Abstract
Matching a regular expression (regex) on a text is widely used in many applications, such as text editing, information extraction and instruction detection (IDS). Traditional algorithms generally compile an equivalent automaton from the regex query, then run it on the text to find all matching results. However, they have to scale linearly with the size of the text. Recent algorithms utilize various filtering techniques to quickly jump to candidate positions in a text where a matching result may appear, then only these candidate positions are verified by the automaton. In this paper, we give a full specification on filtering techniques for the regex matching problem, in which filters for the regex query can be classified into positive factor and negative factor. We review three typical positive factors, including prefix, suffix, and necessary factor and show that negative factors can collaborate with positive factors to significantly improve the filtering ability.
The work is partially supported by the National Natural Science Foundation of China (Nos. 61572122, U1736104, 61532021).
The original version of this chapter was revised: For detailed information please see correction chapter. The correction to this chapter is available at https://doi.org/10.1007/978-3-319-91455-8_24
Access provided by CONRICYT-eBooks. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Regular expression (regex) matching is a fundamental problem that exists in many applications, such as text editing, information extraction, protein sequence matching and instruction detection (IDS). For example, in the domain of bioinformatics, a regex query \(\texttt {TC(T|G)(C|T)A}\) has the language \(\{\texttt {TCTCA,TCTTA,TCGCA,TCGTA}\}\), matching this regex is to find all matchings of the string in this language from a genome sequence.
The classical approaches to match a regex query in a text is that first transforming the regex into an equivalent automaton, and then running it from each position in the text to verify if the substring is an occurrence of the regex. An occurrence is found whenever a final state of the automaton is reached [1, 5, 8]. \(\mathsf {NFAThompson}\) [8] and \(\mathsf {DFAClassical}\) [1] are two typical automaton-based algorithms. \(\mathsf {NFAThompson}\) is the pioneering work that proposes the Thompson NFA to match a regex with time complexity O(mn), where m is the size of the regex query and n is the length of the text. \(\mathsf {DFAClassical}\) realizes regex matching by simulating the DFA, which can guarantee a linear search time of O(n). However, the automaton-based algorithms have to check every character in a text, the matching efficiency is largely limited.
To improve query efficiency, filtering techniques have been proposed for many applications which focus on producing a set of candidates which could be the final query results [3, 4, 7, 10, 13, 14]. To alleviate the above issue in the regex matching problem, many algorithms have been developed under a filtering-and-verification framework, where candidate positions are generated using one or more filters and then verified by an automaton to find the true matching positions [7, 11]. The filters can be divided into two types. The first one, called positive factor, utilizes the substrings extracted from the regex query, including prefix, suffix and necessary factor. \(\mathsf {MultiStringRE}\) [9] computes a set of prefixes for all strings matching the regex query (i.e., the language of a regex), then uses a Commentz-Water-like algorithm to verify the text starting from each occurrence of these prefixes. \(\mathsf {NRGrep}\) [6] gets the candidate positions using the reversed prefixes of the regex and verifies them using a reversed automaton. \(\mathsf {GNU}\) \(\mathsf {grep}\) [2] utilizes the necessary factors to get candidate positions, which are the substring must appear in a regex match. Since a necessary factor could divide a regex into a left and a right part, two automatons are constructed to verify a candidate position in forward and backward directions. The other one is called negative factor and initially proposed in [12], which is the substring that cannot appear in any matching string of a regex. Negative factors can further prune the candidate positions generated by the positive factors.
In this paper, we give a full specification on filtering techniques for the regex matching problem and show different filters of the regex can be used together to improve the filtering ability.
2 Filtering-Based Regular Expression Matching
Let \(\varSigma \) be a finite alphabet. A regular expression (regex) Q is a string over \(\varSigma \cup \{\epsilon ,|,\cdot ,*,(,)\}\), in which \(\{|,\cdot ,*\}\) are the operators that represents disjunction, conjunction and Kleene closure (repeating unit), respectively. We use L(Q) to represent the language of a regex Q. For a text T of the characters in \(\varSigma \), we use |T| to denote its length, T[i] to denote its i-th character (starting from 0), and T[i, j] to denote the substring ranging from its i-th character to its j-th character.
Regular Expression Matching. Given a regex Q and a text T, the regex matching problem is to find matching occurrences of the strings in L(Q) from T.
In the following, we first review the filtering techniques with positive factors, then show negative factors can collaborate with positive factors.
2.1 Computing Candidate Positions Using Positive Factors
Recent techniques have utilized certain features of the regex Q to improve the performance of automaton-based methods [7]. Their main idea is to use positive factors, which are substrings of Q, to identify candidate positions of Q in T. Next, we present three typical positive factors, including prefix, suffix, and necessary factor.
A prefix w.r.t. a regex Q is defined as a prefix of a string in the language L(Q). We use \(l_{pre}\) to denote the length of a prefix. A set of prefixes \(S_P\) can be used as the filters of a regex if and only if there is a prefix in \(S_P\) for any string in L(Q) [9]. For example, for the regex \(Q=\texttt {(A|G)}{} \texttt {T}^*\texttt {AT}^*\texttt {G}\), the prefixes with \(l_{pre}=2\) are \(\{\texttt {AT},\texttt {AA},\texttt {GT},\texttt {GA}\}\). Due to any matching string of Q must start with a prefix in \(S_P\), then the matching positions of the prefixes in \(S_P\) on T are the candidate positions for Q. To compute all matches of Q, we only examine these matching positions of prefixes using the automaton of Q.
Similarly, a suffix w.r.t. a regex Q is defined as a suffix of a string in L(Q), and the length is denoted by \(l_{suf}\). We use \(S_S\) to represent the set of suffixes computed from Q, e.g., for the regex \(Q=\texttt {(A|G)}{} \texttt {T}^*\texttt {AT}^*\texttt {G}\), the suffixes with \(l_{suf}=2\) are \(\{\texttt {TG},\texttt {AG}\}\). Different from prefixes, the ending matching positions of suffixes in \(S_S\) are candidate positions, which are verified by a reversed automaton in the backward direction [6].
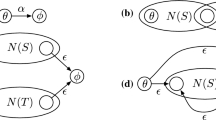
In addition to prefixes and suffixes, the necessary factor is another type of positive factor, which is a substring that must appear in every matching string in L(Q) [2]. For instance, for the regex \(Q=\texttt {(A|G)}{} \texttt {T}^*\texttt {AT}^*\texttt {G}\), \(\{\texttt {A}\}\) is a necessary factor of Q. To verify a candidate position where a necessary factor appears, we can divide Q into a left part and a right part with a corresponding automaton, e.g., two automatons are constructed for the left and right parts of the regex Q (i.e., \(\texttt {(A|G)}{} \texttt {T}^*\) and \(\texttt {AT}^*\texttt {G}\)).
Instead of independently applying each positive factor, all three types of positive factors can also be leveraged together to further improve the filtering ability [11]. \(\mathsf {PS}\) and \(\mathsf {PMS}\) are two typical patterns used to identify candidate positions. \(\mathsf {PS}\) pattern utilizes prefix and suffix which requires a candidate occurrence contains the matchings of a prefix and a suffix simultaneously in T. Likewise, \(\mathsf {PMS}\) pattern requires a candidate occurrence contains all matchings of the three positive factors. Generally, \(\mathsf {PMS}\) pattern can achieve better filtering ability than \(\mathsf {PS}\) pattern since one more positive factor is considered, but it also needs more computational cost for filtering.
Consider the example in Fig. 1, there is a matching result T[6, 10] for the regex \(Q=\texttt {(A|G)}{} \texttt {T}^*\texttt {AT}^*\texttt {G}\). Using prefixes of Q as filters, there are 6 candidate occurrences needed to be verified. \(\mathsf {PS}\) and \(\mathsf {PMS}\) further prune the candidate occurrences when considering more positive factors, and obtain 5 and 4 candidate occurrences, respectively.
2.2 Further Pruning Candidate Positions Using Negative Factors
Although positive factors can be used together to compute candidate occurrences, compared to the single type of positive factors, using more than one type of positive factors obtains few improvements in the filtering ability. Negative factors solve this problem.
A negative factor (also called N-factor) w.r.t. a regex Q is a string w such that there is no string \(\varSigma ^*w\varSigma ^*\) in L(Q) [11, 12]. For example, for the running example, \(\texttt {C}\) is an N-factor since any string in L(Q) does not contain \(\texttt {C}\). Essentially, N-factor is the substring that does not appear in any matching string of Q. Based on this property, given a set of N-factors of Q, a text T can be divided into several disjoint segments and we can get the matching result of Q can only appear within a segment.
At first, we show N-factors can be integrated into the \(\mathsf {PS}\) pattern. According to the definition of N-factor, a candidate occurrence must start with a prefix and end with a suffix, and do not contain any matching of N-factor. We call such candidate occurrences satisfy the \(\mathsf {PNS}\) pattern. For example, as shown in Fig. 2, candidate occurrences T[0, 16] and T[10, 16] obtained by \(\mathsf {PS}\) pattern can be pruned by the \(\mathsf {PNS}\) pattern since they contain the matching of N-factor \(\texttt {C}\).
Similarly, we can get the \(\mathsf {PMNS}\) pattern by integrating N-factors into \(\mathsf {PMS}\) pattern, which requires a candidate occurrence contains the matchings of necessary factors based on the requirements of the \(\mathsf {PNS}\) pattern. Because \(\mathsf {PMNS}\) considers the requirements of all filters computed from the regex, it achieves the best filtering ability. For the example in Fig. 2, compared to the \(\mathsf {PNS}\) pattern, the candidate occurrence T[13, 16] can be further pruned by the \(\mathsf {PMNS}\) pattern since it does not contain the matching of \(\texttt {A}\).
3 Conclusion and Future Work
Regular expression matching is a fundamental problem existing in a diverse range of applications. In this paper, we introduced the filtering techniques for the regex matching problem, in which filters of the regex query can be classified into positive factor and negative factor. We reviewed three typical positive factors, including prefix, suffix, and necessary factor and showed they can be used together to compute candidate occurrences. Furthermore, we showed negative factors can collaborate with positive factors to significantly improve the filtering ability. As parts of future work, we will (i) further investigate the correlation between different filters extracted from the regex query; (ii) balance the filtering cost caused by different filters.
Change history
01 December 2018
The original version of the chapter starting on p. 118 was revised.
References
Aho, A.V., Sethi, R., Ullman, J.D.: Compilers - Principles, Techniques and Tools. Addison-Wesley, Reading (1986)
GNUgrep: ftp://reality.sgiweb.org/freeware/relnotes/fw-5.3/fw_gnugrep/gnugrep.html
Li, B., Yang, X., Wang, B., Cui, W.: Efficiently mining high quality phrases from texts. In: AAAI, pp. 3474–3481 (2017)
Li, B., Yang, X., Zhou, R., Wang, B., Liu, C., Zhang, Y.: An efficient method for high quality and cohesive topical phrase mining. TKDE (2018, to appear)
Mohri, M.: String matching with automata. Nord. J. Comput. 4(2), 217–231 (1997)
Navarro, C.: NR-grep: a fast and flexible pattern matching tool. Softw. Pract. Exp. (SPE) 31, 1265–1312 (2001)
Navarro, C., Raffinot, M.: Flexible Pattern Matching in Strings: Practical On-line Search Algorithms for Texts and Biological Sequences. Cambridge University Press, Reading (1979)
Thomphson, K.: Regular expression search algorithm. Commun. ACM 11, 419–422 (1968)
Watson, B.W.: A new regular grammar pattern matching algorithm. In: Diaz, J., Serna, M. (eds.) ESA 1996. LNCS, vol. 1136, pp. 364–377. Springer, Heidelberg (1996). https://doi.org/10.1007/3-540-61680-2_68
Yang, X., Liu, H., Wang, B.: ALAE: accelerating local alignment with affine gap exactly in biosequence databases. PVLDB 5(11), 1507–1518 (2012)
Yang, X., Qiu, T., Wang, B., Zheng, B., Wang, Y., Li, C.: Negative factor: improving regular-expression matching in strings. ACM Trans. Database Syst. (TODS) 40(4), 25 (2016)
Yang, X., Wang, B., Qiu, T., Wang, Y., Li, C.: Improving regular-expression matching on strings using negative factors. In: SIGMOD, pp. 361–372, June 2013
Yang, X., Wang, B., Yang, K., Liu, C., Zheng, B.: A novel representation and compression for queries on trajectories in road networks. TKDE 30(4), 613–629 (2018)
Yang, X., Wang, Y., Wang, B., Wang, W.: Local filtering: improving the performance of approximate queries on string collections. In: SIGMOD, pp. 377–392. ACM (2015)
Acknowledgment
This paper constituted an invited talk, held at BDQM 2018, a DASFAA 2018 satellite workshop. The main techniques derive from our work cited in [11].
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this paper
Cite this paper
Qiu, T., Yang, X., Wang, B. (2018). Filtering Techniques for Regular Expression Matching in Strings. In: Liu, C., Zou, L., Li, J. (eds) Database Systems for Advanced Applications. DASFAA 2018. Lecture Notes in Computer Science(), vol 10829. Springer, Cham. https://doi.org/10.1007/978-3-319-91455-8_12
Download citation
DOI: https://doi.org/10.1007/978-3-319-91455-8_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-91454-1
Online ISBN: 978-3-319-91455-8
eBook Packages: Computer ScienceComputer Science (R0)