Abstract
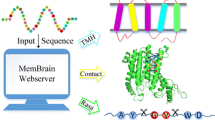
A specific number of chains form alpha-helical membrane protein complexes in order to realize the biochemical function, i.e. as gateways to decide whether specific substances can be transported across the membrane or not. However, few structures of membrane proteins have been solved. The knowledge of protein-protein binding residues can help biologists figure out how the function works and solve the 3D structures.
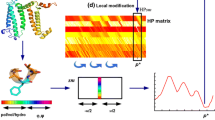
We present a novel, sequence-based method to predict protein-protein binding residues from primary protein sequences by machine learning classifiers. We use a support vector regression model to predict relative solvent accessibility by features based on sequences, including position specific scoring matrix, conserved score, z-coordinate prediction, second structure prediction, physical parameter and sequence length. Afterwards, combining features mentioned above with the predicted solvent accessibility, we use ensemble support vector machines to predict protein-protein binding residues. To the best of our knowledge, there is no method to predict protein-protein binding residues in alpha-helical membrane proteins. Our method outperforms MAdaBoost successfully used in predicting protein-ligand binding residues and random forest used in protein-protein binding residues from surface residues. We also assess the importance of each individual type of features. PSSM profile and conserved score are shown to be more effective to predict protein-protein binding residues in alpha-helical membrane proteins.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
References
Almén, M.S., Nordström, K.J.V., Fredriksson, R., et al.: Mapping the human membrane proteome: a majority of the human membrane proteins can be classified according to function and evolutionary origin. BMC Biology 7(1), 50 (2009)
Kozma, D., Simon, I., Tusnády, G.E.: PDBTM: Protein Data Bank of transmembrane proteins after 8 years. Nucleic Acids Research 41(D1), D524–D529 (2013)
Yarov-Yarovoy, V., Schonbrun, J., Baker, D.: Multipass membrane protein structure prediction using Rosetta. Proteins: Structure, Function, and Bioinformatics 62(4), 1010–1025 (2006)
Nugent, T., Jones, D.T.: Accurate de novo structure prediction of large transmembrane protein domains using fragment-assembly and correlated mutation analysis [J]. Proceedings of the National Academy of Sciences 109(24), E1540–E1547 (2012)
Weiner, B.E., Woetzel, N., Karakaş, M., et al.: BCL: MP-fold: folding membrane proteins through assembly of transmembrane helices. Structure 21(7), 1107–1117 (2013)
Chen, K., Mizianty, M.J., Kurgan, L.: ATPsite: sequence-based prediction of ATP-binding residues. Proteome Sci. 9(suppl. 1), S4 (2011)
Yu, D., Hu, J., Yang, J., et al.: Designing template-free predictor for targeting protein-ligand binding sites with classifier ensemble and spatial clustering (2013)
Bordner, A.J.: Predicting protein-protein binding sites in membrane proteins. BMC Bioinformatics 10(1), 312 (2009)
Adamczak, R., Porollo, A., Meller, J.: Accurate prediction of solvent accessibility using neural networks–based regression. Proteins: Structure, Function, and Bioinformatics 56(4), 753–767 (2004)
Illergård, K., Callegari, S., Elofsson, A.: MPRAP: An accessibility predictor for a-helical transmem-brane proteins that performs well inside and outside the membrane. BMC Bioinformatics 11(1), 333 (2010)
Hubbard, S.J.T.J.: NACCESS, Computer program. Department of Biochemistry and Molecular Biology 1, 1–2 (1993), http://wolf.bi.umist.ac.uk/unix/naccess.html
McGuffin, L.J., Bryson, K., Jones, D.T.: PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000)
Altschul, S.F., Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W., Lipman, D.J.: Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Granseth, E., Viklund, H., Elofsson, A.: ZPRED: predicting the distance to the membrane center for residues in alpha-helical membrane proteins. Bioinformatics 22(14), e191-e196 (2006)
Mayrose, I., Graur, D., Ben-Tal, N., Pupko, T.: Comparison of site-specific rate-inference methods: Bayesian methods are superior. Mol. Biol. Evol. 21, 1781–1791 (2004)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Xiao, F., Shen, HB. (2014). Sequence-Based Prediction of Protein-Protein Binding Residues in Alpha-Helical Membrane Proteins. In: Li, S., Liu, C., Wang, Y. (eds) Pattern Recognition. CCPR 2014. Communications in Computer and Information Science, vol 484. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-662-45643-9_44
Download citation
DOI: https://doi.org/10.1007/978-3-662-45643-9_44
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-662-45642-2
Online ISBN: 978-3-662-45643-9
eBook Packages: Computer ScienceComputer Science (R0)