Abstract
Voltage-gated calcium (Cav) channels are miniature membrane transistors that convert membrane electrical signals to intracellular Ca2+ transients that trigger many physiological events. In mammals, there are ten subtypes of Cav channel, among which Cav1.1 is the first Cavα1 to be cloned. Cav1.1 is specified for the excitation–contraction coupling of skeletal muscles, and has been a prototype in the structural investigations of Cav channels. This article summarized the recent advances in the structural elucidation of Cav1.1 and the mechanistic insights derived from the 3.6 Å structure obtained using single-particle, electron cryomicroscopy. The structure of the Cav1.1 complex established the framework for mechanistic understanding of excitation–contraction coupling and provides the template for molecular interpretations of the functions and disease mechanisms of Cav and Nav channels.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Voltage-gated calcium (Cav) channels are a large family of membrane proteins that are activated upon the change of membrane potentials. The Ca2+ ions permeated by the Cav channel act as second messengers that trigger a cascade of cellular events involved in a multitude of physiological processes such as muscle contraction, synaptic transmission, hormone secretion, gene expression and cell death (Fig. 2.1) [1, 2].
1.1 Classification of Voltage-Gated Calcium Channels
Cav channels exhibit tissue specificity. Ten subtypes of Cav channels have been identified in mammals through the voltage-clamp measurements of Ca2+ currents in distinct tissues and organisms (Table 2.1) [3, 4]. The ten subtypes can be divided into three subfamilies based on their sequence similarity. The Cav1 channels activate at high voltage and conduct large and long lasting ion currents, thus designated as the L-type calcium channels [5]. The Cav1 channels can be specifically inhibited by dihydropyridine and its derivatives. They are thereby also named as the dihydropyridine receptor (DHPR) [6, 7]. In contrast, the Cav3 members are designated as T-type calcium channel for activation at low voltage and conduct tiny single channel current, and the current is transient [4, 8]. The Cav2 channels can be further divided to the P/Q-type, the N-type, and the R-type based on their Ca2+ current properties and inhibition by specific toxins [4, 9,10,11].
1.2 Molecular Properties of Cav
Cav channels belong to the voltage-gated ion channel superfamily, which has a conserved core fold. Unlike their prokaryotic homologues, which are homotetramers, the ion conducting α1 subunit of eukaryotic Cav channels is a single peptide chain with four similar repeats linked by intracellular loops [12]. Each repeat contains six transmembrane helices named S1–S6, among which S1–S4 of each repeat form the voltage sensing domain (VSD) and S5–S6 of the four repeats constitute the ion conducing pore (Fig. 2.2) [14].
Topology of Cav channels. Cav channels contain several subunits: the ion permeation subunit α1, the extracellular α2δ subunits, the cytosolic β subunit and sometimes the transmembrane γ subunit. The topology shown here is of Cav1.1. The upper right pannel is adapted from Wu et al. [13]
Cav1 and Cav2 channels also contain several auxiliary subunits that form a complex with the α1 subunit (Fig. 2.2). The auxiliary subunits include the extracellular α2δ subunit, the cytosolic β subunit and sometimes the transmembrane γ subunit [1]. The auxiliary subunits are not essential for the channel permeation, but they can regulate the membrane trafficking and channel properties of the α1 subunit. For example, the α2δ subunit can facilitate channel translocation to the cell surface, and the β subunit modulates the kinetic properties of the channel [15,16,17]. In skeletal muscle, the β subunit in Cav1.1 plays an essential role in the excitation-contraction coupling [18,19,20]. The γ subunit assists channel inactivation [21, 22].
Not only α subunit has different subtypes, each auxiliary subunit may have distinct subtypes and splicing isoforms. Four genes encoding the β subunit have been identified, and each can yield different isoforms by alternative splicing [16, 23]. The α2 and δ subunits result from proteolytic cleavage of a single gene product. Besides, four genes have been reported encoding the α2δ subunit [24]. Combinations of the distinct α1 subtypes with various auxiliary subunits give rise to the diversity of Cav channels.
1.3 Channelopathies
Voltage-gated calcium channels play essential roles in various physiological processes, and their disorders will trigger serious diseases, such as hypokalemic periodic paralysis (Cav1.1 related), cardiac arrhythmia (Cav1.2 related), autism spectrum disorder (Cav1.2 related), stationary night blindness (Cav1.4), cerebellar ataxia (Cav2.2) [1, 25].
Cav channels represent important drug targets. The α1 subunits are directly targeted by various natural toxins and clinical drugs that treat hypertension, arrhythmia, and angina pectoris. The α2δ-1 subunit is targeted by the gabapentinoid drugs gabapentin and pregabalin [26, 27].
2 Structural Studies of Voltage-Gated Calcium Channels
Because of the physiological and pathological importance, the structural studies of Cav channels have been pursued for decades. However, the long sequence of the single peptide chain with various intracellular linkers makes the channels resistant to recombinant overexpression, nor to say crystallization. Besides, the existence of the auxiliary subunits and various post-translational modifications such as glycosylation add further challenges to obtain a crystal structure of the Cav channel complex. Despite extensive efforts, the structural information of eukaryotic Cav channels have remained elusive. There were only low resolution cryo-EM maps reported (Fig. 2.3a) [28, 30,31,32]. High resolution crystal structures were obtained for the β1 subunit alone and in complex with the α1-interacting domain (AID) motif from the α1 subunit (Fig. 2.3b) [29, 33, 34].
Prokaryotic homologues of voltage-gated calcium or sodium channel are homotetramers, hence relatively easier than the eukaryotic ones for structural pursuit using X-ray crystallography. In the past several years, researchers have been focusing on the structural characterizations of the prokaryotic homologues of Cav or Nav channel. A few crystal structures of bacterial Nav channels were obtained, including NavAb [35], NavRh [36] and NavMs [37] since 2011. Moreover, the crystal structure of a NavAb variant that exhibits calcium selectivity, hence named CavAb, was determined [38]. These structures represent imperative steps towards molecular understanding of the Nav/Cav channels. However, there are still many questions remaining to be answered, including the most straightforward questions like how the four repeats of α1 subunit are organized spatially and how the auxiliary subunits interact with the α1 subunit. A high resolution structure of a eukaryotic Cav channel is required to address these questions.
The recent technological breakthrough in electron microscopy (EM), including the development of direct electron detector for image acquisition and new algorithms for data processing, provided an unprecedented opportunity to solve structures of challenging targets that were nearly insurmountable by X-ray crystallography. Structural biology by single-particle cryo-EM, used to be nick-named ‘blob-ology’, has been used to solve macromolecular structures at resolutions that can resolve side chains. Compared to X-ray crystallography, cryo-EM has two obvious advantages: (1) only several microliters of sample solutions are required, and (2) crystallization is spared. Cryo-EM has become a prevailing approach for structural biology of membrane proteins and macromolecular machineries since 2013 [39, 40]. The eukaryotic Cav channels have molecular weights beyond 300 kDa with the auxiliary subunits, representing suitable targets for cryo-EM.
Cav1.1 is the first Cavα1 to be cloned, and it has been a prototype in the functional, structural, and mechanistic investigations of Cav channels [12, 41]. There is only one isoform of β subunit, β1a, in skeletal muscle [42], and the Cav1.1 complex is exclusively expressed in skeletal muscle with relatively high abundance. It thereby may be easier to purify the Cav1.1 complex to homogeneity for cryo-EM imaging than for other Cav channels from other tissues. Several purification methods for Cav1.1 from rabbit skeletal muscle have been reported in literature [43,44,45]. Recently, a revised strategy for purification of Cav1.1 from rabbit skeletal muscle membrane was developed, using GST-fused β1a subunit to compete with the endogenous β1a subunit to pull down the whole complex [46]. This strategy is simple and more specific. The protein obtained using this protocol is of high purity and well behaved, representing an ideal target for single particle cryo-EM analysis.
In 2015, the cryo-EM structure of rabbit Cav1.1 was obtained at 4.2 Å resolution [46]. Most of the secondary structure elements of the protein complex were revealed in this structure. All the subunits were identified and the arrangement of the four repeats in α1 subunit has been revealed. In the following year, the resolution of the Cav1.1 structure was further improved to 3.6 Å resolution, which allowed assignment of most side chains [13]. This is the first near atomic structure of a eukaryotic Cav channels. The high resolution structure provides important insights into the understanding of the structure-function relationship of Cav and the related Nav channels.
3 Structural Analysis of Cav1.1
3.1 The Overall Architecture of Cav1.1
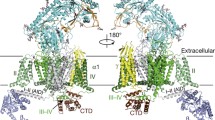
The structure of Cav1.1 is constituted by the ion-permeating core subunit α1, the extracellular α2δ subunit, the intracellular β1a subunit, and the transmembrane γ subunit. The overall structure is approximately 170 Å in height and 100 Å in the longest dimension of the width (Fig. 2.4a).
Structure of the Cav1.1 channel complex from rabbit. (a) Overall structure of Cav1.1. The structure is color-coded for different subunits. The four repeats of the α1 subunit are colored from white to dark green and CTD is colored brown. (b) Spatial arrangement of α1 subunit. The four repeats are arranged clockwisely in the extracellular view. Adapted from Wu et al. [13]
3.2 Structure of the α1 Subunit
The α1 subunit in eukaryotic Cav channels is a single peptide chain that folds into four similar repeats. Therefore, an evident and interesting question arose: how are the four homologous repeats arranged? Clockwise, counterclockwise, or intertwined?
Careful examination of the 4.2 Å cryo-EM reconstruction suggested that the four repeats are arranged in a clockwise manner in the extracellular view, which was then unambiguously confirmed by the 3.6 Å cryo-EM map (Fig. 2.4b). This repeat organization principle may be conserved in all eukaryotic Cav or Nav channels.
3.3 The VSDs of Cav1.1
The VSDs are responsible for voltage sensing. Among the four transmembrane segments S1–S4 in each VSD, S4 contains repetitively aligned basic residues Arg or Lys that occur every three residues. They are named the “gating charges”. The four VSDs of Cav1.1 are similar but non-identical with each other. The S3 segments of prokaryotic homologues NavAb and NavRh are largely unfolded on the extracellular side, whereas the S3 segments of Cav1.1 are full transmembrane helices.
According to the sequence alignment of Cav1.1 and ten human Cav channels, there are up to six gating charge residues on each S4 segment, defined as R1–R6 (Fig. 2.5a). For the rabbit Cav1.1, there are R1–R5 on S4I, R2–R6 on S4II, R1–R5 on S4III, and R2–R5 on S4IV.
VSDIV of Cav1.1. (a) Sequence alignment of S4IV helix. The conserved R/K highlighted blue. (b) Structure of the VSDIV of Cav1.1. (c) Structural superimposition of the four VSDs of Cav1.1. Adapted from Wu et al. [13]
In each VSD, all the gating charges are aligned on one side of the 310 helix of S4. The R5 residues and R6II are below, whereas the R1–R4 residues are above the conserved occluding Phe in the charge transfer centre, representing the depolarized or ‘up’ conformation of VSDs (Fig. 2.5b). Considering the closed pore and the ‘up’ VSDs, the structure of Cav1.1 shown here may represent a potentially inactivated state. The R1–R4 residues are above the conserved occluding Phe in the charge transfer center, representing the depolarized or up conformation of VSDs. This is consistent with the sample conditions of 0 mV membrane potential (depolarized state).
3.4 The Selectivity Filter
The selectivity filter (SF) is responsible for ion selection. In Cav1.1, the SF is constituted by an outer site comprising the side chains of four essential Glu residues and an inner site formed by the carbonyl oxygen of the two preceding residues.
The SF of different voltage gated ion channels exhibits distinct chemical and structural features. The selectivity filter of KV channels is constituted by the carbonyl oxygen of the residues. But in Cav and Nav channels, the side chains of the outer site residues are important for selectivity. In Nav channels, the key residues are Asp/Glu/Lys/Ala (DEKA).
The SF of rCav1.1 is constituted by the side groups of Glu292/614/1014/1323 and the two preceding residues in each repeat that contribute the carbonyl oxygen (C=O). A consecutive stretch of density stands along the selectivity filter vestibule that can be deconvoluted to a round disc in the centre of the four Glu residues and a sphere surrounded by the eight C=O groups. Two Ca2+ ions are tentatively assigned to the middle disc and the inner sphere. The two maps (3.6 Å and 4.2 Å) were from two different batches of protein which was purified in 10 mM and 0.5 mM Ca2+, respectively. The shape and position of the density in the selectivity filter vestibule in the two maps are different (Fig. 2.6a). The heights of the two Ca2+ ions in the 3.6 Å map are similar to those in CavAb [38], except that the inner one is slightly off the central axis and closer to repeats I and II.
Selectivity filter. (a) Structures of the selectivity filter obtained from two different maps. (b) The permeation path of the pore domain. Adapted from Wu et al. [13]
Above the entrance to the Ca2+ permeation path, the extended extracellular loops, which are stabilized by many disulfide bonds, form a windowed dome. The window is enriched in negatively charged residues, which may help attract cations. Below the SF is a hydrophobic cavity with fenestrations. The radius in the inner gate is less than 1 Å too narrow to permeate Ca2+ ions (Fig. 2.6b). Therefore the structure is in a closed conformation. Considering the closed pore and the ‘up’ VSDs, the reported structure of Cav1.1 may represent a potentially inactivated state.
3.5 The Auxiliary Subunits
In the structure of the rabbit Cav1.1 channel complex, four tandem cache domains and one VWA domain are identified in the α2δ subunit. Although these domains are organized separately in space, they involve intertwined sequences. Interestingly, although the δ subunit and α2 subunit are separated in the primary sequence, they co-fold with each other. The δ subunit contributes three β-strands to the fourth Cache domain. Cys1074 in the δ subunit forms a disulfide bond with Cys406 in the VWA domain. The extended conformation of δ is stabilized through multiple intra and inter-subunit disulfide bonds. In total, four disulfide bonds were observed between the α2- and δ-subunits and two within the δ-subunit.
There was a debate on whether the δ subunit is a transmembrane protein or a GPI (glycosylphosphatidylinositol)-anchored protein. In the 3.6 Å resolution EM map, there is no extra density for a TM helix. The density for δ subunit ended after Cys1074, which happened to be the predicted site for GPI modification. Besides, in the related MS result, no peptide can be detected after Cys1074. The structure appeared to support that δ subunit is a membrane anchored protein.
In the Cav1.1 complex structure, four transmembrane helices (named TM1–4), an extracellular β sheet and the cytosolic amino (N)- and C-terminal loops are resolved for the γ subunit. The transmembrane interface between α1 and γ is mediated by TM2 and TM3 of γ and the S3 and S4 segments in α1-VSDIV. All of the residues constituting the interface are hydrophobic residues, which are unlikely to provide the specificity between γ and VSDIV. On the intracellular side, the C-terminus of γ subunit contains some polar residues that are hydrogen-bonded with the III–IV linker in the α1 subunit, which may provide the interaction specificity with VSDIV but not for other VSDs. The direct contact between γ and VSDIV may affect the conformational changes of the latter during voltage dependent activation or inactivation, thereby providing the molecular basis for the antagonistic modulation of the γ subunit.
Before the determination of the structure of Cav1.1 complex, the crystal structure of β has been reported. The different cryo-EM maps of Cav1.1 revealed distinct conformations of the intracellular domains. The AID motif is sandwiched between α1-VSDII and β. Comparison of the different EM 3D reconstructions reveals shifts of the C terminus of S6I and the ensuing I–II helix. Meanwhile, the β-subunit undergoes a pronounced displacement between the two reconstructions.
3.6 Structural Mapping of Disease-Associated Mutations
Cav channels and the closely related Nav channels play a major role in a multitude of physiological and pathological processes. Hundreds of mutations have been identified in these channels. The structure presented here represents the first atomic model of a single-chain eukaryotic Cav or Nav channel. Structural mapping of the disease-associated mutations will greatly foster mechanistic understanding of the related disorders and provide the opportunity for novel drug development targeting these channels (Fig. 2.7).
Structural mapping of disease-associated mutations identified in the human Cav channels. The disease-related mutations identified in six human Cav channels are mapped to the structure of the rCav1.1α1 and color coded for different subtypes of Cav channels (a) or disorders (b). HOKPP1: periodic paralysis hypokalemic 1; MHS5: malignant hyperthermia 5; TCLS&NPP: Transient compartment-like syndrome and normokalaemic periodic paralysis; TS: Timothy syndrome; BRGDA3: Brugada syndrome 3; SANDD: sinoatrial node dysfunction and deafness; PASNA: primary aldosteronism, seizures, and neurologic abnormalities; NIDDM: non-insulin-dependent diabetes mellitus; CSNB2A: night blindness, congenital stationary, 2A; AIED: Aaland island eye disease; SCA6: spinocerebellar ataxia 6; FHM1: migraine, familial hemiplegic, 1; EA2: episodic ataxia 2; ECA6: epilepsy, childhood absence 6
4 Voltage-Gated Calcium Channels in Excitation-Contraction Coupling
Excitation-contraction coupling (E-C coupling) of muscles is an important and fundamental physiological process. Multiple proteins are involved in this signal transduction cascade, among which the ryanodine receptor RyR and Cav channels are two key components [47].
RyRs are high-conductance calcium release channels located on the sarco/endoplasmic (SR/ER) membrane. They are responsible for the rapid release of Ca2+ from intracellular stores during E-C coupling [48,49,50]. As the largest known ion channel, the homotetrameric RyR has a molecular mass of more than 2.2 MDa with each protomer containing about 5000 residues [51, 52]. In mammals, there are three isoforms: RyR1 and RyR2 are primarily expressed in skeletal and cardiac muscles, respectively, and RyR3 was originally found in the brain [53,54,55,56]. Recently, the near atomic 3D structures of RyR1 and RyR2 in multiple conformations have been reported [57,58,59,60]. Details of RyRs are discussed in a later chapter.
Cav channels, which are located in the transverse tubule (or T-tubule) are activated upon plasma membrane depolarization, and then induce the activation of the downstream RyRs. The sudden increase of the cytoplasmic Ca2+ triggers a cascade of cellular events that eventually result in muscle contraction. Ca2+ ions are then pumped back to the SR by the calcium ATPase SERCA, leading to muscle relaxation [47].
Cav1.1 (in skeletal muscle) and Cav1.2 (in cardiac muscle) sense the changes of the membrane potential of the plasma membrane and activate RyR1 and RyR2, respectively, but via different mechanisms (Fig. 2.8). In skeletal muscle, Cav1.1 undergoes conformational changes in response to membrane depolarization, which then activate RyR1 through physical association. It is called the “mechanical coupling” mechanism [61]. In cardiac muscle, it is the Ca2+ influx mediated by Cav1.2 that activates RyR2, a mechanism known as the “calcium induced calcium release” (CICR) [62].
Working models for activation of RyR1/2 by Cav1/2, respectively. In the skeletal muscle, RyR1 is activated by the membrane depolarization-induced conformational changes of Cav1.1 through physical interactions. It is called the “mechanical coupling” mechanism. In cardiac muscle, Cav1.2 permeates Ca2+ ions into the cytosol, leading to the activation of RyR2, a mechanism known as the “calcium induced calcium release”
To understand the activation mechanism of RyR1 by Cav1.1, the accurate interaction between Cav1.1 and RyR1 is needed. Although the near atomic resolution structures of Cav1.1 and RyR1 have been solved, there is no detailed structural information on the Cav1.1 and RyR1 supercomplex. According to the structural analysis, a speculative mechanism is proposed [57]. As reported, RyR1 is activated through direct physical contacts with the Cav1.1 [61, 63,64,65]. Multiple areas, such as the SPRY3 domain of RyR1 and β-subunit of Cav1.1, are involved in their coupling [66]. The conformational changes of the VSDs of Cav1.1α1 may induce shifts of the β-subunit and other cytoplasmic segments of Cav1.1, subsequently triggering the motion of the adjacent RyR1 cytoplasmic domains. Note that the SPRY3 domain of RyR1 is in direct contact with the N-terminal domain (NTD) within the same protomer. Potential shifts of the SPRY3 domain may be translated to the conformational changes of NTD, and subsequently the Handle domain and the Central domain. So the structural shifts triggered by Cav1.1 at the periphery of the RyR1 cytoplasmic region are propagated along the superhelical assemblies of the cytoplasmic domains to the Central domain, eventually leading to the opening of the intracellular gate. Nonetheless, the speculative mechanism still awaits experimental evidence.
5 Perspective
Voltage-gated calcium channels play essential roles in a multitude of physiological processes. The structure of Cav1.1 complex provides clues to the gating mechanism and selectivity, which establishes a foundation for mechanistic understanding of E-C coupling and provides a three-dimensional template for molecular interpretations of the functions and disease mechanisms of Cav and Nav channels.
In the past few decades, most high resolution protein structures were obtained from X-ray crystallography. Until a few years ago, we witnessed the breakthrough of cryo-EM. The resolution was improved to near atomic resolution (beyond 4 Å), a “resolution revolution”. A number of important protein or protein complex structures were solved. However, regardless of single-particle cryo-EM or X-ray crystallography, purification of the protein is required. Therefore, some proteins or protein complexes that are resistant to in vitro purification cannot yield high resolution structure. To visualize the E-C coupling ultrastructure formed by RyR1 and Cav1.1, we may eventually have to employ another technology, electron tomography (ET), which can image the samples in situ. The rapid technological development of ET may allow structural resolution the RyR1-Cav1.1 supercomplex to molecular details in the near future.
Abbreviations
- Cav:
-
Voltage-gated calcium channel
- DHPR:
-
Dihydropyridine receptor
- E-C coupling:
-
Excitation-contraction coupling
- Nav:
-
Voltage-gated sodium channel
- RyR:
-
Ryanodine receptor
- TM:
-
Transmembrane
- VGIC:
-
Voltage-gated ion channel
- VSD:
-
Voltage sensing domain
References
Catterall WA (2011) Voltage-gated calcium channels. Cold Spring Harb Perspect Biol 3:a003947
Clapham DE (2007) Calcium signaling. Cell 131:1047–1058
Ertel EA, Campbell KP, Harpold MM, Hofmann F, Mori Y, Perez-Reyes E, Schwartz A, Snutch TP, Tanabe T, Birnbaumer L et al (2000) Nomenclature of voltage-gated calcium channels. Neuron 25:533–535
Nowycky MC, Fox AP, Tsien RW (1985) Three types of neuronal calcium channel with different calcium agonist sensitivity. Nature 316:440–443
Tsien RW, Lipscombe D, Madison DV, Bley KR, Fox AP (1988) Multiple types of neuronal calcium channels and their selective modulation. Trends Neurosci 11:431–438
Fleckenstein A (1983) History of calcium antagonists. Circ Res 52:I3–16
Kohlhardt M, Fleckenstein A (1977) Inhibition of the slow inward current by nifedipine in mammalian ventricular myocardium. Naunyn Schmiedebergs Arch Pharmacol 298:267–272
Carbone E, Lux HD (1984) A low voltage-activated, fully inactivating Ca channel in vertebrate sensory neurones. Nature 310:501–502
Llinas R, Yarom Y (1981) Electrophysiology of mammalian inferior olivary neurones in vitro. Different types of voltage-dependent ionic conductances. J Physiol 315:549–567
Llinas RR, Sugimori M, Cherksey B (1989) Voltage-dependent calcium conductances in mammalian neurons. The P channel. Ann NY Acad Sci 560:103–111
Randall A, Tsien RW (1995) Pharmacological dissection of multiple types of Ca2+ channel currents in rat cerebellar granule neurons. J Neurosci 15:2995–3012
Tanabe T, Takeshima H, Mikami A, Flockerzi V, Takahashi H, Kangawa K, Kojima M, Matsuo H, Hirose T, Numa S (1987) Primary structure of the receptor for calcium channel blockers from skeletal muscle. Nature 328:313–318
Wu J, Yan Z, Li Z, Qian X, Lu S, Dong M, Zhou Q, Yan N (2016) Structure of the voltage-gated calcium channel Ca(v)1.1 at 3.6 A resolution. Nature 537:191–196
Takahashi M, Seagar MJ, Jones JF, Reber BF, Catterall WA (1987) Subunit structure of dihydropyridine-sensitive calcium channels from skeletal muscle. Proc Natl Acad Sci USA 84:5478–5482
Davies A, Hendrich J, Van Minh AT, Wratten J, Douglas L, Dolphin AC (2007) Functional biology of the alpha(2)delta subunits of voltage-gated calcium channels. Trends Pharmacol Sci 28:220–228
Dolphin AC (2003) Beta subunits of voltage-gated calcium channels. J Bioenerg Biomembr 35:599–620
Dolphin AC (2013) The alpha2delta subunits of voltage-gated calcium channels. Biochim Biophys Acta 1828:1541–1549
Cheng W, Altafaj X, Ronjat M, Coronado R (2005) Interaction between the dihydropyridine receptor Ca2+ channel beta-subunit and ryanodine receptor type 1 strengthens excitation-contraction coupling. Proc Natl Acad Sci USA 102:19225–19230
Gregg RG, Messing A, Strube C, Beurg M, Moss R, Behan M, Sukhareva M, Haynes S, Powell JA, Coronado R et al (1996) Absence of the beta subunit (cchb1) of the skeletal muscle dihydropyridine receptor alters expression of the alpha 1 subunit and eliminates excitation-contraction coupling. Proc Natl Acad Sci USA 93:13961–13966
Schredelseker J, Di Biase V, Obermair GJ, Felder ET, Flucher BE, Franzini-Armstrong C, Grabner M (2005) The beta 1a subunit is essential for the assembly of dihydropyridine-receptor arrays in skeletal muscle. Proc Natl Acad Sci USA 102:17219–17224
Andronache Z, Ursu D, Lehnert S, Freichel M, Flockerzi V, Melzer W (2007) The auxiliary subunit gamma 1 of the skeletal muscle L-type Ca2+ channel is an endogenous Ca2+ antagonist. Proc Natl Acad Sci USA 104:17885–17890
Kang MG, Campbell KP (2003) Gamma subunit of voltage-activated calcium channels. J Biol Chem 278:21315–21318
Olivera BM, Miljanich GP, Ramachandran J, Adams ME (1994) Calcium channel diversity and neurotransmitter release: the omega-conotoxins and omega-agatoxins. Annu Rev Biochem 63:823–867
Klugbauer N, Lacinova L, Marais E, Hobom M, Hofmann F (1999) Molecular diversity of the calcium channel alpha2delta subunit. J Neurosci 19:684–691
Zamponi GW, Striessnig J, Koschak A, Dolphin AC (2015) The physiology, pathology, and pharmacology of voltage-gated calcium channels and their future therapeutic potential. Pharmacol Rev 67:821–870
Bian F, Li Z, Offord J, Davis MD, McCormick J, Taylor CP, Walker LC (2006) Calcium channel alpha2-delta type 1 subunit is the major binding protein for pregabalin in neocortex, hippocampus, amygdala, and spinal cord: an ex vivo autoradiographic study in alpha2-delta type 1 genetically modified mice. Brain Res 1075:68–80
Li Z, Taylor CP, Weber M, Piechan J, Prior F, Bian F, Cui M, Hoffman D, Donevan S (2011) Pregabalin is a potent and selective ligand for alpha(2)delta-1 and alpha(2)delta-2 calcium channel subunits. Eur J Pharmacol 667:80–90
Wolf M, Eberhart A, Glossmann H, Striessnig J, Grigorieff N (2003) Visualization of the domain structure of an L-type Ca2+ channel using electron cryo-microscopy. J Mol Biol 332:171–182
Van Petegem F, Clark KA, Chatelain FC, Minor DL (2004) Structure of a complex between a voltage-gated calcium channel beta-subunit and an alpha-subunit domain. Nature 429:671–675
Hu HL, Wang Z, Wei RS, Fan GZ, Wang QL, Zhang KM, Yin CC (2015) The molecular architecture of dihydropyrindine receptor/L-type Ca2+ channel complex. Sci Rep 5:8370
Murata K, Nishimura S, Kuniyasu A, Nakayama H (2010) Three-dimensional structure of the alpha(1)-beta complex in the skeletal muscle dihydropyridine receptor by single-particle electron microscopy. J Electron Microsc 59:215–226
Serysheva II, Ludtke SJ, Baker MR, Chiu W, Hamilton SL (2002) Structure of the voltage-gated L-type Ca2+ channel by electron cryomicroscopy. Proc Natl Acad Sci USA 99:10370–10375
Chen YH, Li MH, Zhang Y, He LL, Yamada Y, Fitzmaurice A, Shen Y, Zhang HL, Tong L, Yang J (2004) Structural basis of the alpha(1)-beta subunit interaction of voltage-gated Ca2+ channels. Nature 429:675–680
Opatowsky Y, Chen CC, Campbell KP, Hirsch JA (2004) Structural analysis of the voltage-dependent calcium channel beta subunit functional core and its complex with the alpha 1 interaction domain. Neuron 42:387–399
Payandeh J, Scheuer T, Zheng N, Catterall WA (2011) The crystal structure of a voltage-gated sodium channel. Nature 475:353–358
Zhang X, Ren WL, DeCaen P, Yan CY, Tao X, Tang L, Wang JJ, Hasegawa K, Kumasaka T, He JH et al (2012) Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature 486:130–U160
Sula A, Booker J, Ng LCT, Naylor CE, DeCaen PG, Wallace BA (2017) The complete structure of an activated open sodium channel. Nat Commun 8:14205
Tang L, El-Din TMG, Payandeh J, Martinez GQ, Heard TM, Scheuer T, Zheng N, Catterall WA (2014) Structural basis for Ca2+ selectivity of a voltage-gated calcium channel. Nature 505:56–61
Liao M, Cao E, Julius D, Cheng Y (2013) Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504:107–112
Yan C, Hang J, Wan R, Huang M, Wong CC, Shi Y (2015a) Structure of a yeast spliceosome at 3.6-angstrom resolution. Science 349:1182–1191
Bannister RA, Beam KG (2013) Ca(V)1.1: the atypical prototypical voltage-gated Ca(2)(+) channel. Biochim Biophys Acta 1828:1587–1597
Buraei Z, Yang J (2013) Structure and function of the beta subunit of voltage-gated Ca(2)(+) channels. Biochim Biophys Acta 1828:1530–1540
Curtis BM, Catterall WA (1984) Purification of the calcium-antagonist receptor of the voltage-sensitive calcium-channel from skeletal-muscle transverse tubules. Biochemistry 23:2113–2118
Florio V, Striessnig J, Catterall WA (1992) Purification and reconstitution of skeletal-muscle calcium channels. Methods Enzymol 207:529–546
Sharp AH, Imagawa T, Leung AT, Campbell KP (1987) Identification and characterization of the dihydropyridine-binding subunit of the skeletal-muscle dihydropyridine receptor. J Biol Chem 262:12309–12315
Wu J, Yan Z, Li Z, Yan C, Lu S, Dong M, Yan N (2015) Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350:aad2395
Lanner JT, Georgiou DK, Joshi AD, Hamilton SL (2010) Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb Perspect Biol 2:a003996
Inui M, Saito A, Fleischer S (1987) Purification of the ryanodine receptor and identity with feet structures of junctional terminal cisternae of sarcoplasmic reticulum from fast skeletal muscle. J Biol Chem 262:1740–1747
Lai FA, Erickson HP, Rousseau E, Liu QY, Meissner G (1988) Purification and reconstitution of the calcium release channel from skeletal muscle. Nature 331:315–319
Pessah IN, Waterhouse AL, Casida JE (1985) The calcium-ryanodine receptor complex of skeletal and cardiac muscle. Biochem Biophys Res Commun 128:449–456
Bhat MB, Zhao J, Takeshima H, Ma J (1997) Functional calcium release channel formed by the carboxyl-terminal portion of ryanodine receptor. Biophys J 73:1329–1336
Takeshima H, Nishimura S, Matsumoto T, Ishida H, Kangawa K, Minamino N, Matsuo H, Ueda M, Hanaoka M, Hirose T et al (1989) Primary structure and expression from complementary DNA of skeletal muscle ryanodine receptor. Nature 339:439–445
Hakamata Y, Nakai J, Takeshima H, Imoto K (1992) Primary structure and distribution of a novel ryanodine receptor/calcium release channel from rabbit brain. FEBS Lett 312:229–235
Nakai J, Imagawa T, Hakamat Y, Shigekawa M, Takeshima H, Numa S (1990) Primary structure and functional expression from cDNA of the cardiac ryanodine receptor/calcium release channel. FEBS Lett 271:169–177
Otsu K, Willard HF, Khanna VK, Zorzato F, Green NM, MacLennan DH (1990) Molecular cloning of cDNA encoding the Ca2+ release channel (ryanodine receptor) of rabbit cardiac muscle sarcoplasmic reticulum. J Biol Chem 265:13472–13483
Rossi D, Sorrentino V (2002) Molecular genetics of ryanodine receptors Ca2+-release channels. Cell Calcium 32:307–319
Bai XC, Yan Z, Wu JP, Li ZQ, Yan N (2016) The central domain of RyR1 is the transducer for long-range allosteric gating of channel opening. Cell Res 26:995–1006
des Georges A, Clarke OB, Zalk R, Yuan Q, Condon KJ, Grassucci RA, Hendrickson WA, Marks AR, Frank J (2016) Structural basis for gating and activation of RyR1. Cell 167:145–157
Peng W, Shen HZ, Wu JP, Guo WT, Pan XJ, Wang RW, Chen SRW, Yan N (2016) Structural basis for the gating mechanism of the type 2 ryanodine receptor RyR2. Science 354(6310):aah5324
Yan Z, Bai XC, Yan C, Wu J, Li Z, Xie T, Peng W, Yin CC, Li X, Scheres SH et al (2015b) Structure of the rabbit ryanodine receptor RyR1 at near-atomic resolution. Nature 517:50–55
Rios E, Brum G (1987) Involvement of dihydropyridine receptors in excitation-contraction coupling in skeletal muscle. Nature 325:717–720
Endo M (1977) Calcium release from the sarcoplasmic reticulum. Physiol Rev 57:71–108
Franzini-Armstrong C, Protasi F, Ramesh V (1999) Shape, size, and distribution of Ca(2+) release units and couplons in skeletal and cardiac muscles. Biophys J 77:1528–1539
Protasi F, Franzini-Armstrong C, Flucher BE (1997) Coordinated incorporation of skeletal muscle dihydropyridine receptors and ryanodine receptors in peripheral couplings of BC3H1 cells. J Cell Biol 137:859–870
Tanabe T, Beam KG, Adams BA, Niidome T, Numa S (1990) Regions of the skeletal muscle dihydropyridine receptor critical for excitation-contraction coupling. Nature 346:567–569
Perez CF, Mukherjee S, Allen PD (2003) Amino acids 1-1,680 of ryanodine receptor type 1 hold critical determinants of skeletal type for excitation-contraction coupling. Role of divergence domain D2. J Biol Chem 278:39644–39652
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG, part of Springer Nature
About this chapter
Cite this chapter
Wu, J., Yan, N., Yan, Z. (2017). Structure-Function Relationship of the Voltage-Gated Calcium Channel Cav1.1 Complex. In: Krebs, J. (eds) Membrane Dynamics and Calcium Signaling. Advances in Experimental Medicine and Biology, vol 981. Springer, Cham. https://doi.org/10.1007/978-3-319-55858-5_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-55858-5_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55857-8
Online ISBN: 978-3-319-55858-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)