Abstract
Studies reporting molecular analysis of nosocomial outbreaks of Pneumocystis jirovecii pneumonia (PCP) are reviewed. Although the methods currently used for typing detect only the most abundant co-infecting genotypes, these studies have consistently identified a single P. jirovecii genotype as responsible for most or all PCP cases in a given outbreak. A single genotype was responsible for three separate outbreaks in Germany and Switzerland. Together with transmission maps compatible with potential infectious encounters, these results strongly suggest that inter-human transmission was involved. However, infection from a common source or by indirect transmission by asymptomatic carriers of the fungus could not be excluded. The following conditions favoring outbreaks were seen in all settings: (i) anti-PCP prophylaxis was not administered or was inadequate, (ii) there was potential contact between source and susceptible patients, and (iii) there was no isolation of patients with PCP. One study suggested that hospitalized immunosuppressed patients colonized by P. jirovecii might be a potential source of nosocomial infection. Underlying pulmonary disease, transplanted organ dysfunction, and lower CD4 lymphocyte counts were identified as new clinical risk factors for PCP. At present, two non-mutually exclusive hypotheses can explain the selection of the Pneumocystis genotypes that were responsible for the outbreaks: (i) the genotype represented the predominant genotype in the geographical area, or (ii) the genotype has developed pathogenic factors that facilitate infection. Perspectives on further investigation and prevention of outbreaks are discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Clinical Risk Factor
- Pneumocystis Jirovecii
- Indirect Transmission
- Molecular Typing Method
- Major Surface Glycoprotein
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
I. Introduction
Pneumocystis jirovecii is a fungus that causes severe pneumonia in immunocompromised humans. Recognized initially among presumably malnourished infants, and subsequently among immunosuppressed cancer patients, Pneumocystis jirovecii pneumonia (PCP) assumed medical prominence with the advent of the HIV epidemic. With the development and broad utilization of combination antiretroviral therapy, the disease has become sporadic in HIV-infected persons, primarily occurring among those who are unaware of their status, as well as in solid organ transplantation recipients and patients with hemato-oncologic or autoimmune diseases. Notably, multiple nosocomial outbreaks of PCP among renal transplant recipients (RTRs) have been reported, especially over the past 15 years. These outbreaks suggest inter-human transmission of the pathogen or its acquisition from a common source. Because a detailed review of studies of PCP outbreaks has recently been published (de Boer et al. 2011), the present chapter focuses on new insights from more recent publications. In addition, we comment on the limitations of the molecular typing methods available for investigation of PCP outbreaks.
II. Biology
P. jirovecii specifically infects humans, whereas other Pneumocystis species infect other mammals (Thomas and Limper 2007; Cushion and Stringer 2010). P. jirovecii grows extracellularly in the alveoli of human lungs. In the absence of a long-term in vitro culture system, the life cycle remains hypothetical and is postulated to include asexual (trophic cell) and sexual (ascus, or cyst) forms. Although PCR techniques are being increasingly utilized for diagnosis, the gold standard still remains the detection of organisms within respiratory specimens using various staining methods, such as methenamine silver and direct immunofluorescence (examples are shown in Fig. 11.1). A number of biological features of Pneumocystis species suggest that they are obligate parasites without a free-living form: (i) organisms can incorporate host cholesterol (Furlong et al. 1997), (ii) they possess a system of surface antigen variation (Stringer and Keely 2001), (iii) they appear to have co-evolved with their hosts (Demanche et al. 2001), and (iv) they lack most of the enzymes that are dedicated to the biosyntheses of amino acids in fungi (Table 11.1) (Hauser et al. 2010; Cissé et al. 2012). Moreover, the P. jirovecii genome lacks known virulence factors and has a relatively small size and reduced GC content (Cissé et al. 2012). Together with epidemiological studies, which have failed to identify a free-living form of the fungus, these features suggest that P. jirovecii is an obligate parasite that has developed specialized systems for infecting humans but that causes clinically significant disease only in immunocompromised hosts.
(a) P. jirovecii asci visualized in a patient’s bronchoalveolar lavage using Grocott’s methenamine silver staining as described by Churukian and Schenk (1977). The parentheses-like structure in each ascus resembles opercula observed in other asci. Ascomycetous operculum is thought to be involved in the dehiscence (release of endospores) upon contact with humidity, suggesting that asci might be the airborne P. jirovecii infectious forms (picture from Institute of Microbiology, Centre Hospitalier Universitaire Vaudois). (b) P. jirovecii organisms visualized in a patient’s induced sputum using direct immunofluorescence kit BioRad MONOFLUO Pneumocystis (Picture from Critical Care Medicine Department, NIH Clinical Center, National Institutes of Health)
III. Epidemiology
Immunity against P. jirovecii develops during the first years of life, presumably after a primary infection that occurs prior to 2 years of age (Pifer et al. 1978; Vargas et al. 2001). Thus, carriage of the fungus as a latent infection and reactivation upon immunosuppression was the first hypothesis for the development of PCP later in life. However, a number of recent observations have suggested that infection de novo from exogenous sources is responsible for the majority of episodes: (i) the occurrence of PCP outbreaks caused by a single genotype (de Boer et al. 2011), (ii) the elimination of Pneumocystis after experimental infection in the rat model (Vargas et al. 1995), and (iii) the presence of mutations within the dihydropteroate synthase gene in patients who never received sulfa drugs (Huang et al. 2000; Nahimana et al. 2003a). The latter observation provides especially strong evidence supporting de novo infection because these mutations (i) are associated with failure of sulfa prophylaxis, (ii) are absent in P. jirovecii isolated before the use of sulfa drugs for treating or preventing PCP (Meshnick 1999), and (iii) have been shown to be selected by sulfonamide use within single patients (Nahimana et al. 2003b).
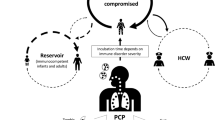
The occurrence of infection de novo raises questions about the source of the pathogen and mode(s) of transmission. Obligate parasitism implies that the reservoir and source of infection are probably only humans.
A number of experiments in animal models have helped in understanding the epidemiology of P. jirovecii in humans. Studies on rats demonstrated that air is the route of transmission for de novo infection (Hughes 1982; Hughes et al. 1983). Horizontal transfer of the fungus can occur from animals with PCP to susceptible animals (Powles et al. 1992; Vogel et al. 1993; Ceré et al. 1997), but also from immunocompetent murine carriers of the fungus following a single day of exposure (Dumoulin et al. 2000; Chabé et al. 2004).
Carriage of P. jirovecii without overt PCP by immunocompromised (Nevez et al. 1999; Hauser et al. 2000) and immunocompetent (Vargas et al. 2000; Miller et al. 2001; Tipirneni et al. 2009) humans has been documented. The genotypes observed in carriers (Hauser et al. 2000) and in infants, presumably experiencing primary infection (Totet et al. 2003, 2004), were not different from those in PCP patients, suggesting that these could be sources of the pathogen. However, one study of larger cohorts found that infants may harbor different genotypes from AIDS patients (Beard et al. 2005). This study suffers from the fact that the infant and adult populations were from different geographical areas, which may have had different predominant genotypes. The dispersion of Pneumocystis organisms in the air around PCP patients (Choukri et al. 2010; Bartlett et al. 1997), as well as around infected rats (Choukri et al. 2011) has been examined by PCR techniques. Significant fungal burdens were detected in the air at 1 m from the source, then decreased with distance but were detected up to 8 m away. For rats, air burdens correlated with lung burdens. The same P. jirovecii genotypes were present in the lungs of PCP patients and in air particles collected in their room (Bartlett et al. 1997; Damiani et al. 2012). Together, these observations suggest that inter-human transmission from infected humans through the air is the main route of dissemination of P. jirovecii. The possible categories of humans that may act as a source include patients with active or developing PCP, infants experiencing primary infection, and colonized immunocompromised patients, but also, possibly, transient carriers such as elderly people (Vargas et al. 2010), pregnant women (Vargas et al. 2003), and health-care workers (Miller et al. 2001; Tipirneni et al. 2009). Nevertheless, the actual human source(s) of infection for sporadic cases and their relative importance remain to be elucidated. We speculate that infants experiencing primary infection might play a major role because, being immunologically naive, multiplication of the fungus might be more productive than in other categories of colonized humans.
IV. Molecular Methods for Investigating PCP Outbreaks
In order to test the hypothesis that inter-human transmission of P. jirovecii is involved in outbreaks, molecular typing methods are crucial because they can document identical P. jirovecii genotypes in patients between whom transmission is suspected. Typing methods for P. jirovecii rely on the natural genetic heterogeneity among isolates. In the absence of a culture method, current typing methods utilize PCR amplification of variable regions of the P. jirovecii genome, followed by the detection of polymorphisms by DNA sequencing, single-strand conformation polymorphism (SSCP) analysis, or restriction fragment length polymorphism (RFLP) analysis. Initially, one to two genetic regions were sequenced, including a variable region of the mitochondrial 26S rRNA gene (hereafter called mt26S) (Keely et al. 1995) or the internal transcribed spacers number 1 and 2 of the nuclear rRNA operon (ITS1 and ITS2; ITSs sequencing) (Lee et al. 1993; Lu et al. 1994; Tsolaki et al. 1996). Subsequently, a multilocus approach was developed that utilized SSCP to analyze four genomic regions (Hauser et al. 1997, 2001a; Nahimana et al. 2000), including ITS1, mt26S, the intron of the nuclear 26S rRNA gene (26S), and the β-tubulin intron 6 region (b-tub). As DNA sequencing prices decreased, SSCP was replaced by direct sequencing, i.e., without cloning PCR products (multilocus sequence typing, MLST) (Schmoldt et al. 2008; Gianella et al. 2010; Pliquett et al. 2012). RFLP analysis of a PCR product from a variant region of the gene family encoding the major surface glycoprotein (RFLP-MSG) was shown to be discriminative and useful for investigating outbreaks (Ripamonti et al. 2009; Sassi et al. 2012). The polymorphisms in all markers that were used for typing appeared to be stable over months (Keely et al. 1995; Hauser et al. 1997, 2001a; Hauser 2004), which is a prerequisite for microbe-based epidemiological studies (Hauser et al. 1998a). The index of discrimination power is the average probability that two unrelated specimens randomly chosen will be assigned to different types by the method. It has been determined only for the SSCP method and was estimated to be 0.93 (Hauser et al. 2001a), which can be considered as satisfactory because the minimal desired level for a typing system to be used as a single method is 0.95 (Struelens and ESGEM 1996). SSCP and ITSs sequencing identified, respectively, 43 (Hauser 2004) and more than 60 different P. jirovecii genotypes (Lu and Lee 2008). The same markers have been analyzed by SSCP and MLST, but the latter is more discriminative because it detects all polymorphisms.
The interpretation of data using the current typing methods for P. jirovecii must take into account two important caveats. First, co-infection of a single patient with two or more P. jirovecii genotypes has been reported using multiple typing methods (Tsolaki et al. 1996; Hauser et al. 1998a; Nahimana et al. 2000; Ma et al. 2002; Ripamonti et al. 2009), which obviously complicates epidemiological studies (Hauser et al. 1998b). The proportion of co-infected patients varies considerably, ranging from a few percent using direct sequencing to ~70% using SSCP. SSCP can detect a co-infecting type if it represents at least 11% of the population (Nahimana et al. 2000). Cloning of PCR products followed by DNA sequencing can also be used to detect co-infections; the minimal proportion detected will depend on the number of clones that are analyzed. Direct sequencing of PCR products without cloning would not be able to detect low-abundance co-infecting types. Geographical variation or other parameters probably account for some of the reported differences in the proportion of co-infected patients. Identifying all co-infecting genotypes present in clinical specimens of infected humans is probably necessary to fully understand P. jirovecii epidemiology, and may also be useful for analysis of outbreaks, because some genotypes might be more transmissible than others. Nonetheless, at present most investigations of outbreaks use methods that detect only the most abundant genotype(s). Accordingly, recent outbreaks have often been investigated by MLST without cloning to detect co-infections, and this method was able to provide evidence that a single P. jirovecii genotype accounted for all or most of the PCP cases in a given outbreak (Schmoldt et al. 2008; Gianella et al. 2010; Phipps et al. 2011; Pliquett et al. 2012). ITSs sequencing with or without cloning PCR products also reached this conclusion (Yazaki et al. 2009; Le Gal et al. 2012).
The second caveat in interpreting data from typing methods is that the P. jirovecii genotypes present in respiratory samples may not include all those present in the patient’s lungs. Indeed, analysis of lungs from three autopsied individuals in one study demonstrated a compartmentalization of different co-infecting P. jirovecii genotypes in the lungs of two patients. Importantly, some genotypes identified in specific regions of the lungs were absent in the corresponding respiratory specimen (see Table 11.2) (Helweg-Larsen et al. 2001a). As previously stressed by Hughes (2007), this raises questions about studies that concluded that recurrent PCP episodes represent de novo infection, because genotype(s) already present in the lungs may have been detected only during the second and not the first episode. On the other hand, this apparently has not caused problems for investigations of outbreaks since the same P. jirovecii genotype was observed in many instances (see following section).
As far as ITSs typing is concerned, amplification by PCR has been reported to create an artifactual diversity of genotypes, i.e., 37% of them were chimeras (Beser et al. 2007). A substantial decrease in the proportion of these artifacts was obtained by increasing elongation time, primer concentration, and annealing temperature, as well as by decreasing the number of cycles and using a proofreading polymerase (Beser et al. 2007). Nevertheless, two recent studies successfully detected the same ITSs genotype in outbreak PCP cases, apparently without using the advised modifications to decrease chimera formation (Yazaki et al. 2009; Le Gal et al. 2012). Infection with a single genotype may account for this because there would be no opportunity for recombination.
V. Molecular Epidemiology of Pneumocystis Outbreaks
In a recent publication, de Boer et al. (2011) reviewed studies of 16 nosocomial PCP outbreaks among RTRs, six of which used molecular typing to characterize the genotype. A transmission map describing timings of outpatient visits and hospitalizations was generally provided in these studies to assess potential occurrence of infectious encounters (an example is shown in Fig. 11.2). Molecular typing revealed in five out of six instances that all or the vast majority of the outbreak cases with suspected infectious contact(s) harbored the same P. jirovecii genotype, strongly suggesting that inter-human transmission of P. jirovecii had occurred. Importantly, all outbreaks shared the three following features, which probably contributed to the outbreaks: (i) cases received no or inadequate anti-PCP prophylaxis, (ii) contact could occur between potential source patients with clinical or incubating PCP and susceptible patients (in outpatient waiting rooms or during a simultaneous hospitalization), and (iii) no isolation procedure of PCP patients was utilized. However, being retrospective, these studies could not firmly exclude that cases acquired the fungus from asymptomatic colonized health-care worker(s), from an unidentified environmental source, or by indirect transmission from an index patient through nosocomial carriers.
Potential encounters compatible with nosocomial inter-human transmission of P. jirovecii in one building of a hospital. Thicker parts of solid lines represent periods of hospitalization. Each encounter or consecutive encounters are figured by an arrow pointing to the direction of the presumed transmission; the number of encounters is indicated close to each arrow. *Anti-PCP prophylaxis with Fansidar was suboptimal (25 mg pyrimethamine and 500 mg sulfadoxine once or twice every 2 weeks). D death, G graft, R rejection episode, RTR renal transplant recipient, PCPnoso nosocomial case, DHPS dihydropteroate synthase, M2 amino acid change within the putative sulfa-binding site at position 57 (Pro to Ser) [Taken from Fig. 4 of Rabodonirina et al. (2004)]
Since the review of de Boer et al. (2011), eight additional PCP outbreaks have been reported in seven studies, nearly all in RTRs. One study provided evidence of transmission only on the basis of possible infectious encounters (Mori et al. 2010). Of note, the latter is the first among outpatients with rheumatoid arthritis. In addition to possible infectious encounters, the other six studies reported that a single molecular P. jirovecii genotype accounted for most or all of the RTR outbreak cases (Phipps et al. 2011; Thomas et al. 2011; Wynckel et al. 2011; Brunot et al. 2012; Pliquett et al. 2012; Le Gal et al. 2012). In all eight outbreaks, patients were not receiving anti-PCP prophylaxis at the time of suspected exposure, and there was no policy to isolate PCP patients. Thus, these recent studies provide further evidence that inter-human transmission is involved in such outbreaks, but they also could not exclude a common source or indirect transmission.
These studies highlight at least three new issues that could help explain PCP outbreaks:
-
(i)
For the first time, three out of ten potential source patients were identified as carriers of P. jirovecii without clinical symptoms of PCP (Le Gal et al. 2012). This is important because carriers are not routinely identified within hospitals and they may be even more numerous than PCP patients, and as such they may constitute an important nosocomial source of P. jirovecii. The risk of transmission by such individuals needs to be determined, potentially by measuring dissemination of the fungus in the air as previously described for PCP patients (Choukri et al. 2010). If such individuals are an important source of infection, approaches to minimizing such transmission would need to be evaluated.
-
(ii)
The new studies identified new clinical risk factors for PCP that may have played a role in the outbreaks. Alteration in immunosuppressive therapy, graft rejection, cytomegalovirus infection, and older age were previously identified (de Boer et al. 2011). A cohort analysis including 14 cases and 324 control patients identified two new independent risk factors: underlying pulmonary disease and transplanted organ dysfunction (Phipps et al. 2011). Lower CD4 lymphocyte counts at 3 months after transplantation (Brunot et al. 2012) or just before PCP diagnosis (Struijk et al. 2011) were also identified as risk factors. Although they need confirmation because they were identified in single studies, these findings may be important for preventing outbreaks by better identifying patients who would benefit from PCP prophylaxis.
-
(iii)
Two different P. jirovecii genotypes were involved in two outbreaks that occurred at a relatively close geographical distance of 50 km (Liverpool and Manchester) (Thomas et al. 2011). This raises new issues for the analysis of the different outbreak-associated P. jirovecii genotypes, which we present in the following section.
VI. P. jirovecii Outbreak Genotypes
An important finding concerning P. jirovecii genotypes involved in outbreaks has been reported recently (Sassi et al. 2012). Genotypes linked to two PCP outbreaks that occurred contemporaneously in two European cities at a distance of 300 km (Zurich and Munich) were found to be identical using both MLST and RFLP-MSG. Subsequently, the same genotype was identified using MLST in a third contemporaneous outbreak at a distance of 400 km from the two previous ones (Frankfurt) (Hauser et al. 2013). However, this genotype was different from that involved in a contemporaneous Japanese outbreak (Nagoya) (Sassi et al. 2012), as well as from that observed in another European outbreak that occurred about 12 years earlier at a distance of 400–800 km (Lyon) (Hauser et al. 2013).
Two epidemiological factors may have played a role in the variation of the genotype involved in the different outbreaks: the period of time and the geographical location (Hauser et al. 2013). However, the presence of the same genotype in three contemporaneous and geographically close outbreaks is particularly intriguing. As previously noted (Sassi et al. 2012; de Boer 2012), two hypotheses could explain this latter finding: (i) the genotype was predominant in an area covering the three European locations and infected index patients of the three outbreaks, or (ii) the genotype carried uncharacterized pathogenic factor(s) and some unidentified contact (e.g., a patient or colonized individual) existed between the three outbreaks. Because the few sporadic contemporaneous cases analyzed in the three outbreaks (six, three, and two in Munich, Zurich, and Frankfurt, respectively) were infected by other genotypes, the outbreak genotype was considered not to be predominant, but was hypothesized to be more pathogenic in RTRs (Sassi et al. 2012).
On the other hand, by analysis of a large number of isolates, the Lyon outbreak genotype was demonstrated to be the predominant one at the time in the affected hospital as well as in the local area (Rabodonirina et al. 2004), and one of the most prevalent in several European locations (Hauser et al. 2001a). Nevertheless, the Lyon genotype may have been not only predominant but also more transmissible. Indeed, the potential infectious encounters during the outbreak also involved other genotypes, but these apparently were not transmitted (Rabodonirina et al. 2004). The increased transmissibility or pathogenicity of the Lyon genotype might have been due to unidentified specific factors, but also potentially to the presence of one or two amino acid changes within the putative sulfa-binding site of P. jirovecii dihydropteroate synthase (Pro to Ser at position 57, mutation M2; Thr to Ala and Pro to Ser at positions 55 and 57, respectively, double mutation M3). All but one potential donor in the encounters harbored M2, whereas M3 was present in the remaining donor (Fig. 11.2). Epidemiological studies have shown that these mutations confer some level of resistance to sulfonamides (Kazanjian et al. 1998; Helweg-Larsen et al. 1999; Nahimana et al. 2003a), and the results were confirmed by functional complementation in Escherichia coli (Iliades et al. 2005) and Saccharomyces cerevisiae (Meneau et al. 2004). Consequently, the Lyon outbreak genotype may have been selected among the different genotypes encountered because several patients were receiving suboptimal prophylaxis (Fig. 11.2). Thus, the observations in Lyon suggest that the two hypotheses, preponderance and increased pathogenicity of the outbreak genotypes, may not be mutually exclusive.
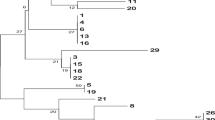
In order to further investigate this issue, we analyzed all outbreak genotypes that could be compared with each other because they shared at least one sequenced marker (see Table 11.3). The results of Wynckel et al. (2011) could not be included because ITS types are not given. Although only one marker could be compared, the ITS1 allele confirmed that the Japanese genotype was different from all others. The ITS1 and mt26S alleles showed that the two genotypes involved in the Manchester and Liverpool outbreaks (Thomas et al. 2011) were also different from all others. The ITS1 allele of the Brest genotype (Le Gal et al. 2012) was the same as that in the three nearly contemporaneous European outbreaks, but the discrimination power of a single marker is not sufficient to draw any conclusions. As revealed by three markers, the Australian outbreak (Phipps et al. 2011) may have harbored the same genotype as that of Lyon. Our analysis demonstrates a great diversity of the outbreak genotypes. Notably, the two contemporaneous English outbreaks involved different genotypes although they occurred at a close geographic distance, which contrasts with the three European outbreaks involving a single genotype. P. jirovecii genotypes from nine (Phipps et al. 2011) and two (Pliquett et al. 2012) unlinked control patients were different from the outbreak genotype, favoring the pathogenicity hypothesis. On the other hand, the Brest outbreak genotype was also predominant among a larger cohort of 22 controls, favoring the preponderance hypothesis. The latter observation also suggests that analysis of numerous controls, as performed only in Brest and Lyon (Table 11.3), may be necessary to confidently determine genotype frequencies.
VII. Biological and Epidemiological Considerations
P. jirovecii has most probably evolved for millions of years as a colonizer of human lungs. Its pathogenic manifestation as acute PCP may have become frequent only recently in history, especially with the advent of HIV-associated immunodeficiency and the increase in the number of immunosuppressed humans due to the progress of medicine. Consequently, P. jirovecii is probably not a primary pathogen like, for example, pathogenic dimorphic fungi, and thus is not expected to harbor virulence factors that have been selected to cause disease. Its potential virulence factors could be considered “accidental”, i.e., selected for improved survival in colonized humans. Pneumocystis has also developed a sophisticated system for antigenic variation that presumably provides a mechanism for evading host immune responses, possibly T cell responses (Bishop et al. 2012). Thus, the biology of P. jirovecii suggests that selection of factors conferring increased pathogenicity is unexpected, which is in favor of the hypothesis of the predominance of the outbreak genotypes. Nevertheless, events that convert non-pathogens to pathogens cannot be excluded.
Studies that reported the association of certain P. jirovecii genotypes with various phenotypes may support the hypothesis of pathogenic factors in outbreak genotypes. Some genotypes were reported to be associated with milder disease (Miller and Wakefield 1999), failure of prophylaxis (Hauser et al. 2001b), failure of PCP treatment (Matos et al. 2003), reduced severity of PCP (van Hal et al. 2009), or increased mortality rate (Rabodonirina et al. 2013). The same genotype was associated with related phenotypes by two independent laboratories (Miller and Wakefield 1999; Hauser et al. 2001b), which strengthens the conclusions drawn. However, in the absence of culture, these phenotypes could not be confirmed and, moreover, other studies failed to confirm some of these associations (Helweg-Larsen et al. 2001b; Valerio et al. 2007). The recent release of the P. jirovecii genome sequence (Cissé et al. 2012) should facilitate studies addressing such issues.
VIII. Conclusions and Perspectives
The data summarized above strongly support the hypothesis that inter-human transmission of P. jirovecii is important in spreading infection in PCP outbreaks. It is noteworthy that outbreaks appeared to occur in settings in which there was no or inadequate use of prophylaxis, contact occurred between susceptible and infected (and infectious) patients, and PCP patients were not placed in respiratory isolation. Thus, host susceptibility and clinical management of PCP patients may play major roles in the outbreaks. How can such outbreaks be prevented in the future? One approach that has been validated in clinical trials is the appropriate use of PCP prophylaxis for at-risk populations. In most of the outbreaks reported to date, widespread implementation of prophylactic measures led to elimination of additional cases. Preventing transmission by other mechanisms, such as isolation of PCP patients, is logical but has not been validated. It is possible that by the time PCP is diagnosed, transmission to susceptible patients has already occurred, given that progressive infection may be present for weeks before diagnosis. Alternatively, attempts to screen all potential carriers of Pneumocystis, including asymptomatic patients as well as medical personnel, does not seem practical, feasible, or cost effective.
The data available to date are insufficient to determine if outbreak genotypes are causing disease because they are the preponderant local genotype, or because they have increased pathogenicity. It is possible that both factors play a role because they are not mutually exclusive, although our current understanding of the biology of the fungus tends to favor the preponderance hypothesis. One approach to addressing this issue would be to sequence the entire genome of multiple genotypes and look for genetic differences that may contribute to pathogenicity. As sequencing techniques are becoming more efficient and cost effective, this may be feasible in the near future.
To better understand PCP outbreaks, the dynamics of populations of P. jirovecii genotypes need to be better understood. Genotypes should ideally be determined for each outbreak in PCP patients and colonized individuals within hospitals, as well as, if possible, in their relatives, including infants as they may represent an important reservoir. The use of a standardized typing method, e.g. MLST, would have the advantage of allowing interlaboratory comparison of different isolates. Parallel DHPS genotyping would allow investigation of the importance of sulfa drug resistance.
The optimal utilization of prophylaxis in RTRs needs to be defined. Currently, anti-PCP prophylaxis is given (if administered) for varying periods at different centers, generally for 3–6 months after transplantation. Given the recent outbreaks, it seems prudent to administer prophylaxis for at least 6 months. However, cases can occur after prophylaxis has been discontinued, with a report up to 15 years after transplantation (Le Gal et al. 2012). As clinical risk factors for development of PCP are identified, prophylaxis can potentially be administered after 6 months to patients with these risk factors, such as in cases of graft rejection.
The occurrence of even two potentially nosocomial cases of PCP should raise concerns and be evaluated as a potential outbreak. P. jirovecii genotyping can determine if in fact an outbreak is occurring and if additional measures to minimize transmission need to be implemented.
References
Bartlett MS, Vermund SH, Jacobs R, Durant PJ, Shaw MM, Smith JW, Tang X, Lu JJ, Li B, Jin S, Lee CH (1997) Detection of Pneumocystis carinii DNA in air samples: likely environmental risk to susceptible persons. J Clin Microbiol 35:2511–2513
Beard CB, Fox MR, Lawrence GG, Guarner J, Hanzlick RL, Huang L, del Rio C, Rimland D, Duchin JS, Colley DG (2005) Genetic differences in Pneumocystis isolates recovered from immunocompetent infants and from adults with AIDS: epidemiological Implications. J Infect Dis 192:1815–1818
Beser J, Hagblom P, Fernandez V (2007) Frequent in vitro recombination in internal transcribed spacers 1 and 2 during genotyping of Pneumocystis jirovecii. J Clin Microbiol 45:881–886
Bishop LR, Helman D, Kovacs JA (2012) Discordant antibody and cellular responses to Pneumocystis major surface glycoprotein variants in mice. BMC Immunol 13:39
Brunot V, Pernin V, Chartier C, Garrigue V, Vetromile F, Szwarc I, Delmas S, Portalès P, Basset D, Mourad G (2012) An epidemic of Pneumocystis jiroveci pneumonia in a renal transplantation center: role of T-cell lymphopenia. Transplant Proc 44:281–2820
Ceré N, Polack B, Chanteloup NK, Coudert P (1997) Natural transmission of Pneumocystis carinii in nonimmunosuppressed animals: early contagiousness of experimentally infected rabbits (Oryctolagus cuniculus). J Clin Microbiol 35:2670–2672
Chabé M, Dei-Cas E, Creusy C, Fleurisse L, Respaldiza N, Camus D, Durand-Joly I (2004) Immunocompetent hosts as a reservoir of Pneumocystis organisms: histological and rt-PCR data demonstrate active replication. Eur J Clin Microbiol Infect Dis 23:89–97
Choukri F, Menotti J, Sarfati C, Lucet JC, Nevez G, Garin YJ, Derouin F, Totet A (2010) Quantification and spread of Pneumocystis jirovecii in the surrounding air of patients with Pneumocystis pneumonia. Clin Infect Dis 51:259–265
Choukri F, el Aliouat M, Menotti J, Totet A, Gantois N, Garin YJ, Bergeron V, Dei-Cas E, Derouin F (2011) Dynamics of Pneumocystis carinii air shedding during experimental pneumocystosis. J Infect Dis 203:1333–1336
Churukian CJ, Schenk EA (1977) Rapid Grocott’s methenamine silver nitrate method for fungi and Pneumocystis carinii. Am J Clin Pathol 68:427–428
Cissé OH, Pagni M, Hauser PM (2012) De novo assembly of the Pneumocystis jirovecii genome from a single bronchoalveolar lavage fluid specimen from a patient. MBio 4:e00428–12. doi:10.1128/mBio.00428-12
Cushion MT, Stringer JR (2010) Stealth and opportunism: alternative lifestyles of species in the fungal genus Pneumocystis. Annu Rev Microbiol 64:431–452
Damiani C, Choukri F, Le Gal S, Menotti J, Sarfati C, Nevez G, Derouin F, Totet A (2012) Possible nosocomial transmission of Pneumocystis jirovecii. Emerg Infect Dis 18:877–878
de Boer MG (2012) Linking Pneumocystis epidemiology, transmission, and virulence. Clin Infect Dis 54:1445–1447
de Boer MGJ, de Fijter JW, Kroon FP (2011) Outbreaks and clustering of Pneumocystis pneumonia in kidney transplant recipients: a systematic review. Med Mycol 49:673–680
Demanche C, Berthelemy M, Petit T, Polack B, Wakefield AE, Dei-Cas E, Guillot J (2001) Phylogeny of Pneumocystis carinii from 18 primate species confirms host specificity and suggests coevolution. J Clin Microbiol 39:2126–2133
Dumoulin A, Mazars E, Seguy N, Gargallo-Viola D, Vargas S, Cailliez JC, Aliouat EM, Wakefield AE, Dei-Cas E (2000) Transmission of Pneumocystis carinii disease from immunocompetent contacts of infected hosts to susceptible hosts. Eur J Clin Microbiol Infect Dis 19:671–678
Furlong ST, Koziel H, Bartlett MS, McLaughlin GL, Shaw MM, Jack RM (1997) Lipid transfer from human epithelial cells to Pneumocystis carinii in vitro. J Infect Dis 175:661–668
Gianella S, Haeberli L, Joos B, Ledergerber B, Wuthrich RP, Weber R, Kuster H, Hauser PM, Fehr T, Mueller NJ (2010) Molecular evidence of interhuman transmission in an outbreak of Pneumocystis jirovecii pneumonia among renal transplant recipients. Transpl Infect Dis 12:1–10
Hauser PM (2004) The development of a typing method for an uncultivable microorganism: the example of Pneumocystis jirovecii. Infect Genet Evol 4:199–203
Hauser PM, Francioli P, Bille J, Telenti A, Blanc DS (1997) Typing of Pneumocystis carinii f. sp. hominis by single-strand conformation polymorphism of four genomic regions. J Clin Microbiol 35:3086–3091
Hauser PM, Blanc DS, Bille J, Francioli P (1998a) Typing methods to approach Pneumocystis carinii genetic heterogeneity. FEMS Immunol Med Microbiol 22:27–35
Hauser PM, Blanc DS, Telenti A, Nahimana A, Bille J, Francioli P (1998b) Potential coinfections complicate typing of Pneumocystis carinii sp.f. hominis. J Clin Microbiol 36:311
Hauser PM, Blanc DS, Bille J, Nahimana A, Francioli P (2000) Carriage of Pneumocystis carinii by immunosuppressed patients and molecular typing of the organisms. AIDS 14:461–463
Hauser PM, Blanc DS, Sudre P, Senggen Manoloff E, Nahimana A, Bille J, Weber R, Francioli P, Study group (2001a) Genetic diversity of Pneumocystis carinii in HIV-positive and -negative patients as revealed by PCR-SSCP typing. AIDS 15:461–466
Hauser PM, Sudre P, Nahimana A, Francioli P, Study group (2001b) Prophylaxis failure is associated with a specific Pneumocystis carinii genotype. Clin Infect Dis 33:1080–1082
Hauser PM, Burdet FX, Cissé OH, Keller L, Taffé P, Sanglard D, Pagni M (2010) Comparative genomics suggests that the fungal pathogen Pneumocystis is an obligate parasite scavenging amino acids from its host’s lungs. PLoS One 5:e15152. doi:10.1371/journal.pone.0015152
Hauser PM, Rabodonirina M, Nevez G (2013) Pneumocystis jirovecii genotypes involved in PCP outbreaks among renal transplant recipients. Clin Infect Dis 56:165–166
Helweg-Larsen J, Benfield TL, Eugen-Olsen J, Lundgren JD, Lundgren B (1999) Effects of mutations in Pneumocystis carinii dihydropteroate synthase gene on outcome of AIDS-associated P. carinii pneumonia. Lancet 354:1347–1351
Helweg-Larsen J, Lundgren B, Lundgren JD (2001a) Heterogeneity and compartmentalization of Pneumocystis carinii f. sp. hominis genotypes in autopsy lungs. J Clin Microbiol 39:3789–3792
Helweg-Larsen J, Lee CH, Jin S, Hsueh JY, Benfield TL, Hansen J, Lundgren JD, Lundgren B (2001b) Clinical correlation of variations in the internal transcribed spacer regions of rRNA genes in Pneumocystis carinii f.sp. hominis. AIDS 15:451–459
Huang L, Beard CB, Creasman J, Levy D, Duchin JS, Lee S, Pieniazek N, Carter JL, del Rio C, Rimland D, Navin TR (2000) Sulfa or sulfone prophylaxis and geographic region predict mutations in the Pneumocystis carinii dihydropteroate synthase gene. J Infect Dis 182:1192–1198
Hughes WT (2007) Transmission of Pneumocystis species among renal transplant recipients. Clin Infect Dis 44:1150–1151
Hughes WT (1982) Natural mode of acquisition for de novo infection with Pneumocystis carinii. J Infect Dis 145:842–848
Hughes WT, Bartley DL, Smith BM (1983) A natural source of infection due to Pneumocystis carinii. J Infect Dis 147:595
Iliades P, Meshnick SR, Macreadie IG (2005) Analysis of Pneumocystis jirovecii DHPS alleles implicated in sulfamethoxazole resistance using an Escherichia coli model system. Microb Drug Resist 11:1–8
Kazanjian P, Locke AB, Hossler PA, Lane BR, Bartlett MS, Smith JW, Cannon M, Meshnick SR (1998) Pneumocystis carinii mutations associated with sulfa and sulfone prophylaxis failures in AIDS patients. AIDS 12:873–878
Keely SP, Stringer JR, Baughman RP, Linke MJ, Walzer PD, Smulian AG (1995) Genetic variation among Pneumocystis carinii hominis isolates in recurrent pneumocystosis. J Infect Dis 172:595–598
Le Gal S, Damiani C, Rouillé A, Grall A, Tréguer L, Virmaux M, Moalic E, Quinio D, Moal MC, Berthou C, Saliou P, Le Meur Y, Totet A, Nevez G (2012) A cluster of Pneumocystis infections among renal transplant recipients: molecular evidence of colonized patients as potential infectious sources of Pneumocystis jirovecii. Clin Infect Dis 54:e62–e71. doi:10.1093/cid/cir996
Lee CH, Lu JJ, Bartlett MS, Durkin MM, Liu TH, Wang J, Jiang B, Smith JW (1993) Nucleotide sequence variation in Pneumocystis carinii strains that infect humans. J Clin Microbiol 31:754–757
Lu JJ, Lee CH (2008) Pneumocystis pneumonia. J Formos Med Assoc 107:830–842
Lu JJ, Bartlett MS, Shaw MM, Queener SF, Smith JW, Ortiz-Rivera M, Leibowitz MJ, Lee CH (1994) Typing of Pneumocystis carinii strains that infect humans based on nucleotide sequence variations of internal transcribed spacers of rRNA genes. J Clin Microbiol 32:2904–2912
Ma L, Kutty G, Jia Q et al (2002) Analysis of variation in tandem repeats in the intron of the major surface glycoprotein expression site of the human form of Pneumocystis carinii. J Infect Dis 186:1647–1654
Matos O, Lee CH, Jin S, Li B, Costa MC, Gonçalves L, Antunes F (2003) Pneumocystis jiroveci in Portuguese immunocompromised patients: association of specific ITS genotypes with treatment failure, bad clinical outcome and childhood. Infect Genet Evol 3:281–285
Meneau I, Sanglard D, Bille J, Hauser PM (2004) Pneumocystis jiroveci dihydropteroate synthase polymorphisms confer resistance to sulfadoxine and sulfanilamide in Saccharomyces cerevisiae. Antimicrob Agents Chemother 48:2610–2606
Meshnick SR (1999) Drug-resistant Pneumocystis carinii. Lancet 354:1318–1319
Miller RF, Wakefield AE (1999) Pneumocystis carinii genotypes and severity of pneumonia. Lancet 353:2039–2040
Miller RF, Ambrose HE, Wakefield AE (2001) Pneumocystis carinii f. sp. hominis DNA in immunocompetent health care workers in contact with patients with P. carinii pneumonia. J Clin Microbiol 39:3877–3882
Mori S, Cho I, Sugimoto M (2010) A cluster of Pneumocystis jirovecii infection among outpatients with rheumatoid arthritis. J Rheumatol 37:1547–1548
Nahimana A, Blanc DS, Francioli P, Bille J, Hauser PM (2000) Typing of Pneumocystis carinii f. sp. hominis by PCR-SSCP to indicate a high frequency of co-infections. J Med Microbiol 49:753–758
Nahimana A, Rabodonirina M, Zanetti G, Meneau I, Francioli P, Bille J, Hauser PM (2003a) Association between a specific Pneumocystis jiroveci dihydropteroate synthase mutation and failure of pyrimethamine/sulfadoxine prophylaxis in human immunodeficiency virus-positive and -negative patients. J Infect Dis 188:1017–1023
Nahimana A, Rabodonirina M, Helweg-Larsen J, Meneau I, Francioli P, Bille J, Hauser PM (2003b) Sulfa resistance and dihydropteroate synthase mutants in recurrent Pneumocystis carinii pneumonia. Emerg Infect Dis 9:864–867
Nevez G, Raccurt C, Jounieaux V, Dei-Cas E, Mazars E (1999) Pneumocystosis versus pulmonary Pneumocystis carinii colonization in HIV-negative and HIV-positive patients. AIDS 13:535–536
Payne SH, Loomis WF (2006) Retention and loss of amino acid biosynthetic pathways based on analysis of whole-genome sequences. Eukaryot Cell 5:272–276
Phipps LM, Chen SC, Kable K, Halliday CL, Firacative C, Meyer W, Wong G, Nankivell BJ (2011) Nosocomial Pneumocystis jirovecii pneumonia: lessons from a cluster in kidney transplant recipients. Transplantation 92:1327–1334
Pifer LL, Hughes WT, Stagno S, Woods D (1978) Pneumocystis carinii infection: evidence for high prevalence in normal and immunosuppressed children. Pediatrics 61:35–41
Pliquett RU, Asbe-Vollkopf A, Hauser PM, Presti LL, Hunfeld KP, Berger A, Scheuermann EH, Jung O, Geiger H, Hauser IA (2012) A Pneumocystis jirovecii pneumonia outbreak in a single kidney-transplant center: role of cytomegalovirus co-infection. Eur J Clin Microbiol Infect Dis 31:2429–2437
Powles MA, McFadden DC, Pittarelli LA, Schmatz DM (1992) Mouse model for Pneumocystis carinii pneumonia that uses natural transmission to initiate infection. Infect Immun 60:1397–1400
Rabodonirina M, Vanhems P, Couray-Targe S, Gillibert RP, Ganne C, Nizard N, Colin C, Fabry J, Touraine JL, van Melle G, Nahimana A, Francioli P, Hauser PM (2004) Molecular evidence of interhuman transmission of Pneumocystis pneumonia among renal transplant recipients hospitalized with HIV-infected patients. Emerg Infect Dis 10:1766–1773
Rabodonirina M, Vaillant L, Taffé P, Nahimana A, Gillibert R-P, Vanhems P, Hauser PM (2013) Pneumocystis jirovecii genotype associated with increased death rate of HIV-infected patients with pneumonia. Emerg Infect Dis 19:21–28
Ripamonti C, Orenstein A, Kutty G, Huang L, Schuhegger R, Sing A, Fantoni G, Atzori C, Vinton C, Huber C, Conville PS, Kovacs JA (2009) Restriction fragment length polymorphism typing demonstrates substantial diversity among Pneumocystis jirovecii isolates. J Infect Dis 200:1616–1622
Sassi M, Ripamonti C, Mueller NJ, Yazaki H, Kutty G, Ma L, Huber C, Gogineni E, Oka S, Goto N, Fehr T, Gianella S, Konrad R, Sing A, Kovacs JA (2012) Outbreaks of Pneumocystis pneumonia in 2 renal transplant centers linked to a single strain of Pneumocystis: implications for transmission and virulence. Clin Infect Dis 54:1437–1444
Schmoldt S, Schuhegger R, Wendler T, Huber I, Söllner H, Hogardt M, Arbogast H, Heesemann J, Bader L, Sing A (2008) Molecular evidence of nosocomial Pneumocystis jirovecii transmission among 16 patients after kidney transplantation. J Clin Microbiol 46:966–971
Stringer JR, Keely SP (2001) Genetics of surface antigen expression in Pneumocystis carinii. Infect Immun 69:627–639
Struelens MJ, Members of the European Study Group on Epidemiological Markers (ESGEM) (1996) Consensus guidelines for appropriate use and evaluation of microbial epidemiologic typing systems. Clin Microbiol Infect 2:2–11
Struijk GH, Gijsen AF, Yong SL, Zwinderman AH, Geerlings SE, Lettinga KD, van Donselaar-van der Pant KA, ten Berge IJ, Bemelman FJ (2011) Risk of Pneumocystis jiroveci pneumonia in patients long after renal transplantation. Nephrol Dial Transplant 26:3391–3398
Thomas CF Jr, Limper AH (2007) Current insights into the biology and pathogenesis of Pneumocystis pneumonia. Nat Rev Microbiol 5:298–308
Thomas S, Vivancos R, Corless C, Wood G, Beeching NJ, Beadsworth MB (2011) Increasing frequency of Pneumocystis jirovecii pneumonia in renal transplant recipients in the United Kingdom: clonal variability, clusters, and geographic location. Clin Infect Dis 53:307–308
Tipirneni R, Daly KR, Jarlsberg LG, Koch JV, Swartzman A, Roth BM, Walzer PD, Huang L (2009) Healthcare worker occupation and immune response to Pneumocystis jirovecii. Emerg Infect Dis 15:1590–1597
Totet A, Respaldiza N, Pautard JC, Raccurt C, Nevez G (2003) Pneumocystis jiroveci genotypes and primary infection. Clin Infect Dis 36:1340–1342
Totet A, Duwat H, Magois E, Jounieaux V, Roux P, Raccurt C, Nevez G (2004) Similar genotypes of Pneumocystis jirovecii in different forms of Pneumocystis infection. Microbiology 150:1173–1178
Tsolaki AG, Miller RF, Underwood AP, Banerji S, Wakefield AE (1996) Genetic diversity at the internal transcribed spacer regions of the rRNA operon among isolates of Pneumocystis carinii from AIDS patients with recurrent pneumonia. J Infect Dis 174:141–156
Valerio A, Tronconi E, Mazza F, Fantoni G, Atzori C, Tartarone F, Duca P, Cargnel A (2007) Genotyping of Pneumocystis jiroveci pneumonia in Italian AIDS patients. Clinical outcome is influenced by dihydropteroate synthase and not by internal transcribed spacer genotype. J Acquir Immune Defic Syndr 45:521–528
van Hal SJ, Gilgado F, Doyle T, Barratt J, Stark D, Meyer W, Harkness J (2009) Clinical significance and phylogenetic relationship of novel Australian Pneumocystis jirovecii genotypes. J Clin Microbiol 47:1818–1823
Vargas SL, Hughes WT, Wakefield AE, Oz HS (1995) Limited persistence in and subsequent elimination of Pneumocystis carinii from the lungs after P. carinii pneumonia. J Infect Dis 172:506–510
Vargas SL, Ponce CA, Gigliotti F, Ulloa AV, Prieto S, Muñoz MP, Hughes WT (2000) Transmission of Pneumocystis carinii DNA from a patient with P. carinii pneumonia to immunocompetent contact health care workers. J Clin Microbiol 38:1536–1538
Vargas SL, Hughes WT, Santolaya ME, Ulloa AV, Ponce CA, Cabrera CE, Cumsille F, Gigliotti F (2001) Search for primary infection by Pneumocystis carinii in a cohort of normal, healthy infants. Clin Infect Dis 32:855–861
Vargas SL, Ponce CA, Sanchez CA, Ulloa AV, Bustamante R, Juarez G (2003) Pregnancy and asymptomatic carriage of Pneumocystis jiroveci. Emerg Infect Dis 9:605–606
Vargas SL, Pizarro P, López-Vieyra M, Neira-Avilés P, Bustamante R, Ponce CA (2010) Pneumocystis colonization in older adults and diagnostic yield of single versus paired noninvasive respiratory sampling. Clin Infect Dis 50:e19–e21. doi:10.1086/649869
Vogel P, Miller CJ, Lowenstine LL, Lackner AA (1993) Evidence of horizontal transmission of Pneumocystis carinii pneumonia in simian immunodeficiency virus-infected rhesus macaques. J Infect Dis 168:836–843
Wynckel A, Toubas D, Noël N, Toupance O, Rieu P (2011) Outbreak of Pneumocystis pneumonia occurring in late post-transplantation period. Nephrol Dial Transplant 26:2417
Yazaki H, Goto N, Uchida K, Kobayashi T, Gatanaga H, Oka S (2009) Outbreak of Pneumocystis jiroveci pneumonia in renal transplant recipients: P. jirovecii is contagious to the susceptible host. Transplantation 88:380–385
Acknowledgments
Work in PMH’s laboratory is supported by the Swiss National Science Foundation grants 310030-124998 and 310030-146135. Supported in part by the Intramural Research Program of the NIH Clinical Center, National Institutes of Health, Bethesda, MD, USA.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer-Verlag Berlin Heidelberg
About this chapter
Cite this chapter
Hauser, P.M., Kovacs, J.A. (2014). 11 Molecular Epidemiology of Pneumocystis Outbreaks. In: Kurzai, O. (eds) Human Fungal Pathogens. The Mycota, vol 12. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-39432-4_11
Download citation
DOI: https://doi.org/10.1007/978-3-642-39432-4_11
Published:
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-642-39431-7
Online ISBN: 978-3-642-39432-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)