyllide oxidoreductase in pea. FEBS Lett 502:11–15
Ayliffe MA, Agostino A, Clarke BC, Furbank R, von Caemmerer S, Pryor AJ (2009) Suppression of the barley uroporphyrinogen III synthase gene by a Ds activation tagging element generates developmental photosensitivity. Plant Cell 21:814–831
Bang WY, Jeong IS, Kim DW, Im CH, Ji C, Hwang SM, Kim SW, Son YS, Jeong J, Shiina T, Bahk JD (2008) Role of Arabidopsis CHL27 protein for photosynthesis, chloroplast development and gene expression profiling. Plant Cell Physiol 49:1350–1363
Barnes SA, Nishizawa NK, Quaggio RB, Whitelam GC, Chua N-H (1996) Far-red light blocks greening of Arabidopsis seedlings via a phytochrome A-mediated change in plastid development. Plant Cell 8:601–615
Bazzaz MB (1981) New chlorophyll chromophores isolate from a chlorophyll deficient mutant of maize. Photobiochem Photobiophys 2:199–207
Beale SI, Weinstein JD (1990) Tetrapyrrole metabolism in photosynthetic organisms. In: Dailey HA (ed) Biosynthesis of heme and chlorophyll. McGraw-Hill, New York, pp 287–291
Begley TP, Young H (1989) Protochlorophyllide reductase. 1. Determination of the regiochemistry and the stereochemistry of the reduction of protochlorophyllide to chlorophyllide. J Am Chem Soc 111:3095–3096
Bengtson C, Klockare B, Klockare R, Larsson S, Sundqvist C (1978) The after-effect of water stress on chlorophyll formation during greening and the levels of abscisic acid and proline in dark-grown wheat seedlings. Physiol Plant 43:205–212
Benz J, Rüdiger W (1981a) Incorporation of 1-14C-isopentenyldiphosphate, geraniol and farnesol into chlorophyll in plastid membrane fractions of Avena sativa L. Z Pflanzenphysiol 102:95–100
Benz J, Rüdiger W (1981b) Chlorophyll biosynthesis: various chlorophyllides as exogenous substrates for chlorophyll synthetase. Z Naturforsch 36c:51–57
Bhardwaj R, Singhal GS (1981) Effect of water stress on photochemical activity of chloroplasts during greening of etiolated barley seedlings. Plant Cell Physiol 22:155–162
Bhattacharjee S, Mukherjee AK (2003) Heavy metals alter photosynthetic pigment profiles as well as activities of chlorophyllase and 5-aminolevulinic acid dehydratase (ALAD) in Amaranthus lividus seedlings. J Environ Biol 24:395–399
Biswal AK, Pattanayak GK, Pandey SS, Leelavathi S, Reddy VS, Govindjee, Tripathy BC (2012) Light intensity-dependent modulation of chlorophyll b biosynthesis and photosynthesis by overexpression of chlorophyllide a oxygenase (CAO) in tobacco. Plant Physiol 159:433–449
Block MA, Tewari AK, Albrieux C, Marechal E, Joyard J (2002) The plant S-adenosyl-L-methionine: Mg-protoporphyrin IX methyl transferase is isolated in both envelope and thylakoid chloroplast membranes. Eur J Biochem 269:240–248
Block MA, Douce R, Joyard J, Rolland N (2007) Chloroplast envelope membranes: a dynamic interface between plastids and the cytosol. Photosynth Res 92:225–244
Böddi B, Lindsten A, Ryberg M, Sundqvist C (1989) On the aggregational states of protochlorophyllide and its protein complexes in wheat etioplasts. Physiol Plant 76:135–143
Böddi B, Lindsten A, Ryberg M, Sundqvist C (1990) Phototransformation of aggregated forms of protochlorophyllide in isolated etioplast inner membranes. Photochem Photobiol 52:83–87
Böddi B, Ryberg M, Sundqvist C (1991) The formation of a short-wavelength chlorophyllide form at partial phototransformation of protochlorophyllide in etioplast inner membranes. Photochem Photobiol 53:667–673
Böddi B, Ryberg M, Sundqvist C (1992) Identification of four universal protochlorophyllide forms in dark-grown leaves by analyses of the 77 K fluorescence emission spectra. J Photochem Photobiol B Biol 12:389–401
Böddi B, Ryberg M, Sundqvist C (1993) Analysis of the 77 K fluorescence emission and excitation spectra of isolated etioplast inner membranes. J Photochem Photobiol B Biol 21:125–133
Bohren KM, Grimshaw CE, Lai CJ, Harrison DH, Ringe D, Petsko GA, Gabbay KH (1994) Tyrosine-48 is the proton donor and histidine-110 directs substrate stereochemical selectivity in the reduction reaction of human aldose reductase: enzyme kinetics and crystal structure of the Y48H mutant enzyme. Biochemistry 33:2021–2032
Bollivar DW (2006) Recent advances in chlorophyll biosynthesis. Photosynth Res 90:173–194
Buhr F, El Bakkouri M, Valdez O, Pollmann S, Lebedev N, Reinbothe S, Reinbothe C (2008) Photoprotective role of NADPH: protochlorophyllide oxidoreductase A. Proc Natl Acad Sci USA 105:12629–12634
Camadro JM, Matringe M, Scalla R, Labbe P (1991) Kinetic studies on protoporphyrinogen oxidase inhibition by diphenyl ether herbicides. Biochem J 277:17–21
Castelfranco PA, Beale SI (1981) Chlorophyll biosynthesis. In: Stumpf PK, Conn EE (eds) The biochemistry of plants: a comprehensive treatise, vol 8. Academic, New York, pp 375–421
Castelfranco PA, Thayer SS, Wilkinson JQ, Bonner BA (1988) Labeling of porphobilinogen deaminase by radioactive 5-aminolevulinic acid in isolated developing pea chloroplasts. Arch Biochem Biophys 266:219–226
Chakraborty N, Tripathy BC (1992) Involvement of singlet oxygen in photodynamic damage of isolated chloroplasts of cucumber (Cucumis sativus L.) cotyledons. Plant Physiol 98:7–11
Che FS, Watnabe N, Iwano M, Inokuchi H, Takayama S, Yoshida S, Isogai A (2000) Molecular characterization and subcellular localization of protoporphyrinogen oxidase in spinach chloroplasts. Plant Physiol 124:59–70
Chen TC, Miller GW (1974) Purification and characterization of uroporphyrinogen decarboxylase from tobacco leaves. Plant Cell Physiol 15:993–1005
Chory J, Reinecke D, Sim S, Washburn T, Brenner M (1994) A role of cytokinins in de-etiolation in Arabidopsis det mutants have an altered response to cytokinins. Plant Physiol 104:339–347
Crockett N, Alefounder PR, Battersby AR, Abell C (1991) Uroporphyrinogen III synthase: studies on its mechanism of action molecular biology and biochemistry. Tetrahedron 47:6003–6014
Dahlin C, Aronsson H, Wilks HM, Lebedev N, Sundqvist C, Timko MP (1999) The role of protein surface charge in catalytic activity and chloroplast membrane association of the pea NADPH: protochlorophyllide oxidoreductase (POR) as revealed by alanine scanning mutagenesis. Plant Mol Biol 39:309–323
Dalal VK, Tripathy BC (2012) Modulation of chlorophyll biosynthesis by water stress in rice seedlings during chloroplast biogenesis. Plant Cell Environ. doi:10.1111/j.1365-3040.2012.02520.x
Davison PA, Schubert HL, Reid JD, Iorg CD, Heroux A, Hill CP, Hunter CN (2005) Structural and biochemical characterization of Gun4 suggests a mechanism for its role in chlorophyll biosynthesis. Biochemistry 44:7603–7612
Dayan FE, Ferreira D, Wang YH, Khan IA, McInroy JA, Pan Z (2008) A pathogenic fungi diphenyl ether phytotoxin targets plant enoyl (acyl carrier protein) reductase. Plant Physiol 147:1062–1071
Dutta S, Mohanty S, Tripathy BC (2009) Role of temperature stress on chloroplast biogenesis and protein import in pea. Plant Physiol 150:1050–1061
Eggink LL, LoBrutto R, Brune DC, Brusslan J, Yamasato A, Tanaka A, Hoober JK (2004) Synthesis of chlorophyll b: localization of chlorophyllide a oxygenase and discovery of a stable radical in the catalytic subunit. BMC Plant Biol 4:5–21
El Hamouri B, Brouers M, Sironval C (1981) Pathway from photoinactive P663–628 protocholorphyllide to the P696–682 chlorophyllide in cucumber etioplast suspensions. Plant Sci Lett 21:375–379
Eskins K, McCarthy SA, Dybas L, Duysen M (1986) Corn chloroplast development in weak fluence rate red light and in weak fluence rate red plus far-red light. Physiol Plant 67:242–246
Espineda CE, Linford AS, Devine D, Brusslan JA (1999) The AtCAO gene, encoding chlorophyll a oxygenase, is required for chlorophyll b synthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 96:10507–10511
Eullaffroy P, Popovic R (1997) Effect of heat treatment on protochlorophyllide phototransformation initiated by different light intensities. J Plant Physiol 151:293–298
Eullaffroy P, Salvetat R, Franck F, Popovic R (1995) Temperature dependence of chlorophyspectral shifts and photoactive protochlorophyllide regeneration after flash in etiolated barley leaves. J Photochem Photobiol B Biol 62:751–756
Fodje MN, Hansson A, Hansson M, Olsen JG, Gough S, Willows RD, Al-Karadaghi S (2001) Interplay between an AAA module and an integrin I domain may regulate the function of magnesium chelatase. J Mol Biol 311:111–122
Forreiter C, van Cleve B, Schmidt A, Apel K (1991) Evidence for a general light-dependent negative control of NADPH–protochlorophyllide oxidoreductase in angiosperms. Planta 183:126–132
Franck F (1993) On the formation of photosystem II chlorophyll-proteins after a short light flash in etiolated barley leaves, as monitored by in vivo fluorescence spectroscopy. J Photochem Photobiol B Biol 18:35–40
Franck F, Bereza B, Böddi B (1999) Protochlorophyllide-NADP + and protochlorophyllide-NADPH complexes and their regeneration after flash illumination in leaves and etioplast membranes of dark-grown wheat. Photosynth Res 59:53–61
Frick G, Su Q, Apel K, Armstrong GA (2003) An Arabidopsis porB porC double mutant lacking light-dependent NADPH: protochlorophyllide oxidoreductases B and C is highly chlorophyll-deficient and developmentally arrested. Plant J 35:141–153
Frustaci JM, Sangwan I, O’Brian MR (1995) gsa1 is a universal tetrapyrrole synthesis gene in soybean and is regulated by a GAGA element. J Biol Chem 270:7387–7393
Fusada N, Masuda T, Kuroda H, Shiraishi T, Shimada H, Ohta H, Takamiya K (2000) NADPH–protochlorophyllide oxidoreductase in cucumber is encoded by a single gene and its expression is transcriptionally enhanced by illumination. Photosynth Res 64:147–154
Fusada N, Masuda T, Kuroda H, Shimada H, Ohta H, Takamiya K (2005) Identification of a novel cis-element exhibiting cytokinin-dependent protein binding in vitro in the 5′-region of NADPH-protochlorophyllide oxidoreductase gene in cucumber. Plant Mol Biol 59:631–645
Gaubier P, Wu HJ, Laudie M, Delseny M, Grellet F (1995) A chlorophyll synthetase gene from Arabidopsis thaliana. Mol Gen Genet 249:673–676
Giannino D, Condello E, Bruno L, Testone G, Tartarini A, Cozza R, Innocenti AM, Bitonti MB, Mariotti D (2004) The gene geranylgeranyl reductase of peach (Prunus persica [L.] Batsch) is regulated during leaf development and responds differentially to distinct stress factors. J Exp Bot 55:2063–2073
Gibson KD, Neuberger A, Tait GH (1963) Studies on the biosynthesis of porphyrin and bacteriochlorophyll by Rhodopseudomonas spheriodes. 4. S-adenosylmethionine-magnesium protoporphyrin methyltransferase. Biochem J 88:325–334
Gibson LC, Marrison JL, Leech RM, Jensen PE, Bassham DC, Gibson M, Hunter CN (1996) A putative Mg chelatase subunit from Arabidopsis thaliana cv C24. Sequence and transcript analysis of the gene, import of the protein into chloroplasts, and in situ localization of the transcript and protein. Plant Physiol 111:61–71
Goslings D, Meskauskiene R, Kim C, Lee KP, Nater M, Apel K (2004) Concurrent interactions of heme and FLU with Glu tRNA reductase (HEMA1), the target of metabolic feedback inhibition of tetrapyrrole biosynthesis, in dark- and light-grown Arabidopsis plants. Plant J 40:957–967
Gough SP, Kannangara CG, von Wettestein D (1992) Glutamate 1-semialdehyde aminotransferase as a target for herbicides. In: Boger P, Sandmann G (eds) Target assays for modern herbicides and related phytotoxic compounds. Lewis, Chelesa, pp 21–27
Gray JC, Sullivan JA, Wang JH, Jerome CA, MacLean D (2003) Coordination of plastid and nuclear gene expression. Philos Trans R Soc Lond B Biol Sci 358:135–144
Guo R, Luo M, Weinstein JD (1998) Magnesium-chealatse from developing pea leaves. Plant Physiol 116:605–615
Hansson A, Jensen PE (2009) Chlorophyll limitation in plants remodels and balances the photosynthetic apparatus by changing the accumulation of photosystems I and II through two different approaches. Physiol Plant 135:214–228
Hansson A, Kannangara CG, von Wettstein D, Hansson M (1999) Molecular basis for semidominance of missense mutations in the XANTHA-H (42-kDa) subunit of magnesium chelatase. Proc Natl Acad Sci USA 96:1744–1749
Harper AL, von Gesjen SE, Linford AS, Peterson MP, Faircloth RS, Thissen MM, Brusslan JA (2004) Chlorophyllide a oxygenase mRNA and protein levels correlate with the chlorophyll a/b ratio in Arabidopsis thaliana. Photosynth Res 79:149–159
Havaux M, Lutz C, Grimm B (2003) Chloroplast membrane photostability in chlP transgenic tobacco plants deficient in tocopherols. Plant Physiol 132:300–310
He ZH, Li J, Sundqvist C, Timko MP (1994) Leaf development age controls expression of genes encoding enzymes of chlorophyll and heme biosynthesis in pea (Pisum sativum L.). Plant Physiol 106:537–546
Helfrich M, Schoch S, Lempert U, Cmiel E, Rudiger W (1994) Chlorophyll synthetase cannot synthesize chlorophyll a. Eur J Biochem 219:267–275
Heyes DJ, Hunter CN (2004) Identification and characterization of the product release steps within the catalytic cycle of protochlorophyllide oxidoreductase. Biochemistry 43:8265–8271
Heyes DJ, Ruban AV, Wilks HM, Hunter CN (2002) Enzymology below 200 K: the kinetics and thermodynamics of the photochemistry catalyzed by protochlorophyllide oxidoreductase. Proc Natl Acad Sci USA 99:11145–11150
Heyes DJ, Ruban AV, Hunter CN (2003) Protochlorophyllide oxidoreductase: spectroscopic characterization of the ‘dark’ reactions. Biochemistry 42:523–528
Heyes DJ, Kruk J, Hunter CN (2006) Spectroscopic and kinetic characterization of the light-dependent enzyme protochlorophyllide oxidoreductase (POR) using monovinyl and divinyl substrates. Biochem J 394:243–248
Heyes DJ, Menon BR, Sakuma M, Scrutton NS (2008) Conformational events during ternary enzyme-substrate complex formation are rate limiting in the catalytic cycle of the light-driven enzyme protochlorophyllide oxidoreductase. Biochemistry 47:10991–10998
Higuchi M, Bogorad L (1975) The purification and properties of uroporphyrinogen I synthase and uroporphyrinogen III cosynthase. Interactions between the enzymes. Annu NY Acad Sci 244:401–418
Hirashima M, Satoh S, Tanaka R, Tanaka A (2006) Pigment shuffling in antenna systems achieved by expressing prokaryotic chlorophyllide a oxygenase in Arabidopsis. J Biol Chem 281:15385–15393
Hiriart JB, Lehto K, Tyystjarvi E, Junttila T, Aro EM (2002) Suppression of a key gene involved in chlorophyll biosynthesis by means of virus-inducing gene silencing. Plant Mol Biol 50:213–224
Hodgins RR, Oquist G (2006) Porphyrin metabolism in chill-stressed seedlings of Scots pine (Pinus sylvestris). Physiol Plant 77:620–624
Hodgins RR, Van Huystee RB (1986) Delta-aminolevulinic acid metabolism in chill stressed maize (Zea mays L.). J Plant Physiol 126:257–268
Holtorf H, Apel K (1996) Transcripts of the two NADPH–protochlorophyllide oxidoreductase genes PorA and PorB are differentially degraded in etiolated barley seedlings. Plant Mol Biol 31:387–392
Holtorf H, Reinbothe S, Reinbothe C, Bereza B, Apel K (1995) Two routes of chlorophyllide synthesis that are differentially regulated by light in barley (Hordeum vulgare L.). Proc Natl Acad Sci USA 92:3254–3258
Hoober JK, Kahn A, Ash DE, Gough SP, Kannagara CG (1988) Biosynthesis of δ-aminolevulinate in greening barley leaves. IX. Structure of the substrate, mode of gabaculine inhibition, and the catalytic mechanism of Glutamate-1-semialdehyde aminotransferase. Carlsberg Res Commun 53:11–25
Horie Y, Ito H, Kusaba M, Tanaka R, Tanaka A (2009) Participation of chlorophyll b reductase in the initial step of the degradation of light-harvesting chlorophyll a/b-protein complexes in Arabidopsis. J Biol Chem 284:17449–17456
Hsu WP, Miller GW (1970) Coproporphyrinogenase in tobacco (Nicotiana tabacum L.). Biochem J 117:215–220
Huang YS, Li HM (2009) Arabidopsis CHLI2 can substitute for CHLI1. Plant Physiol 150:636–645
Huang DD, Wang WY, Gough SP, Kannangara CG (1984) Delta-Aminolevulinic acid-synthesizing enzymes need an RNA moiety for activity. Science 225:1482–1484
Hukmani P, Tripathy BC (1994) Chlorophyll biosynthetic reactions during senescence of excised barley (Hordeum vulgare L. cv IB 65) Leaves. Plant Physiol 105:1295–1300
Ikegami A, Yoshimura N, Motohashi K, Takahashi S, Romano PG, Hisabori T, Takamiya K, Masuda T (2007) The CHLI1 subunit of Arabidopsis thaliana magnesium chelatase is a target protein of the chloroplast thioredoxin. J Biol Chem 282:19282–19291
Ilag LL, Kumar AM, Soll D (1994) Light reduction of chlorophyll biosynthesis at the level of 5-aminolevulinate formation in Arabidopsis. Plant Cell 6:265–275
Ishikawa A, Okamoto H, Iwasaki Y, Asahi T (2001) A deficiency of coproporphyrinogen III oxidase causes lesion formation in Arabidopsis. Plant J 27:89–99
Jacobs NJ, Jacobs JM (1979) Microbial oxidation of protoporphyrinogen: an intermediate in heme and chlorophyll biosynthesis. Arch Biochem Biophys 197:396–403
Jacobs JM, Jacobs NJ (1987) Oxidation of protoporphyrinogen to protoporphyrin, a step in chlorophyll and haem biosynthesis. Purification and partial characterization of the enzyme from barley organelles. Biochem J 244:219–224
Jain M, Gadre RP (2004) Inhibition of 5-amino levulinic acid dehydratase activity by arsenic in excised etiolated maize leaf segments during greening. J Plant Physiol 161:251–255
Jensen PE, Willows RD, Petersen BL, Vothknecht UC, Stummann BM, Kannangara CG, von Wettstein D, Henningsen KW (1996) Structural genes for Mg-chelatase subunits in barley: Xantha-f, -g, and -h. Mol Gen Genet 250:383–394
Jensen PE, Gibson LCD, Hunter CN (1999) ATPase activity associated with the magnesium-protoporphyrin IX chelatase enzyme of Synechocystis PCC6803: evidence for ATP hydrolysis during Mg2+ insertion, and MgATP dependent interaction of ChlI and ChlD subunits. Biochem J 339:127–134
Jung KH, Hur J, Ryu CH, Choi Y, Chung YY, Miyao A, Hirochika H, An G (2003) Characterization of a rice chlorophyll-deficient mutant using the T-DNA gene-trap system. Plant Cell Physiol 44:463–472
Kannangara CG, Gough SP, Oliver RP, Rasmussen SK (1984) Biosynthesis of d-aminolevulinate in greening barley leaves VI. Activation of glutamate by ligation to RNA. Carlsberg Res Commun 49:417–437
Kannangara CG, Gough SP, Bruyant P, Hoober JK, Kahn A, Wettestein DV (1988) tRNAglu as a cofactor on δ-aminolevulinate biosynthesis: steps that regulate chlorophyll synthesis. Trends Biol Sci 13:139–143
Kannangara CG, Andersen RV, Pontoppidan B, Willows R, von Wettstein D (1994) Enzymic and mechanistic studies on the conversion of glutamate to 5-aminolaevulinate. Ciba Found Symp 180:3–20
Keller Y, Bouvier F, D’Harlingue A, Camara B (1998) Metabolic compartmentation of plastid prenyllipid biosynthesis. Evidence for the involvement of a multifunctional geranylgeranyl reductase. Eur J Biochem 251:413–417
Kervinen J, Dunbrack R Jr, Litwin S, Martins J, Scarrow RC, Volin M, Yeung AT, Yoon E, Jaffe EK (2000) Porphobilinogen synthase from pea: expression from an artificial gene, kinetic characterisation and novel implication for subunit interactions. Biochemistry 39:9018–9029
Koch M, Breithaupt C, Kiefersauer R, Freigang J, Huber R, Messerschmidt A (2004) Crystal structure of protoporphyrinogen IX oxidase: a key enzyme in haem and chlorophyll biosynthesis. EMBO J 23:1720–1728
Kolossov VL, Rebeiz CA (2001) Chloroplast biogenesis 84: solubilization and partial purification of membrane-bound [4-vinyl] chlorophyllide a reductase from etiolated barley leaves. Anal Biochem 295:214–219
Kruse E, Mock HP, Grimm B (1995a) Reduction of coproporphyrinogen oxidase level by antisense RNA synthesis leads to deregulated gene expression of plastid proteins and affects the oxidative defense system. EMBO J 14:3712–3720
Kruse E, Mock HP, Grimm B (1995b) Coproporphyrinogen III oxidase from barley and tobacco-sequence analysis and initial expression studies. Planta 196:796–803
Kruse E, Mock HP, Grimm B (1997) Isolation and characterization of tobacco (Nicotiana tabacum) cDNA clones encoding proteins involved in magnesium chelation into protoporphyrin IX. Plant Mol Biol 35:1053–1056
Kuroda H, Masuda T, Ohta H, Shioi Y, Takamiya K (1995) Light-enhanced gene expression of NADPH–protochlorophyllide oxidoreductase in cucumber. Biochem Biophys Res Commun 210:310–316
Kuroda H, Masuda T, Fusada N, Ohta H, Takamiya K (2001) Cytokinin-induced transcriptional activation of NADPH–protochlorophyllide oxidoreductase gene in cucumber. J Plant Res 114:1–7
Kusaba M, Ito H, Morita R, Iida S, Sato Y, Fujimoto M, Kawasaki S, Tanaka R, Hirochika H, Nishimura M, Tanaka A (2007) Rice NON-YELLOW COLORING1 is involved in light-harvesting complex II and grana degradation during leaf senescence. Plant Cell 19:1362–1375
Kusnetsov V, Herrmann RG, Kulaeva ON, Oelmüller R (1998) Cytokinin stimulates and abscisic acid inhibits greening of etiolated Lupinus luteus cotyledons by affecting the expression of the light-sensitive protochlorophyllide oxidoreductase. Mol Gen Genet 259:21–28
Kusumi J, Sato A, Tachidi H (2006) Relaxation of functional constraint on light-independent protochlorophyllide reductase in Thuja. Mol Biol Evol 23:941–948
Larkin RM, Alonso JM, Ecker JR, Chory J (2003) GUN4, a regulator of chlorophyll synthesis and intracellular signaling. Science 299:902–906
Le Lay P, Eullaffroy P, Juneau P, Popovic D (2000) Evidence of chlorophyll synthesis pathway alteration in desiccated barley leaves. Plant Cell Physiol 41:565–570
Le Lay P, Boddi B, Kovacevic D, Juneau P, Dewez D, Popovic P (2001) Spectroscopic analysis of desiccation-induced alterations of the chlorophyllide transformation pathway in etiolated barley leaves. Plant Physiol 127:202–211
Lebedev N, Timko MP (1999) Protochlorophyllide oxidoreductase B-catalysed protochlorophyllide photoreduction in vitro: insight into the mechanism of chlorophyll formation in light-adapted plants. Proc Natl Acad Sci USA 96:9954–9959
Lebedev N, Carcinoma O, McIvor W, Timko MP (2001) Tyr275 and Lys279 stabilize NADPH within the catalytic site of NADPH: protochlorophyllide oxidoreductase and are involved in the formation of the enzyme photoactive state. Biochemistry 40:12562–12574
Lee KP, Kim C, Lee DW, Apel K (2003) TIGRINA d, required for regulating the biosynthesis of tetrapyrroles in barley, is an ortholog of the FLU gene of Arabidopsis thaliana. FEBS Lett 553:119–124
Lee S, Kim JH, Yoo ES, Lee CH, Hirochika H, An G (2005) Differential regulation of chlorophyll a oxygenase genes in rice. Plant Mol Biol 57:805–818
Leon P, Arroyo A, Mackenzie S (1998) Nuclear control of plastid and mitochondrial developments in higher plants. Annu Rev Plant Physiol Plant Mol Biol 49:453–480
Lermontova I, Kruse E, Mock HP, Grimm B (1997) Cloning and characterisation of a plastidal and a mitochondrial isoform of tobacco protoporphyrinogen IX oxidase. Proc Natl Acad Sci USA 94:8895–8900
Li J, Spano AJ, Timko MP (1991) Isolation and characterisation of nuclear genes encoding the ALA dehydratase of pea (Pisum sativum L.). Plant Physiol 96:125–127
Liedgens W, Grutzmann R, Schneider HAW (1980) Highly efficient purification of the labile plant enzyme 5-aminolevulinate dehydratase (EC 4.2.1.24) by means of monoclonal antibodies. Z Naturforsch 35c:958–962
Lim SH, Witty M, Wallace-Cook AD, Ilag LI, Smith AG (1994) Porphobilinogen deaminase is encoded by a single gene in Arabidopsis thaliana and is targeted to the chloroplasts. Plant Mol Biol 26:863–872
Lindstein A, Welch CJ, Schoch S, Ryberg M, Rüdiger W, Sundqvist C (1990) Chlorophyll synthatase is latent in well preserved prolamellar bodies of etiolated wheat. Physiol Plant 80:277–285
Lindsten A, Ryberg M, Sundqvist C (1988) The polypeptide composition of highly purified prolamellar bodies and prothylakoids from wheat (Tritium aestivum) as revealed by silver staining. Physiol Plant 72:167–176
Liu N, Yang YT, Liu HH, Yang GD, Zhang NH, Zheng CC (2004) NTZIP antisense plants show reduced chlorophyll levels. Plant Physiol Biochem 42:321–327
Luo J, Lim CK (1993) Order of urogen III decarboxylation on incubation of PBG and urogen III with erythrocyte UDC. Biochem J 289:529–532
Madsen O, Sandal L, Sandal NN, Marcker KA (1993) A soybean coproporphyrinogen oxidase gene is highly expressed in root nodules. Plant Mol Biol 23:35–43
Manohara MS, Tripathy BC (2000) Regulation of protoporphyrin IX biosynthesis by intraplastidic compartmentalization and adenosine triphosphate. Planta 212:52–59
Martin BM, Grimm B, Mock HP, Huber R, Messerschmidt A (2001) Crystal structure and substrate binding modeling of the uroporphyrinogen-III decarboxylase from Nicotiana tabacum. Implications for the catalytic mechanism. J Biol Chem 276:44108–44116
Martin W, Rujan T, Richly E, Hansen A, Cornelsen S, Lins T, Leister D, Stoebe B, Hasegawa M, Penny D (2002) Evolutionary analysis of Arabidopsis, cyanobacterial, and chloroplast genomes reveals plastid phylogeny and thousands of cyanobacterial genes in the nucleus. Proc Natl Acad Sci USA 99:12246–12251
Masuda T (2008) Recent overview of the Mg branch of the tetrapyrrole biosynthesis leading to chlorophylls. Photosynth Res 96:121–143
Masuda T, Takamiya K (2004) Novel insights into enzymology, regulation and physiological functions of light-dependant protochlorophyllide oxidoreductase in angiosperms. Photosynth Res 81:1–29
Masuda T, Fusada N, Shiraishi T, Kuroda H, Awai K, Shimada H, Ohta H, Takamiya K (2002) Identification of two differentially regulated isoforms of protochlorophyllide oxidoreductase (POR) from tobacco revealed a wide variety of light and development-dependent regulations of POR gene expression among angiosperms. Photosynth Res 74:165–172
Masuda T, Fusada N, Oosawa N, Takamatsu K, Yamamoto YY, Ohto M, Nakamura K, Goto K, Shibata D, Shirano Y, Hayashi H, Kato T, Tabata S, Shimada H, Ohta H, Takamiya K (2003) Functional analysis of isoforms of NADPH: protochlorophyllide oxidoreductase (POR), PORB and PORC, in Arabidopsis thaliana. Plant Cell Physiol 44:963–974
Masuda S, Ikeda R, Masuda T, Hashimoto H, Tsuchiya T, Kojima H, Nomata J, Fujita Y, Mimuro M, Ohta H, Takamiya K (2009) Prolamellar bodies formed by cyanobacterial protochlorophyllide oxidoreductase in Arabidopsis. Plant J 58:952–960
Matringe M, Camadro JM, Block MA, Joyard J, Scalla R, Labbe P, Douce R (1992a) Localisation within the chloroplasts of protoporphyrinogen oxidase, the target enzyme for diphenylether like herbicides. J Biol Chem 267:4646–4651
Matringe M, Mornet R, Scalla R (1992b) Characterization of [3H] acifluorfen binding to purified pea etioplasts and evidence that protogen oxidase specifically binds acifluorfen. Eur J Biochem 209:861–868
Matsumoto F, Obayashi T, Sasaki-Sekimoto Y, Ohta H, Takamiya K, Masuda T (2004) Gene expression profiling of the tetrapyrrole metabolic pathway in Arabidopsis with a mini-array system. Plant Physiol 135:2379–2391
Meskauskiene R, Apel K (2002) Interaction of FLU, a negative regulator of tetrapyrrole biosynthesis, with the glutamyl–tRNA reductase requires the tetratricopeptide repeat domain of FLU. FEBS Lett 532:27–30
Meskauskiene R, Nater M, Goslings D, Kessler F, Op den Camp R, Apel K (2001) FLU: a negative regulator of chlorophyll biosynthesis in Arabidopsis thaliana. Proc Natl Acad Sci USA 98:12826–12831
Mochizuki N, Brusslan JA, Larkin R, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelatase H subunit in plastidto-nucleus signal transduction. Proc Natl Acad Sci USA 98:2053–2058
Mock HP, Trainotti L, Kruse E, Grimm B (1995) Isolation, sequencing and expression of cDNA sequences encoding uroporphyrinogen decarboxylase from tobacco and barley. Plant Mol Biol 28:245–256
Mohanty S, Tripathy BC (2011) Early and late plastid development in response to chill stress and heat stress in wheat seedlings. Protoplasma 248:725–736
Mohanty S, Grimm B, Tripathy BC (2006) Light and dark modulation of chlorophyll biosynthetic genes in response to temperature. Planta 224:692–699
Mosinger E, Batschauer A, Schafer E, Apel K (1985) Phytochrome control of in vitro transcription of specific genes in isolated nuclei from barley (Hordeum vulgare). Eur J Biochem 147:137–142
Nagai S, Koide M, Takahashi S, Kikuta A, Aono M, Sasaki-Sekimoto Y, Ohta H, Takamiya K-I, Masuda T (2007) Induction of isoforms of tetrapyrrole biosynthetic enzymes, AtHEMA2 and AtFC1, under stress conditions and their physiological functions in Arabidopsis. Plant Physiol 144:1039–1051
Nagata N, Satoh S, Tanaka R, Tanaka A (2004) Domain structures of chlorophyllide a oxygenase of green plants and Prochlorothrix hollandica in relation to catalytic functions. Planta 218:1019–1025
Nagata N, Tanaka R, Satoh S, Tanaka A (2005) Identification of a vinyl reductase gene for chlorophyll synthesis in Arabidopsis thaliana and implications for the evolution of prochlorococcus species. Plant Cell 17:233–240
Nagata N, Tanaka R, Tanaka A (2007) The major route for chlorophyll synthesis includes [3,8-divinyl]-chlorophyllide a reduction in Arabidopsis thaliana. Plant Cell Physiol 48:1803–1808
Nakagawara E, Sakuraba Y, Yamasato A, Tanaka R, Tanaka A (2007) Clp protease controls chlorophyll b synthesis by regulating the level of chlorophyllide a oxygenase. Plant J 49:800–809
Nakanishi H, Nozue H, Suzuki K, Kaneko Y, Taguchi G, Hayashida N (2005) Characterization of the Arabidopsis thaliana mutant pcb2 which accumulates divinyl chlorophylls. Plant Cell Physiol 46:467–473
Nakayama M, Masuda T, Bando T, Yamagata H, Ohta H, Takamiya K (1998) Cloning and expression of the soybean chlH gene encoding a subunit of Mg-chelatase and localization of the Mg++ concentration-dependent chlH protein within the chloroplast. Plant Cell Physiol 39:275–284
Narita S, Tanaka R, Ito T, Okada K, Taketani S, Inokuchi H (1996) Molecular cloning and characterisation of a cDNA that encodes protoporphyrinogen oxidase of Arabidopsis thaliana. Gene 182:169–175
Nott A, Jung HS, Koussevitzky S, Chory J (2006) Plastid-to-nucleus retrograde signaling. Annu Rev Plant Biol 57:739–759
Oliver RP, Griffiths WT (1982) Pigment-protein complexes of illuminated etiolated leaves. Plant Physiol 70:1019–1025
Oosawa N, Masuda T, Awai K, Fusada N, Shimada H, Ohta H, Takamiya K (2000) Identification and light-induced expression of a novel gene of NADPH-photochlorophyllide oxidoreductase isoform in Arabidopsis thaliana. FEBS Lett 474:133–136
Oster U, Tanaka R, Tanaka A, Rudiger W (2000) Cloning and functional expression of the gene encoding the key enzyme for chlorophyll b biosynthesis (CAO) from Arabidopsis thaliana. Plant J 21:305–310
Papenbrock J, Grimm B (2001) Regulatory network of tetrapyrrole biosynthesis-studies for interacellular signaling involved in metabolic and developmental control of plastids. Planta 213:667–681
Papenbrock J, Gräfe S, Kruse E, Hanel F, Grimm B (1997) Mg-chelatase of tobacco: identification of a Chl D cDNA sequence encoding a third subunit, analysis of the interaction of the three subunits with the yeast two-hybrid system, and reconstitution of the enzyme activity by co-expression of recombinant CHL D, CHL H and CHL I. Plant J 12:981–990
Papenbrock J, Mock HP, Kruse E, Grimm B (1999) Expression studies on tetrapyrrole biosynthesis: inverse maxima of magnesium chelatase and ferrochelatase activity during cyclic photoperiods. Planta 208:264–273
Papenbrock J, Pfundel E, Mock HP, Grimm B (2000a) Decreased and increased expression of the subunit CHL I diminishes Mg chelatase activity and reduced chlorophyll synthesis in transgenic tobacco plants. Plant J 22:155–164
Papenbrock J, Mock HP, Tanaka R, Kruse E, Grimm B (2000b) Role of magnesium chelatase activity in the early steps of the tetrapyrrole biosynthetic pathway. Plant Physiol 122:1161–1169
Parham R, Rebeiz CA (1995) Chloroplast biogenesis 72: a [4-vinyl] chlorophyllide a reductase assay using divinyl chlorophyllide a as an exogenous substrate. Anal Biochem 231:164–169
Pattanayak GK, Tripathy BC (2002) Catalytic function of a novel protein protochlorophyllide oxidoreductase C of Arabidopsis thaliana. Biochem Biophys Res Commun 291:921–924
Pattanayak GK, Tripathy BC (2011) Overexpression of protochlorophyllide oxidoreductase C regulates oxidative stress in Arabidopsis. PLoS One 6:e26532
Pattanayak GK, Biswal AK, Reddy VS, Tripathy BC (2005) Light-dependent regulation of chlorophyll b biosynthesis in chlorophyllide a oxygenase overexpressing tobacco plants. Biochem Biophys Res Commun 326:466–471
Peter E, Rothbart M, Oelze ML, Shalygo N, Dietz KJ, Grimm B (2010) Mg protoporphyrin monomethylester cyclase deficiency and effects on the tetrapyrrole metabolism in different light conditions. Plant Cell Physiol 51:1229–1241
Phung T, Jung HI, Park J-H, Kim J-G, Back K, Jung S (2012) Porphyrin biosynthesis control under water stress: sustained porphyrin status correlates with drought tolerance in transgenic rice. Plant Physiol 57:1746–1764
Plöscher M, Granvogl B, Reisinger V, Eichacker LA (2009) Identification of the N-termini of NADPH: protochlorophyllide oxidoreductase A and B from barley etioplasts (Hordeum vulgare L.). FEBS J 276:1074–1081
Pontier D, Albrieux C, Joyard J, Lagrange T, Block MA (2007) Knock-out of the magnesium protoporphyrin IX methyltransferase gene in Arabidopsis. Effects on chloroplast development and on chloroplast-to-nucleus signaling. J Biol Chem 282:2297–2304
Pontoppidan B, Kannangara CG (1994) Purification and partial characterization of barley glutamyl-tRNAglu reductase, the enzyme that directs glutamate to chlorophyll biosynthesis. Eur J Biochem 225:529–537
Pontoppodian B, Kannangara CG (1994) Purification and partial characterization of barley glutamyl-tRNAglu reductase, the enzyme that directs glutamate to chlorophyll biosynthesis. Eur J Biochem 225:529–537
Popperl G, Oster U, Blos I, Rudiger W (1997) Magnesium chelatase of Hordeum vulgare L. is not activated by light but inhibited by pheorphorbide. Z Naturforsch 52c:144–152
Porra RJW, Schäfer E, Cmiel IK, Scheer H (1993) Derivation of the formyl group oxygen of chlorophyll b from molecular oxygen in greening leaves of a higher plant (Zea mays). FEBS Lett 371:21–24
Poulson R, Polglasse WJ (1974) Aerobic and anaerobic coproporphyrinogen oxidase activities in extract from Saccharomyces cerevisiae. J Biol Chem 249:6367–6371
Rebeiz CA, Parham R, Fasoula DA, Ioannides IM (1994) Chlorophyll biosynthetic heterogeneity. In: The biosynthesis of the tetrapyrrole pigments. Ciba symposium 180. Wiley, New York, pp 177–193
Reinbothe C, Lebedev N, Reinbothe S (1999) A protochlorophyllide light-harvesting complex involved in de-etiolation of higher plants. Nature 397:80–84
Reinbothe C, Buhr F, Bartsch S, Desvignes C, Quigley F, Pesey H, Reinbothe S (2006) In vitro-mutagenesis of NADPH: protochlorophyllide oxidoreductase B: two distinctive protochlorophyllide binding sites participate in enzyme catalysis and assembly. Mol Genet Genomics 275:540–552
Rissler HM, Collakova E, DellaPenna D, Whelan J, Pogson BJ (2002) Chlorophyll biosynthesis. Expression of a second chl I gene of magnesium chelatase in Arabidopsis supports only limited chlorophyll synthesis. Plant Physiol 128:770–779
Rüdiger W, Benz J, Guthoff C (1980) Detection and partial characterization of activity of chlorophyll synthetase in etioplast membranes. Eur J Biochem 109:193–200
Runge S, Sperling U, Frick G, Apel K, Armstrong GA (1996) Distinct roles for light-dependent NADPH: protochlorophyllide oxidoreductases (POR) A and B during greening in higher plants. Plant J 9:513–523
Rzeznicka K, Walker CJ, Westergren T, Kannangara CG, von Wettstein D, Merchant S, Gough SP, Hansson M (2005) Xantha-l encodes a membrane subunit of the aerobic Mg-protoporphyrin IX monomethyl ester cyclase involved in chlorophyll biosynthesis. Proc Natl Acad Sci USA 102:5886–5891
Sakuraba Y, Yamasato A, Tanaka R, Tanaka A (2007) Functional analysis of N-terminal domains of Arabidopsis chlorophyllide a oxygenase. Plant Physiol Biochem 45:740–749
Sakuraba Y, Yokono M, Akimoto S, Tanaka R, Tanaka A (2010) Deregulated chlorophyll b synthesis reduces the energy transfer rate between photosynthetic pigments and induces photodamage in Arabidopsis thaliana. Plant Cell Physiol 51:1055–1065
Sangwan I, O’Brian MR (1993) Expression of the soybean (Glycine max) glutamate 1-semialdehtde aminotransferase gene in symbiotic root nodules. Plant Physiol 102:829–834
Santel HJ, Apel K (1981) The protochlorophyllide holochrome of barley (Hordeum vulgare L.). The effect of light on the NADPH: protochlorophyllide oxidoreductase. Eur J Biochem 120:95–103
Santos CV (2004) Regulation of chlorophyll biosynthesis and degradation by salt stress in sunflower leaves. Scienia Horticulturae 103:93–99
Santos C, Caldeira G (1999) Comparative responses of Helianthus annuus plants and calli exposed to NaCl: I. Growth rate and osmotic regulation in intact plants and calli. J Plant Physiol 155:769–777
Santos C, Azevedo H, Caldeira G (2001) In situ and in vitro senescence induced by KCl stress: nutritional imbalance, lipid peroxidation and antioxidant metabolism. J Exp Bot 52:351–360
Sato Y, Morita R, Katsuma S, Nishimura M, Tanaka A, Kusaba M (2009) Two short-chain dehydrogenase/reductases, NON-YELLOW COLORING 1 and NYC1-LIKE, are required for chlorophyll b and light-harvesting complex II degradation during senescence in rice. Plant J 57:120–131
Satoh S, Ikeuchi M, Mimuro M, Tanaka A (2001) Chlorophyll b expressed in cyanobacteria functions as a light-harvesting antenna in photosystem I through flexibility of the proteins. J Biol Chem 276:4293–4297
Scheumann V, Ito H, Tanaka A, Schoch S, Rüdiger W (1996) Substrate specificity of chlorophyll(ide) b reductase in etioplasts of barley (Hordeum vulgare L.). Eur J Biochem 242:163–170
Scheumann V, Klement H, Helfrich M, Oster U, Schoch S, Rüdiger W (1999) Protochlorophyllide b does not occur in barley etioplasts. FEBS Lett 445:445–448
Schmid HC, Oster U, Kögel J, Lenz S, Rüdiger W (2001) Cloning and characterisation of chlorophyll synthase from Avena sativa. J Biol Chem 382:903–911
Schmid HC, Rassadina V, Oster U, Schoch S, Rüdiger W (2002) Pre-loading of chlorophyll synthase with tetraprenyl diphosphate is an obligatory step in chlorophyll biosynthesis. J Biol Chem 383:1769–1778
Schoefs B, Franck F (1993) Photoreduction of protochlorophyllide to chlorophyllide in 2-d-old dark-grown bean (Phaseolus vulgaris cv. Commodore) leaves. Comparison with 10-d-old dark-grown (etiolated) leaves. J Expt Bot 44:1053–1057
Schoefs B, Franck F (2003) Protochlorophyllide reduction: mechanism and evolution. Photochem Photobiol 78:543–557
Schoefs B, Garnir HP, Bertrand M (1994) Comparison of the photoreduction protochlorophyllide to chlorophyllide in leaves and cotyledons from dark-grown beans as a function of age. Photosynth Res 41:405–417
Schoefs B, Bertrand M, Franck F (2000a) Spectroscopic properties of protochlorophyllide analyzed in situ in the course of etiolation and in illuminated leaves. Photochem Photobiol 72:85–93
Schoefs B, Bertrand M, Franck F (2000b) Photoactive protochlorophyllide regeneration in cotyledons and leaves from higher plants. Photochem Photobiol 72:660–668
Schon A, Krupp G, Gough S, Berry-Lowe S, Kannangara CG, Soll D (1986) The RNA required in the first step of chlorophyll biosynthesis is a chloroplast glutamate tRNA. Nature 322:281–284
Schon A, Kannangara CG, Gough S, Soll D (1988) Protein biosynthesis in organelles requires misaminoacylation of tRNA. Nature 331:187–190
Scolnik P, Bartley GE (1996) A table of some cloned plant genes involved in isoprenoid biosynthesis. Plant Mol Biol Report 14:305–319
Seyedi M, Selstam E, Timko MP, Sundqvist C (2001a) The cytokinin 2-isopentenyladenine causes partial reversion to skotomorphogenesis and induces formation of prolamellar bodies and protochlorophyllide657 in the lip1 mutant of pea. Physiol Plan 112:261–272
Seyedi M, Timko MP, Sundqvist C (2001b) The distribution of protochlorophyllide and chlorophyll within seedlings of the lip1 mutant of pea. Plant Cell Physiol 42:931–941
Shalygo N, Czarnecki O, Peter E, Grimm B (2009) Expression of chlorophyll synthase is also involved in feedback-control of chlorophyll biosynthesis. Plant Mol Biol 71:425–436
Shashidhara LS, Smith AG (1991) Expression and subcellular location of the tetrapyrrole synthesis enzyme porphobilinogen deaminase in light-grown Euglena gracilis and three nonchlorophyllous cell lines. Proc Natl Acad Sci USA 88:63–67
Shemin D (1976) 5-Aminolevulinic acid dehydratase: structure, function and mechanism. Philos Trans R Soc Lond 273:109–115
Shetty AS, Miller GW (1969) Purification and general properties of 5-aminolevulinate dehydratase from Nicotiana tabaccum L. Biochem J 114:331–337
Shibata K (1957) Spectroscopic studies on chlorophyll formation in intact leaves. J Biochem 44:147–173
Shibata H, Ochiai H (1977) Purification and properties of δ-aminolevulinic acid dehydratase from radish cotyledons. Plant Cell Physiol 18:420–429
Smeller L, Solymosi K, Fidy J, Böddi B (2003) Activation parameters of the blue shift (Shibata shift) subsequent to protochlorophyllide phototransformation. Biochim Biophys Acta 1651:130–138
Smith AG (1988) Subcellular localization of two porphyrin-synthesis enzymes in Pisum sativum (pea) and Arum (cuckoo-pint) species. Biochem J 249:423–428
Smith AG, Francis JE (1981) Investigations of rat liver uroporphyrinogen decarboxylase. comparisons of porphyrinogens I and III as substrate and inhibition by porphyrins. Biochem J 195:241–250
Soll J, Schultz G, Rüdiger W, Benz J (1983) Hydrogenation of geranylgeraniol. Two pathways exist in spinach chloroplasts. Plant Physiol 71:849–854
Solymosi K, Böddi B (2006) Optical properties of bud scales and protochlorophyll(ide) forms in leaf primodia of closed and opened buds. Tree Physiol 26:1075–1085
Solymosi K, Bóka K, Böddi B (2006) Transient etiolation: protochlorophyll(ide) and chlorophyll forms in differentiating plastids of closed and breaking leaf buds of horse chestnut (Aesculus hippocastanum). Tree Physiol 26:1087–1096
Sood S, Tyagi AK, Tripathy BC (2004) Inhibition of Photosystem I and Photosystem II in wheat seedlings with their root–shoot transition zones exposed to red light. Photosynth Res 81:31–40
Sood S, Gupta V, Tripathy BC (2005) Photoregulation of the greening process of wheat seedlings grown in red light. Plant Mol Biol 59:269–287
Spano AJ, Timko MP (1991) Isolation, characterization and partial amino acid sequence of a chloroplast-localized porphobilinogen deaminase from pea (Pisum sativum L.). Biochim Biophys Acta 1076:29–36
Spano AJ, He Z, Michel H, Hunt DF, Timko MP (1992) Molecular cloning, nuclear gene structure, and developmental expression of NADPH: protochlorophyllide oxidoreductase in pea (Pisum sativum L.). Plant Mol Biol 18:967–972
Spencer P, Jordan PM (1995) Characterization of the two 5-aminolevulinic acid binding sites of 5-aminolevulinic acid dehydratase from Escherichia coli. Biochem J 305:151–158
Sperling U, van Cleve B, Frick G, Apel K, Armstrong GA (1997) Overexpression of light-dependent PORA or PORB in plants depleted of endogenous POR by far-red light enhances seedling survival in white light and protects against photooxidative damage. Plant J 12:649–658
Sperling U, Franck F, Cleve BV, Frick G, Apel K, Armstrong GA (1998) Etioplast differentiation in Arabidopsis: Both PORA and PORB restore the prolamellar body and photoactive protochlorophyllide-F655 to the cop1 photomorphogenic mutant. Plant Cell 10:283–296
Staehelin LA, Newcomb EH (2000) Membrane structure and membrane organelles. In: Buchanan B, Gruissem W, Jones RJ (eds) Biochemistry and molecular biology of plants. American Society of Plant Biologists, Rockville, pp 19–22
Stephenson PG, Terry MJ (2008) Light signalling pathways regulating the Mg-chelatase branchpoint of chlorophyll synthesis during de-etiolation in Arabidopsis thaliana. Photochem Photobiol Sci 7:1243–1252
Su Q, Frick G, Armstrong G, Apel K (2001) POR C of Arabidopsis thaliana: a third light- and NADPH-dependent protochlorophyllide oxidoreductase that is differentially regulated by light. Plant Mol Biol 47:805–813
Sytina OA, Heyes DJ, Hunter CN, Alexandre MT, van Stokkum IH, van Grondelle R, Groot ML (2008) Conformational changes in an ultrafast light-driven enzyme determine catalytic activity. Nature 456:1001–1004
Tan FC, Cheng Q, Saha K, Heinemann IU, Jahn M, Jahn D, Smith AG (2008) Identification and characterization of the Arabidopsis gene encoding the tetrapyrrole biosynthesis enzyme uroporphyrinogen III synthase. Biochem J 410:291–299
Tanaka R, Tanaka A (2005) Effects of chlorophyllide a oxygenase overexpression on light acclimation in Arabidopsis thaliana. Photosynth Res 85:327–340
Tanaka R, Tanaka A (2007) Tetrapyrrole biosynthesis in higher plants. Annu Rev Plant Biol 58:321–346
Tanaka R, Yoshida K, Nakayashiki T, Masuda T, Tsuji H, Inokuchi H, Tanaka A (1996) Differential expression of two hemA mRNAs encoding glutamyl-tRNA reductase proteins in greening cucumber seedlings. Plant Physiol 110:1223–1230
Tanaka A, Ito H, Tanaka R, Tanaka NK, Yoshida K, Okada K (1998) Chlorophyll a oxygenase (CAO) is involved in chlorophyll b formation from chlorophyll a. Proc Natl Acad Sci USA 95:12719–12723
Tanaka R, Oster U, Kruse E, Rudiger W, Grimm B (1999) Reduced activity of geranylgeranyl reductase leads to loss of chlorophyll and tocopherol and to partially geranylgeranylated chlorophyll in transgenic tobacco plants expressing antisense RNA for geranylgeranyl reductase. Plant Physiol 120:695–704
Tanaka R, Koshino Y, Sawa S, Ishiguro S, Okada K, Tanaka A (2001) Overexpression of chlorophyllide a oxygenase (CAO) enlarges the antenna size of photosystem II in Arabidopsis thaliana. Plant J 26:365–373
Tewari AK, Tripathy BC (1998) Temperature-stress-induced impairment of chlorophyll biosynthetic reactions in cucumber and wheat. Plant Physiol 117:851–858
Tewari AK, Tripathy BC (1999) Acclimation of chlorophyll biosynthetic reactions to temperature stress in cucumber (Cucumis sativus L.). Planta 208:431–437
Tottey S, Block MA, Allen M, Westergren T, Albrieux C, Scheller HV, Merchant S, Jensen PE (2003) Arabidopsis CHL27, located in both envelope and thylakoid membranes, is required for the synthesis of protochlorophyllide. Proc Natl Acad Sci USA 100:16119–16124
Townley HE, Sessions RB, Clarke AR, Dafforn TR, Griffiths WT (2001) Protochlorophyllide oxidoreductase: a homology model examined by site-directed mutagenesis. Proteins 44:329–335
Tripathy BC, Pattanayak G (2012) Chlorophyll biosynthesis in higher plants. In: Eaton-Rye JJ, Tripathy BC, Sharkey TD (eds) Advances in photosynthesis and photorespiration, vol 34. Springer, Dordrecht, pp 63–94
Tripathy BC, Rebeiz CA (1985) Chloroplast biogenesis: quantitative determination of monovinyl and divinyl Mg-protoporphyrins and protochlorophyll(ides) by spectrofluorometry. Anal Biochem 149:43–61
Tripathy BC, Rebeiz CA (1986) Chloroplast biogenesis. Demonstration of the monovinyl and divinyl monocarboxylic routes of chlorophyll biosynthesis in higher plants. J Biol Chem 261:13556–13564
Tripathy BC, Rebeiz CA (1988) Chloroplast biogenesis 60: conversion of divinyl protochlorophyllide to monovinyl protochlorophyllide in green(ing) barley, a dark monovinyl/light divinyl plant species. Plant Physiol 87:89–94
Tsai S, Bishop DF, Desnick RJ (1987) Purification and properties of uroporphyrinogen III synthase from human erythrocytes. J Biol Chem 262:1268–1273
Virgin HI (1965) Chlorophyll formation and water deficit. Physiol Plant 18:994–1000
Walker CJ, Weinstein JD (1994) The magnesium-insertion step of chlorophyll biosynthesis is a two-stage reaction. Biochem J 299:277–284
Walker CJ, Willows RD (1997) Mechanism and regulation of Mg-chelatase. Biochem J 327:321–333
Walker CJ, Mansfield KE, Smith KM, Castelfranco PA (1989) Incorporation of atmospheric oxygen into the carbonyl functionality of the protochlorophyllide isocyclic ring. Biochem J 257:599–602
Walker CJ, Castelfranco PA, Whyte BJ (1991) Synthesis of divinyl protochlorophyllide. Enzymological properties of the Mg-protoporphyrin IX monomethyl ester oxidative cyclase system. Biochem J 276:691–697
Wang P, Gao J, Wan C, Zhang F, Xu Z, Huang X, Sun X, Deng X (2010) Divinyl chlorophyll (ide) a can be converted to monovinyl chlorophyll (ide) a by a divinyl reductase in rice. Plant Physiol 153:994–1003
Watanabe N, Che F-S, Iwano M, Takayama S, Yoshida S, Isogai A (2001) Dual targeting of spinach protoporphyrinogen oxidase II to mitochondria and chloroplast by alternative use in-frame initiation codons. J Biol Chem 276:20474–20481
Waters M, Pyke K (2005) Plastid development and differentiation. In: Moller SG (ed) Plastids. Blackwell, Oxford, pp 30–59
Whyte BJ, Castelfranco PA (1993) Further observations on the Mg-protoporphryin IX monomethyl ester (oxidative) cyclase system. Biochem J 290:355–359
Wiktorsson B, Engdahl S, Zhong LB, Böddi B, Ryberg M, Sundqvist C (1993) The effect of cross-linking of the subunits of NADPH-protochlorophyllide oxidoreductase of the aggregational state of protochlorophyllide. Photosynthtica 29:205–218
Wilks HM, Timko MP (1995) A light-dependent complementation system for analysis of NADPH: protochlorophyllide oxidoreductase: identification and mutagenesis of two conserved residues that are essential for enzyme activity. Proc Natl Acad Sci USA 92:724–728
Williams P, Hardeman K, Fowler J, Rivin C (2006) Divergence of duplicated genes in maize: evolution of contrasting targeting information for enzymes in the porphyrin pathway. Plant J 45:727–739
Witty M, Wallace-Cook AD, Albrecht H, Spano AJ, Michel H, Shabanowitz J, Hunt DF, Timko MP, Smith AG (1993) Structure and expression of chloroplast-localized porphobilinogen deaminase from pea (Pisum sativum L.) isolated by redundant polymerase chain reaction. Plant Physiol 103:139–147
Wu Z, Zhang X, He B, Diao L, Sheng S, Wang J, Guo X, Su N, Wang L, Jiang L, Wang C, Zhai H, Wan J (2007) A chlorophyll-deficient rice mutant with impaired chlorophyllide esterification in chlorophyll biosynthesis. Plant Physiol 145:29–40
Xu H, Vavilin D, Vermaas W (2001) Chlorophyll b can serve as the major pigment in functional photosystem II complexes of cyanobacteria. Proc Natl Acad Sci USA 98:14168–14173
Yamasato A, Nagata N, Tanaka R, Tanaka A (2005) The N-terminal domain of chlorophyllide a oxygenase confers protein instability in response to chlorophyll b accumulation in Arabidopsis. Plant Cell 17:1585–1597
Yamasato A, Tanaka R, Tanaka A (2008) Loss of the N-terminal domain of chlorophyllide a oxygenase induces photodamage during greening of Arabidopsis seedlings. BMC Plant Biol 8:64
Yaronskaya E, Vershilovskaya I, Poers Y, Alawady AE, Averina N, Grimm B (2006) Cytokinin effects on tetrapyrrole biosynthesis and photosynthetic activity in barley seedlings. Planta 224:700–709
Zhang H, Li J, Yoo JH, Yoo SC, Cho SH, Koh HJ, Seo HS, Paek NC (2006) Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol 62:325–337
Acknowledgments
Supported by a grant from the Department of Biotechnology, Government of India grant (BT/PR14827/BCE/08/841/2010), University Grants Commission capacity build up funds, and Department of Science and Technology purse grant from Jawaharlal Nehru University, New Delhi to BCT.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media Dordrecht
About this chapter
Cite this chapter
Tripathy, B.C., Dalal, V. (2013). Modulation of Chlorophyll Biosynthesis by Environmental Cues. In: Biswal, B., Krupinska, K., Biswal, U. (eds) Plastid Development in Leaves during Growth and Senescence. Advances in Photosynthesis and Respiration, vol 36. Springer, Dordrecht. https://doi.org/10.1007/978-94-007-5724-0_27
Download citation
DOI: https://doi.org/10.1007/978-94-007-5724-0_27
Published:
Publisher Name: Springer, Dordrecht
Print ISBN: 978-94-007-5723-3
Online ISBN: 978-94-007-5724-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)