Abstract
Small RNAs (sRNAs), ~20–25 nucleotide (nt) in size, regulate various biological processes in plants through directing sequence-specific gene silencing. sRNAs are derived from either single- or double-stranded precursor RNAs. Proper levels of sRNAs are crucial for plant growth, development, genomic stability, and adaptation to abiotic and biotic stresses. Studies have identified the machineries controlling sRNA levels through biogenesis and degradation. This chapter covers recent progresses related to mechanisms governing small RNA biogenesis and degradation.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Small RNAs (sRNAs) are repressors of gene expression and play essential roles in various biological processes (Baulcombe 2004; Bologna and Voinnet 2014; Borges and Martienssen 2015). Upon production, sRNAs are sorted into their effector protein called ARGONUATE (AGO) and guide it to recognize target RNAs or DNA loci through sequence complementarity. AGO then silences gene expression at transcriptional levels through directing DNA methylation or histone modification and/or at post-transcriptional levels through target RNA cleavage or translational inhibition. microRNAs (miRNAs) and small interfering RNAs (siRNAs) are two major classes of sRNAs. They are chemically identical, but disguisable at their origin and biogenesis (Chen 2009). miRNAs are derived from primary miRNA transcripts (pri-miRNAs), which contain one or more miRNA-residing imperfect step-loops, while siRNAs are produced from long perfect double-stranded RNAs (dsRNAs) (Chen 2009). Depending on their origin, biogenesis and acting model, endogenous siRNAs can be further divided into several classes: trans-acting siRNAs (ta-siRNAs), phased siRNAs (pha-siRNAs), repeated DNA-derived siRNAs (ra-siRNAs), and natural cis-antisense siRNAs (nat-siRNAs) (Baulcombe 2004; Bologna and Voinnet 2014; Borges and Martienssen 2015). sRNAs are also subject to degradation (Xie et al. 2015). Turnover is also critical for proper function of sRNAs because excess amount of sRNA impairs plant development. Here, we review our current knowledge related to sRNA biogenesis and degradation.
2 miRNA Biogenesis in Plants
Most pri-miRNA-encoding genes (MIR) are localized at intergenic regions and transcribed as independent units by the DNA-dependent RNA polymerase II (Pol II) (Coruh et al. 2014; Nozawa et al. 2012; Xie et al. 2005). However, some MIRs are co-transcribed with host genes, some of which are transposons, as intronic or exonic sequences (Piriyapongsa and Jordan 2008; Yang et al. 2012).Upon transcription, pri-miRNAs are cut by the RNAseIII enzyme called DICER-LIKE 1 (DCL1) at least two times to release the miRNA/miRNA* (passenger strand) duplexes in the nucleus (Fig. 1). HUA1 ENHANCER 1 (HEN1), a small RNA methylase, then deposits a methyl group at the 3′ end of the miRNA duplex to stabilize them (Fig. 1) (Xie et al. 2015; Yu et al. 2005).
The miRNA biogenesis pathway. Most MIRs are transcribed by Pol II to produce pri-miRNAs. CDC5, NOT2, and Mediator interact with Pol II and MIR promoters and are required for MIR transcription. After transcription, DDL-binding and PRL1-binding stabilize pri-miRNAs. Pri-miRNAs are co-transcriptionally processed, which requires the elongator complex. SE and HYL1 form a complex with DCL1 to precisely and efficiently process pri-miRNAs in the nucleus. Additional factors such as TGH, CDC5, PRL1, NOT2, CBP20/80, and DDL associate with the DCL1 complex to facilitate its activity. NOT2, MOS2, and TGH also promote the recruitment of pri-miRNAs to the DCL1 complex. After processing, HEN1, which interacts with HYL1 and DCL1, methylates the miRNA duplex
Many additional factors contribute to miRNA biogenesis by modulating pri-miRNA transcription, processing, and/or stability. Similar to mRNA-coding genes, MIR transcription requires general and specific transcription factors. NOT2a and its homolog NOT2b, which are core subunits of the conserved CARBON CATABOLITE REPRESSION4 (CCR4)-NOT complex, CDC5, which is an atypical MYB transcription factor, and Mediator (a transcription factor) interact with Pol II and MIR promoters, and positively regulate transcription of many MIR genes (Fig. 1) (Kim et al. 2011; Wang et al. 2013; Zhang et al. 2013b). Besides them, the cycling DOF transcription factor (CDF2) binds a subset of MIR promoters to promote or repress their activities (Sun et al. 2015). In addition, the ATP-dependent SWR1 chromatin-remodeling complex (SWR1-C) also positively contributes to the expression of MIRs through changings of the nucleosome dynamics (Choi et al. 2016). Notably, transcription of some MIRs is temporally and spatially regulated. For instance, the transcription factors SCARECROW (SCR) and SHORT ROOT activate the expression of MIR166 in root endodermis (Carlsbecker et al. 2010). After transcription, a 5′ 7-methylguanosine cap and a 3′ polyadenylated tail (poly-A) are added to pri-miRNAs (Jones-Rhoades and Bartel 2004; Xie et al. 2005). 5′ cap likely stabilizes pri-miRNAs since defection in 5′ capping reduces pri-miRNA accumulation (Hajheidari et al. 2012). Besides 5′ cap, two proteins, DAWDLE (DDL) and PLEIOTROPIC REGULATORY LOCUS 1 (PRL1), bind and stabilize pri-miRNAs following transcription (Fig. 1) (Yu et al. 2008; Zhang et al. 2014, 2015).
Pri-miRNAs are co-transcriptionally processed by DCL1, which is evidenced by the involvement of the elongator complex, which is required for the elongation of Pol II-dependent transcripts, in miRNA biogenesis (Fig. 1) (Fang et al. 2015a). Elongator interacts with DCL1 and is required for the association of DCL1 with MIR loci. This observation suggests that DCL1 may be recruited to the nascent pri-miRNAs during transcript elongation (Fang et al. 2015a). The efficient cleavage of miRNA/miRNA duplex from pri-miRNAs by DCL1 requires HYL1 (and sRNA-binding protein), TOUGH (TGH; an RNA-binding protein), and SERRATE (SE; a zinc-finger protein) (Fig. 1) (Dong et al. 2008; Fang and Spector 2007; Fujioka et al. 2007; Ren et al. 2012b; Ren and Yu 2012). HYL1 and SE are also required for precise cleavage of miRNA/miRNA* from pri-miRNAs (Dong et al. 2008), while TGH also modulates the interaction between pri-miRNAs and the processing complex (Ren et al. 2012b). It has been proposed that DCL1, SE, and HYL1 form a Dicing body (D-body) (Fang and Spector 2007), whose formation requires MOS2, an RNA-binding protein (Wu et al. 2013). In addition, NOT2, CDC5, PRL1, and CDF2 also interact with DCL1 and SE to promote pri-miRNA processing (Fig.1) (Sun et al. 2015; Wang et al. 2013; Zhang et al. 2014, 2013b). As these proteins also associate with Pol II, it is possible that they play a role in the co-transcriptional recruitment of DCL1 to pri-miRNAs. Interestingly, the CAP-binding proteins 80 (CBP 80) and CBP20 also associate with the DCL1 complex and are required for miRNA accumulation (Fig. 1) (Gregory et al. 2008; Kim et al. 2008; Laubinger et al. 2008). The recruitment of pri-miRNA to the DCL1 complex involves the THO/TREX complex that functions in the transport of nascent mRNAs from the nucleus towards the cytoplasm (Fig. 1) (Francisco-Mangilet et al. 2015). Furthermore, several additional proteins participate in miRNA biogenesis through the interaction with the accessory factors of DCL1. RECEPTOR FOR ACTIVATED C KINASE 1 (RACK1) that serves as a scaffold for protein bindings interacts with SE to promote pri-miRNA processing (Speth et al. 2013), whereas SICKLE (SIC), a proline-rich protein, co-localizes with HYL1 and is required for the accumulation of a subset of miRNAs (Zhan et al. 2012). Interestingly, GRP7, a homology of human hnRNP A1 involved in splicing, binds a subset of pri-miRNAs to repress their processing (Koster et al. 2014). Notably, REGULATOR OF CBF GENE EXPRESSION 3 (RCF3, also known as HOS5 and SHI1) can bind a subset of pri-miRNAs to regulate their processing in a tissue-specific manner (Chen et al. 2015; Karlsson et al. 2015).
The miRNA biogenesis machinery itself is regulated at both transcriptional and post-transcription levels. The histone acetyltransferase GCN5 promotes the transcription of both DCL1 and HYL1 (Kim et al. 2009). Optimal DCL1 transcription also requires the STA1, a splicing factor, and CAM33/XAP CIRCADIAN TIMEKEEPER (XCT, a nuclear localized protein) (Ben Chaabane et al. 2013; Fang et al. 2015b). Notably, SE and DCL1 are targets of miR863 and miR162, respectively (Niu et al. 2016; Rajagopalan et al. 2006). This suggests that DCL1 and SE transcripts subject to feedback regulation. Furthermore, both DCL1 and HYL1 activities are regulated by protein phosphorylation. HYL1 is phosphorylated by the MITOGEN-ACTIVATED PROTEIN KINASE 3 (MPK3), which inhibits miRNA biogenesis (Raghuram et al. 2015). To counteract MPK3 activity, C-TERMINAL DOMAIN PHOSPHATASE-LIKE 1 and 2 (CPL1 and CPL2) dephosphorylate HYL1 to enhance miRNA biogenesis in an SE-dependent manner (Manavella et al. 2012). Besides HYL1, CPL1 and CPL2 also recognize RCF3 to positively impact its function in miRNA biogenesis (Chen et al. 2015; Karlsson et al. 2015). DCL1 interacts with the forkhead domain (FHA) of DDL, which mediates protein–protein interactions by targeting phospho-threonine containing motifs (Machida and Yuan 2013). The phospho-threonine binding cleft of FHA interacts with the helicase domain of DCL1 that contains potential phospho-threonine motifs, suggesting that DCL1 may be phosphorylated for its optimal activity (Machida and Yuan 2013). Interestingly, ubiquitination also plays a role in miRNA biogenesis. For instance, Constitutive Photomorphogenic 1 (COP1), an E3 ubiquitin ligase, has been recently shown to block an activity degrading HYL1 in light via unknown mechanism (Cho et al. 2014).
3 The Biogenesis of ta-siRNAs and pha-siRNAs
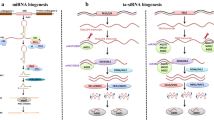
ta-siRNAs refer to a class siRNAs that act on targets other than the genes that derive ta-siRNAs (Allen et al. 2005; Peragine et al. 2004). This distinguishes ta-siRNAs from other endogenous siRNAs, which mostly silence genes that are the same as or homologs to the genes from which they derive. The production of ta-siRNAs requires miRNA-directed cleavage of primary ta-siRNA transcripts (TASs) that have the same structures as mRNAs (Fig. 2a) (Allen et al. 2005; Axtell et al. 2006; Yoshikawa et al. 2005). Two models for ta-siRNA production have been proposed. In the one-hit model, a 22-nt miRNA first directs AGO to cleave TASs (Fig. 2a) (Axtell et al. 2006). The 3′ cleavage products are then used as a template to synthesize dsRNAs by RDR6 (Fig. 2a). In the two-hit model, two 21-nt miRNAs recognize TASs at independent target sites along the transcript (Fig. 2a) (Axtell et al. 2006). RDR6 is then recruited to the cleavage products to generate dsRNAs. In both scenarios, the dsRNAs are cleaved by DCL4 or DCL2 every 21 or 22 nt from the initial miRNA cleavage point, resulting in a phased production of secondary siRNAs (Fig. 2a) (Fei et al. 2013; Ronemus et al. 2006). ta-siRNA biogenesis requires the assistance of SGS3 and DRB4 (Fig. 2a). DRB4 is a HYL1 homolog and interacts with DCL4 (Adenot et al. 2006), while SGS3 binds dsRNAs with 5′ overhang and partners with RDR6 (Fukunaga and Doudna 2009). In Arabidopsis, miR173 and miR828, 22 nt in size, induce the production of ta-siRNAs from TAS1/TAS2 and TAS4, respectively, whereas miR390, 21 nt in size binds AGO7 to target TAS3 (Allen et al. 2005; Axtell et al. 2006; Yoshikawa et al. 2005). The miR390-AGO7-TAS3 combination appears to be conserved among moss, rice, maize, and gymnosperms, suggesting that the two-hit model may be ancestral to the one-hit model (Fei et al. 2013).
The biogenesis pathways for ta-/phas-siRNAs and nat-siRNAs. (a) Proposed biogenesis pathways for phas- and ta-siRNAs. The precursor RNAs of ta- and phas-siRNAs are first targeted by miRNAs through the one-hit or two-hit model, which triggers the production of dsRNAs by RDR6 (R) with the assistance of SGS3 (S). The resulting dsRNAs will be cleaved by DCLs to produce phased siRNAs. Yellow oval indicates AGOs associated with miRNAs. (b) Proposed biogenesis pathways for nat-siRNA production. Pol II- or Pol IV-dependent transcription of convergent genes results in dsRNAs. The dsRNAs will be processed by one or more DCLs to produce primary nat-siRNAs. Primary nat-siRNAs will be loaded into AGO (Yellow oval) to recognize one of original transcripts. This leads to the production of RDR-dependent dsRNAs, which will be further processed by DCLs to generate secondary nat-siRNAs
Unlike the Brassica plants that only encode few TAS loci, most non-brassica plants contain larger number of loci that produce pha-siRNAs (Arikit et al. 2014; Johnson et al. 2009; Shivaprasad et al. 2012; Zhai et al. 2015b). These pha-siRNAs are derived from many mRNAs and long noncoding RNAs called PHAS ncRNAs that are transcribed by Pol II, capped and polyadenylated, resembling mRNAs. However, some pha-siRNAs act in cis rather in trans. The production of pha-siRNAs resembles that of ta-siRNAs and is triggered by miRNA-directed cleavage (Fig. 2a). It has been proposed that RDR6 together with SGS3 use the 3′ cleavage fragments of PHAS transcripts as templates to synthesize dsRNA from the poly-A tail to the cleavage site (Fig. 2a) (Song et al. 2012b). DCL4 and DCL5/DCL3b subsequently process the dsRNAs to generate 21- and 24-nucleotide pha-siRNAs, respectively (Song et al. 2012a). In dicots, there is a conserved miR2118-482 superfamily that triggers the production of pha-siRNAs from transcripts encoding nucleotide-binding/leucine-rich repeat (NB-LRR) proteins (Arikit et al. 2014; Shivaprasad et al. 2012; Zhai et al. 2011). NB-LRR-derived pha-siRNAs act both in cis and in trans and thereby regulate additional members of NB-LRR family (Arikit et al. 2014; Shivaprasad et al. 2012; Zhai et al. 2011). This pha-siRNAs seem to benefit the plant resistance to bacterial infection, as NB-LRRs play essential role in plant immunity. However, NB-LRR pha-siRNAs are lost in grass genomes, which possess anther-specific pha-siRNAs instead (Arikit et al. 2014; Shivaprasad et al. 2012; Zhai et al. 2011).
4 The Biogenesis of Natural cis-antisense siRNAs (nat-siRNAs)
nat-siRNAs are derived from dsRNAs that are formed by convergent bidirectional transcripts generated from two partially overlapping genes (Fig. 2b) (Borsani et al. 2005; Katiyar-Agarwal et al. 2006; Ron et al. 2010; Zhang et al. 2012; Zubko and Meyer 2007). These cis-antisense transcripts are common in plant genome. Notably, nat-siRNAs are often induced by various stresses or at specific developmental stage or tissues and appear to be required for plant immunity and development (Borsani et al. 2005; Katiyar-Agarwal et al. 2006; Ron et al. 2010; Zhang et al. 2012; Zubko and Meyer 2007). DCL1, DCL2, and/or DCL3 cleave the dsRNA formed by the cis-antisense transcripts to initiate the production of primary nat-siRNAs (Fig. 2b) (Borsani et al. 2005; Katiyar-Agarwal et al. 2006; Ron et al. 2010; Zhang et al. 2012; Zubko and Meyer 2007). The primary nat-siRNAs will then guide the cleavage of the complementary transcripts (Fig. 2b). The resulting cleavage products will be used as templates for RDRs to produce dsRNAs, which will be further processed by DCLs into secondary siRNAs, leading to the reinforcement phase (Fig. 2b) (Borsani et al. 2005; Katiyar-Agarwal et al. 2006; Ron et al. 2010; Zhang et al. 2012; Zubko and Meyer 2007). The second phase resembles the production of pha-siRNAs involving RDRs and SGS3. The production of DCL3-dependent nat-siRNAs also requires the RNA-dependent RNA polymerase 2 (RDR2) and plant-specific RNA polymerase IV (Pol IV) (Zhang et al. 2012). In contrast, not all DCL1-dependent nat-siRNAs require RDR6/RDR2 and Pol IV for production (Zhang et al. 2012).
5 The Production of sRNAs Involved in RNA–Direct DNA Methylation (RdDM)
RdDM is a conserved process to silence transposable elements, to direct gene imprinting, and to maintain genome stability in plants and many metazoans (Castel and Martienssen 2013; Law and Jacobsen 2010; Matzke and Mosher 2014). Studies have revealed the presence of both canonical and non-canonical RdDM mechanisms in plants. In canonical RdDM, 24-nt siRNAs derived from repeated DNAs or heterochromatic regions (ra-siRNAs) direct the DNA de novo methyl transferase DOMAINS REARRANGED METHYLTRANSFERASE 2 (DRM2) to recognize RdDM target loci and to catalyze the methylation of previously unmodified cytosine in any sequence context (Fig. 3) (Castel and Martienssen 2013; Law and Jacobsen 2010; Matzke and Mosher 2014). In contrast, in non-canonical RdDM, miRNAs, ta-siRNAs, and 21-nt siRNAs partner with AGO to direct DNA methylation (Xie and Yu 2015). The mechanisms leading to the production of sRNAs involved in canonical and non-canonical RdDM have been proposed.
The biogenesis of canonical ra-siRNAs. SHH1 recognizes the K9 dimethylation (m in red circle) of H3 (shown in green oval) and recruits Pol IV-RDR2 to the RdDM loci, leading to the production of P4R2 transcripts that are converted to dsRNAs by RdR2. CLSY1 helps the correct localization of Pol IV and RDR2. The resulting dsRNAs are processed by DCL3 to produce 24-nt siRNAs, which are loaded into AGO4 (Yellow oval). The AGO4-ra-siRNAs are recruited to chromatin by the Pol V–AGO 4 interaction and the base-pairing between ra-siRNAs and Pol V-dependent transcripts flanking the RdDM loci. AGO4 recruits DRM2 to catalyze the de novo methylation (Red hexagon) of RdDM loci
5.1 The Biogenesis of Canonical ra-siRNAs
The biogenesis of ra-siRNAs starts with Pol IV-dependent transcription from the RdDM loci (Herr et al. 2005; Onodera et al. 2005). Pol IV is a plant-specific DNA-dependent RNA polymerase. It composes of twelve subunits, which are either identical or paralog to subunit of Pol II (Haag et al. 2014; Ream et al. 2009). Pol IV interacts with RDR2 (Haag et al. 2012; Law et al. 2011), which is also required for the production of Pol IV-dependent transcripts (Fig. 3) (P4R2 RNAs). After transcription, RDR2 converts the Pol IV-dependent transcripts into dsRNAs in the nucleus (Fig. 3). DCL3 then cuts the dsRNAs into 24-nt siRNAs (Fig. 3). When DCL3 is defective, DCL2 and DCL4 can process the dsRNAs into 22-nt and 21-nt siRNAs, respectively. 24-nt siRNAs are then loaded into AGO4 or AGO6 and recruited to the RdDM loci through base-pairing with Pol V-dependent transcripts and the interaction between AGO4 and Pol V (Fig. 3). AGO4/AGO6 subsequently recruits DRM2 to catalyze de novo DNA methylation (Fig. 3). Consequently, DNA methylation causes histone deacetylation, histone H3 lysine 9 methylation, and histone H3 lysine 4 demethylation, which repress gene transcription and cause heterochromatin formation.
The nature of Pol IV-dependent transcripts (P4R2 RNAs) has been mystery since the discovery of Pol IV. Recently, several groups identified P4R2 RNAs through sequencing RNAs in the mutant deficient in DCL3 (Blevins et al. 2015; Li et al. 2015; Yang et al. 2016; Ye et al. 2016; Zhai et al. 2015a). The sizes of P4R2 RNAs are ranging from ~26 to 45 nt although long P4R2 RNAs may also exist (Li et al. 2015). Unlike Pol II transcripts, P4R2 RNAs are not polyadenylated and capped (Blevins et al. 2015; Li et al. 2015; Yang et al. 2016; Ye et al. 2016; Zhai et al. 2015a). They often start with a purine (A or G) at their 5′ ends and contain one or two untemplated nucleotide (s) at the 3′ end. Notably, P4R2 RNAs can guide DNA methylation without being diced into siRNAs (Yang et al. 2016; Ye et al. 2016), raising the possibility that ra-siRNAs may not be required for RdDM. Pol IV activity may not require the promoters since Pol IV-associated regions do not contain consensus sequences (Law et al. 2013). Instead, Pol IV needs chromatin marks to define the transcript sites (Blevins et al. 2014; Law et al. 2013; Law et al. 2011; Zhang et al. 2013a). Histone deacetylation, maintenance DNA methylation, and histone H3K9 dimethylation (H3K9me2) have been shown to recruit Pol IV to the chromatin (Blevins et al. 2014; Law et al. 2011, 2013; Zhang et al. 2013a). Consistent with these observations, SAWADEE HOMEODOMAIN HOMOLOGUE 1 (SHH1), which interacts with Pol IV and binds H3K9me2 and unmethylated H3K4 through its unique tandem Tudor-like fold, is required for the recruitment of Pol IV to the chromatin (Fig. 3) (Law et al. 2011; Zhang et al. 2013a). In addition, CLSY1, a putative chromatin-remodeling factor, is required for Pol IV-RDR2 localization and activity (Fig. 3) (Smith et al. 2007). Interestingly, Pol II also assists the recruitment of Pol IV to chromatin at some RdDM loci presumably through its transcription activity, which suggest the interplay among different polymerase may be required for ra-siRNA production (Zheng et al. 2009).
5.2 The Biogenesis of Non-canonical sRNAs Involved in RdDM
Plants also use non-canonical RdDM pathways to defend transposons. In non-canonical RdDM pathways, sRNAs are produced differently from those in canonical pathways. Some Pol II-derived hairpin transcripts and pri-miRNAs can be processed by DCL3 to generate 24-nt siRNAs or miRNAs (Chellappan et al. 2010; Dunoyer et al. 2010; Khraiwesh et al. 2010; Slotkin et al. 2005; Wu et al. 2010). These 24-nt siRNAs and miRNAs are then fed into the canonical RdDM pathway to direct DNA methylation either in trans or in cis. RDR6 also converts TE mRNAs into dsRNAs if these TEs are highly expressed (Gasciolli et al. 2005; Mari-Ordonez et al. 2013). When DCL2 and DCL4 become saturated, DCL3 cuts TE-derived, RDR6-dependent dsRNAs, which are not typically DCL3 substrates, into 24-nt siRNAs (Gasciolli et al. 2005; Mari-Ordonez et al. 2013). This pathway relies on the hierarchical activity of DCLs. Interestingly, RDR6-dependent 21- and 22-nt siRNAs can also participate in RdDM, which is evidenced by the fact that ta-siRNAs can be loaded into AGO4 or AGO6 to direct DNA methylation at TAS loci (McCue et al. 2015; Wu et al. 2012). Notably, this pathway is independent of Pol IV, RDR2, and DCL3 although it requires Pol V and DRM2 (Wu et al. 2012). Subsequent studies show that similar mechanisms are employed to initiate and establish the silencing of transcriptionally active TEs, which are long and autonomous (Nuthikattu et al. 2013). Like ta-siRNAs, the production of TE-derived 21-/22-nt siRNAs depends on Pol II, SGS3, DCL2, and DCL4 (McCue et al. 2015; Wu et al. 2012). Recent studies also uncovered a non-canonical RdDM pathway that targets a subset of non-conserved genomic loci (Garcia et al. 2012; Pontier et al. 2012). This pathway depends on Pol IV, Pol V, AGO2, and a protein named Needed for RDR2-independent DNA methylation (NERD), which contains GW repeats and a PHD finger domain and interacts with Pol V and AGO2 (Garcia et al. 2012; Pontier et al. 2012). The biogenesis pathway of siRNAs that act in NERD-dependent RdDM is not well established. However, it seems that siRNA production in this pathway requires the combination of canonical and non-canonical RdDM proteins since siRNA accumulation is reduced in rdr1, rdr6, sde3, dcl2, dcl3, and nrpd1a (a Pol IV mutant) (Garcia et al. 2012; Pontier et al. 2012).
6 Methylation Stabilizes miRNAs and siRNAs
In plants, the 3′ termini of miRNAs and siRNAs harbor a 2′-O-methyl group (Fig. 1) (Yu et al. 2005). This modification is added by HEN1 following the release of the miRNA or siRNA duplexes (Fig. 1) (Yang et al. 2006; Yu et al. 2005). HEN1 exists in both eukaryotes and prokaryotes (Huang et al. 2009). Plant HEN1 contains several protein domains, including two dsRNA-binding domains (dsRBD1 and dsRBD2), a La-motif containing domain (LCD), and a methyltransferase domain (MTase) (Huang et al. 2009). Among these protein domains, the dsRBD1 and dsRBD2 domains enable HEN1 to recognize dsRNAs, while the distance between the LCD domain and the MTase domain helps HEN1 to determine its substrate length (Huang et al. 2009). As a result, plant HEN1 specifically deposits a methyl group to the 2′ OH position of the 3′ end in each strand of 21–24 base-pair (bp) dsRNAs with 2-nt overhangs (Huang et al. 2009; Yang et al. 2006), which are typical features of the miRNA and siRNA duplexes. HEN1 interacts with HYL1 and DCL1 (Fig. 1) (Baranauske et al. 2015), suggesting that miRNA production and methylation are a coupled process. In contrast, HEN1s from metazoans and bacteria lack the dsRNA-binding domain, and therefore, act on ssRNAs (Chan et al. 2009a; Horwich et al. 2007; Kirino and Mourelatos 2007). Consistent with this, metazoan HEN1 recognizes AGO-bound sRNAs (Ohara et al. 2007; Saito et al. 2007), whereas bacterial HEN1 modifies transfer RNAs (tRNAs) (Chan et al. 2009b). Interestingly, fly HEN1 interacts with PIWI (an AGO protein) (Ohara et al. 2007; Saito et al. 2007), indicating a potential role of AGO in determining substrate specificity for HEN1 in metazoans.
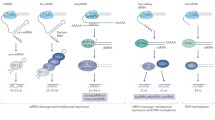
In plant hen1, sRNAs are reduced in abundance and become heterogeneity in size that is caused by untemplated uridine addition at 3′ termini (uridylation) and 3′-to-5′ exonucleolytic trimming activity (Fig. 4) (Abe et al. 2010; Li et al. 2005; Zhai et al. 2013). Consistent with the observation in plant hen1, sRNAs are also subjected to 3′-to-5′ trimming and uridylation in metazoan hen1 (Billi et al. 2012; Horwich et al. 2007; Kamminga et al. 2010; Kamminga et al. 2012; Montgomery et al. 2012). These results demonstrate that methylation is a conserved mechanism to protect sRNAs from degradation and uridylation.
Proposed model for miRNA methylation and degradation. After production, the miRNA/miRNA* duplexes are methylated (Green cycle) by HEN1. Methylated miRNAs are then loaded into AGO1 (Yellow oval) and direct AGO1 to cleave targets. The resulted 5′ cleavage products (5′ CP) are degraded through HESO1/URT1 (Blue hexagon)-mediated uridylation or 3′ trimming. The 2′-O-methylation protects miRNAs from uridylation and 3′ trimming. When methylation is lacking, miRNAs will be attacked by the AGO1-associated uridylation and 3′ trimming activities, leading to degradation. SDN1 may target AGO1-bound miRNAs with the 2′-O-methyl group and lead to their uridylation and degradation
7 Uridylation Triggers the Degradation of siRNAs and miRNAs
In plants, sRNAs become uridylated globally when HEN1 is lacking (Ren et al. 2014a). In Arabidopsis, a terminal uridyl transferase (TUTase) named HEN1 SUPPRESSOR1 (HESO1) catalyzes the uridylation of miRNAs and siRNAs (Ren et al. 2012a; Zhao et al. 2012). HESO1 acts progressively on RNAs in vitro, but its activity is blocked by 2′-O-methylation (Fig. 4) (Ren et al. 2012a; Zhao et al. 2012). Besides HESO1, UTP: RNA uridylyltransferase (URT1), which has been shown to uridylate some mRNAs, also act on miRNAs (Fig. 4) (Tu et al. 2015; Wang et al. 2015). However, unlike HESO1, URT1 does not recognize ra-siRNAs, likely due to its exclusive localization in the cytoplasm (Wang et al. 2015). In addition, URT1 appears to add short U-tail to miRNAs in vivo and seems to have a different preference to 3′ end nucleotides with HESO1 (Tu et al. 2015).
In hen1, heso1 reduces the U-tail length, resulting in increased abundance of most normal-sized, 3′ trimmed, and/or short-tailed sRNAs (Ren et al. 2012a; Zhao et al. 2012), whereas urt1 only affects a few miRNAs (Tu et al. 2015; Wang et al. 2015). However, when both HESO1 and URT1 are lacking, miRNA uridylation is globally abolished in hen1, resulting in elevated abundance of miRNAs and an extensive increase of 3′-to-5′ trimming (Wang et al. 2015). Furthermore, overexpression of HESO1 in hen1 further reduced the accumulation of normal-sized and 3′ trimmed miRNAs (Ren et al. 2012a). Taken together, these results reveal that HESO1 and URT1 synergistically and independently act on miRNAs and that uridylation triggers miRNA degradation and competes with the 3′-to-5′ trimming activity for substrates (Fig. 4) (Tu et al. 2015; Wang et al. 2015). In Chlamydomonas, the MUT68 nucleotidyltransferase uridylates miRNAs and siRNAs to trigger their degradation by the exosome components (Ibrahim et al. 2010). Bedside triggering degradation, uridylation may also block miRNA activity, which is evidenced by the fact that tailing of AGO1-bound miRNA165/6 greatly reduced its cleavage activity on their targets (Tu et al. 2015). Intriguingly, a single U addition in miR171 in hen1 urt1 enables miR171 to trigger the production of secondary siRNAs from its targets (Tu et al. 2015), suggesting that U addition may also alter miRNA activity. sRNA uridylation also exist in metazoans (Burroughs et al. 2010; Wyman et al. 2011). Like in plants, uridylation affects stability and function of metazoan sRNAs. Interestingly, many TUTases act on sRNAs and some of them act on miRNAs in a sequence-specific manner in metazoans (Burroughs et al. 2010; Wyman et al. 2011).
In addition to miRNAs, HESO1 and URT1 also uridylates the 5′ RNA fragments (5′ CP) generated by AGO1 cleavage of target RNAs (Fig. 4) (Ren et al. 2014b; Wang et al. 2015). Uridylation of 5′ CP is also a conserved process in both metazoans and plants (Shen and Goodman 2004). Similar to its effect on sRNAs, uridylation triggers degradation of 5′ CP, but competes with 3′-to-5′ exonucleolytic trimming activity (Fig. 4) (Ren et al. 2014b). Interestingly, MUT68 from Chlamydomonas also acts on 5′ CP (Ibrahim et al. 2006). These results suggest the presence of a common mechanism, by which TUTase recognize both sRNAs and 5′ CP. Indeed, both HESO1 and URT1 interact with AGO1 and add U-tails to AGO1-bound miRNAs (Ren et al. 2014b; Tu et al. 2015; Wang et al. 2015). Furthermore, defection of AGO1 abolishes uridylation of miRNAs in HEN1 (Ren et al. 2014b; Zhai et al. 2013). Thus, it is likely that TUTases recognizes its substrates in the AGO1 complex. These results also answer the question why plant miRNAs, but metazoan miRNAs, require methylation for stability (Ren et al. 2014a). In plants, miRNAs majorly direct target cleavage. The cleavage products need to be further eliminated. Otherwise, they cause lethality of plants. To ensure the rapid degradation of 5′ CP, uridylation and other degradation activities are associated with the AGO1 complex (Fig. 4). However, base-pairing plant miRNAs and their targets may expose miRNA 3′ end to these AGO1-associated activities due to the extensive complementarity (Ren et al. 2014a). Thus, plant miRNAs may need methylation to protect them from such activities. In contrast, metazoan miRNAs are less complementary to their targets and majorly inhibit translation. They consequently may not be exposed to uridylation/degradation activities when meeting with their targets. Consistent with this notion, the high complementarity between aritificial target RNAs and endogenous miRNAs. triggers miRNA tailing and trimming in metazoans (Ameres et al. 2010, 2011).
Similar to uridylation, a common mechanism may exist to degrade both uridylated miRNAs and 5′ CPs, which transiently associate with AGO. Indeed, the exosome has been shown to degrade both miRNAs and 5′ CP in Chlamydomonas (Ibrahim et al. 2006, 2010). However, such enzymes remain to be identified in higher plants. In metazoans, Dis3l2, a paralog of RRP44 that is a core component of the exosome, degrades uridylated precursor of let-7 miRNA (pre-let-7) (Chang et al. 2013; Ustianenko et al. 2013). By analog, plant homologs of Dis3l2 such as SUPPRESSOR OF VARICOSE and RRP44A may act on uridylated sRNAs and 5′ CPs (Ren et al. 2014a). Alternatively, U-tail may disassociate miRNAs and 5′ CPs from the AGO1 complex, causing their rapid degradation. This is supported by the observation that long tails can be added to AGO1-bound miRNAs in vitro (Ren et al. 2014b), but no long-tailed miRNAs/5′ CP can be detected in vivo (Tu et al. 2015; Wang et al. 2015). It is also possible that some unusual exoribonucleases may recognize the 3′ U-overhang of miRNA-5′ CP duplex to trigger their degradation.
8 Exoribonucleases Degrading sRNAs in Plants
In Arabidopsis, a family of 3′-to-5′ exoribonucleases including SMALL RNA DEGRADING NUCLEASE 1 (SDN1), 2, and 3 have been shown to degrade mature miRNAs (Ramachandran and Chen 2008). SDNs appear to act on short single-stranded RNAs, but not sRNA duplexes or pre-miRNAs (Ramachandran and Chen 2008). Furthermore, SDNs act on methylated, but not uridylated miRNAs (Ramachandran and Chen 2008), suggesting that SDNs may function coordinately with HESO1/URT1 to regulate miRNA abundance (Fig. 4). Lack of SDNs increases the abundance of miRNAs and causes pleiotropic development defects, demonstrating that turnover is an essential mechanism to maintain proper miRNA activities (Ramachandran and Chen 2008). Recent studies show that target mimicry, which blocks miRNA-mediate target cleavage, can induce the degradation of miRNAs by SDNs (Yan et al. 2012). This result further reinforces that the function of SDNs is to eliminate the unnecessary miRNAs. AGO10 has been shown to decoy miR165/166 from AGO1, and AGO10 binding also seems to trigger degradation of miR165/166 (Zhu et al. 2011). It is possible that SDNs may play a role in miR165/166 degradation caused by AGO10-decoy.
Unmethylated miRNAs also subjects to 3′-to-5′ trimming activity in higher plants. However, the enzyme remains to be identified. In Chlamydomonas, the RRP6 that possesses 3′-to-5′ exoribonuclease activity can degrade miRNAs (Ibrahim et al. 2006; Ibrahim et al. 2010). Arabidopsis encode three RRP6-LIKE (RRP6L) proteins (Lange et al. 2008). It is possible that RRP6L can act on unmethylated miRNAs in higher plants. In fly, a 3′-to-5′ exoribonuclease named Nibbler, binds AGO to trim miRNAs (Han et al. 2011; Liu et al. 2011). It has two homologs in Arabidopsis (Xie et al. 2015). It is reasonable to speculate that these two enzymes may act on unmethylated miRNAs in Arabidopsis.
9 Perspective
In the past decades, the framework for sRNA biogenesis has been established and progresses have been made towards to sRNA degradation. In addition, factors which play regulatory roles in miRNA biogenesis also have been isolated. These advances have resulted in a better understanding of biological processes involving sRNAs and improved our ability to apply related technologies. However, challenges still exist. Plants employ multiple mechanisms to regulate gene expression and genome stability. How sRNAs are coordinated with other regulatory mechanisms in various biological processes is not well known. In addition, the functional mechanisms of accessory proteins involved in sRNA biogenesis are mostly not well defined. Notably, many of these protein factors also function in transcription, RNA processing, splicing, and RNA decay. Thus, sRNA biogenesis may have interconnections with other RNA metabolisms. Further elucidation of these interconnections is still an obstacle to our understanding of various sRNA pathways. miRNA biogenesis is regulated through transcription, processing, and stability. It remains poorly understood how plants coordinate these processes to ensure proper miRNA levels in response to development and physiological signals. In higher plants, the enzymes degrading modified and unmodified sRNAs are largely unknown, which have greatly limited our understanding of sRNA turn over. Furthermore, the factors regulating siRNA production are largely unknown. Consequently, it is not clear how related biological processes such as DNA methylation are regulated at various developmental stages and in response to biotic and abiotic stresses. Finally, a practical challenge is the optimization of sRNA-based technology and related application used to improve agricultural trait of crops.
References
Abe M, Yoshikawa T, Nosaka M et al (2010) WAVY LEAF1, an ortholog of Arabidopsis HEN1, regulates shoot development by maintaining microRNA and trans-acting small interfering RNA accumulation in rice. Plant Physiol 154:1335–1346
Adenot X, Elmayan T, Lauressergues D et al (2006) DRB4-dependent TAS3 trans-acting siRNAs control leaf morphology through AGO7. Curr Biol 16:927–932
Allen E, Xie Z, Gustafson AM et al (2005) microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell 121:207–221
Ameres SL, Horwich MD, Hung JH et al (2010) Target RNA-directed trimming and tailing of small silencing RNAs. Science 328:1534–1539
Ameres SL, Hung JH, Xu J et al (2011) Target RNA-directed tailing and trimming purifies the sorting of endo-siRNAs between the two Drosophila Argonaute proteins. RNA 17:54–63
Arikit S, Xia R, Kakrana A et al (2014) An atlas of soybean small RNAs identifies phased siRNAs from hundreds of coding genes. Plant Cell 26:4584–4601
Axtell MJ, Jan C, Rajagopalan R et al (2006) A two-hit trigger for siRNA biogenesis in plants. Cell 127:565–577
Baranauske S, Mickute M, Plotnikova A et al (2015) Functional mapping of the plant small RNA methyltransferase: HEN1 physically interacts with HYL1 and DICER-LIKE 1 proteins. Nucleic Acids Res 43:2802–2812
Baulcombe D (2004) RNA silencing in plants. Nature 431:356–363
Ben Chaabane S, Liu R, Chinnusamy V et al (2013) STA1, an Arabidopsis pre-mRNA processing factor 6 homolog, is a new player involved in miRNA biogenesis. Nucleic Acids Res 41:1984–1997
Billi AC, Alessi AF, Khivansara V et al (2012) The Caenorhabditis elegans HEN1 ortholog, HENN-1, methylates and stabilizes select subclasses of germline small RNAs. PLoS Genet 8:e1002617
Blevins T, Pontvianne F, Cocklin R et al (2014) A two-step process for epigenetic inheritance in Arabidopsis. Mol Cell 54:30–42
Blevins T, Podicheti R, Mishra V et al (2015) Identification of Pol IV and RDR2-dependent precursors of 24 nt siRNAs guiding de novo DNA methylation in Arabidopsis. elife 4:e09591
Bologna NG, Voinnet O (2014) The diversity, biogenesis, and activities of endogenous silencing small RNAs in Arabidopsis. Annu Rev Plant Biol 65:473–503
Borges F, Martienssen RA (2015) The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol 16:727–741
Borsani O, Zhu J, Verslues PE et al (2005) Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis. Cell 123:1279–1291
Burroughs AM, Ando Y, de Hoon MJ et al (2010) A comprehensive survey of 3′ animal miRNA modification events and a possible role for 3′ adenylation in modulating miRNA targeting effectiveness. Genome Res 20:1398–1410
Carlsbecker A, Lee JY, Roberts CJ et al (2010) Cell signalling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465:316–321
Castel SE, Martienssen RA (2013) RNA interference in the nucleus: roles for small RNAs in transcription, epigenetics and beyond. Nat Rev Genet 14:100–112
Chan CM, Zhou C, Brunzelle JS et al (2009a) Structural and biochemical insights into 2′-O-methylation at the 3′-terminal nucleotide of RNA by Hen1. Proc Natl Acad Sci USA 106:17699–17704
Chan CM, Zhou C, Huang RH (2009b) Reconstituting bacterial RNA repair and modification in vitro. Science 326:247
Chang HM, Triboulet R, Thornton JE et al (2013) A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28-let-7 pathway. Nature 497:244–248
Chellappan P, Xia J, Zhou X et al (2010) siRNAs from miRNA sites mediate DNA methylation of target genes. Nucleic Acids Res 38:6883–6894
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44
Chen T, Cui P, Xiong L (2015) The RNA-binding protein HOS5 and serine/arginine-rich proteins RS40 and RS41 participate in miRNA biogenesis in Arabidopsis. Nucleic Acids Res 43:8283–8298
Cho SK, Ben Chaabane S, Shah P et al (2014) COP1 E3 ligase protects HYL1 to retain microRNA biogenesis. Nat Commun 5:5867
Choi K, Kim J, Muller SY et al (2016) Regulation of microRNA-mediated developmental changes by the SWR1 chromatin remodeling complex. Plant Physiol 171:1128–1143
Coruh C, Shahid S, Axtell MJ (2014) Seeing the forest for the trees: annotating small RNA producing genes in plants. Curr Opin Plant Biol 18C:87–95
Dong Z, Han MH, Fedoroff N (2008) The RNA-binding proteins HYL1 and SE promote accurate in vitro processing of pri-miRNA by DCL1. Proc Natl Acad Sci USA 105:9970–9975
Dunoyer P, Brosnan CA, Schott G et al (2010) An endogenous, systemic RNAi pathway in plants. EMBO J 29:1699–1712
Fang Y, Spector DL (2007) Identification of nuclear dicing bodies containing proteins for microRNA biogenesis in living Arabidopsis plants. Curr Biol 17:818–823
Fang X, Cui Y, Li Y et al (2015a) Transcription and processing of primary microRNAs are coupled by elongator complex in Arabidopsis. Nat Plants 1:15075
Fang X, Shi Y, Liu X et al (2015b) CMA33/XCT regulates small RNA production through modulating the transcription of dicer-like genes in Arabidopsis. Mol Plant 8:1227–1236
Fei Q, Xia R, Meyers BC (2013) Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell 25:2400–2415
Francisco-Mangilet AG, Karlsson P, Kim MH et al (2015) THO2, a core member of the THO/TREX complex, is required for microRNA production in Arabidopsis. Plant J 82:1018–1029
Fujioka Y, Utsumi M, Ohba Y et al (2007) Location of a possible miRNA processing site in SmD3/SmB nuclear bodies in Arabidopsis. Plant Cell Physiol 48:1243–1253
Fukunaga R, Doudna JA (2009) dsRNA with 5′ overhangs contributes to endogenous and antiviral RNA silencing pathways in plants. EMBO J 28:545–555
Garcia D, Garcia S, Pontier D et al (2012) Ago hook and RNA helicase motifs underpin dual roles for SDE3 in antiviral defense and silencing of nonconserved intergenic regions. Mol Cell 48:109–120
Gasciolli V, Mallory AC, Bartel DP et al (2005) Partially redundant functions of Arabidopsis DICER-like enzymes and a role for DCL4 in producing trans-acting siRNAs. Curr Biol 15:1494–1500
Gregory BD, O'Malley RC, Lister R et al (2008) A link between RNA metabolism and silencing affecting Arabidopsis development. Dev Cell 14:854–866
Haag JR, Ream TS, Marasco M et al (2012) In vitro transcription activities of Pol IV, Pol V, and RDR2 reveal coupling of Pol IV and RDR2 for dsRNA synthesis in plant RNA silencing. Mol Cell 48:811–818
Haag JR, Brower-Toland B, Krieger EK et al (2014) Functional diversification of maize RNA polymerase IV and V subtypes via alternative catalytic subunits. Cell Rep 9:378–390
Hajheidari M, Farrona S, Huettel B et al (2012) CDKF;1 and CDKD protein kinases regulate phosphorylation of serine residues in the C-terminal domain of Arabidopsis RNA polymerase II. Plant Cell 24:1626–1642
Han BW, Hung JH, Weng Z et al (2011) The 3′-to-5′ exoribonuclease Nibbler shapes the 3′ ends of microRNAs bound to Drosophila Argonaute1. Curr Biol 21:1878–1887
Herr AJ, Jensen MB, Dalmay T et al (2005) RNA polymerase IV directs silencing of endogenous DNA. Science 308:118–120
Horwich MD, Li C, Matranga C et al (2007) The Drosophila RNA methyltransferase, DmHen1, modifies germline piRNAs and single-stranded siRNAs in RISC. Curr Biol 17:1265–1272
Huang Y, Ji LJ, Huang QC et al (2009) Structural insights into mechanisms of the small RNA methyltransferase HEN1. Nature 461:823–U886
Ibrahim F, Rohr J, Jeong WJ et al (2006) Untemplated oligoadenylation promotes degradation of RISC-cleaved transcripts. Science 314:1893–1893
Ibrahim F, Rymarquis LA, Kim EJ et al (2010) Uridylation of mature miRNAs and siRNAs by the MUT68 nucleotidyltransferase promotes their degradation in Chlamydomonas. Proc Natl Acad Sci USA 107:3906–3911
Johnson C, Kasprzewska A, Tennessen K et al (2009) Clusters and superclusters of phased small RNAs in the developing inflorescence of rice. Genome Res 19:1429–1440
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Kamminga LM, Luteijn MJ, den Broeder MJ et al (2010) Hen1 is required for oocyte development and piRNA stability in zebrafish. EMBO J 29:3688–3700
Kamminga LM, van Wolfswinkel JC, Luteijn MJ et al (2012) Differential impact of the HEN1 homolog HENN-1 on 21U and 26G RNAs in the germline of Caenorhabditis elegans. PLoS Genet 8:e1002702
Karlsson P, Christie MD, Seymour DK et al (2015) KH domain protein RCF3 is a tissue-biased regulator of the plant miRNA biogenesis cofactor HYL1. Proc Natl Acad Sci USA 112:14096–14101
Katiyar-Agarwal S, Morgan R, Dahlbeck D et al (2006) A pathogen-inducible endogenous siRNA in plant immunity. Proc Natl Acad Sci USA 103:18002–18007
Khraiwesh B, Arif MA, Seumel GI et al (2010) Transcriptional control of gene expression by microRNAs. Cell 140:111–122
Kim S, Yang JY, Xu J et al (2008) Two cap-binding proteins CBP20 and CBP80 are involved in processing primary microRNAs. Plant Cell Physiol 49:1634–1644
Kim W, Benhamed M, Servet C et al (2009) Histone acetyltransferase GCN5 interferes with the miRNA pathway in Arabidopsis. Cell Res 19:899–909
Kim YJ, Zheng B, Yu Y et al (2011) The role of Mediator in small and long noncoding RNA production in Arabidopsis thaliana. EMBO J 30:814–822
Kirino Y, Mourelatos Z (2007) The mouse homolog of HEN1 is a potential methylase for Piwi-interacting RNAs. RNA 13:1397–1401
Koster T, Meyer K, Weinholdt C et al (2014) Regulation of pri-miRNA processing by the hnRNP-like protein AtGRP7 in Arabidopsis. Nucleic Acids Res 42:9925–9936
Lange H, Holec S, Cognat V et al (2008) Degradation of a polyadenylated rRNA maturation by-product involves one of the three RRP6-like proteins in Arabidopsis thaliana. Mol Cell Biol 28:3038–3044
Laubinger S, Sachsenberg T, Zeller G et al (2008) Dual roles of the nuclear cap-binding complex and SERRATE in pre-mRNA splicing and microRNA processing in Arabidopsis thaliana. Proc Natl Acad Sci USA 105:8795–8800
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220
Law JA, Vashisht AA, Wohlschlegel JA et al (2011) SHH1, a homeodomain protein required for DNA methylation, as well as RDR2, RDM4, and chromatin remodeling factors, associate with RNA polymerase IV. PLoS Genet 7:e1002195
Law JA, Du J, Hale CJ et al (2013) Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498:385–389
Li J, Yang Z, Yu B et al (2005) Methylation protects miRNAs and siRNAs from a 3′-end uridylation activity in Arabidopsis. Curr Biol 15:1501–1507
Li S, Vandivier LE, Tu B et al (2015) Detection of Pol IV/RDR2-dependent transcripts at the genomic scale in Arabidopsis reveals features and regulation of siRNA biogenesis. Genome Res 25:235–245
Liu N, Abe M, Sabin LR et al (2011) The exoribonuclease Nibbler controls 3′ end processing of microRNAs in Drosophila. Curr Biol 21:1888–1893
Machida S, Yuan YA (2013) Crystal structure of Arabidopsis thaliana Dawdle forkhead-associated domain reveals a conserved phospho-threonine recognition cleft for dicer-like 1 binding. Mol Plant 6:1290–1300
Manavella PA, Hagmann J, Ott F et al (2012) Fast-forward genetics identifies plant CPL phosphatases as regulators of miRNA processing factor HYL1. Cell 151:859–870
Mari-Ordonez A, Marchais A, Etcheverry M et al (2013) Reconstructing de novo silencing of an active plant retrotransposon. Nat Genet 45:1029–1039
Matzke MA, Mosher RA (2014) RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat Rev Genet 15:394–408
McCue AD, Panda K, Nuthikattu S et al (2015) ARGONAUTE 6 bridges transposable element mRNA-derived siRNAs to the establishment of DNA methylation. EMBO J 34:20–35
Montgomery TA, Rim YS, Zhang C et al (2012) PIWI associated siRNAs and piRNAs specifically require the Caenorhabditis elegans HEN1 ortholog henn-1. PLoS Genet 8:e1002616
Niu D, Lii YE, Chellappan P et al (2016) miRNA863-3p sequentially targets negative immune regulator ARLPKs and positive regulator SERRATE upon bacterial infection. Nat Commun 7:11324
Nozawa M, Miura S, Nei M (2012) Origins and evolution of microRNA genes in plant species. Genome Biol Evol 4:230–239
Nuthikattu S, McCue AD, Panda K et al (2013) The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21-22 nucleotide small interfering RNAs. Plant Physiol 162:116–131
Ohara T, Sakaguchi Y, Suzuki T et al (2007) The 3′ termini of mouse Piwi-interacting RNAs are 2′-O-methylated. Nat Struct Mol Biol 14:349–350
Onodera Y, Haag JR, Ream T et al (2005) Plant nuclear RNA polymerase IV mediates siRNA and DNA methylation-dependent heterochromatin formation. Cell 120:613–622
Peragine A, Yoshikawa M, Wu G et al (2004) SGS3 and SGS2/SDE1/RDR6 are required for juvenile development and the production of trans-acting siRNAs in Arabidopsis. Genes Dev 18:2368–2379
Piriyapongsa J, Jordan IK (2008) Dual coding of siRNAs and miRNAs by plant transposable elements. RNA 14:814–821
Pontier D, Picart C, Roudier F et al (2012) NERD, a plant-specific GW protein, defines an additional RNAi-dependent chromatin-based pathway in Arabidopsis. Mol Cell 48:121–132
Raghuram B, Sheikh AH, Rustagi Y et al (2015) MicroRNA biogenesis factor DRB1 is a phosphorylation target of mitogen activated protein kinase MPK3 in both rice and Arabidopsis. FEBS J 282:521–536
Rajagopalan R, Vaucheret H, Trejo J et al (2006) A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev 20:3407–3425
Ramachandran V, Chen X (2008) Degradation of microRNAs by a family of exoribonucleases in Arabidopsis. Science 321:1490–1492
Ream TS, Haag JR, Wierzbicki AT et al (2009) Subunit compositions of the RNA-silencing enzymes Pol IV and Pol V reveal their origins as specialized forms of RNA polymerase II. Mol Cell 33:192–203
Ren G, Yu B (2012) Critical roles of RNA-binding proteins in miRNA biogenesis in Arabidopsis. RNA Biol 9:1424–1428
Ren G, Chen X, Yu B (2012a) Uridylation of miRNAs by HEN1 SUPPRESSOR1 in Arabidopsis. Curr Biol 22:695–700
Ren G, Xie M, Dou Y et al (2012b) Regulation of miRNA abundance by RNA binding protein TOUGH in Arabidopsis. Proc Natl Acad Sci USA 109:12817–12821
Ren G, Chen X, Yu B (2014a) Small RNAs meet their targets: when methylation defends miRNAs from uridylation. RNA Biol 11:1099–1104
Ren G, Xie M, Zhang S et al (2014b) Methylation protects microRNAs from an AGO1-associated activity that uridylates 5′ RNA fragments generated by AGO1 cleavage. Proc Natl Acad Sci USA 111:6365–6370
Ron M, Alandete Saez M, Eshed Williams L et al (2010) Proper regulation of a sperm-specific cis-nat-siRNA is essential for double fertilization in Arabidopsis. Genes Dev 24:1010–1021
Ronemus M, Vaughn MW, Martienssen RA (2006) MicroRNA-targeted and small interfering RNA-mediated mRNA degradation is regulated by argonaute, dicer, and RNA-dependent RNA polymerase in Arabidopsis. Plant Cell 18:1559–1574
Saito K, Sakaguchi Y, Suzuki T et al (2007) Pimet, the Drosophila homolog of HEN1, mediates 2′-O-methylation of Piwi- interacting RNAs at their 3′ ends. Genes Dev 21:1603–1608
Shen BZ, Goodman HM (2004) Uridine addition after microRNA-directed cleavage. Science 306:997–997
Shivaprasad PV, Chen HM, Patel K et al (2012) A microRNA superfamily regulates nucleotide binding site-leucine-rich repeats and other mRNAs. Plant Cell 24:859–874
Slotkin RK, Freeling M, Lisch D (2005) Heritable transposon silencing initiated by a naturally occurring transposon inverted duplication. Nat Genet 37:641–644
Smith LM, Pontes O, Searle I et al (2007) An SNF2 protein associated with nuclear RNA silencing and the spread of a silencing signal between cells in Arabidopsis. Plant Cell 19:1507–1521
Song X, Li P, Zhai J et al (2012a) Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J 69:462–474
Song X, Wang D, Ma L et al (2012b) Rice RNA-dependent RNA polymerase 6 acts in small RNA biogenesis and spikelet development. Plant J 71:378–389
Speth C, Willing EM, Rausch S et al (2013) RACK1 scaffold proteins influence miRNA abundance in Arabidopsis. Plant J 76:433–445
Sun Z, Guo T, Liu Y et al (2015) The roles of Arabidopsis CDF2 in transcriptional and posttranscriptional regulation of primary microRNAs. PLoS Genet 11:e1005598
Tu B, Liu L, Xu C et al (2015) Distinct and cooperative activities of HESO1 and URT1 nucleotidyl transferases in microRNA turnover in Arabidopsis. PLoS Genet 11:e1005119
Ustianenko D, Hrossova D, Potesil D et al (2013) Mammalian DIS3L2 exoribonuclease targets the uridylated precursors of let-7 miRNAs. RNA 19:1632–1638
Wang L, Song X, Gu L et al (2013) NOT2 proteins promote polymerase II-dependent transcription and interact with multiple MicroRNA biogenesis factors in Arabidopsis. Plant Cell 25:715–727
Wang X, Zhang S, Dou Y et al (2015) Synergistic and independent actions of multiple terminal nucleotidyl transferases in the 3′ tailing of small RNAs in Arabidopsis. PLoS Genet 11:e1005091
Wu L, Zhou H, Zhang Q et al (2010) DNA methylation mediated by a microRNA pathway. Mol Cell 38:465–475
Wu L, Mao L, Qi Y (2012) Roles of dicer-like and argonaute proteins in TAS-derived small interfering RNA-triggered DNA methylation. Plant Physiol 160:990–999
Wu X, Shi Y, Li J et al (2013) A role for the RNA-binding protein MOS2 in microRNA maturation in Arabidopsis. Cell Res 23:645–657
Wyman SK, Knouf EC, Parkin RK et al (2011) Post-transcriptional generation of miRNA variants by multiple nucleotidyl transferases contributes to miRNA transcriptome complexity. Genome Res 21:1450–1461
Xie M, Yu B (2015) siRNA-directed DNA methylation in plants. Curr Genomics 16:23–31
Xie ZX, Allen E, Fahlgren N et al (2005) Expression of Arabidopsis MIRNA genes. Plant Physiol 138:2145–2154
Xie M, Zhang S, Yu B (2015) microRNA biogenesis, degradation and activity in plants. Cell Mol Life Sci 72:87–99
Yan J, Gu Y, Jia X et al (2012) Effective small RNA destruction by the expression of a short tandem target mimic in Arabidopsis. Plant Cell 24:415–427
Yang Z, Ebright YW, Yu B et al (2006) HEN1 recognizes 21-24 nt small RNA duplexes and deposits a methyl group onto the 2′ OH of the 3′ terminal nucleotide. Nucleic Acids Res 34:667–675
Yang GD, Yan K, Wu BJ et al (2012) Genomewide analysis of intronic microRNAs in rice and Arabidopsis. J Genet 91:313–324
Yang DL, Zhang G, Tang K et al (2016) Dicer-independent RNA-directed DNA methylation in Arabidopsis. Cell Res 26:66–82
Ye R, Chen Z, Lian B et al (2016) A dicer-independent route for biogenesis of siRNAs that Direct DNA methylation in Arabidopsis. Mol Cell 61:222–235
Yoshikawa M, Peragine A, Park MY et al (2005) A pathway for the biogenesis of trans-acting siRNAs in Arabidopsis. Genes Dev 19:2164–2175
Yu B, Yang Z, Li J et al (2005) Methylation as a crucial step in plant microRNA biogenesis. Science 307:932–935
Yu B, Bi L, Zheng BL et al (2008) The FHA domain proteins DAWDLE in Arabidopsis and SNIP1 in humans act in small RNA biogenesis. Proc Natl Acad Sci USA 105:10073–10078
Zhai J, Jeong DH, De Paoli E et al (2011) MicroRNAs as master regulators of the plant NB-LRR defense gene family via the production of phased, trans-acting siRNAs. Genes Dev 25:2540–2553
Zhai J, Zhao Y, Simon SA et al (2013) Plant microRNAs display differential 3′ – truncation and tailing, modifications which are ARGONAUTE1-dependent and conserved across species. Plant Cell 25:2417–2428
Zhai J, Bischof S, Wang H et al (2015a) A one precursor one siRNA model for pol IV-dependent siRNA biogenesis. Cell 163:445–455
Zhai J, Zhang H, Arikit S et al (2015b) Spatiotemporally dynamic, cell-type-dependent premeiotic and meiotic phasiRNAs in maize anthers. Proc Natl Acad Sci USA 112:3146–3151
Zhan X, Wang B, Li H et al (2012) Arabidopsis proline-rich protein important for development and abiotic stress tolerance is involved in microRNA biogenesis. Proc Natl Acad Sci USA 109:18198–18203
Zhang X, Xia J, Lii YE et al (2012) Genome-wide analysis of plant nat-siRNAs reveals insights into their distribution, biogenesis and function. Genome Biol 13:R20
Zhang H, Ma ZY, Zeng L et al (2013a) DTF1 is a core component of RNA-directed DNA methylation and may assist in the recruitment of Pol IV. Proc Natl Acad Sci USA 110:8290–8295
Zhang S, Xie M, Ren G et al (2013b) CDC5, a DNA binding protein, positively regulates posttranscriptional processing and/or transcription of primary microRNA transcripts. Proc Natl Acad Sci USA 110:17588–17593
Zhang S, Liu Y, Yu B (2014) PRL1, an RNA-binding protein, positively regulates the accumulation of miRNAs and siRNAs in Arabidopsis. PLoS Genet 10:e1004841
Zhang S, Liu Y, Yu B (2015) New insights into pri-miRNA processing and accumulation in plants. Wiley Interdiscip Rev RNA 6:533–545
Zhao Y, Yu Y, Zhai J et al (2012) The Arabidopsis nucleotidyl transferase HESO1 uridylates unmethylated small RNAs to trigger their degradation. Curr Biol 22:689–694
Zheng B, Wang Z, Li S et al (2009) Intergenic transcription by RNA polymerase II coordinates Pol IV and Pol V in siRNA-directed transcriptional gene silencing in Arabidopsis. Genes Dev 23:2850–2860
Zhu H, Hu F, Wang R et al (2011) Arabidopsis Argonaute10 specifically sequesters miR166/165 to regulate shoot apical meristem development. Cell 145:242–256
Zubko E, Meyer P (2007) A natural antisense transcript of the Petunia hybrida Sho gene suggests a role for an antisense mechanism in cytokinin regulation. Plant J 52:1131–1139
Acknowledgement
This work was supported by Nebraska Soybean Board (Award 16R-05-3/3 #1706 to B.Y) and National Science Foundation (OIA-1557417 to B.Y).
Note
Due to the comprehensive nature of the review topic and limited space allowed for the paper, we apologize to the scientists whose work is not cited here.
No conflict interest declared.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing AG
About this chapter
Cite this chapter
Yu, Q., Liu, Y., Li, M., Yu, B. (2017). Small RNA Biogenesis and Degradation in Plants. In: Rajewsky, N., Jurga, S., Barciszewski, J. (eds) Plant Epigenetics. RNA Technologies. Springer, Cham. https://doi.org/10.1007/978-3-319-55520-1_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-55520-1_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-55519-5
Online ISBN: 978-3-319-55520-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)