Abstract
Mitosis, the process by which one cell divides into two genetically identical daughter cells, is the most basic process for the development and proliferation of living organisms. In eukaryotes, mitosis involves the transient organization of a sophisticated molecular machine, the bipolar spindle that orchestrates the segregation of the genetic material to the daughter cells. The spindle is a microtubule (MT)-based apparatus whose assembly and function rely on the fine modulation of MT intrinsic dynamic properties and on their spatial and temporal organization. In this chapter, we will focus on the mechanisms of spindle assembly and dynamics. We will discuss some current questions in the field and review the consequences of defective MT function in mitotic cells for human health.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1.1 Introduction
Almost 200 years ago, Schwann’s theory stated that all living things are composed of cells. Later in 1857, Rudolf Virchow postulated that “Omnis cellula e cellula,” the generation of new cells from a pre-existing cell, involves a specific process called mitosis.
After the very first division of the fertilized egg, a complex developmental program involving billions of cell divisions takes place to generate the tissues and organs that constitute a full organism. Cell division is also essential in the adult organism. It occurs permanently throughout the life of a human being, playing an essential role in the maintenance and renewal of its tissues and organs.
Flemming in the late nineteenth century described the two main elements in animal cells undergoing mitosis, called “thin filaments” and “chromo elements” (Paweletz 2001): these are microtubules (MTs) and chromosomes. Almost 150 years after this initial description, we have now attained a reasonably good understanding of the mechanism that underlies cell division. In this chapter, we will focus on the general principles underlying spindle assembly and function from an MT centric perspective. For simplicity, we will focus on mitosis in higher eukaryotes, but it is worth keeping in mind that many of the pathways and mechanisms are similar in meiosis, a gamete-specific cell division process.
1.2 Microtubule Basic Properties and Mitosis
Cell division involves the full reorganization of the interphase MT network to assemble the mitotic spindle, a dynamic molecular machine that provides the forces and support for chromosome segregation (Inoue and Sato 1967). To address the mechanism leading to the assembly of the spindle, it is therefore essential to start by revising some essential MT properties.
1.2.1 MT Basics
MTs are hollow tubes of 25 nm in diameter formed by lateral interactions of 13 protofilaments. Each protofilament in turn is formed by head to tail interactions of α-/β-tubulin heterodimers, two closely related tubulin isoforms that bind GTP. This molecular organization defines MT polarity: only α-tubulin subunits are exposed at one extremity called the minus end and only β-tubulin subunits are exposed at the other extremity called the plus end (Fig. 1.1). The resulting polarity along the MT lattice is read by molecular motors that move directionally along the filament either toward the plus or the minus end.
MT basics. Microtubules (MTs) are composed by tubulin heterodimers of α- and β-tubulin. This composition defines a polarity to the MTs; α-tubulin is exposed at the minus end and β-tubulin at the plus end. They are intrinsically dynamic and undergo phase of growth and shrinkage. The switch from a growth state to a shrinking state is called catastrophe, and the opposite switch is the MT rescue
In vitro MTs form spontaneously above a certain concentration threshold of α-/β-tubulin heterodimers and exhibit dynamic properties. In the presence of GTP, MTs grow and shrink, stochastically alternating between these two phases by undergoing catastrophes when switching from the growing state to the shrinking state and rescues for the opposite switch (Wade 2009) (Fig. 1.1). These unique properties define MT dynamic instability as named by Mitchison and Kirschner when they first described it in 1984 (Mitchison and Kirschner 1984). Interestingly, they demonstrated that the dynamic properties and average length of any MT population could be described using four parameters: the velocities of growth and shrinkage, and the catastrophe and rescue frequencies (Mitchison and Kirschner 1984). All these events occur at the MT plus ends, while MT minus ends are more stable. MT polarity therefore is also related to the differential dynamic behavior of the two MT extremities.
MT dynamic properties are intimately related to GTP hydrolysis. Indeed, MTs formed with a slowly hydrolysable GTP analogue GMCPP are remarkably stable in vitro. GTP hydrolysis occurs only at the β-tubulin subunit after incorporation of the α-/β-tubulin heterodimer into the growing MT plus end. MTs are therefore mainly composed by GDP-tubulin with a so-called GTP cap at their plus ends. By promoting a change in conformation of the tubulin dimer, GTP hydrolysis generates a tension within the polymer. When GTP hydrolysis occurs faster than the rate of tubulin incorporation at the plus end, this tension is released by fast depolymerization (Alushin et al. 2014).
MTs therefore have two essential intrinsic properties: they are polarized and they show dynamic instability. These properties are key for most of the events leading to the assembly of the mitotic spindle.
1.2.2 Microtubule Organization in Mitosis
By regulating MT dynamic properties, the cell controls the organization of its MT network. This is key for mitosis when in a timely manner the interphase MT network disassembles to build the mitotic spindle, the essential machinery to segregate the chromosomes. This reorganization is in fact dynamic and occurs through an ordered sequence of phases (Fig. 1.2).
Mitotic phases. In prophase, chromosomes are condensed, and centrosomes have been duplicated and nucleate highly dynamics MTs. Nuclear envelope breaks down. In prometaphase, MTs become specialized. Some of them interact with the cell cortex and are called astral MTs; interpolar MTs form the main part of the bipolar spindle and MTs interacting with the kinetochores become organized into K-fibers. In metaphase, the bipolar spindle is mature and the chromosomes are aligned in the center at the metaphase plate, all of them connected to the two opposite poles through sister K-fibers. In anaphase A, K-fibers shorten and pull the sister chromatids apart. In anaphase B, a new MT-based structure, called the central spindle, is built between the two chromosome masses and helps in separating them. In telophase, nuclear envelope forms again and the midbody will define the site of abscission between the two daughter cells
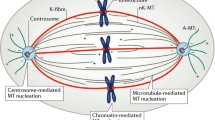
In prophase, the interphase MT network disassembles and the duplicated centrosomes separate to opposite sides of the nucleus along the nuclear envelope. As the centrosome MT nucleation activity increases, they form two asters of highly dynamic MTs. The chromosomes condense inside the nucleus. Prometaphase starts after nuclear envelope breakdown. MTs establish connections with the chromosomes and some of them get stabilized at specific sites on the centromeres, the kinetochores. As these interactions get established, MTs start to organize into a spindle-shaped apparatus (Fig. 1.2). Metaphase is characterized by a mature bipolar spindle in which MT minus ends are focused at the two spindle poles and MT plus ends interdigitate at the center where chromosomes are aligned on the metaphase plate. Once all the chromosomes are correctly attached to both spindle poles, anaphase A starts, and the sister chromatids are pulled apart toward the two opposite poles. In anaphase B, a novel MT-based antiparallel array, the central spindle, assembles in between the separated chromosome masses and promotes their further separation by antiparallel MT sliding. In telophase, two nuclei form and the daughter cells separate completely through cytokinesis and abscission before establishing their interphase MT network (Fig. 1.2).
All the major events driving cell division rely on timely and spatially controlled changes in MT dynamic properties and organization. Based on the MT dynamic instability property, Kirschner and Mitchison (1986) proposed the “search and capture” model for spindle assembly. It postulated that highly dynamic MTs emanating from the separated centrosomes grow and shrink, exploring the cytoplasm until some of them get captured and stabilized by the kinetochores. Since MTs emanate from two centrosomes and each chromosome has two kinetochores, this stochastic process should naturally drive bipolar spindle assembly. Although we know now that the mechanism driving spindle assembly is more complex, the basic principle of this model is still valid: a large number of dynamic MTs are required to efficiently explore the cellular space and attach the chromosomes. However, these MTs may or may not be generated by the centrosomes. In fact, centrosomal MTs alone do not form a functional bipolar spindle (Gruss et al. 2002) and modeling approaches have suggested that a process involving only centrosomal MTs could not account for the rapid attachment of all chromosomes in human cells in the observed time (Wollman et al. 2005). Indeed, we know now that in addition to the centrosomes, other pathways are set up in mitotic cells to promote MT nucleation and assembly in an acentrosomal manner. We will see later in this chapter that these mitotic pathways are essential for spindle assembly. We will also discuss whether their role is merely to provide an efficient “search and capture” mechanism or whether it may go beyond.
1.3 General Principles in Mitotic Spindle Assembly
Since the bipolar spindle is constituted by transient interactions between highly dynamic MTs, it is itself highly dynamic in nature, a property that underlies its self-organization and self-correction properties. In the most extreme case, mitotic cells can rebuild a spindle after the full depolymerization of MTs by cold or drug treatments (Tulu et al. 2003). Under physiological conditions, spindle dynamics enables the correction of erroneous MT-chromosome attachments thereby preventing defects in chromosome segregation or mitotic slippage. However, and despite its highly dynamic nature, the spindle also has to provide mechanical support for the forces required to move and segregate the chromosomes.
In this chapter, we will focus on two general mechanisms essential to understand how MTs organize the bipolar spindle: First, we will focus on the mechanism by which cells promote the assembly of a large number of highly dynamic MTs by upregulating MT nucleation. Second, we will focus on the global and local regulation of MT dynamics.
1.3.1 Control of MT Nucleation in M Phase
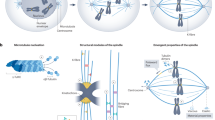
In cells, the tubulin concentration does not reach the critical threshold for spontaneous MT assembly. In fact, cells define and control where and when MT assembly occurs by using a specific mechanism driving MT nucleation. In eukaryotic cells, this involves a major MT nucleation complex called the γ-TuRC (γ-tubulin ring complex). This protein complex is composed by multiple copies of γ-tubulin and five additional proteins called GCPs (γ-tubulin complex proteins) that organize a ringlike structure postulated to act as a template for tubulin dimer addition and MT polymerization (Teixido-Travesa et al. 2010; Kollman et al. 2011). A number of additional proteins associate with the γ-TuRC. One of them, NEDD1 (also called GCP-WD), functions as an adaptor or targeting factor for the γ-TuRC (Haren et al. 2006; Luders et al. 2006; Zhu et al. 2008; Zhang et al. 2009; Sdelci et al. 2012; Pinyol et al. 2013). For a detailed description of this complex and its role in MT nucleation, we refer the reader to excellent reviews (Kollman et al. 2011; Teixido-Travesa et al. 2012) (also see the Chap. 4 by Sánchez-Huertas, Freixo, and Lüders).
In mitosis, MT nucleation increases through different mechanisms that all involve the γ-TuRC (Moudjou et al. 1996; Teixido-Travesa et al. 2012). This in turn defines different MT assembly pathways.
1.3.1.1 Centrosome Maturation and MT Nucleation
Centrosomes are MT-based organelles playing a variety of functions in the cell (Bettencourt-Dias and Glover 2007). In higher eukaryotes, the cell has one centrosome composed by a pair of centrioles surrounded by pericentriolar material (PCM) (see the Chap. 3 by Comartin and Pelletier for a detailed description). In interphase, the centrosome is the main MT organizing center (MTOC), promoting MT nucleation and maintaining in focus most MT minus ends. In cycling cells, the centrosome duplicates during interphase and before mitosis onset, the duplicated centrosomes undergo maturation characterized by the active recruitment of PCM components, in particular MT nucleation factors such as γ-tubulin as part of the γ-TuRC (Khodjakov and Rieder 1999; Piehl et al. 2004). Centrosome maturation therefore leads to a dramatic increase of the MT nucleation activity. This promotes the formation of two asters of dynamic MTs that get positioned on opposite sides of the nucleus through the active separation of the centrosomes before the nuclear envelope breaks down. However, centrosomes are not essential for bipolar spindle assembly (Bettencourt-Dias 2013). Other pathways are specifically set up in dividing cells and activate MT nucleation in a centrosome-independent manner.
1.3.1.2 Chromosome-Dependent MT Assembly
The last 20 years have provided compelling evidence for the existence of a specific MT nucleation and assembly pathway triggered around the chromosomes in dividing cells (Karsenti et al. 1984; Heald et al. 1996). The underlying mechanism involves the activity of Ran, a small GTPase that is essential for nucleocytoplasmic transport in interphase (Clarke and Zhang 2008). In dividing cells, the association of its guanosine exchange factor (GEF) RCC1 with the chromatin results in the formation of a GTP-bound Ran (RanGTP) gradient centered around the chromosomes (Carazo-Salas et al. 1999; Kalab et al. 1999, 2002; Ohba et al. 1999; Zhang et al. 1999; Carazo-Salas et al. 2001; Gruss et al. 2001). Following the same basic mechanism as in interphase, RanGTP induces the release of NLS (nuclear localization signal)-containing proteins from karyopherins. Some of these proteins perform essential functions in spindle assembly. A number of recent reviews cover the principles of the RanGTP gradient in mitosis as well as the current knowledge on the identity and function of RanGTP-regulated proteins (Karsenti and Vernos 2001; Meunier and Vernos 2012).
The mechanism by which RanGTP upregulates de novo MT nucleation in the vicinity of the chromosomes is however not fully understood yet. As for all the other pathways, it requires γ-TuRC activity (Groen et al. 2004; Luders et al. 2006), but it also requires the RanGTP-regulated protein TPX2 (targeting protein for Xklp2) (Wittmann et al. 2000; Gruss et al. 2001, 2002), a specific activator of the Aurora A kinase (Bayliss et al. 2003; Eyers et al. 2003; Tsai et al. 2003). Recently, it was shown that the specific phosphorylation of the γ-TuRC-associated protein NEDD1 by the mitotic kinase Aurora A is essential for this pathway (Pinyol et al. 2013). TPX2 interacts with the mitotic kinase Aurora A and activates it in a RanGTP-dependent manner, promoting NEDD1 phosphorylation and thereby a mechanism for the activation of MT nucleation by RanGTP (Scrofani et al. 2015).
Although many data support a mechanism for RanGTP-/chromosomal-dependent MT nucleation without a predefined site, some reports on the localization of γ-TuRCs at the kinetochores suggest that MTs may also be nucleated at the kinetochore (Torosantucci et al. 2008; Mishra et al. 2010). However, it is still not clear whether such a direct kinetochore-dependent MT nucleation occurs or whether MTs are stabilized in this specific region of the chromosomes (Tulu et al. 2006; Maresca et al. 2009; Needleman et al. 2010) by the chromosomal passenger complex, located at the kinetochores (Sampath et al. 2004; Tseng et al. 2010). The putative nucleation of MTs at the kinetochore would however result in their “reversed” polarity with their minus end at the kinetochore and their plus end extending toward the spindle poles. This orientation has never been observed in animal cells (Euteneuer and McIntosh 1981; Euteneuer et al. 1983; Rieder 2005; Kitamura et al. 2010).
1.3.1.3 MT Amplification
In addition to the activation of MT nucleation at the centrosome and around the chromosomes, the recruitment of γ-TuRCs on pre-existing MTs drives an amplification mechanism that increases MT polymer amounts (Goshima et al. 2007, 2008; Lawo et al. 2009). This pathway relies on the augmin complex, constituted by eight proteins in humans (Lawo et al. 2009). The augmin complex binds the lattice of a pre-existing MT recruiting the γ-TuRC and promotes the nucleation and elongation of a new MT (Kamasaki et al. 2013). This results in MT branching and drives the efficient amplification of the whole MT mass during mitosis (Petry et al. 2013). Augmin-dependent MTs are then sorted toward the spindle poles (Lecland and Luders 2014). The augmin-dependent amplification pathway plays an important role during cell division. Indeed, the silencing of some of its components results in dramatic phenotypes (Uehara et al. 2009; Wainman et al. 2009; Petry et al. 2011). Recently, this pathway has been proposed to be intimately related with the chromosomal, RanGTP-dependent pathway of MT assembly (Petry et al. 2013).
Altogether, various mechanisms boost MT nucleation in the dividing cells actively promoting MT assembly. This activation does not occur simultaneously or at a single site; it starts at the centrosomes and then around the chromosomes and on pre-existing MTs. One common requirement is the γ-TuRC (Moudjou et al. 1996; Teixido-Travesa et al. 2012) and NEDD1 phosphorylation (Haren et al. 2006; Luders et al. 2006; Zhu et al. 2008; Zhang et al. 2009; Johmura et al. 2011; Gomez-Ferreria et al. 2012; Sdelci et al. 2012; Pinyol et al. 2013; Scrofani et al. 2015). This regulatory mechanism involves different mitotic kinases that target specific sites on NEDD1 but interestingly all in close proximity within the protein (Luders et al. 2006; Sdelci et al. 2012; Pinyol et al. 2013). However, the precise mechanisms by which these phosphorylation events participate in the activation of MT nucleation are still not understood.
1.3.2 Balancing High MT Dynamics
Spindle assembly requires abundant dynamic MTs, but their local stabilization is essential to generate a structure that is robust enough to provide support for chromosome movements yet flexible enough to correct erroneous interactions with the chromosomes to ensure their error-free segregation.
The global destabilization of the interphase MTs (turnover in the range of minutes to hours) is triggered by the cell cycle machinery through the activation of the Cdk1 kinase. However, the highly dynamic nature of the mitotic MTs (turnover in the range of seconds to a few minutes) is finely controlled by MT stabilizing and destabilizing activities. This general concept was validated in vitro. Indeed, mitotic MT dynamics could be mimicked in vitro by adding at a certain ratio two proteins with antagonizing MT stabilizing and destabilizing activities (Tournebize et al. 2000). These two proteins are XMAP215/chTOG and MCAK.
chTOG/XMAP215 is an MT-associated protein (MAP) conserved in all eukaryotes (Gard and Kirschner 1987; Vasquez et al. 1994); it binds very efficiently to MTs and promotes their assembly in vitro and in vivo (Brouhard et al. 2008). XMAP215 can bind along the whole MT lattice but has also been characterized as a plus-end binding protein, promoting MT polymerization. Indeed, MT stabilizing factors are MAPs acting through various mechanisms. Some of them as XMAP215 are found along the entire MT length, whereas others have specific plus-end-tracking properties (+TIPs). They localize to the MT growing ends, probably recognizing the GTP-bound state of tubulin, and regulate their dynamic behavior (Mimori-Kiyosue et al. 2000; Maiato et al. 2005; Kronja et al. 2009; Akhmanova and Steinmetz 2010; Maurer et al. 2012; Zanic et al. 2013; Alushin et al. 2014; Zhang et al. 2015). Finally, other MT stabilizing factors seem to function by protecting MT against depolymerization. At MT minus ends, the γ-TuRC or other minus-end binding complexes may stabilize MTs through an end-capping activity (Wiese and Zheng 2000; Goodwin and Vale 2010; Meunier and Vernos 2011; Jiang et al. 2014; Meunier et al. 2015).
Counteracting the action of MT stabilizing factors, three different types of activities promote MT destabilization: catastrophe factors, tubulin sequestering factors, and severing factors. MCAK is a major MT depolymerase belonging to the kinesin 13 family (constituted by KIF2A, KIF2B, and MCAK/KIF2C). In vitro, MCAK was shown to attach to the MT lattice through electrostatic interactions and diffuse along the MT, driving MT depolymerization both at MT plus and minus ends (Walczak et al. 1996; Desai et al. 1999; Hunter et al. 2003). The members of another class of depolymerizing kinesins in the kinesin 8 family (KIF18A and KIF18B) use a distinct mechanism to induce MT destabilization. They act by blocking the incorporation of new tubulin dimers at MT plus ends (Mayr et al. 2007; Du et al. 2010; Walczak et al. 2013). An additional way to regulate MT stability involves the protein Op18 (also called stathmin). Op18 binds free tubulin dimers and impairs their incorporation at the MT plus end (Belmont and Mitchison 1996; Cassimeris 2002; Gupta et al. 2013), leading to a decrease in MT growth and MT destabilization. Finally, a number of MT severing enzymes have been recently identified, affecting MT stability within the mitotic spindle (Sharp and Ross 2012).
Altogether, a large number of mechanisms control MT stabilization and destabilization. They are tightly regulated and coordinated in space and time during mitosis to ensure the correct assembly and function of the bipolar spindle. A spatial control on MT dynamics is provided by the chromosomes through the RanGTP pathway that favors MT stabilization by creating a gradient of MT stabilizing factors (Karsenti and Vernos 2001; Caudron et al. 2005; Clarke and Zhang 2008). The RanGTP gradient is moreover translated into a phosphorylation gradient through the TPX2-dependent activation of the Aurora A kinase, promoting MT stabilization around the chromosomes (Eyers et al. 2003; Tsai et al. 2003; Sardon et al. 2008). The assembly of the full mitotic spindle appears therefore to rely on overlapping regulatory gradients. The RanGTP gradient around the chromosomes favors MT nucleation and stabilization, and the Aurora A kinase acts on MT stabilization at the chromosomes and the centrosomes. A Plk1-dependent phosphorylation gradient around the centrosomes favors MT assembly and is essential for spindle positioning (Kiyomitsu and Cheeseman 2012). An Aurora B-dependent phosphorylation gradient centered at the kinetochores in the first phases of mitosis and at the central spindle in anaphase is involved in multiple functions, including MT dynamic regulation (Carmena et al. 2012). Altogether, these regulatory gradients within the mitotic cell stabilize and orient the nascent MTs in order to organize MTs into a spindle-shaped structure, with MT minus ends focused at the two poles and their plus ends interdigitating or connecting with the chromosomes.
1.4 MT Organization in the Spindle: Different Configurations, Functions, and Properties
While keeping highly dynamic properties, MTs get organized into different structures whose properties and dynamics change throughout cell division. In this section, we will focus on the principles driving the organization of the three most characteristic mitotic MT assemblies: the bipolar spindle, the K-fibers, and the central spindle.
1.4.1 The Bipolar Spindle
The organization of MTs into two interdigitating antiparallel arrays is key to cell division. The main forces driving MT organization into this typical configuration are provided by molecular motors that interact with MTs in an ATP-dependent manner using the energy derived from its hydrolysis to move directionally along the MTs. While some of them move toward MT minus ends (dynein and some kinesins), others move toward the plus ends (most of the kinesins). Recent excellent reviews describe in detail the mechanochemistry of motor movement and force generation (Roberts et al. 2013; Cross and McAinsh 2014).
Mitosis in human cells involves the activity of a large number of kinesins and cytoplasmic dynein. The collective action of these motors drives bipolar spindle assembly by establishing and maintaining three main activities: stable but dynamic interactions between the two antiparallel MT asters, focusing MT minus ends into the spindle poles, and dynamic interactions between MTs and chromosomes. Each one of these functions is related to specific motor organizations and mechanisms of action. Motors can indeed cross-link and move on two antiparallel MTs, link two MTs but move on only one of them, or mediate the MT-chromosome interaction. We will here briefly describe examples of these three kinds of motors.
Eg5 (also called KIF11 or kinesin 5) is an extensively characterized mitotic motor. It is a homotetramer that can interact with two MTs preferentially in an antiparallel configuration. By moving toward the plus ends of the two cross-linked MTs, it drives their separation (van den Wildenberg et al. 2008; Tanenbaum and Medema 2010) (Fig. 1.3). This is an essential mechanism promoting spindle pole separation and bipolarity establishment and maintenance. Eg5 function is essential for bipolar spindle organization. In the absence of Eg5 activity, MTs organize into a monopolar spindle with the MT minus ends in the center and the plus ends at the periphery of a rosette-like structure (Mayer et al. 1999).
Bipolar spindle MT organization. Left: schematic representation of a prometaphase spindle. Right boxes: (upper box) Eg5 is a homotetrameric, plus-end-directed motor. It works separating two antiparallel MTs apart and is essential for spindle bipolarization and cell division. Middle box: HSET and dynein are two types of minus-end-directed motors. They move on one MT and also interact with another, parallel, MT. This process is essential for focusing MT minus ends at the poles. Lower box: KID is one example of chromokinesins. It is a plus-end-directed, homodimeric motor interacting with an MT through its motor domain and with chromosomes through its C-terminal part. Its function is essential in congressing the chromosomes in the metaphase plate
Other MT-cross-linking motors adopt distinct configurations, such as HSET, also known as KIFC1 or XCTK2 (Walczak et al. 1997; Mountain et al. 1999). HSET is a minus-end-directed kinesin, which cross-links two MTs, but moves only on one of them. It is organized as a homodimer that can interact with one MT through its motor domain and with another MT through another ATP-independent MT-binding domain, promoting MT organization (Fig. 1.3). HSET plays a role in spindle length control and, like the dynein complex, in spindle pole focusing (Cai et al. 2009; Hentrich and Surrey 2010). Dynein is a major minus-end-directed motor in the mitotic cell. Depending on its “cargo,” dynein functions by promoting MT movement in relation to another MT (pole focusing), to the nuclear envelope (pole separation in prophase), to the cell cortex (spindle positioning), or to the kinetochore (chromosome positioning) (Roberts et al. 2013) (Fig. 1.3).
Another class of motors includes the chromokinesins that mediate interactions between MTs and the chromosome arms. The forces exerted by chromokinesins are called polar ejection forces that play important roles in chromosome congression (Vanneste et al. 2011). At least two classes of kinesins are able to directly interact with chromosome arms: KIF22 (Kid) and KIF4 (Mazumdar and Misteli 2005). The role of Kid in chromosome congression is essential (Levesque and Compton 2001). In Xenopus egg extracts, Xkid depletion results in the dramatic chromosome scattering phenotype (Antonio et al. 2000; Funabiki and Murray 2000). Chromokinesins share common structural features. They are all plus-end-directed motors and function in homodimers, with a motor MT-binding domain in N-terminal and a DNA-binding motif in the C-terminal part (Vanneste et al. 2011) (Fig. 1.3).
The complexity of motor functions and their interactions is overall a challenge to achieve a full understanding of their roles in bipolar spindle assembly. Their function cannot be limited to focusing minus ends at the poles, congressing chromosomes, and cross-linking antiparallel MTs in the region of their overlap. For example, dynein has also a role in the targeting of a number of essential factors to the spindle poles, including Eg5 and TPX2 (Ma et al. 2010).
1.4.2 The K-Fibers
As the spindle assembles, the dynamic plus ends of some MTs are “captured” by the kinetochore, a specialized region organized as a paired structure on each chromosome. MT plus-end attachment to the kinetochore is mediated by interactions with the Ndc80 and Ska complexes leading to their stabilization (Jeyaprakash et al. 2012; Cheerambathur et al. 2013; Shrestha and Draviam 2013). These MTs form bundles of 20–40 MTs organized in parallel orientation, called the K-fibers. K-fibers have specific dynamic properties and are remarkably more stable than the other spindle MTs (Rieder 1981). They generate pushing forces that contribute to centrosome separation and the establishment of spindle bipolarity and are obviously essential for chromosome movements and segregation (McHedlishvili et al. 2012).
The K-fiber MT bundles require protein complexes forming bridges in between the parallel MTs. One of them is composed by clathrin and TACC3 that interacts with the MT polymerase chTOG/XMAP215 (Booth et al. 2011) (Fig. 1.4). K-fiber organization in bundles is however still poorly understood, since other kinds of inter-MT bridges have been observed (Booth et al. 2011).
K-fiber organization. K-fibers are bundles of 20–40 MTs. Bridges between MTs, in part composed by XMAP214, clathrin, and TACC3, maintain the MTs forming the K-fiber together. Their plus ends are interacting with the kinetochores, and their minus ends are focused in close proximity to the spindle poles. K-fibers are constantly depolymerizing at the minus end and overall incorporate new tubulin dimers at the plus end. This mechanism creates a tubulin poleward flux from the plus to the minus end of the K-fibers. While kinesins 13 work at both ends in depolymerizing K-fiber MTs, KIF18A is a depolymerizing kinesin specific for the plus end. At the plus end, +TIPS factors such as EB1 and CLASPs favor MT polymerization. At the minus end, MCRS1 protects MTs against destabilization
The K-fibers exhibit very specific plus- and minus-end dynamics that are tightly related to attachment error correction and chromosome movements. The MT plus ends of the K-fiber alternate between phases of growth and depolymerization that drive poleward and anti-poleward chromosome movements and oscillations that result in chromosome alignment at the metaphase plate (Magidson et al. 2011). While the plus ends exhibit a “switching” behavior alternating between phases of MT growth and shrinkage coordinated at the two sister kinetochores, K-fiber MT minus ends depolymerize constantly. This results in a characteristic tubulin flux from the plus end toward the minus end (Fig. 1.4) that generates forces within the spindle strong enough to move the chromosomes (Waters et al. 1996).
K-fiber dynamics regulation involves both proteins favoring MT polymerization like CLASP or EB1 and others promoting MT destabilization like the kinesin 13 and 8 family members (Tirnauer et al. 2002; Maiato et al. 2005; Joglekar et al. 2010; Manning et al. 2010) (Fig. 1.4). The mechanism involved in regulating MT depolymerization at the minus end is still unclear. However, the recent identification of novel proteins that specifically protect MT minus ends from depolymerase activities opens the way to a better understanding of K-fiber dynamics (Goodwin and Vale 2010; Meunier and Vernos 2011; Jiang et al. 2014; Meunier et al. 2015) (Fig. 1.4).
A dramatic change in K-fiber dynamics drives K-fiber shortening and thereby chromosome segregation in anaphase (Waters et al. 1996). However, how the shortening of all K-fibers is coordinated, which signal triggers of the process, and what mechanism controls the depolymerization rate are still open questions.
1.4.3 The Central Spindle
During anaphase, a new MT-based structure, called the central spindle, assembles between the two segregating chromosome masses. The central spindle is formed by interdigitating MT arrays that promote the separation of the centrosomes and chromosomes by sliding in an antiparallel manner. MTs in the central spindle are organized into bundles that are remarkably more stable than those forming the metaphase spindle (Saxton and McIntosh 1987).
The mechanism underlying central spindle assembly involves at least two protein complexes. One of them called, centralspindlin, initiates central spindle assembly. It is formed by a kinesin-like protein, MKLP1, and a Rho GTPase-activating protein (RhoGAP), CYK-4. This protein complex is targeted to an antiparallel MT overlapping area immediately after chromosome segregation (Mishima et al. 2002; White and Glotzer 2012) (Fig. 1.5). Centralspindlin plays also essential roles in midbody assembly and abscission, the very last steps in the full separation of the daughter cells.
Central spindle organization. Left: schematic representation of an anaphase B spindle, with the central spindle in the center. Right box: detail of central spindle organization. MTs plus tips are interdigitating in the central spindle. They are cross-linked together though the action of two main complexes: the centralspindlin complex and the complex formed by PRC1 and KIF4. The interaction between PRC1 and KIF4 is essential for maintaining central spindle length and defining the antiparallel, overlapping MT region
The mechanism defining the extent of MT overlap and sliding in the central spindle involves another complex constituted by the protein PRC1 (protein regulator of cytokinesis 1), a MAP with MT bundling activity in vitro (Mollinari et al. 2002), and the kinesin KIF4 (Kurasawa et al. 2004) (Fig. 1.5). In vitro experiments with recombinant PRC1 and KIF4 showed that these two activities are sufficient to reconstitute a central spindle-like organization (Bieling et al. 2010).
Several regulating protein complexes are targeted to the central spindle during anaphase. Among them, the chromosomal passenger complex (CPC) containing the Aurora B kinase relocalizes from the kinetochores to the central spindle and regulates central spindle assembly and function, by phosphorylating a variety of substrates (Guse et al. 2005). Recently, it was also shown that the kinase Aurora A plays a role in central spindle organization and dynamics regulation through the phosphorylation of the dynein complex and TACC3 (Lioutas and Vernos 2013; Reboutier et al. 2013).
The central spindle is constituted by some interpolar MTs, but recent studies have demonstrated that it can be assembled de novo, involving MT nucleation and stabilization. The signals triggering the assembly of these MTs are however still not identified, but the involvement of the RanGTP pathway and the augmin complex is likely (Glotzer 2009; Uehara and Goshima 2010).
1.5 Specifying MT Identities in the Spindle?
MTs during mitosis show a range of dynamic properties, organization, and function that coexist at any given time and evolve as mitosis proceeds. Overall, astral and interpolar MTs are very dynamic mainly at their plus ends. K-fibers are organized into bundles with specific dynamic properties both at the plus and at the minus ends, while being more stable than the other spindle MTs. How these characteristics are specified is still not understood. In fact, it is unclear whether MT organization determines function (for instance, MTs contacting the kinetochore organize into K-fibers) or whether different MTs with specific properties define organization and function. In this context, it may be relevant to consider that mitotic MTs originate at different sites and through different pathways that involve specific components and regulators. Interestingly, some proteins were found to specifically associate with one class of MTs. The RanGTP-regulated protein HURP associates only with the K-fibers in a region close to the chromosomes (Sillje et al. 2006). Another RanGTP-regulated protein, MCRS1, associates exclusively to chromosomal MTs and to those forming the K-fibers (Meunier and Vernos 2011). These data suggest that the chromosomal MTs participate at least in part to K-fiber formation, as they have specific MAPs that confer them properties different to the other MTs. This hypothesis still requires experimental support to be confirmed.
Other mechanisms could potentially confer specific properties to the spindle MTs. Although all MTs are formed by α-/β-tubulin dimers, mammalian cells have several genes for these two proteins (7 α-tubulin and 8 β-tubulin genes in humans). The different tubulin isotypes are extremely conserved (more than 95 % amino acid sequence identity), but their C-terminal tails have more variability. Interestingly, this region is exposed on the MT surface and is responsible for binding MAPs and motors (Sirajuddin et al. 2014). The expression of specific tubulin isotypes has been related with specific MT organizations (Raff et al. 1997) and with adaptation mechanisms. Although it is not yet clear whether they play any role in mitosis, changes in the expression pattern of tubulin isotypes have been reported in some cancer cells that show altered MT dynamics and resistance to antitumor treatments that target tubulin (Wang et al. 2014).
In addition to the expression of specific tubulin isotypes, several posttranslational modifications (PTMs) on the C-terminal tails of the tubulins modulate the binding affinities of MAPs and motors and may even change motor processivity and/or velocity (Janke and Bulinski 2011; Sirajuddin et al. 2014). Many PTMs have been described including detyrosination, mono- or poly-glutamylation, phosphorylation, acetylation, and glycylation. These modifications may constitute a “tubulin code” that could specify which MAPs and which motors would bind to one specific MT (Janke and Bulinski 2011; Magiera and Janke 2014; Barisic et al. 2015). Although this remains speculative, the number of α- and β-tubulin isotypes together with the combinatorial possibilities of PTMs at their C-terminus potentially offers a myriad of possibilities to precisely define the properties of MT subpopulations, and this could play a role in the bipolar spindle.
1.6 Mitotic MTs in Health and Disease
1.6.1 Mitotic MT-Related Disorders and Pathologies
Cell division is fundamental for life. Any error in this process may be fatal or generate cells with an incorrect chromosome number. Aneuploidy, the loss or gain of chromosomes, is the leading genetic cause of miscarriage and congenital birth defects as well as being tightly associated to health-threatening conditions like cancer.
In humans, as many as one in five pregnancies end in miscarriage, the most common complication of early pregnancy. Aneuploidy is the leading known cause of miscarriage, but some of them (as, e.g. trisomy 21 and monosomy X, Down or Turner syndromes, respectively) are compatible with live birth, making aneuploidy the leading cause of congenital birth defects and mental retardation. A common cause for these miscarriages seems to be aneuploidies in human oocytes, which increase dramatically with age (Holubcova et al. 2015). This is associated to the weakening of cohesion between the sister chromatids with time (Chiang et al. 2010; Duncan et al. 2012). This phenomenon was recently described as “chromosome fatigue” (Daum et al. 2011) and is particularly threatening for human reproduction because oocytes are maintained blocked in prophase of meiosis I from birth until maturation is induced after puberty on a monthly basis throughout the reproductive lifespan (Chiang et al. 2010).
A number of pathologies derive from spindle orientation defects that compromise the fate of the daughter cells during development (Noatynska et al. 2012). During development, neural progenitor cells undergo symmetric and asymmetric divisions from a monolayer of stem cells to build the brain. Mutations in genes related with centrosome function and duplication or with astral MT stabilization have been related to brain development defects like microcephaly or lissencephaly, respectively (Fish et al. 2006; Yingling et al. 2008; Chavali et al. 2014). However, the mutated genes have a large range of distinct functions in various processes and organs (Noatynska et al. 2012) and a direct causal relationship between spindle orientation defects and these brain pathologies is currently missing.
Cell division is essential after birth for the growth of organs and body parts and throughout adulthood for the maintenance and renewal of cells and tissues. Mutations in proteins related with mitotic MT regulation, for example, in the PCM component pericentrin, have been linked to a number of pathologies including cancer (Delaval and Doxsey 2010). Most human solid tumors have aneuploid cells due to CIN (chromosome instability), which promotes chromosome missegregation in mitosis. CIN occurs early in tumorigenesis and is associated with poor prognosis. Aneuploid cells may get supernumerary copies of oncogenes and/or insufficient copies of tumor suppressor genes, which could favor the development of tumors (Duijf and Benezra 2013; Salmela and Kallio 2013). CIN can be caused by multiple mechanisms including a weakened or overactivated mitotic spindle assembly checkpoint, sister chromatid cohesion defects, increased merotelic kinetochore‐microtubule attachments, or the presence of extra centrosomes. Although CIN was proposed as a leading cause of tumor progression, recent studies suggest that CIN can either promote or suppress tumor progression, depending on the context.
Aneuploidy or other mechanisms may also be involved with changes in the expression levels of mitotic factors: enzymes involved in the regulation of the cell division like the kinase Aurora A as well as MT-binding proteins like TPX2. These two proteins interact during mitosis and the TPX2-Aurora A complex has been described as an “oncogenic holoenzyme” (Asteriti et al. 2010). Interestingly, TPX2 was found to be the protein with the highest CIN (chromosome instability) score among 10,000 analyzed genes in a number of tumors (Carter et al. 2006).
1.6.2 MTs and Therapeutic Strategies
The highly dynamic properties of the mitotic MTs are essential for the assembly of a functional spindle. At the same time, they render mitotic cells particularly sensitive to factors that alter these properties. In fact, MT-binding agents (TBAs) that alter MT dynamics were the first antitumor compounds used for cancer treatment. There are different classes of TBAs isolated from a broad range of species, such as bacteria, sponges, or plants. They either promote MT stabilization (such as taxanes and epothilones) or MT destabilization (such as vinblastine). The prevailing idea is that TBAs exert an anticancer activity by targeting dividing cells particularly abundant in tumors, although this is under debate (Mitchison 2012; Topham and Taylor 2013). In any case, these drugs are still widely used in the clinic as they have a clear therapeutic value. However, there is a need for novel ways to fight tumors, as cancer cells can evade the effects of compounds and drugs through different mechanisms. Interestingly, one of them is the expression of the neuronal βΙΙΙ-tubulin isotype (Cortes and Vidal 2011). Moreover, TBAs are not specific for the dividing cells, generating secondary effects like neurotoxicity (Harrison et al. 2009; Kavallaris 2010).
Targeting characteristics specific to tumor cells, such as CIN, aneuploidies, and supernumerary centrosomes, is therefore an attractive therapeutic avenue. Compounds targeting mitotic factors such as the kinases Plk1, Aurora A or Aurora B (Salmela and Kallio 2013), or microtubule motors have been developed. Some of them are in different phases of clinical trials (Ding et al. 2014).
1.7 Conclusions
Spindle assembly relies on the coordination of a number of individual events including MT nucleation, stabilization, and organization. The importance of accurate chromosome segregation for the continuity of life is underscored by tightly controlled mechanisms ensuring the interaction between MTs and chromosomes with several layers of regulations and checkpoints that ensure the fidelity of the system. However, errors in cell division can occur, often leading to catastrophic consequences in terms of fertility, development, or tissue maintenance and renewal. Although individual pieces of the puzzle start to be well understood, future work will certainly focus on getting more information on the coordination of all single events and thereby a global view of a system at the basis of life transmission.
References
Akhmanova A, Steinmetz MO (2010) Microtubule + TIPs at a glance. J Cell Sci 123(Pt 20):3415–3419. doi:123/20/3415 [pii] 10.1242/jcs.062414
Alushin GM, Lander GC, Kellogg EH, Zhang R, Baker D, Nogales E (2014) High-resolution microtubule structures reveal the structural transitions in alphabeta-tubulin upon GTP hydrolysis. Cell 157(5):1117–1129. doi:S0092-8674(14)00483-8 [pii] 10.1016/j.cell.2014.03.053
Antonio C, Ferby I, Wilhelm H, Jones M, Karsenti E, Nebreda AR, Vernos I (2000) Xkid, a chromokinesin required for chromosome alignment on the metaphase plate. Cell 102(4):425–435. doi:S0092-8674(00)00048-9 [pii]
Asteriti IA, Rensen WM, Lindon C, Lavia P, Guarguaglini G (2010) The Aurora-A/TPX2 complex: a novel oncogenic holoenzyme? Biochim Biophys Acta 1806(2):230–239. doi:S0304-419X(10)00058-2 [pii] 10.1016/j.bbcan.2010.08.001
Barisic M, Silva e Sousa R, Tripathy SK, Magiera MM, Zaytsev AV, Pereira AL, Janke C, Grishchuk EL, Maiato H (2015) Mitosis. Microtubule detyrosination guides chromosomes during mitosis. Science 348(6236):799–803. doi:science.aaa5175 [pii] 10.1126/science.aaa5175
Bayliss R, Sardon T, Vernos I, Conti E (2003) Structural basis of Aurora-A activation by TPX2 at the mitotic spindle. Mol Cell 12(4):851–862. doi:S1097276503003927 [pii]
Belmont LD, Mitchison TJ (1996) Identification of a protein that interacts with tubulin dimers and increases the catastrophe rate of microtubules. Cell 84(4):623–631. doi:S0092-8674(00)81037-5 [pii]
Bettencourt-Dias M (2013) Q&A: who needs a centrosome? BMC Biol 11:28. doi:1741-7007-11-28 [pii] 10.1186/1741-7007-11-28
Bettencourt-Dias M, Glover DM (2007) Centrosome biogenesis and function: centrosomics brings new understanding. Nat Rev Mol Cell Biol 8(6):451–463. doi:nrm2180 [pii] 10.1038/nrm2180
Bieling P, Telley IA, Surrey T (2010) A minimal midzone protein module controls formation and length of antiparallel microtubule overlaps. Cell 142(3):420–432. doi:S0092-8674(10)00723-3 [pii] 10.1016/j.cell.2010.06.033
Booth DG, Hood FE, Prior IA, Royle SJ (2011) A TACC3/ch-TOG/clathrin complex stabilises kinetochore fibres by inter-microtubule bridging. EMBO J 30(5):906–919. doi:emboj201115 [pii] 10.1038/emboj.2011.15
Brouhard GJ, Stear JH, Noetzel TL, Al-Bassam J, Kinoshita K, Harrison SC, Howard J, Hyman AA (2008) XMAP215 is a processive microtubule polymerase. Cell 132(1):79–88. doi:S0092-8674(07)01547-4 [pii] 10.1016/j.cell.2007.11.043
Cai S, Weaver LN, Ems-McClung SC, Walczak CE (2009) Kinesin-14 family proteins HSET/XCTK2 control spindle length by cross-linking and sliding microtubules. Mol Biol Cell 20(5):1348–1359. doi:E08-09-0971 [pii] 10.1091/mbc.E08-09-0971
Carazo-Salas RE, Gruss OJ, Mattaj IW, Karsenti E (2001) Ran-GTP coordinates regulation of microtubule nucleation and dynamics during mitotic-spindle assembly. Nat Cell Biol 3(3):228–234. doi:10.1038/35060009
Carazo-Salas RE, Guarguaglini G, Gruss OJ, Segref A, Karsenti E, Mattaj IW (1999) Generation of GTP-bound Ran by RCC1 is required for chromatin-induced mitotic spindle formation. Nature 400(6740):178–181. doi:10.1038/22133
Carmena M, Wheelock M, Funabiki H, Earnshaw WC (2012) The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat Rev Mol Cell Biol 13(12):789–803. doi:nrm3474 [pii] 10.1038/nrm3474
Carter SL, Eklund AC, Kohane IS, Harris LN, Szallasi Z (2006) A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nat Genet 38(9):1043–1048. doi:ng1861 [pii] 10.1038/ng1861
Cassimeris L (2002) The oncoprotein 18/stathmin family of microtubule destabilizers. Curr Opin Cell Biol 14(1):18–24
Caudron M, Bunt G, Bastiaens P, Karsenti E (2005) Spatial coordination of spindle assembly by chromosome-mediated signaling gradients. Science 309(5739):1373–1376. doi:309/5739/1373 [pii] 10.1126/science.1115964
Chavali PL, Putz M, Gergely F (2014) Small organelle, big responsibility: the role of centrosomes in development and disease. Philos Trans R Soc Lond B Biol Sci 369(1650). pii:20130468. doi:10.1098/rstb.2013.0468
Cheerambathur DK, Gassmann R, Cook B, Oegema K, Desai A (2013) Crosstalk between microtubule attachment complexes ensures accurate chromosome segregation. Science 342(6163):1239–1242. doi:science.1246232 [pii] 10.1126/science.1246232
Chiang T, Duncan FE, Schindler K, Schultz RM, Lampson MA (2010) Evidence that weakened centromere cohesion is a leading cause of age-related aneuploidy in oocytes. Curr Biol 20(17):1522–1528. doi:S0960-9822(10)00817-1 [pii] 10.1016/j.cub.2010.06.069
Clarke PR, Zhang C (2008) Spatial and temporal coordination of mitosis by Ran GTPase. Nat Rev Mol Cell Biol 9(6):464–477. doi:nrm2410 [pii] 10.1038/nrm2410
Cortes J, Vidal M (2011) Beyond taxanes: the next generation of microtubule-targeting agents. Breast Cancer Res Treat. doi:10.1007/s10549-011-1875-6
Cross RA, McAinsh A (2014) Prime movers: the mechanochemistry of mitotic kinesins. Nat Rev Mol Cell Biol 15(4):257–271. doi:nrm3768 [pii] 10.1038/nrm3768
Daum JR, Potapova TA, Sivakumar S, Daniel JJ, Flynn JN, Rankin S, Gorbsky GJ (2011) Cohesion fatigue induces chromatid separation in cells delayed at metaphase. Curr Biol 21(12):1018–1024. doi:S0960-9822(11)00588-4 [pii] 10.1016/j.cub.2011.05.032
Delaval B, Doxsey SJ (2010) Pericentrin in cellular function and disease. J Cell Biol 188(2):181–190. doi:jcb.200908114 [pii] 10.1083/jcb.200908114
Desai A, Verma S, Mitchison TJ, Walczak CE (1999) Kin I kinesins are microtubule-destabilizing enzymes. Cell 96(1):69–78. doi:S0092-8674(00)80960-5 [pii]
Ding S, Zhao Z, Sun D, Wu F, Bi D, Lu J, Xing N, Sun L, Wu H, Ding K (2014) Eg5 inhibitor, a novel potent targeted therapy, induces cell apoptosis in renal cell carcinoma. Tumour Biol. doi:10.1007/s13277-014-2022-x
Du Y, English CA, Ohi R (2010) The kinesin-8 Kif18A dampens microtubule plus-end dynamics. Curr Biol 20(4):374–380. doi:S0960-9822(09)02212-X [pii] 10.1016/j.cub.2009.12.049
Duijf PH, Benezra R (2013) The cancer biology of whole-chromosome instability. Oncogene 32(40):4727–4736. doi:onc2012616 [pii] 10.1038/onc.2012.616
Duncan FE, Hornick JE, Lampson MA, Schultz RM, Shea LD, Woodruff TK (2012) Chromosome cohesion decreases in human eggs with advanced maternal age. Aging Cell 11(6):1121–1124. doi:10.1111/j.1474-9726.2012.00866.x
Euteneuer U, McIntosh JR (1981) Structural polarity of kinetochore microtubules in PtK1 cells. J Cell Biol 89(2):338–345
Euteneuer U, Ris H, Borisy GG (1983) Polarity of kinetochore microtubules in Chinese hamster ovary cells after recovery from a colcemid block. J Cell Biol 97(1):202–208
Eyers PA, Erikson E, Chen LG, Maller JL (2003) A novel mechanism for activation of the protein kinase Aurora A. Curr Biol 13(8):691–697. doi:S0960982203001660 [pii]
Fish JL, Kosodo Y, Enard W, Paabo S, Huttner WB (2006) Aspm specifically maintains symmetric proliferative divisions of neuroepithelial cells. Proc Natl Acad Sci U S A 103(27):10438–10443. doi:0604066103 [pii] 10.1073/pnas.0604066103
Funabiki H, Murray AW (2000) The Xenopus chromokinesin Xkid is essential for metaphase chromosome alignment and must be degraded to allow anaphase chromosome movement. Cell 102(4):411–424. doi:S0092-8674(00)00047-7 [pii]
Gard DL, Kirschner MW (1987) A microtubule-associated protein from Xenopus eggs that specifically promotes assembly at the plus-end. J Cell Biol 105(5):2203–2215
Glotzer M (2009) The 3Ms of central spindle assembly: microtubules, motors and MAPs. Nat Rev Mol Cell Biol 10(1):9–20. doi:nrm2609 [pii] 10.1038/nrm2609
Gomez-Ferreria MA, Bashkurov M, Helbig AO, Larsen B, Pawson T, Gingras AC, Pelletier L (2012) Novel NEDD1 phosphorylation sites regulate gamma-tubulin binding and mitotic spindle assembly. J Cell Sci 125(Pt 16):3745–3751. doi:jcs.105130 [pii] 10.1242/jcs.105130
Goodwin SS, Vale RD (2010) Patronin regulates the microtubule network by protecting microtubule minus ends. Cell 143(2):263–274. doi:S0092-8674(10)01070-6 [pii] 10.1016/j.cell.2010.09.022
Goshima G, Mayer M, Zhang N, Stuurman N, Vale RD (2008) Augmin: a protein complex required for centrosome-independent microtubule generation within the spindle. J Cell Biol 181(3):421–429. doi:jcb.200711053 [pii] 10.1083/jcb.200711053
Goshima G, Wollman R, Goodwin SS, Zhang N, Scholey JM, Vale RD, Stuurman N (2007) Genes required for mitotic spindle assembly in Drosophila S2 cells. Science 316(5823):417–421. doi:1141314 [pii] 10.1126/science.1141314
Groen AC, Cameron LA, Coughlin M, Miyamoto DT, Mitchison TJ, Ohi R (2004) XRHAMM functions in ran-dependent microtubule nucleation and pole formation during anastral spindle assembly. Curr Biol 14(20):1801–1811. doi:S0960982204007924 [pii] 10.1016/j.cub.2004.10.002
Gruss OJ, Carazo-Salas RE, Schatz CA, Guarguaglini G, Kast J, Wilm M, Le Bot N, Vernos I, Karsenti E, Mattaj IW (2001) Ran induces spindle assembly by reversing the inhibitory effect of importin alpha on TPX2 activity. Cell 104(1):83–93. doi:S0092-8674(01)00193-3 [pii]
Gruss OJ, Wittmann M, Yokoyama H, Pepperkok R, Kufer T, Sillje H, Karsenti E, Mattaj IW, Vernos I (2002) Chromosome-induced microtubule assembly mediated by TPX2 is required for spindle formation in HeLa cells. Nat Cell Biol 4(11):871–879. doi:10.1038/ncb870 ncb870 [pii]
Gupta KK, Li C, Duan A, Alberico EO, Kim OV, Alber MS, Goodson HV (2013) Mechanism for the catastrophe-promoting activity of the microtubule destabilizer Op18/stathmin. Proc Natl Acad Sci U S A 110(51):20449–20454. doi:1309958110 [pii] 10.1073/pnas.1309958110
Guse A, Mishima M, Glotzer M (2005) Phosphorylation of ZEN-4/MKLP1 by aurora B regulates completion of cytokinesis. Curr Biol 15(8):778–786. doi:S0960-9822(05)00335-0 [pii] 10.1016/j.cub.2005.03.041
Haren L, Remy MH, Bazin I, Callebaut I, Wright M, Merdes A (2006) NEDD1-dependent recruitment of the gamma-tubulin ring complex to the centrosome is necessary for centriole duplication and spindle assembly. J Cell Biol 172(4):505–515. doi:jcb.200510028 [pii] 10.1083/jcb.200510028
Harrison MR, Holen KD, Liu G (2009) Beyond taxanes: a review of novel agents that target mitotic tubulin and microtubules, kinases, and kinesins. Clin Adv Hematol Oncol 7(1):54–64
Heald R, Tournebize R, Blank T, Sandaltzopoulos R, Becker P, Hyman A, Karsenti E (1996) Self-organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382(6590):420–425. doi:10.1038/382420a0
Hentrich C, Surrey T (2010) Microtubule organization by the antagonistic mitotic motors kinesin-5 and kinesin-14. J Cell Biol 189(3):465–480. doi:jcb.200910125 [pii] 10.1083/jcb.200910125
Holubcova Z, Blayney M, Elder K, Schuh M (2015) Human oocytes. Error-prone chromosome-mediated spindle assembly favors chromosome segregation defects in human oocytes. Science 348(6239):1143–1147. doi:348/6239/1143 [pii] 10.1126/science.aaa9529
Hunter AW, Caplow M, Coy DL, Hancock WO, Diez S, Wordeman L, Howard J (2003) The kinesin-related protein MCAK is a microtubule depolymerase that forms an ATP-hydrolyzing complex at microtubule ends. Mol Cell 11(2):445–457. doi:S1097276503000492 [pii]
Inoue S, Sato H (1967) Cell motility by labile association of molecules. The nature of mitotic spindle fibers and their role in chromosome movement. J Gen Physiol 50(6):Suppl:259–292
Janke C, Bulinski JC (2011) Post-translational regulation of the microtubule cytoskeleton: mechanisms and functions. Nat Rev Mol Cell Biol 12(12):773–786. doi:10.1038/nrm3227
Jeyaprakash AA, Santamaria A, Jayachandran U, Chan YW, Benda C, Nigg EA, Conti E (2012) Structural and functional organization of the Ska complex, a key component of the kinetochore-microtubule interface. Mol Cell 46(3):274–286. doi:S1097-2765(12)00212-2 [pii] 10.1016/j.molcel.2012.03.005
Jiang K, Hua S, Mohan R, Grigoriev I, Yau KW, Liu Q, Katrukha EA, Altelaar AF, Heck AJ, Hoogenraad CC, Akhmanova A (2014) Microtubule minus-end stabilization by polymerization-driven CAMSAP deposition. Dev Cell 28(3):295–309. doi:S1534-5807(14)00002-1 [pii] 10.1016/j.devcel.2014.01.001
Joglekar AP, Bloom KS, Salmon ED (2010) Mechanisms of force generation by end-on kinetochore-microtubule attachments. Curr Opin Cell Biol 22(1):57–67. doi:S0955-0674(09)00239-7 [pii] 10.1016/j.ceb.2009.12.010
Johmura Y, Soung NK, Park JE, Yu LR, Zhou M, Bang JK, Kim BY, Veenstra TD, Erikson RL, Lee KS (2011) Regulation of microtubule-based microtubule nucleation by mammalian polo-like kinase 1. Proc Natl Acad Sci U S A 108(28):11446–11451. doi:1106223108 [pii] 10.1073/pnas.1106223108
Kalab P, Pu RT, Dasso M (1999) The ran GTPase regulates mitotic spindle assembly. Curr Biol 9(9):481–484. doi:S0960-9822(99)80213-9 [pii]
Kalab P, Weis K, Heald R (2002) Visualization of a Ran-GTP gradient in interphase and mitotic Xenopus egg extracts. Science 295(5564):2452–2456. doi:10.1126/science.1068798 295/5564/2452 [pii]
Kamasaki T, O’Toole E, Kita S, Osumi M, Usukura J, McIntosh JR, Goshima G (2013) Augmin-dependent microtubule nucleation at microtubule walls in the spindle. J Cell Biol 202(1):25–33. doi:jcb.201304031 [pii] 10.1083/jcb.201304031
Karsenti E, Newport J, Kirschner M (1984) Respective roles of centrosomes and chromatin in the conversion of microtubule arrays from interphase to metaphase. J Cell Biol 99(1 Pt 2):47s–54s
Karsenti E, Vernos I (2001) The mitotic spindle: a self-made machine. Science 294(5542):543–547. doi:10.1126/science.1063488 294/5542/543 [pii]
Kavallaris M (2010) Microtubules and resistance to tubulin-binding agents. Nat Rev Cancer 10(3):194–204. doi:nrc2803 [pii] 10.1038/nrc2803
Khodjakov A, Rieder CL (1999) The sudden recruitment of gamma-tubulin to the centrosome at the onset of mitosis and its dynamic exchange throughout the cell cycle, do not require microtubules. J Cell Biol 146(3):585–596
Kirschner M, Mitchison T (1986) Review. Beyond self-assembly: from microtubules to morphogenesis. Cell 45(3):329–342
Kitamura E, Tanaka K, Komoto S, Kitamura Y, Antony C, Tanaka TU (2010) Kinetochores generate microtubules with distal plus ends: their roles and limited lifetime in mitosis. Dev Cell 18(2):248–259. doi:S1534-5807(10)00017-1 [pii] 10.1016/j.devcel.2009.12.018
Kiyomitsu T, Cheeseman IM (2012) Chromosome- and spindle-pole-derived signals generate an intrinsic code for spindle position and orientation. Nat Cell Biol 14(3):311–317. doi:ncb2440 [pii] 10.1038/ncb2440
Kollman JM, Merdes A, Mourey L, Agard DA (2011) Microtubule nucleation by gamma-tubulin complexes. Nat Rev Mol Cell Biol. doi:nrm3209 [pii] 10.1038/nrm3209
Kronja I, Kruljac-Letunic A, Caudron-Herger M, Bieling P, Karsenti E (2009) XMAP215-EB1 interaction is required for proper spindle assembly and chromosome segregation in Xenopus egg extract. Mol Biol Cell 20(11):2684–2696. doi:E08-10-1051 [pii] 10.1091/mbc.E08-10-1051
Kurasawa Y, Earnshaw WC, Mochizuki Y, Dohmae N, Todokoro K (2004) Essential roles of KIF4 and its binding partner PRC1 in organized central spindle midzone formation. EMBO J 23(16):3237–3248. doi:10.1038/sj.emboj.7600347 7600347 [pii]
Lawo S, Bashkurov M, Mullin M, Ferreria MG, Kittler R, Habermann B, Tagliaferro A, Poser I, Hutchins JR, Hegemann B, Pinchev D, Buchholz F, Peters JM, Hyman AA, Gingras AC, Pelletier L (2009) HAUS, the 8-subunit human Augmin complex, regulates centrosome and spindle integrity. Curr Biol 19(10):816–826. doi:S0960-9822(09)01032-X [pii] 10.1016/j.cub.2009.04.033
Lecland N, Luders J (2014) The dynamics of microtubule minus ends in the human mitotic spindle. Nat Cell Biol 16(8):770–778. doi:ncb2996 [pii] 10.1038/ncb2996
Levesque AA, Compton DA (2001) The chromokinesin Kid is necessary for chromosome arm orientation and oscillation, but not congression, on mitotic spindles. J Cell Biol 154(6):1135–1146. doi:10.1083/jcb.200106093
Lioutas A, Vernos I (2013) Aurora A kinase and its substrate TACC3 are required for central spindle assembly. EMBO Rep 14(9):829–836. doi:embor2013109 [pii] 10.1038/embor.2013.109
Luders J, Patel UK, Stearns T (2006) GCP-WD is a gamma-tubulin targeting factor required for centrosomal and chromatin-mediated microtubule nucleation. Nat Cell Biol 8(2):137–147. doi:ncb1349 [pii] 10.1038/ncb1349
Ma N, Tulu US, Ferenz NP, Fagerstrom C, Wilde A, Wadsworth P (2010) Poleward transport of TPX2 in the mammalian mitotic spindle requires dynein, Eg5, and microtubule flux. Mol Biol Cell 21(6):979–988. doi:E09-07-0601 [pii] 10.1091/mbc.E09-07-0601
Magidson V, O’Connell CB, Loncarek J, Paul R, Mogilner A, Khodjakov A (2011) The spatial arrangement of chromosomes during prometaphase facilitates spindle assembly. Cell 146(4):555–567. doi:S0092-8674(11)00773-2 [pii] 10.1016/j.cell.2011.07.012
Magiera MM, Janke C (2014) Post-translational modifications of tubulin. Curr Biol 24(9):R351–R354. doi:S0960-9822(14)00324-8 [pii] 10.1016/j.cub.2014.03.032
Maiato H, Khodjakov A, Rieder CL (2005) Drosophila CLASP is required for the incorporation of microtubule subunits into fluxing kinetochore fibres. Nat Cell Biol 7(1):42–47. doi:ncb1207 [pii] 10.1038/ncb1207
Manning AL, Bakhoum SF, Maffini S, Correia-Melo C, Maiato H, Compton DA (2010) CLASP1, astrin and Kif2b form a molecular switch that regulates kinetochore-microtubule dynamics to promote mitotic progression and fidelity. EMBO J 29(20):3531–3543. doi:emboj2010230 [pii] 10.1038/emboj.2010.230
Maresca TJ, Groen AC, Gatlin JC, Ohi R, Mitchison TJ, Salmon ED (2009) Spindle assembly in the absence of a RanGTP gradient requires localized CPC activity. Curr Biol 19(14):1210–1215. doi:S0960-9822(09)01197-X [pii] 10.1016/j.cub.2009.05.061
Maurer SP, Fourniol FJ, Bohner G, Moores CA, Surrey T (2012) EBs recognize a nucleotide-dependent structural cap at growing microtubule ends. Cell 149(2):371–382. doi:S0092-8674(12)00341-8 [pii] 10.1016/j.cell.2012.02.049
Mayer TU, Kapoor TM, Haggarty SJ, King RW, Schreiber SL, Mitchison TJ (1999) Small molecule inhibitor of mitotic spindle bipolarity identified in a phenotype-based screen. Science 286(5441):971–974. doi:7948 [pii]
Mayr MI, Hummer S, Bormann J, Gruner T, Adio S, Woehlke G, Mayer TU (2007) The human kinesin Kif18A is a motile microtubule depolymerase essential for chromosome congression. Curr Biol 17(6):488–498. doi:S0960-9822(07)01013-5 [pii] 10.1016/j.cub.2007.02.036
Mazumdar M, Misteli T (2005) Chromokinesins: multitalented players in mitosis. Trends Cell Biol 15(7):349–355. doi:10.1016/j.tcb.2005.05.006
McHedlishvili N, Wieser S, Holtackers R, Mouysset J, Belwal M, Amaro AC, Meraldi P (2012) Kinetochores accelerate centrosome separation to ensure faithful chromosome segregation. J Cell Sci 125(Pt 4):906–918. doi:jcs.091967 [pii] 10.1242/jcs.091967
Meunier S, Shvedunova M, Van Nguyen N, Avila L, Vernos I, Akhtar A (2015) An epigenetic regulator emerges as microtubule minus-end binding and stabilizing factor in mitosis. Nat Commun 6:7889. doi:ncomms8889 [pii] 10.1038/ncomms8889
Meunier S, Vernos I (2011) K-fibre minus ends are stabilized by a RanGTP-dependent mechanism essential for functional spindle assembly. Nat Cell Biol 13(12):1406–1414. doi:ncb2372 [pii] 10.1038/ncb2372
Meunier S, Vernos I (2012) Microtubule assembly during mitosis – from distinct origins to distinct functions? J Cell Sci 125(Pt 12):2805–2814. doi:jcs.092429 [pii] 10.1242/jcs.092429
Mimori-Kiyosue Y, Shiina N, Tsukita S (2000) The dynamic behavior of the APC-binding protein EB1 on the distal ends of microtubules. Curr Biol 10(14):865–868. doi:S0960-9822(00)00600-X [pii]
Mishima M, Kaitna S, Glotzer M (2002) Central spindle assembly and cytokinesis require a kinesin-like protein/RhoGAP complex with microtubule bundling activity. Dev Cell 2(1):41–54. doi:S1534580701001101 [pii]
Mishra RK, Chakraborty P, Arnaoutov A, Fontoura BM, Dasso M (2010) The Nup107-160 complex and gamma-TuRC regulate microtubule polymerization at kinetochores. Nat Cell Biol 12(2):164–169. doi:ncb2016 [pii] 10.1038/ncb2016
Mitchison T, Kirschner M (1984) Dynamic instability of microtubule growth. Nature 312(5991):237–242
Mitchison TJ (2012) The proliferation rate paradox in antimitotic chemotherapy. Mol Biol Cell 23(1):1–6. doi:23/1/1 [pii] 10.1091/mbc.E10-04-0335
Mollinari C, Kleman JP, Jiang W, Schoehn G, Hunter T, Margolis RL (2002) PRC1 is a microtubule binding and bundling protein essential to maintain the mitotic spindle midzone. J Cell Biol 157(7):1175–1186. doi:10.1083/jcb.200111052, jcb.200111052 [pii]
Moudjou M, Bordes N, Paintrand M, Bornens M (1996) gamma-Tubulin in mammalian cells: the centrosomal and the cytosolic forms. J Cell Sci 109(Pt 4):875–887
Mountain V, Simerly C, Howard L, Ando A, Schatten G, Compton DA (1999) The kinesin-related protein, HSET, opposes the activity of Eg5 and cross-links microtubules in the mammalian mitotic spindle. J Cell Biol 147(2):351–366
Needleman DJ, Groen A, Ohi R, Maresca T, Mirny L, Mitchison T (2010) Fast microtubule dynamics in meiotic spindles measured by single molecule imaging: evidence that the spindle environment does not stabilize microtubules. Mol Biol Cell 21(2):323–333. doi:E09-09-0816 [pii] 10.1091/mbc.E09-09-0816
Noatynska A, Gotta M, Meraldi P (2012) Mitotic spindle (DIS)orientation and DISease: cause or consequence? J Cell Biol 199(7):1025–1035. doi:jcb.201209015 [pii] 10.1083/jcb.201209015
Ohba T, Nakamura M, Nishitani H, Nishimoto T (1999) Self-organization of microtubule asters induced in Xenopus egg extracts by GTP-bound Ran. Science 284(5418):1356–1358
Paweletz N (2001) Walther Flemming: pioneer of mitosis research. Nat Rev Mol Cell Biol 2(1):72–75. doi:10.1038/35048077 35048077 [pii]
Petry S, Groen AC, Ishihara K, Mitchison TJ, Vale RD (2013) Branching microtubule nucleation in Xenopus egg extracts mediated by augmin and TPX2. Cell 152(4):768–777. doi:S0092-8674(13)00015-9 [pii] 10.1016/j.cell.2012.12.044
Petry S, Pugieux C, Nedelec FJ, Vale RD (2011) Augmin promotes meiotic spindle formation and bipolarity in Xenopus egg extracts. Proc Natl Acad Sci U S A 108(35):14473–14478. doi:1110412108 [pii] 10.1073/pnas.1110412108
Piehl M, Tulu US, Wadsworth P, Cassimeris L (2004) Centrosome maturation: measurement of microtubule nucleation throughout the cell cycle by using GFP-tagged EB1. Proc Natl Acad Sci U S A 101(6):1584–1588. doi:10.1073/pnas.0308205100 0308205100 [pii]
Pinyol R, Scrofani J, Vernos I (2013) The role of NEDD1 phosphorylation by Aurora A in chromosomal microtubule nucleation and spindle function. Curr Biol 23(2):143–149. doi:S0960-9822(12)01390-5 [pii] 10.1016/j.cub.2012.11.046
Raff EC, Fackenthal JD, Hutchens JA, Hoyle HD, Turner FR (1997) Microtubule architecture specified by a beta-tubulin isoform. Science 275(5296):70–73
Reboutier D, Troadec MB, Cremet JY, Chauvin L, Guen V, Salaun P, Prigent C (2013) Aurora A is involved in central spindle assembly through phosphorylation of Ser 19 in P150Glued. J Cell Biol 201(1):65–79. doi:jcb.201210060 [pii] 10.1083/jcb.201210060
Rieder CL (1981) The structure of the cold-stable kinetochore fiber in metaphase PtK1 cells. Chromosoma 84(1):145–158
Rieder CL (2005) Kinetochore fiber formation in animal somatic cells: dueling mechanisms come to a draw. Chromosoma 114(5):310–318. doi:10.1007/s00412-005-0028-2
Roberts AJ, Kon T, Knight PJ, Sutoh K, Burgess SA (2013) Functions and mechanics of dynein motor proteins. Nat Rev Mol Cell Biol 14(11):713–726. doi:nrm3667 [pii] 10.1038/nrm3667
Salmela AL, Kallio MJ (2013) Mitosis as an anti-cancer drug target. Chromosoma 122(5):431–449. doi:10.1007/s00412-013-0419-8
Sampath SC, Ohi R, Leismann O, Salic A, Pozniakovski A, Funabiki H (2004) The chromosomal passenger complex is required for chromatin-induced microtubule stabilization and spindle assembly. Cell 118(2):187–202. doi:10.1016/j.cell.2004.06.026 S0092867404006178 [pii]
Sardon T, Peset I, Petrova B, Vernos I (2008) Dissecting the role of Aurora A during spindle assembly. EMBO J 27(19):2567–2579. doi:emboj2008173 [pii] 10.1038/emboj.2008.173
Saxton WM, McIntosh JR (1987) Interzone microtubule behavior in late anaphase and telophase spindles. J Cell Biol 105(2):875–886
Scrofani J, Sardon T, Meunier S, Vernos I (2015) Microtubule nucleation in mitosis by a RanGTP-dependent protein complex. Curr Biol 25(2):131–140. doi:S0960-9822(14)01486-9 [pii] 10.1016/j.cub.2014.11.025
Sdelci S, Schutz M, Pinyol R, Bertran MT, Regue L, Caelles C, Vernos I, Roig J (2012) Nek9 phosphorylation of NEDD1/GCP-WD contributes to Plk1 control of gamma-tubulin recruitment to the mitotic centrosome. Curr Biol 22(16):1516–1523. doi:S0960-9822(12)00672-0 [pii] 10.1016/j.cub.2012.06.027
Sharp DJ, Ross JL (2012) Microtubule-severing enzymes at the cutting edge. J Cell Sci 125(Pt 11):2561–2569. doi:jcs.101139 [pii] 10.1242/jcs.101139
Shrestha RL, Draviam VM (2013) Lateral to end-on conversion of chromosome-microtubule attachment requires kinesins CENP-E and MCAK. Curr Biol 23(16):1514–1526. doi:S0960-9822(13)00765-3 [pii] 10.1016/j.cub.2013.06.040
Sillje HH, Nagel S, Korner R, Nigg EA (2006) HURP is a Ran-importin beta-regulated protein that stabilizes kinetochore microtubules in the vicinity of chromosomes. Curr Biol 16(8):731–742. doi:S0960-9822(06)01277-2 [pii] 10.1016/j.cub.2006.02.070
Sirajuddin M, Rice LM, Vale RD (2014) Regulation of microtubule motors by tubulin isotypes and post-translational modifications. Nat Cell Biol 16(4):335–344. doi:ncb2920 [pii] 10.1038/ncb2920
Tanenbaum ME, Medema RH (2010) Mechanisms of centrosome separation and bipolar spindle assembly. Dev Cell 19(6):797–806. doi:S1534-5807(10)00538-1 [pii] 10.1016/j.devcel.2010.11.011
Teixido-Travesa N, Roig J, Luders J (2012) The where, when and how of microtubule nucleation – one ring to rule them all. J Cell Sci 125(Pt 19):4445–4456. doi:jcs.106971 [pii] 10.1242/jcs.106971
Teixido-Travesa N, Villen J, Lacasa C, Bertran MT, Archinti M, Gygi SP, Caelles C, Roig J, Luders J (2010) The gammaTuRC revisited: a comparative analysis of interphase and mitotic human gammaTuRC redefines the set of core components and identifies the novel subunit GCP8. Mol Biol Cell 21(22):3963–3972. doi:E10-05-0408 [pii] 10.1091/mbc.E10-05-0408
Tirnauer JS, Canman JC, Salmon ED, Mitchison TJ (2002) EB1 targets to kinetochores with attached, polymerizing microtubules. Mol Biol Cell 13(12):4308–4316. doi:10.1091/mbc.E02-04-0236
Topham CH, Taylor SS (2013) Mitosis and apoptosis: how is the balance set? Curr Opin Cell Biol 25(6):780–785. doi:S0955-0674(13)00117-8 [pii] 10.1016/j.ceb.2013.07.003
Torosantucci L, De Luca M, Guarguaglini G, Lavia P, Degrassi F (2008) Localized RanGTP accumulation promotes microtubule nucleation at kinetochores in somatic mammalian cells. Mol Biol Cell 19(5):1873–1882. doi:E07-10-1050 [pii] 10.1091/mbc.E07-10-1050
Tournebize R, Popov A, Kinoshita K, Ashford AJ, Rybina S, Pozniakovsky A, Mayer TU, Walczak CE, Karsenti E, Hyman AA (2000) Control of microtubule dynamics by the antagonistic activities of XMAP215 and XKCM1 in Xenopus egg extracts. Nat Cell Biol 2(1):13–19. doi:10.1038/71330
Tsai MY, Wiese C, Cao K, Martin O, Donovan P, Ruderman J, Prigent C, Zheng Y (2003) A Ran signalling pathway mediated by the mitotic kinase Aurora A in spindle assembly. Nat Cell Biol 5(3):242–248. doi:10.1038/ncb936, ncb936 [pii]
Tseng BS, Tan L, Kapoor TM, Funabiki H (2010) Dual detection of chromosomes and microtubules by the chromosomal passenger complex drives spindle assembly. Dev Cell 18(6):903–912. doi:S1534-5807(10)00254-6 [pii] 10.1016/j.devcel.2010.05.018
Tulu US, Fagerstrom C, Ferenz NP, Wadsworth P (2006) Molecular requirements for kinetochore-associated microtubule formation in mammalian cells. Curr Biol 16(5):536–541. doi:S0960-9822(06)01127-4 [pii] 10.1016/j.cub.2006.01.060
Tulu US, Rusan NM, Wadsworth P (2003) Peripheral, non-centrosome-associated microtubules contribute to spindle formation in centrosome-containing cells. Curr Biol 13(21):1894–1899. doi:S0960982203007449 [pii]
Uehara R, Goshima G (2010) Functional central spindle assembly requires de novo microtubule generation in the interchromosomal region during anaphase. J Cell Biol 191(2):259–267. doi:jcb.201004150 [pii] 10.1083/jcb.201004150
Uehara R, Nozawa RS, Tomioka A, Petry S, Vale RD, Obuse C, Goshima G (2009) The augmin complex plays a critical role in spindle microtubule generation for mitotic progression and cytokinesis in human cells. Proc Natl Acad Sci U S A 106(17):6998–7003. doi:0901587106 [pii] 10.1073/pnas.0901587106
van den Wildenberg SM, Tao L, Kapitein LC, Schmidt CF, Scholey JM, Peterman EJ (2008) The homotetrameric kinesin-5 KLP61F preferentially crosslinks microtubules into antiparallel orientations. Curr Biol 18(23):1860–1864. doi:S0960-9822(08)01395-X [pii] 10.1016/j.cub.2008.10.026
Vanneste D, Ferreira V, Vernos I (2011) Chromokinesins: localization-dependent functions and regulation during cell division. Biochem Soc Trans 39(5):1154–1160. doi:BST0391154 [pii] 10.1042/BST0391154
Vasquez RJ, Gard DL, Cassimeris L (1994) XMAP from Xenopus eggs promotes rapid plus end assembly of microtubules and rapid microtubule polymer turnover. J Cell Biol 127(4):985–993
Wade RH (2009) On and around microtubules: an overview. Mol Biotechnol 43(2):177–191. doi:10.1007/s12033-009-9193-5
Wainman A, Buster DW, Duncan T, Metz J, Ma A, Sharp D, Wakefield JG (2009) A new Augmin subunit, Msd1, demonstrates the importance of mitotic spindle-templated microtubule nucleation in the absence of functioning centrosomes. Genes Dev 23(16):1876–1881. doi:23/16/1876 [pii] 10.1101/gad.532209
Walczak CE, Gayek S, Ohi R (2013) Microtubule-depolymerizing kinesins. Annu Rev Cell Dev Biol 29:417–441. doi:10.1146/annurev-cellbio-101512-122345
Walczak CE, Mitchison TJ, Desai A (1996) XKCM1: a Xenopus kinesin-related protein that regulates microtubule dynamics during mitotic spindle assembly. Cell 84(1):37–47. doi:S0092-8674(00)80991-5 [pii]
Walczak CE, Verma S, Mitchison TJ (1997) XCTK2: a kinesin-related protein that promotes mitotic spindle assembly in Xenopus laevis egg extracts. J Cell Biol 136(4):859–870
Wang H, Vo T, Hajar A, Li S, Chen X, Parissenti AM, Brindley DN, Wang Z (2014) Multiple mechanisms underlying acquired resistance to taxanes in selected docetaxel-resistant MCF-7 breast cancer cells. BMC Cancer 14:37. doi:1471-2407-14-37 [pii] 10.1186/1471-2407-14-37
Waters JC, Mitchison TJ, Rieder CL, Salmon ED (1996) The kinetochore microtubule minus-end disassembly associated with poleward flux produces a force that can do work. Mol Biol Cell 7(10):1547–1558
White EA, Glotzer M (2012) Centralspindlin: at the heart of cytokinesis. Cytoskeleton (Hoboken) 69(11):882–892. doi:10.1002/cm.21065
Wiese C, Zheng Y (2000) A new function for the gamma-tubulin ring complex as a microtubule minus-end cap. Nat Cell Biol 2(6):358–364. doi:10.1038/35014051
Wittmann T, Wilm M, Karsenti E, Vernos I (2000) TPX2, A novel xenopus MAP involved in spindle pole organization. J Cell Biol 149(7):1405–1418
Wollman R, Cytrynbaum EN, Jones JT, Meyer T, Scholey JM, Mogilner A (2005) Efficient chromosome capture requires a bias in the ‘search-and-capture’ process during mitotic-spindle assembly. Curr Biol 15(9):828–832. doi:S0960-9822(05)00284-8 [pii] 10.1016/j.cub.2005.03.019
Yingling J, Youn YH, Darling D, Toyo-Oka K, Pramparo T, Hirotsune S, Wynshaw-Boris A (2008) Neuroepithelial stem cell proliferation requires LIS1 for precise spindle orientation and symmetric division. Cell 132(3):474–486. doi:S0092-8674(08)00126-8 [pii] 10.1016/j.cell.2008.01.026
Zanic M, Widlund PO, Hyman AA, Howard J (2013) Synergy between XMAP215 and EB1 increases microtubule growth rates to physiological levels. Nat Cell Biol 15(6):688–693. doi:ncb2744 [pii] 10.1038/ncb2744
Zhang C, Hughes M, Clarke PR (1999) Ran-GTP stabilises microtubule asters and inhibits nuclear assembly in Xenopus egg extracts. J Cell Sci 112(Pt 14):2453–2461
Zhang R, Alushin GM, Brown A, Nogales E (2015) Mechanistic origin of microtubule dynamic instability and its modulation by EB proteins. Cell 162(4):849–859. doi:S0092-8674(15)00849-1 [pii] 10.1016/j.cell.2015.07.012
Zhang X, Chen Q, Feng J, Hou J, Yang F, Liu J, Jiang Q, Zhang C (2009) Sequential phosphorylation of Nedd1 by Cdk1 and Plk1 is required for targeting of the gammaTuRC to the centrosome. J Cell Sci 122(Pt 13):2240–2251. doi:jcs.042747 [pii] 10.1242/jcs.042747
Zhu H, Coppinger JA, Jang CY, Yates JR 3rd, Fang G (2008) FAM29A promotes microtubule amplification via recruitment of the NEDD1-gamma-tubulin complex to the mitotic spindle. J Cell Biol 183(5):835–848. doi:jcb.200807046 [pii] 10.1083/jcb.200807046
Acknowledgments
We apologize to all scientists whose primary work could not be cited due to space limitations. We thank all members of the Vernos lab for helpful comments and critical reading of this manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer-Verlag Wien
About this chapter
Cite this chapter
Meunier, S., Vernos, I. (2016). Microtubule Organization in Mitotic Cells. In: Lüders, J. (eds) The Microtubule Cytoskeleton. Springer, Vienna. https://doi.org/10.1007/978-3-7091-1903-7_1
Download citation
DOI: https://doi.org/10.1007/978-3-7091-1903-7_1
Published:
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-1901-3
Online ISBN: 978-3-7091-1903-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)