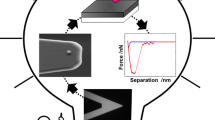
Abstract
Binnig et al. invented atomic force microscope (AFM) in 1986. It is a scanning tool for nanostructure investigations. It is now considered to be one of the landmarks of modern sciences, for citations of the first article increase more than 13,500 times [1]. The AFM has come up as a new addition to macroscopic and microscopic techniques since it has benefits in sample preparation and the ability of high-resolution imaging in both liquid and air environment if compared to standard light microscopy techniques. This sophisticated instrument has successfully been used in all branches of science like material science [2], food science [3], nanofabrication [4], and microbiology, for nearly two decades after its invention [5]. The microbiology field has been revolutionized by AFM. It has enriched the realm of sample preparation and microbial surface analysis in particular during the last two decades [6].
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
8.1 Introduction
Binnig et al. invented atomic force microscope (AFM) in 1986. It is a scanning tool for nanostructure investigations. It is now considered to be one of the landmarks of modern sciences, for citations of the first article increase more than 13,500 times [1]. The AFM has come up as a new addition to macroscopic and microscopic techniques since it has benefits in sample preparation and the ability of high-resolution imaging in both liquid and air environment if compared to standard light microscopy techniques. This sophisticated instrument has successfully been used in all branches of science like material science [2], food science [3], nanofabrication [4], and microbiology, for nearly two decades after its invention [5]. The microbiology field has been revolutionized by AFM. It has enriched the realm of sample preparation and microbial surface analysis in particular during the last two decades [6].
AFM has basically initiated as imaging instrument and provides topographic analysis of microbial surfaces in air with three-dimensional structural details with high resolution [7]. It was reformed in such a way that it can image the microbial surfaces in their natural environmental conditions [8] and to determine the interactive forces of these biological systems within 4 years of AFM invention [9]. For performance optimization, the AFM instrument has been continuously modified with improved sensing and originating features. Its basic purpose is to emerge new techniques like single-molecule force spectroscopy (SMFS), single-cell force spectroscopy (SCFS), and chemical force microscopy (CFM) [10].
It has thoroughly been observed that the AFM-based methods are sensitive enough to allow detection of forces ranging from ∼5 pN, that is approximately equal to the bond strength between a single receptor and ligand, to ∼100 nN, approximately equal to the force with which cells are stuck together [5, 11]. The spatial resolution of AFM approaches to ∼1 nm by using immobilized and extracted cell membranes patches [12], whereas AFM resolution is limited to ∼10 nm for dynamic and crenelated microbial surfaces [13].
The morphology and physicochemical properties of microbial systems are important in characterization of microbial interactions with different interfaces and surfaces. The surface morphology of microbial systems is determined with conventional microscopic techniques which require considerable sample preparation as a result of which the sample surface morphology can damage and produce various defects. In addition to it, the macroscopic techniques used for the determination of physicochemical properties of cell surface like contact angle measurements and zeta potential. These techniques provide averaged data on wide assemblages of cells. Fluorescence imaging is influential technique used for localization of single cells in intricate microbial systems [14], but it should be kept in mind that image resolution is dependent to the wavelength of light source. High-resolution images of microbial systems are obtained from electron microscopy techniques. In particular, images of the whole bacterial cells are acquired with three-dimensional electron microscopy or cryo-electron tomography at higher resolution up to two times higher than light microscopy [15]. High-resolution images of microbial systems can also be provided with transmission electron microscopy and scanning electron microscopy techniques, no doubt they are advantageous to some extent, but their limitations may not be neglected ever regarding complex sample preparation including metal coating, dehydration, chemical fixing, and ultrathin section can significantly deteriorate the sample. The above given limitations were mostly required to be elaborated with sophisticated technique so that its consequence may be clear and obvious. It was the need of the researchers to start getting their questions and answers with the invention of AFM. It is known that the AFM images were obtained through direct contact between the sample and the tip, for no or less sample preparation is required. Moreover, true three-dimensional images are provided with AFM, but it only focuses on the limited ranges in heights at one time. AFM is considered to be one of the most important techniques for biological applications, besides the microbial systems can be examined in buffer solution, which are very important for keeping an eye on the live cells in real-time scanning.
Atomic force microscopy has brought great changes in the way the researchers probe the microbial systems. AFM measures the small forces acting between the sample and a sharp tip instead of an incident beam of light [10, 16]. The tip is attached to a cantilever by using a piezoelectric scanner that bends under force and is moved in three dimensions to generate a topographic image. Cantilever’s bending is measured by a laser beam focused on the free end of the cantilever and reflected into a photodiode at the time of scanning the sample surface. In contrast to other microscopy techniques, three-dimensional images of microbial membranes and cells are acquired without fixation, staining, or labeling at high resolution, so in physiological conditions. The microbial systems comprise of highly complex cells which are present in heterogeneous systems. Normally, microbial cells are divided into subclasses which reveal different resistance to stress and growth rate [17]. Furthermore, these subclasses of cells are spatially organized in such a way that they can perform different key functions [18]. To understand such complex microbial systems required single-cell analysis techniques in microbial research. The mechanical and spatial properties of the molecules present on the surface of individual microbial cell are measured by AFM through single-molecule force spectroscopy, in which the cantilever deflection is recorded to be a function of the vertical displacement of the scanner (as the sample is pressed toward the tip) [16]. As a result, it is observed that there is a cantilever deflection versus scanner displacement curve and is transformed into a force distance curve by using suitable modifications. The distinctive adhesive forces between sample and tip are determined during retraction which is used to probe the mechanics and distribution of single molecules like microbial cell surface receptors.
Besides topographic imaging, AFM can also be operated in force spectroscopy mode to determine the physical properties and interactive forces of the microbial systems. A force distance curve is obtained in this operational mode, in which the force signal is determined by recording the cantilever deflection as the AFM tip reaches the sample and then withdraws from it. Furthermore, having generated a force volume image through obtaining force distance curves at different locations, spatial resolution can be accomplished. To study protein folding and unfolding mechanisms and to determine which molecular groups are present on microbial cell surface, functionalized AFM tips are being used [5, 19].
To comprehend cell surface interactions, these innovative AFM techniques are used for analyzing microbial cell walls and providing new opportunities for better understanding of microbial systems. In this chapter, a flavor of the different applications of AFM in microbiology is provided, and the key breakthroughs in pathogen research have been highlighted. This chapter will facilitate the reader and researcher in multidimensional disciplines.
8.2 Atomic Force Microscope: Instrumentation
The AFM instrument comprises of four major parts including a tip, a piezoelectric scanner, an optical deflection system comprising of photo detector and laser diode, and an electrical feedback system. The probing tip (made from silicon or silicon nitride) mounting at adjustable cantilever’s end is the heart of the AFM. High-resolution three-dimensional images are obtained by movement of either the sample or the cantilever mounting on a piezoelectric scanner by using AFM.
The basic principle of AFM microscopy technique is a sharp AFM tip raster over the surface during scan, at the time of sensing the interaction between the tip and the sample. The optical deflection system detects the cantilever deflection, and cantilever deflection is a result caused by small interactions forces between the AFM tip and the sample. The backside of the cantilever reflects the laser beam and stored into a four-quadrant photodetector, to record the position of the reflected laser beam. The interactive forces between the tip and the sample are responsible to bring changes in magnitude of deflected laser beam. The deflection signals are automatically administered to restructure an interaction force profile or a topographic image of the sample. To obtain a high-resolution image with simple protocol for sample preparation of the microbial systems, both in liquid and in air environments, AFM topographic analysis has emerged as an essential instrument in addition to standard electron and light microscopy techniques in the past years.
AFM images are generated in three different operational modes like contact, noncontact, and intermittent or tapping modes. The tip movement across the sample during analysis is the only difference among all these operational modes. In the analysis of biological samples, the contact and intermittent or tapping modes are mostly applied. Contact operational mode is most widely used in AFM analysis. In this operational mode, the tip of cantilever directly touches the sample during analysis. Contact mode generates two different types of images, height and deflection. The cantilever deflection is recorded to generate height image when the tip moves across the sample during scanning. Height image gives information about surface roughness and height of the analyzed samples. The sample height is adjusted by using feedback loop in order to keep constant the cantilever deflection when the tip moves across the sample during scanning to generate the deflection image. It is very difficult to keep constant the cantilever deflections as the feedback loop is not perfectly adjusted, so a constant signal error is always present in the deflection image. The deflection image is more sensitive to the fine surface information.
High lateral forces are associated in contact operational mode, as a result soft biological samples can be removed from substrate or damaged. Therefore, to prevent the sample damage, it is necessary to minimize the lateral forces generated from AFM tips. The lateral forces are minimized by using the tapping or intermittent operational mode. In this AFM operational mode, phase and amplitude of the cantilever are administered when the probe of AFM is externally excited around its resonating frequency when AFM tip moves across the sample surface during scanning. In intermittent mode, the tip of AFM comes in contact with surface periodically when the tip moves downward, and as a result sample damage is minimized. The three types of different images can be obtained in intermittent or tapping operational mode including height amplitude, and phase images. Topographic information like contact mode are obtained from height image. Fine structural details and main features of the edges of the sample surface are obtained at the sacrifice of height information in amplitude image which is just like deflection image in contact operational mode [20]. The phase image is generated by recording the phase lagging of cantilever oscillation relative to driving signal. It provides information about difference in properties and sample composition like adhesion and elasticity [21].
AFM force spectroscopy offers new and exciting opportunities to probe the local properties and ultrastructure of biological samples in physiological environment along with positional precision and high force sensitivity. In this operational mode, deflection of cantilever is monitored as a function when sample is vertically displaced and the tip is pushed forward toward the sample and then pushed back from it. This force curve is useful to gain observations about different surface properties like cell surface charges, cell surface, cell surface hydrophobicity, and cell wall elasticity. Remarkably, the curve between force and distance is recorded at different locations of the x-y plane which gives spatially resolved maps of physicochemical properties of microbial systems with lateral resolution in nanometer scale [22]. The data obtained in this way are then represented either in elasticity maps [22] or in adhesions maps [23].
The force spectroscopy has emerged into three different force modes including single-molecule force spectroscopy (SMFS), chemical force microscopy (CFM), and single-cell force spectroscopy (SCFS) over the years. The AFM tips are functionalized in all these techniques with biological molecules, viruses, chemical groups, or a living cell or replaced by particle. The key parameter which gives information about a specific ligand-receptor interaction (SMFS) [24], on the chemical group’s distribution (CFM) [25], about interactional forces which control cell-substrate and cell-cell interrelations (SCFS) [5, 11] is the characteristics force involve in unbinding of AFM tip which is observed during retraction of AFM tip from sample. CFM is used to probe charges of individual cells and local surface hydrophobicity. CFM operational mode of force spectroscopy gives a means for resolution of the interactions of living microorganisms and local chemical properties with nanoscale resolution [26]. The AFM tips chemically functionalized with a specific functional group give naval approaches for better understanding of structure activity relationship of the biological cell surfaces. This technique has also provided new avenues for interactions between the antimicrobial drugs and the microbial cell surfaces [27].
A pivotal provocation in cell biology is to understand how molecules bound to the cell surfaces are organized and interact with their surrounding environment [28]. Answers to these questions are provided by SMFS, which is used to analyze the complex biological systems with individual molecules by simultaneously manipulating and localizing single molecules present in live microbial cells, which then reveal properties and proceedings that on other hand would be unattainable. The AFM tip binds to molecule of interest through the complimentary pair and distensible cross linkers to the substrate in SMFS technique [29]. The force involved in the unbinding of the AFM tip on retraction from attached molecule present in the substrate is recorded in real time [30]. In SMFS the force curve is recorded to show a force with nonlinear elongation which reflects stretching of flexible molecules present on the AFM tip and on the sample, until the “full-off” force of adhesion is observed. The worm-like chain (WLC) [31] model and freely jointed chain (FJC) [32] model from statistical mechanics are used to describe the elongation force. Polymers are described as an irregular curved filament in WLC model as a linear on the scale of persistence length, which shows the stiffness of molecules. While, as polymers are described in FJC model as an array of rigid independent statistical segments (Kuhn), arranged via flexible-joints [33]. WLC model generally well describes the proteins which function as a continuous deformable rods [34], while FJC model is useful in describing polysaccharides which behave as a series of loosely connected segments [35]. Instead of AFM tip, microbial cell is fixed on cantilever in SCFS technique and force between the cell present on the probe and other cells present in the substrate is determined [36].
8.3 Cell Fixation
One of the most important difficult encounters in the microbial research which limit the AFM application is how to fix microbial cells without changing the variability or surface properties to flat surface. Firm fixation of the live microbial cells on the flat solid surface is obligatory for fruitful AFM topographic analysis and the force determinations. The method used for cell fixation should be selected in such a way that the cells should be fixed strongly enough to tolerate the lateral frictional forces applied by AFM tip without degrading microbial cell surfaces during scanning [37]. Analysis of microbial samples in liquid faces a major problem as microbial cells are flexible and soft, causing an additional hurdle in fixing. In such condition, to ensure firm fixation of microbial sample, surface modification of the substrate is necessary. One of the primary challenges in analysis of microbial sample is therefore microbial cell fixation.
In the beginning studies, AFM images of microbial cells were recorded after drying, as microbial sample preparation was easy in dry condition, but dried microbial cell can generate misleading information. A tremendous progress has been seen in the past two decades, in improving the existing and developing new protocols for cell fixation procedures of microbial cells. One of the most interesting applications of AFM in biological analysis deals with topographic analysis of the living cells in their native liquid environment, where microbial cell interaction with their surrounding environment can be studied [38].
Some of the most commonly used cell fixation techniques which do not require drying of the microbial cells include physical confinement of cells in a porous membrane; aluminum oxide filters; fixation of cells to surfaces coated with positively charged substances including gelatin [39], polyethyleneimine (PEI) [40], or poly-l-lysine [22, 41], by electrostatic interactions which are covalently bound to carboxyl- or amine-terminated surfaces [42]; and adsorption to coated surfaces with adhesively polyphenolic proteins [43]. All these cell fixation methods are used for cell immobilization, but cells attached via electrostatic interactions are generally detached during scanning, whereas reagents used for cross-linking and reactive groups used for covalent binding are known to affect the cells activity. Adsorption and entrapment techniques are effectively used for viable cell fixation. However, coccoid cells are physically confined in membranes effectively.
In physiological conditions, the living microorganisms are successfully immobilized using adhesively polyphenolic proteins, which strongly fix the microorganism without affecting cells’ viability [43]. A universal method for the cell fixation of living microorganism reported by Dague et al., where the living cells are assembled within the arranged microstructured, using capillary/convective deposition with functionalized poly-dimethysiloxane (PDMS) brands. The results of Dague et al. demonstrate the great potential for assemblance of microorganism on microstructure, by using capillary/convective deposition with functionalized PDMS brands for performing precise and accurate experimentations on the living microbial cells using AFM [37].
8.4 Functionalized AFM Tips
Silicon or silicon nitride-made tips are often used as probe in AFM microscopy. The AFM probe may be bare or chemically functionalized and often attached with a fixed cell or particle onto an AFM tip or tipless cantilever. The colloidal probe technique is introduced by Ducker et al. in 1991, consisting of the AFM tipless cantilevers to an attached micron-sized spherical particle, after that it measures the force between the sample and the sphere [44]. Though this protocol provides controlled surface geometry and chemistry, but to get images with high resolution and to map heterogeneous chemical surfaces, application of this large colloidal probes is not useful [45].
Vakarelski and Higashitani used the colloidal probe technique remarkably well by functionalizing the AFM tips with nanoparticles by using wet chemistry application. They successfully submitted a single-gold nanoparticles of the size 10–40 nm at the end of AFM tip, to avoid the limitation applied by either glue contamination or micromanipulation [46]. A new protocol for microbial cells fixation is reported by Razatos et al. involving coating of silicon nitride cantilevers with connecting bacterial layer, followed by covalent crosslinking between the tip and the cells via glutaraldehyde [47].
The bacteria were attached to 3-MPTS-coated glass beads, and furthermore, cell-coated beads were also attached to a tipless cantilever by Lower et al. using epoxy resin [48]. The silicon nitride cantilevers were pretreated with poly-l-lysine, followed by coating these cantilevers with bacterial layer by Touhami et al. [49] Various strategies have been reported for cell probe preparation, to analyze a single cell of bacterium after the pioneer work of above research groups.
It is worth noting that some limitation exists in cell probe method like attachment of cell with AFM probe may either results in denaturation of cell or may change the cell structure and properties which limits high lateral resolution during analysis via AFM [50]. Avoiding these shortcomings, a novel preparation protocol for live single-cell probe using adhesive bioinspired polydopamine is introduced by Kang and Elimelech [51]. Their findings manipulate that after performing force measurements following the bacterium attachment to tipless cantilevers, bacterium was still alive. Single molecules are fixed to the AFM tips by universally available cross-linkers like polyethylene glycol to prevent denaturation of sample and motional freedom [52]. Primarily, the AFM tips were functionalized with amino groups which were then reacted with benzaldehyde functional groups carrying polyethylene glycol cross-linkers which via lysine residues attach directly to proteins [53].
Self-assembled monolayers (SAMs) of alkanethiols on gold-functionalized AFM tips are attached to single molecules [54]. At molecular level, alkanethiolate SAMs functionalized with gold are well defined structurally. The alkyl chain residues play a vital role in the stabilization through its lateral hydrophobic interactions.
8.5 Microbial Cells Imaging
AFM imaging is an influential supplementary tool if compared to electron microscopy, offering new paradigm for visualization of supramolecular organizations of the microbial systems in liquid or air, in actual time [55].
Primarily, the AFM was used predominantly to encounter the difficulties involved in air, by fixation of delicate samples of microbes on different substrates. In air, the AFM imaging is also useful to minimize the contact area between the substrate and the cell as a result of tumid state of the living cell of bacterium. While scanning, the AFM probe detaches the cell from the substrates [56].
Hayhurst et al. present an interesting AFM imaging of broken sacculi in Bacillus subtilis (Fig. 8.1) in air with very high resolution [57].
According to Gillis et al. , air AFM imaging is simpler and more reliable approach than liquid AFM imaging to monitor morphologies of bacterial surfaces and to measure dimensions and densities of bacterial flagella. Figure 8.2 focuses on the deflective images in air of mutant strains of Bacillus thuringiensis elaborating different levels of flagellation [58].
Having the ability to image specimen in liquid, a new era has been opened by AFM to observe live cells in the nanoregime, and as a result various structural informations can be achieved surpassing the electron and optical microscopy.
An interesting AFM analysis showing images in real time, in liquid is represented by Plomp et al. in a dynamic way of Bacillus atrophaeus spore germination; which reveals changes in spore coat structure and topography induced by germination as shown in the Fig. 8.3.
Diaz et al. claimed that AFM analysis of Pseudomonas fluorescens on organized substrates is a useful tool in air with (70%) humidity contents, to observe orientation of flagella on substrates containing metal without the use of any chemical during the primary level of the biofilm formation. They reported a random orientation of flagella on noble surfaces, like organized gold with sub-microtrenches (Fig. 8.4a) and nano-oriented gold with random arrangements as shown in Fig. 8.4b. Flagella are used for different functions like like movements, keeping cells in contact, oscillating them, possibly involving in signaling functions, and furthermore, it facilitates the accumulating process at initial stage of biofilm formation [59].
Andre et al. have figured out by using AFM topographic imaging in buffers that mutant Lactobacillus plantarum cells lacking in wall-teichoic-acids (WTAs) which reveals rough morphology of cells surfaces (Fig. 8.5 a and b), However, strains of Lactococcus plantarum showing WTAs, the surface morphology was highly polarized, according to AFM images the side walls were less smooth than the pores (Fig. 8.5 d and e) [60].
AFM topographic images of cells of mutant L. plantarum. (a, b) Deflection images in sodium acetate buffer of single WTA-deficient cells, showing rough surface morphology, lateral (a) and polar (b) regions. (d, e) Deflection images in sodium acetate buffer of cell wall polysaccharide-deficient cells, showing polarized surface morphology
Dorobantu et al. pictured the Acinetobacter venetianus RAG-1 surface using chemically functionalized AFM tips and AFM imaging terminating in hydrophilic and hydrophobic groups in buffer [42]. It can be witnessed in Fig. 8.6a, obtained with hydrophilic AFM tip, the existence of the cell surrounded by pili that are thin fibrils, in contrast to image shown in Fig. 8.6b, obtained with hydrophobic AFM tip, where no extracellular structure is seen around A. venetianus RAG-1 cell.
The thin fibrils (pili) are clearly visible in AFM analysis carried out with hydrophilic AFM tips, but when the AFM analysis is carried out with hydrophobic AFM tips, these thin fibrils are not visible. Such investigations potently propose that these thin fibrils are totally hydrophobic in nature. As soon as the hydrophilic tip reaches near these thin fibrils, their structure remained fixed on substrate, and hydrophilic AFM tips record their morphologies. In opposite to it, as the hydrophobic tip touches the structures, they stick to the hydrophobic AFM tip and move along it. Specially, regarding this case, the AFM tip gets failed to search the presence of these structures [42].
AFM imaging of Staphylococcus aureus trapped in filters is used by Turner et al. (etched for 4 h having 990 ± 20 diameters). To draw the AFM image of the division of microorganism with high respect, it is absorbed in brain heart infusion broth (BHI) (Fig. 8.7) [61]. They presented the dynamic way for higher molecular resolution which can be practiced in those cases where microorganisms are fully fixed on the substrate.
The surface of mycobacteria was analyzed by Alsteens et al., with four antimicrobial drugs prior to and after incubation [62].
Shah et al. reported recently the antibacterial potential of silver and gold nanoparticles stabilized with ceftriaxone against Escherichia coli ATCC 8739 with respect to kinetics and surface roughness. Conjugation of ceftriaxone with silver and gold nanoparticles improves the kinetics of antibiotics, and cell morphologies were completely disrupted in just 2 h of incubation, which was far better than the bare ceftriaxone (Fig. 8.8) [63].
8.6 Force Spectroscopy
The AFM instrument’s force measurement potential compounds the importance of AFM spectroscopy; it keeps on going to avail unique visions into the interactions and mechanical property of microbial world.
SMFS used by Gad et al. for the first time in the domain of research points to specific interaction forces between pairs of receptor ligand. The credit also goes to the research group to draw the position of polysaccharide on the live cell surface of microbes. Gold-coated AFM tips fuctionalized with concanavalin-A made possible to determine binding forces between mannan polymers and concanavalin-A on the yeast Saccharomyces cerevisiae surface [64].
Next to them Lower et al. is one of the successful scientists who used SCFS to determine the interactive forces between goethite (a-FeOOH) and Shewanella oneidensis: anaerobic and aerobic conditions were used for the above given process. From the force measurements energy value was derived, it showed that in anaerobic conditions S. oneidensis gives response to the surface of goethite with the help of rapid developing stronger adhesive allergies at boundary if compared to the aerobic environment [65].
Pseudomonas aeruginosa is attached to AFM tips by Touhami et al. The bacterial cell is fastened on mica surface by thin fibril tethers. Soon after, it was observed by them that interactive forces are associated with elastic properties of thin fibrils between these thin fibrils and hard surfaces of mica. Force extension curves of the thin fibrils obtained from P. aeruginosa were qualitatively determined and were found to be similar to the different variety of biopolymers nonlinear stretching curves [49]. To probe complexity in connection with two bacterial surfaces, Rhodococcus erythropolis 20S-E1-c and A. venetianus RAG-1, Dorobantu et al. practiced AFM tips derivatized with alkanethiols finishing at hydrophobic and hydrophilic functional groups in CFM operational mode [42]. There was no difference in the connection involved for the two different species of bacteria observed at the time of hydrophilic tips that were in action, distribution of interactive forces of a patchy surface was clearly observed between two different cell surfaces, as well as highest forces were collected with hydrophobic AFM tips in one end of cell for R. erythropolis 20S-E1-c. In a nut shell, CFM with derivatized hydrophobic AFM tips is allowed to differentiate between microbial surface adhesion differences that could have meaningful significance for the cell knitting to hydrophobic interfaces or surfaces.
Recently, Alsteens et al. used single molecule force spectroscopy to unfold single cell adhesion proteins (Als5p), found on Candida albicans surface (Fig. 8.9) [66]. The WLC model elaborates the force extension curve which shows saw-toothed pattern with well-defined force peak signals. The core purpose of these single-molecule measurements is to avail inside into mechanical properties of adhesive molecules. It will assist the researchers of the pure scientific community to elaborate and elucidate their potential application and implications in different kind of diseases.
The unfolding mechanism of adhesive cell proteins on C. albicans surface. (a) Representation of an AlS molecule projecting outward from the C. albicans cell wall by means of the stalk region and (b) principle of the SMFS experiments. Ig-T-TR6 fragments were attached on a gold surface and stretched via their Ig domains using an Ig-T tip; (c) force extension curves obtained by stretching single Ig-T-TR6 showed periodic features reflecting the sequential unfolding of the TR domains (upper traces). Force peaks were well described by the WLC model (inset, red line). Addition of urea dramatically altered the unfolding peaks (lower traces) [66]
SMFS is used by Andre et al. for the investigations to find out if peptidoglycan is really concealed by polymers present in cell surface of Lactococcus lactis wild type (WT) [67]. Andre et al. depict the proof of their hypothesis that L. lactis mutants VES5751 and VES5748 which clarify confusion in the domain of research will encode synthesis of polysaccharides present in cell wall. In the synthesis of cell wall, the use of mutant strains is vital; polysaccharides (WPS) made it possible for SMFS to manipulate original architecture of peptidoglycan present in cell wall. Force curves analyzed spatially were obtained across surfaces of L. lactis WT cells. Curves associated to interacting forces were obtained spatially by analyzing the surfaces of L. lactis WT cells and the two mutant lacking cells WPS with the help of lysine functionalized AFM tips. Lysin is basically a protein that specifically attached to peptidoglycan in particular (Fig. 8.10).
Single-molecule recognition imaging of peptidoglycan. (a, d, g) Deflection images recorded with silicon nitride tips on L. lactis WT (a), VES5748 WPS mutant, (d) and VES5751 WPS mutant (g). (b, e, b) Adhesion force maps (400 × 400 nm) recorded on the three strains with LysM tips in the square areas shown in the deflection images, using a maximum applied force of 250 pN. (c, f, i) Force curves along with adhesion histograms
Adhesion histogram, deflection image, and typical force curve are being showed with the help of Fig. 8.10a–c, drawn on wild-type L. lactis with a LysM tip at low applied forces (250 pN). No binding events were showed by most force curves. This experiment affirms that other cell wall components are involved in covering of peptidoglycan. A considerable fractional curve having different behavior is presented by the VES5751 WPS mutant (Fig. 8.10g–i), which shows single well-pronounced maximum at 71 ± 16 pN. The rupture of specific LysM peptidoglycan complexes is reflected by this process.
The analysis of mutant VES5751 WPS shows the same conclusion, but binding forces were regularly observed in 100–200 pN range (Fig. 8.10g–i), predicting that concurrent detection at the same time of two or three molecules was more frequently occurred. Finally, the detection of peptidoglycan on enriched mutant cell surfaces reflects absence of outer polysaccharide layers. From these results, a new era has been opened for understanding the assembly and architecture of peptidoglycan in gram-positive bacteria [67].
For the first time in literature, Park and Abu-Lail used AFM measurements and published heterogeneities in the adhesive energies that can be measured between bacterial cells and silicon nitride model surfaces which represent all species of Listeria genus in water. Larger adhesive energies to silicon nitride characterize the pathogenic species of Listeria as a result. More heterogeneities can characterize them if compared to the nonpathogenic species [68].
AFM force measurements of two biofilm positive and two nonbiofilm forming strains of Staphylococcus epidermidis in the pore confined state were measured directly in aqueous media by Hu et al. To probe adhesion on living cells at the nano-level was their core objective. They used AFM tips functionalized with hydrophobic functional groups in CFM operational mode. Their main objective was to list the minute differences among the four strains of Staphylococcus epidermidis. No detectable difference is shown by AFM topographic analysis. Bare hydrophilic silicon nitride tips are used to perform force measurements. They disclose similar adhesive characteristics for every strain; nevertheless, hydrophobic tip showed that hydrophobic interactions are not the main operation in analysis of four different strains of Staphylococcus epidermidis.
In a certain degree, the presence of modular proteins is observed as a result of the effect of saw-toothed force distance arrangements on biofilm surface of positive strains registered which may mediate the process of cell adhesion. This observation is considerably remarkable, as the dynamic silent features are shown, which can offer more confirmation tests for the surface adhesion if compared to the static effect that have been reflected in AFM analysis. [69]
8.7 Conclusion
A treasure of new opportunities is provided by AFM technique for imaging and for using living microbial cells. AFM technique provides innovative intuitions into structure-activity relationship of microbial cells and also images the microbial cell up to single molecular resolution even in physiological environment. Modern development in AFM technology in the field of microbiology has made the scientists able enough to answer related questions concerning microbiology field, like cell signaling and adhesion, shape, tissue and embryonic development, microbial pathogenesis, and cell division [70]. AFM technique has facilitated the direct observations of dynamic structural changes in live microbial systems. In addition to it, the instrumental developments have made possible the dynamic interactions between individual biological macromolecules and cells; they were not getable with other visualization techniques [71].
In this chapter, the principle of AFM is described in detail and discussed in outline; the successes of the present era have been made in AFM spectroscopy regarding topographic analysis of microbial cells and force measurements. Highlighting the core of this chapter is to ask how recent technology has enhanced our view of molecular organization, interaction mechanics, and mechanisms of microbial cell surfaces. The drawbacks and advantages of AFM techniques are also presented in this chapter. Furthermore the challenges will be addressed in the next research of microbial systems.
There is a need for high speed AFM instruments for topographic analysis of microbial cells and to study interactions between the microbial cells in real time with high resolution [72], and their intercommunications in actual time with high resolution will be focused. The AFM ability to pave surface morphology of an individual living cell at the time of growing or interacting with different drugs, for instance, AFM imaging in actual time, discloses new opportunities for studying, remodeling, and assembling of cell walls. It is also worth notable to understand the mode of action of antibiotics [19]. High-speed AFM instrument is therefore expected, which will revolutionize the biological world and will impact physiological process in disease diagnosis and treatments [71].
The applications of SCFS are still under action; it has possessed too much potentiality for betterment. The potential of SCFS is founded on multifunctional applications. It has a magnificent variety of applications in medical as well as in microbiology fields to which it is applicable [36].
The AFM instrumental developments are used to couple with other microscopic techniques or scanning probe microscopy instruments, which will represent a way for instrumental development in the future that will revolutionize the biological sciences in multidimensions [73].
References
Binnig G, Quate CF, Gerber C (1986) Atomic force microscope. Phys Rev Lett 56(9):930
(a) Bhushan B, Kwak KJ, Palacio M (2008) Nanotribology and nanomechanics of AFM probe-based data recording technology. J Phys Condens Mat 20(36):365207; (b) Johnson LL (2008) Atomic force microscopy (AFM) for rubber. Rubber Chem Technol 81(3):359–383; (c) Withers JR, Aston DE (2006) Nanomechanical measurements with AFM in the elastic limit. Adv Colloid Interface Sci 120(1):57–67
Yang H, Wang Y, Lai S, An H, Li Y, Chen F (2007) Application of atomic force microscopy as a nanotechnology tool in food science. J Food Sci 72(4):R65–R75
(a) Simeone FC, Albonetti C, Cavallini M (2009) Progress in micro-and nanopatterning via electrochemical lithography. J Phys Chem C 113(44):18987–18994; (b) Tseng AA, Jou S, Notargiacomo A, Chen T (2008) Recent developments in tip-based nanofabrication and its roadmap. J Nanosci Nanotechnol 8(5):2167–2186
(a) Cohen SR, Bitler A (2008) Use of AFM in bio-related systems. Curr Opin Colloid Interface Sci 13(5):316–325; (b) Dufrêne YF (2003) Recent progress in the application of atomic force microscopy imaging and force spectroscopy to microbiology. Curr Opin Microbiol 6(3):317–323; (c) Müller DJ, Krieg M, Alsteens D, Dufrêne YF (2009) New frontiers in atomic force microscopy: analyzing interactions from single-molecules to cells. Curr Opin Biotechnol 20(1):4–13
Dufrêne YF (2004) Refining our perception of bacterial surfaces with the atomic force microscope. J Bacteriol 186(11):3283–3285
Gould S, Drake B, Prater C, Weisenhorn A, Manne S, Hansma H, Hansma P, Massie J, Longmire M, Elings V (1990) From atoms to integrated circuit chips, blood cells, and bacteria with the atomic force microscope. J Vac Sci Technol A 8(1):369–373
Weisenhorn A, Drake B, Prater C, Gould S, Hansma P, Ohnesorge F, Egger M, Heyn S, Gaub H (1990) Immobilized proteins in buffer imaged at molecular resolution by atomic force microscopy. Biophys J 58(5):1251–1258
Weisenhorn A, Hansma P, Albrecht T, Quate C (1989) Forces in atomic force microscopy in air and water. Appl Phys Lett 54(26):2651–2653
Dorobantu LS, Goss GG, Burrell RE (2012) Atomic force microscopy: a nanoscopic view of microbial cell surfaces. Micron 43(12):1312–1322
Müller DJ, Helenius J, Alsteens D, Dufrêne YF (2009) Force probing surfaces of living cells to molecular resolution. Nat Chem Biol 5(6):383–390
Müller DJ, Engel A (2007) Atomic force microscopy and spectroscopy of native membrane proteins. Nat Protoc 2(9):2191–2197
Plomp M, Leighton TJ, Wheeler KE, Hill HD, Malkin AJ (2007) In vitro high-resolution structural dynamics of single germinating bacterial spores. Proc Natl Acad Sci 104(23):9644–9649
(a) Daniel RA, Errington J (2003) Control of cell morphogenesis in bacteria: two distinct ways to make a rod-shaped cell. Cell 113(6):767–776; (b) Gitai Z (2009) New fluorescence microscopy methods for microbiology: sharper, faster, and quantitative. Curr Opin Microbiol 12(3):341–346
Milne JL, Subramaniam S (2009) Cryo-electron tomography of bacteria: progress, challenges and future prospects. Nat Rev Microbiol 7(9):666–675
(a) Engel A, Müller DJ (2000) Observing single biomolecules at work with the atomic force microscope. Nat Struct Mol Biol 7(9):715–718; (b) Dufrêne YF (2008) Towards nanomicrobiology using atomic force microscopy. Nat Rev Microbiol 6(9):674–680
Lidstrom ME, Konopka MC (2010) The role of physiological heterogeneity in microbial population behavior. Nat Chem Biol 6(10):705–712
Campos M, Jacobs-Wagner C (2013) Cellular organization of the transfer of genetic information. Curr Opin Microbiol 16(2):171–176
Dufrêne YF (2008) AFM for nanoscale microbe analysis. Analyst 133(3):297–301
Vié V, Giocondi M-C, Lesniewska E, Finot E, Goudonnet J-P, Le Grimellec C (2000) Tapping-mode atomic force microscopy on intact cells: optimal adjustment of tapping conditions by using the deflection signal. Ultramicroscopy 82(1):279–288
Dorobantu LS, Gray MR (2010) Application of atomic force microscopy in bacterial research. Scanning 32(2):74–96
Schaer-Zammaretti P, Ubbink J (2003) Imaging of lactic acid bacteria with AFM—elasticity and adhesion maps and their relationship to biological and structural data. Ultramicroscopy 97(1):199–208
Radmacher M, Cleveland JP, Fritz M, Hansma HG, Hansma PK (1994) Mapping interaction forces with the atomic force microscope. Biophys J 66(6):2159–2165
Hinterdorfer P, Dufrêne YF (2006) Detection and localization of single molecular recognition events using atomic force microscopy. Nat Methods 3(5):347–355
Frisbie CD, Rozsnyai LF, Noy A, Wrighton MS, Lieber CM (1994) Functional group imaging by chemical force microscopy. Science 265(5181):2071–2074
(a) Dorobantu LS, Bhattacharjee S, Foght JM, Gray MR (2009) Analysis of force interactions between AFM tips and hydrophobic bacteria using DLVO theory. Langmuir 25(12):6968–6976; (b) Noy A (2006) Chemical force microscopy of chemical and biological interactions. Surf Interface Anal 38(11):1429–1441
Dague E, Alsteens D, Latgé J-P, Verbelen C, Raze D, Baulard AR, Dufrêne YF (2007) Chemical force microscopy of single live cells. Nano Lett 7(10):3026–3030
Dupres V, Alsteens D, Andre G, Verbelen C, Dufrêne YF (2009) Fishing single molecules on live cells. Nano Today 4(3):262–268
Francius G, Alsteens D, Dupres V, Lebeer S, De Keersmaecker S, Vanderleyden J, Gruber HJ, Dufrêne YF (2009) Stretching polysaccharides on live cells using single molecule force spectroscopy. Nat Protocols 4(6):939–946
Horejs C, Ristl R, Tscheliessnig R, Sleytr UB, Pum D (2011) Single-molecule force spectroscopy reveals the individual mechanical unfolding pathways of a surface layer protein. J Biol Chem 286(31):27416–27424
Bustamante C, Bryant Z, Smith SB (2003) Ten years of tension: single-molecule DNA mechanics. Nature 421(6921):423–427
Rief M, Oesterhelt F, Heymann B, Gaub HE (1997) Single molecule force spectroscopy on polysaccharides by atomic force microscopy. Science 275(5304):1295–1297
Marszalek PE, Dufrêne YF (2012) Stretching single polysaccharides and proteins using atomic force microscopy. Chem Soc Rev 41(9):3523–3534
Oberhauser AF, Marszalek PE, Erickson HP, Fernandez JM (1998) The molecular elasticity of the extracellular matrix protein tenascin. Nature 393(6681):181–185
Marszalek PE, Oberhauser AF, Pang Y-P, Fernandez JM (1998) Polysaccharide elasticity governed by chair–boat transitions of the glucopyranose ring. Nature 396(6712):661–664
Helenius J, Heisenberg C-P, Gaub HE, Muller DJ (2008) Single-cell force spectroscopy. J Cell Sci 121(11):1785–1791
Dague E, Jauvert E, Laplatine L, Viallet B, Thibault C, Ressier L (2011) Assembly of live micro-organisms on microstructured PDMS stamps by convective/capillary deposition for AFM bio-experiments. Nanotechnology 22(39):395102
Fukuma T, Kobayashi K, Matsushige K, Yamada H (2005) True molecular resolution in liquid by frequency-modulation atomic force microscopy. Appl Phys Lett 86(19):193108
(a) Beckmann M, Venkataraman S, Doktycz MJ, Nataro JP, Sullivan CJ, Morrell-Falvey JL, Allison DP (2006) Measuring cell surface elasticity on enteroaggregative Escherichia coli wild type and dispersin mutant by AFM. Ultramicroscopy 106(8):695–702; (b) Doktycz M, Sullivan C, Hoyt P, Pelletier D, Wu S, Allison D (2003) AFM imaging of bacteria in liquid media immobilized on gelatin coated mica surfaces. Ultramicroscopy 97(1):209–216
(a) D’souza S, Melo J, Deshpande A, Nadkarni G (1986) Immobilization of yeast cells by adhesion to glass surface using polyethylenimine. Biotechnol Lett 8(9):643–648; (b) Velegol SB, Logan BE (2002) Contributions of bacterial surface polymers, electrostatics, and cell elasticity to the shape of AFM force curves. Langmuir 18(13):5256–5262.
Bolshakova A, Kiselyova O, Filonov A, Frolova OY, Lyubchenko YL, Yaminsky I (2001) Comparative studies of bacteria with an atomic force microscopy operating in different modes. Ultramicroscopy 86(1):121–128
(a) Camesano TA, Natan MJ, Logan BE (2000) Observation of changes in bacterial cell morphology using tapping mode atomic force microscopy. Langmuir 16(10):4563–4572; (b) Dorobantu LS, Bhattacharjee S, Foght JM, Gray MR (2008) Atomic force microscopy measurement of heterogeneity in bacterial surface hydrophobicity. Langmuir 24(9):4944–4951
Louise Meyer R, Zhou X, Tang L, Arpanaei A, Kingshott P, Besenbacher F (2010) Immobilisation of living bacteria for AFM imaging under physiological conditions. Ultramicroscopy 110(11):1349–1357
Ducker WA, Senden TJ, Pashley RM (1991) Direct measurement of colloidal forces using an atomic force microscope. Nature 353(6341):239–241
Takano H, Kenseth JR, Wong S-S, O'Brie JC, Porter MD (1999) Chemical and biochemical analysis using scanning force microscopy. Chem Rev 99(10):2845–2890
Vakarelski IU, Higashitani K (2006) Single-nanoparticle-terminated tips for scanning probe microscopy. Langmuir 22(7):2931–2934
Razatos A, Ong Y-L, Sharma MM, Georgiou G (1998) Molecular determinants of bacterial adhesion monitored by atomic force microscopy. Proc Natl Acad Sci 95(19):11059–11064
Lower SK, Tadanier CJ, Hochella MF Jr (2000) Measuring interfacial and adhesion forces between bacteria and mineral surfaces with biological force microscopy. Geochim Cosmochim Acta 64(18):3133–3139
Touhami A, Jericho MH, Boyd JM, Beveridge TJ (2006) Nanoscale characterization and determination of adhesion forces of Pseudomonas aeruginosa pili by using atomic force microscopy. J Bacteriol 188(2):370–377
Dufrêne YF (2002) Atomic force microscopy, a powerful tool in microbiology. J Bacteriol 184(19):5205–5213
Kang S, Elimelech M (2009) Bioinspired single bacterial cell force spectroscopy. Langmuir 25(17):9656–9659
Verbelen C, Dupres V, Alsteens D, Andre G, Dufrene YF (2011) Single-molecule force spectroscopy of microbial cell envelope proteins. In: Dufrene, Y.F. (Ed.), Life at the Nanoscale – Atomic Force Microscopy of Live Cells. Pan Stanford Publishing Pte. Ltd., Singapore, pp. 317–334
Riener CK, Stroh CM, Ebner A, Klampfl C, Gall AA, Romanin C, Lyubchenko YL, Hinterdorfer P, Gruber HJ (2003) Simple test system for single molecule recognition force microscopy. Anal Chim Acta 479(1):59–75
Dupres V, Menozzi FD, Locht C, Clare BH, Abbott NL, Cuenot S, Bompard C, Raze D, Dufrêne YF (2005) Nanoscale mapping and functional analysis of individual adhesins on living bacteria. Nat Methods 2(7):515–520
Francius G, Lebeer S, Alsteens D, Wildling L, Gruber HJ, Hols P, Keersmaecker SD, Vanderleyden J, Dufrêne YF (2008) Detection, localization, and conformational analysis of single polysaccharide molecules on live bacteria. ACS Nano 2(9):1921–1929
Kailas L, Ratcliffe E, Hayhurst E, Walker M, Foster S, Hobbs J (2009) Immobilizing live bacteria for AFM imaging of cellular processes. Ultramicroscopy 109(7):775–780
Hayhurst EJ, Kailas L, Hobbs JK, Foster SJ (2008) Cell wall peptidoglycan architecture in Bacillus subtilis. Proc Natl Acad Sci 105(38):14603–14608
Gillis A, Dupres V, Delestrait G, Mahillon J, Dufrêne YF (2012) Nanoscale imaging of Bacillus thuringiensis flagella using atomic force microscopy. Nanoscale 4(5):1585–1591
Díaz C, Schilardi P, Salvarezza R, Fernández Lorenzo de Mele M (2011) Have flagella a preferred orientation during early stages of biofilm formation?: AFM study using patterned substrates. Colloids Surf B Biointerfaces 82(2):536–542
Andre G, Deghorain M, Bron PA, van Swam II, Kleerebezem M, Hols P, Dufrêne YF (2011) Fluorescence and atomic force microscopy imaging of wall teichoic acids in Lactobacillus plantarum. ACS Chem Biol 6(4):366–376
Turner R, Thomson N, Kirkham J, Devine D (2010) Improvement of the pore trapping method to immobilize vital coccoid bacteria for high-resolution AFM: a study of Staphylococcus aureus. J Microsc 238(2):102–110
Alsteens D, Verbelen C, Dague E, Raze D, Baulard AR, Dufrêne YF (2008) Organization of the mycobacterial cell wall: a nanoscale view. Pflüg Arch Eur J Phys 456(1):117–125
Shah MR, Ali S, Ateeq M, Perveen S, Ahmed S, Bertino MF, Ali M (2014) Morphological analysis of the antimicrobial action of silver and gold nanoparticles stabilized with ceftriaxone on Escherichia coli using atomic force microscopy. New J Chem 38(11):5633–5640
Gad M, ITOH A, IKAI A (1997) Mapping cell wall polysaccharides of living microbial cells using atomic force microscopy. Cell Biol Int 21(11):697–706
Lower BH, Yongsunthon R, Shi L, Wildling L, Gruber HJ, Wigginton NS, Reardon CL, Pinchuk GE, Droubay TC, Boily J-F (2009) Antibody recognition force microscopy shows that outer membrane cytochromes OmcA and MtrC are expressed on the exterior surface of Shewanella oneidensis MR-1. Appl Environ Microbiol 75(9):2931–2935
Alsteens D, Dupres V, Klotz SA, Gaur NK, Lipke PN, Dufrêne YF (2009) Unfolding individual Als5p adhesion proteins on live cells. ACS Nano 3(7):1677–1682
Andre G, Kulakauskas S, Chapot-Chartier M-P, Navet B, Deghorain M, Bernard E, Hols P, Dufrêne YF (2010) Imaging the nanoscale organization of peptidoglycan in living Lactococcus lactis cells. Nat Commun 1:27
Park B-J, Abu-Lail NI (2011) Atomic force microscopy investigations of heterogeneities in the adhesion energies measured between pathogenic and non-pathogenic Listeria species and silicon nitride as they correlate to virulence and adherence. Biofouling 27(5):543–559
Hu Y, Ulstrup J, Zhang J, Molin S, Dupres V (2011) Adhesive properties of Staphylococcus epidermidis probed by atomic force microscopy. Phys Chem Chem Phys 13(21):9995–10003
Müller DJ, Dufrêne YF (2011) Atomic force microscopy: a nanoscopic window on the cell surface. Trends Cell Biol 21(8):461–469
Ando T, Uchihashi T, Fukuma T (2008) High-speed atomic force microscopy for nano-visualization of dynamic biomolecular processes. Prog Surf Sci 83(7):337–437
Shibata M, Yamashita H, Uchihashi T, Kandori H, Ando T (2010) High-speed atomic force microscopy shows dynamic molecular processes in photoactivated bacteriorhodopsin. Nat Nanotechnol 5(3):208–212
Francis LW, Lewis PD, Wright CJ, Conlan RS (2010) Atomic force microscopy comes of age. Biol Cell 102(2):133–143
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG
About this chapter
Cite this chapter
Shah, M.R., Ateeq, M. (2018). Atomic Force Microscopy for Microbial Cell Surfaces. In: Jackson, M., Ahmed, W. (eds) Micro and Nanomanufacturing Volume II. Springer, Cham. https://doi.org/10.1007/978-3-319-67132-1_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-67132-1_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-67130-7
Online ISBN: 978-3-319-67132-1
eBook Packages: EngineeringEngineering (R0)