Abstract
The number of reports involving the new tools of optogenetics is increasing exponentially to yield detailed insights into anatomical, physiological, and pathological issues. These tools help us to tackle major questions regarding the function of neural circuits in the mammalian brain, which possesses uncountable combinations of neurons. Moreover, rapid progress in diverse collaborations between optogenetics and optical imaging technologies will allow us to analyze, simultaneously, the activities of multiple neurons and glial cells. As well as activity analysis, optogenetics is developing rapidly to support the analysis of stimulation in neuronal function. We can now stimulate multiple cell types independently using selective molecular tools, such as promoters and gene delivery systems. In addition, optical properties also help us to discriminate among subpopulations of cells in neuronal networks. The use of light to study the brain has proved to be a remarkably fruitful strategy, and indeed optogenetics has given us a green light for the future.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Optogenetics has been applied to various neurobiological questions to elucidate in detail the mechanisms underlying behavior. Optogenetics was established in 2006 by Karl Deisseroth and colleagues, who expressed a photoactivatable protein called channelrhodopsin-2 (ChR2) in particular neurons to stimulate them specifically by light. ChR2 thus allows us to control neurons with high spatiotemporal resolution. As with calcium imaging, optogenetics provides a molecular tool with which single neurons, or populations of neurons, within a network of interest can be studied.
1.1 Optogenetics
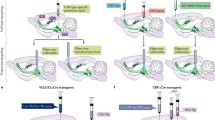
Trillions of synapses connecting neurons are included in the whole network of the mammalian brain (Luo et al. 2008). The components of this huge network, such as neurons and glias, are well characterized in all areas of the brain. These components build up not only a relatively simple system but also an incredibly complex system, corresponding to sensory or motor processing and higher cognitive functions, such as the circuits that mediate vision, motor movements, breathing/respiration, and sleep/wake architecture. At both levels of complexity, the relative contributions of individual cells and the synaptic connections between them should be elucidated to understand the mechanisms of information processing in the brain. However, precise manipulation of the activities of the network has been extremely challenging for traditional electrophysiological, pharmacological, and genetic methods (Luo et al. 2008; Carter and Shieh 2010). While numerous successes have been reported with these classical methods, we have faced considerable obstacles to achieving spatiotemporal precision in the study of neural circuits in vivo. Conventional electrical and physical techniques are spatially imprecise, since surrounding cells are also stimulated or inhibited. Pharmacological and genetic methods yield better data than electrical and physical approaches in terms of spatial selectivity, but temporal resolution is still insufficient for single action potentials. To overcome these limitations, optogenetics has been developed as a new set of tools to precisely stimulate (Boyden et al. 2005; Zhang et al. 2008; Berndt et al. 2009; Lin et al. 2009; Gunaydin et al. 2010; Li et al. 2005; Nagel et al. 2003), inhibit neural activity (Chow et al. 2010; Gradinaru et al. 2010; Zhao et al. 2008; Gradinaru et al. 2008; Zhang et al. 2007a, b), or alter biochemical activity in specific cells (Airan et al. 2009; Oh et al. 2010) with high temporal precision and rapid reversibility. These tools are activated by light (“opto-”) and are genetically encoded (“-genetics”) to give us specific control over particular populations of cells in vitro and in vivo (Fig. 8.1) (Zhang et al. 2006; Gradinaru et al. 2007; Zhang et al. 2007a, b; Zhang et al. 2010; Cardin et al. 2010). The precise manipulation that these new tools permit has facilitated further progress in elucidating structural and functional aspects of neural circuits.
Concept of optogenetics. Rhodopsin molecules (here, ChR2, shown on the membrane in the foreground and on the neuron just behind it) are expressed in a particular type of neurons using molecular tools such as viral vectors or transgenic animals. When a neuron is stimulated by light, cations (green) enter the cell, which results in excitement of the ChR2-expressing neurons (shown as bright spots in the dendrites). Optical fibers, through which a stimulus light source is introduced, are available for in vivo experiments
2 Revealing Dynamism Based on Physiological and Pathological Neuronal Networks
2.1 Retina
Since retinal degeneration is characterized by the progressive loss of rod and cone photoreceptors, targeting ChR2 to retinal ganglion cells (RGCs) or bipolar cells could theoretically restore light sensitivity to mammalian models of retinal degeneration in which most rods and cones are genetically ablated (Tomita et al. 2009). Expression of ChR2 in either RGCs (Bi et al. 2006) or bipolar cells (Lagali et al. 2008) could indeed restore visual sensitivity in mice with mutations causing rod and cone degeneration, as well as in rat models of retinal degeneration caused either by genetic mutation (Tomita et al. 2010) or toxic light exposure (Tomita et al. 2009). Halorhodopsin (NpHR) could potentially be used in these cells to restore vision, because photons naturally cause hyperpolarization of photoreceptors. Interestingly, expression of NpHR in the inner retinal layer could restore OFF responses, whereas combined expression of NpHR and ChR2 in RGCs could cause them to respond as ON, OFF, or even ON-OFF cells depending on the wavelength of light used (Zhang et al. 2009). The expression of “enhanced” NpHR (eNpHR) in light-insensitive cone photoreceptors could substitute for the native phototransduction cascade and restore light sensitivity in two mouse models of retinal degeneration (Busskamp et al. 2010). Importantly, this treatment leads to normal activity in cone photoreceptors and RGCs in response to yellow-light stimulation and allows mice to respond to changes in light intensity and the direction of motion of visual stimulation.
Even in human retinas, eNpHR expression was nontoxic and could rescue light-insensitive human photoreceptors ex vivo (Busskamp et al. 2010). These results demonstrate that optogenetics may be useful for treating various forms of blindness in humans. Optogenetics may also be utilized to establish various prosthetic devices, such as specialized glasses that increase light intensity, which have been proposed to enhance environmental visual stimuli specifically for ChR2- or eNpHR-transduced neurons in the retina (Cepko 2010).
2.2 Reflex and Innate Behavior
2.2.1 Breathing/Respiration
Brainstem or spinal cord injury may result in paralysis, which leads to the inability to breathe in severe cases. Although the neuronal mechanism of respiration is not sufficiently well understood to restore function to damaged circuits, ChR2-mediated photostimulation of motor neurons has recently been reported to recover respiratory diaphragmatic motor activity (Alilain et al. 2008). In the brainstem, stimulation of the retrotrapezoid nucleus produced long-lasting activation of breathing (Abbott et al. 2009a), whereas stimulation of the ventrolateral medulla increased sympathetic nerve activity and blood pressure (Abbott et al. 2009b). These studies indicate that neural and non-neural cell types in the brainstem and spinal cord contribute to the regulation of a central autonomic process.
2.2.2 Sleep/Wake Circuitry
The sleep/wake cycle is one of the most well-defined behaviors whose underlying principles have been dissected by optogenetic tools. The first use of optogenetics to study this system used lentivirus-mediated gene delivery to target ChR2 to hypocretin-expressing neurons in the lateral hypothalamus (Adamantidis et al. 2007). Dysregulation of the hypocretin system, either of the peptide or its receptor, causes the sleep disorder narcolepsy. Photostimulation on ChR2-expressing hypocretin neurons increased the probability of sleep/wake transitions (Adamantidis et al. 2007) and increased neuronal activity in downstream wake-promoting nuclei (Carter et al. 2009). It has also been shown that optogenetic stimulation of the locus coeruleus (LC) produces immediate sleep-to-wake transitions, whereas optogenetic inhibition causes a decrease in wakefulness (Carter et al. 2010). In addition, tonic stimulation of the LC led the mice to a cataplexy-like state, which is a sign of narcolepsy.
2.3 Motor Behavior
The selective loss of dopaminergic neurons in the substantia nigra pars compacta leads to a severe neurodegenerative disorder known as Parkinson’s disease, characterized by muscle rigidity and uncoordinated physical movements. Two major output routes from striatal neurons, known as the direct and indirect pathways, are thought to be related to this disease. The indirect pathway output neurons express a specific dopamine receptor called dopamine receptor type 2 (D2R), a G-protein-coupled receptor that inhibits adenylate cyclase and thus suppresses calcium signaling. Stimulation of D2R by dopamine leads to suppression of the indirect pathway. Under normal circumstances, the indirect pathway suppresses the inhibitory output of the external capsule of globus pallidus (GPe), which is involved in suppressing the subthalamic nucleus (STN). It is generally thought that losing dopamine results in increased indirect pathway activity, due to the loss of D2R-mediated suppression of this pathway. When GPe can no longer function as a major inhibitor of the STN, the ensuing net increase in STN activity may explain the motor symptoms of Parkinson’s disease. Optogenetics has recently clarified that activation of the indirect pathway indeed mimics the parkinsonian state (Kravitz et al. 2010). In this study, the relative contributions of dopamine D1-receptor- and D2-receptor-expressing neurons in the striatum were investigated by selectively targeting each type with ChR2. Stimulation of D1-expressing neurons in the striatum reduced parkinsonian symptoms in a mouse model of the disease, whereas stimulation of D2-expressing neurons—which mimics dopamine depletion of indirect pathway—caused symptoms in wild-type mice.
In addition to deepening our understanding of Parkinson’s disease circuitry, another recent optogenetics study shed light on how one of the traditional treatments for this disease may be relieving the symptoms. A crude technique called deep-brain electrical stimulation within STN causes a reversal of Parkinson’s disease symptoms. To understand the underlying principles of this phenomenon, optogenetic probes were used to systematically stimulate or inhibit a mixture of distinct circuit elements containing neurons, glia, and fiber projections in the STN of freely moving rodent models of Parkinson’s disease (Gradinaru et al. 2009). When the excitatory afferent axons projecting to the STN—such as those of motor neurons in M1—were optogenetically stimulated, the parkinsonian symptoms were relieved. This study demonstrated that the excitatory afferent regulation of STN is at the heart of the beneficial outcome of deep-brain stimulation. Taken together, these results promote our understanding of the functional connections within the basal ganglia and may contribute to current therapeutic strategies to ameliorate parkinsonian motor deficits (Bernstein et al. 2008).
2.4 Memory Formation and Reinforcement
The neural circuits based on reinforcement have been well characterized with optogenetic tools. Stimulation of dopaminergic neurons in the ventral tegmental area co-released glutamate as well as dopamine into the nucleus accumbens, demonstrating that mesolimbic reward signaling involves glutamatergic transmission (Tecuapetla et al. 2010; Stuber et al. 2010). Optical stimulation of α1-adrenergic receptors in the nucleus accumbens, but not of β2-adrenergic receptors, led to a robust increase in place preference during conditioning (Airan et al. 2009). In addition, the relative contributions of distinct tonic versus phasic activity patterns in participating brain structures, as well as the relative contributions of modulatory systems with various neurotransmitters, are unknown (Stuber 2010). By means of optogenetics, selective phasic photostimulation of dopaminergic neurons in the ventral tegmental area was shown to be sufficient to establish association learning, whereas tonic activation was not (Tsai et al. 2009).
2.5 Anxiety and Aggression
Even a fairly deep part of the brain, the ventromedial hypothalamus (VMH), which is thought to be closely related to instinctive behaviors, could be photostimulated through an implanted optical fiber. Optogenetic, but not electrical, stimulation of neurons in the VMH ventrolateral subdivision (VMHvl) causes male mice to attack both females and inanimate objects, as well as males (Lin et al. 2011).
Another group, using of ChR2-assisted circuit mapping in amygdala slices and cell-specific viral tracing, has reported that protein kinase C-d (PKC-d)1 neurons inhibit output neurons in the medial central amygdala (CEm) and also make reciprocal inhibitory synapses with PKC-d2 neurons in the lateral subdivision of the central amygdala (CEl). These results, together with behavioral data, define an inhibitory microcircuit in CEl that gates CEm output to control the level of conditioned freezing (Haubensak et al. 2010). In another study, specific optogenetic stimulation of oxytocinergic axons in the amygdala was shown to reduce freezing responses in fear-conditioned rats, illuminating the mechanisms by which oxytocin modifies emotional circuitry in a positive manner (Knobloch et al. 2012).
2.6 Balance Between Excitatory and Inhibitory Networks
Various oscillatory fluctuations have been observed in the cortex, which is thought to be associated with particular behavior. Cortical gamma oscillations (30–100 Hz) have been well elucidated, but the neural basis of these rhythms, and their role in animal behavior, remain unknown. ChR2-mediated photostimulation of parvalbumin (PV)-expressing interneurons amplified gamma oscillations, whereas eNpHR-mediated photoinhibition suppressed them (Sohal et al. 2009; Cardin et al. 2009). Furthermore, γ-frequency modulation of excitatory input enhanced signal transduction in cortical regions, reducing circuit noise and amplifying circuit signals. These studies provide the first causal evidence that distinct network activity states can be induced in vivo by cell-type-specific activation of PV neurons, and also suggest a potential mechanism for the altered γ-frequency synchronization and cognition in schizophrenia and autism (Sohal et al. 2009; Cardin et al. 2009). In another recent study, optogenetic stimulation of layer VI excitatory neurons was shown to reduce firing within the upper layers of the mouse visual cortex. This suggests that activation of layer VI excitatory neurons plays an essential role in gain control within cortical sensory networks (Olsen et al. 2012). In addition to the neurological studies described above, optogenetic probes have been used to investigate many other aspects of health and disease, including associative fear memory (Haubensak et al. 2010; Ciocchi et al. 2010), epilepsy (Tonnesen et al. 2009), and the blood oxygen level-dependent (BOLD) effect during functional magnetic resonance imaging (Lee et al. 2010).
3 Technical Aspects
3.1 Molecular Aspects
Although there are notable exceptions, the most commonly used optogenetic probes are gene-engineered versions of natural opsins, which are light-sensitive membrane proteins through which ions are translocated in response to light stimulation at specific wavelengths (Kramer et al. 2009). These probes can be utilized either to excite the cells, to inhibit their activity, or to change intracellular signaling (Fig. 8.2).
Examples of optogenetics. Membrane excitability is manipulated using various optogenetic tools. ChR2 and NpHR allow us to excite and inhibit particular neurons, respectively, and thereby to control excitability by applying different excitation wavelengths even when these modified neurons coexist in a particular region of interest. Similarly, intracellular signaling can also be modified with light-triggered G-proteins, “OptoXRs.” The colored balls in blue, yellow, and green indicate cation, anion, and intracellular signal, respectively
3.1.1 Probes for Stimulating Neurons
ChR2 is a nonspecific cation channel naturally expressed in the alga Chlamydomonas reinhardtii (Nagel et al. 2003). On absorbing blue light at an absorption peak of 480 nm, ChR2 undergoes a conformational change from the all-trans-retinal chromophore complex to 13-cis-retinal (Bamann et al. 2008). This switch causes a subsequent conformational change in the channel protein to open the pore, allowing various cations—such as H+, Na+, K+, and Ca2+—to flow (Nagel et al. 2003; Bamann et al. 2008). ChR2 has several features that make it particularly attractive as a neuroscience probe to depolarize neurons. First, the channel can be activated very rapidly and closes quickly upon light offset. 13-cis-retinal relaxes back to the all-trans form within milliseconds, closing the pore and stopping the flow of ions into or out of the cell. Therefore, single action potentials can be generated with a brief pulse of blue light, without any accompanying inappropriate effects of stimulation. Second, retinal is already present in most vertebrate cells in the form of vitamin A, which allows the ChR2 apoprotein to become a light-sensitive holoprotein. Therefore, the extraretinal is not needed when ChR2 is used in vertebrate neural systems, although exogenous application is necessary for invertebrate systems. Finally, as a genetically encoded protein, ChR2 permits cell-specific targeting with defined promoter and enhancer elements. Taken together, these properties of ChR2 allow researchers to stimulate particular neurons of interest with millisecond-level temporal resolution (Boyden et al. 2005; Li et al. 2005).
Recently, various opsins have been developed that can help us to analyze brain functions effectively. The red-shifted opsin VChR1, for example, is useful when an additional wavelength of light is required for excitation, while step-function opsins (SFOs) can inject a stable step current to allow a positive shift in membrane potential for up to 30–60 s with a single brief pulse, after which channel closure can be triggered by a brief exposure to yellow light.
3.1.2 Probes for Inhibiting Neurons
NpHR was the first molecule shown to inhibit neural activity (Zhang et al. 2007a, b; Han and Boyden 2007) and is naturally expressed in the halobacterium Natronomonas pharaonis (Schobert and Lanyi 1982; Bamberg et al. 1993). NpHR is a light-driven pump, which actively pumps Cl ions into cells in response to yellow light at the peak absorption wavelength of 570 nm. Like ChR2, NpHR utilizes retinal as its chromophore and therefore can also be used in vertebrate systems without extra cofactors. Substantial mutagenesis was required to achieve high levels of expression in neurons. Enhanced halorhodopsin (eNpHR), a second-generation NpHR, possesses an endoplasmic reticulum export signal and thus displays improved translocation to the plasma membrane (Zhao et al. 2008; Gradinaru et al. 2008). Third-generation constructs (eNpHR3.0) have been generated to improve photocurrent increasing in membrane hyperpolarization over eNpHR, with additional membrane-trafficking sequences (Gradinaru et al. 2010). Other proteins related to bacteriorhodopsins have been discovered recently and can be used to inhibit neural activity in response to light. Unlike NpHR, these proteins function as light-driven proton pumps. Archaerhodopsin-3 (Arch) proteins are derived from Halorubrum sodomense and allow near-100 % silencing of neurons in vivo in response to yellow light, with an efficiency comparable to that of eNpHR3.0 (Chow et al. 2010). Two other bacterial rhodopsins, Mac proteins from Leptosphaeria maculans and bacteriorhodopsin (BR) from Halobacterium salinarum (and its enhanced, second-generation derivative, eBR), allow silencing of neurons in response to blue-green light (Chow et al. 2010; Gradinaru et al. 2010). Therefore, hyperpolarizing optogenetic tools now exist that respond to blue– green and yellow light, allowing for combinatorial dissection of two neural subtypes in the same preparation. High-throughput genomic screens should reveal additional channels and thereby increase the diversity of inhibitory optogenetic tools for future use.
Interestingly, in a recent set of experiments, a proton-pumping archeorhodopsin was shown to allow high-speed imaging of individual action potentials. When a single amino acid residue was mutated, the resulting structure prevented proton flow across the membrane, allowing the channel to be used solely as an indicator. This genetically encoded voltage indicator exhibited an approximately tenfold improvement in sensitivity and speed over existing protein-based voltage indicators, with a roughly linear twofold increase in brightness between −150 and +150 mV and a sub-millisecond response time (Kralj et al. 2011). Therefore, optogenetics allows not only selective control of neural circuitry but also readout of its signals.
3.1.3 Probes for Manipulation of Intracellular Signaling
Neurons are also modulated by intracellular signaling events, initiated by cell surface receptors that culminate in a change in neuronal electrical activity instead of electrical signals through ion channels, as well as changes in secondary messenger pathways leading to gene expression and downstream protein cascades. Because rhodopsins are members of the GPCR family, it is theoretically possible to design synthetic rhodopsin-GPCR chimeras that combine the light-responsive elements of rhodopsin with the machinery of biochemical signaling of specific GPCRs (Kramer et al. 2009; Karnik et al. 2003; Kim et al. 2005).
3.1.4 Optogenetic Gene Delivery Strategies
The commonest method of delivering optogenetic transgenes into the nervous system is to infect cells with a self-inactivating virus, typically a lentivirus or adeno-associated virus (AAV), that contains the transgene of interest driven by a short promoter or enhancer element (Luo et al. 2008). Cell-type-specific promoters that have been used to drive optogenetic transgenes include EF1a (strong, ubiquitous expression) (Tsai et al. 2009; Cardin et al. 2009), CAMKII-α (expression limited to excitatory neurons) (Boyden et al. 2005; Gradinaru et al. 2009), synapsin I (limited to neurons) (Zhang et al. 2007a, b), and GFAP (limited to astrocytes) (Gradinaru et al. 2009). Several other promoters are useful for targeting to specific cell types in the brain, such as the ppHcrt promoter that targets hypocretin-expressing neurons in the lateral hypothalamus (Adamantidis et al. 2007), the oxytocin promoter that targets oxytocin peptide hormone-releasing neurons of the hypothalamus (Knobloch et al. 2012), and the synthetic PRSx8 promoter, which targets noradrenergic and adrenergic neurons that express dopamine beta hydroxylase (Abbott et al. 2009a; Abbott et al. 2009b). In utero electroporation is also useful for introducing optogenetic transgenes at specific developmental stages. Transgenes can be delivered to specific cortical layers of the brain by electroporating mice at embryonic day E12.5 (layers V and VI), E13.5 (layer IV), or E15.5 (layers II and III) (Zhang et al. 2010). Several studies have used this approach to deliver ChR2 to specific cortical layers for subsequent photostimulation when the mice reach adulthood (Gradinaru et al. 2007; Hull et al. 2009; Adesnik and Scanziani 2010).
Viral, transgenic, and in utero electroporation strategies can be combined to overcome the weak transcriptional activity of most endogenous promoters (Zhang et al. 2010). AAV vectors expressing Cre-dependent transgene cassettes under the control of strong, ubiquitous promoters such as EF1a have been developed which allow us to utilize numerous transgenic mouse lines from individual labs, such as GENSAT (Gong et al. 2007) and the Allen Institute for Brain Science (Madisen et al. 2010), that express Cre recombinase in specific cell types. Indeed, many optogenetic studies have capitalized on these excellent systems (Tsai et al. 2009; Cardin et al. 2009). Additionally, it is possible to specify the expression of optogenetic transgenes using anatomical-based cell targeting (Gradinaru et al. 2010). Rabies viral vectors or proteins such as wheat germ agglutinin or tetanus toxin fragment C help the gene of interest or Cre recombinase to be transported anterogradely and retrogradely (Luo et al. 2008; Gradinaru et al. 2010; Zhang et al. 2010; Osakada et al. 2011; Wall et al. 2010). Thus, it is possible to restrict the expression of optogenetic transgenes in a particular neural circuit even if its cells do not express unique genetic regulatory elements. Finally, cellular indicators of functional activity, including the immediate early genes zif268 (egr1), c-fos, and arc, help us to regulate the activated network relating to particular behaviors (Covington et al. 2010; Liu et al. 2012). The combination of these technologies provides scientists with multiple strategies for expressing optogenetic probes in specific neural networks, and the technologies also offer greater flexibility to express the probes in various model animals, such as rats and primates.
3.2 Optical Aspects
To manipulate neural activities in a specific population of cells, it is necessary to deliver light properly. A light delivery system for in vitro cultured neurons or brain slices can be established using conventional light sources such as halogen/xenon arc lamps, light-emitting diodes (LEDs), and lasers, all of which can be directly built into the light path of a microscope (Zhang et al. 2010). In vivo light delivery, however, is more challenging because surgical implants of optical fibers are required to place them stereotaxically around targeted regions. Light needs to be delivered very close to target regions because brain tissue scatters light exponentially, with only 10 % of light intensity remaining at a distance of 500 μm from the light source (Adamantidis et al. 2007; Aravanis et al. 2007). Regarding freely moving animals, the delivery device must be light enough to be carried easily and should not interfere with natural behaviors. At present, the commonest method for delivering light in vivo is to implant a guide cannula to place a fiber optical cable (Fig. 8.1) (Aravanis et al. 2007). This procedure allows cells in deep-brain structures to be targeted and is applicable for mice with up to 300-μm diameter fibers and for rats with up to 400 μm fibers. Optical fibers are typically connected to a laser diode, although it is also possible to connect them to an LED. To modulate the neural activities of superficial cortical neurons, small LEDs can be mounted above a glass over a cranial window (Gradinaru et al. 2007; Huber et al. 2008, Wentz et al. 2011). Recently, an array of optical fibers or an electrocorticogram (ECoG) equipped with LEDs has been established to modulate simultaneously multiple sites in the brain (Bernstein and Boyden 2011; Sawadsaringkarn et al. 2012). These technologies help us to analyze information processing throughout the brain.
The two-photon microscope is one of the most widespread tools to observe neurons of interest, because it allows deeper areas of the brain to be visualized than does conventional microscopy. However, its adoption for optogenetics presents technical difficulties. The typical diffraction-limited focal volumes required to achieve the conditions of multiphoton excitation are about 1/1,000 of the volume of a typical cell body. This strategy enables us to activate only a tiny fraction of the available channels on the cell’s plasma membrane, but would yield a degree of stimulation insufficient to depolarize most cells to the action potential threshold. Trying to increase this focal volume by reducing the effective numerical aperture of the optical system fails to overcome the problem because the lateral and axial dimensions of the excited focal spot are coupled: a tenfold decrease in lateral resolution corresponds to a 100-fold loss of axial resolution. A simple solution to this resolution versus effectiveness tradeoff is to move the stimulation spot very rapidly across the cell membrane, integrating the cumulative effect of many locations (Rickgauer and Tank 2009) for two-photon ChR2 neural stimulation with a 30 ms spiral scan.
To identify functional connections in the real brain, the cell-type- and site-specific causal controls provided by optogenetics and fMRI in mice have been combined to test the linearity of BOLD signals driven by locally induced excitatory activity (Lee et al. 2010; Kahn et al. 2011; Desai et al. 2011). This strategy helps us to estimate how linear the response to sensory stimuli is, which is essential for the design and interpretation of in vivo fMRI experiments.
Additional information about optogenetics, including technical details about genes and light delivery systems, can be found in other excellent reviews and protocols (Zhang et al. 2006; Cardin et al. 2010). More information about available optogenetic transgenes can be found at the Optogenetics Resource Center Web page (http://www.stanford.edu/group/dlab/optogenetics/) maintained by the laboratory of Karl Deisseroth. Details of tools for gene and light delivery are available on the Web page (http://syntheticneurobiology.org/protocols) maintained by the laboratory of Ed Boyden.
4 Conclusion
Although optogenetics has been established for only 6 years, the number of reports exploiting these tools is increasing exponentially to provide further answers to anatomical, physiological, and pathological issues. These tools allow us to address the complexity of neural circuits, for not only in vitro but also in vivo studies. However, there remains much scope for further refinement that would allow us, for example, to stimulate multiple cell types at the same time by introducing various mutations in a particular domain, as is the case with the multiple available versions of green fluorescent protein. It will also be desirable to increase the conductance of various channels so that less light stimulation is necessary, to avoid the expected side effects of heat on the cell. Finally, it should also be possible to record patterns or sequences of neural firing and then use these recordings to mimic the recorded neural activities with light pulses. Regarding the therapeutic use of optogenetics, it will be necessary to develop safe and reversible gene delivery strategies and light delivery devices that are adaptable to human patients. The use of light to study the brain has proved to be a remarkably fruitful strategy, and indeed optogenetics has given us a green light for the future.
References
Abbott SB, Stornetta RL, Fortuna MG et al (2009a) Photostimulation of retrotrapezoid nucleus phox2b-expressing neurons in vivo produces long-lasting activation of breathing in rats. J Neurosci 29:5806–5819
Abbott SB, Stornetta RL, Socolovsky CS et al (2009b) Photostimulation of channelrhodopsin-2 expressing ventrolateral medullary neurons increases sympathetic nerve activity and blood pressure in rats. J Physiol 587:5613–5631
Adamantidis AR, Zhang F, Aravanis AM et al (2007) Neural substrates of awakening probed with optogenetic control of hypocretin neurons. Nature 450:420–424
Adesnik H, Scanziani M (2010) Lateral competition for cortical space by layer-specific horizontal circuits. Nature 464:1155–1160
Airan RD, Thompson KR, Fenno LE et al (2009) Temporally precise in vivo control of intracellular signalling. Nature 458:1025–1029
Alilain WJ, Li X, Horn KP et al (2008) Light-induced rescue of breathing after spinal cord injury. J Neurosci 28:11862–11870
Aravanis AM, Wang LP, Zhang F et al (2007) An optical neural interface: in vivo control of rodent motor cortex with integrated fiberoptic and optogenetic technology. J Neural Eng 4:S143–56
Bamann C, Kirsch T, Nagel G et al (2008) Spectral characteristics of the photocycle of channelrhodopsin-2 and its implication for channel function. J Mol Biol 375:686–694
Bamberg E, Tittor J, Oesterhelt D (1993) Light-driven proton or chloride pumping by halorhodopsin. Proc Natl Acad Sci USA 90:639–643
Berndt A, Yizhar O, Gunaydin LA et al (2009) Bi-stable neural state switches. Nat Neurosci 12:229–234
Bernstein JG, Han X, Henninger MA et al (2008) Prosthetic systems for therapeutic optical activation and silencing of genetically-targeted neurons. Proc Soc Photo Opt Instrum Eng 6854:68540H
Bernstein JG, Boyden ES (2011) Optogenetic tools for analyzing the neural circuits of behavior. Trends Cogn Sci 15:592–600
Bi A, Cui J, Ma YP et al (2006) Ectopic expression of a microbial-type rhodopsin restores visual responses in mice with photoreceptor degeneration. Neuron 50:23–33
Boyden ES, Zhang F, Bamberg E et al (2005) Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci 8:1263–1268
Busskamp V, Duebel J, Balya D et al (2010) Genetic reactivation of cone photoreceptors restores visual responses in retinitis pigmentosa. Science 329:413–417
Cardin JA, Carlen M, Meletis K et al (2010) Targeted optogenetic stimulation and recording of neurons in vivo using cell-type-specific expression of Channelrhodopsin-2. Nat Protoc 5:247–254
Cardin JA, Carlen M, Meletis K et al (2009) Driving fast-spiking cells induces gamma rhythm and controls sensory responses. Nature 459:663–667
Carter ME, Adamantidis A, Ohtsu H et al (2009) Sleep homeostasis modulates hypocretin-mediated sleep-to-wake transitions. J Neurosci 29:10939–10949
Carter ME, Yizhar O, Chikahisa S et al (2010) Tuning arousal with optogenetic modulation of locus coeruleus neurons. Nat Neurosci 13:1526–1533
Carter M, Shieh JC (2010) Guide to research techniques in neuroscience. Elsevier/Academic, Amsterdam/Boston
Cepko C (2010) Neuroscience. Seeing the light of day. Science 329:403–404
Chow BY, Han X, Dobry AS et al (2010) High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature 463:98–102
Ciocchi S, Herry C, Grenier F et al (2010) Encoding of conditioned fear in central amygdala inhibitory circuits. Nature 468:277–282
Covington HE 3rd, Lobo MK, Maze I et al (2010) Antidepressant effect of optogenetic stimulation of the medial prefrontal cortex. J Neurosci 30:16082–16090
Desai M, Kahn I, Knoblich U et al (2011) Mapping brain networks in awake mice using combined optical neural control and fMRI. J Neurophysiol 105:1393–1405
Gong S, Doughty M, Harbaugh CR et al (2007) Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs. J Neurosci 27:9817–9823
Gradinaru V, Mogri M, Thompson KR et al (2009) Optical deconstruction of parkinsonian neural circuitry. Science 324:354–359
Gradinaru V, Thompson KR, Deisseroth K (2008) eNpHR: a Natronomonas halorhodopsin enhanced for optogenetic applications. Brain Cell Biol 36:129–139
Gradinaru V, Thompson KR, Zhang F et al (2007) Targeting and readout strategies for fast optical neural control in vitro and in vivo. J Neurosci 27:14231–14238
Gradinaru V, Zhang F, Ramakrishnan C et al (2010) Molecular and cellular approaches for diversifying and extending optogenetics. Cell 141:154–165
Gunaydin LA, Yizhar O, Berndt A et al (2010) Ultrafast optogenetic control. Nat Neurosci 13:387–392
Han X, Boyden ES (2007) Multiple-color optical activation, silencing, and desynchronization of neural activity, with single-spike temporal resolution. PLoS One 2:e299
Haubensak W, Kunwar PS, Cai H et al (2010) Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature 468:270–276
Huber D, Petreanu L, Ghitani N et al (2008) Sparse optical microstimulation in barrel cortex drives learned behaviour in freely moving mice. Nature 451:61–64
Hull C, Adesnik H, Scanziani M (2009) Neocortical disynaptic inhibition requires somatodendritic integration in interneurons. J Neurosci 29:8991–8995
Kahn I, Desai M, Knoblich U et al (2011) Characterization of the functional MRI response temporal linearity via optical control of neocortical pyramidal neurons. J Neurosci 31:15086–15091
Karnik SS, Gogonea C, Patil S et al (2003) Activation of G-protein-coupled receptors: a common molecular mechanism. Trends Endocrinol Metab 14:431–437
Kim JM, Hwa J, Garriga P et al (2005) Light-driven activation of beta 2-adrenergic receptor signaling by a chimeric rhodopsin containing the beta 2-adrenergic receptor cytoplasmic loops. Biochemistry 44:2284–2292
Kralj JM, Douglass AD, Hochbaum DR et al (2011) Optical recording of action potentials in mammalian neurons using a microbial rhodopsin. Nat Methods 9:90–95
Kramer RH, Fortin DL, Trauner D (2009) New photochemical tools for controlling neuronal activity. Curr Opin Neurobiol 19:544–552
Kravitz AV, Freeze BS, Parker PR et al (2010) Regulation of parkinsonian motor behaviours by optogenetic control of basal ganglia circuitry. Nature 466:622–626
Lagali PS, Balya D, Awatramani GB et al (2008) Light-activated channels targeted to ON bipolar cells restore visual function in retinal degeneration. Nat Neurosci 11:667–675
Lee JH, Durand R, Gradinaru V et al (2010) Global and local fMRI signals driven by neurons defined optogenetically by type and wiring. Nature 465:788–792
Li X, Gutierrez DV, Hanson MG et al (2005) Fast noninvasive activation and inhibition of neural and network activity by vertebrate rhodopsin and green algae channelrhodopsin. Proc Natl Acad Sci USA 102:17816–17821
Lin D, Boyle MP, Dollar P et al (2011) Functional identification of an aggression locus in the mouse hypothalamus. Nature 470:221–226
Lin JY, Lin MZ, Steinbach P et al (2009) Characterization of engineered channelrhodopsin variants with improved properties and kinetics. Biophys J 96:1803–1814
Luo L, Callaway EM, Svoboda K (2008) Genetic dissection of neural circuits. Neuron 57:634–660
Madisen L, Zwingman TA, Sunkin SM et al (2010) A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 13:133–140
Nagel G, Szellas T, Huhn W et al (2003) Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc Natl Acad Sci USA 100:13940–13945
Oh E, Maejima T, Liu C et al (2010) Substitution of 5-HT1A receptor signaling by a light-activated G protein-coupled receptor. J Biol Chem 285:30825–30836
Osakada F, Mori T, Cetin AH et al (2011) New rabies virus variants for monitoring and manipulating activity and gene expression in defined neural circuits. Neuron 71:617–631
Rickgauer JP, Tank DW (2009) Two-photon excitation of channelrhodopsin-2 at saturation. Proc Natl Acad Sci USA 106:15025–15030
Sawadsaringkarn Y, Kimura H, Maezawa Y et al (2012) CMOS on-chip optoelectronic neural interface device with integrated light source for optogenetics. J Phys Conf Ser 352:012004
Schobert B, Lanyi JK (1982) Halorhodopsin is a light-driven chloride pump. J Biol Chem 257:10306–10313
Sohal VS, Zhang F, Yizhar O et al (2009) Parvalbumin neurons and gamma rhythms enhance cortical circuit performance. Nature 459:698–702
Stuber GD (2010) Dissecting the neural circuitry of addiction and psychiatric disease with optogenetics. Neuropsychopharmacology 35:341–342
Stuber GD, Hnasko TS, Britt JP et al (2010) Dopaminergic terminals in the nucleus accumbens but not the dorsal striatum corelease glutamate. J Neurosci 30:8229–8233
Tecuapetla F, Patel JC, Xenias H et al (2010) Glutamatergic signaling by mesolimbic dopamine neurons in the nucleus accumbens. J Neurosci 30:7105–7110
Tomita H, Sugano E, Fukazawa Y et al (2009) Visual properties of transgenic rats harboring the channelrhodopsin-2 gene regulated by the thy-1.2 promoter. PLoS One 4:e7679
Tomita H, Sugano E, Isago H et al (2010) Channelrhodopsin-2 gene transduced into retinal ganglion cells restores functional vision in genetically blind rats. Exp Eye Res 90:429–436
Tonnesen J, Sorensen AT, Deisseroth K et al (2009) Optogenetic control of epileptiform activity. Proc Natl Acad Sci USA 106:12162–12167
Tsai HC, Zhang F, Adamantidis A et al (2009) Phasic firing in dopaminergic neurons is sufficient for behavioral conditioning. Science 324:1080–1084
Wall NR, Wickersham IR, Cetin A et al (2010) Monosynaptic circuit tracing in vivo through Cre-dependent targeting and complementation of modified rabies virus. Proc Natl Acad Sci USA 107:21848–21853
Wentz CT, Bernstein JG, Monahan P et al (2011) A wirelessly powered and controlled device for optical neural control of freely-behaving animals. J Neural Eng 8:046021
Zhang F, Aravanis AM, Adamantidis A et al (2007a) Circuit-breakers: optical technologies for probing neural signals and systems. Nat Rev Neurosci 8:577–581
Zhang F, Gradinaru V, Adamantidis AR et al (2010) Optogenetic interrogation of neural circuits: technology for probing mammalian brain structures. Nat Protoc 5:439–456
Zhang F, Prigge M, Beyriere F et al (2008) Red-shifted optogenetic excitation: a tool for fast neural control derived from Volvox carteri. Nat Neurosci 11:631–633
Zhang F, Wang LP, Boyden ES et al (2006) Channelrhodopsin-2 and optical control of excitable cells. Nat Methods 3:785–792
Zhang F, Wang LP, Brauner M et al (2007b) Multimodal fast optical interrogation of neural circuitry. Nature 446:633–639
Zhang Y, Ivanova E, Bi A et al (2009) Ectopic expression of multiple microbial rhodopsins restores ON and OFF light responses in retinas with photoreceptor degeneration. J Neurosci 29:9186–9196
Zhao S, Cunha C, Zhang F et al (2008) Improved expression of halorhodopsin for light-induced silencing of neuronal activity. Brain Cell Biol 36:141–154
Acknowledgements
We thank all members of our laboratories for useful discussion and support. Fig. 8.1 was adopted from the front page of BioGARAGE, published on March 2012 by Leave a Nest Co., Ltd. The images in both figures were designed by Science Graphics Co., Ltd.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer
About this chapter
Cite this chapter
Cetin, A., Komai, S. (2013). Controlling Behavior Using Light to Excite and Silence Neuronal Activity. In: Ogawa, H., Oka, K. (eds) Methods in Neuroethological Research. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54331-2_8
Download citation
DOI: https://doi.org/10.1007/978-4-431-54331-2_8
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54330-5
Online ISBN: 978-4-431-54331-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)