Abstract
In the last decade, the nuclear envelope (NE) has emerged as an important regulator of genome architecture and a central player in gene expression regulation. Nuclear pore complexes (NPCs), the channels that penetrate the NE connecting the nucleus to the cytoplasm, are the largest protein complexes of the NE. Built by multiple copies of roughly 30 different proteins, NPCs were traditionally studied for their role in controlling nucleocytoplasmic transport. But accumulating evidence shows that these massive molecular structures play multiple transport-independent roles that are key for the maintenance of cellular physiology and tissue homeostasis. In this chapter, we will focus on the current knowledge of the role of mammalian NPCs in the regulation of genome organization and gene expression. The recent findings showing that NPCs regulate the activity of specific genes either at the nuclear periphery or inside the nucleus point towards these structures as critical controllers of genome function. Deciphering the molecular mechanism employed by NPCs to modulate specific gene expression programs and to maintain genome integrity are our main challenges for the next decade.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
7.1 Introduction
In eukaryotic cells the nucleus is the organelle where the genome is housed. Discovered in 1833 by botanist Robert Brown (Oliver 1913), the nucleus serves as the control center of the cell where all the genetic information is stored and translated. The nucleus is characterized by a double-membrane structure, known as the nuclear envelope (NE), that separates the chromosomes from the cytoplasm. In metazoans, the nuclear lamina, which is a filamentous protein meshwork situated underneath the NE, provides mechanical stability to the nucleus and aids in the regulation of many nuclear processes, including genome organization and gene expression regulation (Stancheva and Schirmer 2014). The NE is perforated by large multiprotein channels known as nuclear pore complexes or NPCs. Discovered in 1950 (Callan and Tomlin 1950), NPCs represent the sole gateway into the nucleus and, thus, they are responsible for the entry and exit of most molecules from and into this compartment. Although NPCs have been historically studied for their essential role in controlling nucleocytoplasmic molecule exchange, they have recently emerged as important regulators of diverse cellular processes in a transport-independent manner (Raices and D’Angelo 2012). One of the most studied functions of NPCs is their role in the regulation of genome integrity. Increasing evidence supports a model in which NPCs not only act as organizers of the cellular genome but also as scaffolds for the regulation of specific gene groups confined to the nuclear periphery. Moreover, several nucleoporins have been now found to localized to the nuclear interior where they assist the transcriptional machinery in regulating gene activity. In this chapter, we describe our current knowledge of the role of mammalian NPCs in the regulation of genome organization and gene expression. While our understanding of these functions of NPCs in mammalian cells is not yet as extensive as in yeast, increasing evidence indicates a strong conservation in the genomic processes and mechanisms regulated by NPCs between these organisms.
7.2 Stability, Mobility and Lifespan of Mammalian NPCs
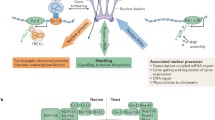
NPCs are one of the largest protein complexes of eukaryotic cells (Raices and D’Angelo 2012). These channels have an eight-fold-symmetrical structure that consists of a nuclear envelope-embedded scaffold that is built to surround a central transport channel through which all nucleocytoplasmic transport takes place (Frenkiel-Krispin et al. 2010; Beck et al. 2004, 2007; Maimon et al. 2012). Attached to this scaffold are two rings, the cytoplasmic and nuclear rings, from where eight filaments emanate (Fig. 7.1a). On the nuclear side, the filaments are joined in a distal ring to form the nuclear basket of the NPC. Even though the overall structure of NPC is conserved among species, its size varies, being the mammalian NPCs the largest complexes of all (estimated molecular mass = 60–125 MDa) (Suntharalingam and Wente 2003; Yang et al. 1998). Interestingly, even though NPCs are massive protein complexes, these channels are composed of roughly 30 different proteins known as nucleoporins or Nups (Rout et al. 2000; Cronshaw et al. 2002; Yang et al. 1998; Reichelt et al. 1990; Hoelz et al. 2016) (Fig. 7.1b). Despite being a membrane embedded structure, most NPCs components are soluble proteins and in mammalian cells only three nucleoporins are transmembrane (D’Angelo and Hetzer 2008). Most nucleoporins have been found to associate in biochemically stable subcomplexes that are believed to act as the building blocks of nuclear pores (Fig. 7.1b) (Hoelz et al. 2016). Due to the eight-fold rotational symmetry of these structures, nucleoporins and subcomplexes are present in eight or multiples of eight copies, and each pore has an estimated 500–1000 total proteins. In addition to their localization within the NPC structure, nucleoporins have also been classified depending on their residence time (Rabut et al. 2004). Stable nucleoporins are those that show very low exchange rates from NPCs and are mostly components of the pore scaffold. Dynamic nucleoporins, on the other hand, are constantly exchanged from NPCs, having residence times at the structure that range from a few seconds to a few hours (Rabut et al. 2004). These dynamic nucleoporins are mostly members of the pore peripheral structures that include the nuclear basket, central channel and filaments.
Mammalian cells divide through open mitosis, a process in which the nucleus is disassembled during the M-phase of the cell cycle to allow the separation of sister chromatids (Kutay and Hetzer 2008). In each cell division, when the nucleus breaks down , NPCs are disassembled in their stable subcomplexes, which are recycled at the end of M-phase to assemble new channels at the daughters’ nuclear envelopes (D’Angelo and Hetzer 2008). This disassembly–reassembly cycle ensures that in dividing mammalian cells NPCs are renewed in each cell-division. But NPCs behave differently in non-dividing and postmitotic cells (D’Angelo et al. 2009). As mentioned above, the scaffold components of NPCs have very long residence times at this structure. In fact, studies in dividing cells have identified that these nucleoporins have residence times at NPCs that are longer than the cell cycle (Rabut et al. 2004; Daigle et al. 2001). This provided the first evidence that the core NPC proteins would only exchange when pores disassemble during mitosis. Support for this model came from recent studies showing that scaffold nucleoporins indeed have extremely long lives at NPCs of postmitotic cells (D’Angelo et al. 2009; Savas et al. 2012). Analysis of nucleoporin turnover in postmitotic cells and tissues uncovered extremely low rates of exchange for these nucleoporins, suggesting minimal turnover of NPC structural components. These findings indicate that when mammalian cells exit the cell cycle they maintain their NPC scaffold structures for almost their entire life (D’Angelo et al. 2009; Savas et al. 2012; Toyama et al. 2013). This long life of NPCs does not come without cost, and nuclear pores have been shown to deteriorate as postmitotic cells age, leading to the loss of nuclear compartmentalization in old cells (D’Angelo et al. 2009).
Mammalian NPCs are not only incredibly stable structures at the NE, they are also immobile. Studies using Fluorescent Recovery After Photobleaching (FRAP) have identified that NPCs do not move independently at the NE (Daigle et al. 2001). In fact, in the same way as the nuclear lamina, nuclear pores move as large arrays in response to changes in nuclear shape (Daigle et al. 2001). The low turnover of NPCs, their large size and potential anchor sites for DNA and proteins, and their lack of mobility at the nuclear periphery suggest that these structures play a role in nuclear organization by acting as stationary, long-lived, positional markers at the NE (D’Angelo and Hetzer 2008).
7.3 Chromatin Interactions with Mammalian NPCs
It is has now become clear that chromosomes are not randomly dispersed inside the nucleus (Misteli 2007). A significant amount of data shows that genes and chromosomal domains have unique relative positons within the nucleus of different cell types, and several studies have demonstrated that intranuclear gene position can affect gene activity/regulation (Nguyen and Bosco 2015; Talamas and Capelson 2015; Stancheva and Schirmer 2014). These findings have uncovered that the three-dimensional (3D) organization of the genome plays a key role in gene expression control. However, how genome architecture is faithfully maintained in mammalian cells remains poorly comprehended.
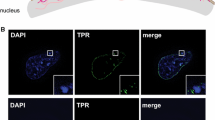
The first studies of the association of mammalian NPCs with the genome identified that the Nup93 nucleoporin associates with chromatin regions enriched in heterochromatin markers (Brown et al. 2008). Nup93 is a scaffold component of NPCs that shows a low exchange rate from nuclear pores during interphase and plays a key role in the maintenance of the nuclear permeability barrier (Rabut et al. 2004; Galy et al. 2003; D’Angelo et al. 2009). The finding that Nup93 associates with heterochromatin fueled the original idea that the nuclear periphery was a repressive environment mostly associated with chromatin condensation and gene silencing (Towbin et al. 2009). Interestingly, in this original study, the genome regions associated with NPCs were found to change when global histone acetylation was modified, indicating that NPC-genome interactions are dynamic (Brown et al. 2008). More recent studies have identified that NPCs also bind active genes (Kehat et al. 2011; Raices et al. 2017), open chromatin domains (Ibarra et al. 2016), and enhancer regions (Ibarra et al. 2016), which is more consistent with the long-time observation by electron microscopy that differently from the nuclear lamina, NPCs are surrounded by euchromatin (Lemaitre and Bickmore 2015; Capelson and Hetzer 2009) (Fig. 7.2). These studies point to NPC surroundings as regions of decondensed, transcription-permissive, chromatin, and suggest that these channels play a role in the positive regulation of gene expression. Notably, the maintenance of these NPC-associated decondensed domains, also known as heterochromatin exclusion zones, has been shown to depend on the nuclear basket nucleoporin Tpr (Krull et al. 2010). This indicates that nuclear pores have an active role in regulating the state of their surrounding chromatin. The direct binding of different genomic regions, their active role in the regulation of the neighboring chromatin environment, and the fact that mammalian NPCs are immobile due to their interactions with the nuclear lamina, support the current model in which these large structures play a critical role in genome architecture.
Chromatin organization around nuclear pore complexes Electron micrograph image of a resting T lymphocyte showing the distribution of euchromatin (light gray) and heterochromatin (dark gray) within the nucleus. Right panel: Schematic representation of a zoomed-in region of the nuclear envelope depicting NPCs and their surrounding decondensed chromatin
7.4 Gene Expression Regulation at NPCs
7.4.1 Genes that Associate with Mammalian NPCs
In budding yeast, many genes have been found to relocate to NPCs when activated, and it is well established that NPC-gene association is important for their efficient expression as well as for their transcriptional memory (Schneider et al. 2015; Sood and Brickner 2014; D’Urso and Brickner 2017). In mammals, the regulation of gene expression by NPCs is just beginning to be exposed, and emerging evidence suggest that similar to its yeast counterparts, nuclear pores play a critical role in the regulation of gene expression at the nuclear periphery. The first description of active genes requiring NPC-association for their efficient transcription came from a study of cardiomyocyte hypertrophic growth (Kehat et al. 2011). This work showed that when cardiomyocyte hypertrophic growth is induced, the proper transcription of several genes, including sarcomeric and calcium-handling genes, requires their relocation to NPCs (Kehat et al. 2011) (Fig. 7.3a). Similarly, the regulation of multiple structural and contraction genes in skeletal muscle has been recently shown to take place at NPCs and to require the presence of a tissue-specific nucleoporin known as Nup210 (Fig. 7.3b). Nup210 was the first nucleoporin identified (Gerace et al. 1982). This transmembrane protein shows cell type- and tissue-specific expression (Olsson et al. 1999, 2004; D’Angelo et al. 2012). By regulating gene expression, Nup210 has been shown to be important for the differentiation of muscle progenitors and for the maturation and survival of differentiated muscle cells (D’Angelo et al. 2012; Raices et al. 2017). In the absence of this nucleoporin, the activity of many structural genes becomes misregulated, although their association with NPCs is not affected (Raices et al. 2017). The findings of these studies indicate that in cardiac and skeletal muscle the positioning of specific genes at NPCs is critical for their proper regulation and reveal that NPCs act as scaffolds for the regulation of specific gene groups. Notably, Nup210 is not expressed in muscle progenitor cells (Raices et al. 2017; D’Angelo et al. 2012). Its expression is induced and Nup210 is added to NPCs during differentiation (Raices et al. 2017; D’Angelo et al. 2012). This indicates that gene regulation at NPCs can be modulated by changing the composition of this structure (Fig. 7.3b).
Gene regulation at NPCs (a) In resting cardiomyocytes, the histone deacetylase 4 (HDAC4) binds to Nup155 at NPCs and prevents the association of sarcomeric and calcium-handling genes with nuclear pores. When hypertrophic growth is induced in these cells, HDAC4 is exported from the nucleus, which allows the association of muscle genes with NPCs, promoting their proper activation. (b) During myoblast differentiation, the expression of the transmembrane nucleoporin Nup210 is induced. Nup210 is added to NPCs of differentiating muscle cells, where it recruits the transcription factor Mef2C to regulate the activity of muscle structural genes that were already associated with nuclear pores. (c) In mammalian cells, Nup98 has been shown to bind inactive genes (gray) or genes that have basal/low (light pink) activity at NPCs. (d) Analysis of the genome domains that associate with nucleoporins Nup93 and Nup153 uncovered an enrichment of superenhancer sequences. The binding to superenhancerrs to Nup93 and Nup153 was found to take place at NPCs. (e) Members of the Nup93-Nup205 NPC subcomplex, including Nup93, Nup188 and Nup205, associate with HoxA genes and repress their activity at NPCs. In this figure, silent genes are shown in gray, active genes in pink
The association of genes with NPCs is not restricted to muscle cells and has also been described during neuronal differentiation. Chromatin immunoprecipitation (ChIP) studies of the Nup98 nuclear pore complex member showed that this nucleoporin binds many developmental and cell differentiation genes (Liang et al. 2013). Nup98 is a phenylalanine-glycine (FG) repeat-containing dynamic nucleoporin that also localizes to the nuclear interior (Griffis et al. 2002, 2003). It is expressed from the NUP98 gene as a 98 kDa protein or as a larger 195-kDa precursor encoding Nup98-Nup96 that is autoproteolytically cleaved to produce both nuclear pore complex components (Fontoura et al. 1999; Ratner et al. 2007). At NPCs, Nup98 localizes at the cytoplasmic and nuclear sides (Griffis et al. 2003). Inside the nucleus, the localization of Nup98 varies among different cell types, but cells that have high levels of this nucleoporin, as well as cells overexpressing it, show its accumulation in intranuclear foci known as GLFG bodies, due to the glycine-lysine-phenylalanine-glycine repeats that Nup98 contains (Griffis et al. 2002, 2003). The function of Nup98-containing intranuclear foci is yet to be elucidated. A large amount of evidence accumulated in the past decade indicates that Nup98 plays an important role in the regulation of gene expression at NPCs and inside the nucleus (Franks and Hetzer 2013). During the early stages of neuronal differentiation Nup98 has been shown to bind a subset of non-active or low activity genes at NPCs (Liang et al. 2013) (Fig. 7.3c). On the other hand, the binding of Nup98 to genes that are highly activated during this process occurs in the nucleoplasm and away from NPCs as described below (see Sect. 7.5)(Liang et al. 2013).
Consistent with NPCs acting as hubs for transcriptional regulation, a recent study identified an enrichment of superenhancer sequences within the genomic regions associated with nuclear pores (Ibarra et al. 2016) (Fig. 7.3d). Superenhancers are domains of the genome that contain clusters of enhancers in close proximity (Niederriter et al. 2015) and generally play a role in the regulation genes involved in cell identity/cell type specification. This study found that depletion of Nup153 or Nup93 dramatically affects the transcription of genes regulated by NPC-associated superenhancers (Ibarra et al. 2016) (Fig. 7.3d).
Even though these studies have identified a positive role of NPCs in transcriptional regulation, the Nup93 nucleoporin was found to bind genome regions enriched in silent chromatin markers (Brown et al. 2008). This suggests a role for NPCs in gene repression. Consistent with this idea, a recent study uncovered that several members of the Nup93-Nup205 NPC subcomplex (including Nup93, Nup188 and Nup205) bind to the promoter of HOXA genes and silence their expression (Labade et al. 2016) (Fig. 7.3e). Fluorescent in situ hybridization analyses of the genes regulated by these nucleoporins confirmed their association with the nuclear periphery and uncovered that Nup93 is required for the tethering of HOXA genes to NPCs (Labade et al. 2016).
All these studies demonstrate that similar to the yeast NPCs, mammalian NPCs can bind active and silent chromatin, and act as positive or negative regulators of gene expression. But more importantly, that mammalian NPCs are key players in the regulation of developmental and cell type-specific gene expression by acting as protein scaffolds that allow the local regulation of specific genes confined to the nuclear periphery.
7.4.2 Mechanisms of Gene-Expression Regulation by NPCs
A large amount of evidence has shown that in yeast the dynamic association of genes with NPCs is regulated by transcription factors (Randise-Hinchliff et al. 2016; Brickner et al. 2012). Transcription factors not only regulate NPC-tethering in these organisms but they are also required for the clustering of genes that present their binding sites (Randise-Hinchliff et al. 2016; Brickner et al. 2012). The role of transcription factors in regulating gene expression at NPCs has also been described in flies, where the Ecdysone receptor, a nuclear hormone receptor, is recruited to NPCs upon stimulation to regulate the activity of NPC-associated genes (Pascual-Garcia et al. 2017). How NPCs regulate gene expression at the nuclear periphery in mammals is just starting to be unraveled. Recent findings showed that the nucleoporin Nup210 recruits the transcription factor Mef2C to NPCs to modulate the activity of several muscle structural genes (Raices et al. 2017). This indicates that the role of transcription factors in regulating gene expression at the nuclear periphery might be conserved in mammals. But even though Nup210 is important for the efficient expression of NPC-associated genes, it is not required for gene localization to the nuclear periphery (Raices et al. 2017). Interestingly, during cardiomyocyte hypertrophic growth, the association of genes with NPCs is negatively modulated by the histone deacetylase HDAC4 (Kehat et al. 2011). In these cells, when HDAC4 is anchored to NPCs, it prevents the association and transcription of several sarcomeric genes (Kehat et al. 2011). When hypertrophic growth is stimulated, the release of HDAC4 from NPCs leads to the recruitment and activation of these genes. Because HDAC4 is a key negative regulator of Mef2C activity that is exported from the nucleus during myogenesis (Clocchiatti et al. 2013; McKinsey et al. 2000), these findings indicate that the interplay between transcription factors and chromatin modulators might regulate NPC-gene association and gene expression regulation. Altogether, the existing data allows to propose that: (1) by interacting and recruiting transcriptional modulators, including transcription factors and chromatin regulators, (2) by concentrating super-enhancers in their vicinity, and (3) by tethering genes that share common regulatory domains, NPCs act as hubs for the transcriptional regulation of specific gene groups at the nuclear periphery.
7.5 Gene Expression Regulation by Nucleoporins in the Nuclear Interior
7.5.1 Genes that Associate with Intranuclear Nucleoporins
Despite the emerging evidence that the NPC structure is itself important for gene expression regulation, most nucleoporin-genome interactions described so far in metazoans take place in the nuclear interior and away from NPCs (Kalverda et al. 2010; Capelson et al. 2010; Liang et al. 2013) (Fig. 7.4). This phenomenon was initially identified in flies, where several nucleoporins were found to bind chromatin inside the nucleus and to play an important role in the regulation of developmental gene expression (Kalverda et al. 2010; Capelson et al. 2010). In mammalian cells, a few nucleoporins have also been shown to regulate gene expression inside the nucleus. The clearest example is Nup98 (Fig. 7.4). As mentioned, Nup98 is a dynamic nucleoporin that shuttles between NPCs and the nuclear interior (Griffis et al. 2002). In mammalian cells, the mobility of Nup98 inside the nucleus has been shown to be transcription-dependent (Griffis et al. 2002), and this nucleoporin has been found to bind developmentally regulated genes during embryonic stem cell (ESC) to neuronal differentiation (Liang et al. 2013). The genes bound by Nup98 in ESCs, which include active cell cycle and nucleic acid metabolism genes as well as some silent genes, differ from the ones bound in neuroprogenitors, which are mostly genes that are activated during neural differentiation (Liang et al. 2013). These findings indicate that Nup98 genome association is cell type-specific and developmentally regulated. Consistent with this idea, in lung fibroblasts Nup98 was found to associate with silent chromatin domains (Liang et al. 2013). Functionally, inhibition of Nup98 activity by expression of a dominant negative mutant has been found to affect the expression of the Nup98-bound developmental genes (Liang et al. 2013). Interestingly, two modes of gene regulation during neuronal differentiation have been described for Nup98. This nucleoporin has been shown to bind genes that are in the initial stage of induction at NPCs (on-pore), while the association of Nup98 with genes that are strongly induced during neuronal differentiation has been found to take place in the nuclear interior (off-pore) (Liang et al. 2013) (Figs. 7.3 and 7.4). These findings indicate that nuclear pore complex components might modulate different genes/gene expression programs depending on their spatial location within the nuclear space.
Gene regulation by nucleoporins inside the nucleus. In mammalian cells, several nucleoporins have been found to regulate gene expression away from NPCs. (a) Nup98 has been found to bind multiple genes, including cell cycle, metabolic, cell differentiation, developmental, and INFγ-regulated genes. Some of these genes are co-bound by a soluble version of the transmembrane nucleoporin Pom121 (sPom121) (b). (c) Nup153 has also been shown to bind several developmental/differentiation genes in the nuclear interior. But contrary to Nup98, Nup153 recruits the polycomb complex PRC1 to their promoters to repress their activity in pluripotent stem cells. (d) The oncogenic Nup98-HoxA9 fusion has been found to induce the abnormal expression of HOX genes by recruiting different chromatin modulators to HOX clusters, including NSL, MLL, HDAC1 and CBP/p300. The association of this fusion protein with HOX genes is mediated by the export receptor CRM1, which has been found to be prebound to these genes
A detailed analysis of the DNA bound by Nup98 in mammals showed enrichment for GA-box DNA motifs. GA repeat motifs are bound by the GAGA factor (Liang et al. 2013), which in Drosophila and mammals regulates boundary activity at HOX clusters (Srivastava et al. 2015; Adkins et al. 2006; Granok et al. 1995) and modulates the expression of homeotic genes (Adkins et al. 2006; Granok et al. 1995). These findings might help to explain why several of the abnormal Nup98 fusion proteins that result from chromosomal translocations result in the alteration of HOX gene expression (see Sect. 7.6).
In addition to its role in regulating the activity of developmental genes, the binding of Nup98 to interferon gamma (INFγ) target genes is required for transcriptional memory (Light et al. 2013). In human cells, many genes that are induced by INFγ retain a “memory” of the activation and are turned on at a faster rate if cells are re-exposed to the cytokine. This transcriptional memory is maintained for several generations and depends on epigenetic modifications (D’Urso and Brickner 2017). Nup98 has been found to be recruited to the promoter of several INFγ target genes, such as HLA-DRA, only after removal of the cytokine (Light et al. 2013). The association of Nup98 with the promoter of these recently expressed genes is required for their proper re-activation upon INFγ re-exposure (transcriptional memory) (Light et al. 2013). The function of Nup98 in transcriptional memory is conserved in yeast, as its homologue Nup100 is required for the transcriptional memory of the inositol-responsive gene INO1 (Light et al. 2013; D’Urso and Brickner 2014). But differently from yeast, the regulation of transcriptional memory by nucleoporins in human cells takes place inside the nucleus and not at NPCs (Fig. 7.4). Interestingly, ChIP studies using the antibody mAb414, which recognizes the Nup62, Nup153, Nup214 and Nup358 nucleoporins, showed that one or more of these nucleoporins also associate with the HLA-DRA locus. But differently from Nup98, this association is also observed when the gene is activated by INFγ (Light et al. 2013). What role does the association of the mAb414-recognized nucleoporins with this gene play in its transcriptional regulation remains to be determined.
The NPC Nup153 member is a main component of the nuclear basket (Sukegawa and Blobel 1993; Pante et al. 1994), and another dynamic nucleoporin that shows transcription-dependent mobility (Griffis et al. 2004). Nup153 has been found to interact with the nuclear lamina (Al-Haboubi et al. 2011; Smythe et al. 2000) and has been associated with intranuclear filaments of the Tpr nucleoporin that emanate from NPCs into the nuclear interior (Hase and Cordes 2003; Simon and Wilson 2011). Nup153 has several zinc fingers motifs in its N-terminal region (Sukegawa and Blobel 1993) and has been shown to bind DNA (Sukegawa and Blobel 1993) and RNA (Ullman et al. 1999; Dimaano et al. 2001; Ball et al. 2007). It also has multiple FG repeats in its C-terminal domain through which it interacts with transport receptors (Shah et al. 1998; Moroianu et al. 1995; Nakielny et al. 1999). Several functions have been attributed to Nup153. These include the regulation of mRNA export (Bastos et al. 1996; Ullman et al. 1999), importin α/β-mediated nuclear import (Walther et al. 2001; Shah and Forbes 1998; Ogawa et al. 2012; Makise et al. 2012), NPC assembly (Walther et al. 2001; Vollmer et al. 2015), mitotic checkpoint regulation (Mackay et al. 2009; Lussi et al. 2010), HIV infection and replication, DNA damage repair (Mackay et al. 2017; Duheron et al. 2017; Chow et al. 2012; Lemaitre et al. 2012) and gene expression regulation (Vaquerizas et al. 2010; Mendjan et al. 2006; Jacinto et al. 2015; Nanni et al. 2016). A role for Nup153 in gene expression regulation was originally identified in Drosophila, where Nup153 together with Megator, the homolog of human Tpr and another nuclear basket nucleoporin, were found to be required for the transcriptional regulation of dosage compensation (Mendjan et al. 2006). These proteins were later shown to bind a great portion of the genome (~25%) in continuous domains of 10–500 kilobases that present chromatin markers of active transcription (Vaquerizas et al. 2010). Consistent with a role in transcriptional regulation, downregulation of Nup153 was found to affect the expression a large number of genes (~5,700) in flies (Vaquerizas et al. 2010). In mouse ESCs, Nup153 was recently identified to bind to the transcription start site of several developmental genes (Jacinto et al. 2015). Interestingly, in these cells Nup153 was found to act as a repressor for differentiation genes (Fig. 7.4). The repression by Nup153 is required for the maintenance of the pluripotent state of ESCs, and depletion of this nuclear pore complex component results in early cell differentiation into different linages (Jacinto et al. 2015). The role of Nup153 in gene expression regulation is not restricted to ESCs. In mouse cardiomyocytes, Nup153 has also been found to associate with, and to regulate the activity of, genes involved in cardiac remodeling (Nanni et al. 2016). In this case, the binding of Nup153 correlates with markers of active chromatin but whether they occur inside the nucleus or at the nuclear periphery has not been investigated (Nanni et al. 2016). These findings further support the concept that nucleoporins can regulate different subsets of genes in distinct cell types.
The idea that nuclear pore complex components might play multiple functions depending on their intracellular localization is further reinforced by the findings that a soluble isoform of the transmembrane nucleoporin Pom121 (sPom121) regulates gene expression inside the nucleus (Franks et al. 2016) (Fig. 7.4). sPom121 is a consequence of a genomic rearrangement during mammalian evolution that generated an alternative transcription initiation in the POM121 loci. The product of this alternative start site is spliced so that it loses exon 4 encoding the transmembrane domain of Pom121. This results in a soluble isoform of Pom121 that does not associate with NPCs (Franks et al. 2016). The soluble sPom121 uses its nuclear localization signal to access the nuclear interior where it interacts with Nup98 at many gene promoters and cooperates to regulate multiple target genes (Fig. 7.4). Like Nup98 and Nup153, sPom121 mobility is affected by the transcriptional state of the cell, and the transcriptional inhibitor Actinomycin D strongly slows down the exchange of Pom121 inside the nucleus (Franks et al. 2016).
7.5.2 Mechanisms of Gene-Expression Regulation by Intranuclear Nucleoporins
To date, we have a very limited knowledge of the mechanisms through which intranuclear nucleoporins regulate gene expression in mammalian cells. In the case of Nup98, studies using the Nup98 fusion proteins that result from chromosomal translocations uncovered that through its FG-rich repeats this nucleoporin interacts with several transcriptional and chromatin modulators including CREB binding protein (CBP)/p300 (Kasper et al. 1999), histone deacetylase 1 (HDAC1) (Bai et al. 2006), and mixed lineage leukemia (MLL) (Shima et al. 2017) (Fig. 7.4). Many of these interactions have been shown to play a key role in the deregulation of HOX gene expression that is associated with the malignant transformation of hematopoietic progenitors expressing Nup98 fusion proteins (see Sect. 7.6). Interestingly, wild type Nup98 has also been found to interact with Trx/MLL and NSL in Drosophila, and to regulate HOX gene expression in this organism (Pascual-Garcia et al. 2014). Nup98 interactors suggest that this nucleoporin helps to recruit chromatin modifiers to specific loci, particularly developmental genes, influencing their expression. This may also hold true for Nup98-regulated INFγ target genes. Although the mechanisms through which this nucleoporin regulates transcriptional memory have not been identified, this process requires specific changes in chromatin modifications (D’Urso and Brickner 2017, 2014) that suggest that Nup98 might also work by modulating the activity of chromatin modifying complexes. But how Nup98, which does not contain DNA binding domains per se, recruits these transcriptional regulators to specific DNA sites is still unknown. A key player in this process might be the nuclear export factor Crm1. Crm1 is the major transport receptor for the export of proteins from the nucleus (Fung and Chook 2014). Recently, it was found that in leukemic cells Crm1 is prebound to HOX gene clusters and helps to recruit the Nup98-HoxA9 and CALM-AF10 aberrant fusion proteins to regulate HOX gene expression (Conway et al. 2015; Oka et al. 2016) (Fig. 7.4). These findings are very exciting because they identify that the coordinated activity of nuclear transport receptors and nuclear pore complex proteins is not just restricted to the regulation of nucleo-cytoplasmic transport, but is also critical for the assembly of transcriptional complexes that modulate the activity of the mammalian genome.
The regulation of gene expression by Nup98 is not restricted to its function on specific loci. Nup98 has also been shown to bind to the 3’ end of a distinct set of p53 target genes and to regulate mRNA stability (Singer et al. 2012). For example, Nup98 binding to the 3’UTR of p21 mRNA prevents its degradation by the exosome and increases its levels in cells. Because certain cancers, such as hepatocellular carcinoma (HCC) show reduced levels of Nup98, this nucleoporin has been suggested to act as a tumor suppressor required for the proper function of p53 in cells (Singer et al. 2012).
Recently, a proteomic screen identified the DExH/D-box helicase DHX9 as a binding partner for Nup98 (Capitanio et al. 2017). Helicases are enzymes that catalyze nucleic acid remodeling. The DHX9 helicase is able to unwind RNA as well as DNA, and has been shown to play critical roles in gene transcription and RNA processing (Lee and Pelletier 2016). Nup98 has been found to recruit DHX9 to specific foci within the nucleus and modifying Nup98 levels affects the intranuclear localization of the enzyme (Capitanio et al. 2017). Interestingly, Nup98 and DHX9 co-bind a subset of messenger RNAs and genomic loci; and the interaction of Nup98 with DHX9 has been shown to stimulate the transcriptional function of the enzyme (Capitanio et al. 2017). Because Nup98 binding to DHX9 increases its ATPase activity, it has been proposed that this nucleoporin acts as a cofactor for DHX9 during transcription. The identification of several additional helicases as interaction partners for Nup98 suggest that a Nup98-helicase complex may be responsible for regulating the activity of a subset of Nup98 target genes, either by modulating the activity of gene loci themselves or by regulating the processing of their transcripts (Capitanio et al. 2017).
In contrast to the transcriptional activator function of Nup98, Nup153 has been shown to negatively regulate the activity of differentiation/developmental genes, promoting in this manner the pluripotency of ESCs (Jacinto et al. 2015). The way Nup153 represses such genes is by directly interacting and recruiting the polycomb repressive complex 1 (PRC1) to their promoters (Jacinto et al. 2015) (Fig. 7.4). The Polycomb-group (PcG) protein complexes, which include PRC1 and PRC2 among others, are responsible for creating and maintaining a repressive chromatin environment that ensures the silencing of many developmental genes (Aloia et al. 2013; Margueron and Reinberg 2011). Polycomb proteins mediate gene silencing mainly by modulating chromatin structure through histone post-translational modifications. The PRC1 complex, for example, induces chromatin condensation by monoubiquitylation of histone H2A (Wang et al. 2004), which leads to the repression of PRC1 target genes. By bringing the PRC1 complex to differentiation-inducing genes, the chromatin-associated Nup153 induces epigenetic gene silencing that maintains stem cell pluripotency.
7.6 Gene Regulation by Abnormal Nucleoporin Fusion Proteins
Many cancers are characterized by chromosomal translocations that lead to gene fusion encoding chimeric proteins with aberrant functions (Zheng 2013). Because chromosomal rearrangements require the interaction of the two translocating chromosomes, it is considered that the nonrandom distribution of chromosomes inside the cell nucleus is a key determinant of this process (Nikiforova et al. 2000; Roix et al. 2003; Soutoglou and Misteli 2008; Zheng 2013). Chromosomal rearrangements are particularly represented in blood cancers, and several chimeric fusion proteins have been found to play critical roles in the transformation of hematopoietic progenitors and to significantly contribute to the development of blood malignancies. Notably, several nucleoporins were found to be part of oncogenic fusions in blood malignancies, particularly in acute myeloid leukemia (AML), chronic myelogenous leukemia (CML), T-cell acute lymphoblastic leukemia (T-ALL) and myelodysplastic syndrome (MDS). These include Nup98, Nup160, Nup214, Nup358 and Tpr (Simon and Rout 2014; Shimozono et al. 2015; Fahrenkrog 2014).
Most studies involving nucleoporin fusions have been centered on Nup98 fusions, and to a minor extent, on Nup214 fusions. Nup98 is the most frequently translocated nucleoporin in leukemia, and at least 30 different fusions partners for this nucleoporin have been identified so far (Fahrenkrog 2014; Saw et al. 2013). The oncogenic strength and the mechanisms of function of different Nup98 fusions depend on the fusion partner (Saw et al. 2013). Leukemias that have Nup98 chimeric fusions are generally highly aggressive and very resistant to therapies (Moore et al. 2007; Gough et al. 2011). In most cases, NUP98 participates in balanced chromosomal translocation that result in fusion proteins of the N-terminal domain of Nup98, which contains its GLFG repeats, and the C-terminal domain of a fusion partner (Gough et al. 2011; Moore et al. 2007). The GLFG repeats of Nup98 have been shown to be essential for the transformation of hematopoietic progenitors by Nup98 fusions (Kasper et al. 1999); and although wild type Nup98 localizes to NPCs and the nuclear interior, Nup98 fusions have been described as intranuclear proteins that do not associate with NPCs (Kasper et al. 1999; Xu and Powers 2010; Fahrenkrog et al. 2016). Of all Nup98 fusions, approximately one third are proteins that contain homeodomains (HD), which are helix-turn-helix DNA-binding domains (Fahrenkrog 2014; Moore et al. 2007; Gough et al. 2011). This class mostly include transcription factors that play key roles in blood development. The rest include non-HD proteins, the majority of which are involved in epigenetic regulation and chromatin remodeling. The ectopic expression of several Nup98 fusion proteins has been shown to promote the proliferation and prevent the differentiation of hematopoietic progenitors (Chung et al. 2006; Calvo et al. 2002; Choi et al. 2009; Yassin et al. 2009; Takeda et al. 2006).
Studies of Nup98 fusion proteins in leukemia have uncovered alterations in the NE structure (Fahrenkrog et al. 2016), nucleocytoplasmic transport (Funasaka et al. 2011; Saito et al. 2017; Takeda et al. 2010), cell signaling (Qiu et al. 2015), and mitosis (Salsi et al. 2014, 2016). But probably the most common abnormalities observed with the ectopic expression of Nup98 chimeric proteins are the deregulation of gene expression, particularly of HOX genes, and epigenetic alterations (Kasper et al. 1999; Ghannam et al. 2004; Bai et al. 2006; Calvo et al. 2002; Saw et al. 2013; Oka et al. 2016; Rio-Machin et al. 2017). This suggest, that key mechanisms though which these aberrant proteins affect normal cell physiology are by modulating the expression of key differentiation/developmental genes and through changes in chromatin organization. How Nup98 fusions perform these functions is an area of active research. Recently, several Nup98 fusion proteins were shown to interact with the histone modifying complexes mixed lineage leukemia 1 (MLL1) and the non-specific lethal (NSL) (Xu et al. 2016), being these interactions required for leukemogenesis (Xu et al. 2016; Shima et al. 2017). These include NUP98-HOXA9, NUP98-HOXD13, NUP98-NSD1, NUP98-PHF23, and NUP98-TOP1 (Xu et al. 2016). As mentioned, wild type Nup98 in drosophila also interacts with Trx/MLL and NSL to regulate HOX gene expression (Pascual-Garcia et al. 2014). This indicates that the regulation of histone modifications is a normal function of Nup98 that gets hijacked by its fusion partners. Nup98 fusion proteins have also been shown to associate with other chromatin modifiers including the histone deacetylase HDAC1 and CBP/p300 (Kasper et al. 1999; Bai et al. 2006; Rio-Machin et al. 2017). For many interactions between Nup98 and chromatin modulators, the FG repeats of Nup98 have been shown to be essential for protein association and for chromatin modifications (Bai et al. 2006; Kasper et al. 1999). This also supports the idea that wild type Nup98 has a role in epigenetic modulation. Some of these endogenous Nup98 functions might be further potentiated by its fusion with other chromatin modifiers such as the histone methyl transferases NSD1, NSD3 or MLL. Conversely, the endogenous functions of transcriptional/chromatin modulators might be enhanced by fusion with Nup98. For example, in some cases of AML, Nup98 is fused to the plant homeodomain (PHD) domains of JARID1A and NSD1. PHD fingers, which are present in many chromatin-remodeling proteins, bind to specific histone/epigenetic marks and help to recruit transcription factors and chromatin modulators (Musselman and Kutateladze 2009). At the HOXA locus, PHD domains have been found to prevent the spreading of polycomb repressive complexes which promote gene silencing. Consistent with a chromatin boundary activity, NUP98–PHD fusions were found to prevent polycomb-mediated gene silencing and to help maintain chromatin in an active state, stimulating in this manner HOX gene expression (Wang et al. 2009).
As mentioned before, several other nucleoporins participate in cancer associated chromosomal translocations. For most of them, the mechanisms through which this nucleoporin chimeric proteins deregulate cellular physiology is still unknown. So far, the only other nucleoporin fusion that has been shown to also affect gene expression directly is the SET-NUP214 chimera. Nup214 (also known as CAN) is an FG repeat-containing nucleoporin component of the cytoplasmic filaments (Kraemer et al. 1994; Napetschnig et al. 2009). This large nucleoporin plays important roles in nuclear import and export. Similar to Nup98, the fusions of Nup214 maintain its FG domains and localize to intranuclear foci instead of NPCs (Fornerod et al. 1995; Saito et al. 2004; Simon and Rout 2014). Also like Nup98, Set-Nup214 chimeras bind to the promoter of HOXA genes and deregulate their expression, promoting cell proliferation and inhibiting cell differentiation. At HOXA gene promoters Set-Nup214 interacts with the transport receptor CRM1 and the histone methyltransferase DOTL1 (Van Vlierberghe et al. 2008). This suggests that its mechanisms of recruitment and gene expression regulation might be conserved with the Nup98-HoxA9 fusion (see Sect. 7.5.2) (Conway et al. 2015; Oka et al. 2016).
Another fusion of Nup214, Dek-Nup214, has been shown to affect gene expression but at the translation, instead of transcriptional, level (Ageberg et al. 2008). By stimulating hyperphosphorylation of the oncogene Elf4E, Dek-Nup214 increases overall protein synthesis (Ageberg et al. 2008). This is interesting, because increased activity of Elf4E has been found to modify the configuration of NPCs to stimulate the export of oncogenic RNAs and promote cell proliferation (Culjkovic-Kraljacic et al. 2012). It is worth mentioning, that even though Set- and Dek-Nup214 fusions do not localize to NPCs, they still affect nucleocytoplasmic transport (Saito et al. 2016; Port et al. 2016). The way these fusion proteins affect nuclear transport is through the sequestration of transport receptors and nucleoporins into their highly dynamic intranuclear foci (Saito et al. 2016; Port et al. 2016). Naturally, by affecting nuclear transport, and also by tethering transcription factors to these foci, Nup214 fusions have an indirect impact on gene expression.
7.7 Conclusions
Since their discovery, NPCs have continuously amazed scientist for their unique features. First was their exceptional structure, then was their essential role in controlling the exchange of all molecules between the nucleus and the cytoplasm, and now their emerging contributions to genome integrity and gene expression regulation. Though almost 70 years have passed from their first observation, there are still many mysteries that need to be elucidated. What is the organization of the central channel? Which is/are the definite mechanism/s of nucleo cytoplasmic transport? How many partners work together with NPCs to regulate genome function? are some of many questions that still remain to be answered.
But what has become clear with a large amount of work from many different labs is that NPCs are not just mere channels that sit at the NE passively allowing the exchange of molecules between the nucleus and the cytoplasm. NPCs are highly dynamic and plastic structures that can be modified to change their properties, that can have distinct composition in different cell types, that play multiple transport-independent functions, and that they are central regulators of cellular physiology. The recent findings that NPC components also have off-pore functions not only extends the potential processes modulated by nucleoporins but also provides a novel perspective on how these structures might contribute to regulate essential cellular functions.
The role of NPCs as critical regulators of genome organization and gene expression was originally described in yeast and flies. Although understanding the function of mammalian NPCs in these processes lagged behind for some time, it has now started to catch up with the emerging roles of mammalian NPCs in all aspects of genome integrity, including genome organization, transcriptional regulation, DNA repair, DNA replication, chromosome segregation, and others (see (Bukata et al. 2013; Raices and D’Angelo 2012) for additional information). The central role that NPCs and nucleoporins play in many of these processes explains the increasing number of alterations in these structures that are being linked to diseases such as neurodegeneration and cancer. Understanding the modes of action of NPCs and nucleoporins in regulating and coordinating genome functions will help us elucidate how these structures contribute to the faithful translation and transmission of the genetic information, and will result in a better understanding of how alterations in their function contribute to disease development.
References
Adkins NL, Hagerman TA, Georgel P (2006) GAGA protein: a multi-faceted transcription factor. Biochem Cell Biol 84(4):559–567. https://doi.org/10.1139/o06-062
Ageberg M, Drott K, Olofsson T et al (2008) Identification of a novel and myeloid specific role of the leukemia-associated fusion protein DEK-NUP214 leading to increased protein synthesis. Genes Chromosomes Cancer 47(4):276–287. https://doi.org/10.1002/gcc.20531
Al-Haboubi T, Shumaker DK, Koser J et al (2011) Distinct association of the nuclear pore protein Nup153 with A- and B-type lamins. Nucleus 2(5):500–509. https://doi.org/10.4161/nucl.2.5.17913
Aloia L, Di Stefano B, Di Croce L (2013) Polycomb complexes in stem cells and embryonic development. Development 140(12):2525–2534. https://doi.org/10.1242/dev.091553
Bai XT, Gu BW, Yin T et al (2006) Trans-repressive effect of NUP98-PMX1 on PMX1-regulated c-FOS gene through recruitment of histone deacetylase 1 by FG repeats. Cancer Res 66(9):4584–4590. https://doi.org/10.1158/0008-5472.CAN-05-3101
Ball JR, Dimaano C, Bilak A et al (2007) Sequence preference in RNA recognition by the nucleoporin Nup153. J Biol Chem 282(12):8734–8740. https://doi.org/10.1074/jbc.M608477200
Bastos R, Lin A, Enarson M et al (1996) Targeting and function in mRNA export of nuclear pore complex protein Nup153. J Cell Biol 134(5):1141–1156
Beck M, Forster F, Ecke M et al (2004) Nuclear pore complex structure and dynamics revealed by cryoelectron tomography. Science 306(5700):1387–1390
Beck M, Lucic V, Forster F et al (2007) Snapshots of nuclear pore complexes in action captured by cryo-electron tomography. Nature 449(7162):611–615
Brickner DG, Ahmed S, Meldi L et al (2012) Transcription factor binding to a DNA zip code controls interchromosomal clustering at the nuclear periphery. Dev Cell 22(6):1234–1246. https://doi.org/10.1016/j.devcel.2012.03.012
Brown CR, Kennedy CJ, Delmar VA et al (2008) Global histone acetylation induces functional genomic reorganization at mammalian nuclear pore complexes. Genes Dev 22(5):627–639
Bukata L, Parker SL, D’Angelo MA (2013) Nuclear pore complexes in the maintenance of genome integrity. Curr Opin Cell Biol 25(3):378–386. https://doi.org/10.1016/j.ceb.2013.03.002
Callan HG, Tomlin SG (1950) Experimental studies on amphibian oocyte nuclei. I. Investigation of the structure of the nuclear membrane by means of the electron microscope. Proc R Soc Lond B Biol Sci 137(888):367–378
Calvo KR, Sykes DB, Pasillas MP et al (2002) Nup98-HoxA9 immortalizes myeloid progenitors, enforces expression of Hoxa9, Hoxa7 and Meis1, and alters cytokine-specific responses in a manner similar to that induced by retroviral co-expression of Hoxa9 and Meis1. Oncogene 21(27):4247–4256. https://doi.org/10.1038/sj.onc.1205516
Capelson M, Hetzer MW (2009) The role of nuclear pores in gene regulation, development and disease. EMBO Rep 10(7):697–705. https://doi.org/10.1038/embor.2009.147
Capelson M, Liang Y, Schulte R et al (2010) Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. Cell 140(3):372–383. https://doi.org/10.1016/j.cell.2009.12.054
Capitanio JS, Montpetit B, Wozniak RW (2017) Human Nup98 regulates the localization and activity of DExH/D-box helicase DHX9. Elife 6. https://doi.org/10.7554/eLife.18825
Choi CW, Chung YJ, Slape C et al (2009) A NUP98-HOXD13 fusion gene impairs differentiation of B and T lymphocytes and leads to expansion of thymocytes with partial TCRB gene rearrangement. J Immunol 183(10):6227–6235. https://doi.org/10.4049/jimmunol.0901121
Chow KH, Elgort S, Dasso M et al (2012) Two distinct sites in Nup153 mediate interaction with the SUMO proteases SENP1 and SENP2. Nucleus 3(4):349–358. https://doi.org/10.4161/nucl.20822
Chung KY, Morrone G, Schuringa JJ et al (2006) Enforced expression of NUP98-HOXA9 in human CD34(+) cells enhances stem cell proliferation. Cancer Res 66(24):11781–11791. https://doi.org/10.1158/0008-5472.CAN-06-0706
Clocchiatti A, Di Giorgio E, Demarchi F et al (2013) Beside the MEF2 axis: unconventional functions of HDAC4. Cell Signal 25(1):269–276. https://doi.org/10.1016/j.cellsig.2012.10.002
Conway AE, Haldeman JM, Wechsler DS et al (2015) A critical role for CRM1 in regulating HOXA gene transcription in CALM-AF10 leukemias. Leukemia 29(2):423–432. https://doi.org/10.1038/leu.2014.221
Cronshaw JM, Krutchinsky AN, Zhang W et al (2002) Proteomic analysis of the mammalian nuclear pore complex. J Cell Biol 158(5):915–927
Culjkovic-Kraljacic B, Baguet A, Volpon L et al (2012) The oncogene eIF4E reprograms the nuclear pore complex to promote mRNA export and oncogenic transformation. Cell Rep 2(2):207–215. https://doi.org/10.1016/j.celrep.2012.07.007
D’Angelo MA, Gomez-Cavazos JS, Mei A et al (2012) A change in nuclear pore complex composition regulates cell differentiation. Dev Cell 22(2):446–458. https://doi.org/10.1016/j.devcel.2011.11.021
D’Angelo MA, Hetzer MW (2008) Structure, dynamics and function of nuclear pore complexes. Trends Cell Biol 18(10):456–466. https://doi.org/10.1016/j.tcb.2008.07.009
D’Angelo MA, Raices M, Panowski SH et al (2009) Age-dependent deterioration of nuclear pore complexes causes a loss of nuclear integrity in postmitotic cells. Cell 136(2):284–295. https://doi.org/10.1016/j.cell.2008.11.037
D’Urso A, Brickner JH (2014) Mechanisms of epigenetic memory. Trends Genet 30(6):230–236. https://doi.org/10.1016/j.tig.2014.04.004
D’Urso A, Brickner JH (2017) Epigenetic transcriptional memory. Curr Genet 63(3):435–439. https://doi.org/10.1007/s00294-016-0661-8
Daigle N, Beaudouin J, Hartnell L et al (2001) Nuclear pore complexes form immobile networks and have a very low turnover in live mammalian cells. J Cell Biol 154(1):71–84
Dimaano C, Ball JR, Prunuske AJ et al (2001) RNA association defines a functionally conserved domain in the nuclear pore protein Nup153. J Biol Chem 276(48):45349–45357. https://doi.org/10.1074/jbc.M102592200
Duheron V, Nilles N, Pecenko S et al (2017) Localisation of Nup153 and SENP1 to nuclear pore complexes is required for 53BP1-mediated DNA double-strand break repair. J Cell Sci 130(14):2306–2316. https://doi.org/10.1242/jcs.198390
Fahrenkrog B (2014) Nucleoporin Gene Fusions and Hematopoietic Malignancies, New Journal of Science, vol. 2014, Article ID 468306, 18 pages. https://doi.org/10.1155/2014/468306
Fahrenkrog B, Martinelli V, Nilles N et al (2016) Expression of leukemia-associated Nup98 fusion proteins generates an aberrant nuclear envelope phenotype. PLoS One 11(3):e0152321. https://doi.org/10.1371/journal.pone.0152321
Fontoura BM, Blobel G, Matunis MJ (1999) A conserved biogenesis pathway for nucleoporins: proteolytic processing of a 186-kilodalton precursor generates Nup98 and the novel nucleoporin, Nup96. J Cell Biol 144(6):1097–1112
Fornerod M, Boer J, van Baal S et al (1995) Relocation of the carboxyterminal part of CAN from the nuclear envelope to the nucleus as a result of leukemia-specific chromosome rearrangements. Oncogene 10(9):1739–1748
Franks TM, Benner C, Narvaiza I et al (2016) Evolution of a transcriptional regulator from a transmembrane nucleoporin. Genes Dev 30(10):1155–1171. https://doi.org/10.1101/gad.280941.116
Franks TM, Hetzer MW (2013) The role of Nup98 in transcription regulation in healthy and diseased cells. Trends Cell Biol 23(3):112–117. https://doi.org/10.1016/j.tcb.2012.10.013
Frenkiel-Krispin D, Maco B, Aebi U et al (2010) Structural analysis of a metazoan nuclear pore complex reveals a fused concentric ring architecture. J Mol Bio 395(3):578–586. https://doi.org/10.1016/j.jmb.2009.11.010
Funasaka T, Nakano H, Wu Y et al (2011) RNA export factor RAE1 contributes to NUP98-HOXA9-mediated leukemogenesis. Cell Cycle 10(9):1456–1467
Fung HY, Chook YM (2014) Atomic basis of CRM1-cargo recognition, release and inhibition. Semin Cancer Biol 27:52–61. https://doi.org/10.1016/j.semcancer.2014.03.002
Galy V, Mattaj IW, Askjaer P (2003) Caenorhabditis elegans nucleoporins Nup93 and Nup205 determine the limit of nuclear pore complex size exclusion in vivo. Mol Biol Cell 14(12):5104–5115
Gerace L, Ottaviano Y, Kondor-Koch C (1982) Identification of a major polypeptide of the nuclear pore complex. J Cell Biol 95(3):826–837
Ghannam G, Takeda A, Camarata T et al (2004) The oncogene Nup98-HOXA9 induces gene transcription in myeloid cells. J Biol Chem 279(2):866–875. https://doi.org/10.1074/jbc.M307280200
Gough SM, Slape CI, Aplan PD (2011) NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood 118(24):6247–6257. https://doi.org/10.1182/blood-2011-07-328880
Granok H, Leibovitch BA, Shaffer CD et al (1995) Chromatin. Ga-ga over GAGA factor. Curr Biol 5(3):238–241
Griffis ER, Altan N, Lippincott-Schwartz J et al (2002) Nup98 is a mobile nucleoporin with transcription-dependent dynamics. Mol Biol Cell 13(4):1282–1297
Griffis ER, Craige B, Dimaano C et al (2004) Distinct functional domains within nucleoporins Nup153 and Nup98 mediate transcription-dependent mobility. Mol Biol Cell 15(4):1991–2002
Griffis ER, Xu S, Powers MA (2003) Nup98 localizes to both nuclear and cytoplasmic sides of the nuclear pore and binds to two distinct nucleoporin subcomplexes. Mol Biol Cell 14(2):600–610
Hase ME, Cordes VC (2003) Direct interaction with nup153 mediates binding of Tpr to the periphery of the nuclear pore complex. Mol Biol Cell 14(5):1923–1940
Hoelz A, Glavy JS, Beck M (2016) Toward the atomic structure of the nuclear pore complex: when top down meets bottom up. Nat Struct Mol Biol 23(7):624–630. https://doi.org/10.1038/nsmb.3244
Ibarra A, Benner C, Tyagi S et al (2016) Nucleoporin-mediated regulation of cell identity genes. Genes Dev 30(20):2253–2258. https://doi.org/10.1101/gad.287417.116
Jacinto FV, Benner C, Hetzer MW (2015) The nucleoporin Nup153 regulates embryonic stem cell pluripotency through gene silencing. Genes Dev 29(12):1224–1238. https://doi.org/10.1101/gad.260919.115
Kalverda B, Pickersgill H, Shloma VV et al (2010) Nucleoporins directly stimulate expression of developmental and cell-cycle genes inside the nucleoplasm. Cell 140(3):360–371. https://doi.org/10.1016/j.cell.2010.01.011
Kasper LH, Brindle PK, Schnabel CA et al (1999) CREB binding protein interacts with nucleoporin-specific FG repeats that activate transcription and mediate NUP98-HOXA9 oncogenicity. Mol Cell Biol 19(1):764–776
Kehat I, Accornero F, Aronow BJ et al (2011) Modulation of chromatin position and gene expression by HDAC4 interaction with nucleoporins. J Cell Biol 193(1):21–29. https://doi.org/10.1083/jcb.201101046
Kraemer D, Wozniak RW, Blobel G et al (1994) The human CAN protein, a putative oncogene product associated with myeloid leukemogenesis, is a nuclear pore complex protein that faces the cytoplasm. Proc Natl Acad Sci U S A 91(4):1519–1523
Krull S, Dorries J, Boysen B et al (2010) Protein Tpr is required for establishing nuclear pore-associated zones of heterochromatin exclusion. EMBO J 29(10):1659–1673. https://doi.org/10.1038/emboj.2010.54
Kutay U, Hetzer MW (2008) Reorganization of the nuclear envelope during open mitosis. Curr Opin Cell Biol 20(6):669–677. https://doi.org/10.1016/j.ceb.2008.09.010
Labade AS, Karmodiya K, Sengupta K (2016) HOXA repression is mediated by nucleoporin Nup93 assisted by its interactors Nup188 and Nup205. Epigenetics Chromatin 9:54. https://doi.org/10.1186/s13072-016-0106-0
Lee T, Pelletier J (2016) The biology of DHX9 and its potential as a therapeutic target. Oncotarget 7(27):42716–42739. https://doi.org/10.18632/oncotarget.8446
Lemaitre C, Bickmore WA (2015) Chromatin at the nuclear periphery and the regulation of genome functions. Histochem Cell Biol 144(2):111–122. https://doi.org/10.1007/s00418-015-1346-y
Lemaitre C, Fischer B, Kalousi A et al (2012) The nucleoporin 153, a novel factor in double-strand break repair and DNA damage response. Oncogene 31(45):4803–4809. https://doi.org/10.1038/onc.2011.638
Liang Y, Franks TM, Marchetto MC et al (2013) Dynamic association of NUP98 with the human genome. PLoS Genet 9(2):e1003308. https://doi.org/10.1371/journal.pgen.1003308
Light WH, Freaney J, Sood V et al (2013) A conserved role for human Nup98 in altering chromatin structure and promoting epigenetic transcriptional memory. PLoS Biol 11(3):e1001524. https://doi.org/10.1371/journal.pbio.1001524
Lussi YC, Shumaker DK, Shimi T et al (2010) The nucleoporin Nup153 affects spindle checkpoint activity due to an association with Mad1. Nucleus 1(1):71–84. https://doi.org/10.4161/nucl.1.1.10244
Mackay DR, Elgort SW, Ullman KS (2009) The nucleoporin Nup153 has separable roles in both early mitotic progression and the resolution of mitosis. Mol Biol Cell 20(6):1652–1660. https://doi.org/10.1091/mbc.E08-08-0883
Mackay DR, Howa AC, Werner TL et al (2017) Nup153 and Nup50 promote recruitment of 53BP1 to DNA repair foci by antagonizing BRCA1-dependent events. J Cell Sci. https://doi.org/10.1242/jcs.203513
Maimon T, Elad N, Dahan I et al (2012) The human nuclear pore complex as revealed by cryo-electron tomography. Structure 20(6):998–1006. https://doi.org/10.1016/j.str.2012.03.025
Makise M, Mackay DR, Elgort S et al (2012) The Nup153-Nup50 protein interface and its role in nuclear import. J Biol Chem 287(46):38515–38522. https://doi.org/10.1074/jbc.M112.378893
Margueron R, Reinberg D (2011) The Polycomb complex PRC2 and its mark in life. Nature 469(7330):343–349. https://doi.org/10.1038/nature09784
McKinsey TA, Zhang CL, Lu J et al (2000) Signal-dependent nuclear export of a histone deacetylase regulates muscle differentiation. Nature 408(6808):106–111. https://doi.org/10.1038/35040593
Mendjan S, Taipale M, Kind J et al (2006) Nuclear pore components are involved in the transcriptional regulation of dosage compensation in Drosophila. Mol Cell 21(6):811–823. https://doi.org/10.1016/j.molcel.2006.02.007
Misteli T (2007) Beyond the sequence: cellular organization of genome function. Cell 128(4):787–800
Moore MA, Chung KY, Plasilova M et al (2007) NUP98 dysregulation in myeloid leukemogenesis. Ann N Y Acad Sci 1106:114–142. https://doi.org/10.1196/annals.1392.019
Moroianu J, Hijikata M, Blobel G et al (1995) Mammalian karyopherin alpha 1 beta and alpha 2 beta heterodimers: alpha 1 or alpha 2 subunit binds nuclear localization signal and beta subunit interacts with peptide repeat-containing nucleoporins. Proc Natl Acad Sci U S A 92(14):6532–6536
Musselman CA, Kutateladze TG (2009) PHD fingers: epigenetic effectors and potential drug targets. Mol Interv 9(6):314–323. https://doi.org/10.1124/mi.9.6.7
Nakielny S, Shaikh S, Burke B et al (1999) Nup153 is an M9-containing mobile nucleoporin with a novel Ran-binding domain. EMBO J 18(7):1982–1995. https://doi.org/10.1093/emboj/18.7.1982
Nanni S, Re A, Ripoli C et al (2016) The nuclear pore protein Nup153 associates with chromatin and regulates cardiac gene expression in dystrophic mdx hearts. Cardiovasc Res 112(2):555–567. https://doi.org/10.1093/cvr/cvw204
Napetschnig J, Kassube SA, Debler EW et al (2009) Structural and functional analysis of the interaction between the nucleoporin Nup214 and the DEAD-box helicase Ddx19. Proc Natl Acad Sci U S A 106(9):3089–3094. https://doi.org/10.1073/pnas.0813267106
Nguyen HQ, Bosco G (2015) Gene positioning effects on expression in eukaryotes. Annu Rev Genet 49:627–646. https://doi.org/10.1146/annurev-genet-112414-055008
Niederriter AR, Varshney A, Parker SC et al (2015) Super enhancers in cancers, complex disease, and developmental disorders. Genes (Basel) 6(4):1183–1200. https://doi.org/10.3390/genes6041183
Nikiforova MN, Stringer JR, Blough R et al (2000) Proximity of chromosomal loci that participate in radiation-induced rearrangements in human cells. Science 290(5489):138–141
Ogawa Y, Miyamoto Y, Oka M et al (2012) The interaction between importin-alpha and Nup153 promotes importin-alpha/beta-mediated nuclear import. Traffic 13(7):934–946. https://doi.org/10.1111/j.1600-0854.2012.01367.x
Oka M, Mura S, Yamada K et al (2016) Chromatin-prebound Crm1 recruits Nup98-HoxA9 fusion to induce aberrant expression of Hox cluster genes. Elife 5:e09540. https://doi.org/10.7554/eLife.09540
Oliver F (ed) (1913) Makers of British Botany: A Collection of Biographies by Living Botanists (Cambridge Library Collection - Botany and Horticulture). Cambridge: Cambridge University Press. https://doi.org/10.1017/CBO9780511710902
Olsson M, Ekblom M, Fecker L et al (1999) cDNA cloning and embryonic expression of mouse nuclear pore membrane glycoprotein 210 mRNA. Kidney Int 56(3):827–838. https://doi.org/10.1046/j.1523-1755.1999.00618.x
Olsson M, Scheele S, Ekblom P (2004) Limited expression of nuclear pore membrane glycoprotein 210 in cell lines and tissues suggests cell-type specific nuclear pores in metazoans. Exp Cell Res 292(2):359–370
Pante N, Bastos R, McMorrow I et al (1994) Interactions and three-dimensional localization of a group of nuclear pore complex proteins. J Cell Biol 126(3):603–617
Pascual-Garcia P, Debo B, Aleman JR et al (2017) Metazoan nuclear pores provide a scaffold for poised genes and mediate induced enhancer-promoter contacts. Mol Cell 66(1):63–76. https://doi.org/10.1016/j.molcel.2017.02.020. e66
Pascual-Garcia P, Jeong J, Capelson M (2014) Nucleoporin Nup98 associates with Trx/MLL and NSL histone-modifying complexes and regulates Hox gene expression. Cell Rep 9(2):433–442. https://doi.org/10.1016/j.celrep.2014.09.002
Port SA, Mendes A, Valkova C et al (2016) The oncogenic fusion proteins SET-Nup214 and sequestosome-1 (SQSTM1)-Nup214 form dynamic nuclear bodies and differentially affect nuclear protein and poly(A)+ RNA export. J Biol Chem 291(44):23068–23083. https://doi.org/10.1074/jbc.M116.735340
Qiu JJ, Zeisig BB, Li S et al (2015) Critical role of retinoid/rexinoid signaling in mediating transformation and therapeutic response of NUP98-RARG leukemia. Leukemia 29(5):1153–1162. https://doi.org/10.1038/leu.2014.334
Rabut G, Doye V, Ellenberg J (2004) Mapping the dynamic organization of the nuclear pore complex inside single living cells. Nat Cell Biol 6(11):1114–1121
Raices M, Bukata L, Sakuma S et al (2017) Nuclear pores regulate muscle development and maintenance by assembling a localized Mef2C complex. Dev Cell 41(5):540–554. https://doi.org/10.1016/j.devcel.2017.05.007. e547
Raices M, D’Angelo MA (2012) Nuclear pore complex composition: a new regulator of tissue-specific and developmental functions. Nat Rev Mol Cell Biol 13(11):687–699. https://doi.org/10.1038/nrm3461
Randise-Hinchliff C, Coukos R, Sood V et al (2016) Strategies to regulate transcription factor-mediated gene positioning and interchromosomal clustering at the nuclear periphery. J Cell Biol 212(6):633–646. https://doi.org/10.1083/jcb.201508068
Ratner GA, Hodel AE, Powers MA (2007) Molecular determinants of binding between Gly-Leu-Phe-Gly nucleoporins and the nuclear pore complex. J Biol Chem 282(47):33968–33976. https://doi.org/10.1074/jbc.M707911200
Reichelt R, Holzenburg A, Buhle Jr. EL et al (1990) Correlation between structure and mass distribution of the nuclear pore complex and of distinct pore complex components. J Cell Biol 110(4):883–894
Rio-Machin A, Gomez-Lopez G, Munoz J et al (2017) The molecular pathogenesis of the NUP98-HOXA9 fusion protein in acute myeloid leukemia. Leukemia. https://doi.org/10.1038/leu.2017.194
Roix JJ, McQueen PG, Munson PJ et al (2003) Spatial proximity of translocation-prone gene loci in human lymphomas. Nat Genet 34(3):287–291. https://doi.org/10.1038/ng1177
Rout MP, Aitchison JD, Suprapto A et al (2000) The yeast nuclear pore complex: composition, architecture, and transport mechanism. J Cell Biol 148(4):635–651
Saito S, Cigdem S, Okuwaki M et al (2016) Leukemia-associated Nup214 fusion proteins disturb the XPO1-mediated nuclear-cytoplasmic transport pathway and thereby the NF-kappaB signaling pathway. Mol Cell Biol 36(13):1820–1835. https://doi.org/10.1128/MCB.00158-16
Saito S, Miyaji-Yamaguchi M, Nagata K (2004) Aberrant intracellular localization of SET-CAN fusion protein, associated with a leukemia, disorganizes nuclear export. Int J Cancer 111(4):501–507. https://doi.org/10.1002/ijc.20296
Saito S, Yokokawa T, Iizuka G et al (2017) Function of Nup98 subtypes and their fusion proteins, Nup98-TopIIbeta and Nup98-SETBP1 in nuclear-cytoplasmic transport. Biochem Biophys Res Commun 487(1):96–102. https://doi.org/10.1016/j.bbrc.2017.04.024
Salsi V, Fantini S, Zappavigna V (2016) NUP98 fusion oncoproteins interact with the APC/C(Cdc20) as a pseudosubstrate and prevent mitotic checkpoint complex binding. Cell Cycle 15(17):2275–2287. https://doi.org/10.1080/15384101.2016.1172156
Salsi V, Ferrari S, Gorello P et al (2014) NUP98 fusion oncoproteins promote aneuploidy by attenuating the mitotic spindle checkpoint. Cancer Res 74(4):1079–1090. https://doi.org/10.1158/0008-5472.CAN-13-0912
Savas JN, Toyama BH, Xu T et al (2012) Extremely long-lived nuclear pore proteins in the rat brain. Science 335(6071):942. https://doi.org/10.1126/science.1217421
Saw J, Curtis DJ, Hussey DJ et al (2013) The fusion partner specifies the oncogenic potential of NUP98 fusion proteins. Leuk Res 37(12):1668–1673. https://doi.org/10.1016/j.leukres.2013.09.013
Schneider M, Hellerschmied D, Schubert T et al (2015) The nuclear pore-associated TREX-2 complex employs mediator to regulate gene expression. Cell 162(5):1016–1028. https://doi.org/10.1016/j.cell.2015.07.059
Shah S, Forbes DJ (1998) Separate nuclear import pathways converge on the nucleoporin Nup153 and can be dissected with dominant-negative inhibitors. Curr Biol 8(25):1376–1386
Shah S, Tugendreich S, Forbes D (1998) Major binding sites for the nuclear import receptor are the internal nucleoporin Nup153 and the adjacent nuclear filament protein Tpr. J Cell Biol 141(1):31–49
Shima Y, Yumoto M, Katsumoto T, Kitabayashi I (2017) MLL is essential for NUP98-HOXA9-induced leukemia. Leukemia 31:2200–2210. https://doi.org/10.1038/leu.2017.62
Shimozono N, Jinnin M, Masuzawa M et al (2015) NUP160-SLC43A3 is a novel recurrent fusion oncogene in angiosarcoma. Cancer Res 75(21):4458–4465. https://doi.org/10.1158/0008-5472.CAN-15-0418
Simon DN, Rout MP (2014) Cancer and the nuclear pore complex. Adv Exp Med Biol 773:285–307. https://doi.org/10.1007/978-1-4899-8032-8_13
Simon DN, Wilson KL (2011) The nucleoskeleton as a genome-associated dynamic ‘network of networks’. Nat Rev Mol Cell Biol 12(11):695–708. https://doi.org/10.1038/nrm3207
Singer S, Zhao R, Barsotti AM et al (2012) Nuclear pore component Nup98 is a potential tumor suppressor and regulates posttranscriptional expression of select p53 target genes. Mol Cell 48(5):799–810. https://doi.org/10.1016/j.molcel.2012.09.020
Smythe C, Jenkins HE, Hutchison CJ (2000) Incorporation of the nuclear pore basket protein nup153 into nuclear pore structures is dependent upon lamina assembly: evidence from cell-free extracts of Xenopus eggs. EMBO J 19(15):3918–3931. https://doi.org/10.1093/emboj/19.15.3918
Sood V, Brickner JH (2014) Nuclear pore interactions with the genome. Curr Opin Genet Dev 25:43–49. https://doi.org/10.1016/j.gde.2013.11.018
Soutoglou E, Misteli T (2008) On the contribution of spatial genome organization to cancerous chromosome translocations. J Natl Cancer Inst Monogr 39:16–19. https://doi.org/10.1093/jncimonographs/lgn017
Srivastava S, Dhawan J, Mishra RK (2015) Epigenetic mechanisms and boundaries in the regulation of mammalian Hox clusters. Mech Dev 138(Pt 2):160–169. https://doi.org/10.1016/j.mod.2015.07.015
Stancheva I, Schirmer EC (2014) Nuclear envelope: connecting structural genome organization to regulation of gene expression. Adv Exp Med Biol 773:209–244. https://doi.org/10.1007/978-1-4899-8032-8_10
Sukegawa J, Blobel G (1993) A nuclear pore complex protein that contains zinc finger motifs, binds DNA, and faces the nucleoplasm. Cell 72(1):29–38
Suntharalingam M, Wente SR (2003) Peering through the pore: nuclear pore complex structure, assembly, and function. Dev Cell 4(6):775–789
Takeda A, Goolsby C, Yaseen NR (2006) NUP98-HOXA9 induces long-term proliferation and blocks differentiation of primary human CD34+ hematopoietic cells. Cancer Res 66(13):6628–6637. https://doi.org/10.1158/0008-5472.CAN-06-0458
Takeda A, Sarma NJ, Abdul-Nabi AM et al (2010) Inhibition of CRM1-mediated nuclear export of transcription factors by leukemogenic NUP98 fusion proteins. J Biol Chem 285(21):16248–16257. https://doi.org/10.1074/jbc.M109.048785
Talamas JA, Capelson M (2015) Nuclear envelope and genome interactions in cell fate. Front Genet 6:95. https://doi.org/10.3389/fgene.2015.00095
Towbin BD, Meister P, Gasser SM (2009) The nuclear envelope--a scaffold for silencing? Curr Opin Genet Dev 19(2):180–186. https://doi.org/10.1016/j.gde.2009.01.006
Toyama BH, Savas JN, Park SK et al (2013) Identification of long-lived proteins reveals exceptional stability of essential cellular structures. Cell 154(5):971–982. https://doi.org/10.1016/j.cell.2013.07.037
Ullman KS, Shah S, Powers MA et al (1999) The nucleoporin nup153 plays a critical role in multiple types of nuclear export. Mol Biol Cell 10(3):649–664
Van Vlierberghe P, van Grotel M, Tchinda J et al (2008) The recurrent SET-NUP214 fusion as a new HOXA activation mechanism in pediatric T-cell acute lymphoblastic leukemia. Blood 111(9):4668–4680. https://doi.org/10.1182/blood-2007-09-111872
Vaquerizas JM, Suyama R, Kind J et al (2010) Nuclear pore proteins nup153 and megator define transcriptionally active regions in the Drosophila genome. PLoS Genet 6(2):e1000846. https://doi.org/10.1371/journal.pgen.1000846
Vollmer B, Lorenz M, Moreno-Andres D et al (2015) Nup153 recruits the Nup107-160 complex to the inner nuclear membrane for interphasic nuclear pore complex assembly. Dev Cell 33(6):717–728. https://doi.org/10.1016/j.devcel.2015.04.027
Walther TC, Fornerod M, Pickersgill H et al (2001) The nucleoporin Nup153 is required for nuclear pore basket formation, nuclear pore complex anchoring and import of a subset of nuclear proteins. Embo J 20(20):5703–5714
Wang GG, Song J, Wang Z et al (2009) Haematopoietic malignancies caused by dysregulation of a chromatin-binding PHD finger. Nature 459(7248):847–851. https://doi.org/10.1038/nature08036
Wang H, Wang L, Erdjument-Bromage H et al (2004) Role of histone H2A ubiquitination in Polycomb silencing. Nature 431(7010):873–878. https://doi.org/10.1038/nature02985
Xu H, Valerio DG, Eisold ME et al (2016) NUP98 fusion proteins interact with the NSL and MLL1 complexes to drive leukemogenesis. Cancer Cell 30(6):863–878. https://doi.org/10.1016/j.ccell.2016.10.019
Xu S, Powers MA (2010) Nup98-homeodomain fusions interact with endogenous Nup98 during interphase and localize to kinetochores and chromosome arms during mitosis. Mol Biol Cell 21(9):1585–1596. https://doi.org/10.1091/mbc.E09-07-0561
Yang Q, Rout MP, Akey CW (1998) Three-dimensional architecture of the isolated yeast nuclear pore complex: functional and evolutionary implications. Mol Cell 1(2):223–234
Yassin ER, Sarma NJ, Abdul-Nabi AM et al (2009) Dissection of the transformation of primary human hematopoietic cells by the oncogene NUP98-HOXA9. PLoS One 4(8):e6719. https://doi.org/10.1371/journal.pone.0006719
Zheng J (2013) Oncogenic chromosomal translocations and human cancer (review). Oncol Rep 30(5):2011–2019. https://doi.org/10.3892/or.2013.2677
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG
About this chapter
Cite this chapter
Raices, M., D’Angelo, M.A. (2018). Nuclear Pore Complexes in the Organization and Regulation of the Mammalian Genome. In: D’Angelo, M. (eds) Nuclear Pore Complexes in Genome Organization, Function and Maintenance. Springer, Cham. https://doi.org/10.1007/978-3-319-71614-5_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-71614-5_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-71612-1
Online ISBN: 978-3-319-71614-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)