Abstract
Cells display a high degree of functional organization, largely attributed to the intracellular biopolymer scaffold known as the cytoskeleton. This inherently complex structure drives the system out of equilibrium by constantly consuming energy to conserve or reorganize its structure. Thus, the active, structurally organized cytoskeleton is the key player for the emergent mechanical properties of cells, which further determine properties of cell clusters and even multicellular organisms. In this spirit, this chapter introduces the physical principles on the different levels of biological complexity ranging from single biopolymers to tissues. The emergent mechanical properties and their respective effects on each level will be highlighted with a strong emphasis on their intertwined nature.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- Cytoskeleton
- Rheology
- Collective behavior
- Tissue mechanics
- Cell mechanics
- Probing techniques
- Tissue dynamics
- Mechanical properties
1 Introduction
The tremendous complexity of biological matter emerges from the interplay between intertwined levels or scales, with each level contributing a rich repertoire of physical principles. To uncover these principles and their interplay has proven to be a nontrivial task since processes, which we consider the fundamentals of life, exist far from thermodynamic equilibrium. Thus, traditional, purely reductionist approaches are unsuitable to fully elucidate and describe biological soft matter [1,2,3]. Generally, complex systems are difficult to capture by intuitive understanding, which rather impedes an abstraction of the system in form of a model. When dealing with living matter, we can refer to different levels of complexity, which can be assigned to physical scales [4]. In this framework, a higher level contains the lower level, and complexity necessarily increases, which can even lead to entirely new properties [1]. If principles of the macrostate are absent in the underlying microstate, they are considered emergent (e.g., a single fish cannot exhibit swarm behavior). The term emergence describes the process leading to novel emergent properties, with the prefixes “micro” and “macro” not referring to definite length scales but to different levels of biological complexity [4]. For biological matter, a possible hierarchy might be described as
protein → filament → network → cell → tissue.
Using this hierarchy as a point of departure, we can aim at describing a given system only on the basis of the next underlying level of complexity, an approach termed hierarchical reductionism or coarse-graining [5, 6]. A biological tissue, for example, can be described as an accumulation of cells and extracellular matrix without considering subcellular structures. Further, cell mechanics can be described in terms of principles of networks of filaments. In this process, the problem is reduced, losing the detail of lower levels, similar to the use of computers, where not every single transistor has to be considered to operate the machine.
This chapter describes the physical principles emerging at the different levels of complexity and how they can be scaled up. In this context, it is important to clearly distinguish the concepts of self-organization (processes driven by energy dissipation) and self-assembly (processes driven by minimization of free energy, i.e., no energy is dissipated) [7,8,9,10]. With these terms at hand, we begin by introducing the lowest level of complexity, i.e., monomers and filaments, and proceed to the higher levels, successively describing the physical principles of cells, cell clusters, and tissues.
2 The Cytoskeleton
The cytoskeleton is a scaffold lending cells mechanical integrity and stability. It consists of three main constituents: actin, intermediate filaments (IFs), and microtubules (MTs). These components form fibers in the micrometer range by polymerizing their monomers into specific arrangements, resulting in a different intrinsic filament stiffness for each class [11] (Fig. 5.1). The stiffness is commonly characterized via the so-called persistence length (l p ) [12, 13]. This material-specific parameter is a measure of the fluctuation correlation along the filament backbone, quantifying over which distance an oscillation at a specific point (S 0) at the backbone becomes uncorrelated to the movement of another point (S 2) at the filament (Fig. 5.1). The persistence length can be directly observed, for instance, by analyzing the average transverse fluctuations of filaments observed over time or by evaluating their tangent–cosine correlation function [14]. Based on these methods, actin filaments have an l p of ~10 μm [15, 16], while MTs have a much longer l p in the range of millimeters [17]. Note that l p of natural biopolymers cannot be freely tuned and new model systems have to be used to derive the respective scaling laws [13, 18,19,20]. Since l p is derived via thermal fluctuations (imagine a fluctuating cooked spaghetti), it is a temperature-dependent parameter and cannot be considered a material-defining constant. However, by multiplying thermal energy, k B T, with the temperature, T, and the Boltzmann constant, k B , a new temperature-independent parameter called bending stiffness, κ = k B Tl p , can be derived.
(a) Points (S) along the contour of a semiflexible polymer have different tangent vectors (t). If points are close to each other (S 0 and S 1), their tangent vectors are correlated and roughly point in the same direction. When points are further apart (S 0 and S 2), their tangent vectors are uncorrelated and point in different directions. (b) illustrates the stiffness regimes of the three major cytoskeletal components—microtubules (MTs), actin, and intermediate filaments (IFs). Different mechanical properties are a direct result of the differing filament architectures. l denotes the length of the filament and l p the persistence length [7]
Besides their mechanical properties, which have a static function and serve as the cellular skeleton, the cytoskeletal filaments are also very dynamic structures, enabling rapid adaptive organization of the entire cytoskeleton to fulfill functions such as cell migration or division. Fascinatingly, the cell can use the same components for these somehow contradictory tasks, which is only possible because of permanent energy dissipation, permitting rapid transition between different states. Furthermore, although the cytoskeletal building blocks are preserved in almost every eukaryotic cell, induced morphologies vary substantially among different cell types. Even within a single cell, the cytoskeleton spatially organizes into various different structures responsible for differing sets of functions—a strategy known as multifunctionality.
The different filament architectures not only result in a wide range of different bending rigidities but also determine their role in dynamic processes. MTs, for instance, are very rigid and thus typically appear as individual fibers, extending from the cell interior to the membrane (Fig. 5.2). Due to their outreach and rather straight structures, they are especially well suited for intracellular transport and for providing directed forces during mitosis and for organelle positioning [7]. Actin filaments, on the other hand, are semiflexible polymers and are typically arranged into networks and bundles driving processes such as cell migration. Actin filaments polymerize near the membrane (leading to high local concentrations as illustrated in Fig. 5.2) and effectively push the boundary of the cell forward. In this dynamic process, actin monomers depolymerize at the filament ends pointing toward the cell center. These monomers subsequently travel to the front to reenter the polymerization cycle, a process called treadmilling [7, 21]. Due to their highly dynamic nature, actin filaments can trigger rapid cellular changes. Additional components such as cross-linking proteins or active myosin motors substantially enrich both their mechanical and dynamic phase spaces. It should be noted here that biological force generation is commonly attributed to the activity of molecular motors [11]. However, recent studies have demonstrated that actin as well as MT-based force generation can be driven solely by entropic arguments without requiring any energy dissipation [22,23,24,25,26,27,28].
Schematic drawing of a crawling cell on a 2D substrate showing the most prominent locations for the three types of cytoskeleton biopolymers. MTs are typically nucleated at the centrosome and span the largest portion of the cell. IFs are most commonly found around the cell nucleus, whereas actin filaments form dense networks close to the cell membrane. Particularly, dense and dynamic actin networks are found at the leading edge of migrating cells (forming lamellipodia and filopodia) [7]
In general, actin turnover and interactions with molecular motors are persistent processes in biological matter, resulting in substantial energy consumption. In eukaryotic cells, for instance, actin turnover alone can reach up to ∼50% of the total energy consumption [29, 30], which in turn indicates that minimizing energy consumption has not been the most dominant evolutionary factor. Apart from molecular motors, all other actin accessory proteins influence the filament or network properties without consuming energy in a form of ATP or GTP. Accordingly, regulatory functions can be roughly classified as modifications of either polymerization dynamics, cross-linking, or filament nucleation [7, 21].
IFs are much less studied than the other two main components of the cytoskeleton. Additionally, IFs describe not only one specific polymer but a rather heterogeneous class of biopolymers, which form extended networks and thus substantially contribute to cell mechanics [31, 32]. Different types of IFs performing specific cellular tasks have been identified [33]. However, a general feature of all these cytoskeletal biopolymers is that they undergo growth and shrinkage by addition or subtraction of monomers. Therefore, their length is adjustable in a dynamic fashion and highly depends on stochastic fluctuations [32, 34, 35]. Further, their dynamic organization is largely determined by a complex interplay with a multitude of molecular accessory proteins, which nucleate, sever, cross-link, weaken, strengthen, or transport individual filaments [11]. The dynamic, self-organizing cytoskeleton is powered by energy-dissipating ATP or GTP consumption, mainly fueling two key processes: hydrolysis-powered depolymerization/polymerization of filaments and molecular motor-driven filament/motor transport [7]. Unlike IFs, MTs and actin are polar structures due to their asymmetrical polymerization and depolymerization dynamics (treadmilling) caused by their differing critical concentrations at the two ends. These two different critical concentrations are a direct result of ATP or GTP consumption, thus reflecting the intrinsic non-equilibrium process, which exerts substantial pushing forces [36]. The arising polarity is crucial for molecular motors to be able to move in a specific direction, enabling controlled cargo transport as well as directed pulling forces [37].
3 Rheology
Rheology is the study of deformation responses of materials to applied forces. The deformation response to constantly acting forces depends on whether the material is categorized as a solid or a fluidlike material. In solid materials, the magnitude of the deformation, typically elongation, scales with the applied force, e.g., an elastic spring under tension. Solid responses may also include plastic deformations such as overstretching a spring beyond its elastic limit, which permanently deforms it. The so-called viscous deformation response of a fluidlike material describes how the deformation rate scales with the applied force, e.g., ketchup flowing out of a bottle or squeezing glue out of a tube.
For biological samples—in this chapter single cells and soft tissues—viscoelastic responses to small forces have two distinct time scales: on short time scales, from split seconds to minutes, tissue deformation is proportional to applied forces and will recover to return to its initial form after stress release. This is easily confirmed by pressing against muscular or fatty tissue, where responses are nearly elastic. On long time scales (days to months), tissues tend to behave like highly viscous fluids, enabling body modification such as stretching lips by inserting lip discs, as, for example, practiced by the tribes of Mursi and Surma residing in Ethiopia [38] and the south American peoples of the Kayapo and Botocudo [39], or earspools (“flesh tunnel”) in western subcultures and various African and American tribes [38, 40]. Materials which are governed by both elastic and viscous behaviors, such as most biological tissues, are considered viscoelastic materials. Examples of deformation responses are presented in Fig. 2.5 of Chap. 2.
Besides these passive responses, cells and tissues can actively react to environmental cues. Well-known active force generators are myosin motors, which exert pulling forces on actin filaments. This interaction is crucial on the cellular level for processes such as single-cell motility as well as on the tissue level, e.g., for muscle contraction. Many active processes are complexly intertwined with cellular pathways or immune responses and can highly influence the material properties of cells and tissues.
While the qualitative description of biological materials is straightforward, quantitative descriptions involve a profound theoretical background and mathematical models. The main goal of quantitative description is to gather material parameters from biological tissues. Since material parameters should describe intrinsic properties, they should be—in the best possible case—independent of features of the experimental setup such as applied forces, size of the tissue, or type of experiment conducted. This chapter will outline the theoretical minimum needed to adequately describe single cells and biological tissues later on and will introduce the concepts and terminology needed to understand the following chapters. Special cases such as nonlinearity and temperature dependencies are deliberately neglected here and are partially discussed later.
3.1 Step Experiment
As a starting point, consider a cuboid of tissue. The deformation response will depend on the strength of the force and how it is applied, i.e., on which side and in which direction. Vice versa, if a given deformation is forced upon the material, an internal force will arise accordingly. For the sake of simplicity—mathematical and explanatory—we restrict the possible types of force and deformation application to the types illustrated in Fig. 5.3: a longitudinal sudden force experiment and a transverse shear experiment.
The two archetypes of deformation response experiments in rheology. (a) shows the stretching mode, where two counter-facing sides are pulled apart perpendicular to the surface. The material elongates in the direction of the applied force and contracts perpendicular to it. (b) shows the shear mode. A strain is applied on the upper side, and the strain response is measured on the lower side
In the stretching experiment, the force is applied equally on two counter-facing sides (red-dashed lines), resulting in an applied stress σ (force per area). The material will expand by Δx in the stretching direction and retract by –Δy perpendicular to it. The resulting elongation is measured as strain, i.e., relative extension γ = Δx/x (a tensor notation of linear strain is given in Eq. (2.6) of Chap. 2). Contraction occurs due to internal forces of the material, usually since many materials, such as water, are nearly incompressible. The relation between axial strain and transverse strain is captured in the Poisson ratio ν given by
with volume V of the cuboid and all further parameters as sketched in Fig. 5.3a. The Poisson ratio is a dimensionless unit and typically ranges from 0 to 0.5. A value close to 0 means nearly no lateral contraction upon pulling, while a value close to 0.5 means that the material is nearly incompressible. The Poisson ratio for biological microtissue samples was found to be on the order of ν = 0.45 [41]. Lower values can be found in multiphasic tissues, in which a fluid phase is allowed to freely move (see Chaps. 3 and 4).
For simplicity, only constant step stresses, as illustrated in Fig. 2.5 of Chap. 2, are considered in the following. When applying such a constant stress profile, σ 0, starting at t = 0, strain γ can be expressed as
where t denotes time and D(t) denotes tensile creep compliance. This parameter is usually unique to the type of material measured. For a perfect elastic, springlike material, tensile creep compliance reduces to a constant, D(t) = 1/E, where Young’s modulus E describes the stiffness of a solid material under forces very similar to the spring constant of an elastic spring. The higher it is, the stiffer the material. Muscle tissue, for example, has an average Young’s modulus of 2.12 ± 0.91 kPa [42], while cancellous/trabecular bone has 14.8 ± 1.4 GPa [43, 44]. An overview of elastic properties of tissues has been presented in a review by Akhtar et al. [45].
For a perfect viscous, fluidlike material, the tensile creep compliance will follow D(t) = t/η, where η is the viscosity of the fluid. The higher the viscosity of a material, the slower the flow speed for given forces will be. Honey, for instance, has a viscosity between 2.54 and 23.4 Pa · s (at 25°C, depending on moisture and sugar composition) [46], while blood has 4 mPa · s [47].
For viscoelastic materials, the detailed time-dependent response will be more complicated. Examples for illustration are presented in Fig. 2.5 and Fig. 5.4.
Graph of the strain response of cells (blue, including confidence interval) under a step stress in an optical stretcher. The applied stress is proportional to the laser power (green). After 2 s, the stress is released and the cell relaxes again. A detailed description of the optical stretcher can be found in the next chapter
If a step strain γ 0 is applied, instead of a step stress, the corresponding stress response σ(t) will be given by
with the elastic modulus E(t), also known as time-dependent Young’s modulus, which is in principle the inverted tensile creep compliance. The two material parameters—tensile creep compliance and elastic modulus—are not independent. In fact, one can translate tensile creep compliance to elastic modulus and vice versa. The conversion can be done using Laplace transform ℒ:
with the complex frequency parameter s = σ + iω. Because the interpretation of elastic modulus E(t) is more straightforward—the higher the modulus, the stiffer the material—it is more commonly used in the scientific community than creep compliance.
3.2 Oscillatory Experiment
The second measurement archetype is the shear experiment (Fig. 5.3b), in which a cuboid of material is fixed between two plates. On one plate, a shear stress is applied, and, in the opposite plate, the strain response is measured. Vice versa, applying a strain and measuring a stress response would give the same qualitative result. Commonly, forced strain γ in is a sinusoidal alternation at frequency ω with a chosen maximum strain amplitude γ 0:
The response to stress on the second plate will depend on the material, either elastic, viscous, or viscoelastic. In Fig. 5.5 the three types of responses are illustrated for applying a shear strain and measuring the stress response and vice versa. In either case, the response is also sinusoidal but phase shifted, if the material is not purely elastic. Since cases (a) and (b) in Fig. 5.5 give principally the same results, but with inverted cause and effect, we henceforth focus on case (a), applying strain and measuring stress, as this is the more common experimental method.
Sketches of the shear–strain relation in the shear experiment. In (a), a shear strain is applied (gray), and the stress response is measured, while in (b) a shear stress is applied, and strain is measured. An elastic response (blue and red, respectively) is in phase, while a viscous response is phase shifted by 90° (positive direction in (a), negative direction in (b)). The phase shift will be between 0° and 90° for a viscoelastic response
As the viscoelastic stress response will be sinusoidal with an added phase shift—the strain lags behind the stress—it is possible to use some basic trigonometric identities for dealing with sines and cosines in an elegant way. Nonetheless, the solution for viscoelastic materials can already be explained qualitatively since the response of elastic and viscous materials is already known.
For elastic materials, the resulting stress, σ ela , is proportional to the applied strain, γ in, giving
since γ in is also a cosine function. In contrast, for a viscous material, the strain rate is proportional to stress, meaning that stress will be highest when strain changes the fastest and stress will be zero when strain is constant. Therefore, the viscous stress response will be out of phase by 90°:
Intuitively, the stress response of a viscoelastic material can be found as the sum of the elastic and the viscous response:
Analogous to the tensile creep compliance presented in the previous chapter, we can define the complex shear modulus given by
with storage modulus G ′ and loss modulus G ′′.
A viscoelastic material can therefore be characterized by the ratio of the viscous and the elastic stress response amplitude for a given frequency [13]. The higher the viscous amplitude relative to the elastic amplitude, the more viscous than elastic a material is at that given frequency. The elastic and viscous response can be different for varying shear frequencies (see Fig. 5.6. The ratio of both amplitudes also defines phase shift angle δ, like in Fig. 5.5, given by the following equation (see Eq. (2.44) in Chap. 2):
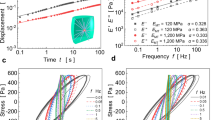
Graph of the elastic and viscous shear modulus (extracted from the complex shear modulus) and phase angle of an actin polymer network measured with a rotation shear rheometer. For low frequencies (10−2 Hz and lower), actin is easier to deform on long time scales of 100 s, which cause a significantly lower storage (blue) and modulus (green). Deformability, both elastic and viscous, increases with frequency. As the phase angle increases with frequency, actin becomes more and more viscous in its response
For the quantitative part, the basic trigonometric identities mentioned above are needed. As the elastic and viscous amplitudes cannot be measured independently in shear experiments, we have to convert G ∗ from a sum of a sine and a cosine to a single cosine (or sine) including phase shift. After conversion, we obtain
where |G ∗(ω)| denotes the measured absolute amplitude given by\( \mid {G}^{\ast}\left(\omega \right)\mid =\sqrt{G^{\prime }{\left(\omega \right)}^2+{G}^{{\prime\prime} }{\left(\omega \right)}^2}. \)The storage and loss modulus can be recovered via G ′(ω) = |G ∗| cos δ and G ′′(ω) = |G ∗| sin δ.
This representation of complex shear modulus G ∗ is most suited for experimental data since it includes absolute amplitude |G ∗(ω)| and phase shift δ, both of which are quantities that are easily measurable in any oscillatory shear experiment for any frequency. Complex shear modulus G ∗(ω) is the favored material parameter in the scientific community since it has a more convenient interpretation, basically the same as elastic modulus E. Also, rotation shear rheometers are predominantly used for oscillatory shear experiments, which can directly measure the storage and loss modulus for a broad frequency range. If instead a shear stress is applied and a strain response is measured (Fig. 5.5a), the calculations above can be done analogously and will result in the complex inverse of G ∗(ω), i.e., complex compliance J ∗(ω) = 1/G ∗(ω). A typical complex shear modulus graph is depicted in Fig. 5.6, showing the frequency-dependent response of an actin network under strain load.
In addition, the previously introduced elastic modulus E(t) and tensile creep compliance D(t) can also be converted to complex shear modulus G ∗(ω). Details of the conversion will be omitted since it involves advanced calculus and will not give more insight into the material properties of biological tissues.
3.3 Modeling Viscoelasticity
Modeling the complex time dependence of tensile creep compliance or the elastic modulus of viscoelastic materials is often done via constitutive equations and/or models. In a simplified, coarse-grained vision, cells can be considered as polymer scaffolds (cytoskeleton) filled with a viscous fluid (cytosol) and functional entities (organelles, which are obstacles in the polymer meshwork), or, put simply, the cytoplasm responds like a water-filled sponge [48]. The cytoskeleton is the main contributor to elastic behavior, while the flow and friction of the cytosol and organelles contribute to the viscous response. The combination of both results in an overall viscoelastic response.
An ideal spring with its elastic response, simulating the cytoskeleton, and an ideal dashpot, simulating the viscosity of the cytosol, can be interconnected to set up toy models simulating viscoelastic responses. We will give here only a short and shallow overview on how viscoelastic properties can arise from the combination of perfectly elastic and viscous subunits. When combining a spring and a dashpot in parallel in the so-called Kelvin–Voigt model, the applied stress is distributed between the spring and the dashpot as captured in the following simple equation:
In addition, the strain of the spring and dashpot will be the same as the total strain:
With this set of equations, the time-dependent strain response γ(t) of the Kelvin–Voigt model (and analogously for the Maxwell model) for any given time-dependent stress σ(t) can be calculated. With the recipe given above, more complex models, possibly featuring more material details for different time scales, can be set up using more than one spring and dashpot. Introduction of another dashpot in series, for instance, accounts for permanent plastic deformation. Furthermore, these models can be applied to any type of experiment—shear and pulling/pushing mode, stress or strain application, and oscillatory and stepping mode—rendering them universally applicable. These models are therefore widely used in the scientific community as a first top–down approach.
Another modeling approach originates from polymer physics allowing to derive many scaling laws from basic principles [12]. Scaling laws are powerful, predictive tools and are generally found in biological systems [49,50,51]. For instance, the basal metabolic rate P of mammals is approximately proportional to their mass M to the power of three fourths (P ∝ M 3/4). For single cells and tissues, scaling laws can be found for the strain response under stress load (γ ∝ σ 0 · t α) [52,53,54]. For a scaling exponent α between 0 and 1, a viscoelastic response can be modeled, including the limit cases of α = 0 and α = 1, corresponding to a purely elastic and purely viscous response, respectively. Indeed, scaling behaviors of material parameters, such as G ∗ and E, can be found in various biological systems [55]. They can be also derived from more fundamental principles of polymer interactions [12] and hold even for advanced theories, i.e., the glassy wormlike chain [56, 57] including nonlinearities like strain hardening and softening [58]. More modeling approaches for cells and tissues can be found in [59, 60].
However, many approaches assume a passive material, which might not be the case for biological matter on time scales of minutes or longer [60]. Introducing active responses, and therefore active force generation, in models is a challenging task since many active processes cannot be described with ease in a coarse-grained manner. Modeling force generation of myosin motors, however, has made significant advances, and appropriate models have been introduced [61,62,63,64,65].
Besides active responses, when probing the mechanical properties of biological matter, the effect of temperature should not be neglected. The temperature should always be in a physiological range since many processes in organisms are highly temperature dependent, e.g., polymerization and depolymerization rates of actin and microtubules [66, 67] as well as motor activity of myosin, dynein, and kinesin [68]. Temperature also affects passive material properties as many materials become less viscous at higher temperatures. Honey, for instance, is much more viscous at lower temperatures [46]. In detail, the viscosity of honey follows an Arrhenius law [69]:
where η(T ref) is the viscosity of the material at a given reference temperature, k B the Boltzmann constant, and E A the activation energy of a transition of states, usually energy barriers of chemical reactions or binding energies. This effect, commonly known as time–temperature superposition, can be observed for single cells [70]. However, many cells do not show this clear relation, and the temperature dependency of their responses is more complex [71].
4 Mechanics on the Cellular Level
Modeling is often limited by strong interactions across multiple levels of complexity. Already on a cellular level, the many cell organelles and functional groups make it difficult to grasp the cell “as a whole” in terms of a coarse-grained system. As the internal structures of cells are already highly anisotropically distributed, it might appear counterintuitive to describe the behavior of a whole cell as a coarse-grained system. It turns out, however, that many of the introduced concepts are no oversimplifications per se and that biological systems can often be described in such a simplified manner. As long as cells (and/or tissues) are not actively reacting to applied forces and deformations, cell mechanics can be understood as an emergent consequence of the cytoskeletal network level. To push this conceptual approach even further, key aspects of cell migration [72] or cell shape [73] can be described without considering details of the filamentous or the molecular level by solely using very fundamental hydrodynamics-based descriptions [74].
Biological cells can be structurally regarded as polymer-filled entities enclosed by a nearly impenetrable membrane. Due to the considerable backbone stiffness of actin filaments and microtubules, mechanical integrity is already given for relatively large mesh sizes and low-volume fraction. An analogy for this concept is a tent, which mechanically stabilizes a certain volume with enough space for passive and active molecular transport. As described in the cytoskeleton chapter, the three main components, actin, microtubules, and intermediate filaments, can form emergent structures such as networks and bundles [7, 75]. Although cells come in a broad phenotypic diversity, including keratinocytes, fibroblasts, and neurons, the cytoskeletal composition does not necessarily need a thorough overhaul. Usually, slight compositional variation or introduction of active processes, i.e., molecular motors, is often sufficient to generate this rich pool of structural appearances. Additionally, many other cellular components contribute to the mechanical behavior.
The main actors in this scenario are the nucleus, the cytoskeleton, and the cell membrane. While these structures are functionally and mechanically intertwined, effects can be specifically attributed to certain subcellular structures. Whole-cell deformations like squeezing or stretching of the cell body are mostly affected by the cytoskeleton as it is the most extensive structure in the cell. Small deformations in the linear regime (up to 5% [58]) are dominated by actin and intermediate filaments [76]. For larger deformations between 5 and 25% strains, nonlinear effects of the actin cytoskeleton can result in nonlinear responses, e.g., strain stiffening [77,78,79] and—paradoxically—also strain softening [80, 81] (the paradox is resolved in [58]). Even larger elongations are intercepted by microtubules and ultimately limited by the integrity of the membrane. Lamellipodia and other protrusions also rely on the stabilization by cytoskeletal filaments. Nuclear mechanics come into play when cells are moving in narrow spaces or are heavily compressed [82, 83]. Passing of narrow channels is dependent on the viscous response of the nucleus to such confinement [83,84,85], and the nuclear lamina will actively respond to environmental stiffness [86, 87]. The mechanical response of cells is also influenced by their membrane, and diseases such as cancer can often cause membrane alterations [88, 89]. Small indentations (tenth of cell diameter) and pulling forces (sub-nano-Newton) are a matter of membrane bending rigidity, while global (and quasi-global) indentations are influenced by effective membrane tension [89, 90]. Furthermore, membrane rigidity influences the extent of mechanosensing signaling pathways. This also holds for self-induced invaginations and blebbing, i.e., exo- and endocytosis, receptor binding, and treadmilling. When discussing processes beyond these circumstances and on the scale of a whole cell, the entanglement of these differently behaving structures has to be taken into account. Still, the cytoskeleton can be considered the most influential part of the cell regarding mechanical behavior.
The strong influence of the cytoskeleton on the overall mechanical properties is striking when looking at the elastic modulus E of most cells. It ranges from unexpected low values [52] of some hundred Pa in glial or neuronal cells [91] to tens of kPa in human thrombocytes [92], illustrating the high variability and adaptivity of cells compared to classical and synthetic materials. As parts of the cytoskeleton are in constant treadmilling, appearance and mechanical structure are not persistent and allow the cell to adapt to its environment, rendering the cytoskeleton self-regulatory.
From the broad range of elastic moduli of different cell types and completely different functions, it is apparent that cell mechanics is a vital component of cellular functioning [93,94,95,96,97] including mitosis, where the cytoskeleton undergoes a significant overhaul enabling controlled cell division [98].
4.1 Probing Techniques
Cell rheology probes the response of cells to applied disturbances. As already explained in detail in the rheology chapter, responses and disturbances are usually forces and deformation or vice versa. Since material parameters like the complex shear modulus are in principle independent of the probing technique, a variety of techniques have been established based on different physical concepts to generate stresses or strains. Table 5.1 summarizes the most common probing techniques for single cells. Another reason for this variety is that every technique has its own working range of stresses and strains and temporal resolution and probes a cell either locally or globally. Nonetheless, due to the high structural heterogeneity of single cells including local mechanical alterations, it has turned out that directly comparable, consistent results are difficult to obtain. One eminent question is what exactly is probed since the main components of the cytoskeleton already differ in their mechanical properties. Also, adherent cells, which form prominent stress fibers on substrates [99], differ in their responses from suspended cells, in which actin conglomerates to a shell-like cortex below the membrane [100, 101]. It remains an open question whether (and how) results obtained for adhered and suspended cells compare.
The comparably simple and inexpensive micropipettes were one of the first tools for characterizing cell mechanics via micropipette aspiration [117], albeit it is limited by inherently lower throughput due to long preparation and measurement times. In the earliest application of this method, blood cells with different diameters were used and analyzed with regard to their response to higher or lower suction pressure (=stress). In general, any suspended cell can be probed including isolated cells from tissues [103]. If a very small pipette diameter is chosen, the local mechanical properties can be probed, whereas larger pipettes can be used to suck in cells for global probing on time scales from seconds to hours [102], and deformations far from the linear regime can be obtained.
Another well-established method to determine mechanical properties of cells, which can be also extended to small tissue samples, is atomic (scanning) force microscope (AFM). Depending on the geometry of the cantilever, cells can be probed very locally by using a pointy tip [93], broadly locally by using a beaded tip [91, 104, 118], or globally by using a flat tip [119]. In terms of stress and strain application, the AFM is versatile, allowing indention times from milliseconds to minutes, only limited by drifting of experimental stages (which can be stabilized with slight sophistication [120]). The force application ranges from pN to nN, including forces beyond the linear limit [121, 122]. Furthermore, using an oscillating cantilever to induce oscillatory stresses allows complex shear modulus measurements [41, 105]. One drawback of the AFM is the comparably low throughput due to possible long preparation times of the experiments, and since adherent cells are very flat, substrate stiffness and roughness affect the results and have to be considered [104]. Stretching of cells can also be done by letting the cell adhere to the cantilever first and subsequently pulling it away [123].
To circumvent the bottleneck of low throughput, further techniques make use of parallel preparation of cells by incorporating tracing beads and using passive Brownian motion (passive microrheology) or probing beads and actively displacing them (active microrheology) [124, 125]. The established method of analysis is based on cross-correlation of the motion of different beads and correction for local heterogeneities [109, 126]. Since artificial beads may be invasive, naturally present “tracers” such as storage granules, mitochondria, and other submicron particles were used with success [108, 127].
With the small size of the beads relative to cell size, both active and passive microrheologies are best suited to probe local rather than global mechanical properties. As microrheology is a contact method, it is highly dependent on the type and strength of linkage between the beads and the surrounding cellular structures [124, 128] and, due to the heterogeneity of cells, a controlled binding affinity is yet to be achieved [124].
The most common active microrheological method is magnetic twisting cytometry, which involves manipulation (usually twisting) of magnetic beads with an external magnetic field [54, 110, 129]. Since oscillating magnetic fields can be easily generated, many cells can be probed in parallel at once across a range of over four decades of frequencies. Limiting factors degrading measurement accuracy, however, include the exact determination of the bead’s magnetic moment along with bead-to-bead variation and the applied external magnetic field.
All probing techniques discussed so far are contact based and are therefore prone to be invasive and might measure cell mechanics in an altered state. Furthermore, for all techniques, cells have to be at least weakly adherent, introducing additional problems due to substrate influences even for techniques based on optically trapped probing beads [106]. These limitations and difficulties can be overcome by using optical manipulation and microfluidic techniques to measure single suspended cells.
For optical manipulation, the optical stretcher has been established. This technique is based on the momentum transfer of photons on interfaces with changing refractive indices. Two antiparallel laser fibers with divergent beam profiles can generate an optical pressure force [130] enabling optical trapping at lower laser powers and deformation of cells at higher powers [101]. The optical stretcher can apply step stresses over a time-range from 0.1 s to tens of seconds, enabling creep compliance measurements [111, 112]. Applied stresses are in the Pa range, corresponding to sub-nano-Newton forces, which depend on fiber-to-fiber distance and cell size. Cell mechanics can be probed locally for large cells and small fiber distances or globally for small cells and increased fiber distances. Since cells can be optically trapped for a prolonged period of time, active responses of single cells can be observed, for instance, active contractions of cells under force load [131]. Oscillatory stress application is also possible, however, with the restriction that only positive stresses (stretching of cells) and no negative stresses (squeezing of cells) can be applied [132, 133]. Therefore, the stress pattern will include an offset stress: σ(t) = σ 0 + σ 0 sin(ωt). Embedded in a microfluidic setup, the optical stretcher allows serial measurements of up to 300 cells per hour and subsequent sorting, which can be challenged by the global heterogeneity of cell stiffness [97, 101].
Related techniques are able to deform cells in a similar contact-free manner but are based on hydrodynamic instead of optical forces [116]. Here, cells are pushed through capillaries in a continuous flow. Sudden changes in capillary geometry alter the flow profile locally. Shear flow velocities are applied that are sufficient to generate force differences large enough to deform whole-cell bodies. With these techniques, immense throughputs can be achieved [114,115,116]. However, the very limited observation time for one cell (millisecond range) impedes long-term deformation measurement, reducing measurable cell mechanics to the relative deformation of the cells after entering the measurement channel, i.e., the (time-independent) elastic modulus E.
Further details and comparisons of the different commonly applied probing techniques can be found in [52, 55, 60, 121, 134, 135].
4.2 Comparability and Interpretation
The broad range of experimental techniques and their intrinsic advantages and disadvantages make it challenging to compare results obtained with different techniques, especially quantitative results. Responses of suspended and adherent cells (as well as resting and migrating) will inherently differ from each other due to their altered geometries. Furthermore, probing a cell locally might not yield the same results as probing it globally. Even focusing on a single technique, defining the mechanical properties of a certain cell type is already nontrivial as cell-type stiffness follows a broad, non-Gaussian distribution (which can be tackled by averaging over many cells in large cell monolayer shear measurements [136]).
Despite the given quantitative challenges, the broad range of experimental techniques has yielded a comprehensive qualitative picture of cell mechanics covering various orders of stress and strain regimes [52, 124]. For instance, measurements with different techniques show a common power-law behavior of the complex shear modulus with only a slightly varying power-law exponent [107, 124, 134, 137, 138].
Nonetheless, in order to compare experimental techniques, one has to overcome some drawbacks of these techniques, which often involve poor statistics and the lack of standardized measurement protocols. For some of the methods presented here, it is difficult to obtain enough data during the short time in which the mechanical properties are altered actively in response to environmental changes. On the other hand, the diversity of probing techniques is also an advantage. As different cell types occur in different environments, cell mechanics differ to suit their environment, e.g., red blood cells are more elastic since they have to squeeze through capillaries [11]. Thus, the most suitable technique for a given cell type can be chosen. Suspended cells like RBCs, for instance, can be measured more easily and rapidly in an optical stretcher or real-time deformation cytometer [83, 112, 115, 139].
5 Tissues
The mechanics of systems consisting of multiple cells are changed drastically by two elements that are not present at the single-cell level: adhesive contacts between cells and between cells and the extracellular matrix (ECM), enabling active responses of cells to their neighbors. Cell–cell communication via very basic interactions, such as chemical feedback loops, can already lead to collective behavior of cells, e.g., swarm-like collective migration of keratinocytes [140] and collective cell migration during invasion and metastatic spread of malignant tumors [141]. Furthermore, strength and type of adhesive contacts direct force transmission and modulate cell migration and motility. As a consequence, cells can show collective structural behavior on different length scales ranging from organization of smaller subdomains in tissues with individual properties up to global quasi-frozen, glass-like states with no (relative) cell migration as in epithelial layers [142], which commonly occurs as soon as increased adhesion force and cell density are introduced. Since active processes of tissues are emergent phenomena, they can strongly influence mechanical responses. Adhesive cell–cell contacts and ECM can highly contribute to the stiffness and fluidity of tissues and span a phase space ranging from quasi-solid responses, e.g., epithelial tissue, to quasi-fluid materials such as migratory cells in mesenchymal tissues.
5.1 Cell–Cell and Cell–Matrix Adhesion
Most types of cell adhesion are mediated by proteins of the family of cell adhesion molecules (CAMs). CAMs are typically transmembrane proteins with binding sites for the cytoskeleton as well as sites for trans- and cis-interactions with other CAMs or the ECM in the extracellular domain. In addition to adhesion, they function as cytoskeletal anchors and play significant roles in mechanosignaling [11], which elegantly illustrates the intertwined nature of the different levels of complexity.
CAMs are often calcium dependent, and one of the most prominent CAM families is cadherins (a portmanteau word combining calcium and adhering). Cadherins come in three flavors and are usually associated with certain tissues (but are not restricted to these): E-cad, epithelial cells; N-cad, neuronal cells, and P-cad, pancreatic cells. They can appear as single free molecules but are usually ordered as nanoclusters linked to actin or as more complex desmosomes linked to the keratin cytoskeleton [143]. Differences in the function of these proteins are still under investigation, but changes in their expression rate can be correlated to changes of phenotype and behavior. In epithelial-to-mesenchymal transition (EMT), for example, cells lose their well-structured belt-like distribution of E-cadherins in favor of P- and N-cadherins [144]. At the same time, cells will regain their ability to move through tissue, a hallmark step in tumorigenesis and metastasis [145, 146]. In malignant tumors, this switch can promote directed invasion into the surrounding tissue of small cell clusters, often regulated by messenger RNA (mRNA). For instance, upregulation of miR-9, a short, noncoding RNA gene involved in gene regulation, leads to increased cell motility and invasiveness [147]. The increased expression of P-cad triggers polarization and directed movement [148]. This results in cells moving in single file (Indian filing) out of the tumor, with a tumor cell or fibroblast as the leading cell [149]. The polarization of the fibroblast that moves away from the cancer cells is mediated by N-cad adhesive sites [150].
Contact to the ECM is mainly established by integrins, which consist of two subunits, the α- and β-units, and can bind collagen, glycoproteins of the ECM (e.g., fibronectin), or both. There are several types of α- and β-units and consequentially a broad range of different integrins. Like cadherins, they cluster into functional domains by focal adhesions. Binding of integrins to extracellular structures induces signaling cascades that intervene with basic cellular functions such as cell growth and apoptosis. Depending on the range of expression of different integrins, and subsequently the composition of focal adhesions, signaling pathways can be promoted or suppressed. Integrins such as α v β 3 or α 5 β 1, for instance, are often found in cancer cells and seem play a role in cancer development. They influence the mechanical behavior [151], invasiveness in ECM-rich surroundings [152], and cellular survival [153]. In addition, some integrins are known to form complexes with growth factors that induce EMT [154] and increase proliferation [155].
The mechanical feedback of cell–cell and cell–ECM interactions is a fundamental parameter for cellular regulation and shows its drastic influence in diseases such as cancer [156]. Cells can modify their morphology and mechanical properties in response to a changing microenvironment [99]. Especially since cells can and do modulate their ECM and CAMs, they can be a strong promoter of metastasis formation [157]. This active reaction of cells to their microenvironment becomes stronger with increasing malignancy, e.g., metastatic cells can mimic mechanical properties of neuronal cells [158] by reactivating (epigenetically) silenced genes [159,160,161].
5.2 Tissue Dynamics and Collective Cell Behavior
It is a great challenge to define simple mechanical parameters for tissues. At small time scales and low strains, tissues show frequency-dependent complex shear moduli [162]. At larger time scales, tissues can lose their viscoelastic behavior and show fluidlike mechanics—usually coupled to movement of single cells in the tissue and in a regime far from equilibrium dynamics. The two established main models describe tissues as glass-like, amorphous materials or as yield stress fluids. With both models, the active contribution of cells in the system is of interest. Softness of the cell body, adhesion strength to neighboring cells, and matrix as well as active forces determine whether a cell is able to migrate or not [163].
Analogies to glassy materials can be found in liquid- to solid-like glass transition. When the temperature of a glassy material drops below the glass transition point, molecules are strongly confined in their motion and fixed in a chaotic lattice. A cell system that reaches a certain density due to proliferation will exhibit similar behavior. Additionally, density-independent transitions can be observed when adhesion and stiffness of cells are modulated [164]. Inhibited migration and proliferation mark the point at which the system goes into a static (glassy) state [165, 166], a concept which can be transferred to tissues under high stresses. Epithelial layers, for example, have been shown to exert constant pulling forces that can influence their behavior [167].
Yield stress fluids show viscoelastic behavior to the point where a certain energy barrier has been overcome, when the material will start to flow. In this model, single-cell mechanics and stresses acting on the tissue are the main contributors [168]. Stress on the tissue, intrinsic or extrinsic, interacts with local adhesive mechanics, giving rise to either fluidlike or solid-like behavior. Especially homeostatic stresses determine the flow of the tissue, and a tissue with higher homeostatic stress will invade surrounding tissues either as small, separate islands or as a front [169] (Fig. 5.7A1, A2).
In both cases, transitions can occur for the whole tissue at once, cell clusters in confinement, or for single cells within a tissue. Even when a tissue is above the transition point and remains static, some cells might be in a different state and still able to pass through it (Fig. 5.7).
Wound-healing assay of two different cell types. In the upper panel, A1 and A2, an epithelial cell layer starts at a certain time (A1) and closes the wound after 30 h (A2). The epithelial cell layer maintains its cell front and shows a coordinated, collective motion. A mesenchymal cell layer, B1 and B2, loses its front (B1) in this process and shows randomly walking single cells and no coordinated, collective motion after 30 h (B2)
A rheological approach to access the different states of tissues is measuring phase angle δ. Although this approach does not account for active cell migration, it allows estimating the fluidity of the tissue. δ near 0° (purely elastic) indicates a jammed state, while viscous, flowing tissue approaches 90°. To probe local properties of tissues, scanning force microscopy can be employed to create a map of viscoelasticity of a tissue slice that can be attributed to processes in the sample [170]. With standard bulk shear rheometers, global measurements can be performed to determine the overall viscoelastic properties [171]. Methods such as magnetic resonance elastography enable direct measurement of complex shear moduli of a whole tissue or with spatial resolution [162, 172].
When parameters of single cells, such as adhesive and viscoelastic properties, are known, these can be used to draw conclusions regarding tissue dynamics. The differential adhesion hypothesis (DAH) states that a mixture of cells with different adhesiveness develops into an ordered state, separating subpopulations accordingly [173]. The DAH holds true in morphogenesis, where cells demix like fluids with different surface tensions [174], but fails for cancer development. When cells undergo EMT, demixing can be observed, which seems to follow a more complex behavior [175]. Local jamming, unjamming, and tumor cell invasion are strongly related to these observations. When a malignant neoplasm forms, it constitutes a clearly separated bulk of tissue within the healthy stroma. Cells in the tumor exhibit altered mechanical behavior and heavily remodel the ECM, but there is no obvious reason why a distinct boundary exists. As soon as the cells lose their epithelial phenotype, they show increased invasion [176]. Still, the primary tumor grows to a certain size before cells escape, which might have its origins in cellular jamming. Fibrosis, for instance, leads to a very stiff ECM [177], and the growth of the tumor creates pressure on the surrounding tissue and the tumor itself [178]. In other words, the tumor embeds itself in a strong matrix. This goes hand in hand with a tumor being a rigid mass, although single malignant cells tend to be softer when becoming more invasive [179]. While the self-driven confinement creates a jammed state within the tumor [180], the mechanical feedback leads to further transitioning toward more malignant phenotypes [181]. Over time, the distribution of cellular softness becomes broader [97], and expression of CAMs is altered [144] up to the point where some cells undergo an unjamming transition and start to move out of the tumor [182]. It has to be noted that this cellular escape does not occur in a random pattern. Similar to embryogenesis, where cells follow the DAH, self-organization within the tissue is the first step. Collective migration in 2D assays can be observed, when the confluency of the layer confines cells, before the layer becomes jammed [183] (Fig. 5.7 A2). In 3D, this cellular streaming has also been observed: small conglomerates of cells in an unjammed state form and migrate collectively, often following a leader cell [184]. This marks the start of metastatic spread, and the moment at which invasion begins heavily depends on the individual neoplasm. Some tumors will grow to immense sizes over months or even years before cells pass the boundaries, while others metastasize within weeks of the original tumor formation.
Conclusion
The eukaryotic cell is well studied with decades of research dedicated to various branches and aspects ranging from classification of whole-cell types down to molecular details of protein folding processes [11, 185]. With the advancement of techniques and detailed insights into biological matter, the cause and effect of many diseases could be attributed to certain functional or structural units and levels of complexity within the cell; for instance, sickle cell disease is often caused by only a single-nucleotide polymorphism (SNP) of the beta-globin gene, which results in strand-like clustering of defective hemoglobin and consequently stiffening of red blood cells [186, 187]. Especially cancer—as one of the most prominent maladies—is well studied on many levels of complexity [188], and crucial developmental steps and many biological changes in tumorigenesis were found and described [145, 146]. The development of cancer is accompanied by major changes of the cytoskeleton, which are necessary for cancer cells to migrate and invade other tissues [97, 189]. The cytoskeleton usually gets softer with increased malignancy, as shown with different cellular probing techniques [115, 116, 179, 188,189,190,191,192,193,194]. These important insights render rheological material characterization of a viable tumor marker. At the tissue level, however, tumors are found to be stiffer than healthy surrounding tissue due to a stiffer stroma and elevated cytoskeletal tension [177] although they are constituted of softer cells. While many emergent phenomena on the microtissue level will influence mechanical properties, many of them are physically characterized and quantified. While the biophysics of tumorigenesis and tissue mechanics is qualitatively well studied. Still, the quantitative description of demixing, jamming, and surface tension is under current investigation and remains promising with ongoing research on this frontier of science [163, 164, 183, 195,196,197,198,199,200,201].
References
Anderson PW. More is different. Science. 1972;177(4047):393–6. https://doi.org/10.1126/science.177.4047.393.
Laughlin RB, Pines D. The theory of everything. Proc Natl Acad Sci U S A. 2000;97(1):28–31. https://doi.org/10.1073/pnas.97.1.28.
Schrödinger E. What is life?: the physical aspect of the living cell, Canto. Cambridge: Cambridge University Press; 2010.
Ryan AJ. Emergence is coupled to scope, not level. Complexity. 2007;13(2):67–77. https://doi.org/10.1002/cplx.20203.
Dawkins R The blind watchmaker: why the evidence of evolution reveals a universe without design. New edition [reissue] ed. New York: W. W. Norton; 1996.
Schuster P. A beginning of the end of the holism versus reductionism debate?: molecular biology goes cellular and organismic. Complexity. 2007;13(1):10–3. https://doi.org/10.1002/cplx.20193.
Huber F, Schnauss J, Ronicke S, et al. Emergent complexity of the cytoskeleton: from single filaments to tissue. Adv Phys. 2013;62(1):1–112. https://doi.org/10.1080/00018732.2013.771509.
Huber F, Kas J. Self-regulative organization of the cytoskeleton. Cytoskeleton. 2011;68(5):259–65. https://doi.org/10.1002/cm.20509.
Halley JD, Winkler DA. Classification of emergence and its relation to self-organization. Complexity. 2008;13(5):10–5. https://doi.org/10.1002/cplx.20216.
Halley JD, Winkler DA. Consistent concepts of self-organization and self-assembly. Complexity. 2008;14(2):10–7. https://doi.org/10.1002/cplx.20235.
Alberts B. Molecular biology of the cell: [MBOC]. 6th ed. New York: GS Garland Science; 2015.
Doi M, Edwards SF. The theory of polymer dynamics, The international series of monographs on physics, vol. 73. Oxford: Clarendon Press; 2003.
Schuldt C, Schnauß J, Händler T, et al. Tuning synthetic semiflexible networks by bending stiffness. Phys Rev Lett. 2016;117(19):197801. https://doi.org/10.1103/PhysRevLett.117.197801.
Isambert H, Venier P, Maggs A, et al. Flexibility of actin filaments derived from thermal fluctuations. Effect of bound nucleotide, phalloidin, and muscle regulatory proteins. J Biol Chem. 1995;270(19):11437–44. https://doi.org/10.1074/jbc.270.19.11437.
Isambert H, Maggs AC. Dynamics and rheology of actin solutions. Macromolecules. 1996;29(3):1036–40. https://doi.org/10.1021/ma946418x.
Greenberg MJ, Wang C-LA, Lehman W, et al. Modulation of actin mechanics by caldesmon and tropomyosin. Cell Motil Cytoskeleton. 2008;65(2):156–64. https://doi.org/10.1002/cm.20251.
Janson ME, Dogterom M. A bending mode analysis for growing microtubules: evidence for a velocity-dependent rigidity. Biophys J. 2004;87(4):2723–36. https://doi.org/10.1529/biophysj.103.038877.
Yin P, Hariadi RF, Sahu S, et al. Programming DNA tube circumferences. Science. 2008;321(5890):824–6. https://doi.org/10.1126/science.1157312.
Schiffels D, Liedl T, Fygenson DK. Nanoscale structure and microscale stiffness of DNA nanotubes. ACS Nano. 2013;7(8):6700–10. https://doi.org/10.1021/nn401362p.
Glaser M, Schnauß J, Tschirner T, et al. Self-assembly of hierarchically ordered structures in DNA nanotube systems. New J Phys. 2016;18(5):55001. https://doi.org/10.1088/1367-2630/18/5/055001.
Pollard TD, Blanchoin L, Mullins RD. Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu Rev Biophys Biomol Struct. 2000;29:545–76. https://doi.org/10.1146/annurev.biophys.29.1.545.
Schnauß J, Händler T, Käs J. Semiflexible biopolymers in bundled arrangements. Polymers. 2016;8(8):274. https://doi.org/10.3390/polym8080274.
Schnauss J, Golde T, Schuldt C, et al. Transition from a linear to a harmonic potential in collective dynamics of a multifilament actin bundle. Phys Rev Lett. 2016;116(10):108102. https://doi.org/10.1103/PhysRevLett.116.108102.
Lansky Z, Braun M, Ludecke A, et al. Diffusible crosslinkers generate directed forces in microtubule networks. Cell. 2015;160(6):1159–68. https://doi.org/10.1016/j.cell.2015.01.051.
Braun M, Lansky Z, Hilitski F, et al. Entropic forces drive contraction of cytoskeletal networks. BioEssays. 2016;38(5):474–81. https://doi.org/10.1002/bies.201500183.
Ward A, Hilitski F, Schwenger W, et al. Solid friction between soft filaments. Nat Mater. 2015;14(6):583–8. https://doi.org/10.1038/nmat4222.
Hilitski F, Ward AR, Cajamarca L, et al. Measuring cohesion between macromolecular filaments one pair at a time: depletion-induced microtubule bundling. Phys Rev Lett. 2015;114(13):138102. https://doi.org/10.1103/PhysRevLett.114.138102.
Huber F, Strehle D, Schnauß J, et al. Formation of regularly spaced networks as a general feature of actin bundle condensation by entropic forces. New J Phys. 2015;17(4):43029. https://doi.org/10.1088/1367-2630/17/4/043029.
Daniel JL, Molish IR, Robkin L, et al. Nucleotide exchange between cytosolic ATP and F-actin-bound ADP may be a major energy-utilizing process in unstimulated platelets. Eur J Biochem. 1986;156(3):677–83. https://doi.org/10.1111/j.1432-1033.1986.tb09631.x.
Bernstein BW, Bamburg JR. Actin-ATP hydrolysis is a major energy drain for neurons. J Neurosci. 2003;23(1):1–6.
Block J, Schroeder V, Pawelzyk P, et al. Physical properties of cytoplasmic intermediate filaments. Biochim Biophys Acta. 2015;1853(11 Pt B):3053–64. https://doi.org/10.1016/j.bbamcr.2015.05.009.
Herrmann H, Bar H, Kreplak L, et al. Intermediate filaments: from cell architecture to nanomechanics. Nat Rev Mol Cell Biol. 2007;8(7):562–73. https://doi.org/10.1038/nrm2197.
Herrmann H, Aebi U. Intermediate filaments: molecular structure, assembly mechanism, and integration into functionally distinct intracellular scaffolds. Annu Rev Biochem. 2004;73:749–89. https://doi.org/10.1146/annurev.biochem.73.011303.073823.
Howard J, Hyman AA. Dynamics and mechanics of the microtubule plus end. Nature. 2003;422(6933):753–8. https://doi.org/10.1038/nature01600.
Vavylonis D, Yang Q, O’Shaughnessy B. Actin polymerization kinetics, cap structure, and fluctuations. Proc Natl Acad Sci U S A. 2005;102(24):8543–8. https://doi.org/10.1073/pnas.0501435102.
Footer MJ, Kerssemakers JWJ, Theriot JA, et al. Direct measurement of force generation by actin filament polymerization using an optical trap. Proc Natl Acad Sci U S A. 2007;104(7):2181–6. https://doi.org/10.1073/pnas.0607052104.
Howard J. Mechanics of motor proteins and the cytoskeleton. Sunderland: Sinauer Associates; 2006.
Lockot HW, Uhlig S. Bibliographia aethiopica, Aethiopistische forschungen, vol. 41. Wiesbaden: Steiner; 1998.
Bell A, Macfarquhar C. Encyclopaedia britannica: or, a dictionary of arts and sciences, three volumes. Scotland: Edinburgh; 1771.
Borel F, Taylor JB, Paris IM. The splendor of ethnic jewelry: from the Colette and Jean-Pierre Ghysels collection, Pbk. ed. New York: H. N. Abrams; 2001
Mahaffy RE, Shih CK, MacKintosh FC, et al. Scanning probe-based frequency-dependent microrheology of polymer gels and biological cells. Phys Rev Lett. 2000;85(4):880–3. https://doi.org/10.1103/PhysRevLett.85.880.
Chen EJ, Novakofski J, Jenkins WK, et al. Young’s modulus measurements of soft tissues with application to elasticity imaging. IEEE Trans Ultrason Ferroelect Freq Contr. 1996;43(1):191–4. https://doi.org/10.1109/58.484478.
Rho J-Y, Kuhn-Spearing L, Zioupos P. Mechanical properties and the hierarchical structure of bone. Med Eng Phys. 1998;20(2):92–102. https://doi.org/10.1016/S1350-4533(98)00007-1.
Rho JY, Ashman RB, Turner CH. Young’s modulus of trabecular and cortical bone material: ultrasonic and microtensile measurements. J Biomech. 1993;26(2):111–9. https://doi.org/10.1016/0021-9290(93)90042-D.
Akhtar R, Sherratt MJ, Cruickshank JK, et al. Characterizing the elastic properties of tissues. Mater Today. 2011;14(3):96–105. https://doi.org/10.1016/S1369-7021(11)70059-1.
Yanniotis S, Skaltsi S, Karaburnioti S. Effect of moisture content on the viscosity of honey at different temperatures. J Food Eng. 2006;72(4):372–7. https://doi.org/10.1016/j.jfoodeng.2004.12.017.
Kesmarky G, Kenyeres P, Rabai M, et al. Plasma viscosity: a forgotten variable. Clin Hemorheol Microcirc. 2008;39(1-4):243–6.
Zhou EH, Martinez FD, Fredberg JJ. Cell rheology: mush rather than machine. Nat Mater. 2013;12(3):184. https://doi.org/10.1038/nmat3574.
Spence AJ. Scaling in biology. Curr Biol. 2009;19(2):R57–61. https://doi.org/10.1016/j.cub.2008.10.042.
Brown JH, West GB, editors. Scaling in biology. Santa Fe Institute studies in the sciences of complexity. Oxford: Oxford University Press; 2000.
West GB. A general model for the origin of allometric scaling laws in biology. Science. 1997;276(5309):122–6. https://doi.org/10.1126/science.276.5309.122.
Kollmannsberger P, Fabry B. Linear and nonlinear rheology of living cells. Annu Rev Mater Res. 2011;41(1):75–97. https://doi.org/10.1146/annurev-matsci-062910-100351.
Sandersius SA, Newman TJ. Modeling cell rheology with the subcellular element model. Phys Biol. 2008;5(1):15002. https://doi.org/10.1088/1478-3975/5/1/015002.
Fabry B, Maksym GN, Butler JP, et al. Scaling the microrheology of living cells. Phys Rev Lett. 2001;87(14):148102. https://doi.org/10.1103/PhysRevLett.87.148102.
Chen DT, Wen Q, Janmey PA, et al. Rheology of Soft materials. Annu Rev Condens Matter Phys. 2010;1(1):301–22. https://doi.org/10.1146/annurev-conmatphys-070909-104120.
Kroy K. Dynamics of wormlike and glassy wormlike chains. Soft Matter. 2008;4(12):2323. https://doi.org/10.1039/B807018K.
Kroy K, Glaser J. The glassy wormlike chain. New J Phys. 2007;9(11):416. https://doi.org/10.1088/1367-2630/9/11/416.
Wolff L, Fernandez P, Kroy K. Resolving the stiffening-softening paradox in cell mechanics. PLoS One. 2012;7(7):e40063. https://doi.org/10.1371/journal.pone.0040063.
Rodriguez ML, McGarry PJ, Sniadecki NJ. Review on cell mechanics: experimental and modeling approaches. Appl Mech Rev. 2013;65(6):60801. https://doi.org/10.1115/1.4025355.
Lim CT, Zhou EH, Quek ST. Mechanical models for living cells—a review. J Biomech. 2006;39(2):195–216. https://doi.org/10.1016/j.jbiomech.2004.12.008.
Herant M, Marganski WA, Dembo M. The mechanics of neutrophils: synthetic modeling of three experiments. Biophys J. 2003;84(5):3389–413. https://doi.org/10.1016/S0006-3495(03)70062-9.
Dai J, Ting-Beall HP, Hochmuth RM, et al. Myosin I contributes to the generation of resting cortical tension. Biophys J. 1999;77(2):1168–76.
Peskin CS, Odell GM, Oster GF. Cellular motions and thermal fluctuations: the Brownian ratchet. Biophys J. 1993;65(1):316–24. https://doi.org/10.1016/S0006-3495(93)81035-X.
Mogilner A, Oster G. Cell motility driven by actin polymerization. Biophys J. 1996;71(6):3030–45. https://doi.org/10.1016/S0006-3495(96)79496-1.
Kuusela E, Alt W. Continuum model of cell adhesion and migration. J Math Biol. 2009;58(1-2):135–61. https://doi.org/10.1007/s00285-008-0179-x.
Zimmerle CT, Frieden C. Effect of temperature on the mechanism of actin polymerization. Biochemistry. 1986;25(21):6432–8.
Kis A, Kasas S, Kulik AJ, et al. Temperature-dependent elasticity of microtubules. Langmuir. 2008;24(12):6176–81. https://doi.org/10.1021/la800438q.
Yengo CM, Takagi Y, Sellers JR. Temperature dependent measurements reveal similarities between muscle and non-muscle myosin motility. J Muscle Res Cell Motil. 2012;33(6):385–94. https://doi.org/10.1007/s10974-012-9316-7.
Oroian M, Amariei S, Escriche I, et al. A viscoelastic model for honeys using the time–temperature superposition principle (TTSP). Food Bioprocess Technol. 2013;6(9):2251–60. https://doi.org/10.1007/s11947-012-0893-7.
Kießling TR, Stange R, Käs JA, et al. Thermorheology of living cells—impact of temperature variations on cell mechanics. New J Phys. 2013;15(4):45026. https://doi.org/10.1088/1367-2630/15/4/045026.
Schmidt BUS, Kießling TR, Warmt E, et al. Complex thermorheology of living cells. New J Phys. 2015;17(7):73010. https://doi.org/10.1088/1367-2630/17/7/073010.
Joanny J, Prost J. Active gels as a description of the actin-myosin cytoskeleton. HFSP J. 2009;3(2):94–104. https://doi.org/10.2976/1.3054712.
Joanny J-F, Ramaswamy S. A drop of active matter. J Fluid Mech. 2012;705:46–57. https://doi.org/10.1017/jfm.2012.131.
Pearson JE. Complex patterns in a simple system. Science. 1993;261(5118):189–92. https://doi.org/10.1126/science.261.5118.189.
Strehle D, Schnauss J, Heussinger C, et al. Transiently crosslinked F-actin bundles. Eur Biophys J. 2011;40(1):93–101. https://doi.org/10.1007/s00249-010-0621-z.
Goldman RD, Khuon S, Chou YH, et al. The function of intermediate filaments in cell shape and cytoskeletal integrity. J Cell Biol. 1996;134(4):971–83.
Pourati J, Maniotis A, Spiegel D, et al. Is cytoskeletal tension a major determinant of cell deformability in adherent endothelial cells? Am J Physiol. 1998;274(5 Pt 1):C1283–9.
Wang N, Im T-N, Chen J, et al. Cell prestress. I. Stiffness and prestress are closely associated in adherent contractile cells. Am J Physiol Cell Physiol. 2002;282(3):C606–16. https://doi.org/10.1152/ajpcell.00269.2001.
Fernandez P, Pullarkat PA, Ott A. A master relation defines the nonlinear viscoelasticity of single fibroblasts. Biophys J. 2006;90(10):3796–805. https://doi.org/10.1529/biophysj.105.072215.
Trepat X, Deng L, An SS, et al. Universal physical responses to stretch in the living cell. Nature. 2007;447(7144):592–5. https://doi.org/10.1038/nature05824.
Krishnan R, Park CY, Lin YC, et al. Reinforcement versus fluidization in cytoskeletal mechanoresponsiveness. PLoS One. 2009;4(5):e5486. https://doi.org/10.1371/journal.pone.0005486.
Wolf K, Te Lindert M, Krause M, et al. Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force. J Cell Biol. 2013;201(7):1069–84. https://doi.org/10.1083/jcb.201210152.
Lange JR, Steinwachs J, Kolb T, et al. Microconstriction arrays for high-throughput quantitative measurements of cell mechanical properties. Biophys J. 2015;109(1):26–34. https://doi.org/10.1016/j.bpj.2015.05.029.
Friedl P, Wolf K, Lammerding J. Nuclear mechanics during cell migration. Curr Opin Cell Biol. 2011;23(1):55–64. https://doi.org/10.1016/j.ceb.2010.10.015.
Dahl KN, Ribeiro AJ, Lammerding J. Nuclear shape, mechanics, and mechanotransduction. Circ Res. 2008;102(11):1307–18. https://doi.org/10.1161/CIRCRESAHA.108.173989.
Swift J, Discher DE. The nuclear lamina is mechano-responsive to ECM elasticity in mature tissue. J Cell Sci. 2014;127(Pt 14):3005–15. https://doi.org/10.1242/jcs.149203.
Harada T, Swift J, Irianto J, et al. Nuclear lamin stiffness is a barrier to 3D migration, but softness can limit survival. J Cell Biol. 2014;204(5):669–82. https://doi.org/10.1083/jcb.201308029.
Händel C, Schmidt BUS, Schiller J, et al. Cell membrane softening in human breast and cervical cancer cells. New J Phys. 2015;17(8):83008. https://doi.org/10.1088/1367-2630/17/8/083008.
Braig S, Schmidt BUS, Stoiber K, et al. Pharmacological targeting of membrane rigidity: implications on cancer cell migration and invasion. New J Phys. 2015;17(8):83007. https://doi.org/10.1088/1367-2630/17/8/083007.
Gracià RS, Bezlyepkina N, Knorr RL, et al. Effect of cholesterol on the rigidity of saturated and unsaturated membranes: fluctuation and electrodeformation analysis of giant vesicles. Soft Matter. 2010;6(7):1472. https://doi.org/10.1039/b920629a.
Lu Y-B, Franze K, Seifert G, et al. Viscoelastic properties of individual glial cells and neurons in the CNS. Proc Natl Acad Sci U S A. 2006;103(47):17759–64. https://doi.org/10.1073/pnas.0606150103.
Radmacher M, Fritz M, Kacher CM, et al. Measuring the viscoelastic properties of human platelets with the atomic force microscope. Biophys J. 1996;70(1):556–67. https://doi.org/10.1016/S0006-3495(96)79602-9.
Radmacher M. Studying the mechanics of cellular processes by atomic force microscopy. In:Cell mechanics, vol. 83. London: Elsevier; 2007. p. 347–72.
Janmey PA, Winer JP, Murray ME, et al. The hard life of soft cells. Cell Motil Cytoskeleton. 2009;66(8):597–605. https://doi.org/10.1002/cm.20382.
Mierke CT, Rosel D, Fabry B, et al. Contractile forces in tumor cell migration. Eur J Cell Biol. 2008;87(8-9):669–76. https://doi.org/10.1016/j.ejcb.2008.01.002.
Engler AJ, Sen S, Sweeney HL, et al. Matrix elasticity directs stem cell lineage specification. Cell. 2006;126(4):677–89. https://doi.org/10.1016/j.cell.2006.06.044.
Fritsch A, Höckel M, Kiessling T, et al. Are biomechanical changes necessary for tumour progression? Nat Phys. 2010;6(10):730–2. https://doi.org/10.1038/nphys1800.
Thery M, Bornens M. Cell shape and cell division. Curr Opin Cell Biol. 2006;18(6):648–57. https://doi.org/10.1016/j.ceb.2006.10.001.
Yeung T, Georges PC, Flanagan LA, et al. Effects of substrate stiffness on cell morphology, cytoskeletal structure, and adhesion. Cell Motil Cytoskeleton. 2005;60(1):24–34. https://doi.org/10.1002/cm.20041.
Thoumine O, Cardoso O, Meister J-J. Changes in the mechanical properties of fibroblasts during spreading: a micromanipulation study. Eur Biophys J. 1999;28(3):222–34. https://doi.org/10.1007/s002490050203.
Wottawah F, Schinkinger S, Lincoln B, et al. Optical rheology of biological cells. Phys Rev Lett. 2005;94(9):98103. https://doi.org/10.1103/PhysRevLett.94.098103.
Schmid-Schönbein GW, Sung KL, Tözeren H, et al. Passive mechanical properties of human leukocytes. Biophys J. 1981;36(1):243–56. https://doi.org/10.1016/S0006-3495(81)84726-1.
Thoumine O, Ott A. Comparison of the mechanical properties of normal and transformed fibroblasts. Biorheology. 1997;34(4-5):309–26. https://doi.org/10.1016/S0006-355X(98)00007-9.
Mahaffy RE, Park S, Gerde E, et al. Quantitative analysis of the viscoelastic properties of thin regions of fibroblasts using atomic force microscopy. Biophys J. 2004;86(3):1777–93. https://doi.org/10.1016/S0006-3495(04)74245-9.
Alcaraz J, Buscemi L, Grabulosa M, et al. Microrheology of human lung epithelial cells measured by atomic force microscopy. Biophys J. 2003;84(3):2071–9. https://doi.org/10.1016/S0006-3495(03)75014-0.
Mizuno D, Bacabac R, Tardin C, et al. High-resolution probing of cellular force transmission. Phys Rev Lett. 2009;102(16):168102. https://doi.org/10.1103/PhysRevLett.102.168102.
Hoffman BD, Massiera G, van Citters KM, et al. The consensus mechanics of cultured mammalian cells. Proc Natl Acad Sci U S A. 2006;103(27):10259–64. https://doi.org/10.1073/pnas.0510348103.
Yamada S, Wirtz D, Kuo SC. Mechanics of living cells measured by laser tracking microrheology. Biophys J. 2000;78(4):1736–47. https://doi.org/10.1016/S0006-3495(00)76725-7.
Crocker JC, Valentine MT, Weeks ER, et al. Two-point microrheology of inhomogeneous soft materials. Phys Rev Lett. 2000;85(4):888–91. https://doi.org/10.1103/PhysRevLett.85.888.
Fabry B, Maksym GN, Butler JP, et al. Time scale and other invariants of integrative mechanical behavior in living cells. Phys Rev E Stat Nonlin Soft Matter Phys. 2003;68(4 Pt 1):41914. https://doi.org/10.1103/PhysRevE.68.041914.
Guck J, Ananthakrishnan R, Moon TJ, et al. Optical deformability of soft biological dielectrics. Phys Rev Lett. 2000;84(23):5451–4. https://doi.org/10.1103/PhysRevLett.84.5451.
Guck J, Ananthakrishnan R, Mahmood H, et al. The optical stretcher: a novel laser tool to micromanipulate cells. Biophys J. 2001;81(2):767–84. https://doi.org/10.1016/S0006-3495(01)75740-2.
Brunner C, Niendorf A, Käs JA. Passive and active single-cell biomechanics: a new perspective in cancer diagnosis. Soft Matter. 2009;5(11):2171. https://doi.org/10.1039/b807545j.
Mietke A, Otto O, Girardo S, et al. Extracting cell stiffness from real-time deformability cytometry: theory and experiment. Biophys J. 2015;109(10):2023–36. https://doi.org/10.1016/j.bpj.2015.09.006.
Otto O, Rosendahl P, Mietke A, et al. Real-time deformability cytometry: on-the-fly cell mechanical phenotyping. Nat Methods. 2015;12(3):199–202. https://doi.org/10.1038/nmeth.3281.
Gossett DR, Tse HTK, Lee SA, et al. Hydrodynamic stretching of single cells for large population mechanical phenotyping. Proc Natl Acad Sci U S A. 2012;109(20):7630–5. https://doi.org/10.1073/pnas.1200107109.
Evans EA. Bending elastic modulus of red blood cell membrane derived from buckling instability in micropipet aspiration tests. Biophys J. 1983;43(1):27–30. https://doi.org/10.1016/S0006-3495(83)84319-7.
Schulze C, Muller K, Kas JA, et al. Compaction of cell shape occurs before decrease of elasticity in CHO-K1 cells treated with actin cytoskeleton disrupting drug cytochalasin D. Cell Motil Cytoskeleton. 2009;66(4):193–201. https://doi.org/10.1002/cm.20341.
Jonas O, Duschl C. Force propagation and force generation in cells. Cytoskeleton. 2010;67(9):555–63. https://doi.org/10.1002/cm.20466.
Fuhs T, Reuter L, Vonderhaid I, et al. Inherently slow and weak forward forces of neuronal growth cones measured by a drift-stabilized atomic force microscope. Cytoskeleton. 2013;70(1):44–53. https://doi.org/10.1002/cm.21080.
Thoumine O, Ott A, Cardoso O, et al. Microplates: a new tool for manipulation and mechanical perturbation of individual cells. J Biochem Biophys Methods. 1999;39(1-2):47–62. https://doi.org/10.1016/S0165-022X(98)00052-9.
Fernandez P, Ott A. Single cell mechanics: stress stiffening and kinematic hardening. Phys Rev Lett. 2008;100(23):238102. https://doi.org/10.1103/PhysRevLett.100.238102.
Benoit M, Gabriel D, Gerisch G, et al. Discrete interactions in cell adhesion measured by single-molecule force spectroscopy. Nat Cell Biol. 2000;2(6):313–7. https://doi.org/10.1038/35014000.
Hoffman BD, Crocker JC. Cell mechanics: dissecting the physical responses of cells to force. Annu Rev Biomed Eng. 2009;11:259–88. https://doi.org/10.1146/annurev.bioeng.10.061807.160511.
Golde T, Schuldt C, Schnauss J, et al. Fluorescent beads disintegrate actin networks. Phys Rev E Stat Nonlinear Soft Matter Phys. 2013;88(4):44601. https://doi.org/10.1103/PhysRevE.88.044601.
Levine L. One- and two-particle microrheology. Phys Rev Lett. 2000;85(8):1774–7. https://doi.org/10.1103/PhysRevLett.85.1774.
Lau AWC, Hoffman BD, Davies A, et al. Microrheology, stress fluctuations, and active behavior of living cells. Phys Rev Lett. 2003;91(19):198101. https://doi.org/10.1103/PhysRevLett.91.198101.
Mijailovich SM, Kojic M, Zivkovic M, et al. A finite element model of cell deformation during magnetic bead twisting. J Appl Physiol. 2002;93(4):1429–36. https://doi.org/10.1152/japplphysiol.00255.2002.
Massiera G, van Citters KM, Biancaniello PL, et al. Mechanics of single cells: rheology, time dependence, and fluctuations. Biophys J. 2007;93(10):3703–13. https://doi.org/10.1529/biophysj.107.111641.
Kreysing MK, Kießling T, Fritsch A, et al. The optical cell rotator. Opt Express. 2008;16(21):16984. https://doi.org/10.1364/OE.16.016984.
Gyger M, Stange R, Kiessling TR, et al. Active contractions in single suspended epithelial cells. Eur Biophys J. 2014;43(1):11–23. https://doi.org/10.1007/s00249-013-0935-8.
Maloney JM, Lehnhardt E, Long AF, et al. Mechanical fluidity of fully suspended biological cells. Biophys J. 2013;105(8):1767–77. https://doi.org/10.1016/j.bpj.2013.08.040.
Maloney JM, van Vliet KJ. Chemoenvironmental modulators of fluidity in the suspended biological cell. Soft Matter. 2014;10(40):8031–42. https://doi.org/10.1039/C4SM00743C.
van Vliet K, Bao G, Suresh S. The biomechanics toolbox: Experimental approaches for living cells and biomolecules. Acta Mater. 2003;51(19):5881–905. https://doi.org/10.1016/j.actamat.2003.09.001.
Pullarkat P, Fernandez P, Ott A. Rheological properties of the eukaryotic cell cytoskeleton. Phys Rep. 2007;449(1-3):29–53. https://doi.org/10.1016/j.physrep.2007.03.002.
Fernández P, Heymann L, Ott A, et al. Shear rheology of a cell monolayer. New J Phys. 2007;9(11):419. https://doi.org/10.1088/1367-2630/9/11/419.
Deng L, Trepat X, Butler JP, et al. Fast and slow dynamics of the cytoskeleton. Nat Mater. 2006;5(8):636–40. https://doi.org/10.1038/nmat1685.
Weihs D, Mason TG, Teitell MA. Bio-microrheology: a frontier in microrheology. Biophys J. 2006;91(11):4296–305. https://doi.org/10.1529/biophysj.106.081109.
Roth KB, Neeves KB, Squier J, et al. High-throughput linear optical stretcher for mechanical characterization of blood cells. Cytometry A. 2016;89(4):391–7. https://doi.org/10.1002/cyto.a.22794.
Szabó B, Szöllösi GJ, Gönci B, et al. Phase transition in the collective migration of tissue cells: experiment and model. Phys Rev E. 2006;74(6):61908. https://doi.org/10.1103/PhysRevE.74.061908.
Deisboeck TS, Couzin ID. Collective behavior in cancer cell populations. BioEssays. 2009;31(2):190–7. https://doi.org/10.1002/bies.200800084.
Sander EE, van Delft S, ten KJP, et al. Matrix-dependent Tiam1/Rac signaling in epithelial cells promotes either cell-cell adhesion or cell migration and is regulated by phosphatidylinositol 3-kinase. J Cell Biol. 1998;143(5):1385–98.
Wu Y, Kanchanawong P, Zaidel-Bar R. Actin-delimited adhesion-independent clustering of E-cadherin forms the nanoscale building blocks of adherens junctions. Dev Cell. 2015;32(2):139–54. https://doi.org/10.1016/j.devcel.2014.12.003.
Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009;119(6):1420–8. https://doi.org/10.1172/JCI39104.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57–70. https://doi.org/10.1016/S0092-8674(00)81683-9.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. https://doi.org/10.1016/j.cell.2011.02.013.
Ma L, Young J, Prabhala H, et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat Cell Biol. 2010;12(3):247–56. https://doi.org/10.1038/ncb2024.
Plutoni C, Bazellieres E, Le Borgne-Rochet M, et al. P-cadherin promotes collective cell migration via a Cdc42-mediated increase in mechanical forces. J Cell Biol. 2016;212(2):199–217. https://doi.org/10.1083/jcb.201505105.
Gaggioli C, Hooper S, Hidalgo-Carcedo C, et al. Fibroblast-led collective invasion of carcinoma cells with differing roles for RhoGTPases in leading and following cells. Nat Cell Biol. 2007;9(12):1392–400. https://doi.org/10.1038/ncb1658.
Ladoux B, Mege R-M, Trepat X. Front-rear polarization by mechanical cues: from single cells to tissues. Trends Cell Biol. 2016;26(6):420–33. https://doi.org/10.1016/j.tcb.2016.02.002.
Mierke CT. The integrin alphav beta3 increases cellular stiffness and cytoskeletal remodeling dynamics to facilitate cancer cell invasion. New J Phys. 2013;15(1):15003. https://doi.org/10.1088/1367-2630/15/1/015003.
Seftor RE, Seftor EA, Gehlsen KR, et al. Role of the alpha v beta 3 integrin in human melanoma cell invasion. Proc Natl Acad Sci U S A. 1992;89(5):1557–61. https://doi.org/10.1073/pnas.89.5.1557.
Aoudjit F, Vuori K. Integrin signaling inhibits paclitaxel-induced apoptosis in breast cancer cells. Oncogene. 2001;20(36):4995–5004. https://doi.org/10.1038/sj.onc.1204554.
Munger JS, Huang X, Kawakatsu H, et al. A mechanism for regulating pulmonary inflammation and fibrosis: the integrin αvβ6 binds and activates latent TGF β1. Cell. 1999;96(3):319–28. https://doi.org/10.1016/S0092-8674(00)80545-0.
Soldi R, Mitola S, Strasly M, et al. Role of alphavbeta3 integrin in the activation of vascular endothelial growth factor receptor-2. EMBO J. 1999;18(4):882–92. https://doi.org/10.1093/emboj/18.4.882.
Bissell MJ, Radisky DC, Rizki A, et al. The organizing principle: microenvironmental influences in the normal and malignant breast. Differentiation. 2002;70(9-10):537–46. https://doi.org/10.1046/j.1432-0436.2002.700907.x.
Dvorak HF, Weaver VM, Tlsty TD, et al. Tumor microenvironment and progression. J Surg Oncol. 2011;103(6):468–74. https://doi.org/10.1002/jso.21709.
Heine P, Ehrlicher A, Käs J. Neuronal and metastatic cancer cells: unlike brothers. Biochim Biophys Acta Mol Cell Res. 2015;1853(11):3126–31. https://doi.org/10.1016/j.bbamcr.2015.06.011.
Ibragimova I, de Cáceres II, Hoffman AM, et al. Global Reactivation of Epigenetically Silenced Genes in Prostate Cancer. Cancer Prev Res (Phila). 2010;3(9):1084–92. https://doi.org/10.1158/1940-6207.CAPR-10-0039.
Karpf AR, Jones DA. Reactivating the expression of methylation silenced genes in human cancer. Oncogene. 2002;21(35):5496–503. https://doi.org/10.1038/sj.onc.1205602.
Cameron EE, Bachman KE, Myohanen S, et al. Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nat Genet. 1999;21(1):103–7. https://doi.org/10.1038/5047.
Riek K, Klatt D, Nuzha H, et al. Wide-range dynamic magnetic resonance elastography. J Biomech. 2011;44(7):1380–6. https://doi.org/10.1016/j.jbiomech.2010.12.031.
Bi D, Yang X, Marchetti MC, et al. Motility-driven glass and jamming transitions in biological tissues. Phys Rev X. 2016;6(2):021011. https://doi.org/10.1103/PhysRevX.6.021011.
Bi D, Lopez JH, Schwarz JM, et al. A density-independent rigidity transition in biological tissues. Nat Phys. 2015;11(12):1074–9. https://doi.org/10.1038/NPHYS3471.
Angelini TE, Hannezo E, Trepat X, et al. Glass-like dynamics of collective cell migration. Proc Natl Acad Sci U S A. 2011;108(12):4714–9. https://doi.org/10.1073/pnas.1010059108.
Bi D, Lopez JH, Schwarz JM, et al. Energy barriers and cell migration in densely packed tissues. Soft Matter. 2014;10(12):1885–90. https://doi.org/10.1039/c3sm52893f.
Farhadifar R, Roper J-C, Aigouy B, et al. The influence of cell mechanics, cell-cell interactions, and proliferation on epithelial packing. Curr Biol. 2007;17(24):2095–104. https://doi.org/10.1016/j.cub.2007.11.049.
Basan M, Prost J, Joanny J-F, et al. Dissipative particle dynamics simulations for biological tissues: rheology and competition. Phys Biol. 2011;8(2):26014. https://doi.org/10.1088/1478-3975/8/2/026014.
Podewitz N, Jülicher F, Gompper G, et al. Interface dynamics of competing tissues. New J Phys. 2016;18(8):83020. https://doi.org/10.1088/1367-2630/18/8/083020.
Zhu Y, Dong Z, Wejinya UC, et al. Determination of mechanical properties of soft tissue scaffolds by atomic force microscopy nanoindentation. J Biomech. 2011;44(13):2356–61. https://doi.org/10.1016/j.jbiomech.2011.07.010.
Schuldt C, Karl A, Korber N, et al. Dose-dependent collagen cross-linking of rabbit scleral tissue by blue light and riboflavin treatment probed by dynamic shear rheology. Acta Ophthalmol. 2015;93(5):e328–36. https://doi.org/10.1111/aos.12621.
Reiss-Zimmermann M, Streitberger K-J, Sack I, et al. High resolution imaging of viscoelastic properties of intracranial tumours by multi-frequency magnetic resonance elastography. Clin Neuroradiol. 2015;25(4):371–8. https://doi.org/10.1007/s00062-014-0311-9.
Steinberg MS. On the mechanism of tissue reconstruction by dissociated cells. I. Population kinetics, differential adhesiveness. and the absence of directed migration. Proc Natl Acad Sci. 1962;48(9):1577–82. https://doi.org/10.1073/pnas.48.9.1577.
Foty RA, Steinberg MS. The differential adhesion hypothesis: a direct evaluation. Dev Biol. 2005;278(1):255–63. https://doi.org/10.1016/j.ydbio.2004.11.012.
Pawlizak S, Fritsch AW, Grosser S, et al. Testing the differential adhesion hypothesis across the epithelial−mesenchymal transition. New J Phys. 2015;17(8):83049. https://doi.org/10.1088/1367-2630/17/8/083049.
Albini A, Iwamoto Y, Kleinman HK, et al. A rapid in vitro assay for quantitating the invasive potential of tumor cells. Cancer Res. 1987;47(12):3239–45.
Paszek MJ, Zahir N, Johnson KR, et al. Tensional homeostasis and the malignant phenotype. Cancer Cell. 2005;8(3):241–54. https://doi.org/10.1016/j.ccr.2005.08.010.
Young JS, Llumsden CE, Stalker AL. The significance of the “tissue pressure” of normal testicular and of neoplastic (Brown-Pearce carcinoma) tissue in the rabbit. J Pathol. 1950;62(3):313–33. https://doi.org/10.1002/path.1700620303.
Plodinec M, Loparic M, Monnier CA, et al. The nanomechanical signature of breast cancer. Nat Nanotechnol. 2012;7(11):757–65. https://doi.org/10.1038/nnano.2012.167.
Nnetu KD, Knorr M, Käs J, et al. The impact of jamming on boundaries of collectively moving weak-interacting cells. New J Phys. 2012;14(11):115012. https://doi.org/10.1088/1367-2630/14/11/115012.
Mouw JK, Yui Y, Damiano L, et al. Tissue mechanics modulate microRNA-dependent PTEN expression to regulate malignant progression. Nat Med. 2014;20(4):360–7. https://doi.org/10.1038/nm.3497.
Haeger A, Krause M, Wolf K, et al. Cell jamming: collective invasion of mesenchymal tumor cells imposed by tissue confinement. Biochim Biophys Acta. 2014;1840(8):2386–95. https://doi.org/10.1016/j.bbagen.2014.03.020.
Park J-A, Kim JH, Bi D, et al. Unjamming and cell shape in the asthmatic airway epithelium. Nat Mater. 2015;14(10):1040–8. https://doi.org/10.1038/nmat4357.
Farina KL, Wyckoff JB, Rivera J, et al. Cell motility of tumor cells visualized in living intact primary tumors using green fluorescent protein. Cancer Res. 1998;58(12):2528–32.
Wang Y-L, Discher DE, editors. Cell mechanics, Methods in cell biology. London: Elsevier; 2007.
Diggs LW. Pathology of sickle cell disease. JAMA. 1971;218(4):600. https://doi.org/10.1001/jama.1971.03190170078040.
Platt OS. Sickle cell anemia as an inflammatory disease. J Clin Invest. 2000;106(3):337–8. https://doi.org/10.1172/JCI10726.
Weinberg RA. The biology of cancer. 2nd ed. New York: Garland Science; 2014.
Seltmann K, Fritsch AW, Käs JA, et al. Keratins significantly contribute to cell stiffness and impact invasive behavior. Proc Natl Acad Sci U S A. 2013;110(46):18507–12. https://doi.org/10.1073/pnas.1310493110.
Cross SE, Jin Y-S, Tondre J, et al. AFM-based analysis of human metastatic cancer cells. Nanotechnology. 2008;19(38):384003. https://doi.org/10.1088/0957-4484/19/38/384003.
Lichtman MA. Rheology of leukocytes, leukocyte suspensions, and blood in leukemia possible relationship to clinical manifestations. J Clin Invest. 1973;52(2):350–8.
Baker EL, Bonnecaze RT, Zaman MH. Extracellular matrix stiffness and architecture govern intracellular rheology in cancer. Biophys J. 2009;97(4):1013–21. https://doi.org/10.1016/j.bpj.2009.05.054.
Mofrad MR. Rheology of the cytoskeleton. Annu Rev Fluid Mech. 2009;41(1):433–53. https://doi.org/10.1146/annurev.fluid.010908.165236.
Guck J, Schinkinger S, Lincoln B, et al. Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence. Biophys J. 2005;88(5):3689–98. https://doi.org/10.1529/biophysj.104.045476.
Merkel M, Manning ML. Using cell deformation and motion to predict forces and collective behavior in morphogenesis. Semin Cell Dev Biol. 2016;67:167. https://doi.org/10.1016/j.semcdb.2016.07.029.
Manning ML, Collins E-MS. Focus on physical models in biology: multicellularity and active matter. New J Phys. 2015;17(4):40201. https://doi.org/10.1088/1367-2630/17/4/040201.
Pegoraro AF, Fredberg JJ, Park J-A. Problems in biology with many scales of length: cell–cell adhesion and cell jamming in collective cellular migration. Exp Cell Res. 2016;343(1):54–9. https://doi.org/10.1016/j.yexcr.2015.10.036.
Weigelin B, Friedl P. Cancer cells: stemness shaped by curvature. Nat Mater. 2016;15(8):827–8. https://doi.org/10.1038/nmat4711.
Te Boekhorst V, Preziosi L, Friedl P. Plasticity of cell migration in vivo and in silico. Annu Rev Cell Dev Biol. 2016;32:491–526. https://doi.org/10.1146/annurev-cellbio-111315-125201.
Collins C, Nelson WJ. Running with neighbors: coordinating cell migration and cell–cell adhesion. Cell Adhes Migr. 2015;36:62–70. https://doi.org/10.1016/j.ceb.2015.07.004.
Kashef J, Franz CM. Quantitative methods for analyzing cell–cell adhesion in development. Dev Biol. 2015;401(1):165–74. https://doi.org/10.1016/j.ydbio.2014.11.002.
Acknowledgments
We thank Till Möhn, Jürgen Lippoldt, Martin Glaser and Benjamin Wolf for their very helpful comments, discussions, advices for illustrations, and language editing.
Glossary
Self-organization: Self-organization is an active, non-equilibrium process of an open system where energy is constantly dissipated and needs to be resupplied, for instance, to generate forces (such as the actin–myosin power stroke) or to organize dynamic structures (such as the lamellipodium for cell migration) far from the thermodynamic equilibrium [10].
Self-assembly: Self-assembly processes are solely based on equilibrium dynamics and are independent of energy dissipation. They occur spontaneously and tend to minimize the free energy of the system driving it toward its thermodynamic equilibrium without an additional energy source such as ATP or GTP. Self-assembly can occur in closed systems [10].
Persistence Length: Mechanical property quantifying the stiffness of a polymer relative to its length. The length scale on which the direction vectors of both ends of a filament lose their correlation.
Bending Stiffness: The resistance of a beam with unit diameter and length while undergoing bending. The higher the bending stiffness, the harder to flex the unit beam.
Molecular Motors: Molecular machines which consume energy (e.g., ATP, GTP) and convert it into motion or mechanical work.
Treadmilling: Steady-state phenomenon of cytoskeletal filaments, mostly actin, where one filament end depolymerizes and the other end polymerizes, leading to shrinkage and growth at the ends with no net length change of the filament.
Tensile Creep Compliance: The magnitude of the creep response of a unit bulk material for a given unit force load. The higher the tensile creep compliance, the easier it deforms under force load.
Elastic Modulus (Young’s Modulus) or Shear Modulus: The resistance of a unit bulk material under axial load or under shearing load, respectively. The higher the elastic modulus or shear modulus, the harder to deform the material. In incompressible materials, the elastic modulus is three times the shear modulus.
Exocytosis: Active transport of molecules out of the cell via a secretory vesicle as transport carrier.
Endocytosis: Active transport of molecules into the cell via encapsulation of the molecules with the cell membrane and formation of a vesicle as transport carrier.
Receptor Binding: Binding of signaling molecules to transmembrane proteins used for cellular and tissue response.
Mechanosignaling: Sensing and signaling of cells induce a response to mechanical, environmental cues.
Glass-like Material: Solid-phase state of a material, where the strong, noncrystalline entanglement of the molecules, usually polymer chains, prevents an unhindered liquid-like flow and movement of the molecules for low thermal energy. Above the glass transition temperature, the material can flow again.
Amorphous Material: Noncrystalline solid with no long-range order, usually consisting of many clustered domains with different (crystalline) orientations and substructures.
Yield Stress Fluid: A fluid which only starts to flow above a critical stress, the yield stress. For stresses below the yield stress, it behaves like a solid.
Homeostatic Stress: Internal stress of a tissue generated by adhesion forces, which is actively regulated to remain close to constant.
Differential Adhesion Hypothesis: Hypothesis for cellular movement in tissues of different cell types based on thermodynamic principles. Cells with different adhesion forces will minimize their free energy by moving to other cells with similar adhesion forces in order to maximize bonding strength.
Jamming: A quasi-phase transition of a material, where rigidity suddenly increases and fluidity suddenly decreases when the density of cells (or molecules) increases above a critical level.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG
About this chapter
Cite this chapter
Kubitschke, H., Morawetz, E.W., Käs, J.A., Schnauß, J. (2018). Physical Properties of Single Cells and Collective Behavior. In: Sack, I., Schaeffter, T. (eds) Quantification of Biophysical Parameters in Medical Imaging. Springer, Cham. https://doi.org/10.1007/978-3-319-65924-4_5
Download citation
DOI: https://doi.org/10.1007/978-3-319-65924-4_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-65923-7
Online ISBN: 978-3-319-65924-4
eBook Packages: MedicineMedicine (R0)