Abstract
Lysophosphatidylserine (LysoPS), a deacylated form of phosphatidylserine (PS), has been assumed to serve as a bioactive lysophospholipid mediator. However, LysoPS has received little attention because its mode of actions as well as its synthetic pathways have been obscure. Recently G protein-coupled receptors (GPCRs) specific for LysoPS, and enzymes responsible for the generation of LysoPS have been identified. These LysoPS-related molecules are mainly expressed on immune cells, which led us to assume that LysoPS may have some roles in inflammation. Here we summarised the newly discovered LysoPS receptors and enzymes including our recent works.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Lysophospholipid (LPL, 1-acyl-2-LPLs or 2-acyl-1-LPLs) is a deacylated form of phospholipids with a single fatty acid chain. Various LPLs, including lysophosphatidylcholine (LPC), lysophosphatidic acid (LPA) , lysophosphatidylethanolamine (LPE), lysophosphatidylserine (LysoPS) , lysophosphatidylinositol (LPI), lysophosphatidylglycerol (LPG), sphingosine-1-phosphate (S1P) , and sphingosylphosphorylcholine (SPC) are present. They serve as precursors for diacyl phospholipids, and at least some of them also have roles as lipid mediators . Among them, LPA and S1P have been studied intensively on their receptors, producing enzymes, and catabolic enzymes (Mendelson et al. 2014; Yung et al. 2014). Studies using knockout mice and human hereditary diseases of receptors and synthetic enzymes of LPA and S1P have revealed their in vivo significance. Furthermore some of them were found to be drug targets and have received a lot of attention in terms of drug discovery. Notably FTY720 (Fingolimod), an S1P1 functional antagonist, has been approved for treatment of multiple sclerosis (Groves et al. 2013).
On the other hand, lysophosphatidylserine (LysoPS) is known to induce various cellular responses when applied in vivo or in vitro, but has received little attention because its receptors, producing pathways, and its in vivo occurrence have remained unknown. Recently LysoPS receptors and producing enzymes have been identified, which enables us to examine the role of LysoPS in vivo using knockout mice. In addition, the new methods using LC-MS/MS can sensitively detect LysoPS in vivo. In this review we summarise the recent advance in the study of LysoPS in terms of receptors, synthetic enzymes, and detection, with particular focus on the relevance of LysoPS to both physiological and pathological states.
2 LysoPS Receptors
Lysophosphatidylserine is known to induce several cellular responses both in vitro and in vivo. The most characterised response has been the stimulatory response of mast cell degranulation. In vitro, LysoPS enhances histamine release from peritoneal rodent mast cells triggered by the crosslinking of high-affinity IgE receptors (FcεRI) (Martin and Lagunoff 1979). It also induces rapid degranulation of mast cells and a consequent anaphylactic shock and hypothermia when administered i.v. in rodents. The mast cell degranulation-stimulating activity is not induced by other LPLs and strictly requires serine residue of LysoPS. This suggests the presence of a receptor for LysoPS on mast cells , although it remains unidentified.
In the course of a ligand fishing study of orphan GPCRs , Sugo et al. found that LysoPS is a ligand for GPR34, which is thought to be a member of P2Y family (Sugo et al. 2006). The P2Y family includes receptors for nucleotide, UDP-glucose, LPA , and the orphan GPCRs. They showed that LysoPS caused a dose-dependent inhibition of forskolin-stimulated cAMP accumulation in human GPR34-expressing Chinese hamster ovary (CHO) cells. They also showed that LysoPS induced phosphorylation of ERK in GPR34-expressing CHO cells. These responses were completely abolished by treatment of pertussis toxin (PTX), which indicates that GPR34 couples to a Gi/o-type G-protein. Later, Liebscher et al. and Makide et al. demonstrated that GPR34 are conserved in a wide range of vertebrates from fish to mammals by showing that GPR34 from carp and zebrafish react with LysoPS (Liebscher et al. 2011; Makide et al. 2014). Kitamura et al. used two assays for measuring GPR34 activation, a Ca2+ mobilisation assay and a newly developed transforming growth factor-α (TGFα) shedding assay, and confirmed that mammalian GPR34s from human, rat, and mouse origins reacted specifically with LysoPS but not with other LPLs (Kitamura et al. 2012). Notably, GPR34 reacted strongly with LysoPS species with an unsaturated fatty acid at the sn-2 position, showing that sn-2 unsaturated LysoPS is a preferable ligand for GPR34. GPR34 mRNA is expressed in many tissues but is most highly expressed in mast cells. Thus, it was once proposed that LysoPS enhances mast cell degranulation through GPR34. However, peritoneal mast cells from GPR34-deficient mice still responded to LysoPS, which suggested that GPR34 is not involved in the mast cell degranulation response induced by LysoPS. Until now the LysoPS receptor on mast cells has remained unknown.
Frasch et al. showed that LysoPS was generated in neutrophils by an oxidation-dependent mechanism and that is served as an endogenous anti-inflammatory mediator by stimulating the clearance of recruited neutrophils by macrophages, contributing to the resolution of inflammation (Frasch et al. 2008; Frasch and Bratton 2012). In addition, their results suggest that G2A (GPR132) on macrophages is responsible for the clearance of neutrophils by macrophages, raising the possibility that LysoPS is an endogenous ligand for G2A. G2A was once proposed as a receptor for LPC but the proposal was later retracted. Now many reports have confirmed that G2A is a receptor for protons (Murakami et al. 2004) and for 9-hydroxyoctadecadienoic acid (9-HODE), a kind of oxidised fatty acid (linoleic acid) (Obinata et al. 2005). We observed that G2A responded to 9-HODE, but not to LysoPS using a TGFα shedding assay (unpublished data). Thus, it is currently not clear if G2A directly recognises LysoPS.
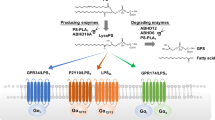
Recently, we identified three additional GPCRs (P2Y10, A630033H20, and GPR174) as novel LysoPS receptors through a ligand screening of orphan GPCRs using a TGFα shedding assay (Inoue et al. 2012). These receptors are members of the P2Y family, and are located in tandem in a clustered locus on the X chromosome. We propose that, according to the nomenclature of lysophospholipid receptors, GPR34, P2Y10, A630033H20, and GPR174 be designated as LPS1, LPS2, LPS2-like (LPS2L), and LPS3, respectively (Fig. 38.1).
P2Y10 is coupled with Gα12/13 but not with other G proteins. Expression of P2Y10 is restricted to lymphoid organs such as spleen, thymus , and lymph node. The expression of P2Y10 is dependent on PU.1 and Spi-B, which are closely related Ets transcription factors (Rao et al. 1999). In PU.1 +/−Spi-B−/− mice, the expression of P2Y10 is dramatically reduced. These Ets transcription factors have a role in the signal transduction of B-cell receptors (BCRs), which suggests that P2Y10 has a role in regulating BCR signalling. In addition, G12/13 signalling in B cells has a role in regulating the marginal zone B cells and germinal center B cells (Muppidi et al. 2014; Rieken et al. 2006), which raises the possibility that LysoPS functions in B cells through P2Y10.
Although human A630033H20 is truncated in the open reading frame and becomes a pseudogene, mouse and rat A630033H20 function as LysoPS receptors. A630033H20 is the closest homologue of P2Y10, with a 75 % homology to P2Y10 at the amino acid level, coupled with G12/13, and its expression pattern (it is highly expressed in lymphoid tissue) is similar to that of P2Y10.
GPR174 shows the highest homology to P2Y10 and A630033H20 with ~50 % identity at the amino acid level. Like P2Y10 and A630033H20, GPR174 is activated by LysoPS. The expression pattern of GPR174 is similar to the expression pattern of P2Y10 and A630033H20 with high expression in lymphoid tissues. However, GPR174 is also strongly expressed in some melanoma cells (Qin et al. 2011). GPR174 mainly coupled with both Gαs and Gα13. Given that Gα13 signalling is induced by the three LysoPS receptors (P2Y10, A630033H20, and GPR174), which have similar expression patterns, it is likely that these three LysoPS receptors share redundant functions in activating the Gα13 pathway. On the other hand, inasmuch as only one LysoPS receptor (GPR174) is coupled with Gαs, GPR174 may have a unique role in regulating Gαs signalling. Recent genome-wide association studies show that single nucleotide polymorphisms (SNPs) of GPR174 are associated with the risk for Graves’ disease (Chu et al. 2013; Szymanski et al. 2014) or Addison’s disease (Napier et al. 2015), autoimmune diseases. The expression of GPR174 is elevated in blood cells in vasovagal syncope patients (Huang et al. 2015). Thus, LysoPS may serve as an immunomodulator through GPR174.
Although P2Y10 was reported to be a receptor for LPA and S1P (Murakami et al. 2008), we and other researchers were unable to confirm it. In recent studies, we throughly examined the ligand specificity of cloned LysoPS receptors, that is, LPS1/GPR34, LPS2/P2Y10, and LPS3/GPR174, using chemically synthesised LysoPS analogues (Ikubo et al. 2015; Iwashita et al. 2009; Kitamura et al. 2012; Uwamizu et al. 2015). These studies revealed that modifications of the serine residue resulted in the loss of agonistic activity for all three LysoPS receptors, demonstrating that LysoPS receptors strictly recognise the head group of LysoPS, that is, serine. They also revealed that some modifications conferred certain selectivity toward each LysoPS receptor to the LysoPS analogues. For example, deoxy LysoPS, in which an sn-2 hydroxy group (−OH) was removed, was found to be selective for LPS2/P2Y10 and lysophosphatidylallothreonine, in which a methyl group (−CH3) was introduced in the serine residue, was found to be selective for LPS3/GPR174. These ligand specificities suggest that LysoPS is the real ligand for the LysoPS receptors. This is supported by the fact that all four LysoPS receptors belong to the P2Y family, to which the LPA receptors (LPA4–6) belong.
3 Generation of LysoPS
The main pathway of LysoPS production is probably the hydrolysis of PS by phospholipase (Figure 38.2). We recently established a method to detect LysoPS with high sensitivity (Okudaira et al. 2014). In this method, acyl migration reaction, in which an acyl chain of 2-acyl-1-LPLs is quickly moved to the sn-1 position, generating 1-acyl-2-LPLs, was completely suppressed by lowering the pH. With this method, we detected two types of LysoPS (1-acyl-2-LysoPS and 2-acyl-1-LysoPS) in various tissues and cells. The results, together with previous knowledge, suggest that both phospholipase A1 (PLA1) and A2 (PLA2) are involved in the production of LysoPS. In fact, rat platelets express two extracellular PLA enzymes, secretory PLA2 group IIA (sPLA2IIA) and PS-specific PLA1 (PS-PLA1), and secrete them upon activation. In the course of activation of rat platelets, sPLA2-IIA and PS-PLA1 produce 1-acyl LysoPS and 2-acyl LysoPS, respectively.
PS-PLA1 is stored in α granules of rat platelets and is secreted into the medium when activated. Although PS-PLA1 is structurally homologous to triglyceride (TG) lipase, it selectively hydrolyses PS and doesn’t have lipase activity for TG. PS-PLA1 expression is dramatically induced at the mRNA and protein levels by various inflammatory stimuli. Under some inflammatory conditions, we detected an increase of LysoPS in parallel with the induction of PS-PLA1 and found that the increase was partially abolished in PS-PLA1 knockout mice.
Because secretory PLA2 and PS-PLA1 are secreted proteins, they should act on PS extracellularly. On the contrary, PS localises exclusively to the inner leaflet of the plasma membrane. Recently, TMEM16F and Xkr8 were identified as scramblases that trigger exposure of PS in activated platelets and apoptotic cells, respectively (Suzuki et al. 2010; 2013). It is likely that secretory PLAs such as PS-PLA1 and sPLA2 deacylate PS exposed by TMEM16F or Xkr8 during platelet activation or apoptosis.
Recently ABHD16A was identified as a PS lipase that generates LysoPS (Kamat et al. 2015). ABHD16A, also known as lymphocyte antigen B-associated transcript 5 (BAT5), is a member of the alpha beta hydrolase domain (ABHD) enzyme family and is predicted to be a multipass membrane protein. In mice, ABHD16A mRNA is abundantly and ubiquitously expressed with highest expression in skeletal muscle and brain. The number of newborn ABHD16A−/− mice was much less than the value expected from Mendelian ratio. The body weight of ABHD16A−/− mice is smaller than wild-type mice, however, their behavior and survival rate appeared to be normal. The amount of various species of LysoPS in ABHD16A−/− brain were reduced, which suggests that ABHD16A does not discriminate between saturated fatty acids (mainly in the sn-1 position) and polyunsaturated fatty acids (mainly in the sn-2 position). Thioglycollate-elicited peritoneal macrophages derived from ABHD16A−/− mice have less LysoPS and release fewer inflammatory cytokines following stimulation with lipopolysaccharide. It is not known whether ABHD16A produces LysoPS intracellularly or extracellularly. If LysoPS is produced intracellularly, LysoPS must be released through some transporter on the plasma membrane such as Spns2 for S1P.
The degradation pathways of LysoPS are also important, because degradation is involved in the termination of LysoPS signaling. ABHD12 is reported to have lysoPS lipase activity in the mammalian brain (Blankman et al. 2013). In addition to having elevated brain LysoPS and microglial activation, ABHD12−/− mice show the phenotype of the human neurogenerative disorder PHARC (polyneuropathy, hearing loss, ataxia, retinosis pigmentosa , and cataract). Taken together, these results suggest that the interplay of ABHD12 and ABHD16A regulates the LysoPS level in neuronal diseases.
4 Conclusion
The LysoPS field has advanced with an expanding repertoire of receptors and metabolic enzymes. Recent studies strongly suggest that LysoPS plays important roles in processes related to inflammation. Further studies are needed to identify LysoPS-generating cells and enzymes, and LysoPS signaling pathways. The results of these studies will help to develop drugs that target LysoPS enzymes and receptors.
References
Blankman JL, Long JZ, Trauger SA, Siuzdak G, Cravatt BF (2013) ABHD12 controls brain lysophosphatidylserine pathways that are deregulated in a murine model of the neurodegenerative disease PHARC. Proc Natl Acad Sci U S A 110(4):1500–1505. doi:10.1073/pnas.1217121110
Chu X, Shen M, Xie F, Miao XJ, Shou WH, Liu L, Yang PP, Bai YN, Zhang KY, Yang L, Hua Q, Liu WD, Dong Y, Wang HF, Shi JX, Wang Y, Song HD, Chen SJ, Chen Z, Huang W (2013) An X chromosome-wide association analysis identifies variants in GPR174 as a risk factor for Graves’ disease. J Med Genet 50(7):479–485. doi:10.1136/jmedgenet-2013-101595
Frasch SC, Bratton DL (2012) Emerging roles for lysophosphatidylserine in resolution of inflammation. Prog Lipid Res 51(3):199–207. doi:10.1016/j.plipres.2012.03.001
Frasch SC, Berry KZ, Fernandez-Boyanapalli R, Jin HS, Leslie C, Henson PM, Murphy RC, Bratton DL (2008) NADPH oxidase-dependent generation of lysophosphatidylserine enhances clearance of activated and dying neutrophils via G2A. J Biol Chem 283(48):33736–33749. doi:10.1074/jbc.M807047200
Groves A, Kihara Y, Chun J (2013) Fingolimod: direct CNS effects of sphingosine 1-phosphate (S1P) receptor modulation and implications in multiple sclerosis therapy. J Neurol Sci 328(1–2):9–18. doi:10.1016/j.jns.2013.02.011
Huang YJ, Zhou ZW, Xu M, Ma QW, Yan JB, Wang JY, Zhang QQ, Huang M, Bao L (2015) Alteration of gene expression profiling including GPR174 and GNG2 is associated with vasovagal syncope. Pediatr Cardiol 36(3):475–480. doi:10.1007/s00246-014-1036-x
Ikubo M, Inoue A, Nakamura S, Jung S, Sayama M, Otani Y, Uwamizu A, Suzuki K, Kishi T, Shuto A, Ishiguro J, Okudaira M, Kano K, Makide K, Aoki J, Ohwada T (2015) Structure-activity relationships of lysophosphatidylserine analogs as agonists of G-protein-coupled receptors GPR34, P2Y10, and GPR174. J Med Chem 58(10):4204–4219. doi:10.1021/jm5020082
Inoue A, Ishiguro J, Kitamura H, Arima N, Okutani M, Shuto A, Higashiyama S, Ohwada T, Arai H, Makide K, Aoki J (2012) TGFalpha shedding assay: an accurate and versatile method for detecting GPCR activation. Nat Methods 9(10):1021–1029. doi:10.1038/nmeth.2172
Iwashita M, Makide K, Nonomura T, Misumi Y, Otani Y, Ishida M, Taguchi R, Tsujimoto M, Aoki J, Arai H, Ohwada T (2009) Synthesis and evaluation of lysophosphatidylserine analogues as inducers of mast cell degranulation. Potent activities of lysophosphatidylthreonine and its 2-deoxy derivative. J Med Chem 52(19):5837–5863. doi:10.1021/jm900598m
Kamat SS, Camara K, Parsons WH, Chen DH, Dix MM, Bird TD, Howell AR, Cravatt BF (2015) Immunomodulatory lysophosphatidylserines are regulated by ABHD16A and ABHD12 interplay. Nat Chem Biol 11(2):164–171. doi:10.1038/nchembio.1721
Kitamura H, Makide K, Shuto A, Ikubo M, Inoue A, Suzuki K, Sato Y, Nakamura S, Otani Y, Ohwada T, Aoki J (2012) GPR34 is a receptor for lysophosphatidylserine with a fatty acid at the sn-2 position. J Biochem 151(5):511–518. doi:10.1093/jb/mvs011
Liebscher I, Muller U, Teupser D, Engemaier E, Engel KM, Ritscher L, Thor D, Sangkuhl K, Ricken A, Wurm A, Piehler D, Schmutzler S, Fuhrmann H, Albert FW, Reichenbach A, Thiery J, Schoneberg T, Schulz A (2011) Altered immune response in mice deficient for the G protein-coupled receptor GPR34. J Biol Chem 286(3):2101–2110. doi:10.1074/jbc.M110.196659
Makide K, Uwamizu A, Shinjo Y, Ishiguro J, Okutani M, Inoue A, Aoki J (2014) Novel lysophosphoplipid receptors: their structure and function. J Lipid Res 55(10):1986–1995. doi:10.1194/jlr.R046920
Martin TW, Lagunoff D (1979) Interactions of lysophospholipids and mast cells. Nature 279(5710):250–252
Mendelson K, Evans T, Hla T (2014) Sphingosine 1-phosphate signalling. Development 141(1):5–9. doi:10.1242/dev.094805
Muppidi JR, Schmitz R, Green JA, Xiao W, Larsen AB, Braun SE, An J, Xu Y, Rosenwald A, Ott G, Gascoyne RD, Rimsza LM, Campo E, Jaffe ES, Delabie J, Smeland EB, Braziel RM, Tubbs RR, Cook JR, Weisenburger DD, Chan WC, Vaidehi N, Staudt LM, Cyster JG (2014) Loss of signalling via Galpha13 in germinal centre B-cell-derived lymphoma. Nature 516(7530):254–258. doi:10.1038/nature13765
Murakami N, Yokomizo T, Okuno T, Shimizu T (2004) G2A is a proton-sensing G-protein-coupled receptor antagonized by lysophosphatidylcholine. J Biol Chem 279(41):42484–42491. doi:10.1074/jbc.M406561200
Murakami M, Shiraishi A, Tabata K, Fujita N (2008) Identification of the orphan GPCR, P2Y(10) receptor as the sphingosine-1-phosphate and lysophosphatidic acid receptor. Biochem Biophys Res Commun 371(4):707–712. doi:10.1016/j.bbrc.2008.04.145
Napier C, Mitchell AL, Gan E, Wilson I, Pearce SH (2015) Role of the X-linked gene GPR174 in autoimmune Addison’s disease. J Clin Endocrinol Metab 100(1):E187–E190. doi:10.1210/jc.2014-2694
Obinata H, Hattori T, Nakane S, Tatei K, Izumi T (2005) Identification of 9-hydroxyoctadecadienoic acid and other oxidized free fatty acids as ligands of the G protein-coupled receptor G2A. J Biol Chem 280(49):40676–40683. doi:10.1074/jbc.M507787200
Okudaira M, Inoue A, Shuto A, Nakanaga K, Kano K, Makide K, Saigusa D, Tomioka Y, Aoki J (2014) Separation and quantification of 2-acyl-1-lysophospholipids and 1-acyl-2-lysophospholipids in biological samples by LC-MS/MS. J Lipid Res 55(10):2178–2192. doi:10.1194/jlr.D048439
Qin Y, Verdegaal EM, Siderius M, Bebelman JP, Smit MJ, Leurs R, Willemze R, Tensen CP, Osanto S (2011) Quantitative expression profiling of G-protein-coupled receptors (GPCRs) in metastatic melanoma: the constitutively active orphan GPCR GPR18 as novel drug target. Pigment Cell Melanoma Res 24(1):207–218. doi:10.1111/j.1755-148X.2010.00781.x
Rao S, Garrett-Sinha LA, Yoon J, Simon MC (1999) The Ets factors PU.1 and Spi-B regulate the transcription in vivo of P2Y10, a lymphoid restricted heptahelical receptor. J Biol Chem 274(48):34245–34252
Rieken S, Sassmann A, Herroeder S, Wallenwein B, Moers A, Offermanns S, Wettschureck N (2006) G12/G13 family G proteins regulate marginal zone B cell maturation, migration, and polarization. J Immunol 177(5):2985–2993
Sugo T, Tachimoto H, Chikatsu T, Murakami Y, Kikukawa Y, Sato S, Kikuchi K, Nagi T, Harada M, Ogi K, Ebisawa M, Mori M (2006) Identification of a lysophosphatidylserine receptor on mast cells. Biochem Biophys Res Commun 341(4):1078–1087
Suzuki J, Umeda M, Sims PJ, Nagata S (2010) Calcium-dependent phospholipid scrambling by TMEM16F. Nature 468(7325):834–838. doi:10.1038/nature09583
Suzuki J, Denning DP, Imanishi E, Horvitz HR, Nagata S (2013) Xk-related protein 8 and CED-8 promote phosphatidylserine exposure in apoptotic cells. Science 341(6144):403–406. doi:10.1126/science.1236758
Szymanski K, Miskiewicz P, Pirko K, Jurecka-Lubieniecka B, Kula D, Hasse-Lazar K, Krajewski P, Bednarczuk T, Ploski R (2014) rs3827440, a nonsynonymous single nucleotide polymorphism within GPR174 gene in X chromosome, is associated with Graves’ disease in Polish Caucasian population. Tissue Antigens 83(1):41–44. doi:10.1111/tan.12259
Uwamizu A, Inoue A, Suzuki K, Okudaira M, Shuto A, Shinjo Y, Ishiguro J, Makide K, Ikubo M, Nakamura S, Jung S, Sayama M, Otani Y, Ohwada T, Aoki J (2015) Lysophosphatidylserine analogues differentially activate three LysoPS receptors. J Biochem 157(3):151–160. doi:10.1093/jb/mvu060
Yung YC, Stoddard NC, Chun J (2014) LPA receptor signaling: pharmacology, physiology, and pathophysiology. J Lipid Res 55(7):1192–1214. doi:10.1194/jlr.R046458
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Japan
About this chapter
Cite this chapter
Makide, K., Inoue, A., Aoki, J. (2016). Lysophosphatidylserine as an Inflammatory Mediator. In: Miyasaka, M., Takatsu, K. (eds) Chronic Inflammation. Springer, Tokyo. https://doi.org/10.1007/978-4-431-56068-5_38
Download citation
DOI: https://doi.org/10.1007/978-4-431-56068-5_38
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-56066-1
Online ISBN: 978-4-431-56068-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)