Abstract
Over the last decades, the study of congenital heart disease (CHD) has benefited from various model systems and the development of molecular biological techniques enabling the analysis of single gene as well as global effects. In this chapter, we first describe different models including CHD patients and their families, animal models ranging from invertebrates to mammals, and various cell culture systems. Moreover, techniques to experimentally manipulate these models are discussed. Secondly, we introduce cardiac phenotyping technologies comprising the analysis of mouse and cell culture models, live imaging of cardiogenesis, and histological methods for fixed hearts. Finally, the most important and latest molecular biotechniques are described. These include genotyping technologies, different applications of next-generation sequencing, as well as the analysis of the transcriptome, epigenome, proteome, and metabolome. In summary, the models and technologies presented in this chapter are essential to study the function and development of the heart and to understand the molecular pathways underlying CHD.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Genotyping technologies
- Next-generation sequencing
- Transcriptome
- Epigenome
- Proteome
- Metabolome
- Animal models
- Nematode
- Caenorhabditis elegans
- Fruit fly
- Drosophila melanogaster
- Mouse
- Mus musculus
- Rat
- Rattus norvegicus
- Zebrafish
- NKX2-5
- Clawed frog
- Xenopus laevis
- Chicken
- Gallus gallus
- Embryonic stem cells
- ES cells
- Cell culture
- Cardiomyocytes
- Induced pluripotent ES cells
- iPSCs
- Transcription activator-like effector nucleases
- TALENs
- Zinc finger nucleases
- ZFN
- CRISPR
- Morpholino oligonucleotides
- Imaging
- Phenotyping
- Micro-computed tomography
- Micro-CT
- Magnetic resonance imaging
- MRI
- Histological analysis
- In situ hybridization
- Immunohistochemistry
- Fluorescence-activated cell sorting
- FACS
- Single-nucleotide polymorphisms
- SNPs
- Sanger sequencing
- Fluorescence in situ hybridization
- FISH
- Array comparative genomic hybridization
- Array-CGH
- NGS
- Genome-wide association studies
- GWAS
- RNA-seq
- Chromatin immunoprecipitation
- ChIP
- ChIP-seq
- DNA methylation
- MBD
- ATAC-seq
- CLIP
- PAR-CLIP
- Mass spectrometry
- MS
- Nuclear magnetic resonance spectroscopy
- NMR
- SILAC
- Yeast-two-hybrid
1 Introduction
Understanding the genetic alterations and molecular pathways underlying congenital heart disease (CHD) is essential to develop novel therapeutic strategies. Besides genomic mutations, CHD is characterized by multiple changes in epigenetic marks as well as in the expression and modification of RNAs and proteins. Some of these changes have strong effects on the protein structure, function, or localization, while others result in more subtle differences. Over the last decade, advanced high-throughput technologies have been developed that allow the study of CHD from single locus effects to the global level. Here, we describe the most important model systems and techniques to explore the changes in the different regulatory layers affecting cardiac function and development.
2 Model Systems
2.1 Animal Models
A large variety of animal models are used in cardiovascular research ranging from invertebrates like the nematode Caenorhabditis elegans and the fruit fly Drosophila melanogaster to mammals like the mouse Mus musculus and the laboratory rat Rattus norvegicus. Although it has no heart or vascular system, C. elegans has become a useful model that is easy to culture and manipulate by RNA interference or mutation. Its body wall muscle cells are well suited to gain insights into human cardiomyocytes; as they form, striated muscle and many structures and proteins like sarcomeric components are highly conserved [1]. The fruit fly is characterized by a simple tube-shaped heart, also called the dorsal vessel, which consists of a single layer of cardiomyocytes and pumps the hemolymph around the body. Drosophila can be used easily for genetic screens and for crossing experiments, for which several large collections of mutant stocks are available. Due to few genetic redundancies, it is a widely used model of regulatory pathways and essential players of heart development [2, 3]. For example, the important role of the cardiac transcription factor NKX2-5 (tinman) was first demonstrated in Drosophila [4]. However, the fly might not be well suited for studies of buffering effects, which are essential for the understanding of human cardiac disease [3].
The zebrafish (Danio rerio) has a two-chambered heart, and the transparency of the developing embryo allows the in vivo study of heart size, shape, and function [5]. Due to its small size, fecundity, and brief generation time, the zebrafish can easily be used for forward genetic perturbation screens [6] and moreover, allows the study of otherwise lethal disturbances because no functional cardiovascular system is required during embryogenesis [3]. In addition, the zebrafish represents a valuable model for cardiac muscle regeneration, as it has the ability to replace massive sections of damaged myocardium [7]. In contrast to the zebrafish, the African clawed frog (Xenopus laevis) has a three-chambered heart with two atria and one ventricle. Its large embryo allows surgical manipulations of the developing heart and moreover, is useful for genetic screens [8]. Like the human, the chicken (Gallus gallus) develops a four-chambered heart and is a powerful model for experimental embryology as its imaging during development requires no surgical procedures [9].
The two essential mammalian animal models are the mouse and the laboratory rat. The cardiovascular system of the latter shares a high similarity to human physiology and is widely used for pharmaceutical testing [10]. Recently, the genetic manipulation limits for the rat have been overcome by the development of gene-targeting approaches for gene knockout and replacement [3]. However, the mouse remains the most important model for genetic studies, and transgenic mice have become a valuable human CHD pathology model [11]. The International Knockout Mouse Consortium aims to generate a major resource of knockout mouse embryonic stem cells (ES cells). So far, more than 17,400 mutant murine embryonic stem cell clones have been generated, which provide the opportunity of systematic screens of gene functions [12]. Currently, more than 500 genes are known to be causative for heart defects when mutated in mice, while only about 50 CHD genes are known in humans [13]. In addition, mouse strains with different genetic backgrounds offer the opportunity to study genetic buffering effects, as recently shown for NKX2-5 [14]. Finally, the mouse is also a useful model for gene expression studies, and several large-scale projects aim to systematically determine gene expression patterns during mouse embryogenesis [15].
2.2 Cell Culture Models
The heart comprises a mixture of different cell types including cardiac fibroblasts and cardiomyocytes. In vitro studies using largely homogeneous cell populations enable the analysis of spatiotemporal changes and distinct molecular pathways. Moreover, cell culture models are easier to manipulate than animal models, can be sorted for surface markers, and provide the opportunity to produce enough material for downstream experiments.
Isolated cardiomyocytes cultured in primary cell culture are one of the established models to investigate cardiac function and closely reflect in vivo physiology. In addition, several stable, immortalized cell lines have been generated. The HL-1 cell line, derived from mouse atrial cardiomyocytes, maintains the ability to contract, shows a gene expression pattern similar to adult cardiomyocytes [16], and can be used in tissue engineering applications (3D cell culture) [17]. In contrast, the H9C2 cell line, which was established from embryonic rat heart tissue, shows cardiac as well as skeletal muscle properties [18]. To study myogenesis, the mouse myogenic cell line C2C12 is frequently used, since it is capable of differentiation into myotubes [19]. Furthermore, embryonic cell lines like the mouse P19 embryonic stem (ES) cells [20] and the human H1 or hES2 ES cells [21] can be differentiated into mesodermal cells including cardiac and skeletal muscle cells and have been used for the quantification of cardiomyocyte differentiation [22]. Finally, the generation of cardiomyocytes from induced pluripotent stem cells (iPSCs) and the direct conversion of fibroblasts into cardiomyocytes have recently been established as additional approaches that enable the study of patient-specific cells and provide novel perspectives for regenerative medicine [23, 24].
2.3 Patients with CHD
Patients with CHD and their families are a unique resource to gain insights into cardiac functional properties and molecular pathways. For example, linkage analysis in CHD families has led to the identification of single-gene defects like mutations in NKX2-5 [25] and GATA4 (GATA binding protein 4) [26]. Several national registries like the CONCOR registry of the Netherlands or the National Register for Congenital Heart Defects in Germany aim to establish comprehensive collections of biomaterial like blood or cardiac biopsies. While genomic DNA isolated from blood can be used for genetic studies, cardiac biopsies offer the opportunity to study gene expression profiles and epigenetic mechanisms, providing insights into regulatory relationships. Besides the direct analysis of patient material, the generation of patient-specific iPSCs offers a new and very promising possibility to study human diseases [27]. For example, the power of iPSCs has already been shown for the analysis of congenital arrhythmia and malformations [28, 29].
2.4 Techniques to Induce Perturbations
To enable the study of cause-effect relationships in different model systems, in vivo gene targeting techniques play an essential role. For example, the generation of designed mouse mutants relies on gene targeting in ES cells. Representing appropriate genetic models of inherited diseases, knockout mice with a null allele in their germline often exhibit embryonic or early postnatal lethality [30]. To study cell type or stage-specific gene functions, a system based on the DNA recombinase Cre and its recognition sites (loxP) has been established [30]. Other faster methods for targeted genome editing, which are also used in cell culture models, include transcription activator-like effector nucleases (TALENs), zinc finger nucleases (ZFN), and the recently established clustered regulatory interspaced short palindromic repeat (CRISPR)/Cas-based RNA-guided DNA endonucleases [31]. Besides the targeted perturbation of selected genes in a reverse genetics approach, random mutation screens enable the discovery of novel gene functions in an unbiased manner. For example, treatment with the supermutagen N-ethyl-N-nitrosourea (ENU) is the most potent method to induce random point mutations throughout the mouse genome. The offspring of treated animals can then be screened for autosomal dominant or recessive phenotypes and used for the identification of the causal mutation in a forward genetics approach [32].
In addition to the perturbation of the genomic DNA sequence, knockdown of a gene at the transcriptional level provides another valuable approach for studying gene functions and can be used for large genome-scale high-throughput screens. RNA interference (RNAi) is a widely used approach for screens in cell culture systems, C. elegans and Drosophila [33], while the most common antisense knockdown technique in zebrafish are morpholino oligonucleotides (MOs) [34]. Moreover, the overexpression of genes, for example, by the transfection of expression vectors, not only allows the study of wild-type gene function but also enables the analysis of mutant alleles. Finally, knockdown in combination with overexpression can be applied to perform rescue experiments to demonstrate the specificity of the observed phenotype.
Besides genetic alterations, environmental influences also play an important role in the etiology of CHD. Thus, a variety of stress models have been established to analyze gene–environment interactions. For example, a maternal high fat diet increases the penetrance of CHD in heterozygous Cited2 knockout mice [35]. Moreover, a commonly used experimental model to induce hemodynamic stress in the mouse heart is the transverse aortic constriction (TAC), which results in a pressure overload-induced cardiac hypertrophy [36]. In cell culture models, hypertrophic agents like phenylephrine (PE) and endothelin 1 (ET1) are commonly used to study signaling pathways involved in cardiomyocyte hypertrophy [37].
3 Cardiac Phenotyping
3.1 Systematic Phenotyping of Mouse Models
The International Knockout Mouse Consortium has established thousands of mutant mouse models [12], whose characterization, archiving, and distribution are realized by phenotyping centers (mouse clinics) organized in large consortia like the International Mouse Phenotyping Consortium [38] or the European Mouse Disease Clinic (EUMODIC) [39]. The phenotyping pipelines systematically assess the stage of a potential embryonic lethality and a broad range of physiological parameters such as metabolic functions, fertility, behavior, body composition, and immune function in adult animals. Cardiac phenotyping includes the measurement of heart weight, electrocardiography, and echocardiography as well as histological analysis [38, 40].
3.2 Imaging Cardiogenesis in Live Animals
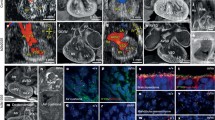
Embryogenesis is based on three fundamental processes, namely, growth, differentiation, and organization. Thus, in vivo imaging of these processes is important for understanding the structural formation and function of the heart and its molecular background. The main quantitative in vivo imaging techniques are optical imaging, ultrasound, micro-computed tomography (micro-CT), and magnetic resonance imaging (MRI) [41].
Confocal microscopy is a routine method to study the transparent zebrafish embryo, and multiple fluorescent transgenic lines have been established to label, for example, cardiac myocytes and endothelial cells [5]. Ultrasound is another inherently real-time imaging technique and is useful to analyze aberrant morphologies and hemodynamic phenotypes in developing embryos [41]. The small size and rapid motions of the embryo make ultrasound imaging challenging in mice and thus the hemodynamics of the dorsal aorta is often used as a surrogate for intracardiac hemodynamics. In contrast, avian embryos allow the direct measurement of chamber-specific blood flow and valve motions [41]. The highest possible imaging resolution is theoretically provided by micro-CT. However, X-rays are attenuated poorly by soft tissues and necessitate the application of contrast agents, which are mostly unsuitable for live embryonic imaging. Nevertheless, recent studies have evaluated different agents with a reduced toxicity [41]. Finally, MRI has mainly been used for the imaging of fixed murine embryos, as technical limitations hinder its application for live embryonic imaging. It offers excellent tissue contrast and the ability of three-dimensional image reconstruction [42], but high field strengths of 7–11 T and long acquisition times (6–24 h) are required for the high spatial resolution needed for embryonic imaging (25–50 μm) [41, 43]. Therefore, multiple embryos are often analyzed at once to increase the throughput of phenotyping. Live imaging applications of MRI with a reduced spatial resolution include the study of mouse embryos in utero and chicken embryos in ovo [41].
3.3 Histological Analysis of Fixed Hearts
Classical methods for the histological assessment of fixed hearts are hematoxylin and eosin (HE) staining for morphological inspection, in situ hybridization for transcript expression, and immunohistochemistry for protein expression. For the visualization of subcellular structures, electron microscopy is the gold standard [3].
To identify cell nuclei and cytoplasm, HE staining is the most widely used method and provides a general overview of the sample. Moreover, a large variety of special stains is available to visualize, for example, polysaccharides, glycoproteins, and glycolipids (periodic acid–Schiff; PAS stain) or collagen fibers (trichome staining) [44]. In situ hybridization is based on the binding of an approximately 20-bp-long oligonucleotide probe, usually labeled with digoxigenin or fluorescent dyes, to the complementary mRNA. In contrast, immunohistochemistry is based on the application of labeled antibodies. To reconstruct the 3D structure of embryonic hearts and to visualize cardiac gene expression patterns, techniques like high-resolution episcopic microscopy (HREM) and optical projection tomography (OPT) [45, 46] have proved to be very useful tools and enhance the phenotyping of mouse embryos. Finally, electron microscopes provide the highest resolution in the range of picometers [47], which is needed for studying the ultrastructure of a wide variety of biological and inorganic specimens.
3.4 Phenotyping of Cell Culture Models
The easy handling of cell culture models allows the application of various phenotyping techniques. Immunohistochemistry provides the opportunity to detect the subcellular localization of proteins within the cell under defined conditions. Moreover, co-staining with different antibodies allows visualization of protein–protein interactions and thus, is a useful method to study molecular signaling pathways. Fluorescence-activated cell sorting (FACS) combines specific immunostaining and single-cell analysis using flow cytometry. For example, in a heterogeneous cell population like blood, distinct cell types can be distinguished based on their expressed surface markers [48]. Other cell-based assays include the measurement of proliferation, viability, apoptosis, autophagy, production of reactive oxygen species (ROS), mitochondrial function, cell migration, and cytotoxicity of chemical compounds. Finally, cultures of beating cardiomyocytes are an important model for electrophysiology studies, and the patch clamp technique enables the measurement of ionic currents on isolated cells down to the single-channel level [49].
4 Molecular Biological Techniques
4.1 Genotyping Techniques
One of the first methods to identify disease genes in affected families was the mapping of genes relative to known genetic markers by linkage analysis. This method is based on the recombination between homologous chromosomes, which occurs randomly. Thus, two genomic positions are less likely to undergo recombination if they are located in close proximity to each other. Different genetic markers including single-nucleotide polymorphisms (SNPs), microsatellites (short repeat sequences of variable length), or restriction fragment length polymorphisms (RFLPs) can be used for linkage analysis. RFLPs are sequence polymorphisms that cause differences in enzymatic cleavage sites between alleles, resulting in DNA fragments of unequal lengths that can be detected by probe hybridization [50]. For the genotyping of SNPs, microsatellites, and other short variations, direct sequencing or denaturing high-performance liquid chromatography (DHPLC) has been frequently used. DHPLC is based on the formation of heteroduplexes between chromosomes and can identify the presence but not the exact position and nature of a mismatch [51].
The gold standard for direct DNA sequencing is the Sanger method, reaching read lengths of up to 1000 bp and a per-base sequencing accuracy as high as 99.999 % [52]. It is based on the incorporation of dideoxynucleotides (ddNTPs) into the DNA that act as specific chain-terminating inhibitors of the DNA polymerase [53]. The introduction of shotgun sequencing, fluorescent labeling, and capillary gel electrophoresis significantly increased the throughput of Sanger sequencing and enabled the deciphering of the complete human genome in 2001 [54, 55]. The sequencing biochemistry is performed in a cycle sequencing reaction, which is stochastically terminated by the incorporation of fluorescently labeled ddNTPs. This results in a mixture of end-labeled products, and the final sequence is determined by electrophoretic separation of the products and laser excitation of the four different fluorescent labels [52] (Fig. 18.1a).
Workflow of Sanger sequencing and two next-generation sequencing platforms. (a) Sanger sequencing. (b) Genome Sequencer FLX from Roche/454. (c) Genome Analyzer from Illumina. dNTP deoxynucleotide, ddNTP dideoxynucleotides, NGS next-generation sequencing, nts nucleotides, Pol DNA polymerase (Figure adapted from Etheridge [56], Shendure and Ji [52], and Mardis [57])
A variety of methods also have been developed for the detection of chromosomal abnormalities. Giemsa staining is a simple and rapid technique for conventional karyotyping and can identify many chromosomal changes including balanced chromosomal aberrations [58]. A higher resolution from tens of kilobases up to several megabases is offered by fluorescence in situ hybridization (FISH), which uses fluorescently labeled probes that hybridize to their complementary chromosomal sequences [58]. As an alternative to these microscopy-based methods, multiplex ligation-dependent probe amplification (MLPA) can be applied, which is based on a multiplexed PCR and can detect copy number changes of up to 50 different loci in parallel [58].
New possibilities for the analysis of genetic variations were provided by microarray-based genotyping, which offers high-resolution genome-wide variation detection and is based on the hybridization of a DNA sample to oligonucleotide probes that have been immobilized on a glass or silicon surface [59]. Array comparative genomic hybridization (array-CGH) is used to identify chromosomal aberrations by comparing a DNA sample to a reference sample. Moreover, DNA microarrays enable the analysis of disease-specific or even genome-wide SNP panels (SNP arrays) [58]. Thus, they allow the detection of known disease-causing mutations in individual patients or the identification of novel associations between SNPs and complex traits in genome-wide association studies (GWAS) [60].
4.2 Next-Generation Sequencing
The development of novel high-throughput sequencing technologies has revolutionized biomedical research. These next-generation sequencing (NGS) technologies, first introduced in 2005 [61, 62], have evolved rapidly, and the costs have been reduced from $1000 per megabase to less than $0.1 in 2014 [63]. Thus, it is much more cost efficient than Sanger sequencing ($500 per megabase) and allows a higher degree of parallelization [52]. In contrast to microarrays, NGS is not dependent on DNA hybridization to preselected probes, enabling the identification of novel variations at a single-base resolution without a priori sequence information.
Different NGS platforms have been established, and the companies Roche/454 (Fig. 18.1b), Illumina (Fig. 18.1c), and Life Technologies have set the standard for high-throughput sequencing [64]. Although their systems vary in their chemistry, they are all based on the principle of cyclic-array sequencing. Here, a dense array of DNA features is iteratively enzymatically sequenced combined with imaging-based data collection [52]. In general, a sequencing run generates reads that randomly cover the genome [65]. The coverage describes the average number of times a single base is read during a sequencing run. A higher number of sequence reads result in greater sequencing depth and thus, in higher sequence confidence. For example, within the 1000 Genomes Project, the coverage ranges from low (2–6×) for whole genome sequencing to high (50–100×) for exome sequencing [66].
For the sequencing of genomic DNA, three basic approaches are available [64]. Whole-genome sequencing allows the determination of all genomic variations but is relatively cost intensive. Here, useful alternatives are provided by whole exome and targeted re-sequencing approaches, which require sequence enrichment technologies such as array-based sequence capturing. Whole-exome sequencing enables the sequencing of almost all protein-coding regions (optionally including untranslated regions or long non-coding RNAs), often combined with a high coverage. When knowledge about possible candidate regions (e.g., genes, promoters, and enhancers) and disease pathways is already available, the targeted re-sequencing of these regions is a promising option. The selection of genomic targets for re-sequencing can be based on data from previous projects like sequencing analyses, GWAS, animal models, as well as publicly available databases [64]. Moreover, disease-specific Web resources like the CHDWiki [67] and the Cardiovascular Gene Annotation Initiative, which has annotated more than 4000 cardiovascular-associated proteins [68], provide useful information for candidate gene selection.
Several large cohorts of CHD patients already are under investigation by NGS [64]. For example, the Congenital Heart Disease Genetic Network Study established by the Pediatric Cardiac Genomics Consortium enrolled more than 3700 patients with a diverse range of CHD [69], and so far, whole-exome sequencing data for a subset of 362 patients and their parents is available [70]. Having a broader focus on undiagnosed children with developmental disorders, the Deciphering Developmental Disorders (DDD) study headed by the Wellcome Trust Sanger Institute aims to recruit 12,000 patients and their parents [71]. Recently, exome sequencing and array-CGH were performed for 1113 children and their parents, with CHD occurring in 11 % of the patients [72, 73].
4.3 Transcriptome and Epigenome Analysis
Both NGS and array-based technologies are extensively used for transcriptome and epigenome analysis. In addition, quantitative real-time PCR is a useful low- to medium-throughput application. The study of gene expression has been revolutionized by RNA sequencing (RNA-seq), which enables the discovery, profiling, and quantification of RNA transcripts across the entire transcriptome without prior knowledge about the probed sequences. Applications of RNA-seq comprise total RNA-seq (coding and non-coding RNA above a certain size), mRNA-seq (including mRNAs and long non-coding RNAs with a poly-A tail), and small RNA-seq (including microRNAs and other small non-coding RNAs). Novel applications of RNA-seq include de novo transcriptome assembly [74], single-cell transcriptomics [75], and tomography sequencing to determine spatially resolved transcription profiles in whole embryos or isolated organs [76].
A powerful technique for the genome-wide identification of protein–DNA interactions such as transcription factor binding sites or chromatin histone marks is chromatin immunoprecipitation (ChIP). In ChIP, the protein of interest is cross-linked to the DNA, either in cultured cells or in tissue samples. After cross-linking, the chromatin is sheared and an antibody is used to enrich for DNA fragments bound to the protein. Immunoprecipitation and reverse cross-linking isolate the DNA enriched in the binding sites, and finally, the enriched DNA fragments can further be analyzed by hybridization to microarrays (ChIP-chip) or NGS (ChIP-seq) [77, 78] (Fig. 18.2). If candidate target genes or potential sites are available, ChIP-qPCR represents an alternative strategy. To investigate the co-localization of proteins on the DNA, ChIP-reChIP (sequential ChIP) has been developed using two independent rounds of immunoprecipitation [80]. An alternative method used to map protein–genome interactions is DamID, which does not require the use of antibodies. This technique is based on the fusion of the protein of interest to Escherichia coli DNA adenine methyltransferase (dam) and the resulting methylation of adenines in DNA surrounding the native binding sites of the dam fusion partner. In most eukaryotes, adenine methylation does not occur endogenously. Thus, it provides a unique tag to mark protein interaction sites, which can further be identified by array hybridization or NGS [81].
Schematic representation of a chromatin immunoprecipitation (ChIP) experiment followed by microarray detection (ChIP-chip) or next-generation sequencing (ChIP-seq) (Figure adapted from Visel et al. [79])
In addition to histone modifications, DNA methylation occurring on cytosine residues in the context of CpG dinucleotides is also an important epigenetic mark. Altered DNA methylation has been shown to play a role in various diseases, including CHD [82]. Three methods are commonly used to detect genome-wide DNA methylation levels. Two techniques are based on the isolation of methylated DNA fragments by methylated DNA immunoprecipitation (MeDIP) or methyl-CpG binding domain-based (MBD) proteins. Subsequently, the enriched DNA fragments can be detected by arrays or NGS [83]. The third technique applies the treatment of DNA with sodium bisulfite, which converts all non-methylated cytosines to uracil. These will finally be detected as thymine residues, analogous to a C to T SNP, by, for example, pyrosequencing [84] or NGS.
Several techniques are available to assess chromatin structure and regulatory interactions. Chromatin that has lost its condensed structure is sensitive to cleavage by the DNase I enzyme (DNase I hypersensitive sites). Thus, the enzymatic degradation of DNA can be used to identify regions of open chromatin, representing cis-regulatory elements including promoters, enhancers, insulators, and silencers [85]. An alternative method to DNase-seq is the assay of transposase-accessible chromatin (ATAC-seq), which uses an engineered Tn5 transposase to cleave DNA in open chromatin and to integrate primer DNA sequences into the cleaved genomic DNA. Furthermore, a commonly used method to identify the exact positions of nucleosomes is the treatment with micrococcal nuclease (MNase), an endo-exonuclease that processively digests DNA until it is blocked, for example, by a nucleosome [86]. To study interactions between regulatory elements, including long-range interactions between different chromosomes, the chromosome conformation capture (3C) and various derivatives (4C, 5C, and Hi-C) have been developed. They are all based on the cross-linking of interacting DNA fragments and their subsequent restriction digest [87]. Using an additional ChIP step, chromatin analysis by paired-end tag sequencing (ChIA-PET) allows the identification of long-range interactions mediated by target proteins of interest [88].
The interaction of RNAs and proteins is also an important layer for the co-transcriptional and posttranscriptional regulation of gene expression. Genome-wide protein–RNA interaction can be identified based on ultraviolet cross-linking and immunoprecipitation (CLIP). To reach a base-pair resolution, this method was further developed to photoactivatable ribonucleoside-enhanced CLIP (PAR-CLIP), which relies on the incorporation of photoactivatable nucleotide analogues into the RNA. Here, reverse transcription results in a T to C base transition at the cross-link site, detectable as SNPs in the subsequent NGS analysis. However, the need to incorporate photoactivatable nucleotides restricts PAR-CLIP to cultured cells [89]. Finally, a highly sensitive method for the general profiling of RNA-induced silencing complexes (RISC) and individual microRNA target identification is RISC-seq [90].
4.4 Proteome and Metabolome Analysis
The quantitative and qualitative large-scale study of proteins (proteomics) and small-molecule metabolites such as alcohols, amino acids, and nucleotides (metabolomics) has undergone great developments over recent years. However, these new technologies have only begun to be applied in CHD research [91, 92], where they have the potential to boost our knowledge of molecular mechanisms underlying heart disease from the pharmacological viewpoint and to enable the discovery of novel biomarkers.
The core technologies for both proteome and metabolome studies are mass spectrometry (MS) and nuclear magnetic resonance (NMR) spectroscopy. Most approaches are based on the analysis of peptides, which are frequently generated by enzymatic digestion of proteins. A key step for MS analysis is the selection and enrichment of the proteins/peptides of interest, which can be achieved by subcellular fractioning (e.g., membrane enrichment, nucleus precipitation, or mitochondria separation), by co-immunoprecipitation (e.g., for a protein and its interaction partners), or by enrichment for proteins with particular modifications (e.g., phosphorylation). Furthermore, the development of stable isotope labeling enabled the generation of relative quantitative information [93]. An important technique that can be applied to cell culture studies and more recently, also to studying mouse and drosophila models [94, 95] is stable isotope labeling by amino acids in cell culture (SILAC). Here, two cell populations are cultured in the presence of heavy or light amino acid (e.g., lysine or arginine, respectively) and are further combined for MS analysis [96]. In addition to the metabolic labeling used in SILAC, other methods have been established, including chemical (ICAT and iTRAQ) [97, 98] or enzymatic labeling (18O) [99].
A common method in metabolomics is NMR, which in contrast to MS does not require analyte separation and allows the recovery of the sample for further analyses. It can provide detailed information on the molecular structure of compounds found in complex mixtures like biofluids as well as cell and tissue extracts. NMR offers a high analytical reproducibility and easy sample preparation but is relatively insensitive in comparison to MS [100].
Methods suitable for the high-throughput analysis of protein–protein interactions are the yeast-two-hybrid (Y2H) and the mammalian-two-hybrid (M2H) systems. Both are based on the expression of the two proteins of interest, one fused to the DNA-binding domain and the other to the transactivation domain of a transcription factor, typically Gal4. The binding of the two proteins leads to the complementation of the TF, which activates the expression of a reporter gene (e.g., LacZ). For example, Y2H experiments have been used to identify a large and highly connected network comprising over 3000 interactions between 1705 human proteins [101]. Moreover, a M2H study provided a map of physical interactions within 762 human and 877 mouse DNA-binding transcription factors [102]. In addition to the two-hybrid systems, peptide microarrays have been employed to study protein–protein interactions [103]. However, they have been implemented much slower than DNA arrays due to technical challenges including the high-throughput and economic synthesis of peptides.
An overview of the various molecular biological techniques to study the different regulatory layers that control the gene and protein expression is given in Fig 18.3.
Overview of various molecular biological techniques to study the different regulatory layers controlling gene and protein expression (Figure adapted from Lara-Pezzi et al. [104])
Conclusion
In this chapter, we described various model systems and biotechniques to study the different regulatory levels affecting congenital heart defects. In particular, the application of NGS techniques has revolutionized biomedical research and is still rapidly developing, enabling its application to a wide range of scientific questions. Thus, these high-throughput techniques will enhance our understanding of CHD and will hopefully accelerate the development of novel therapeutic and preventive strategies.
References
Benian GM, Epstein HF (2011) Caenorhabditis elegans muscle: a genetic and molecular model for protein interactions in the heart. Circ Res 109:1082–1095
Reim I, Frasch M (2010) Genetic and genomic dissection of cardiogenesis in the Drosophila model. Pediatr Cardiol 31:325–334
Sperling SR (2011) Systems biology approaches to heart development and congenital heart disease. Cardiovasc Res 91:269–278
Bodmer R (1993) The gene tinman is required for specification of the heart and visceral muscles in Drosophila. Development 118:719–729
Schoenebeck JJ, Yelon D (2007) Illuminating cardiac development: advances in imaging add new dimensions to the utility of zebrafish genetics. Semin Cell Dev Biol 18:27–35
Molina G, Vogt A, Bakan A et al (2009) Zebrafish chemical screening reveals an inhibitor of Dusp6 that expands cardiac cell lineages. Nat Chem Biol 5:680–687
Major RJ, Poss KD (2007) Zebrafish heart regeneration as a model for cardiac tissue repair. Drug Discov Today Dis Models 4:219–225
Warkman AS, Krieg PA (2007) Xenopus as a model system for vertebrate heart development. Semin Cell Dev Biol 18:46–53
Kain KH, Miller JWI, Jones-Paris CR et al (2014) The chick embryo as an expanding experimental model for cancer and cardiovascular research. Dev Dyn 243:216–228
Gill TJ, Smith GJ, Wissler RW et al (1989) The rat as an experimental animal. Science 245:269–276
Snider P, Conway SJ (2011) Probing human cardiovascular congenital disease using transgenic mouse models. Prog Mol Biol Transl Sci 100:83–110
Bradley A, Anastassiadis K, Ayadi A et al (2012) The mammalian gene function resource: the International Knockout Mouse Consortium. Mamm Genome 23:580–586
Andersen TA, Troelsen Kde LL, Larsen LA (2014) Of mice and men: molecular genetics of congenital heart disease. Cell Mol Life Sci 71:1327–1352
Winston JB, Erlich JM, Green CA et al (2010) Heterogeneity of genetic modifiers ensures normal cardiac development. Circulation 121:1313–1321
Siddiqui AS, Khattra J, Delaney AD et al (2005) A mouse atlas of gene expression: large-scale digital gene-expression profiles from precisely defined developing C57BL/6J mouse tissues and cells. Proc Natl Acad Sci U S A 102:18485–18490
Claycomb WC, Lanson NA, Stallworth BS et al (1998) HL-1 cells: a cardiac muscle cell line that contracts and retains phenotypic characteristics of the adult cardiomyocyte. Proc Natl Acad Sci U S A 95:2979–2984
Gonnerman EA, Kelkhoff DO, McGregor LM, Harley BAC (2012) The promotion of HL-1 cardiomyocyte beating using anisotropic collagen-GAG scaffolds. Biomaterials 33:8812–8821
Kimes BW, Brandt BL (1976) Properties of a clonal muscle cell line from rat heart. Exp Cell Res 98:367–381
Yaffe D, Saxel O (1977) Serial passaging and differentiation of myogenic cells isolated from dystrophic mouse muscle. Nature 270:725–727
McBurney MW, Jones-Villeneuve EM, Edwards MK, Anderson PJ (1982) Control of muscle and neuronal differentiation in a cultured embryonal carcinoma cell line. Nature 299:165–167
Yang L, Soonpaa MH, Adler ED et al (2008) Human cardiovascular progenitor cells develop from a KDR+ embryonic-stem-cell-derived population. Nature 453:524–528
Moore JC, Spijker R, Martens AC et al (2004) A P19Cl6 GFP reporter line to quantify cardiomyocyte differentiation of stem cells. Int J Dev Biol 48:47–55
Dambrot C, Passier R, Atsma D et al (2011) Cardiomyocyte differentiation of pluripotent stem cells and their use as cardiac disease models. Biochem J 434:25–35
Wada R, Muraoka N, Inagawa K et al (2013) Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc Natl Acad Sci U S A 110:12667–12672
Schott JJ, Benson DW, Basson CT et al (1998) Congenital heart disease caused by mutations in the transcription factor NKX2-5. Science 281:108–111
Garg V, Kathiriya IS, Barnes R et al (2003) GATA4 mutations cause human congenital heart defects and reveal an interaction with TBX5. Nature 424:443–447
Park I-H, Arora N, Huo H et al (2008) Disease-specific induced pluripotent stem cells. Cell 134:877–886
Moretti A, Bellin M, Welling A et al (2010) Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N Engl J Med 363:1397–1409
Carvajal-Vergara X, Sevilla A, D’Souza SL et al (2010) Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature 465:808–812
Friedel RH, Wurst W, Wefers B et al (2011) Generating conditional knockout mice. Methods Mol Biol 693:205–231
Gaj T, Gersbach CA, Barbas CF (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31:397–405
Probst FJ, Justice MJ (2010) Mouse mutagenesis with the chemical supermutagen ENU. Methods Enzymol 477:297–312
Mohr SE, Perrimon N (2012) RNAi screening: new approaches, understandings, and organisms. Wiley Interdiscip Rev RNA 3:145–158
Bedell VM, Westcot SE, Ekker SC (2011) Lessons from morpholino-based screening in zebrafish. Brief Funct Genomics 10:181–188
Bentham J, Michell AC, Lockstone H et al (2010) Maternal high-fat diet interacts with embryonic Cited2 genotype to reduce Pitx2c expression and enhance penetrance of left-right patterning defects. Hum Mol Genet 19:3394–3401
Rockman HA, Ross RS, Harris AN et al (1991) Segregation of atrial-specific and inducible expression of an atrial natriuretic factor transgene in an in vivo murine model of cardiac hypertrophy. Proc Natl Acad Sci U S A 88:8277–8281
Yue TL, Gu JL, Wang C et al (2000) Extracellular signal-regulated kinase plays an essential role in hypertrophic agonists, endothelin-1 and phenylephrine-induced cardiomyocyte hypertrophy. J Biol Chem 275:37895–37901
Brown SDM, Moore MW (2012) The International Mouse Phenotyping Consortium: past and future perspectives on mouse phenotyping. Mamm Genome 23:632–640
Ayadi A, Birling M-C, Bottomley J et al (2012) Mouse large-scale phenotyping initiatives: overview of the European Mouse Disease Clinic (EUMODIC) and of the Wellcome Trust Sanger Institute Mouse Genetics Project. Mamm Genome 23:600–610
Gates H, Mallon A-M, Brown SDM, EUMODIC Consortium (2011) High-throughput mouse phenotyping. Methods 53:394–404
Gregg CL, Butcher JT (2012) Quantitative in vivo imaging of embryonic development: opportunities and challenges. Differentiation 84:149–162
Bamforth SD, Schneider JE, Bhattacharya S (2012) High-throughput analysis of mouse embryos by magnetic resonance imaging. Cold Spring Harb Protoc 2012:93–101
Phoon CKL (2006) Imaging tools for the developmental biologist: ultrasound biomicroscopy of mouse embryonic development. Pediatr Res 60:14–21
Veuthey T, Herrera G, Dodero VI (2014) Dyes and stains: from molecular structure to histological application. Front Biosci (Landmark Ed) 19:91–112
Mohun TJ, Weninger WJ (2011) Imaging heart development using high-resolution episcopic microscopy. Curr Opin Genet Dev 21:573–578
Norris FC, Wong MD, Greene NDE et al (2013) A coming of age: advanced imaging technologies for characterising the developing mouse. Trends Genet 29:700–711
Erni R, Rossell MD, Kisielowski C, Dahmen U (2009) Atomic-resolution imaging with a sub-50-pm electron probe. Phys Rev Lett 102:096101
Herzenberg LA, Parks D, Sahaf B et al (2002) The history and future of the fluorescence activated cell sorter and flow cytometry: a view from Stanford. Clin Chem 48:1819–1827
Bébarová M (2012) Advances in patch clamp technique: towards higher quality and quantity. Gen Physiol Biophys 31:131–140
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Xiao W, Oefner PJ (2001) Denaturing high-performance liquid chromatography: a review. Hum Mutat 17:439–474
Shendure J, Ji H (2008) Next-generation DNA sequencing. Nat Biotechnol 26:1135–1145
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A 74:5463–5467
Lander ES, Linton LM, Birren B et al (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Venter JC, Adams MD, Myers EW et al (2001) The sequence of the human genome. Science 291:1304–1351
Etheridge S. What’s so special about next generation sequencing? Available at: www.oxbridgebiotech.com/review/research-and-policy/whats-so-special-about-next-generation-sequencing. Accessed 05 Feb 2015
Mardis ER (2008) The impact of next-generation sequencing technology on genetics. Trends Genet 24:133–141
Gijsbers ACJ, Ruivenkamp CAL (2011) Molecular karyotyping: from microscope to SNP arrays. Horm Res Paediatr 76:208–213
Maskos U, Southern EM (1992) Oligonucleotide hybridizations on glass supports: a novel linker for oligonucleotide synthesis and hybridization properties of oligonucleotides synthesised in situ. Nucleic Acids Res 20:1679–1684
Visscher PM, Brown MA, McCarthy MI, Yang J (2012) Five years of GWAS discovery. Am J Hum Genet 90:7–24
Shendure J, Porreca GJ, Reppas NB et al (2005) Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309:1728–1732
Margulies M, Egholm M, Altman WE et al (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Wetterstrand KA. DNA sequencing costs: data from the NHGRI genome sequencing program (GSP). Available at: www.genome.gov/sequencingcosts. Accessed 05 Feb 2015
Dorn C, Grunert M, Sperling SR (2013) Application of high-throughput sequencing for studying genomic variations in congenital heart disease. Brief Funct Genomics 13:51–65
Lander ES, Waterman MS (1988) Genomic mapping by fingerprinting random clones: a mathematical analysis. Genomics 2:231–239
1000 Genomes Project Consortium, Abecasis GR, Auton A, et al (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65
Barriot R, Breckpot J, Thienpont B et al (2010) Collaboratively charting the gene-to-phenotype network of human congenital heart defects. Genome Med 2:16
Cardiovascular gene annotation. Cardiovascular Gene Annotation group. Available at: www.ucl.ac.uk/functional-gene-annotation/cardiovascular. Accessed 05 Feb 2015
Pediatric Cardiac Genomics Consortium, Gelb B, Brueckner M, et al (2013) The Congenital Heart Disease Genetic Network Study: rationale, design, and early results. Circ Res 112:698–706
Zaidi S, Choi M, Wakimoto H et al (2013) De novo mutations in histone-modifying genes in congenital heart disease. Nature 498:220–223
Firth HV, Wright CF, DDD Study (2011) The Deciphering Developmental Disorders (DDD) study. Dev Med Child Neurol 53:702–703
The Deciphering Developmental Disorders Study (2015) Large-scale discovery of novel genetic causes of developmental disorders. Nature 519(7542):223–8
Wright CF, Fitzgerald TW, Jones WD et al (2015) Genetic diagnosis of developmental disorders in the DDD study: a scalable analysis of genome-wide research data. Lancet 385(9975):1305–14
Martin JA, Wang Z (2011) Next-generation transcriptome assembly. Nat Rev Genet 12:671–682
Saliba A-E, Westermann AJ, Gorski SA, Vogel J (2014) Single-cell RNA-seq: advances and future challenges. Nucleic Acids Res 42:8845–8860
Junker JP, Noël ES, Guryev V et al (2014) Genome-wide RNA Tomography in the zebrafish embryo. Cell 159:662–675
Johnson DS, Mortazavi A, Myers RM, Wold B (2007) Genome-wide mapping of in vivo protein-DNA interactions. Science 316:1497–1502
Han Y, Garcia BA (2013) Combining genomic and proteomic approaches for epigenetics research. Epigenomics 5:439–452
Visel A, Rubin EM, Pennacchio LA (2009) Genomic views of distant-acting enhancers. Nature 461:199–205
Furlan-Magaril M, Rincón-Arano H, Recillas-Targa F (2009) Sequential chromatin immunoprecipitation protocol: ChIP-reChIP. Methods Mol Biol 543:253–266
Greil F, Moorman C, van Steensel B (2006) DamID: mapping of in vivo protein-genome interactions using tethered DNA adenine methyltransferase. Methods Enzymol 410:342–359
Serra-Juhé C, Cuscó I, Homs A et al (2015) DNA methylation abnormalities in congenital heart disease. Epigenetics
Hsu H-K, Weng Y-I, Hsu P-Y et al (2014) Detection of DNA methylation by MeDIP and MBDCap assays: an overview of techniques. Methods Mol Biol 1105:61–70
Ronaghi M, Uhlén M, Nyrén P (1998) A sequencing method based on real-time pyrophosphate. Science 281:363–365
Thurman RE, Rynes E, Humbert R et al (2012) The accessible chromatin landscape of the human genome. Nature 489:75–82
Meyer CA, Liu XS (2014) Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. Nat Rev Genet 15:709–721
Belmont AS (2014) Large-scale chromatin organization: the good, the surprising, and the still perplexing. Curr Opin Cell Biol 26:69–78
Fullwood MJ, Liu MH, Pan YF et al (2009) An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462:58–64
König J, Zarnack K, Luscombe NM, Ule J (2011) Protein-RNA interactions: new genomic technologies and perspectives. Nat Rev Genet 13:77–83
Matkovich SJ, Van Booven DJ, Eschenbacher WH, Dorn GW (2011) RISC RNA sequencing for context-specific identification of in vivo microRNA targets. Circ Res 108:18–26
Xia Y, Hong H, Ye L et al (2013) Label-free quantitative proteomic analysis of right ventricular remodeling in infant Tetralogy of Fallot patients. J Proteomics 84:78–91
Bahado-Singh RO, Ertl R, Mandal R et al (2014) Metabolomic prediction of fetal congenital heart defect in the first trimester. Am J Obstet Gynecol 211:240.e1–240.e14
Aebersold R, Mann M (2003) Mass spectrometry-based proteomics. Nature 422:198–207
Krüger M, Moser M, Ussar S et al (2008) SILAC mouse for quantitative proteomics uncovers kindlin-3 as an essential factor for red blood cell function. Cell 134:353–364
Sury MD, Chen J-X, Selbach M (2010) The SILAC fly allows for accurate protein quantification in vivo. Mol Cell Proteomics 9:2173–2183
Ong S-E, Blagoev B, Kratchmarova I et al (2002) Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteomics 1:376–386
Gygi SP, Rist B, Gerber SA et al (1999) Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol 17:994–999
Ross PL, Huang YN, Marchese JN et al (2004) Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol Cell Proteomics 3:1154–1169
Bonzon-Kulichenko E, Pérez-Hernández D, Núñez E et al (2011) A robust method for quantitative high-throughput analysis of proteomes by 18O labeling. Mol Cell Proteomics 10:M110.003335
Beckonert O, Keun HC, Ebbels TMD et al (2007) Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat Protoc 2:2692–2703
Stelzl U, Worm U, Lalowski M et al (2005) A human protein-protein interaction network: a resource for annotating the proteome. Cell 122:957–968
Ravasi T, Suzuki H, Cannistraci CV et al (2010) An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140:744–752
Volkmer R, Tapia V, Landgraf C (2012) Synthetic peptide arrays for investigating protein interaction domains. FEBS Lett 586:2780–2786
Lara-Pezzi E, Dopazo A, Manzanares M (2012) Understanding cardiovascular disease: a journey through the genome (and what we found there). Dis Model Mech 5:434–443
Acknowledgements
This work was supported by the European Community’s Seventh Framework Programme contract (“CardioNeT”) grant 289600 to S.R.S and the German Research Foundation (Heisenberg professorship and grant 574157 to S.R.S.). This work was also supported by the Berlin Institute of Health (BIH-CRG2-ConDi to S.R.S.).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer-Verlag Wien
About this chapter
Cite this chapter
Dorn, C. et al. (2016). Technologies to Study Genetics and Molecular Pathways. In: Rickert-Sperling, S., Kelly, R., Driscoll, D. (eds) Congenital Heart Diseases: The Broken Heart. Springer, Vienna. https://doi.org/10.1007/978-3-7091-1883-2_18
Download citation
DOI: https://doi.org/10.1007/978-3-7091-1883-2_18
Publisher Name: Springer, Vienna
Print ISBN: 978-3-7091-1882-5
Online ISBN: 978-3-7091-1883-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)