Abstract
Many bacteria communicate with each other through the action of diffusible signal molecules, a process that has been termed quorum sensing (QS). QS acts to regulate diverse processes in different bacteria, to include the formation of biofilms, cellular differentiation, synthesis of antibiotics and other secondary metabolites, and the production of virulence factors in pathogens. Many bacteria use amphiphilic lipids of different chemical classes as signal molecules. N-acyl homoserine lactones, cis-2-unsaturated fatty acids, methyl esters of hydroxylated fatty acids, and tridecanone derivatives have been described in different Gram-negative organisms and gamma-butyrolactones in Gram-positive streptomycetes. A diverse range of mechanisms for perception and transduction of these signals has been described. Here we review these different signals, their mode of biosynthesis, and transduction pathways before going on to discuss interference of QS as a strategy for the control of bacterial disease.
Access provided by CONRICYT-eBooks. Download reference work entry PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Many bacteria communicate with each other through the action of diffusible signal molecules. Such cell-cell communication allows organisms to monitor aspects of their environment such as population density, a process that has been termed quorum sensing (QS). The elevated levels of signal concentration resulting from a higher local population density lead to activation of specific QS-regulated functions. Other growth conditions such as confinement to particular niches in which diffusion may be limited or exposure to conditions that affect signal production or stability can also affect the local concentration of signal molecules. The term QS is now used to describe cell-cell signaling in this range of different contexts (Platt and Fuqua 2010). QS allows a colony or group of organisms to act in a coordinated fashion to regulate diverse processes such as the formation of biofilms, cellular differentiation, synthesis of antibiotics and other secondary metabolites, and the production of virulence factors in pathogenic bacteria. The signal molecules synthesized by bacteria belong to a wide range of chemical classes to include amphiphilic lipids as well as peptides and carbohydrate derivatives. Multiple systems using different types of signal can often occur within a single organism. Equally, a diverse range of mechanisms for signal perception and transduction has been described.
As noted above, different amphiphilic lipids have been shown to act as bacterial cell-to-cell signals (Fig. 1). Indeed the most common signal molecules found in Gram-negative bacteria are the N-acyl homoserine lactones (N-AHLs) (Waters and Bassler 2005). The plant pathogen Ralstonia solanacearum has an additional signaling system mediated by methyl esters of 3-OH palmitic or myristic acids (Kai et al. 2015). Gram-negative bacteria from the order Xanthomonadales, which includes important plant and human pathogens, utilize cis-unsaturated fatty acids of the DSF (diffusible signal factor) family as signals (Ryan et al. 2015), whereas Vibrio cholerae utilizes (S)-3-hydroxytridecan-4-one (Higgins et al. 2007). Although many Gram-positive bacteria use amino acids or modified peptides as signals, actinomycetes such as Streptomyces species use gamma-butyrolactones (GBLs) (Takano 2006). In this overview, we will discuss each of these lipid-based signals, the pathways for their synthesis and turnover, and the signal transduction pathways that lead to specific alteration in bacterial behavior. We will go on to briefly discuss interference of QS as a strategy for the control of disease as part of a consideration of outstanding research questions. The reader is also directed to several other reviews in this area that focus on particular signaling systems, strategies for interference, or their applications in synthetic biology (Deng et al. 2011; LaSarre and Federle 2013; Biarnes-Carrera et al. 2015).
2 N-AHL-Mediated Signaling
N-AHLs comprise an invariant homoserine lactone ring attached to a fatty acid residue through an amide bond (see Fig. 1). The fatty acid moieties differ in chain length from 4 to 18 carbons, some can be unsaturated and often occur with hydroxy or keto groups at position 3 (Ng and Bassler 2009). These signals were first described in the marine bioluminescent bacterium Vibrio fischeri (now Photobacterium fischeri) in which QS regulates light production. A new class of N-AHL first described in Rhodopseudomonas palustris has p-coumaric acid instead of a fatty acid (Schaefer et al. 2008) but will not be considered further here.
2.1 The Archetypal LuxIR QS System
The archetypal N-AHL QS system comprises two proteins belonging to the LuxI and LuxR families, respectively (Fuqua et al. 2001; Whitehead et al. 2001; Ng and Bassler 2009; Fig. 2). N-AHLs are synthesized by cytoplasmic LuxI family proteins using S-adenosyl methionine and a fatty acyl-acyl carrier protein (fatty acyl-ACP) as substrates. The reaction generates the N-AHL and 5′-methylthioadenosine as products. LuxI proteins do not have a strict substrate specificity and will generate a range of N-AHLs with fatty acid substituents of similar chain length or additional substitution. The precise mode of action of several LuxI family proteins has been described (Pappas et al. 2004). After synthesis, the signal can move across the bacterial membranes and accumulates both intra- and extracellularly in proportion to cell number. It is not clear whether movement across the cytoplasmic membrane requires facilitation by transporter proteins.
The sensing of the N-AHL signal is mediated by a transcriptional regulator of the LuxR family. These proteins comprise two domains: an amino-terminal region involved in N-AHL binding and a C-terminal domain implicated in DNA binding (Pappas et al. 2004). Preferential binding of a particular N-AHL by its cognate LuxR family protein ensures a good degree of specificity in signal transduction and in most cases results in the formation of homodimers. These complexes can then bind at specific promoter DNA sequences called lux boxes affecting the expression of target QS-regulated genes.
The luxI and luxR genes are in most cases linked within the bacterial genome. Some organisms have several LuxI family proteins that direct the synthesis of N-AHL signal molecules sometimes with diverse acyl moieties. Each of these LuxI proteins has an associated LuxR protein, and the different LuxI/R systems usually interact extensively and are hierarchically organized. The best studied of these hierarchical systems is probably that of Pseudomonas aeruginosa (Jimenez et al. 2012). This organism has two N- AHL-QS systems, the Las and Rhl systems. LasI directs the synthesis of N-(3-oxo-dodecanoyl)-L-homoserine lactone (3-oxo-C12-HSL) which interacts with LasR and activates target promoters. RhlI directs the synthesis of N-(butanoyl)-L-homoserine lactone (C4-HSL) which interacts with the cognate regulator RhlR, thus activating its gene promoters. The Las and Rhl systems are connected and regulate the production of multiple virulence factors such as rhamnolipid, elastase, and pyocyanin production as well as biofilm formation (Jimenez et al. 2012).
The LasIR and RhlIR system also regulate the synthesis of PQS, (for Pseudomonas quinolone signal; 2-heptyl-3-hydroxy-4 (1H)-quinolone) and its precursor 2-heptyl-3-hydroxy-4(1H)-hydroxyquinolone (HHQ). Both PQS and HHQ act as QS signals in Pseudomonas aeruginosa, whereas other species of Pseudomonas and Burkholderia species do not synthesize PQS but use HHQ as a QS signal.
2.2 LuxMN Is a Second QS System
A second pathway of N-AHL-dependent QS that is distinct from LuxIR has been described in Vibrio harveyi (Waters and Bassler 2005). In this organism, synthesis of N-AHL (specifically 3-hydroxy butanoyl-homoserine lactone; 3-OH-BHL) is catalyzed by LuxM, which is unrelated to LuxI and sensed by a periplasmic loop of LuxN, a histidine kinase in the cytoplasmic membrane. This pathway acts in concert with two other QS pathways mediated by CAI-I, which in V. harveyi is (Z)-3-aminoundecan-4-one (see below) and AI-2, a furanosyl borate diester, to activate bioluminescence and inhibit exopolysaccharide production and type III secretion at high cell density. Each pathway involves a different histidine kinase, but they converge at the cytoplasmic phosphotransfer protein LuxU. This protein can exchange phosphoryl groups with the σ54 -dependent activator LuxO. At low cell density, signal concentration is low, and LuxN acts as a kinase resulting in autophosphorylation. The phosphoryl group is then transferred via LuxU to LuxO. This leads to activation of synthesis of small RNA species that together with the protein Hfq inhibit the transcription of luxR, which encodes an activator of bioluminescence (hence light is not produced) (Fig. 3).
QS circuitry in Vibrio harveyi. The major QS signals in Vibrio harveyi are CAI-1, which is (S)-3-hydroxytridecan-4-one, AI-2, a furanosyl borate diester, and 3-OH butanoyl homoserine lactone (3-OH-BHL). Synthesis of CAI-1 requires CqsA, whereas signal perception and transduction require the sensor kinase CqsS. These signaling pathways act in concert to repress virulence factor synthesis and promote bioluminescence when the cognate signals are present, an action mediated by the LuxR regulator. All three systems act via the LuxU phosphotransfer protein and the σ54 - dependent activator LuxO to modulate luxR expression. A “P” next to an arrow indicates the transfer of phosphoryl groups (see text for details)
At high cell density, the 3-hydroxy butanoyl-homoserine lactone, AI-2, and CAI-1 signal molecules are produced at a high level and interact with their cognate sensors. (In the case of AI-2, the signal binds to LuxP, a periplasmic protein that is associated with the sensor kinase LuxQ.) These interactions convert the sensor proteins (LuxN, LuxQ, and CqsS) from kinases to phosphatases, resulting in loss of phosphoryl groups from LuxU and LuxO. Consequently LuxO is inactivated, the small RNAs are not synthesized, and LuxR synthesis can proceed. This allows the LuxR transcriptional activator (which is unrelated to LuxR of Vibrio fischeri) to activate expression of bioluminescence genes.
2.3 Enzymatic Degradation of N-AHL Signals
A number of enzymes capable of degradation of N-AHL signals have been described (LaSarre and Federle 2013). The two principal classes are acylases that release the fatty acid from the homoserine moiety by hydrolysis and lactonases that open the homoserine lactone ring. A cytochrome P450 oxidoreductase from Bacillus megaterium can also act to oxidize N-AHLs at the ω-1, ω-2, or ω-3 position, as a first step to their further degradation (Chowdhary et al. 2007). These quorum-quenching enzymes can have diverse roles within the producing organisms. They may be involved in degradation of signals within a species, so that the organism is no longer subject to QS control. In contrast, they may also be involved in competition with other bacteria, where the enzymes degrade the N-AHL signals of the competitor to provide an advantage to the producing organism. There is a substantial interest in engineering the use of such enzymes in the control of bacterial disease. The first description of such quorum quenching was the expression of the lactonase AiiA in tobacco and potato to control of symptoms caused by Erwinia carotovora (now Pectobacterium carotovorum) (Dong et al. 2001). The expression of AiiA reduced the maceration symptoms which are normally caused by extracellular enzymes (pectinases and cellulases) that are under QS control. Further examples are discussed by LaSarre and Federle (2013).
3 DSF-Mediated Signaling
Cell-cell signals of the DSF (diffusible signal factor) family are cis-2-unsaturated fatty acids of different chain lengths and branching (Fig. 1). The first of these to be described was 11-methyl-cis-2-dodecenoic acid (which was named DSF) from the plant pathogen Xanthomonas campestris (Barber et al. 1997; Wang et al. 2004; Ryan et al. 2015). The cis-unsaturated double bond at the 2 position is a key structural feature for the signaling and regulatory activities; trans isomers and saturated derivatives have little or no activity (Wang et al. 2004). Different family members have been described in Burkholderia cenocepacia (cis-2-dodecenoic acid; BDSF), Pseudomonas aeruginosa (cis-2-decenoic acid), Xylella fastidiosa (cis-2-tetradecenoic acid; XfDSF), and Xanthomonas oryzae (cis,cis-11 methyldodeca-2,5-dienoic acid; CDSF) (Fig. 1).
3.1 The Rpf Proteins and DSF Signaling in Xanthomonads
In Xanthomonas, the synthesis and perception of the DSF signal require products of genes within the rpf cluster (for regulation of pathogenicity factors) (Barber et al. 1997; Slater et al. 2000). DSF signaling in Xanthomonas positively regulates the synthesis of multiple virulence factors including extracellular enzymes and extracellular polysaccharides but negatively regulates biofilm formation/aggregation. The synthesis of DSF is totally dependent on RpfF, which has amino acid sequence relatedness to enoyl CoA hydratase and is a member of the crotonase superfamily of proteins (Barber et al. 1997). The rpfF gene is downstream of and transcriptionally linked to rpfB, which encodes a long chain fatty acyl CoA ligase, although rpfF also has its own promoter. RpfB does not have a role in DSF synthesis but rather in DSF turnover (see below).
The sensing and transduction of the DSF signal in Xanthomonas depend upon a two-component regulatory system encoded by the rpfGHC operon, which is adjacent to rpfF but convergently transcribed (Ryan et al. 2015). RpfC is a complex sensor kinase comprising an N-terminal membrane-associated sensory input domain, histidine kinase and histidine kinase acceptor (HisKA) domains, a CheY-like receiver (REC) domain, and a C-terminal histidine phosphotransfer (HPT) domain. The RpfG regulator comprises a REC domain and an HD-GYP domain, which is a phosphodiesterase involved in degradation of the second messenger cyclic di-GMP (Ryan et al. 2015). RpfH is a novel protein with four transmembrane helices that has amino acid sequence similarity to the sensory input domain of RpfC but no known function (Slater et al. 2000). DSF signal transduction is believed to involve autophosphorylation of RpfC, followed by phosphorelay and finally phosphotransfer to the REC domain of the RpfG regulator. Phosphorylation of RpfG leads to its activation as a cyclic di-GMP phosphodiesterase and consequent alterations in the level of cyclic di-GMP in the cell, which influences the synthesis of virulence factors by diverse mechanisms (Ryan et al. 2015). A simplified scheme is shown in Fig. 4a. RpfC acts to positively regulate synthesis of virulence factors, but to negatively regulate DSF synthesis (Slater et al. 2000; Wang et al. 2004), although this requires neither phosphorelay nor involvement of RpfG (reviewed in He and Zhang 2008).
Two “core” pathways of signaling involving DSF family signals are exemplified by Xanthomonas and Burkholderia species (a, b, respectively). In both cases, the DSF signal is synthesized by an RpfF homolog and is linked to the turnover of the second messenger cyclic di-GMP. In xanthomonads (a), signal perception involves the sensor kinase RpfC and two-component regulator RpfG, which is an HD-GYP domain cyclic di-GMP phosphodiesterase. A “P” next to the arrow indicates the transfer of phosphoryl groups. In Burkholderia (b) signal sensing involves RpfR, a cytoplasmic GGDEF-EAL domain protein implicated in cyclic di-GMP degradation (see text for details)
Genome sequencing indicates the presence of a largely conserved rpf gene cluster in all xanthomonads as well as in Stenotrophomonas spp. some of which are opportunistic human pathogens. The rpfH gene is not fully conserved however. Furthermore a role for DSF signaling in the virulence of a number of these other bacteria has now been described.
Some of these organisms have an alternate sensor for DSF called RpfS, a histidine kinase that is predicted to be cytoplasmic and recognizes DSF through an N-terminal PAS sensor domain (An et al. 2014). RpfS is not widely conserved, however, and has been considered accessory to the core pathway in DSF transduction involving RpfGC .
3.2 A Second Core Pathway of DSF Signaling
A second core pathway in DSF family signal transduction was first identified in Burkholderia. The synthesis of the Burkholderia signal (BDSF) depends on a homolog of RpfF, but in this case signal perception depends upon RpfR, a protein with PAS, GGDEF, and EAL domains (Deng et al. 2012; Fig. 4b). GGDEF and EAL domains are implicated in the synthesis and degradation, respectively, of the second messenger cyclic di-GMP (Römling et al. 2013). In vitro, RpfR exhibits cyclic di-GMP phosphodiesterase activity that is modulated by binding of BDSF to the N-terminal PAS domain (Deng et al. 2012). The rpfF and rpfR genes are adjacent and convergently transcribed. The RpfR-RpfF system is widely conserved not only in Burkholderia species but also in bacteria from related genera such as Achromobacter and unrelated Enterobacteriaceae including Enterobacter, Cronobacter, Yersinia, and Serratia (Deng et al. 2012). Recently it has been reported that the RpfFR system of Cronobacter regulates a diverse range of functions (Suppiger et al. 2016). A second sensing system for BDSF in B. cenocepacia involves BCAM0227, a complex sensor kinase that is not a homolog of RpfC of Xanthomonas (McCarthy et al. 2010). However unlike RpfR, BCAM0227 is restricted to B. cenocepacia suggesting that it is an accessory sensor. Notably the two “core” pathways exemplified by RpfFR in B. cenocepacia and RpfFGC in X. campestris both link sensing of a DSF family signal to cyclic di-GMP turnover, but the mechanisms are completely different.
P. aeruginosa produces cis-2-decenoic acid (Fig. 1), a factor that can induce dispersion of biofilms produced by P. aeruginosa as well as other bacteria (Davies and Marques 2009). The enzyme responsible for the synthesis of cis-2-decenoic acid is an RpfF homolog called DspI (Amari et al. 2013). The dspI gene is located in a cluster of genes encoding enzymes implicated in fatty acid metabolism. P. aeruginosa does not have an rpfF-rpfC-rpfG or rpfF-rpfR gene cluster, and the identity of the sensor for this signal is not known. Homologs of DspI occur in over ten Pseudomonas species.
3.3 RpfF and Signal Synthesis
In vitro studies of the RpfF homolog from B. cenocepacia have shown that it is a bifunctional crotonase having both desaturase and thioesterase activity (Bi et al. 2012). This B. cenocepacia enzyme acts upon 3-hydroxylated fatty acyl-ACP, an intermediate of fatty acid biosynthesis, to produce BDSF (Bi et al. 2012). The Xanthomonas RpfF enzyme also exhibits activity as a broad specificity thioesterase and desaturase in vitro. RpfF is the only member of the crotonase superfamily with both desaturase and thioesterase activity. A model for the action of RpfF is that the enzyme first works as a dehydratase to convert 3-hydroxydodecanoyl-ACP to cis-2-dodecenoyl-ACP and then as a thioesterase to release free BDSF (cis-2-dodecenoic acid). RpfF can generate free saturated fatty acids from any fatty acyl-ACP substrate through its thioesterase activity. The synthesis of BDSF in the in vitro assay requires the addition of an exogenous acyl-ACP synthetase to reverse the thioesterase reaction. It is unclear how the two actions of dehydratase and thiosterase are coordinated in vivo to produce BDSF (Bi et al. 2012). The dual nature of RpfF is consistent with observations that mutation of rpfF affects the appearance in culture supernatants of saturated fatty acids as well as unsaturated fatty acids of the DSF family.
In vivo, individual bacteria can produce multiple DSF family signals that are all dependent on RpfF for their synthesis (see, e.g., He et al. 2010; Zhou et al. 2015a). This suggests that the enzyme does not have a strict specificity for a particular substrate. The available evidence suggests that the pattern of signals produced is not regulated by differences in specificity of different RpfF synthases but rather by the supply of different substrates. Consistent with this contention, in X. oryzae pv. oryzae, the rice pathogen produces three signals (DSF, BDSF, CDSF) with different time courses during growth and are present in different ratios depending on the culture medium (He et al. 2010). The substrate for DSF synthesis in vivo must be 11-methyl-3-hydroxydodecanoyl-ACP, with the 11-methyl substitution derived from leucine via the branched chain fatty acid synthetic pathway (Bi et al. 2012; Zhou et al. 2015a). Recent work has identified minor signals of the DSF family in Xanthomonas including cis-9-methyl-2-dodecenoic acid and cis-10-methyl-2-dodecenoic acid, where the 10-methyl substitution is probably derived from isoleucine (Deng et al. 2015; Zhou et al. 2015a).
Whether production of multiple DSF family signals by one organism has any biological relevance is unclear. For example, there are no reports that different signals induce different responses in the producing organisms. However the systems for sensing DSF family signals within a particular organism appear to be attuned to the major signal produced by that organism. For example, Xanthomonas and Xylella fastidiosa generate cis-11-methyl-dodecenoic acid and cis-2-tetradecenoic acid, respectively, as major signals, each is more responsive to its own signal than the heterologous one (Beaulieu et al. 2013).
3.4 RpfB and Signal Degradation
Although RpfB was originally thought to be involved in DSF synthesis, more recent work has established that it has a different role, acting in the mobilization of (saturated) free fatty acids generated by the thioesterase action of RpfF and in the degradation of DSF (Bi et al. 2014). Work in Xanthomonas has shown that RpfB, which is a predicted fatty acid CoA ligase, activates free saturated fatty acids allowing their use in phospholipid biosynthesis. In this way RpfB counteracts the thioesterase activity of RpfF. Although RpfB has little activity against BDSF or DSF in vitro (Bi et al. 2014; Zhou et al. 2015b), Xanthomonas cells can degrade exogenous DSF, and RpfB has a role in this process (Zhou et al. 2015b). It is suggested that in vivo, the substrate specificity of RpfB can be modulated by additional factors (cofactors, salts, or metals) or by an alteration in conformation which could conceivably involve interactions with other protein (Zhou et al. 2015b).
4 Gamma-Butyrolactone-Mediated Signaling
Gamma-butyrolactones (GBLs) have been identified as signaling molecules in Actinobacteria and principally in Streptomyces species (Takano 2006; Polkade et al. 2016). GBL signaling within different streptomycetes acts to regulate morphological differentiation and antibiotic production (Takano 2006). The first such molecule described was the autoregulatory factor or A-factor (2-isocapryloyl-3R-hydroxymethyl-gamma-butyrolactone) from Streptomyces griseus (Fig. 1). A-factor regulates streptomycin production and sporulation. The determination of the structure of a number of GBLs involved in signaling shows that these share the 3R-hydroxymethyl-gamma-butyrolactone moiety but differ in the length, branching, and stereochemistry of the fatty acid side chain which in general is specific for each species (Polkade et al. 2016). Nevertheless most of these molecules regulate the production of different antibiotics.
Different GBL signaling circuits occur in different Streptomyces species (Biarnes-Carrera et al. 2015). In S. griseus, A-factor synthesis is catalyzed by AfsA, and signal recognition involves ArpA, a repressor of the TetR family. Binding of the A-factor to the ArpA homodimer causes its release from the promoter of adpA, which encodes a master regulator of streptomycin synthesis and morphological differentiation (Fig. 5). The genes encoding the synthase and receptor for A-factor (afsA and arpA, respectively) are separated by 100 kb in the genome. This contrasts with what is seen in other Streptomyces species such as S. coelicolor, where the genes encoding the synthase and receptor are divergently transcribed and have overlapping promoters (Biarnes-Carrera et al. 2015).
A-factor (Gamma-butyrolactone) signaling in Streptomyces griseus. A-factor synthesis is catalyzed by AfsA, and signal recognition involves ArpA, a repressor of the TetR family. Binding of the A-factor to the ArpA homodimer causes its release from the promoter of adpA, which encodes a master regulator of streptomycin synthesis and morphological differentiation (see text for details)
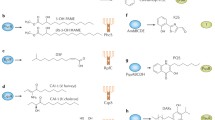
The proposed synthetic mechanism of A-factor synthesis by AfsA involves 8-methyl-3-oxononanoyl-acyl carrier protein and dihydroxyacetone phosphate (DHAP) as substrates (Kato et al. 2007). Beta-ketoacyl transfer to the hydroxyl group of DHAP catalyzed by AfsA produces 8-methyl-3-oxononanoyl-DHAP ester as a product. This ester is nonenzymatically converted to a butenolide phosphate by intramolecular aldol condensation. The butenolide phosphate is then reduced by BprA that was encoded just downstream of afsA. The phosphate group on the resultant butanolide is finally removed by a phosphatase, resulting in formation of A-factor. The 8-methyl-3-oxononanoyl-DHAP ester is also converted to A-factor by a second pathway. In this scheme, the phosphate group on the ester is first removed by a phosphatase, and the dephosphorylated product is converted nonenzymatically to a butenolide. Reduction of this butenolide by a reductase (different from BprA) generates A-factor. The ability of cloned afsA to direct production of an A-factor activity in Escherichia coli (Kato et al. 2007) suggests that AfsA is the key enzyme for the biosynthesis of GBLs and that the reductase(s) and phosphatase(s) are commonly present in bacteria and hence are not specific for A-factor biosynthesis.
As outlined above, the organization of genes involved in GBL signaling is different within different Streptomyces genomes. For example, in S. coelicolor, the gene encoding the receptor (scbR) is divergently transcribed from scbA, which encodes the synthase (Takano et al. 2001). Furthermore, the promoters of the two genes overlap, indicating the possible occurrence of transcriptional interference. Such an organization may be required to allow tight regulation of synthesis of prodigiosin and actinorhodin at relatively low GBL concentration.
5 Other Lipid-Based QS Systems
5.1 Hydroxylated Fatty Acid Esters in Ralstonia
The plant pathogen Ralstonia solanacearum produces the fatty acid esters 3-OH palmitic acid methyl ester (3-OH PAME) and (R)-methyl 3-hydroxymyristate [(R)-3-OH MAME] as QS signals (Clough et al. 1997; Kai et al. 2015). 3-OH PAME was originally detected in strain AW1 (Clough et al. 1997) but is not found in other strains that produce (R)-3-OH MAME instead (Kai et al. 2015). QS mediated by these molecules acts to regulate the synthesis of extracellular enzymes, extracellular polysaccharides, and secondary metabolites called ralfuranones, all which are virulence factors. The signals are synthesized by the methyltransferase PhcB, using S-adenosyl methionine and the 3-hydroxylated fatty acid-ACP as substrates. Signal sensing and transduction involve the membrane-associated histidine kinase PhcS and the two-component regulator PhcR. Interaction of PhcR and the transcriptional regulator PhcA acts to regulate virulence factor synthesis. The available evidence suggests marked differences in mechanistic detail between the AW1 strain, which responds to 3-OH PAME, and strains that respond to (R)-3-OH MAME. In the former, it is proposed that in the absence of signal, PhcS acts to phosphorylate PhcR thus negatively regulating PhcA. The presence of a threshold level of 3-OH PAME reduces the kinase activity of PhcS, so that PhcR becomes dephosphorylated thus relieving repression of PhcA. By contrast, in strains that respond to (R)-3-OH MAME, it is proposed that sensing of the signal leads to activation of autophosphorylation and phosphotransfer to PhcR. Phosphorylated PhcR is suggested to activate PhcA although the mechanism remains obscure. Intriguingly, the phylogenetic trees of the Phc proteins from R. solanacearum strains were divided into two groups, according to their QS signal types: (R)-3-OH MAME or (R)-3-OH PAME. An added complexity is that in the AW1 strain, PhcA also regulates an N-AHL-based QS system called SolIR (Whitehead et al. 2001).
5.2 (S)-3-hydroxytridecan-4-one in Vibrio cholerae
The major QS signal in Vibrio cholerae designated CAI-1 has been identified as (S)-3-hydroxytridecan-4-one (Higgins et al. 2007). This CAI-1-mediated system works in concert with a QS system mediated by AI-2 (a furanosyl borate diester) to regulate biofilm formation and virulence factor production. Synthesis of CAI-1 requires CqsA, whereas signal perception and transduction require the sensor kinase CqsS. This signaling system influences the phosphorylation level of LuxU and LuxO, which are also involved in AI-2 signaling. These signaling pathways act in concert to repress virulence factor synthesis and promote biofilm formation when the cognate signals are present (Fig. 6). These effects are exerted through an influence on transcription of the HapR regulator, which occurs in an analogous fashion to that described above for LuxR regulation in Vibrio harveyi.
QS circuitry in Vibrio cholerae. The major QS signals in Vibrio cholerae are CAI-1, which is (S)-3-hydroxytridecan-4-one and AI-2, a furanosyl borate diester. Synthesis of CAI-1 requires CqsA, whereas signal perception and transduction require the sensor kinase CqsS. This signaling system influences the phosphorylation level of LuxU and LuxO, which are also involved in AI-2 signaling. A “P” next to an arrow indicates the transfer of phosphoryl groups These signaling pathways act in concert to repress virulence factor synthesis and promote biofilm formation when the cognate signals are present, an action mediated by the HapR regulator (see text for details)
CqsA uses S-adenosyl methionine and decanoyl-CoA to produce 3-aminotridec-2-en-4-one, a reaction that depends upon pyridoxal phosphate as a cofactor (Wei et al. 2011). 3-aminotridec-2-en-4-one is converted to CAI-1 in two steps: spontaneous conversion to tridecane-3,4-dione followed by an enzyme-catalyzed conversion to CAI-1. Intriguingly, the CAI-1 signal produced by V. harveyi has been isolated as the CqsS ligand and identified as (Z)-3-aminoundecan-4-one (Ng et al. 2011). V. harveyi CqsA and CqsS are extremely selective for the production and detection, respectively, of this molecule, whereas the V. cholerae CqsA/CqsS can produce and sense both (Z)-3-aminoundecan-4-one and (S)-3-hydroxytridecan-4-one (Ng et al. 2011).
The examples discussed above (summarized in Table 1) deal with QS within individual organisms. In the next section we will consider the role of these molecules in interspecies and inter-kingdom signaling.
6 Interspecies and Inter-kingdom Signaling
It is now appreciated that many bacteria occur in polymicrobial communities; many human diseases are polymicrobial in nature (Short et al. 2014). Interactions between organisms in such communities can be mediated by a number of factors, to include the same signals that individual bacteria use for QS. The possibility of interplay between species that utilize the same class of QS signal is evident. However some bacteria can sense signals that they do not themselves synthesize, a process that has been termed eavesdropping. Such interspecies signaling can act to influence bacterial behavior, including biofilm formation and antibiotic tolerance, reflecting the impact that interspecies signaling may have on the efficacy of antibiotic therapies. It is anticipated that research in the next few years will reveal more of these mechanisms.
The next few years should also see an expansion of understanding of the role of QS signals in inter-kingdom signaling. This can occur between bacteria and microbes such as yeast, plants, or mammalian cells. For example, QS signals can act to inhibit morphological transitions in the dimorphic fungus Candida albicans (Hogan et al. 2004; Boon et al. 2008), can trigger or modulate host defense responses in both mammalian and plant cells (Teplitski et al. 2011; Schenk et al. 2014), and in some cases can act as virulence factors in their own right (Cooley et al. 2008).
7 Research Needs
Since QS acts in regulation of biofilm formation, antibiotic tolerance, and virulence factor synthesis in many pathogenic bacteria, interference with QS has received a great deal of attention as a route toward reducing the virulence of pathogens or making them more susceptible to existing antibiotics. Inhibition of QS could be effected at several steps: inhibition of signal synthesis, signal sequestration, signal degradation by enzymes (quorum quenching), or the inhibition of signal perception and transduction by small molecules. It has been proposed that since small molecule QS inhibitors would not kill bacterial cells, the target organisms would not develop resistance, although this view has been recently challenged (García-Contreras et al. 2016).
The determination of the structure of different signal synthases may further the rational design of inhibitors of their action. Currently a number of structures of LuxI family N-AHL synthases have been described; there is some information for DSF synthases of the RpfF family and a report on the structure CqsA.
In the field of quorum-quenching enzymes, interest has been largely focused on those degrading N-AHLs. Some work on degradation of DSF family signals has been reported, where a role for RpfB in Xanthomonas and CarAB in Pseudomonas has been indicated, although mechanistic details are sketchy. Strains that can degrade DSF can reduce severity of disease symptoms caused by Xanthomonas in brassica when applied to the leaves (Newman et al. 2008). Overexpression of QS signals causing pathogen confusion has also been suggested as a route to disease control (Fray et al. 1999; Mae et al. 2001). Accordingly, expression of RpfF in grape and citrus can reduce virulence and symptom production by Xylella fastidiosa and Xanthomonas citri pv. citri, respectively (Caserta et al. 2014; Lindow et al. 2014). Deployment of such methods (transgenic plants, strains with enhanced capacity for production or degradation of QS signals) in agriculture must take into account the broader issues of environmental impact, including beneficial plant-microbe interactions.
In addition to inhibition of signal synthesis, the identification of small molecule inhibitors of key steps in signal sensing and transduction may define lead compounds for new drugs (Curtis et al. 2014). Such compounds have been identified by screening libraries of structural analogues of inter- and intracellular signal molecules for action against particular signaling components or by screening of much larger libraries of chemical compounds. As indicated above, these molecules may not act as antibiotics per se but rather as inhibitors of virulence factor synthesis or potentiators of antibiotic action.
Finally the definition of elements of QS circuitry may have applications in synthetic biology (see, e.g., Wu et al. 2014; Biarnes-Carrera et al. 2015). In many nonpathogens, QS acts to regulate the synthesis of important secondary metabolites. N-AHL- and GBL-based circuitry may find applications in the field of synthetic biology in approaches for tight control and improved production of important and valuable microbial products.
References
Amari DT, Marques CNH, Davies DG (2013) The putative enoyl-coenzyme A hydratase DspI is required for production of the Pseudomonas aeruginosa biofilm dispersion autoinducer cis-2-decenoic acid. J Bacteriol 195:4600–4610
An SQ, Allan JH, McCarthy Y, Febrer M, Dow JM, Ryan RP (2014) The PAS domain-containing histidine kinase RpfS is a second sensor for the diffusible signal factor of Xanthomonas campestris. Mol Microbiol 92:586–597
Barber CE, Tang JL, Feng JX, Pan MQ, Wilson TJ, Slater H, Dow JM, Williams P, Daniels MJ (1997) A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol Microbiol 24:555–566
Beaulieu ED, Ionescu M, Chatterjee S, Yokota K, Trauner D, Lindow S (2013) Characterization of a diffusible signaling factor from Xylella fastidiosa. MBio 4:e00539–e00512
Bi H, Christensen QH, Feng Y, Wang H, Cronan JE (2012) The Burkholderia cenocepacia BDSF quorum sensing fatty acid is synthesized by a bifunctional crotonase homologue having both dehydratase and thioesterase activities. Mol Microbiol 83:840–855
Bi H, Yu Y, Dong H, Wang H, Cronan JE (2014) Xanthomonas campestris RpfB is a fatty acyl-CoA ligase required to counteract the thioesterase activity of the RpfF diffusible signal factor (DSF) synthase. Mol Microbiol 93:262–275
Biarnes-Carrera M, Breitling R, Takano E (2015) Butyrolactone signalling circuits for synthetic biology. Curr Opin Chem Biol 28:91–98
Boon C, Deng Y, Wang LH, He Y, Xu JL, Fan Y, Pan SQ, Zhang LH (2008) A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J 2:27–36
Caserta R, Picchi SC, Takita MA, Tomaz JP, Pereira WE, Machado MA, Ionescu M, Lindow S, De Souza AA (2014) Expression of Xylella fastidiosa RpfF in citrus disrupts signaling in Xanthomonas citri subsp. citri and thereby its virulence. Mol Plant-Microbe Interact 27:1241–1252
Chowdhary PK, Keshavan N, Nguyen HQ, Peterson JA, González JE, Haines DC (2007) Bacillus megaterium CYP102A1 oxidation of acyl homoserine lactones and acyl homoserines. Biochemistry 46:14429–14437
Clough SJ, Lee KE, Schell MA, Denny TP (1997) A two-component system in Ralstonia (Pseudomonas) solanacearum modulates production of PhcA-regulated virulence factors in response to 3-hydroxypalmitic acid methyl ester. J Bacteriol 179:3639–3648
Cooley M, Chhabra SR, Williams P (2008) N-acylhomoserine lactone-mediated quorum sensing: a twist in the tail and a blow for host immunity. Chem Biol 15:1141–1147
Curtis MM, Russell R, Moreira CG, Adebesin AM, Wang C, Williams NS, Taussig R, Stewart D, Zimmern P, Lu B, Prasad RN, Zhu C, Rasko DA, Huntley JF, Falck JR, Sperandio V (2014) QseC inhibitors as an antivirulence approach for gram-negative pathogens. MBio 5:e02165
Davies DG, Marques CNH (2009) A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J Bacteriol 191:1393–1403
Deng Y, Wu J, Tao F, Zhang LH (2011) Listening to a new language: DSF-based quorum sensing in gram-negative bacteria. Chem Rev 111:160–173
Deng Y, Schmid N, Wang C, Wang J, Pessi G, Wu D, Lee J, Aguilar C, Ahrens CH, Chang C, Song H, Eberl L, Zhang LH (2012) Cis-2-dodecenoic acid receptor RpfR links quorum-sensing signal perception with regulation of virulence through cyclic dimeric guanosine monophosphate turnover. Proc Natl Acad Sci U S A 109:15479–15484
Deng Y, Liu X, Wu J, Lee J, Chen S, Cheng Y, Zhang C, Zhang LH (2015) The host plant metabolite glucose is the precursor of diffusible signal factor (DSF) family signals in Xanthomonas campestris. Appl Environ Microbiol 81:2861–2868
Dong YH, Wang LH, Xu JL, Zhang HB, Zhang XF, Zhang LH (2001) Quenching quorum-sensing-dependent bacterial infection by an N-acyl homoserine lactonase. Nature 411:813–817
Fray RG, Throup JP, Daykin M, Wallace A, Williams P, Stewart GS, Grierson D (1999) Plants genetically modified to produce N-acylhomoserine lactones communicate with bacteria. Nat Biotechnol 17:1017–1020
Fuqua C, Parsek MR, Greenberg EP (2001) Regulation of gene expression by cell-to-cell communication: acyl-homoserine lactone quorum sensing. Annu Rev Genet 35:439–468
García-Contreras R, Maeda T, Wood TK (2016) Can resistance against quorum-sensing interference be selected? ISME J 10:4–10
He YW, Zhang LH (2008) Quorum sensing and virulence regulation in Xanthomonas campestris. FEMS Microbiol Rev 32:842–857
He Y-W, Je W, Cha J-S, Zhang L-H (2010) Rice bacterial blight pathogen Xanthomonas oryzae pv. oryzae produces multiple DSF-family signals in regulation of virulence factor production. BMC Microbiol 10:187
Higgins DA, Pomianek ME, Kraml CM, Taylor RK, Semmelhack MF, Bassler BL (2007) The major Vibrio cholerae autoinducer and its role in virulence factor production. Nature 450:883–886
Hogan DA, Vik A, Kolter R (2004) A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol Microbiol 54:1212–1223
Jimenez PN, Koch G, Thompson JA, Xavier KB, Cool RH, Quax WJ (2012) The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol Mol Biol Rev 76:46–65
Kai K, Ohnishi H, Shimatani M, Ishikawa S, Mori Y, Kiba A, Ohnishi K, Tabuchi M, Hikichi Y (2015) Methyl 3-hydroxymyristate, a diffusible signal mediating phc quorum sensing in Ralstonia solanacearum. Chembiochem 16:2309–2318
Kato JY, Funa N, Watanabe H, Ohnishi Y, Horinouchi S (2007) Biosynthesis of gamma-butyrolactone autoregulators that switch on secondary metabolism and morphological development in Streptomyces. Proc Natl Acad Sci U S A 104:2378–2383
LaSarre B, Federle MJ (2013) Exploiting quorum sensing to confuse bacterial pathogens. Microbiol Mol Biol Rev 77:73–111
Lindow S, Newman K, Chatterjee S, Baccari C, Lavarone AT, Ionescu M (2014) Production of Xylella fastidiosa diffusible signal factor in transgenic grape causes pathogen confusion and reduction in severity of Pierce’s disease. Mol Plant-Microbe Interact 27:244–254
Mae A, Montesano M, Koiv V, Palva ET (2001) Transgenic plants producing the bacterial pheromone N-acyl-homoserine lactone exhibit enhanced resistance to the bacterial phytopathogen Erwinia carotovora. Mol Plant-Microbe Interact 14:1035–1042
McCarthy Y, Yang L, Twomey KB, Sass A, Tolker-Nielsen T, Mahenthiralingam E, Dow JM, Ryan RP (2010) A sensor kinase recognizing the cell-cell signal BDSF (cis-2-dodecenoic acid) regulates virulence in Burkholderia cenocepacia. Mol Microbiol 77:1220–1236
Newman KL, Chatterjee S, Ho KA, Lindow SE (2008) Virulence of plant pathogenic bacteria attenuated by degradation of fatty acid cell-to-cell signaling factors. Mol Plant-Microbe Interact 21:326–334
Ng WL, Bassler BL (2009) Bacterial quorum-sensing network architectures. Annu Rev Genet 43:197–222
Ng WL, Perez LJ, Wei Y, Kraml C, Semmelhack MF, Bassler BL (2011) Signal production and detection specificity in Vibrio CqsA/CqsS quorum-sensing systems. Mol Microbiol 79:1407–1417
Pappas KM, Weingart CL, Winans SC (2004) Chemical communication in proteobacteria: biochemical and structural studies of signal synthases and receptors required for intercellular signalling. Mol Microbiol 53:755–769
Platt TG, Fuqua C (2010) What’s in a name? The semantics of quorum sensing. Trends Microbiol 18:383–387
Polkade AV, Mantri SS, Patwekar UJ, Jangid K (2016) Quorum sensing: an under-explored phenomenon in the phylum actinobacteria. Front Microbiol 7:131
Römling U, Galperin MY, Gomelsky M (2013) Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol Mol Biol Rev 77:1–52
Ryan RP, An SQ, Allan JH, McCarthy Y, Dow JM (2015) The DSF family of cell-cell signals: an expanding class of bacterial virulence regulators. PLoS Pathog 11:e1004986
Schaefer AL, Greenberg EP, Oliver CM, Oda Y, Huang JJ, Bittan-Banin G, Peres CM, Schmidt S, Juhaszova K, Sufrin JR, Harwood CS (2008) A new class of homoserine lactone quorum-sensing signals. Nature 454:595–599
Schenk ST, Hernández-Reyes C, Samans B, Stein E, Neumann C, Schikora M, Reichelt M, Mithöfer A, Becker A, Kogel KH, Schikora A (2014) N-acyl-homoserine lactone primes plants for cell wall reinforcement and induces resistance to bacterial pathogens via the salicylic acid/oxylipin pathway. Plant Cell 26:2708–2723
Short FL, Murdoch SL, Ryan RP (2014) Polybacterial human disease: the ills of social networking. Trends Microbiol 22:508–516
Slater H, Alvarez-Morales A, Barber CE, Daniels MJ, Dow JM (2000) A two-component system involving an HD-GYP domain protein links cell-cell signalling to pathogenicity gene expression in Xanthomonas campestris. Mol Microbiol 38:986–1003
Suppiger A, Eshwar AK, Stephan R, Kaever V, Eberl L, Lehner A (2016) The DSF type quorum sensing signalling system RpfF/R regulates diverse phenotypes in the opportunistic pathogen cronobacter. Sci Rep 6:18753
Takano E (2006) Gamma-butyrolactones: streptomyces signalling molecules regulating antibiotic production and differentiation. Curr Opin Microbiol 9:287–294
Takano E, Chakraburtty R, Nihira T, Yamada Y, Bibb MJ (2001) A complex role for the gamma-butyrolactone SCB1 in regulating antibiotic production in Streptomyces coelicolor A3(2). Mol Microbiol 41:1015–1028
Teplitski M, Mathesius U, Rumbaugh KP (2011) Perception and degradation of N-acyl homoserine lactone quorum sensing signals by mammalian and plant cells. Chem Rev 111:100–116
Wang LH, He Y, Gao Y, Wu JE, Dong YH, He C, Wang SX, Weng LX, Xu JL, Tay L, Fang RX, Zhang LH (2004) A bacterial cell-cell communication signal with cross-kingdom structural analogues. Mol Microbiol 51:903–912
Waters CM, Bassler BL (2005) Quorum sensing: cell-to-cell communication in bacteria. Annu Rev Cell Dev Biol 21:319–346
Wei Y, Perez LJ, Ng WL, Semmelhack MF, Bassler BL (2011) Mechanism of Vibrio cholerae autoinducer-1 biosynthesis. ACS Chem Biol 6:356–365
Whitehead NA, Barnard AM, Slater H, Simpson NJ, Salmond GP (2001) Quorum-sensing in gram-negative bacteria. FEMS Microbiol Rev 25:365–404
Wu F, Menn DJ, Wang X (2014) Quorum-sensing crosstalk-driven synthetic circuits: from unimodality to trimodality. Chem Biol 21:1629–1638
Zhou L, Yu Y, Chen X, Diab AA, Ruan L, He J, Wang H, He YW (2015a) The multiple DSF-family QS signals are synthesized from carbohydrate and branched-chain amino acids via the FAS elongation cycle. Sci Rep 5:13294
Zhou L, Wang XY, Sun S, Yang LC, Jiang BL, He YW (2015b) Identification and characterization of naturally occurring DSF-family quorum sensing signal turnover system in the phytopathogen Xanthomonas. Environ Microbiol 17:4646–4658
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer International Publishing AG, part of Springer Nature
About this entry
Cite this entry
Dow, M., Naughton, L.M. (2018). Amphiphilic Lipids, Signaling Molecules, and Quorum Sensing. In: Krell, T. (eds) Cellular Ecophysiology of Microbe: Hydrocarbon and Lipid Interactions. Handbook of Hydrocarbon and Lipid Microbiology . Springer, Cham. https://doi.org/10.1007/978-3-319-50542-8_31
Download citation
DOI: https://doi.org/10.1007/978-3-319-50542-8_31
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-50540-4
Online ISBN: 978-3-319-50542-8
eBook Packages: Biomedical and Life SciencesReference Module Biomedical and Life Sciences