Abstract
The liver can regenerate itself in response to acute liver damage. However, chronically induced liver dysfunction interferes with the liver regeneration process and increases the risk of onset of more severe hepatic failure, including hepatic cirrhosis and liver cancer. To develop more efficient therapeutics for chronic liver diseases, cell-based regenerative therapies using functional hepatocyte-like cells derived from pluripotent stem cells are actively under investigation. In addition to such stem cell–based approaches, recent studies have revealed that direct cell-fate conversion from fibroblasts into hepatocyte-like cells can be induced by forced expression of particular sets of transcription factors in fibroblasts. This phenomenon is known as “direct reprogramming” and is expected to be a complementary or alternative technology to the stem cell-based regenerative therapies. In this chapter, we briefly summarize the recent progress and future perspectives of studies on reprogramming technologies, which are directed at the development of cell-based regenerative therapies for liver diseases.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
The Liver and Disease
The liver is the central organ for metabolism, by which it produces energy for life activities and detoxifies various extrinsic and intrinsic harmful substances. In addition, the liver creates endocrine and exocrine materials, such as serum albumin, growth factors, and bile acids. The liver is developed from the foregut endoderm [1] and consists of parenchymal cells, such as hepatocytes, and various non-parenchymal cells, such as biliary epithelial cells, Kupffer cells, pit cells, hepatic stellate cells, mesothelial cells, and sinusoid endothelial cells [2]. Among the diverse kinds of cell populations, the major cell population in the liver is composed of hepatocytes, which play primary physiological roles in this organ.
The liver is one of the few organs capable of regeneration in the body. Since the ancient Greek era, the liver has been known to have a strong intrinsic regenerative ability in vivo. The famous “Prometheus” myth represents the rapid regeneration of this organ [3]. Indeed, deletion of two thirds of the total mass of the liver can be recovered completely in vivo [2]. However, chronic hepatitis generates a highly serious situation for the liver, and often leads to more severe hepatic failure, including hepatic cirrhosis and liver cancer. Thus, various conditions are attributed etiologically to chronic hepatitis. The most common cause of chronic hepatitis is infection by hepatotropic viruses, followed by alcoholic hepatitis, non-alcoholic fatty liver disease, drug-induced hepatitis, and autoimmune hepatitis [4].
Expectations for Cell-Based Regenerative Therapies
Many types of medical treatments have been applied to patients with chronic hepatitis. Although anti-inflammatory drugs, such as ursodeoxycholic acid, glycyrrhizin, and liver extracts, are widely used to treat all types of chronic hepatitis, such treatments are only symptomatic therapies. For patients with viral hepatitis, antiviral drugs, such as interferon, corticosteroid, ribavirin, lamivudine, and azathioprine, have been specifically employed. However, several side effects associated with these drugs, including fever and drug-resistant strains, appear with high frequency [4].
In the treatment of severe liver failure, organ transplantation is the ultimate solution and has a relatively high postoperative survival rate. However, there are many problems to be solved, including a chronic donor shortage, ethics, and immune rejection [4]. Therefore, cell-based regenerative therapies, which employ in vitro-expanded or newly generated hepatocytes, are expected to be the next-generation therapies.
Although hepatocytes have a high proliferative ability in vivo, this potential disappears immediately upon in vitro culture. Consequently, innovative technologies that can expand, maintain, mature, and create hepatocytes in vitro are required for future cell-based therapies for liver disorders.
Pluripotent Stem Cell–Based Approaches to Treat Liver Failure
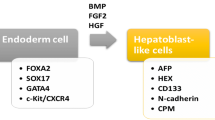
Among the current technologies aimed toward regenerative therapies for liver diseases, pluripotent stem cell–based approaches could be considered to show the most promise. Pluripotent stem cells can differentiate into any of the cell types responsible for the formation of particular tissues and organs, with the exception of extraembryonic tissues. Embryonic stem (ES) cells have been established as pluripotent stem cells and employed in basic research toward regenerative therapies for liver diseases. Hepatocyte differentiation from murine ES cells is generally induced via methods involving embryoid body (EB) formation [5–8]. Although EB-derived hepatocyte-like cells survive and function in vivo, their low differentiation frequency and associated teratoma formation remain as serious problems [9]. Several studies have tried to purify definitely differentiated hepatocytes or hepatic progenitor cells, using reporter expressions that are regulated by the promoters or enhancers of hepatocyte-specific genes, including albumin and α-fetoprotein (Afp) [10, 11]. Meanwhile, other studies have revealed the molecular mechanisms that regulate hepatocyte differentiation. In particular, employment of a liquidity factor, activin A, was found to efficiently improve both the ratio of hepatocyte differentiation and the quality of parental ES cells [12–14]. Other regulatory molecules required for hepatocyte differentiation have also been discovered [15–17]. By improving the in vitro hepatocyte differentiation of ES cells, many researchers have succeeded in transplantation of mouse/human ES cell–derived hepatocyte-like cells [18–20].
In 2006, Takahashi and Yamanaka [21] reported epoch-making artificial stem cells, termed “induced pluripotent stem” (iPS) cells. These novel stem cells can be expected to overcome the ethical and immunological problems associated with the use of ES cells [21, 22]. Soon after the publication of their study on iPS cells, many research groups started to employ these novel stem cells for in vitro hepatocyte differentiation using methods and knowledge accumulated in studies on ES cells [23–26]. Most recently, Takebe et al. [27] reported that vascularized and functional human liver bud-like structures could be formed by human iPS cells in culture and regenerate a part of the liver tissue upon transplantation. This technology has brought iPS cells closer to clinical reality. Nonetheless, iPS cells still have some problems to be solved, including tumorigenesis after transplantation and substantial costs. Thus, more vigorous basic studies are required for clinical-level applications using iPS cell technology.
Direct Reprogramming
Development of Direct Reprogramming Technologies
Recently, the strategy of direct cell-fate conversion from one cell type into another cell type, termed “direct reprogramming”, has rapidly expanded worldwide and is expected to be a complementary or alternative technology for future cell-based regenerative therapies using pluripotent stem cells. This method can generate both mature and progenitor-like cells in specific lineages from other types of cells. Thus, the cells required for treatment of particular diseases could be provided from an alternative source of cells and used as donor cells for transplantation therapy.
The first evidence for direct reprogramming was reported in the 1980s. Davis et al. [28] discovered a single transcription factor, Myod, that could induce fate conversion of fibroblasts into myoblasts. That sensational report encouraged researchers to look for single master transcription factors that could specify and govern the fate of cells in each lineage. However, studies on direct reprogramming subsequently went into decline for a long time, because further master transcription factors were not easy to discover.
In the last decade, the emergence of iPS cells has completely changed the situation for direct reprogramming studies [21, 22]. After the discovery of iPS cells, it was found that sets of transcription factors, rather than single transcription factors, could be successively identified as master regulators capable of inducing fate conversion of cells. At present, various types of cells have been induced from fibroblasts, including neurons [29], cardiomyocytes [30], hepatocytes [31, 32], adipocytes [33], Sertoli cells [34], and chondrocytes [35]. Furthermore, not only fibroblasts but also other types of cells, such as hepatocytes [36], astrocytes [37], Sertoli cells [38], and B cells [39], have been employed as sources of cells for direct reprogramming. These findings imply that direct reprogramming technology will become a universal method for almost all kinds of cells. Although the recent studies on direct reprogramming are summarized in Table 1, studies on direct reprogramming are continuing to increase year by year.
Regarding successful cases of direct reprogramming, three different cell lineages, cardiomyocytes, neurons, and hepatocytes, can be considered to be well-investigated targets for the following reasons: (1) functional failure of these cells is critical for the survival of individuals; (2) a number of patients suffer from diseases associated with malfunction of these cells; and (3) a large number of cells should be prepared in vitro prior to application of these cells for transplantation therapies. Among these three cell types, cardiomyocytes could be considered to be the most widely studied cells in the field of direct reprogramming. Following the publication of the first report by Ieda et al. [30], several groups reported similar methods for cardiomyocyte reprogramming [40–47]. However, the transcription factors used for the induction of cardiomyocyte-like cells were different in each study (Table 1), and the properties of the cells were also inhomogeneous [30, 40–47]. The obtained evidence suggested flexibility in the molecular mechanisms underlying direct reprogramming and a necessity for standardized protocols toward therapeutic applications.
The molecular machinery for the direct induction of neuronal cells in vitro can be considered to be the most deeply investigated example in the field of direct reprogramming. Vierbuchen et al. [29] showed that three transcription factors, Brm2, Ascl1, and Myt1l, induced conversion of mouse fibroblasts into neuron-like cells, designated induced neuronal (iN) cells. iN cells had neuron-specific characteristics, including neurite outgrowth, expression of specific neuronal markers, and electrophysiological activities. In addition to the discovery of iN cells, the same group revealed that remarkable epigenetic remodeling occurred in iN cells, and that Ascl1 acted as a pioneering factor that could activate closed chromatins during the reprogramming [48]. A better understanding of the molecular mechanisms controlling direct cell-fate conversion could provide great advantages for the development and application of direct reprogramming technology.
Direct Reprogramming of Fibroblasts to Hepatocytes
As mentioned above, functionally mature hepatocytes are strongly demanded for clinical use. To supply hepatocytes safely and stably without any ethical problems, direct reprogramming of patient-derived non-hepatic cells into hepatocytes appears to be a preferable method. Such newly generated hepatocytes could be technologically and economically useful for future cell-based regenerative therapies for liver diseases. The first studies showing direct conversion of non-hepatic cells into hepatocyte-like cells were independently reported by two groups: one was from our group [31] and the other was from Hui’s group [31, 32] (Fig. 1). Although both groups induced conversion of mouse fibroblasts into hepatocyte-like cells, the transcription factors used were different. Our group found that three combinations of two transcription factors, comprising a hepatocyte nuclear receptor, Hnf4α [49], and members of the forkhead domain-containing transcription factor family, Foxa1, Foxa2, and Foxa3 [50], could induce conversion of fibroblasts into hepatocyte-like cells, designated induced hepatocyte-like (iHep) cells [31]. In contrast, Hui’s group used a combination of three transcription factors, Gata4, Hnf1α, and Foxa3, with siRNA-based suppression of the expression of a tumor suppressor, p19Arf [32]. Although the reprogramming factors were different, iHep cells could be induced in both methods, and their morphological and functional properties were similar [31, 32]. These findings suggest the presence of lineage-specific core transcriptional networks that can determine the fate of cells.
iHep cells have multiple hepatocyte-specific properties: (1) the morphology and gene expression pattern of iHep cells resemble those of epithelial cells; (2) iHep cells express hepatocyte-specific genes and proteins; (3) iHep cells have functional features of hepatocytes, including glycogen storage, LDL uptake, ammonium metabolism, urea production, cytochrome P450 activity, and drug metabolism; and (4) iHep cells can reconstitute liver tissues, ameliorate hepatic functions of recipient mice, and rescue mice from a deadly hepatic disorder, hereditary tyrosinemia type I, upon transplantation into the liver of a mouse model of the disease. In addition to these features, we recently showed that iHep cells possess the potential to be involved in lipid metabolism, similar to hepatocytes [51]. These findings suggest that iHep cells are useful not only for screening of drugs, but also for the treatment of patients with fatty liver diseases. The morphology and function of iHep cells do not become attenuated during culture or after freeze-preservation, which could be great advantages in the application of iHep cells to cell-based regenerative therapies for liver diseases.
The above-mentioned characteristics suggest that iHep cells could be a potent cell source for future cell transplantation therapies. However, three problems remain to be resolved. First, the level of hepatic function is lower in iHep cells than in primary hepatocytes. Therefore, for use of iHep cells in cell transplantation therapies, the level of hepatic function in iHep cells needs to be improved by inducing the maturation of these cells. Second, viral vectors need to be excluded from the reprogramming procedures, because integration of the virus genomes into the host genomes may induce malignant transformation of cells. Thus, other reprogramming strategies using chemical compounds, growth factors, or non-integrative vectors are expected to be practical. Recently, Simeonov and Uppal [52] demonstrated the potential utility of an mRNA transfection-based technique for the generation of iHep cells. This technology may contribute to safer induction of iHep cells. Third, application of iHep cell technology to human cells is still in the development stage. Recently, two different groups reported data on the induction of human iHep cells [53, 54]. These studies showed that the technology for iHep cell generation in mouse cells could be reproducible in human cells. However, there are some critical differences between the methods for mouse and human iHep cell induction. In the case of human iHep cells, activation of MYC and SV40 large T antigen and suppression of TP53 are included in the methods for iHep cell generation, which have a risk for inducing malignant transformation of these cells (Fig. 1). Thus, it is expected that human iHep cells capable of proliferation without growth boosters, which can avoid the risk of tumorigenesis, should be generated for the application of iHep cells in transplantation therapies.
Conversely, Zhu et al. [55] reported a completely different strategy for induction of hepatocytes in vitro. They initially used OCT4, KLF4, SOX2, and MYC to induce iPS cells. However, they interrupted the conversion process of iPS cells and continued the culture of partially reprogrammed intermediate cells in the presence of various growth factors and chemical compounds that are known inducers of hepatic differentiation. The resultant hepatocyte-like cells showed hepatocyte-specific features like iHep cells. Thus, this method could be another alternative to stem cell–based regenerative therapies for liver failures, as well as the technology of iHep cell generation.
Conclusions
Cell-based regenerative medicine is one of the most promising clinical technologies in the treatment of liver disorders. However, it has remained unclear whether cell-based therapies are actually available in clinical settings without any risk for patients. To reduce the associated risks, the methods for cellular reprogramming and differentiation procedures should be improved in ES cell, iPS cell, and iHep cell technologies. As one of the important approaches, we need to understand the molecular mechanisms underlying the generation of iHep cells and the differentiation of hepatocytes, with a view to updating the technology for direct reprogramming and actualizing the functional maturation of iHep cells for use in cell-based regenerative therapies toward liver diseases. We believe that many current efforts in basic research will open up a new horizon for next-generation therapies using directly reprogrammed cells.
References
Zaret KS (2002) Regulatory phases of early liver development: paradigms of organogenesis. Nat Rev Genet 3:499–512. doi:10.1038/nrg837
Fausto N (2002) Regeneration of liver. eLS. doi:10.1038/npg.els.0001108
Taub R (2004) Liver regeneration: from myth to mechanism. Nat Rev Mol Cell Biol 5:836–847. doi:10.1038/nrm1489
Monga SPS (2011) Molecular pathology of liver diseases. Springer, New York
Hamazaki T, Iiboshi Y, Oka M, Papst PJ, Meacham AM, Zon LI, Terada N (2001) Hepatic maturation in differentiating embryonic stem cells in vitro. FEBS Lett 497:15–19. doi:10.1016/S0014-5793(01)02423-1
Chinzei R, Tanaka Y, Shimizu-Saito K, Hara Y, Kakinuma S, Watanabe M, Teramoto K, Arii S, Takase K, Sato C, Terada N, Teraoka H (2002) Embryoid-body cells derived from a mouse embryonic stem cell line show differentiation into functional hepatocytes. Hepatology 36:22–29. doi:10.1053/jhep.2002.34136
Yamamoto H, Quinn G, Asari A, Yamanokuchi H, Teratani T, Terada M, Ochiya T (2003) Differentiation of embryonic stem cells into hepatocytes: biological functions and therapeutic application. Hepatology 37:983–993. doi:10.1053/jhep.2003.50202
Imamura T, Cui L, Teng R, Johkura K, Okouchi Y, Asanuma K, Ogiwara N, Sasaki K (2004) Embryonic stem cell-derived embryoid bodies in three-dimensional culture system form hepatocyte-like cells in vitro and in vivo. Tissue Eng 10:1716–1724. doi:10.1089/ten.2004.10.1716
Kumashiro Y, Asahina K, Ozeki R, Shimizu-Saito K, Tanaka Y, Kida Y, Inoue K, Kaneko M, Sato T, Teramoto K, Arii S, Teraoka H (2005) Enrichment of hepatocytes differentiated from mouse embryonic stem cells as a transplantable source. Transplantation 79:550–557. doi:10.1097/01.TP.0000153637.44069.C6
Heo J, Factor VM, Uren T, Takahama Y, Lee J-S, Major M, Feinstone SM, Thorgeirsson SS (2006) Hepatic precursors derived from murine embryonic stem cells contribute to regeneration of injured liver. Hepatology 44:1478–1486. doi:10.1002/hep.21441
Drobinskaya I, Linn T, Saric T, Bretzel RG, Bohlen H, Hescheler J, Kolossov E (2008) Scalable selection of hepatocyte- and hepatocyte precursor-like cells from culture of differentiating transgenically modified murine embryonic stem cells. Stem Cells 26:2245–2256. doi:10.1634/stemcells.2008-0387
Tada S, Era T, Furusawa C, Sakurai H, Nishikawa S, Kinoshita M, Nakao K, Chiba T, Nishikawa S-I (2005) Characterization of mesendoderm: a diverging point of the definitive endoderm and mesoderm in embryonic stem cell differentiation culture. Development 132:4363–4374. doi:10.1242/dev.02005
Yasunaga M, Tada S, Torikai-Nishikawa S, Nakano Y, Okada M, Jakt LM, Nishikawa S, Chiba T, Era T, Nishikawa S-I (2005) Induction and monitoring of definitive and visceral endoderm differentiation of mouse ES cells. Nat Biotechnol 23:1542–1550. doi:10.1038/nbt1167
D’Amour KA, Agulnick AD, Eliazer S, Kelly OG, Kroon E, Baetge EE (2005) Efficient differentiation of human embryonic stem cells to definitive endoderm. Nat Biotechnol 23:1534–1541. doi:10.1038/nbt1163
Gouon-Evans V, Boussemart L, Gadue P, Nierhoff D, Koehler CI, Kubo A, Shafritz DA, Keller G (2006) BMP-4 is required for hepatic specification of mouse embryonic stem cell-derived definitive endoderm. Nat Biotechnol 24:1402–1411. doi:10.1038/nbt1258
Kubo A, Kim YH, Irion S, Kasuda S, Takeuchi M, Ohashi K, Iwano M, Dohi Y, Saito Y, Snodgrass R, Keller G (2010) The homeobox gene Hex regulates hepatocyte differentiation from embryonic stem cell-derived endoderm. Hepatology 51:633–641. doi:10.1002/hep.23293
Han S, Dziedzic N, Gadue P, Keller GM, Gouon-Evans V (2011) An endothelial cell niche induces hepatic specification through dual repression of Wnt and Notch signaling. Stem Cells 29:217–228. doi:10.1002/stem.576
Ishii T, Yasuchika K, Machimoto T, Kamo N, Komori J, Konishi S, Suemori H, Nakatsuji N, Saito M, Kohno K, Uemoto S, Ikai I (2007) Transplantation of embryonic stem cell-derived endodermal cells into mice with induced lethal liver damage. Stem Cells 25:3252–3260. doi:10.1634/stemcells.2007-0199
Sharma AD, Cantz T, Vogel A, Schambach A, Haridass D, Iken M, Bleidissel M, Manns MP, Schöler HR, Ott M (2008) Murine embryonic stem cell-derived hepatic progenitor cells engraft in recipient livers with limited capacity of liver tissue formation. Cell Transplant 17:313–323, http://dx.doi.org/10.3727/096368908784153896
Basma H, Soto-Gutiérrez A, Yannam GR, Liu L, Ito R, Yamamoto T, Ellis E, Carson SD, Sato S, Chen Y, Muirhead D, Navarro-Alvarez N, Wong RJ, Roy-Chowdhury J, Platt JL, Mercer DF, Miller JD, Strom SC, Kobayashi N, Fox IJ (2009) Differentiation and transplantation of human embryonic stem cell-derived hepatocytes. Gastroenterology 136:990–999. doi:10.1053/j.gastro.2008.10.047
Takahashi K, Yamanaka S (2006) Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126:663–676. doi:10.1016/j.cell.2006.07.024
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, Yamanaka S (2007) Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131:861–872. doi:10.1016/j.cell.2007.11.019
Song Z, Cai J, Liu Y, Zhao D, Yong J, Duo S, Song X, Guo Y, Zhao Y, Qin H, Yin X, Wu C, Che J, Lu S, Ding M, Deng H (2009) Efficient generation of hepatocyte-like cells from human induced pluripotent stem cells. Cell Res 19:1233–1242. doi:10.1038/cr.2009.107
Sullivan GJ, Hay DC, Park I-H, Fletcher J, Hannoun Z, Payne CM, Dalgetty D, Black JR, Ross JA, Samuel K, Wang G, Daley GQ, Lee J-H, Church GM, Forbes SJ, Iredale JP, Wilmut I (2010) Generation of functional human hepatic endoderm from human induced pluripotent stem cells. Hepatology 51:329–335. doi:10.1002/hep.23335
Li W, Wang D, Qin J, Liu C, Zhang Q, Zhang X, Yu X, Lahn BT, Mao FF, Xiang AP (2010) Generation of functional hepatocytes from mouse induced pluripotent stem cells. J Cell Physiol 222:492–501. doi:10.1002/jcp.22000
Espejel S, Roll GR, McLaughlin KJ, Lee AY, Zhang JY, Laird DJ, Okita K, Yamanaka S, Willenbring H (2010) Induced pluripotent stem cell-derived hepatocytes have the functional and proliferative capabilities needed for liver regeneration in mice. J Clin Invest 120:3120–3126. doi:10.1172/JCI43267
Takebe T, Sekine K, Enomura M, Koike H, Kimura M, Ogaeri T, Zhang R-R, Ueno Y, Zheng Y-W, Koike N, Aoyama S, Adachi Y, Taniguchi H (2013) Vascularized and functional human liver from an iPSC-derived organ bud transplant. Nature 499:481–484. doi:10.1038/nature12271
Davis RL, Weintraub H, Lassar AB (1987) Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell 51:987–1000. doi:10.1016/0092-8674(87)90585-X
Vierbuchen T, Ostermeier A, Pang ZP, Kokubu Y, Südhof TC, Wernig M (2010) Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463:1035–1041. doi:10.1038/nature08797
Ieda M, Fu J-D, Delgado-Olguin P, Vedantham V, Hayashi Y, Bruneau BG, Srivastava D (2010) Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell 142:375–386. doi:10.1016/j.cell.2010.07.002
Sekiya S, Suzuki A (2011) Direct conversion of mouse fibroblasts to hepatocyte-like cells by defined factors. Nature 475:390–393. doi:10.1038/nature10263
Huang P, He Z, Ji S, Sun H, Xiang D, Liu C, Hu Y, Wang X, Hui L (2011) Induction of functional hepatocyte-like cells from mouse fibroblasts by defined factors. Nature 475:386–389. doi:10.1038/nature10116
Zhu J, Pang D, Zhou Y, Tang X, Huang Y, Xie W, Gao F, Lai L, Zhang M, Ouyang H (2012) Direct conversion of porcine embryonic fibroblasts into adipocytes by chemical molecules. Cell Reprogram 14:99–105. doi:10.1089/cell.2011.0074
Buganim Y, Itskovich E, Hu Y-C, Cheng AW, Ganz K, Sarkar S, Fu D, Welstead GG, Page DC, Jaenisch R (2012) Direct reprogramming of fibroblasts into embryonic Sertoli-like cells by defined factors. Cell Stem Cell 11:373–386. doi:10.1016/j.stem.2012.07.019
Tam WL, DF O, Hiramatsu K, Tsumaki N, Luyten FP, Roberts SJ (2014) Sox9 reprogrammed dermal fibroblasts undergo hypertrophic differentiation in vitro and trigger endochondral ossification in vivo. Cell Reprogram 16:29–39. doi:10.1089/cell.2013.0060
Marro S, Pang ZP, Yang N, Tsai M-C, Qu K, Chang HY, Südhof TC, Wernig M (2011) Direct lineage conversion of terminally differentiated hepatocytes to functional neurons. Cell Stem Cell 9:374–382. doi:10.1016/j.stem.2011.09.002
Potts MB, Siu JJ, Price JD, Salinas RD, Cho MJ, Ramos AD, Hahn J, Margeta M, Oldham MC, Lim DA (2014) Analysis of Mll1-deficiency Identifies Neurogenic Transcriptional Modules and Brn4 as a Factor for Direct Astrocyte-to-neuron Reprogramming. Neurosurgery. doi:10.1227/NEU.0000000000000452
Sheng C, Zheng Q, Wu J, Xu Z, Wang L, Li W, Zhang H, Zhao X-Y, Liu L, Wang Z, Guo C, Wu H-J, Liu Z, Wang L, He S, Wang X-J, Chen Z, Zhou Q (2012) Direct reprogramming of Sertoli cells into multipotent neural stem cells by defined factors. Cell Res 22:208–218. doi:10.1038/cr.2011.175
Sadahira K, Fukuchi Y, Kunimono H, Sakurai M, Ikeda Y, Okamoto S, Nakajima H (2012) Direct reprogramming of terminally differentiated B cells into erythroid lineage. FEBS Lett 586:3645–3652. doi:10.1016/j.febslet.2012.08.019
Jayawardena TM, Egemnazarov B, Finch EA, Zhang L, Payne JA, Pandya K, Zhang Z, Rosenberg P, Mirotsou M, Dzau VJ (2012) MicroRNA-mediated in vitro and in vivo direct reprogramming of cardiac fibroblasts to cardiomyocytes. Circ Res 110:1465–1473. doi:10.1161/CIRCRESAHA.112.269035
Song K, Nam Y-J, Luo X, Qi X, Tan W, Huang GN, Acharya A, Smith CL, Tallquist MD, Neilson EG, Hill JA, Bassel-Duby R, Olson EN (2012) Heart repair by reprogramming non-myocytes with cardiac transcription factors. Nature 485:599–604. doi:10.1038/nature11139
Chen JX, Krane M, Deutsch M-A, Wang L, Rav-Acha M, Gregoire S, Engels MC, Rajarajan K, Karra R, Abel ED, Wu JC, Milan D, Wu SM (2012) Inefficient reprogramming of fibroblasts into cardiomyocytes using Gata4, Mef2c, and Tbx5. Circ Res 111:50–55. doi:10.1161/CIRCRESAHA.112.270264
Protze S, Khattak S, Poulet C, Lindemann D, Tanaka EM, Ravens U (2012) A new approach to transcription factor screening for reprogramming of fibroblasts to cardiomyocyte-like cells. J Mol Cell Cardiol 53:323–332. doi:10.1016/j.yjmcc.2012.04.010
Hirai H, Katoku-Kikyo N, Keirstead SA, Kikyo N (2013) Accelerated direct reprogramming of fibroblasts into cardiomyocyte-like cells with the MyoD transactivation domain. Cardiovasc Res 100:105–113. doi:10.1093/cvr/cvt167
Wada R, Muraoka N, Inagawa K, Yamakawa H, Miyamoto K, Sadahiro T, Umei T, Kaneda R, Suzuki T, Kamiya K, Tohyama S, Yuasa S, Kokaji K, Aeba R, Yozu R, Yamagishi H, Kitamura T, Fukuda K, Ieda M (2013) Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc Natl Acad Sci U S A 110:12667–12672. doi:10.1073/pnas.1304053110
Fu J-D, Stone NR, Liu L, Spencer CI, Qian L, Hayashi Y, Delgado-Olguin P, Ding S, Bruneau BG, Srivastava D (2013) Direct Reprogramming of Human Fibroblasts toward a Cardiomyocyte-like State. Stem Cell Rep 1:235–247. doi:10.1016/j.stemcr.2013.07.005
Muraoka N, Yamakawa H, Miyamoto K, Sadahiro T, Umei T, Isomi M, Nakashima H, Akiyama M, Wada R, Inagawa K, Nishiyama T, Kaneda R, Fukuda T, Takeda S, Tohyama S, Hashimoto H, Kawamura Y, Goshima N, Aeba R, Yamagishi H, Fukuda K, Ieda M (2014) MiR-133 promotes cardiac reprogramming by directly repressing Snai1 and silencing fibroblast signatures. EMBO J 33:1565–1581. doi:10.15252/embj.201387605
Wapinski OL, Vierbuchen T, Qu K, Lee QY, Chanda S, Fuentes DR, Giresi PG, Ng YH, Marro S, Neff NF, Drechsel D, Martynoga B, Castro DS, Webb AE, Südhof TC, Brunet A, Guillemot F, Chang HY, Wernig M (2013) Hierarchical mechanisms for direct reprogramming of fibroblasts to neurons. Cell 155:621–635. doi:10.1016/j.cell.2013.09.028
Hwang-Verslues WW, Sladek FM (2010) HNF4α–role in drug metabolism and potential drug target? Curr Opin Pharmacol 10:698–705. doi:10.1016/j.coph.2010.08.010
Lam EW-F, Brosens JJ, Gomes AR, Koo C-Y (2013) Forkhead box proteins: tuning forks for transcriptional harmony. Nat Rev Cancer 13:482–495. doi:10.1038/nrc3539
Miura S, Suzuki A (2014) Acquisition of lipid metabolic capability in hepatocyte-like cells directly induced from mouse fibroblasts. Front Cell Dev Biol 2:43
Simeonov KP, Uppal H (2014) Direct reprogramming of human fibroblasts to hepatocyte-like cells by synthetic modified mRNAs. PLoS One 9, e100134. doi:10.1371/journal.pone.0100134
Huang P, Zhang L, Gao Y, He Z, Yao D, Wu Z, Cen J, Chen X, Liu C, Hu Y, Lai D, Hu Z, Chen L, Zhang Y, Cheng X, Ma X, Pan G, Wang X, Hui L (2014) Direct reprogramming of human fibroblasts to functional and expandable hepatocytes. Cell Stem Cell 14:370–384. doi:10.1016/j.stem.2014.01.003
Du Y, Wang J, Jia J, Song N, Xiang C, Xu J, Hou Z, Su X, Liu B, Jiang T, Zhao D, Sun Y, Shu J, Guo Q, Yin M, Sun D, Lu S, Shi Y, Deng H (2014) Human hepatocytes with drug metabolic function induced from fibroblasts by lineage reprogramming. Cell Stem Cell 14:394–403. doi:10.1016/j.stem.2014.01.008
Zhu S, Rezvani M, Harbell J, Mattis AN, Wolfe AR, Benet LZ, Willenbring H, Ding S (2014) Mouse liver repopulation with hepatocytes generated from human fibroblasts. Nature 508:93–97. doi:10.1038/nature13020
Berninger B, Costa MR, Koch U, Schroeder T, Sutor B, Grothe B, Götz M (2007) Functional properties of neurons derived from in vitro reprogrammed postnatal astroglia. J Neurosci 27:8654–8664. doi:10.1523/JNEUROSCI.1615-07.2007
Heinrich C, Blum R, Gascón S, Masserdotti G, Tripathi P, Sánchez R, Tiedt S, Schroeder T, Götz M, Berninger B (2010) Directing astroglia from the cerebral cortex into subtype specific functional neurons. PLoS Biol 8, e1000373. doi:10.1371/journal.pbio.1000373
Caiazzo M, Dell’Anno MT, Dvoretskova E, Lazarevic D, Taverna S, Leo D, Sotnikova TD, Menegon A, Roncaglia P, Colciago G, Russo G, Carninci P, Pezzoli G, Gainetdinov RR, Gustincich S, Dityatev A, Broccoli V (2011) Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature 476:224–227. doi:10.1038/nature10284
Pang ZP, Yang N, Vierbuchen T, Ostermeier A, Fuentes DR, Yang TQ, Citri A, Sebastiano V, Marro S, Südhof TC, Wernig M (2011) Induction of human neuronal cells by defined transcription factors. Nature 476:220–223. doi:10.1038/nature10202
Pfisterer U, Kirkeby A, Torper O, Wood J, Nelander J, Dufour A, Björklund A, Lindvall O, Jakobsson J, Parmar M (2011) Direct conversion of human fibroblasts to dopaminergic neurons. Proc Natl Acad Sci U S A 108:10343–10348. doi:10.1073/pnas.1105135108
Cobaleda C, Jochum W, Busslinger M (2007) Conversion of mature B cells into T cells by dedifferentiation to uncommitted progenitors. Nature 449:473–477. doi:10.1038/nature06159
Suzuki T, Nakano-Ikegaya M, Yabukami-Okuda H, de Hoon M, Severin J, Saga-Hatano S, Shin JW, Kubosaki A, Simon C, Hasegawa Y, Hayashizaki Y, Suzuki H (2012) Reconstruction of monocyte transcriptional regulatory network accompanies monocytic functions in human fibroblasts. PLoS One 7, e33474. doi:10.1371/journal.pone.0033474
Laiosa CV, Stadtfeld M, Xie H, de Andres-Aguayo L, Graf T (2006) Reprogramming of committed T cell progenitors to macrophages and dendritic cells by C/EBP alpha and PU.1 transcription factors. Immunity 25:731–744. doi:10.1016/j.immuni.2006.09.011
Feng R, Desbordes SC, Xie H, Tillo ES, Pixley F, Stanley ER, Graf T (2008) PU.1 and C/EBPalpha/beta convert fibroblasts into macrophage-like cells. Proc Natl Acad Sci U S A 105:6057–6062. doi:10.1073/pnas.0711961105
Szabo E, Rampalli S, Risueño RM, Schnerch A, Mitchell R, Fiebig-Comyn A, Levadoux-Martin M, Bhatia M (2010) Direct conversion of human fibroblasts to multilineage blood progenitors. Nature 468:521–526. doi:10.1038/nature09591
Ono Y, Wang Y, Suzuki H, Okamoto S, Ikeda Y, Murata M, Poncz M, Matsubara Y (2012) Induction of functional platelets from mouse and human fibroblasts by p45NF-E2/Maf. Blood 120:3812–3821. doi:10.1182/blood-2012-02-413617
Bredenkamp N, Ulyanchenko S, O’Neill KE, Manley NR, Vaidya HJ, Blackburn CC (2014) An organized and functional thymus generated from FOXN1-reprogrammed fibroblasts. Nat Cell Biol. doi:10.1038/ncb3023
Ginsberg M, James D, Ding B-S, Nolan D, Geng F, Butler JM, Schachterle W, Pulijaal VR, Mathew S, Chasen ST, Xiang J, Rosenwaks Z, Shido K, Elemento O, Rabbany SY, Rafii S (2012) Efficient direct reprogramming of mature amniotic cells into endothelial cells by ETS factors and TGFβ suppression. Cell 151:559–575. doi:10.1016/j.cell.2012.09.032
Zhou Q, Brown J, Kanarek A, Rajagopal J, Melton DA (2008) In vivo reprogramming of adult pancreatic exocrine cells to beta-cells. Nature 455:627–632. doi:10.1038/nature07314
Kajimura S, Seale P, Kubota K, Lunsford E, Frangioni JV, Gygi SP, Spiegelman BM (2009) Initiation of myoblast to brown fat switch by a PRDM16-C/EBP-beta transcriptional complex. Nature 460:1154–1158. doi:10.1038/nature08262
Desgraz R, Herrera PL (2009) Pancreatic neurogenin 3-expressing cells are unipotent islet precursors. Development 136:3567–3574. doi:10.1242/dev.039214
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution Noncommercial License, which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Copyright information
© 2015 The Author(s)
About this paper
Cite this paper
Horisawa, K., Suzuki, A. (2015). Cell-Based Regenerative Therapy for Liver Disease. In: Nakao, K., Minato, N., Uemoto, S. (eds) Innovative Medicine. Springer, Tokyo. https://doi.org/10.1007/978-4-431-55651-0_27
Download citation
DOI: https://doi.org/10.1007/978-4-431-55651-0_27
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-55650-3
Online ISBN: 978-4-431-55651-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)