Abstract
Plant transformation technology offers unique prospects to transfer a wide spectrum of functionally relevant genes in plants. Expression of genes is regulated by a number of factors among which promoter strength, specificity, and cis- and trans-acting elements play a critical role. The choice of promoter is a key determinant for the levels and specificities of gene expression. In sugarcane, the maize ubiquitin promoter has been the workhorse promoter for decades. The availability of limited promoters for sugarcane transformation is critical in sugarcane crop improvement through genetic engineering. However, recent advancements in biotechnology have provided greater insights into promoter validation from wild and commercially cultivated sugarcane, which is evident from an array of different promoters reported. This review describes the various promoters isolated from sugarcane and its wild relatives that would benefit future genetic engineering studies in sugarcane. In addition, the challenges ahead and improved strategies for sugarcane transformation are discussed.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
6.1 Introduction
Sugarcane (Saccharum spp. hybrid) belonging to Poaceae family is an economically important food and energy crop grown worldwide. Large genome size, polyploidy, low fertility, complex environmental interactions, slow breeding advances, and nobilization hinder the breeding for this crop. In addition, several issues like low cane and sugar yields; susceptibility to abiotic stresses such as drought, cold, and salinity; and biotic stresses such as pest insects and fungal diseases are the major constraints in sugarcane cultivation (Tiwari et al. 2010). Transgenic technology provides an effective tool for sugarcane crop improvement. Both biolistic and Agrobacterium-mediated transformation methods have been well established and are widely used to develop transgenic sugarcane. Several factors have to be considered in the development of transgenic sugarcane among which the choice of promoter plays a crucial role.
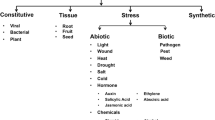
Promoters are the regulatory sequences present upstream of the genes and are involved in the regulation of the gene expression. In recent years, an array of promoters from plant and viral origin have been characterized and extensively used in transgene expression in plants. They can be broadly grouped into constitutive, spatiotemporal, and inducible promoters based on their activity. The CaMV 35S promoter has been well described and widely used for constitutive transgene expression (Potenza et al. 2004). However, in monocots, the CaMV 35S promoter confers lower levels of transgene expression. The maize ubiquitin promoter (M-ubi1) is the promoter of choice for sugarcane transformation and has been widely used for over two decades. Sugarcane ubiquitin promoters (Ubi4 and Ubi9) when expressed in sugarcane have led to posttranscriptional gene silencing (PTGS) (Wei et al. 2003). The promoters used currently for the development of transgenic sugarcane are limited in number and very few provide tissue-specific expression. Limited tissue-specific promoters have been characterized so far for sugarcane transformation (Damaj et al. 2010). Hence, there is a need for identification of more promoters for specific applications, and from unrelated sources, which would be of great value for future genetic engineering studies in sugarcane. This review discusses the various advancements that took place in the past decade pertaining to sugarcane transformation and promoter validation which will benefit researchers aiming to develop transgenic sugarcane with desirable traits.
6.2 Plant Promoters: Structure and Function
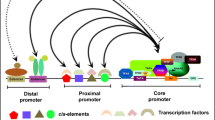
Promoters are defined as regions upstream of a gene’s coding region and are involved in the regulation of the frequency of transcription. They usually contain specific DNA sequences and regulatory elements and are the key regulators of transcription, also called as molecular switches. Promoters can be broadly classified into constitutive, tissue specific, and inducible based on their activity. Another type of promoters which are currently of importance is the synthetic promoters that combine the available core promoters with different motifs and are designed for specific expression. A typical plant promoter is composed of a transcription start site (TSS) , the core promoter region, the proximal region (or upstream regulatory elements), and the distal regulatory region (or long-range regulatory elements). Plant promoters and their cis-acting regulatory elements have been reviewed extensively (Hernandez-Garcia and Finer 2014; Porto et al. 2014; Grunennvaldt et al. 2015; Shah et al. 2015; Naqvi et al. 2016; Biłas et al. 2016) and hence are not focused in this review.
6.3 Promoters for Enhanced Transgene Expression in Sugarcane
In sugarcane , the maize ubiquitin (Zmubi1) promoter is being used worldwide for more than two decades for genetic engineering. However, it has failed to drive sustained transgene expression throughout the sugarcane growth cycle (Wang et al. 2005). Promoters of viral origin such as sugarcane bacilliform virus promoter (Braithwaite et al. 2004), banana streak virus promoter (Schenk et al. 1999), and CaMV35S:Zmubi1 tandem promoter (Groenewald and Botha 2008) conferred enhanced expression in mature canes. Some of the other promoters used were the enhanced maize ubiquitin promoter and maize carboxylase promoter (Kinkema et al. 2014a, b) which conferred enhanced transgene expression than the Zmubi1 promoter (five and fourfold). The advent of novel tools for promoter discovery, next-generation sequencing boom, and advanced bioinformatics techniques have led to isolation of new promoters from sugarcane and its wild relatives that could drive enhanced transgene expression than the routine promoters. Table 6.1 lists the different promoters characterized from sugarcane and its wild relatives that would enable researchers to develop GM sugarcane with enhanced transgene expression. Yet, the number of promoters is very limited when compared to other plant species. Mudge et al. (2013) characterized three promoters from sugarcane which conferred preferential transgene expression in mature stems and thus have practical application in sucrose-targeted metabolic engineering.
6.4 Challenges Ahead in Sugarcane Biotechnology
Sugarcane is affected by several biotic and biotic stresses which lead to losses in productivity. Using conventional breeding practices, the release of a new variety typically takes 12–15 years after rigorous testing of performance, sugar content, agronomic traits, and genetic stability (Gazaffi et al. 2010). With the advent of transgenic technology, considerable progress has been made in the recent years. Biolistic and Agrobacterium methods of transformation, though routinely used, have several disadvantages. Biolistic bombardment method usually generates multiple transgene integration sites. Agrobacterium-mediated transformation is rather a time-consuming laborious process, has low transformation efficiency (Joyce et al. 2010), has high variability between experiments, and is genotype dependent (Anderson and Birch 2012). In fact, the time taken from DNA delivery till whole-plant regeneration is longer than any other crop plants.
Another major challenge in sugarcane transformation is the transgene inactivation/silencing problem. Several promoters failed to drive transgene expression in mature canes despite showing activity in callus (Wei et al. 2003). Moreover both transcriptional and posttranscriptional gene silencing has been reported in sugarcane (Ingelbrecht et al. 1999). Mudge et al. (2009) characterized eight distinct promoters of MYB family of which three were expressive. Interestingly, their results illustrated that multiple copies of promoter do not trigger silencing and polyploids may have intrinsic silencing mechanisms that are yet to be deciphered. Birch et al. (2010) reported that silencing in sugarcane is 5′-sequence specific, independent of copy number, developmentally regulated, and posttranscriptional in T0 transgenic lines. Transgenes fused with strong tissue-specific promoters may alleviate the silencing problem .
Sugarcane genome is about 10 Gb size with homologous genes ranging from 8 to 12 copies (Souza et al. 2011) and the monoploid genome size being 750–930 Mb (D’Hont and Glaszmann 2001). Currently there is a lack of whole genomic data in sugarcane. The major factors that make the whole-genome sequencing of sugarcane difficult are (1) polyploidy—80% of sugarcane genome is inherited from S. officinarum and 10% from S. spontaneum; (2) high level of recombination—more than 10% of sugarcane genome is mosaic and unknown; (3) heterozygosity—leads to variations that deter genome assembly; and (4) repeats—high number of repetitive sequences present throughout the genome . Moreover, sugarcane lacks diploid progenitors that aid in a faster and easier genome assembly (Garcia et al. 2013) unlike banana (D’hont et al. 2012). In addition, it is difficult to employ shot-gun sequencing such as Illumina which generates shorter reads. Thus, the large and complex genome, high ploidy levels, and high content of repetitive DNA make sugarcane an unusually recalcitrant crop species for both forward and reverse genetic studies.
6.5 Improved Strategies for Sugarcane Transformation
Several researchers are striving hard to overcome the challenges in sugarcane transformation which has led to a breakthrough with an array of advanced techniques, modified protocols, and strategies for efficient sugarcane transformation. This section of the review discusses some of the significant studies which will have a greater impact on sugarcane transgenic research in the near future. Joyce et al. (2010) optimized different parameters for Agrobacterium transformation in sugarcane and observed that selection and cocultivation systems were critical factors that affected sugarcane transformation. Jackson et al. (2013) compared both the methods using whole plasmids and minimal cassettes and observed that both the procedures were high expressing and yielded single-gene insertions at a reasonable transformation efficiency (TE).
Taparia et al. (2012a, b) used minimal expression cassettes for biolistic gene transfer and with reduced plasmid concentration and achieved simple transgene integration and stable transgene expression . They also described a rapid transformation procedure that only needs 3 months from culture initiation to potting of transgenic sugarcane. Use of minimal cassettes has shown to be effective since they are devoid of prokaryotic backbone sequences that may contribute to recombination or induce methylation, thereby leading to transgene silencing .
Anderson and Birch (2012) studied several parameters that are critical for transformation of sugarcane variety Q117. They reported that the key factors influencing transformation efficiency in Agrobacterium method were minimal handling of callus during cocultivation and the use of a super-binary vector in AGL Agrobacterium strain which led to the highest transformation efficiency reported so far for Agrobacterium-mediated transformation in sugarcane. Recently, Mayavan et al. (2015) have developed a rapid, efficient, and genotype-independent in planta transformation protocol using sugarcane setts as explants. They have claimed a maximum of 32.6% TE which is so far the highest TE in sugarcane. Their group had earlier developed a seed-based transformation protocol which also proved to be efficient to develop transgenic sugarcane in a shorter duration (Mayavan et al. 2013).
Dong et al. (2014) developed a robust protocol that could be applied on a larger industrial scale for sugarcane improvement through genetic engineering. This protocol employs desiccation during cocultivation that leads to higher TE and has also been tested in several varieties and in several laboratories proving its versatility. In addition, the transgenes were stable across multiple generations and growing seasons that further proves the great utility of the protocol. Sandhu et al. (2016) have recently reported single-step direct transgenic plant regeneration from agro-infected spindle leaf roll segments of sugarcane with a very short period of 8 weeks since it avoids the callusing phase. Stable integration was observed in the transgenics making the protocol reliable for sugarcane transformation.
Jackson et al. (2014) presented a set of rules to achieve sustained transgene expression and validated them in sugarcane. They used the following methods independently or in combination—removal of rare codons, removal of RNA instability sequences, blocking of putative endogenous sRNA-binding sites, and randomization of non-rare codons. This technique can be applied in sugarcane effectively to alleviate transgene silencing . Recently, Lowe et al. (2016) reported an efficient monocot transformation strategy wherein they over-expressed the maize morphogenic regulators Baby boom (Bbm) and Wuschel2 (Wus2) genes in previously non-transformable maize inbred lines and achieved high transformation frequencies. They also successfully employed this approach to enhance transformation frequency in sorghum, sugarcane, and rice.
Other notable advances worth mentioning are (1) development of synthetic reporter genes in order to alleviate silencing effects to validate promoter expression in sugarcane (Chou and Moyle 2014); (2) use of alternate monocot models such as Setaria viridis that yields higher transformation efficiency, has shorter duration, and contains a similar cell wall composition to that of sugarcane. Hence it can be used as an alternate model plant for sugarcane-applied research for stress resistance, improved biomass , and bioethanol production (Martins et al. 2015); (3) use of novel promoters that drive higher levels of transgene expression than the routine promoters and exploiting codon-optimized target genes specific for sugarcane to enhance transgene expression (Kinkema et al. 2014a); (4) application of RNAi technology to develop improved sugarcane for desired traits (Gan et al. 2010; Gao et al. 2013; Jung et al. 2012); (5) use of a combinatorial approach wherein multiple promoters/enhancers/terminators/5′UTRs are employed to achieve higher transgene expression (unpublished data); and (6) use of systems biology and metabolic modeling approach to unravel gene regulatory networks underlying key mechanisms such as sucrose synthesis and accumulation .
6.6 Conclusion and Future Perspectives
Sugarcane biotechnology has advanced rapidly over the years and transgenic lines for various biotic and abiotic stresses have been developed and are being tested in laboratories worldwide. Commercial testing of transgenic sugarcane has already been approved in Indonesia and is in pipeline in several other countries. Several recombinant proteins have already been produced using sugarcane as a bio-factory. With the advances in transgenic technology, genome sequencing tools, and systems biology coupled with bioinformatics, it is now feasible to manipulate the metabolic pathways in sugarcane, thereby enhancing the crop productivity and increased sugar content. Although obstacles including transgene inactivation, lack of whole genome, and long duration for transformation are certainly a hindrance, genetic engineering combined with the novel advanced strategies would undoubtedly be instrumental in helping the sugarcane industries develop into a stronger bio-economy.
References
Abraha TG (2005) Isolation and characterization of a culm-specific promoter element from sugarcane. MSc Dissertation, Department of Botany and Zoology, Institute of Biotechnology, Stellenbosch University, South Africa
Anderson DJ, Birch RG (2012) Minimal handling and super-binary vectors facilitate efficient, Agrobacterium-mediated, transformation of sugarcane (Saccharum spp. hybrid). Trop Plant Biol 5(2):183–192
Biłas R, Szafran K, Hnatuszko-Konka K, Kononowicz AK (2016) Cis-regulatory elements used to control gene expression in plants. Plant Cell Tiss Organ Cult 127:269–287. doi:10.1007/s11240-016-1057-7
Birch RG, Bower RS, Elliott AR (2010) Highly efficient, 5′-sequence-specific transgene silencing in a complex polyploid. Trop Plant Biol 3(2):88–97
Braithwaite KS, Geijskes RJ, Smith GR (2004) A variable region of the sugarcane bacilliform virus (SCBV) genome can be used to generate promoters for transgene expression in sugarcane. Plant Cell Rep 23:319–326
Chakravarthi M, Philip A, Subramonian N (2015) Truncated ubiquitin 5′ regulatory region from Erianthus arundinaceus drives enhanced transgene expression in heterologous systems. Mol Biotechnol 57(9):820–835
Chakravarthi M, Syamaladevi DP, Harunipriya P, Augustine SM, Subramonian N (2016) A novel PR10 promoter from Erianthus arundinaceus directs high constitutive transgene expression and is enhanced upon wounding in heterologous plant systems. Mol Biol Rep 43(1):17–30
Chou TC, Moyle RL (2014) Synthetic versions of firefly luciferase and Renilla luciferase reporter genes that resist transgene silencing in sugarcane. BMC Plant Biol 14(1):92
D’Hont A, Glaszmann JC (2001) Sugarcane genome analysis with molecular markers, a first decade of research. Proc Int Soc Sug Cane Tech 24:556–559
D’hont A, Denoeud F, Aury JM, Baurens FC, Carreel F et al (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488:213–217
Damaj MB, Kumpatla SP, Emani C, Beremand PD, Reddy AS, Rathore KS, Buenrostro-Nava MT, Curtis IS, Thomas TL, Mirkov TE (2010) Sugarcane DIRIGENT and O-METHYLTRANSFERASE promoters confer stem-regulated gene expression in diverse monocots. Planta 231(6):1439–1458
Dong S, Delucca P, Geijskes RJ, Ke J, Mayo K, Mai P, Sainz M, Caffall K, Moser T, Yarnall M, Setliff K (2014) Advances in Agrobacterium-mediated sugarcane transformation and stable transgene expression. Sugar Tech 16(4):366–371
Gan D, Zhang J, Jiang H, Jiang T, Zhu S, Cheng B (2010) Bacterially expressed dsRNA protects maize against SCMV infection. Plant Cell Rep 29(11):1261–1268
Gao SJ, Damaj MB, Park JW, Beyene G, Buenrostro-Nava MT, Molina J, Wang X, Ciomperlik JJ, Manabayeva SA, Alvarado VY, Rathore KS (2013) Enhanced transgene expression in sugarcane by co-expression of virus-encoded RNA silencing suppressors. PLoS One 8(6):e66046
Garcia AA, Mollinari M, Marconi TG, Serang OR, Silva RR et al (2013) SNP genotyping allows an in depth characterisation of the genome of sugarcane and other complex autopolyploids. Sci Rep 3:3399
Gazaffi R, Oliveira KM, Souza AP, Garcia AAF (2010) The importance of the germplasm in developing agro-energetic profile sugarcane cultivars. In: Cortez LAB (ed) Sugar cane bioethanol: R&D for productivity and sustainability. Blucher, São Paulo, pp 333–343. ISBN: 9788521205302
Groenewald JH, Botha FC (2008) Down-regulation of pyrophosphate: fructose 6-phosphate 1-phosphotransferase (PFP) activity in sugarcane enhances sucrose accumulation in immature internodes. Trans Res 17(1):85–92
Grunennvaldt RL, Goldbach JD, Grhardt IR, Quoirin M (2015) Promoters used in genetic transformation of plants. Res J Biol Sci 10(1–2):1–9
Hernandez-Garcia CM, Finer JJ (2014) Identification and validation of promoters and cis-acting regulatory elements. Plant Sci 217:109–119
Ingelbrecht IL, Irvine JE, Mirkov TE (1999) Posttranscriptional gene silencing in transgenic sugarcane dissection of homology dependent virus resistance in a monocot that has a complex polyploid genome. Plant Physiol 119:1187–1197
Jackson MA, Anderson DJ, Birch RG (2013) Comparison of Agrobacterium and particle bombardment using whole plasmid or minimal cassette for production of high-expressing low-copy transgenic plants. Trans Res 22(1):143–151
Jackson MA, Sternes PR, Mudge SR, Graham MW, Birch RG (2014) Design rules for efficient transgene expression in plants. Plant Biotechnol J 12(7):925–933
Joyce P, Kuwahata M, Turner N, Lakshmanan P (2010) Selection system and co-cultivation medium are important determinants of Agrobacterium-mediated transformation of sugarcane. Plant Cell Rep 29(2):173–183
Jung JH, Fouad WM, Vermerris W, Gallo M, Altpeter F (2012) RNAi suppression of lignin biosynthesis in sugarcane reduces recalcitrance for biofuel production from lignocellulosic biomass. Plant Biotechnol J 10(9):1067–1076
Kharte SB, Watharkar AD, Shingote PR, Chandrashekharan S, Pagariya MC, Kawar PG, Govindwar SP (2016) Functional characterization and expression study of sugarcane MYB transcription factor gene PEaMYBAS1 promoter from Erianthus arundinaceus that confers abiotic stress tolerance in tobacco. RSC Adv 6(23):19576–19586
Kinkema M, Geijskes J, Palupe A, Shand K, Coleman HD, Brinin A, Williams B, Sainz M, Dale JL (2014a) Improved molecular tools for sugar cane biotechnology. Plant Mol Biol 84(4–5):497–508
Kinkema M, Geijskes RJ, Shand K, Coleman HD, De Lucca PC, Palupe A et al (2014b) An improved chemically inducible gene switch that functions in the monocotyledonous plant sugar cane. Plant Mol Biol 84(4–5):443–454
Li HJ, Qi NJ, Hui Z, Tao YL, Rui LY, Qin WA (2013) Cloning and sequence analysis of promoter in sugarcane ethylene receptor gene (Sc-ERS). J South Agric 44(5):722–729
Lowe K, Wu E, Wang N, Hoerster G, Hastings C, Cho MJ et al (2016) Morphogenic regulators baby boom and Wuschel improve monocot transformation. Plant Cell 28(9):1998–2015
Martins PK, Ribeiro AP, da Cunha BADB, Kobayashi AK, Molinari HBC (2015) A simple and highly efficient Agrobacterium-mediated transformation protocol for Setaria viridis. Biotechnol Rep 6:41–44
Mayavan S, Subramanyam K, Arun M, Rajesh M, Dev GK, Sivanandhan G, Jaganath B, Manickavasagam M, Selvaraj N, Ganapathi A (2013) Agrobacterium tumefaciens-mediated in planta seed transformation strategy in sugarcane. Plant Cell Rep 32(10):1557–1574
Mayavan S, Subramanyam K, Jaganath B, Sathish D, Manickavasagam M, Ganapathi A (2015) Agrobacterium-mediated in planta genetic transformation of sugarcane setts. Plant Cell Rep 34(10):1835–1848
Moyle RL, Birch RG (2013) Sugarcane loading stem gene promoters drive transgene expression preferentially in the stem. Plant Mol Biol 82(1–2):51–58
Mudge SR, Osabe K, Casu RE, Bonnett GD, Manners JM, Birch RG (2009) Efficient silencing of reporter transgenes coupled to known functional promoters in sugarcane a highly polyploid crop species. Planta 229(3):549–558
Mudge SR, Basnayake SW, Moyle RL, Osabe K, Graham MW, Morgan TE, Birch RG (2013) Mature-stem expression of a silencing-resistant sucrose isomerase gene drives isomaltulose accumulation to high levels in sugarcane. Plant Biotechnol J 11(4):502–509
Naqvi RZ, Mubeen H, Raza S (2016) Role of plant promoters and their cis regulatory elements in gene expression regulation. Eur J Pharm Med Res 3(1):347–352
Niu JQ, Wang AQ, Huang JL, Yang LT, Li YR (2015) Isolation characterization and promoter analysis of cell wall invertase gene SoCIN1 from sugarcane (Saccharum spp). Sugar Tech 17(1):65–76
Porto MS, Pinheiro MPN, Batista VGL, dos Santos RC, de Albuquerque Melo Filho P, de Lima LM (2014) Plant promoters: an approach of structure and function. Mol Biotechnol 56(1):38–49
Potenza C, Aleman L, Sengupta-Gopalan C (2004) Targeting transgene expression in research agricultural and environmental applications: promoters used in plant transformation. In Vitro Cell Dev Biol Plant 40(1):1–22
Prabu G, Prasad DT (2012) Functional characterization of sugarcane MYB transcription factor gene promoter (PScMYBAS1) in response to abiotic stresses and hormones. Plant Cell Rep 31(4):661–669
Sandhu JS, Kaur M, Kaur A, Kalia A (2016) Single step direct transgenic plant regeneration from adventive embryos of agro-infected sugarcane (Saccharum spp.) spindle leaf roll segments with assured genetic fidelity. Plant Cell Tissue Org Cult 125(1):149–162
Schenk PM, Sagi L, Remans T, Dietzgen RG, Bernard MJ, Graham MW, Manners JM (1999) A promoter from sugarcane bacilliform badnavirus drives transgene expression in banana and other monocot and dicot plants. Plant Mol Biol 39:1221–1230
Shah SH, Jan SA, Ahmad N, Khan SU, Kumar T, Iqbal A, Nasir F (2015) Use of different promoters in transgenic plant development: current challenges and future perspectives. Am Eurasian J Agric Environ Sci 15:664–675
Souza GM, Berges H, Bocs S, Casu R, D’Hont A, Ferreira JE et al (2011) The sugarcane genome challenge: strategies for sequencing a highly complex genome. Trop Plant Biol 4:145–156
Taparia Y, Gallo M, Altpeter F (2012a) Comparison of direct and indirect embryogenesis protocols, biolistic gene transfer and selection parameters for efficient genetic transformation of sugarcane. Plant Cell Tissue Organ Cult 111(2):131–141
Taparia Y, Fouad WM, Gallo M, Altpeter F (2012b) Rapid production of transgenic sugarcane with the introduction of simple loci following biolistic transfer of a minimal expression cassette and direct embryogenesis. In Vitro Cell Dev Biol Plant 48(1):15–22
Tiwari AK, Bharti YP, Tripathi S, Mishra N, Lal M, Rao GP, Sharma PK, Sharma ML (2010) Biotechnological approaches to improve sugarcane crop with special reference to disease resistance. Acta Phytopath et Entom Hung 45(2):235–249
Wang ML, Goldstein C, Su W, Moore PH, Albert HH (2005) Production of biologically active GM-CSF in sugarcane: a secure biofactory. Trans Res 14(2):167–178
Wei H, Wang ML, Moore PH, Albert HH (2003) Comparative expression analysis of two sugarcane polyubiquitin promoters and flanking sequences in transgenic plants. J Plant Physiol 160(10):1241–1251
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Mohan, C., Schneider, V.K., Henrique-Silva, F. (2017). Novel Potential Candidate Promoters and Advanced Strategies for Sugarcane Transformation. In: Mohan, C. (eds) Sugarcane Biotechnology: Challenges and Prospects. Springer, Cham. https://doi.org/10.1007/978-3-319-58946-6_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-58946-6_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-58945-9
Online ISBN: 978-3-319-58946-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)