Abstract
Heart disease remains a leading cause of death due to limited therapeutic regiments. Human and murine adult hearts suffer from limited regenerative capacity of cardiomyocytes after injury such as myocardial infarction. The advances of cellular reprogramming that leveraged knowledge from decades of research in developmental biology provide a promising approach to convert cell fate and generate reprogrammed cells in situ. Production of induced cardiomyocytes raises hopes for potential usage in regenerative medicine and disease modeling. Here we reviewed latest advances in reprogramming fibroblasts to cardiomyocyte-like cells and summarized the challenges that must be overcome to move this filed closer for clinic applications.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
- iCM Generation
- Reprogramming Efficiency
- Direct Cardiac Reprogramming
- TGFβ Inhibition
- Converting Fibroblasts
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
4.1 Introduction
Heart disease is the leading cause of adult mortality in the developed world and continues to be a heavy burden to health care systems [1]. Resulting from the limited regenerative capacity of adult cardiomyocytes, it’s difficult for heart to functionally recover after lesions such as myocardium infarction (MI). The lost cardiomyocytes in the injured area are replaced by activated cardiac fibroblasts (CFs) that proliferate and secrete excessive extracellular matrix to form scar tissues and pathologically remodel the myocardium. Although recently studies showed that mammalian hearts possess modest self-renewal and turnover under certain scenarios [2,3,4,5], it is still insufficient to regenerate a damaged heart.
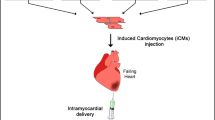
Recent development of direct reprogramming, which directly converts cells from one differentiated phenotype to another without transitioning through the intermediate pluripotent state, offers a promising alternative approach for regenerative medicine. A single or multiple transcription factors have been shown to drive cell fata conversion from fibroblast into neuron like cells, hepatocyte like cells and many other somatic cell types [6]. As for heart, the activated residential CFs upon injury could serve as an endogenous source of new CMs for regenerative purpose if they could be directly reprogrammed into functional CMs. Several groups have successfully converted fibroblasts in to induced CM-like cells (iCMs) using a cocktail of transcription factors that reside at top of developmental regulatory hierarchy for cardiogenesis, both in vitro and in vivo. Alternatively, combinations of small molecules and microRNAs have been developed to either directly reprogram or enhance reprogramming of iCMs (Fig. 4.1). Now more effort has been taken into studying the mechanisms underlying this process. Here we will summarize current advances in direct iCM reprogramming and discuss its challenges and further applications for regenerative medicine.
4.2 Direct Reprogramming of Mouse Fibroblasts into iCMs
4.2.1 Transcription Factors
Starting from 14 transcription factors, Ieda et al. discovered that a specific combination of three factors, Gata 4 (G), Mef2c (M) and Tbx5 (T) (collectively referred to as GMT) was sufficient to transform mouse Thy1+ dermal or cardiac fibroblasts into iCMs [7]. The iCMs exhibit similar global gene expression and epigenetic imprinting as endogenous CMs, whereas the fibroblasts program is significantly repressed. Functionally, iCMs show calcium oscillation and spontaneous beating. Importantly, iCMs do not pass through a cardiac progenitor stage (in particular Mesp1+ and Isl1+ lineages), suggesting iCM generation is a direct conversion from one somatic cell type to another. In accordance with this observation, the fully reprogrammed iCMs morphologically and functionally resemble neonatal cardiomyocytes.
The following in vivo studies using genetic linage tracing strategies demonstrated the regenerative capacity of iCM reprogramming. It has been demonstrated that retroviral delivery of GMT after coronary ligation produced iCMs characterized with mature CM features including bi-nucleation, well-organized sarcomere structures as well as similar gene expression and electrophysiological properties [8]. Importantly, the in vivo reprogramming efficiency is much higher than the in vitro one, suggesting the environmental factors may contribute to the enhancement of this conversion process. In vivo delivery of GMT also improved heart function, decreased infarction size and reduced fibrosis in mice with myocardial infarction [8]. Similarly, injection of GMT retroviruses into immunosuppressed mouse heart suffering MI resulted in newly emerged iCMs 2 weeks after surgery [9]. To overcome the disadvantages of the retro- and lenti-viral infection (integration and chronic expression), Mathison et al. generated replication-deficient adenovirus expressing GMT (Ad-GMT) [10]. These Ad-GMTs appeared to be as efficient as lentiviral GMT for rat iCM reprogramming both in vitro and in vivo [10].
In addition, other combinations of transcription factors have been reported to succeed in converting fibroblasts into iCMs. It has been shown that another transcription factor, Hand2, can function together with GMT (referred to as HGMT) to improve iCM reprogramming efficiency of adult fibroblasts in vitro and produce iCMs in vivo to attenuate heart dysfunction after myocardial injury [11]. In comparison with GMT, HGMT appears to generate diverse cell types including atrial, ventricular and pacemaker cardiomyocytes in vitro [12]. Protze et al. screened a pool of 10 transcription factors in MEFs and found another three factor combination (Mef2c, Tbx5 and Myocd) could induce iCMs with cardiac program and functionally these cells are more mature than GMT derived iCMs [13].
4.2.2 MicroRNAs
MicroRNAs (miRNAs) are a class of small noncoding RNAs of 21–25 nt in length that in general repress gene expression at the posttranscriptional level by degrading their target mRNAs and/or inhibiting their translation [14, 15]. MiRNAs play pivotal roles in governing gene expression during cardiovascular development and disease [16, 17]. For example, miR-1 was the first reported miRNA to be involved in regulation of heart development through targeting Hand2 [18, 19]. Recent studies imply additional import roles of miRNA in controlling cell fate conversion. Using combination of miRNAs, both mouse and human fibroblasts could be directly converted into induced pluripotent cells and neurons [20,21,22]. Based on the potential roles of miRNAs, Jayawardena et al. identified a combination of miRNAs 1, 133, 208, and 499 (referred to as miR combo) that are capable of inducing iCMs both in vitro and in vivo [23,24,25]. iCMs generated by miR combo are characterized with similar gene expression as endogenous CMs, spontaneous calcium flux and contraction. Mice harboring miRNA combo after MI showed newly derived iCMs originated from fibroblasts, and resulted improvement in cardiac function. JAK inhibitor I treatment further increased miR combo mediated iCM reprogramming efficiency [23]. Mechanistically, removal of tri-methylation of the lysine 27 of histone H3 (H3K27me3) is essential for miR combo to initiate the reprogramming [26]. Most recently, with the development of three-dimensional (3D) tissue-engineered cardiac hydrogel patches, miR combo directed iCM reprogramming was further enhanced with observation of strong environmental matrix metalloproteinases expression [27].
4.2.3 Small Molecules
Canonical reprogramming utilizes retroviral or lentiviral based strategies to deliver transcription factors in vitro and in vivo. Application of virus inevitably brings up the challenge of viral integration into the host genome and thus limits the clinical translation. An alternative approach is to use small chemical compounds, which are cell permeable, nonimmunogenic and could be easily handled for the delivery procedure to be standardized. Proof-of-concept studies have demonstrated that combinations of small molecules can replace master transcription genes to initiate iPS production [28]. With this concept, Want et al. demonstrated the transdifferentiation of mouse fibroblasts into cardiomyocytes with a single transcription factor Oct4 and a defined small molecule pool consisting SB431542 (ALK4/5/7 inhibitor), CHIR99021 (GSK3 inhibitor), parnate (LSD1/KDM1 inhibitor), and forskolin (adenylyl cyclase activator) [29]. Fu et al. developed a full chemical approach to generate chemical induced cardiomyocyte-like cells (CiCMs). Using compounds (CRFVPTZ (C, CHIR99021; R, RepSox; F, Forskolin; V, VPA; P, Parnate; and T, TTNPB) together with optimized culture medium, MEFs were amenable to become contractile cardiomyocytes [30]. Of note, different from GMT induced direct reprogramming, CiCMs pass through cardiac progenitor stage with high expression of progenitor markers Msp1 and Isl1 [29, 30].
4.3 Enhancement of Mouse iCM Generation
4.3.1 Optimization of Transcription Factors
Three transcription factors GMT are sufficient to induce cell fate conversion from fibroblasts to iCMs. Suffering from the relatively low efficiency and incomplete reprogramming, several studies aimed at improving reprogramming efficiency through harnessing the transcription factor pool. Addition of MYOCD and SRF alone or in combination with Mesp1 and SMARCD3 enhanced GMT activated basal cardiac gene expression, though no significant difference was observed in terms of myocyte functionality [31]. Taking advantage of a transgenic calcium florescent reporter system driven by cardiac specific Troponin T promoter, Addis et al. evaluated several transcription factor combinations for their capacity to produce functional iCMs [32]. Interestingly, they found that addition of Nkx2-5 to HGMT cocktail (referred to as HNGMT) resulted in highest reprogramming efficiency [32]. There are also studies attempting at modifying activity of one reprogramming factor Mef2c. MyoD was one of the skeletal muscle master genes which has been identified to transform fibroblasts into myoblasts [33]. Hirai et al. fused MyoD transactivation domain to Mef2c and demonstrated that chimeric Mef2c together with Gata4, Tbx5 and Hand2 (referred to as MM3-GHT) yields larger contractile iCM clusters with shortened time window in comparison with traditional HGMT [34]. Alternatively, Abad et al. enhanced binding of Mef2c to the promoter region of cardiac genes, which also resulted in higher reprogramming efficiency [35].
4.3.2 Stoichiometry of Transcription Factors
The use of transcription factor cocktails raised a critical question on relationship between expression level of each exogenous factor and outcome of iCM conversion. During heart development, delicate regulation and dose-spatial-temporal balance of these transcription factors are required to initiate and maintain cardiac specification and differentiation properly [36,37,38,39]. To address this question, Wang et al. manipulated expression level of GMT using polycistronic constructs and showed distinctive protein expression based on splicing orders among identical self-cleaving 2A sequences. They further demonstrated that relative ratio of G, M, T protein was crucial for efficient iCM reprogramming. An optimal expression of GMT with relative high level of M and low levels of G and T achieved by using polycistronic MGT vector (hereafter refer to as MGT) significantly increased reprogramming efficiency and improved iCM quantity and quality in vitro [40, 41]. Moreover, in vivo MGT delivery generated more iCMs and further improved heart functions than traditional delivery of GMT separate viruses [42, 43]. Another two polycistronic constructs encoding GMT were also reported to improve in vivo reprogramming [9, 44]. These reports emphasized the importance of stoichiometric expression of transcription factors and established a single vector platform to facilitate consistent and reproducible iCM reprogramming and further moved the basic and translational research on iCMs.
4.3.3 Addition of Small Molecules
Enhancement of iCM reprogramming could be achieved through addition of small molecules and microRNAs to base transcription factor cocktails (Table 4.1). In brief, there are three main groups of the supplements. First group consists of proteins and peptides like Thymosin β4, Akt and growth factors such as VEGF and FGFs. Thymosin β4 is a natural peptide that has been implied to be critical for cardiac development and play cardio-protective roles upon heart injury [45,46,47,48]. Co-administration of thymosin β4 resulted in a better delivery of GMT to re-activated fibroblasts, hence increased iCM numbers and resulted in further functional improvement in ejection fraction and cardiac output in vivo [8, 49]. Another in vivo research demonstrated that preconditioning rats with VEGF promoted iCM generation and improved cardiac function after MI [50], suggesting the possible important role of angiogenesis for heart repair. In vitro, treatment of serum free medium containing VEGF with FGFs leads to faster maturation of iCMs induced by GMT, possibly through activation of Akt [51]. Overexpression of Akt also led to more efficient generation of contractile iCMs [52]. Moreover, VEGF and FGFs can substitute Gata4 and contribute to direct iCM generation with M and T [51].
The second group consists of chemical compounds targeting epigenetic modifiers and signaling pathways that will be discussed in later section. Noticeably, almost all the drugs were tested in dish, only a very recent study from Srivastava group showed the applicability of small molecules to enhance in vivo reprogramming [53]. They first depicted the reinforcement of in vitro reprogramming by using TGFβ inhibitor and WNT inhibitor with GMT (referred to as GMTc) characterized with shortened duration and enhanced iCM quantity and quality. Mice exposed to GMTc developed smaller scar size, thicker re-muscularized myocardium and further improvement in heart function than GMT alone. At cellular level, the number of ex vivo isolated iCMs from GMTc group was five-fold higher than that from GMT group, and GMTc-iCMs are functionally closer to adult cardiomyocytes in terms of their electrophysiological properties [53].
The third group includes microRNAs with or without chemical compounds. Although miRNA itself could generate iCMs in vitro and in vivo [23, 24], addition of miRNAs to cardiac transcription factors enables higher iCM reprogramming efficiency and better cellular quality. Ectopic expression of miR-133 alone with GMT increased beating iCMs by sevenfold and noticeably enhanced the speed (from 30 to 10 days) for iCM maturation [54]. Overexpression of miR-1 and miR-133 together with GHMT (referred to as GHMT2m) induced more matured iCMs that started to beat by day 8 [55]. In combination with these two miRNAs, ROCK inhibitor and/or TGFβ inhibitor converted fibroblasts into functional iCMs with the efficiency over 60% when quantifying the percentage of beating cells [55].
4.4 Molecular Mechanisms Underlying iCM Generation
4.4.1 Epigenetic Regulation of iCM
Epigenetic regulation plays fundamental roles in cellular specification and lineage commitment during development. Emerging evidence indicates that dysregulated epigenetic landscape contributes to cardiomyopathy and heart failure [59, 60]. Recent studies on cellular reprogramming also demonstrated the dynamic alternation of epigenetic modifications [61,62,63,64]. In the first iCM paper, Ieda et al. discovered that trimethylation of histone H3 at lysine 27 (H3K27me3), a commonly used marker to mark transcriptionally inactive chromatin, was significantly reduced at the promoter region of several cardiac specific genes in iCMs 4 weeks after GMT induction [7]. Whereas, trimethylation of histone H3 at lysine 4 (H3K4me3), which labels an open chromatin, was increased at the same promoter region in iCMs compared to fibroblasts. Liu et al. further analyzed the repatterning of H3K27me3, H3K4me3 at cardiac and fibroblast loci at the MGT mediated reprogramming day3 and day10 [65]. Loss of H3K27me3 at cardiac gene loci appeared as early as day3, suggesting the rapid suppression of fibroblast signatures and early activation of cardiac program. Furthermore, data from ChIP-Seq revealed that upon transduction of GHMT, H3K4 dimethylation (H3K4me2, a general marker of both promoter and enhancer regions [66, 67]) peak shifted from fibroblast toward myocyte status at reprogramming day 7, indicating the existence of epigenetic orchestration at gene regulatory regions during early phase of iCM reprogramming [55].
To explore the underlying mechanism and identify potential epigenetic barriers to iCM reprogramming, our lab performed the first loss of function screen with a shRNA pool consisting 35 components that were involved in chromatin remodeling and modification and identified several factors that could either facilitate or blunt iCM reprogramming [68]. In particular, Bmi1, an important component of the Polycomb repressive complex 1 (PRC1) [69, 70], functioned as a major epigenetic barrier at the early stage of iCM reprogramming. Bmi1 suppressed iCM reprogramming through direct binding to a battery of cardiogenic loci including Gata4, Nkx2.5, Isl1, Pitx2, Tbx20, and Hand2. Furthermore, we demonstrated that Bmi1 depletion could replace Gata4 and convert fibroblasts into iCMs together with a single vector encoding Mef2c and Tbx5.
Liu et al. adopted a gain-of-function approach and identified Men1 and Suv39h1 as epigenetic inhibitors of iCM reprogramming [58]. Men1 is an essential component of a MLL/SET1 histone methyltransferase (HMT) complex responsible for H3K4 methylation and H3K9 methylation [71,72,73]. Suv39h1 also mediates H3K9 methylation [71, 74]. Chemical inhibitors targeting MLL1 complex to repress H3K4 methyltransferase activity significantly enhanced reprogramming efficiency, indicating that Men1 regulate iCM generation through modifying H3K4m3 instead of H3K9m3.
Enhancer of Zeste Homolog 2 (Ezh2), a catalytic subunit of PRC2 complex for H3K27me2 and H3K27me3, behaved as one of the epigenetic barriers for MM3-GHT mediated reprogramming [56]. Exposure to Ezh2 inhibitor GSK126 resulted in a decrease of H3K27me3 and an increase of beating iCM clusters. Similarly, UNC0638, an inhibitor to G9a and GLP that mainly controls H3K9me and H3K9me2, led to a higher iCM reprogramming efficiency in association with lower level of H3K9me2 in iCMs [56].
Taken together, iCM reprogramming is largely guided by specific cardiac transcription factors and the associated chromatin modifiers to establish authentic myocyte cell fate in another distinct cell type.
4.4.2 Suppression of Fibroblast Program
During reprograming, transcription factors drive fibroblast toward a differentiated cardiomyocyte lineage. Genome wide transcriptome research demonstrated that iCM reprogramming requires depletion of the original fibroblast signatures and de novo establishment of myocyte programs such as the contractile machinery, sarcomere structures, high mass of mitochondria and the metabolic switches [35, 51,52,53,54,55]. Suppression of fibroblast program has been shown to fundamentally affect iCM reprogramming. Overexpression of miR133 with GMT repressed Snail1 to silence fibroblast signatures and activates cardiac programs [54]. Snail1 is one of the major mediators of epithelial-mesenchymal transition (EMT) that contributes to cardiac fibrosis [75, 76]. MiR133 directly targeted Snail1 for degradation and overexpression of Snail1 inhibited iCM reprogramming.
Accumulating studies revealed the pivotal role of TGFβ signaling pathway during iCM conversion [53, 55, 57]. TGFβ signals activate cardiac microvascular endothelial cells to undergo endothelial-to-mesenchymal transformation and contributes to cardiac fibrosis [77, 78]. TGFβ also behaves as a repressor for embryonic cells differentiation toward cardiomyocytes [79]. Generally, TGFβ superfamily members bind to TGFβ type II receptor, which subsequently recruits and triggers phosphorylation of TGFβ type I receptor. Phosphorylated type I receptor activates SMAD molecules and leads to formation of SMAD complex. Activated SMAD complex translocates into nucleus and interacts with other DNA binding factors, transcription factors, thus regulates the transcription of target gene [80]. One of the most commonly used TGFβ inhibitors is SB-431542 that selectively blocks the TGF-β type I receptor including ALK4 and ALK5, as well as ALK7 [81]. Both Srivastava group [53] and Gearhart [57] group screened out this inhibitor for its application in iCM reprogramming, while Song group [55] identified another TGF-β inhibitor termed as A8301 that inactivates similar receptors for its use in enhancing iCM induction. Through disturbing TGFβ signaling with chemical inhibitors, all three researches achieved much greater reprogramming quality. Most iCMs generated with the help of TGFβ inhibitors were relatively more reprogrammed beating cells with transcriptome more similar to adult cardiomyocytes [53, 55, 57].
4.5 Direct Cardiac Reprograming in Human Cells
Compared to the rapid advances of murine iCM programming, generation of human iCMs in vitro is more complicated thus much delayed. Neither GMT nor GHMT was sufficient to induce human iCMs [82,83,84]. Screening additional transcription factors finally led to successful induction of cardiomyocyte-like cells from human fibroblasts. Fu et al. discovered that the combination of GMT with ESRRG, MESP1 was sufficient to turn on cardiac specific markers in transduced human fibroblasts. Addition of myocardin and ZFPM2 further enhanced the reprogramming and resulted in iCMs exhibiting calcium flux and action potential [82]. More recently, with the help of two chemical inhibitors (TGFβ inhibitor and WMT inhibitor), the seven transcription factors (7c) induced reprogramming was further accelerated [53]. In addition, the 7c cocktail could be cut down to a four-factor recipe (GMT plus myocardin) with the two inhibitors, indicating the critical role of TGFβ and WNT signaling for human iCM reprogramming. Wada et al. showed that addition of Mesp1 and Myocd to GMT (referred to as GMTMM) cocktail transformed HCF (human cardiac fibroblasts) and HDF (human dermal fibroblasts) to iCMs that expressed a broad panel of cardiac markers, exhibited calcium oscillation and contracted synchronously when co-cultured murine primary CMs [84]. Later they demonstrated that miR-133 mediated snail1 inhibition in human fibroblasts is as important as that for mouse iCM reprogramming. Inclusion of miR-133 or snail1 depletion promoted GMTMM induced human iCM reprogramming [54]. Nam et al. showed that combination of GHMT with myocardin generates few beating cells after 11 weeks in culture [83]. Addition of miR590 to GHMT with myocardin upregulated cardiac gene expression and further suppressed fibroblast marker genes by directly inhibition of Sp1 (specificity protein 1) expression [85].
Generation of expandable cardiac progenitor cells (CPC) from fibroblasts shed lights on the production of CMs through de-differentiation. CPCs can differentiate into three major cell types of heart- endothelial cells, CMs and smooth muscle cells [86, 87]. Transcription factors ETS2 and MESP1 have been reported to transdifferentiate HDF into cardiac progenitors [88]. Lalit et al. reprogrammed adult mouse fibroblasts into induced CPCs (iCPCs) with a cocktail of five transcription factors (Mesp1, Tbx5, Gata4, Nkx2.5, and Baf60c) with two compounds (BIO and LIF) [89]. iCPCs were capable of proliferation and differentiation into all three cell types and generated new myocardium post MI [89]. In comparison, Zhang et al. transiently expressed four Yamanaka factors (Oct4, Sox2, Klf4 and c-myc) in fibroblasts, cultured these primed cells in conditioned medium and ended with acquisition of iCPCs [90]. However, the precise outcome of the final differentiation is difficult to be controlled, with the likelihood of contamination of cells from non-cardiac lineages. Independent of transcription factors, Cao et al. demonstrated that 9 chemical compounds, called 9c, successfully transformed human foreskin fibroblasts to cardiomyocyte like cells [91]. The chemically induced iCMs (ciCM) sequentially expressed mesoderm, CPC and CM genes and eventually became spontaneously beating cells. It appears that ciCMs acquire similar transcriptional and epigenetic signatures, as well as functional properties that are similar to human CMs.
In summary, current studies (summarized in Table 4.2) paved a great foundation for future translational applications that require generation of mature human cardiomyocytes from different resources. Induction of human iCMs takes longer time and requires more factors; in addition human iCMs are far less mature, all of which suggest the need of further refinement from laboratory work. Research using cells from large animals in particular non-human primates could serve as an alternative to study the combination of factors with small molecules on the outcomes of iCM reprogramming, the outcome of which may facilitate the ultimate goal of effectively generating human iCMs.
4.6 Conclusions and Perspectives
Direct cardiac reprogramming holds great potential for regenerative medicine by offering an alternative strategy for treatment of heart disease and disease modeling. Recent studies indicated that the reprogramming efficiency is steadily increased by utilizing multiple strategies and through understanding the molecular mechanisms. Although it has been progressed rapidly, there are still challenges for this field.
First, the reprogramming efficiency is still low and varies between labs. The majority of reprogramed cells are not functionally fully matured cardiomyocytes. It’s undoubtedly required to further optimize the platform and remove the molecular barriers so as to obtain sufficient highly matured iCMs for drug screening and disease modeling. One of the intriguing areas is to identify the contributing factors by evolving “omics” technologies, such as genomics, transcriptomics, proteomics and metabolics, which would reveal the genetic and epigenetic regulation, protein interaction network and metabolite profiles. Interrogating data from these techniques can help to understand more thoroughly about the events and mechanisms underlying the iCM reprogramming. Recent single cell RNA sequencing analysis (scRNA-seq) offers another opportunity to dissect the reprogramming trajectory, profile dynamic gene expression and identify cell fate determinants. Researches from Treutlein et al. set an example as how to use scRNA-seq to gain mechanistic understanding for induced neuron reprogramming [93].
Second, most studies used virus-based strategies to deliver reprogramming factors, which inevitably raises up the safety issues such as genomic integration and subsequent tumorigenesis [6]. To address this concern, safer delivery vectors, such as bio-safe Adeno-associated virus (AAV) based vectors [94], could be developed and optimized of reprogramming. Moreover, the use of small molecules would be another promising way for the clinical application of iCM reprogramming.
Last but not least, direct in vivo reprogramming yields iCMs at higher efficiency, better quantity and quality than iCMs generated from in vitro [8], clearly suggesting the fundamental effective amelioration of environmental niches. Mechanical forces, inflammatory responses, angiogenesis and extracellular matrix could be potential contributors for iCM maturation. Studies using growth factors and small peptide to mimic the environmental changes showed the enhancement of iCM reprogramming, however, the detailed mechanism remains unclear [8, 49, 51]. Identification and understanding the role of these environmental factors could further benefit iCM production and harnessing this approach for regenerative purpose. An alternative approach to identify microenvironmental clues is the application of recently advanced bioengineered materials. Biomaterials have been shown to influence cell fate and behavior through mutual interaction between cells and their environment [95]. Manipulation of the biophysical and biochemical properties of certain biomaterials leads to an improvement of iCM reprogramming [27, 96,97,98], indicating that it can be used to engineer the niches and realize the controllable release of reprogramming factors and small molecules in situ for in vivo reprogramming. One more consideration for experimental biologists is the utilization of large animal species for in vivo iCM reprogramming, such as the porcine model that raises exciting prospect for future iCM based therapies. Compared to murine models, pigs are anatomically and physiologically more similar to humans in cardiovascular, skeletal muscle, immune, and metabolic systems. It can not only solve the aforementioned biosafety issues and explore environmental niches, but also allows researchers to overcome the possible adverse effects of arrhythmia caused by newly formed iCMs in the scar region of myocardium. In particular, efficient and improved genetic engineering approaches for pigs are now available, facilitating the establishment of tailored animal models for mimicking human diseases.
In summary, direct cardiac reprogramming converts injury-activated fibroblast into terminally differentiated cardiomyocytes in situ, holding tremendous potential for healing the injured heart. After better understanding the molecular mechanisms and overcoming the obstacles discussed, we anticipate that we can ultimately harness the iCM reprogramming and translate it to mend the broken heart.
References
Benjamin EJ, Blaha MJ, Chiuve SE, Cushman M, Das SR, Deo R, de Ferranti SD, Floyd J, Fornage M, Gillespie C, Isasi CR, Jimenez MC, Jordan LC, Judd SE, Lackland D, Lichtman JH, Lisabeth L, Liu S, Longenecker CT, Mackey RH, Matsushita K, Mozaffarian D, Mussolino ME, Nasir K, Neumar RW, Palaniappan L, Pandey DK, Thiagarajan RR, Reeves MJ, Ritchey M, Rodriguez CJ, Roth GA, Rosamond WD, Sasson C, Towfighi A, Tsao CW, Turner MB, Virani SS, Voeks JH, Willey JZ, Wilkins JT, Wu JH, Alger HM, Wong SS, Muntner P. Heart disease and stroke statistics-2017 update: a report from the American Heart Association. Circulation. 2017;135(10):e146–603. doi:10.1161/CIR.0000000000000485.
Nakada Y, Canseco DC, Thet S, Abdisalaam S, Asaithamby A, Santos CX, Shah AM, Zhang H, Faber JE, Kinter MT, Szweda LI, Xing C, Hu Z, Deberardinis RJ, Schiattarella G, Hill JA, Oz O, Lu Z, Zhang CC, Kimura W, Sadek HA. Hypoxia induces heart regeneration in adult mice. Nature. 2017;541(7636):222–7. doi:10.1038/nature20173.
Senyo SE, Steinhauser ML, Pizzimenti CL, Yang VK, Cai L, Wang M, Wu TD, Guerquin-Kern JL, Lechene CP, Lee RT. Mammalian heart renewal by pre-existing cardiomyocytes. Nature. 2013;493(7432):433–6. doi:10.1038/nature11682.
Bergmann O, Zdunek S, Felker A, Salehpour M, Alkass K, Bernard S, Sjostrom SL, Szewczykowska M, Jackowska T, Dos Remedios C, Malm T, Andra M, Jashari R, Nyengaard JR, Possnert G, Jovinge S, Druid H, Frisen J. Dynamics of cell generation and turnover in the human heart. Cell. 2015;161(7):1566–75. doi:10.1016/j.cell.2015.05.026.
Kimura W, Xiao F, Canseco DC, Muralidhar S, Thet S, Zhang HM, Abderrahman Y, Chen R, Garcia JA, Shelton JM, Richardson JA, Ashour AM, Asaithamby A, Liang H, Xing C, Lu Z, Zhang CC, Sadek HA. Hypoxia fate mapping identifies cycling cardiomyocytes in the adult heart. Nature. 2015;523(7559):226–30. doi:10.1038/nature14582.
Xu J, Du Y, Deng H. Direct lineage reprogramming: strategies, mechanisms, and applications. Cell Stem Cell. 2015;16(2):119–34. doi:10.1016/j.stem.2015.01.013.
Ieda M, Fu JD, Delgado-Olguin P, Vedantham V, Hayashi Y, Bruneau BG, Srivastava D. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell. 2010;142(3):375–86. doi:10.1016/j.cell.2010.07.002.
Qian L, Huang Y, Spencer CI, Foley A, Vedantham V, Liu L, Conway SJ, Fu JD, Srivastava D. In vivo reprogramming of murine cardiac fibroblasts into induced cardiomyocytes. Nature. 2012;485(7400):593–8. doi:10.1038/nature11044.
Inagawa K, Miyamoto K, Yamakawa H, Muraoka N, Sadahiro T, Umei T, Wada R, Katsumata Y, Kaneda R, Nakade K, Kurihara C, Obata Y, Miyake K, Fukuda K, Ieda M. Induction of cardiomyocyte-like cells in infarct hearts by gene transfer of Gata4, Mef2c, and Tbx5. Circ Res. 2012;111(9):1147–56. doi:10.1161/CIRCRESAHA.112.271148.
Mathison M, Singh VP, Chiuchiolo MJ, Sanagasetti D, Mao Y, Patel VB, Yang J, Kaminsky SM, Crystal RG, Rosengart TK. In situ reprogramming to transdifferentiate fibroblasts into cardiomyocytes using adenoviral vectors: implications for clinical myocardial regeneration. J Thorac Cardiovasc Surg. 2017;153(2):329–339. e323. doi:10.1016/j.jtcvs.2016.09.041.
Song K, Nam YJ, Luo X, Qi X, Tan W, Huang GN, Acharya A, Smith CL, Tallquist MD, Neilson EG, Hill JA, Bassel-Duby R, Olson EN. Heart repair by reprogramming non-myocytes with cardiac transcription factors. Nature. 2012;485(7400):599–604. doi:10.1038/nature11139.
Nam YJ, Lubczyk C, Bhakta M, Zang T, Fernandez-Perez A, McAnally J, Bassel-Duby R, Olson EN, Munshi NV. Induction of diverse cardiac cell types by reprogramming fibroblasts with cardiac transcription factors. Development. 2014;141(22):4267–78. doi:10.1242/dev.114025.
Protze S, Khattak S, Poulet C, Lindemann D, Tanaka EM, Ravens U. A new approach to transcription factor screening for reprogramming of fibroblasts to cardiomyocyte-like cells. J Mol Cell Cardiol. 2012;53(3):323–32. doi:10.1016/j.yjmcc.2012.04.010.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281–97.
Ling H, Fabbri M, Calin GA. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat Rev Drug Discov. 2013;12(11):847–65. doi:10.1038/nrd4140.
Liu N, Olson EN. MicroRNA regulatory networks in cardiovascular development. Dev Cell. 2010;18(4):510–25. doi:10.1016/j.devcel.2010.03.010.
Cordes KR, Srivastava D. MicroRNA regulation of cardiovascular development. Circ Res. 2009;104(6):724–32. doi:10.1161/CIRCRESAHA.108.192872.
Zhao Y, Samal E, Srivastava D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature. 2005;436(7048):214–20. doi:10.1038/nature03817.
Zhao Y, Ransom JF, Li A, Vedantham V, von Drehle M, Muth AN, Tsuchihashi T, McManus MT, Schwartz RJ, Srivastava D. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell. 2007;129(2):303–17. doi:10.1016/j.cell.2007.03.030.
Yoo AS, Sun AX, Li L, Shcheglovitov A, Portmann T, Li Y, Lee-Messer C, Dolmetsch RE, Tsien RW, Crabtree GR. MicroRNA-mediated conversion of human fibroblasts to neurons. Nature. 2011;476(7359):228–31. doi:10.1038/nature10323.
Anokye-Danso F, Trivedi CM, Juhr D, Gupta M, Cui Z, Tian Y, Zhang Y, Yang W, Gruber PJ, Epstein JA, Morrisey EE. Highly efficient miRNA-mediated reprogramming of mouse and human somatic cells to pluripotency. Cell Stem Cell. 2011;8(4):376–88. doi:10.1016/j.stem.2011.03.001.
Judson RL, Babiarz JE, Venere M, Blelloch R. Embryonic stem cell-specific microRNAs promote induced pluripotency. Nat Biotechnol. 2009;27(5):459–61. doi:10.1038/nbt.1535.
Jayawardena TM, Egemnazarov B, Finch EA, Zhang L, Payne JA, Pandya K, Zhang Z, Rosenberg P, Mirotsou M, Dzau VJ. MicroRNA-mediated in vitro and in vivo direct reprogramming of cardiac fibroblasts to cardiomyocytes. Circ Res. 2012;110(11):1465–73. doi:10.1161/CIRCRESAHA.112.269035.
Jayawardena TM, Finch EA, Zhang L, Zhang H, Hodgkinson CP, Pratt RE, Rosenberg PB, Mirotsou M, Dzau VJ. MicroRNA induced cardiac reprogramming in vivo: evidence for mature cardiac myocytes and improved cardiac function. Circ Res. 2015;116(3):418–24. doi:10.1161/CIRCRESAHA.116.304510.
Jayawardena T, Mirotsou M, Dzau VJ. Direct reprogramming of cardiac fibroblasts to cardiomyocytes using microRNAs. Methods Mol Biol. 2014;1150:263–72. doi:10.1007/978-1-4939-0512-6_18.
Dal-Pra S, Hodgkinson CP, Mirotsou M, Kirste I, Dzau VJ. Demethylation of H3K27 is essential for the induction of direct cardiac reprogramming by miR combo. Circ Res. 2017;120:1403. doi:10.1161/CIRCRESAHA.116.308741.
Li Y, Dal-Pra S, Mirotsou M, Jayawardena TM, Hodgkinson CP, Bursac N, Dzau VJ. Tissue-engineered 3-dimensional (3D) microenvironment enhances the direct reprogramming of fibroblasts into cardiomyocytes by microRNAs. Sci Rep. 2016;6:38815. doi:10.1038/srep38815.
Hou P, Li Y, Zhang X, Liu C, Guan J, Li H, Zhao T, Ye J, Yang W, Liu K, Ge J, Xu J, Zhang Q, Zhao Y, Deng H. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science. 2013;341(6146):651–4. doi:10.1126/science.1239278.
Wang H, Cao N, Spencer CI, Nie B, Ma T, Xu T, Zhang Y, Wang X, Srivastava D, Ding S. Small molecules enable cardiac reprogramming of mouse fibroblasts with a single factor, Oct4. Cell Rep. 2014;6(5):951–60. doi:10.1016/j.celrep.2014.01.038.
Fu Y, Huang C, Xu X, Gu H, Ye Y, Jiang C, Qiu Z, Xie X. Direct reprogramming of mouse fibroblasts into cardiomyocytes with chemical cocktails. Cell Res. 2015;25(9):1013–24. doi:10.1038/cr.2015.99.
Christoforou N, Chellappan M, Adler AF, Kirkton RD, Wu T, Addis RC, Bursac N, Leong KW. Transcription factors MYOCD, SRF, Mesp1 and SMARCD3 enhance the cardio-inducing effect of GATA4, TBX5, and MEF2C during direct cellular reprogramming. PLoS One. 2013;8(5):e63577. doi:10.1371/journal.pone.0063577.
Addis RC, Ifkovits JL, Pinto F, Kellam LD, Esteso P, Rentschler S, Christoforou N, Epstein JA, Gearhart JD. Optimization of direct fibroblast reprogramming to cardiomyocytes using calcium activity as a functional measure of success. J Mol Cell Cardiol. 2013;60:97–106. doi:10.1016/j.yjmcc.2013.04.004.
Davis RL, Weintraub H, Lassar AB. Expression of a single transfected cDNA converts fibroblasts to myoblasts. Cell. 1987;51(6):987–1000.
Hirai H, Katoku-Kikyo N, Keirstead SA, Kikyo N. Accelerated direct reprogramming of fibroblasts into cardiomyocyte-like cells with the MyoD transactivation domain. Cardiovasc Res. 2013;100(1):105–13. doi:10.1093/cvr/cvt167.
Abad M, Hashimoto H, Zhou H, Morales MG, Chen B, Bassel-Duby R, Olson EN. Notch inhibition enhances cardiac reprogramming by increasing MEF2C transcriptional activity. Stem Cell Rep. 2017;8:548. doi:10.1016/j.stemcr.2017.01.025.
Lin Q, Schwarz J, Bucana C, Olson EN. Control of mouse cardiac morphogenesis and myogenesis by transcription factor MEF2C. Science. 1997;276(5317):1404–7.
Grepin C, Robitaille L, Antakly T, Nemer M. Inhibition of transcription factor GATA-4 expression blocks in vitro cardiac muscle differentiation. Mol Cell Biol. 1995;15(8):4095–102.
Kuo CT, Morrisey EE, Anandappa R, Sigrist K, Lu MM, Parmacek MS, Soudais C, Leiden JM. GATA4 transcription factor is required for ventral morphogenesis and heart tube formation. Genes Dev. 1997;11(8):1048–60.
Molkentin JD, Lin Q, Duncan SA, Olson EN. Requirement of the transcription factor GATA4 for heart tube formation and ventral morphogenesis. Genes Dev. 1997;11(8):1061–72.
Wang L, Liu Z, Yin C, Asfour H, Chen OM, Li Y, Bursac N, Liu J, Qian L. Stoichiometry of Gata4, Mef2c, and Tbx5 influences the efficiency and quality of induced cardiac Myocyte reprogramming. Circ Res. 2014;116:237. doi:10.1161/CIRCRESAHA.116.305547.
Wang L, Liu Z, Yin C, Zhou Y, Liu J, Qian L. Improved generation of induced cardiomyocytes using a polycistronic construct expressing optimal ratio of Gata4, Mef2c and Tbx5. J Vis Exp. 2015;105:e53426. https://doi.org/10.3791/53426.
Ma H, Wang L, Yin C, Liu J, Qian L. In vivo cardiac reprogramming using an optimal single polycistronic construct. Cardiovasc Res. 2015;108(2):217–9. doi:10.1093/cvr/cvv223.
Ma H, Wang L, Liu J, Qian L. Direct cardiac reprogramming as a novel therapeutic strategy for treatment of myocardial infarction. Methods Mol Biol. 2017;1521:69–88. doi:10.1007/978-1-4939-6588-5_5.
Mathison M, Singh VP, Gersch RP, Ramirez MO, Cooney A, Kaminsky SM, Chiuchiolo MJ, Nasser A, Yang J, Crystal RG, Rosengart TK. “Triplet” polycistronic vectors encoding Gata4, Mef2c, and Tbx5 enhances postinfarct ventricular functional improvement compared with singlet vectors. J Thorac Cardiovasc Surg. 2014;148(4):1656–1664. e1652. doi:10.1016/j.jtcvs.2014.03.033.
Gajzer DC, Balbin J, Chaudhry HW. Thymosin beta4 and cardiac regeneration: are we missing a beat? Stem Cell Rev. 2013;9(3):303–12. doi:10.1007/s12015-012-9378-3.
Bock-Marquette I, Saxena A, White MD, Dimaio JM, Srivastava D. Thymosin beta4 activates integrin-linked kinase and promotes cardiac cell migration, survival and cardiac repair. Nature. 2004;432(7016):466–72. doi:10.1038/nature03000.
Smart N, Risebro CA, Melville AA, Moses K, Schwartz RJ, Chien KR, Riley PR. Thymosin beta4 induces adult epicardial progenitor mobilization and neovascularization. Nature. 2007;445(7124):177–82. doi:10.1038/nature05383.
Peng H, Xu J, Yang XP, Dai X, Peterson EL, Carretero OA, Rhaleb NE. Thymosin-beta4 prevents cardiac rupture and improves cardiac function in mice with myocardial infarction. Am J Physiol Heart Circ Physiol. 2014;307(5):H741–51. doi:10.1152/ajpheart.00129.2014.
Srivastava D, Ieda M, Fu J, Qian L. Cardiac repair with thymosin beta4 and cardiac reprogramming factors. Ann N Y Acad Sci. 2012;1270:66–72. doi:10.1111/j.1749-6632.2012.06696.x.
Mathison M, Gersch RP, Nasser A, Lilo S, Korman M, Fourman M, Hackett N, Shroyer K, Yang J, Ma Y, Crystal RG, Rosengart TK. In vivo cardiac cellular reprogramming efficacy is enhanced by angiogenic preconditioning of the infarcted myocardium with vascular endothelial growth factor. J Am Heart Assoc. 2012;1(6):e005652. doi:10.1161/JAHA.112.005652.
Yamakawa H, Muraoka N, Miyamoto K, Sadahiro T, Isomi M, Haginiwa S, Kojima H, Umei T, Akiyama M, Kuishi Y, Kurokawa J, Furukawa T, Fukuda K, Ieda M. Fibroblast growth factors and vascular endothelial growth factor promote cardiac reprogramming under defined conditions. Stem Cell Rep. 2015;5(6):1128–42. doi:10.1016/j.stemcr.2015.10.019.
Zhou H, Dickson ME, Kim MS, Bassel-Duby R, Olson EN. Akt1/protein kinase B enhances transcriptional reprogramming of fibroblasts to functional cardiomyocytes. Proc Natl Acad Sci U S A. 2015;112(38):11864–9. doi:10.1073/pnas.1516237112.
Mohamed TM, Stone NR, Berry EC, Radzinsky E, Huang Y, Pratt K, Ang YS, Yu P, Wang H, Tang S, Magnitsky S, Ding S, Ivey KN, Srivastava D. Chemical enhancement of in vitro and in vivo direct cardiac reprogramming. Circulation. 2017;135(10):978–95. doi:10.1161/CIRCULATIONAHA.116.024692.
Muraoka N, Yamakawa H, Miyamoto K, Sadahiro T, Umei T, Isomi M, Nakashima H, Akiyama M, Wada R, Inagawa K, Nishiyama T, Kaneda R, Fukuda T, Takeda S, Tohyama S, Hashimoto H, Kawamura Y, Goshima N, Aeba R, Yamagishi H, Fukuda K, Ieda M. MiR-133 promotes cardiac reprogramming by directly repressing Snai1 and silencing fibroblast signatures. EMBO J. 2014;33(14):1565–81. doi:10.15252/embj.201387605.
Zhao Y, Londono P, Cao Y, Sharpe EJ, Proenza C, O’Rourke R, Jones KL, Jeong MY, Walker LA, Buttrick PM, McKinsey TA, Song K. High-efficiency reprogramming of fibroblasts into cardiomyocytes requires suppression of pro-fibrotic signalling. Nat Commun. 2015;6:8243. doi:10.1038/ncomms9243.
Hirai H, Kikyo N. Inhibitors of suppressive histone modification promote direct reprogramming of fibroblasts to cardiomyocyte-like cells. Cardiovasc Res. 2014;102(1):188–90. doi:10.1093/cvr/cvu023.
Ifkovits JL, Addis RC, Epstein JA, Gearhart JD. Inhibition of TGFbeta signaling increases direct conversion of fibroblasts to induced cardiomyocytes. PLoS One. 2014;9(2):e89678. doi:10.1371/journal.pone.0089678.
Liu L, Lei I, Karatas H, Li Y, Wang L, Gnatovskiy L, Dou Y, Wang S, Qian L, Wang Z. Targeting Mll1 H3K4 methyltransferase activity to guide cardiac lineage specific reprogramming of fibroblasts. Cell Discov. 2016;2:16036. doi:10.1038/celldisc.2016.36.
Han P, Hang CT, Yang J, Chang CP. Chromatin remodeling in cardiovascular development and physiology. Circ Res. 2011;108(3):378–96. doi:10.1161/CIRCRESAHA.110.224287.
Wamstad JA, Alexander JM, Truty RM, Shrikumar A, Li F, Eilertson KE, Ding H, Wylie JN, Pico AR, Capra JA, Erwin G, Kattman SJ, Keller GM, Srivastava D, Levine SS, Pollard KS, Holloway AK, Boyer LA, Bruneau BG. Dynamic and coordinated epigenetic regulation of developmental transitions in the cardiac lineage. Cell. 2012;151(1):206–20. doi:10.1016/j.cell.2012.07.035.
Luna-Zurita L, Bruneau BG. Chromatin modulators as facilitating factors in cellular reprogramming. Curr Opin Genet Dev. 2013;23(5):556–61. doi:10.1016/j.gde.2013.07.002.
Onder TT, Kara N, Cherry A, Sinha AU, Zhu N, Bernt KM, Cahan P, Marcarci BO, Unternaehrer J, Gupta PB, Lander ES, Armstrong SA, Daley GQ. Chromatin-modifying enzymes as modulators of reprogramming. Nature. 2012;483(7391):598–602. doi:10.1038/nature10953.
Apostolou E, Hochedlinger K. Chromatin dynamics during cellular reprogramming. Nature. 2013;502(7472):462–71. doi:10.1038/nature12749.
Shchuka VM, Malek-Gilani N, Singh G, Langroudi L, Dhaliwal NK, Moorthy SD, Davidson S, Macpherson NN, Mitchell JA. Chromatin dynamics in lineage commitment and cellular reprogramming. Genes (Basel). 2015;6(3):641–61. doi:10.3390/genes6030641.
Liu Z, Chen O, Zheng M, Wang L, Zhou Y, Yin C, Liu J, Qian L. Re-patterning of H3K27me3, H3K4me3 and DNA methylation during fibroblast conversion into induced cardiomyocytes. Stem Cell Res. 2016;16(2):507–18. doi:10.1016/j.scr.2016.02.037.
Bernstein BE, Kamal M, Lindblad-Toh K, Bekiranov S, Bailey DK, Huebert DJ, McMahon S, Karlsson EK, Kulbokas EJ 3rd, Gingeras TR, Schreiber SL, Lander ES. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell. 2005;120(2):169–81. doi:10.1016/j.cell.2005.01.001.
Heintzman ND, Stuart RK, Hon G, Fu Y, Ching CW, Hawkins RD, Barrera LO, Van Calcar S, Qu C, Ching KA, Wang W, Weng Z, Green RD, Crawford GE, Ren B. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat Genet. 2007;39(3):311–8. doi:10.1038/ng1966.
Zhou Y, Wang L, Vaseghi HR, Liu Z, Lu R, Alimohamadi S, Yin C, Fu JD, Wang GG, Liu J, Qian L. Bmi1 is a key epigenetic barrier to direct cardiac reprogramming. Cell Stem Cell. 2016;18(3):382–95. doi:10.1016/j.stem.2016.02.003.
Cao R, Tsukada Y, Zhang Y. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol Cell. 2005;20(6):845–54. doi:10.1016/j.molcel.2005.12.002.
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, Zhang Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004;431(7010):873–8. doi:10.1038/nature02985.
Yang YJ, Song TY, Park J, Lee J, Lim J, Jang H, Kim YN, Yang JH, Song Y, Choi A, Lee HY, Jo CH, Han JW, Kim ST, Youn HD, Cho EJ. Menin mediates epigenetic regulation via histone H3 lysine 9 methylation. Cell Death Dis. 2013;4:e583. doi:10.1038/cddis.2013.98.
Nakamura T, Mori T, Tada S, Krajewski W, Rozovskaia T, Wassell R, Dubois G, Mazo A, Croce CM, Canaani E. ALL-1 is a histone methyltransferase that assembles a supercomplex of proteins involved in transcriptional regulation. Mol Cell. 2002;10(5):1119–28.
Milne TA, Briggs SD, Brock HW, Martin ME, Gibbs D, Allis CD, Hess JL. MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol Cell. 2002;10(5):1107–17.
Sidler C, Woycicki R, Li D, Wang B, Kovalchuk I, Kovalchuk O. A role for SUV39H1-mediated H3K9 trimethylation in the control of genome stability and senescence in WI38 human diploid lung fibroblasts. Aging (Albany NY). 2014;6(7):545–63. doi:10.18632/aging.100678.
Liu Y, Du J, Zhang J, Weng M, Li X, Pu D, Gao L, Deng S, Xia S, She Q. Snail1 is involved in de novo cardiac fibrosis after myocardial infarction in mice. Acta Biochim Biophys Sin Shanghai. 2012;44(11):902–10. doi:10.1093/abbs/gms085.
Nieto MA. The snail superfamily of zinc-finger transcription factors. Nat Rev Mol Cell Biol. 2002;3(3):155–66. doi:10.1038/nrm757.
Weber KT, Sun Y, Bhattacharya SK, Ahokas RA, Gerling IC. Myofibroblast-mediated mechanisms of pathological remodelling of the heart. Nat Rev Cardiol. 2013;10(1):15–26. doi:10.1038/nrcardio.2012.158.
Zeisberg EM, Tarnavski O, Zeisberg M, Dorfman AL, McMullen JR, Gustafsson E, Chandraker A, Yuan X, Pu WT, Roberts AB, Neilson EG, Sayegh MH, Izumo S, Kalluri R. Endothelial-to-mesenchymal transition contributes to cardiac fibrosis. Nat Med. 2007;13(8):952–61. doi:10.1038/nm1613.
Willems E, Cabral-Teixeira J, Schade D, Cai W, Reeves P, Bushway PJ, Lanier M, Walsh C, Kirchhausen T, Izpisua Belmonte JC, Cashman J, Mercola M. Small molecule-mediated TGF-beta type II receptor degradation promotes cardiomyogenesis in embryonic stem cells. Cell Stem Cell. 2012;11(2):242–52. doi:10.1016/j.stem.2012.04.025.
Ikushima H, Miyazono K. TGFbeta signalling: a complex web in cancer progression. Nat Rev Cancer. 2010;10(6):415–24. doi:10.1038/nrc2853.
Inman GJ, Nicolas FJ, Callahan JF, Harling JD, Gaster LM, Reith AD, Laping NJ, Hill CS. SB-431542 is a potent and specific inhibitor of transforming growth factor-beta superfamily type I activin receptor-like kinase (ALK) receptors ALK4, ALK5, and ALK7. Mol Pharmacol. 2002;62(1):65–74.
Fu JD, Stone NR, Liu L, Spencer CI, Qian L, Hayashi Y, Delgado-Olguin P, Ding S, Bruneau BG, Srivastava D. Direct reprogramming of human fibroblasts toward a cardiomyocyte-like state. Stem Cell Rep. 2013;1(3):235–47. doi:10.1016/j.stemcr.2013.07.005.
Nam YJ, Song K, Luo X, Daniel E, Lambeth K, West K, Hill JA, Dimaio JM, Baker LA, Bassel-Duby R, Olson EN. Reprogramming of human fibroblasts toward a cardiac fate. Proc Natl Acad Sci U S A. 2013;110(14):5588–93. doi:10.1073/pnas.1301019110.
Wada R, Muraoka N, Inagawa K, Yamakawa H, Miyamoto K, Sadahiro T, Umei T, Kaneda R, Suzuki T, Kamiya K, Tohyama S, Yuasa S, Kokaji K, Aeba R, Yozu R, Yamagishi H, Kitamura T, Fukuda K, Ieda M. Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc Natl Acad Sci U S A. 2013;110(31):12667–72. doi:10.1073/pnas.1304053110.
Singh VP, Mathison M, Patel V, Sanagasetti D, Gibson BW, Yang J, Rosengart TK. MiR-590 promotes transdifferentiation of porcine and human fibroblasts toward a cardiomyocyte-like fate by directly repressing specificity protein 1. J Am Heart Assoc. 2016;5(11):e003922. doi:10.1161/JAHA.116.003922.
Sturzu AC, Wu SM. Developmental and regenerative biology of multipotent cardiovascular progenitor cells. Circ Res. 2011;108(3):353–64. doi:10.1161/CIRCRESAHA.110.227066.
Martin-Puig S, Wang Z, Chien KR. Lives of a heart cell: tracing the origins of cardiac progenitors. Cell Stem Cell. 2008;2(4):320–31. doi:10.1016/j.stem.2008.03.010.
Islas JF, Liu Y, Weng KC, Robertson MJ, Zhang S, Prejusa A, Harger J, Tikhomirova D, Chopra M, Iyer D, Mercola M, Oshima RG, Willerson JT, Potaman VN, Schwartz RJ. Transcription factors ETS2 and MESP1 transdifferentiate human dermal fibroblasts into cardiac progenitors. Proc Natl Acad Sci U S A. 2012;109(32):13016–21. doi:10.1073/pnas.1120299109.
Lalit PA, Salick MR, Nelson DO, Squirrell JM, Shafer CM, Patel NG, Saeed I, Schmuck EG, Markandeya YS, Wong R, Lea MR, Eliceiri KW, Hacker TA, Crone WC, Kyba M, Garry DJ, Stewart R, Thomson JA, Downs KM, Lyons GE, Kamp TJ. Lineage reprogramming of fibroblasts into proliferative induced cardiac progenitor cells by defined factors. Cell Stem Cell. 2016;18(3):354–67. doi:10.1016/j.stem.2015.12.001.
Zhang Y, Cao N, Huang Y, Spencer CI, Fu JD, Yu C, Liu K, Nie B, Xu T, Li K, Xu S, Bruneau BG, Srivastava D, Ding S. Expandable cardiovascular progenitor cells reprogrammed from fibroblasts. Cell Stem Cell. 2016;18(3):368–81. doi:10.1016/j.stem.2016.02.001.
Cao N, Huang Y, Zheng J, Spencer CI, Zhang Y, Fu JD, Nie B, Xie M, Zhang M, Wang H, Ma T, Xu T, Shi G, Srivastava D, Ding S. Conversion of human fibroblasts into functional cardiomyocytes by small molecules. Science. 2016;352(6290):1216–20. doi:10.1126/science.aaf1502.
Christoforou N, Chakraborty S, Kirkton RD, Adler AF, Addis RC, Leong KW. Core transcription factors, MicroRNAs, and small molecules drive transdifferentiation of human fibroblasts towards the cardiac cell lineage. Sci Rep. 2017;7:40285. doi:10.1038/srep40285.
Treutlein B, Lee QY, Camp JG, Mall M, Koh W, Shariati SA, Sim S, Neff NF, Skotheim JM, Wernig M, Quake SR. Dissecting direct reprogramming from fibroblast to neuron using single-cell RNA-seq. Nature. 2016;534(7607):391–5. doi:10.1038/nature18323.
Maguire AM, Simonelli F, Pierce EA, Pugh EN Jr, Mingozzi F, Bennicelli J, Banfi S, Marshall KA, Testa F, Surace EM, Rossi S, Lyubarsky A, Arruda VR, Konkle B, Stone E, Sun J, Jacobs J, Dell’Osso L, Hertle R, Ma JX, Redmond TM, Zhu X, Hauck B, Zelenaia O, Shindler KS, Maguire MG, Wright JF, Volpe NJ, McDonnell JW, Auricchio A, High KA, Bennett J. Safety and efficacy of gene transfer for Leber’s congenital amaurosis. N Engl J Med. 2008;358(21):2240–8. doi:10.1056/NEJMoa0802315.
Wong SY, Soto J, Li S. Biophysical regulation of cell reprogramming. Curr Opin Chem Eng. 2017;15:95–101. doi:10.1016/j.coche.2017.01.001.
Kong YP, Carrion B, Singh RK, Putnam AJ. Matrix identity and tractional forces influence indirect cardiac reprogramming. Sci Rep. 2013;3:3474. doi:10.1038/srep03474.
Sia J, Yu P, Srivastava D, Li S. Effect of biophysical cues on reprogramming to cardiomyocytes. Biomaterials. 2016;103:1–11. doi:10.1016/j.biomaterials.2016.06.034.
Morez C, Noseda M, Paiva MA, Belian E, Schneider MD, Stevens MM. Enhanced efficiency of genetic programming toward cardiomyocyte creation through topographical cues. Biomaterials. 2015;70:94–104. doi:10.1016/j.biomaterials.2015.07.063.
Acknowledgement
J. Liu is supported by American Heart Association (AHA) 15GRNT25530005, L. Qian is supported by AHA Scientist Development Grant 13SDG17060010, the Ellison Medical Foundation (EMF) New Scholar Grant AG-NS-1064-13, and NIH/NHLBI R01HL128331.
Compliance with Ethics Guidelines
Li Wang, Jiandong Liu, and Li Qian declare that they have no conflict of interest. All institutional and national guidelines for the care and use of laboratory animals were followed.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Wang, L., Liu, J., Qian, L. (2017). In Vivo Lineage Reprogramming of Fibroblasts to Cardiomyocytes for Heart Regeneration. In: Yilmazer, A. (eds) In Vivo Reprogramming in Regenerative Medicine. Stem Cell Biology and Regenerative Medicine. Humana Press, Cham. https://doi.org/10.1007/978-3-319-65720-2_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-65720-2_4
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-65719-6
Online ISBN: 978-3-319-65720-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)