Abstract
For the past three decades, many ageing-regulatory pathways have been identified using C. elegans as a model organism. The insulin/insulin-like growth factor (IGF)-1 signalling (IIS) pathway is one of the most evolutionarily well-conserved ageing-regulatory pathways ranging from worms to mammals. Here, we review the molecular mechanism and the functional significance of IIS in C. elegans ageing. Specifically, we describe the roles of key components of IIS in ageing, systemic ageing regulation by IIS, and other known physiological functions of IIS that contribute to longevity. We also discuss possible implications of IIS in mammalian health and ageing.
*Author contributed equally with all other contributors.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
C. elegans insulin/insulin-like growth factor (IGF)-1 signalling (IIS) is one of the most established ageing -regulatory pathways, whose components have been extensively studied. In C. elegans , IIS is also important for resistance against various stresses, and this is consistent with many findings showing that enhanced stress resistance contributes to longevity. In addition, decreased levels of IIS prevent protein aggregation and delay the onset of many ageing-associated disease models in C. elegans. The function of IIS as a lifespan -regulatory pathway is evolutionarily conserved in Drosophila, mice, and very likely, in humans [1, 2]. In this chapter, we will describe mechanisms by which IIS plays roles in the regulation of ageing, stress resistance, and age-associated disease models . Further, we will discuss the implications that these findings in C. elegans have on human ageing.
4.2 Components That Influence Lifespan in the Insulin/IGF-1 Signalling Pathway
The IIS pathway is composed of various signal-transducing factors, and the role of each component in lifespan regulation is relatively well-characterized in C. elegans (Fig. 4.1). age-1 mutants were the first long-lived IIS mutants identified through a genetic screen [3, 4]. Subsequently, daf-2 mutants, which have been known to display phenotypes in the development of dauer (an alternative diapause larva, discussed in Chap. 3), were shown to live twice as long as wild-type C. elegans [5]. age-1 and daf-2 were eventually shown to encode a phosphoinositide-3 kinase (PI3K) and an insulin/IGF-1 receptor, respectively [6, 7]; these are the key upstream components of IIS. Since then, many more factors that act downstream of the IIS pathway have been identified in C. elegans.
Reduced IIS increases lifespan in C . elegans . Inhibition of DAF-2/insulin/IGF-1 receptor decreases the PI(3,4,5)P3/PI(4,5)P2 ratio through down-regulation of AGE-1/PI3 kinase, whose function is antagonized by the activation of DAF-18/PTEN. This decrease leads to the inactivation of PDK-1 and AKT-1/2, which subsequently promotes the nuclear translocation and activation of DAF-16/FOXO , and SKN-1/NRF2 transcription factors . HSF-1/heat shock factor 1 also collaborates with DAF-16 in the nucleus. These transcription factors regulate the expression of various genes that contribute to longevity in C. elegans
Inhibition of IIS promotes long lifespan in C. elegans. Specifically, the reduced function of DAF-2 results in the inactivation of the downstream kinase cascade, starting from AGE-1/PI3K [[8]; reviewed in [9]]. Down-regulation of AGE-1 then leads to the inactivation of 3-phosphoinositide-dependent kinase 1 (PDK-1) [10], likely through a decrease in the PI(3, 4, 5)P3/PI(4, 5)P2 ratio [11]. This, in turn, down-regulates the Akt/protein kinase B (PKB) family members, AKT-1 and AKT-2 [10, 12]. The PI(3, 4, 5)P3/PI(4, 5)P2 ratio can also be decreased by the activation of DAF-18/phosphatase and tensin (PTEN) phosphatase, which mediates dephosphorylation of PI(3, 4, 5)P3 and increases lifespan [8, 13–17]. Down-regulation of IIS also leads to the activation of transcription factors , which up-regulate the expression of various target genes that contribute to longevity, including chaperones, antioxidants , and antimicrobials. The representative longevity transcription factors downstream of IIS are DAF-16/Forkhead box O (FOXO), heat-shock transcription factor-1 (HSF-1), and skinhead-1 (SKN-1)/Nuclear factor-erythroid-related factor (Nrf).
DAF-16
DAF-16 is a FOXO transcription factor homologue [18, 19] that mediates a diverse array of cellular processes by regulating the expression of numerous genes, including those involved in ageing [20–25]. A variety of post-transcriptional regulators of this protein, including protein kinases and phosphatases, have been identified. Both AKT-1 and AKT-2 phosphorylate and inactivate DAF-16 by preventing nuclear translocation [26–30]. Phosphorylation of DAF-16 by serum/glucocorticoid-inducible kinase 1 (SGK-1)/SGK was also shown to obstruct the translocation into the nucleus [30]. However, subsequent studies using a sgk-1 gain-of-function mutant or overexpression of sgk-1 indicate that SGK-1 may activate DAF-16 [31, 32]. AMP (5′ adenosine monophosphate)-activated protein kinase (AAK-2) can also activate DAF-16 by phosphorylation and increases lifespan [33–36]. Similarly, CST-1/MST kinase and JNK-1/c-Jun N-terminal kinase phosphorylate and up-regulate DAF-16 to extend lifespan [37, 38]. Protein phosphatases also appear to regulate the activity of DAF-16 directly or indirectly. For example, SMK-1/suppressor of MEK null (SMEK), a homologue of the protein phosphatase 4 regulatory subunit, is required for the long lifespan of daf-2 mutants in a daf-16-dependent manner [39]. PPTR-1/protein phosphatase 2A regulatory subunit (PP2A) decreases the phosphorylation of AKT-1 and leads to both activation of DAF-16 and increased longevity in daf-2 mutants [40].
Other regulatory modes for DAF-16 include protein acetylation, protein stability control, protein-protein interactions, and transcriptional control of its isoforms. CBP-1/CREB-binding protein (CBP), which is an acetyl-transferase, contributes to the longevity of daf-2 mutants [41], likely via acetylating and activating DAF-16 [42]. DAF-16 is also required for the long lifespan conferred by the overexpression of sir-2.1/NAD-dependent protein deacetylases [[43–45] but see also [46]]. Components of the ubiquitin proteasome system regulate the stability and activity of DAF-16. Specifically, an E3 ligase, RLE-1/RC3H1, ubiquitinates DAF-16, and consequently, rle-1 mutants live long due to increased stability of DAF-16 [47]. MATH-33/deubiquitylase counteracts the RLE-1-dependent degradation of DAF-16 and extends lifespan [48]. In addition, components of the Skp1-Cul1-F-Box E3 ligase complex contribute to the longevity of daf-2 mutants, perhaps by indirectly up-regulating DAF-16 [49]. Additionally, proteasome activation promotes long lifespan by increasing DAF-16 activity [50]. Scaffold proteins are also important for DAF-16 regulation. Genetic inhibition of the 14-3-3 scaffold protein, PAR-5 or FTT-2, up-regulates DAF-16 by promoting its nuclear translocation [44, 51]. However, overexpression of these proteins paradoxically extends lifespan in a daf-16-dependent manner [52]. Another scaffold protein, SHC-1/Shc-like protein, promotes the nuclear localization of DAF-16 by acting upstream of JNK-1 [53]. In addition to these post-translational modes for regulation, the expression of different DAF-16 isoforms can be regulated at the transcription level [54, 55].
DAF-16 regulates the expression of its target genes by binding to specific DNA motifs: the DAF-16-binding element (DBE) and the DAF-16-associated element (DAE). The DBE was first identified using an iterative in vitro method, and the core sequence, TTGTTTAC, is located upstream of DAF-16 target genes [56]. DAE is a GATA sequence, CTTATCA, which is located within the promoters of many DAF-16 target genes [21, 57–59].
Several factors affect the downstream targets of DAF-16. For example, the PQM-1, a C2H2-type zinc finger and leucine zipper-containing transcriptional activator, increases the expression of DAF-16 targets by translocating in the opposite direction of DAF-16 in cells, and contributes to daf-2 mutant longevity [59]. The ELT-2 and ELT-3/GATA factors, and MDT-15/mediator 15, also induce the expression of DAF-16 target genes [57, 60]. The XBP-1/bZIP transcription factor , along with DAF-16, enhances the expression of the DOX-1/Zn-finger protein [61]. Conversely, the ETS-4/ETS transcription factor alters the expression of a subset of DAF-16 target genes to promote longevity via a non-canonical IIS [62]. In addition, DAF-16 requires other cofactors to induce target gene expression; these include the HEL-1/RNA helicase [63], the PRMT-1/type I protein arginine methyltransferase [64], and the SWI/SNF/chromatin remodeler [65].
HSF-1
HSF-1 is a heat-shock transcription factor that induces transcription of chaperone genes and proteasome-related genes in response to various stresses, including heat [reviewed in [66]]. HSF-1 collaborates with DAF-16 to promote longevity that results from reduced IIS activity [67]. Inhibition of hsf-1 decreases the long lifespan of daf-2 and age-1 mutants, and conversely overexpression of hsf-1 is sufficient to increase lifespan [67, 68]. Neuron -, muscle -, or intestine -specific overexpression of hsf-1 is also sufficient to extend lifespan [68, 69]. Experiments involving the temporal knockdown of hsf-1 indicate that HSF-1 expression during larval stages is more crucial than during adulthood [70]; this result, however, is in contrast to the observation that DAF-16 is required during adulthood for daf-2 mutant longevity [71]. HSF-1 regulates the expression of its target genes by binding to the heat-shock element (HSE), GAANNTTCNNGAA [72]. Together with DAF-16, HSF-1 regulates the expression of chaperone genes, including small heat-shock protein-encoding genes, which contribute to the longevity of daf-2 mutants [21, 67, 68, 73, 74]. Moreover, truncated HSF-1 overexpression increases lifespan by improving actin cytoskeletal integrity, independently of typical molecular chaperone functions [75].
Several regulators of HSF-1 in IIS have been discovered. These include daf-16-dependent longevity-1 and -2 (DDL-1 and -2), which inhibit HSF-1 activity through the formation of a DDL-1-containing HSF-1-inhibitory complex (DHIC) [74]. Under reduced IIS conditions, DDL-1 is phosphorylated, and DHIC is dissociated to activate HSF-1 for lifespan extension [74]. DAF-41/co-chaperone p23 regulates lifespan via HSF-1, as well as DAF-16, at high temperature [76]. HSF-1 has also been shown to act as a hub protein that mediates crosstalk between IIS and target of rapamycin (TOR) signalling pathways [77]. Overall, HSF-1 is a key regulator for IIS-mediated longevity and appears to be as important as DAF-16.
SKN-1
Another crucial longevity-promoting transcription factor in IIS is SKN-1 [reviewed in [78]], an oxidative stress-responsive Nrf transcription factor [79]. Genetic inhibition of skn-1 largely suppresses the long lifespan of daf-2 mutants [80], and skn-1 overexpression is sufficient to promote long lifespan [80]. Elimination of a putative AKT phosphorylation site enhances the nuclear translocation of SKN-1. Therefore, similar to DAF-16, dephosphorylated and nuclear-localized SKN-1 appears to promote longevity under conditions of reduced IIS [80]. SKN-1 regulates the expression of a number of genes involved in several stress responses [80–83] and protein translation [84, 85], many of which overlap with DAF-16 target genes [80, 84]. SKN-1 also up-regulates collagens to promote longevity by extracellular matrix (ECM) remodelling [86].
Various additional factors that affect the activity of DAF-16, HSF-1, and SKN-1, or the expression of their target genes, have been identified. Many of these additional factors work together to regulate the activity of the transcription factors in IIS-mediated longevity. Some of the molecular mechanisms by which these transcription factors are regulated have been revealed; however, most remain incompletely understood. Therefore, further research on these crucial transcription factors will be required to understand the fundamental mechanisms of IIS-mediated ageing regulation in C. elegans .
4.3 Sensory Neural Regulation of Longevity
C. elegans has a simple nervous system , comprised of 302 neurons, which have been mapped in detail [87] (see also Chaps. 2 and 8). Well-known functions of sensory neurons include the perception of environmental stimuli and the transmission of signals for proper physiological responses. Interestingly, sensory neurons in C. elegans also contribute to lifespan regulation [reviewed in [88]]. Chemosensory neurons appear to affect lifespan mostly by acting through IIS [89], whereas thermosensory neurons regulate lifespan via steroid signalling at high temperature [90]. Impairment of general chemosensory neuronal functions leads to the activation of DAF-16 and longevity via modulating the expression of insulin-like peptides (ILPs); chemosensory mutations also do not further extend the longevity of daf-2 mutants [27, 89, 91–93]. Thus, it is likely that the inhibition of chemosensory neurons down-regulates IIS activity, and this may in turn activate DAF-16 to promote longevity (Fig. 4.2).
Neuroendocrine regulation of IIS and longevity. Inhibition of sensory neural functions leads to down-regulation of IIS. This inhibition modulates the expression of hormonal insulin-like peptides that are secreted from sensory neurons , triggering the activation of DAF-16 in non-neuronal tissues, such as the intestine. Activated DAF-16 then translocates into the nucleus, where it induces the expression of target genes that confer organismal longevity
Inhibition of various components required for chemosensory neural function increases lifespan. These include the calcium-regulated neurosecretory factors, G-protein coupled receptors, G-proteins, cyclic nucleotide-gated channel subunits , and proteins that function in sensory signal transduction and synaptic transmission [89, 91, 92, 94–99]. Additionally, it has been shown that the induction of mct-1, a putative monocarboxylate transporter for small molecule trafficking, mediates the long lifespan of sensory mutants [100]. Further, a thermosensitive TRP channel, TRPA-1, increases lifespan by activating DAF-16 at lower temperatures in C. elegans [32, 101]. A recent study also demonstrated that food-derived chemosensory cues decrease lifespan via stimulating sensory neurons, which in turn increases the expression of an ILP/INS-6 that acts as an endocrine IIS-activating signal [93].
4.4 Endocrine Signalling and Tissue Specificity for IIS-Mediated Longevity Regulation
The discovery of the IIS-mediated longevity pathway in C. elegans , combined with the fact that mammalian IIS is regulated by insulin and IGF hormones, implies the presence of endocrine-mediated ageing regulation (Fig. 4.2). Extensive genetic and bioinformatic studies have identified 40 members of the ILP superfamily in C. elegans, including insulin (INS)-1 through INS-39, and DAF-28 [102–107]. C. elegans ILPs are structurally different from mammalian insulins, since most lack a connecting peptide (C-peptide), which is a typical feature of the mammalian counterparts. In addition, some C. elegans ILPs have a different inter-chain disulphide bond conformation between conserved cysteine residues [102, 105]. Interestingly, INS-6, which lacks the C-peptide, can bind to the human insulin receptor [108]. Thus, C. elegans ILPs may function as ligands for the DAF-2, despite the structural divergence.
Among the 40 ILPs that have been identified to date, only a few have been functionally characterized in depth, perhaps because of their redundancy and/or complexity [93, 104–106, 109–117]. ILPs are known to modulate the activity of DAF-2 by acting as either agonists (e.g., INS-6 and DAF-28) or antagonists (e.g., INS-1) [21, 93, 105, 106, 111, 117–120]. However, some ILPs, such as INS-18 and INS-7, can serve as both agonists and antagonists of DAF-2 in a context-dependent manner [104, 105, 109, 112, 116, 121]. Recent studies have characterized the expression patterns and functions of all ILPs systematically [121, 122]. In contrast to the previous notion that ILPs function redundantly [[117, 122] also reviewed in [123]], these studies have suggested that ILPs can constitute combinatorial codes for the regulation of development and physiology in C. elegans [121]. Thus, ILPs appear to have distinct roles as individuals and to regulate various physiological outputs as members of an intricate ILP-regulatory network.
Various tissues in C. elegans express ILPs and appear to regulate IIS in an endocrine manner. ILPs are mainly expressed in neurons, although a few have also been shown to be expressed in other tissues, such as the intestine and the hypodermis [93, 105, 106, 109, 111, 115–117, 119, 120, 122, 124]. These expression patterns of ILPs imply that the nervous system of C. elegans may be a key regulatory centre for endocrine IIS. Consistent with this idea, neuronal IIS has a large impact on organismal physiology. For example, DAF-2, AGE-1, and DAF-18 regulate lifespan cell non-autonomously in the nervous system [125–127]. In addition, disruption of sensory neurons increases lifespan and up-regulates DAF-16 in the intestine and the hypodermis by decreasing the expression of INS-6 and DAF-28 [93]. Neuronal daf-16 contributes to the long lifespan of daf-2 mutants [128], again pointing to the important role of the nervous system in endocrine regulation of IIS-induced longevity.
Tissues other than neurons also play substantial roles in the endocrine IIS-regulated lifespan in C. elegans. The intestine of C. elegans is the major digestive organ [129] and serves as a signalling centre for nutritional status. Thus, IIS in the intestine may transmit signals regarding nutritional status to regulate organismal physiology. In fact, intestine-specific expression of daf-16 substantially restores the longevity of daf-2 mutants [128]. The intestine also regulates the expression of ILPs, in particular ins-7, to modulate IIS in distant tissues via a positive feedback loop [109]. In addition, intestinal daf-16 prevents age-dependent deterioration of muscle [60]. Overall, this endocrine IIS system appears to coordinate the rates of ageing among different C. elegans tissues.
4.5 The Role of IIS in Stress Resistance and Age-Related Disease Models
In addition to lifespan, the C. elegans IIS pathway regulates various other physiological processes . For example, reduced IIS enhances resistance to a number of stresses, including heat [130, 131], oxidative stress [132–134], and osmotic stress [135], as well as hypoxia [136, 137]. Reduced IIS also allows C. elegans to successfully cope with heavy metal toxicity [138], ultraviolet (UV) radiation [139], endoplasmic reticulum (ER) stress [61], and cytosolic proteotoxicity [67, 68, 140]. This signifies the importance of IIS pathway-regulated mechanisms for healthy ageing .
Stress resistance resulting from reduced IIS is mediated by a variety of factors, including longevity-promoting transcription factors DAF-16, HSF-1, and SKN-1 (see Sect. 4.2). For example, DAF-16 contributes to enhanced thermotolerance and resistance to hypertonicity, UV, heavy metals, and hypoxia conferred by reduced IIS [67, 130, 131, 135–139, 141–143]. Reduced IIS also protects against oxidative stress by triggering the activation of DAF-16 and SKN-1 [26–28, 39, 79, 80, 86, 132–134, 144]. The SMK-1 and EGL-27/GATA transcription factor promote UV resistance in daf-2 mutants [39, 142, 145]. XBP-1, a key mediator of the ER unfolded protein response (UPRER), collaborates with DAF-16 to enhance UPRER in daf-2 mutants [61]. Additionally, HSF-1, together with DAF-16, contributes to enhanced cytosolic protein homeostasis conferred by reduced IIS [67, 68]. The decreased levels of IIS also protect somatic cells from various stresses by equipping these cells with many characteristics of germline stem cells [146]. Overall, IIS-mediated stress resistance contributes to the proper management of stresses through a variety of factors, which are also essential for longevity.
Innate immunity ensures survival in the presence of pathogenic threats. C. elegans has an innate immune system that is regulated by evolutionarily conserved signalling pathways, one of which is the IIS pathway. Reduced IIS activity increases resistance to various fungal and bacterial pathogens via DAF-16 [147, 148], in parallel to the well-known immune regulator, p38 MAP kinase [147–151]. The transcription factors SKN-1 and HSF-1 also mediate the enhanced pathogen resistance under conditions of reduced IIS [152, 153]. daf-2 mutants display mitigated internal bacterial colonization, enhanced bacterial clearance, and increased expression of antimicrobial genes [21, 151]. Moreover, daf-2 mutants display enhanced efficiency in RNA interference (RNAi) [154], which is important for antiviral defence in C. elegans [155–157]. Thus, it will be interesting to test whether daf-2 mutants are resistant to viral infections as well.
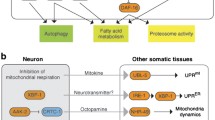
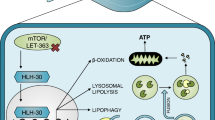
Importantly, reduced IIS has been shown to alleviate the pathological features of various disease models in C. elegans, including Huntington’s disease [140, 158], Alzheimer’s disease [159, 160], Parkinson’s disease [161], and amyotrophic lateral sclerosis (ALS) [162] (Fig. 4.3). In a Huntington’s disease model, reduced IIS ameliorates the polyglutamine (polyQ) aggregation mediated by CAG repeats in a DAF-16 - and HSF-1-dependent manner [67, 140, 163, 164]. In a model for Alzheimer’s disease, reduced IIS protects C. elegans from the toxicity caused by Aβ1−42 expression via DAF-16, HSF-1, and autophagy [165, 166]. In a Parkinson’s disease (PD) model, C. elegans expressing human α-synuclein in neurons displays both a motor deficit and progressive degeneration of dopaminergic neurons [161]; however, daf-2 mutations result in complete retention of these dopaminergic neurons [164]. ALS originates from mutations in various genes, including superoxide dismutase 1 (SOD1) [167]. In a C. elegans model, daf-2 mutations protect against the toxic mutant SOD1-induced motor neuron dysfunction by decreasing protein aggregation [168]. Overall, it appears that the enhanced protein homeostasis conferred by reduced IIS underlies the protective mechanisms against these degenerative disease models in C. elegans [169, 170]. It is noteworthy that daf-2 mutations delay age-dependent neuronal degeneration [171] and neurite branching [172]. Mutations in daf-2 also enhance memory and learning capacity in early adulthood, and delay an age-dependent decline in short-term memory in a DAF-16-dependent manner [173]. These data strongly suggest that proper manipulation of the evolutionarily conserved IIS pathway in C. elegans may shed light on the molecular basis of age- and/or disease-induced defects. Further, this pathway may hold therapeutic potential for the treatment of various degenerative diseases .
The role of IIS in stress resistance and human disease models. Reduced IIS confers enhanced resistance against a variety of stresses, including heat, hypoxia , high osmolarity, heavy metals, UV radiation, proteotoxicity, and pathogens. Reduced IIS also ameliorates the impact of age-related human disease models in C. elegans , including those for Huntington’s disease, amyloid lateral sclerosis (ALS), Alzheimer’s disease, and Parkinson’s disease. These features correlate with healthy ageing and longevity
4.6 Conclusions
In this chapter, we reviewed the functions of IIS and the mechanisms by which it influences C. elegans longevity. The entire IIS pathway appears to play a central role in linking environmental signals, such as food availability and stresses, to various physiological outputs, including ageing , reproduction , and development. Therefore, one possible reason why the IIS pathway has a huge impact on ageing is because this system responds to changes in environmental conditions and alters physiological outputs accordingly. Thus, under favourable conditions, IIS may be activated to promote growth and reproduction, which may lead to normal or shortened lifespan. Conversely, under unfavourable conditions, such as food shortages, IIS is down-regulated and activates genetic programmes to promote organism-wise maintenance, rather than growth and reproduction; this may lead to a longer lifespan. Therefore, enhanced longevity may be associated with slow growth and reduced reproduction. Indeed mutations in many components of IIS result in developmental arrest (see Chap. 3) and reduced fecundity, as well as longevity. However, it is worth pointing out that the regulation of organismal development and ageing by IIS can be uncoupled by temporally modulating the signalling [71]. Further dissection of the pleiotropic aspects of IIS will be crucial for understanding the specific contribution of IIS to ageing regulation.
The establishment of the role of IIS in ageing has paved the way for discoveries showing that various IIS components, such as insulin receptor and IGF-1 receptor, as well as the AKT kinases and FOXO transcription factors , regulate mammalian longevity. These findings have further led to the identification of genetic variants of IGF-1 receptor and FOXO3A that are associated with human longevity [1, 174]. Therefore, the conservation between invertebrate models and mammals, including humans, will help us to understand the biology of human ageing. Ultimately, what we have learned from C. elegans IIS can potentially lead to therapies aimed at delaying the onset of ageing-associated diseases and achieving a healthier and longer life in humans .
References
Kenyon CJ (2010) The genetics of ageing. Nature 464(7288):504–512
Fontana L, Partridge L, Longo VD (2010) Extending healthy life span – from yeast to humans. Science (New York, NY) 328(5976):321–326
Klass MR (1983) A method for the isolation of longevity mutants in the nematode C. elegans and initial results. Mech Ageing Dev 22(3–4):279–286
Friedman DB, Johnson TE (1988) Three mutants that extend both mean and maximum life span of the nematode, C. elegans, define the age-1 gene. J Gerontol 43(4):B102–B109
Kenyon C, Chang J, Gensch E, Rudner A, Tabtiang R (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366(6454):461–464
Morris JZ, Tissenbaum HA, Ruvkun G (1996) A phosphatidylinositol-3-OH kinase family member regulating longevity and diapause in C. elegans. Nature 382(6591):536–539
Kimura KD, Tissenbaum HA, Liu Y, Ruvkun G (1997) daf-2, an insulin receptor-like gene that regulates longevity and diapause in C. elegans. Science (New York, NY) 277(5328):942–946
Dorman JB, Albinder B, Shroyer T, Kenyon C (1995) The age-1 and daf-2 genes function in a common pathway to control the lifespan of C. elegans. Genetics 141(4):1399–1406
Murphy CT, Hu PJ (2013) Insulin/insulin-like growth factor signaling in C. elegans. WormBook Online Rev C elegans Biol:1–43
Paradis S, Ailion M, Toker A, Thomas JH, Ruvkun G (1999) A PDK1 homolog is necessary and sufficient to transduce AGE-1 PI3 kinase signals that regulate diapause in C. elegans. Genes Dev 13(11):1438–1452
Zhou K, Pandol S, Bokoch G, Traynor-Kaplan AE (1998) Disruption of Dictyostelium PI3K genes reduces [32P]phosphatidylinositol 3,4 bisphosphate and [32P]phosphatidylinositol trisphosphate levels, alters F-actin distribution and impairs pinocytosis. J Cell Sci 111(Pt 2):283–294
Paradis S, Ruvkun G (1998) C. elegans Akt/PKB transduces insulin receptor-like signals from AGE-1 PI3 kinase to the DAF-16 transcription factor. Genes Dev 12(16):2488–2498
Larsen PL, Albert PS, Riddle DL (1995) Genes that regulate both development and longevity in C. elegans. Genetics 139(4):1567–1583
Gil EB, Malone Link E, Liu LX, Johnson CD, Lees JA (1999) Regulation of the insulin-like developmental pathway of C. elegans by a homolog of the PTEN tumor suppressor gene. Proc Natl Acad Sci U S A 96(6):2925–2930
Ogg S, Ruvkun G (1998) The C. elegans PTEN homolog, DAF-18, acts in the insulin receptor-like metabolic signaling pathway. Mol Cell 2(6):887–893
Mihaylova VT, Borland CZ, Manjarrez L, Stern MJ, Sun H (1999) The PTEN tumor suppressor homolog in C. elegans regulates longevity and dauer formation in an insulin receptor-like signaling pathway. Proc Natl Acad Sci U S A 96(13):7427–7432
Solari F, Bourbon-Piffaut A, Masse I, Payrastre B, Chan AM, Billaud M (2005) The human tumour suppressor PTEN regulates longevity and dauer formation in C. elegans. Oncogene 24(1):20–27
Ogg S, Paradis S, Gottlieb S, Patterson GI, Lee L, Tissenbaum HA, Ruvkun G (1997) The Fork head transcription factor DAF-16 transduces insulin-like metabolic and longevity signals in C. elegans. Nature 389(6654):994–999
Lin K, Dorman JB, Rodan A, Kenyon C (1997) daf-16: an HNF-3/forkhead family member that can function to double the life-span of C. elegans. Science (New York, NY) 278(5341):1319–1322
Lee SS, Kennedy S, Tolonen AC, Ruvkun G (2003) DAF-16 target genes that control C. elegans life-span and metabolism. Science (New York, NY) 300(5619):644–647
Murphy CT, McCarroll SA, Bargmann CI, Fraser A, Kamath RS, Ahringer J, Li H, Kenyon C (2003) Genes that act downstream of DAF-16 to influence the lifespan of C. elegans. Nature 424(6946):277–283
McElwee J, Bubb K, Thomas JH (2003) Transcriptional outputs of the C. elegans forkhead protein DAF-16. Aging Cell 2(2):111–121
McElwee JJ, Schuster E, Blanc E, Thomas JH, Gems D (2004) Shared transcriptional signature in C. elegans Dauer larvae and long-lived daf-2 mutants implicates detoxification system in longevity assurance. J Biol Chem 279(43):44533–44543
Shaw WM, Luo S, Landis J, Ashraf J, Murphy CT (2007) The C. elegans TGF-beta Dauer pathway regulates longevity via insulin signaling. Curr Biol 17(19):1635–1645
Halaschek-Wiener J, Khattra JS, McKay S, Pouzyrev A, Stott JM, Yang GS, Holt RA, Jones SJ, Marra MA, Brooks-Wilson AR, Riddle DL (2005) Analysis of long-lived C. elegans daf-2 mutants using serial analysis of gene expression. Genome Res 15(5):603–615
Henderson ST, Johnson TE (2001) daf-16 integrates developmental and environmental inputs to mediate aging in the nematode C. elegans. Curr Biol 11(24):1975–1980
Lin K, Hsin H, Libina N, Kenyon C (2001) Regulation of the C. elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat Genet 28(2):139–145
Lee RY, Hench J, Ruvkun G (2001) Regulation of C. elegans DAF-16 and its human ortholog FKHRL1 by the daf-2 insulin-like signaling pathway. Curr Biol 11(24):1950–1957
Cahill CM, Tzivion G, Nasrin N, Ogg S, Dore J, Ruvkun G, Alexander-Bridges M (2001) Phosphatidylinositol 3-kinase signaling inhibits DAF-16 DNA binding and function via 14-3-3-dependent and 14-3-3-independent pathways. J Biol Chem 276(16):13402–13410
Hertweck M, Gobel C, Baumeister R (2004) C. elegans SGK-1 is the critical component in the Akt/PKB kinase complex to control stress response and life span. Dev Cell 6(4):577–588
Chen AT, Guo C, Dumas KJ, Ashrafi K, Hu PJ (2013) Effects of C. elegans sgk-1 mutations on lifespan, stress resistance, and DAF-16/FoxO regulation. Aging Cell 12(5):932–940
Xiao R, Zhang B, Dong Y, Gong J, Xu T, Liu J, Xu XZ (2013) A genetic program promotes C. elegans longevity at cold temperatures via a thermosensitive TRP channel. Cell 152(4):806–817
Apfeld J, O’Connor G, McDonagh T, DiStefano PS, Curtis R (2004) The AMP-activated protein kinase AAK-2 links energy levels and insulin-like signals to lifespan in C. elegans. Genes Dev 18(24):3004–3009
Curtis R, O’Connor G, DiStefano PS (2006) Aging networks in C. elegans: AMP-activated protein kinase (aak-2) links multiple aging and metabolism pathways. Aging Cell 5(2):119–126
Tullet JM, Araiz C, Sanders MJ, Au C, Benedetto A, Papatheodorou I, Clark E, Schmeisser K, Jones D, Schuster EF, Thornton JM, Gems D (2014) DAF-16/FoxO directly regulates an atypical AMP-activated protein kinase gamma isoform to mediate the effects of insulin/IGF-1 signaling on aging in C. elegans. PLoS Genet 10(2), e1004109
Greer EL, Dowlatshahi D, Banko MR, Villen J, Hoang K, Blanchard D, Gygi SP, Brunet A (2007) An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans. Curr Biol 17(19):1646–1656
Lehtinen MK, Yuan Z, Boag PR, Yang Y, Villen J, Becker EB, DiBacco S, de la Iglesia N, Gygi S, Blackwell TK, Bonni A (2006) A conserved MST-FOXO signaling pathway mediates oxidative-stress responses and extends life span. Cell 125(5):987–1001
Oh SW, Mukhopadhyay A, Svrzikapa N, Jiang F, Davis RJ, Tissenbaum HA (2005) JNK regulates lifespan in C. elegans by modulating nuclear translocation of forkhead transcription factor/DAF-16. Proc Natl Acad Sci U S A 102(12):4494–4499
Wolff S, Ma H, Burch D, Maciel GA, Hunter T, Dillin A (2006) SMK-1, an essential regulator of DAF-16-mediated longevity. Cell 124(5):1039–1053
Padmanabhan S, Mukhopadhyay A, Narasimhan SD, Tesz G, Czech MP, Tissenbaum HA (2009) A PP2A regulatory subunit regulates C. elegans insulin/IGF-1 signaling by modulating AKT-1 phosphorylation. Cell 136(5):939–951
Zhang M, Poplawski M, Yen K, Cheng H, Bloss E, Zhu X, Patel H, Mobbs CV (2009) Role of CBP and SATB-1 in aging, dietary restriction, and insulin-like signaling. PLoS Biol 7(11), e1000245
Chiang WC, Tishkoff DX, Yang B, Wilson-Grady J, Yu X, Mazer T, Eckersdorff M, Gygi SP, Lombard DB, Hsu AL (2012) C. elegans SIRT6/7 homolog SIR-2.4 promotes DAF-16 relocalization and function during stress. PLoS Genet 8(9), e1002948
Tissenbaum HA, Guarente L (2001) Increased dosage of a sir-2 gene extends lifespan in C. elegans. Nature 410(6825):227–230
Berdichevsky A, Viswanathan M, Horvitz HR, Guarente L (2006) C. elegans SIR-2.1 interacts with 14-3-3 proteins to activate DAF-16 and extend life span. Cell 125(6):1165–1177
Rizki G, Iwata TN, Li J, Riedel CG, Picard CL, Jan M, Murphy CT, Lee SS (2011) The evolutionarily conserved longevity determinants HCF-1 and SIR-2.1/SIRT1 collaborate to regulate DAF-16/FOXO. PLoS Genet 7(9):e1002235
Burnett C, Valentini S, Cabreiro F, Goss M, Somogyvari M, Piper MD, Hoddinott M, Sutphin GL, Leko V, McElwee JJ, Vazquez-Manrique RP, Orfila AM, Ackerman D, Au C, Vinti G, Riesen M, Howard K, Neri C, Bedalov A, Kaeberlein M, Soti C, Partridge L, Gems D (2011) Absence of effects of Sir2 overexpression on lifespan in C. elegans and Drosophila. Nature 477(7365):482–485
Li W, Gao B, Lee SM, Bennett K, Fang D (2007) RLE-1, an E3 ubiquitin ligase, regulates C. elegans aging by catalyzing DAF-16 polyubiquitination. Dev Cell 12(2):235–246
Heimbucher T, Liu Z, Bossard C, McCloskey R, Carrano AC, Riedel CG, Tanasa B, Klammt C, Fonslow BR, Riera CE, Lillemeier BF, Kemphues K, Yates JR 3rd, O’Shea C, Hunter T, Dillin A (2015) The deubiquitylase MATH-33 controls DAF-16 stability and function in metabolism and longevity. Cell Metab 22(1):151–163
Ghazi A, Henis-Korenblit S, Kenyon C (2007) Regulation of C. elegans lifespan by a proteasomal E3 ligase complex. Proc Natl Acad Sci U S A 104(14):5947–5952
Chondrogianni N, Georgila K, Kourtis N, Tavernarakis N, Gonos ES (2015) 20S proteasome activation promotes life span extension and resistance to proteotoxicity in C. elegans. FASEB J 29(2):611–622
Li J, Tewari M, Vidal M, Lee SS (2007) The 14-3-3 protein FTT-2 regulates DAF-16 in C. elegans. Dev Biol 301(1):82–91
Wang Y, Oh SW, Deplancke B, Luo J, Walhout AJ, Tissenbaum HA (2006) C. elegans 14-3-3 proteins regulate life span and interact with SIR-2.1 and DAF-16/FOXO. Mech Ageing Dev 127(9):741–747
Neumann-Haefelin E, Qi W, Finkbeiner E, Walz G, Baumeister R, Hertweck M (2008) SHC-1/p52Shc targets the insulin/IGF-1 and JNK signaling pathways to modulate life span and stress response in C. elegans. Genes Dev 22(19):2721–2735
Kwon ES, Narasimhan SD, Yen K, Tissenbaum HA (2010) A new DAF-16 isoform regulates longevity. Nature 466(7305):498–502
Bansal A, Kwon ES, Conte D Jr, Liu H, Gilchrist MJ, MacNeil LT, Tissenbaum HA (2014) Transcriptional regulation of C. elegans FOXO/DAF-16 modulates lifespan. Longev Healthspan 3:5
Furuyama T, Nakazawa T, Nakano I, Mori N (2000) Identification of the differential distribution patterns of mRNAs and consensus binding sequences for mouse DAF-16 homologues. Biochem J 349(Pt 2):629–634
Budovskaya YV, Wu K, Southworth LK, Jiang M, Tedesco P, Johnson TE, Kim SK (2008) An elt-3/elt-5/elt-6 GATA transcription circuit guides aging in C. elegans. Cell 134(2):291–303
Schuster E, McElwee JJ, Tullet JM, Doonan R, Matthijssens F, Reece-Hoyes JS, Hope IA, Vanfleteren JR, Thornton JM, Gems D (2010) DamID in C. elegans reveals longevity-associated targets of DAF-16/FoxO. Mol Syst Biol 6:399
Tepper RG, Ashraf J, Kaletsky R, Kleemann G, Murphy CT, Bussemaker HJ (2013) PQM-1 complements DAF-16 as a key transcriptional regulator of DAF-2-mediated development and longevity. Cell 154(3):676–690
Zhang P, Judy M, Lee SJ, Kenyon C (2013) Direct and indirect gene regulation by a life-extending FOXO protein in C. elegans: roles for GATA factors and lipid gene regulators. Cell Metab 17(1):85–100
Henis-Korenblit S, Zhang P, Hansen M, McCormick M, Lee SJ, Cary M, Kenyon C (2010) Insulin/IGF-1 signaling mutants reprogram ER stress response regulators to promote longevity. Proc Natl Acad Sci U S A 107(21):9730–9735
Thyagarajan B, Blaszczak AG, Chandler KJ, Watts JL, Johnson WE, Graves BJ (2010) ETS-4 is a transcriptional regulator of life span in C. elegans. PLoS Genet 6(9), e1001125
Seo M, Seo K, Hwang W, Koo HJ, Hahm JH, Yang JS, Han SK, Hwang D, Kim S, Jang SK, Lee Y, Nam HG, Lee SJ (2015) RNA helicase HEL-1 promotes longevity by specifically activating DAF-16/FOXO transcription factor signaling in C. elegans. Proc Natl Acad Sci U S A 112(31):E4246–E4255
Takahashi Y, Daitoku H, Hirota K, Tamiya H, Yokoyama A, Kako K, Nagashima Y, Nakamura A, Shimada T, Watanabe S, Yamagata K, Yasuda K, Ishii N, Fukamizu A (2011) Asymmetric arginine dimethylation determines life span in C. elegans by regulating forkhead transcription factor DAF-16. Cell Metab 13(5):505–516
Riedel CG, Dowen RH, Lourenco GF, Kirienko NV, Heimbucher T, West JA, Bowman SK, Kingston RE, Dillin A, Asara JM, Ruvkun G (2013) DAF-16 employs the chromatin remodeller SWI/SNF to promote stress resistance and longevity. Nat Cell Biol 15(5):491–501
Morimoto RI (2011) The heat shock response: systems biology of proteotoxic stress in aging and disease. Cold Spring Harb Symp Quant Biol 76:91–99
Hsu AL, Murphy CT, Kenyon C (2003) Regulation of aging and age-related disease by DAF-16 and heat-shock factor. Science (New York, NY) 300(5622):1142–1145
Morley JF, Morimoto RI (2004) Regulation of longevity in C. elegans by heat shock factor and molecular chaperones. Mol Biol Cell 15(2):657–664
Douglas PM, Baird NA, Simic MS, Uhlein S, McCormick MA, Wolff SC, Kennedy BK, Dillin A (2015) Heterotypic signals from neural HSF-1 separate thermotolerance from longevity. Cell Rep 12(7):1196–1204
Volovik Y, Maman M, Dubnikov T, Bejerano-Sagie M, Joyce D, Kapernick EA, Cohen E, Dillin A (2012) Temporal requirements of heat shock factor-1 for longevity assurance. Aging Cell 11(3):491–499
Dillin A, Crawford DK, Kenyon C (2002) Timing requirements for insulin/IGF-1 signaling in C. elegans. Science (New York, NY) 298(5594):830–834
Amin J, Ananthan J, Voellmy R (1988) Key features of heat shock regulatory elements. Mol Cell Biol 8(9):3761–3769
Walker GA, Lithgow GJ (2003) Lifespan extension in C. elegans by a molecular chaperone dependent upon insulin-like signals. Aging Cell 2(2):131–139
Chiang WC, Ching TT, Lee HC, Mousigian C, Hsu AL (2012) HSF-1 regulators DDL-1/2 link insulin-like signaling to heat-shock responses and modulation of longevity. Cell 148(1–2):322–334
Baird NA, Douglas PM, Simic MS, Grant AR, Moresco JJ, Wolff SC, Yates JR 3rd, Manning G, Dillin A (2014) HSF-1-mediated cytoskeletal integrity determines thermotolerance and life span. Science (New York, NY) 346(6207):360–363
Horikawa M, Sural S, Hsu AL, Antebi A (2015) Co-chaperone p23 regulates C. elegans lifespan in response to temperature. PLoS Genet 11(4), e1005023
Seo K, Choi E, Lee D, Jeong DE, Jang SK, Lee SJ (2013) Heat shock factor 1 mediates the longevity conferred by inhibition of TOR and insulin/IGF-1 signaling pathways in C. elegans. Aging Cell 12(6):1073–1081
Blackwell TK, Steinbaugh MJ, Hourihan JM, Ewald CY, Isik M (2015) SKN-1/Nrf, stress responses, and aging in C. elegans. Free Radic Biol Med
An JH, Blackwell TK (2003) SKN-1 links C. elegans mesendodermal specification to a conserved oxidative stress response. Genes Dev 17(15):1882–1893
Tullet JM, Hertweck M, An JH, Baker J, Hwang JY, Liu S, Oliveira RP, Baumeister R, Blackwell TK (2008) Direct inhibition of the longevity-promoting factor SKN-1 by insulin-like signaling in C. elegans. Cell 132(6):1025–1038
An JH, Vranas K, Lucke M, Inoue H, Hisamoto N, Matsumoto K, Blackwell TK (2005) Regulation of the C. elegans oxidative stress defense protein SKN-1 by glycogen synthase kinase-3. Proc Natl Acad Sci U S A 102(45):16275–16280
Kahn NW, Rea SL, Moyle S, Kell A, Johnson TE (2008) Proteasomal dysfunction activates the transcription factor SKN-1 and produces a selective oxidative-stress response in C. elegans. Biochem J 409(1):205–213
Oliveira RP, Porter Abate J, Dilks K, Landis J, Ashraf J, Murphy CT, Blackwell TK (2009) Condition-adapted stress and longevity gene regulation by C. elegans SKN-1/Nrf. Aging Cell 8(5):524–541
Wang J, Robida-Stubbs S, Tullet JM, Rual JF, Vidal M, Blackwell TK (2010) RNAi screening implicates a SKN-1-dependent transcriptional response in stress resistance and longevity deriving from translation inhibition. PLoS Genet 6 (8)
Li X, Matilainen O, Jin C, Glover-Cutter KM, Holmberg CI, Blackwell TK (2011) Specific SKN-1/Nrf stress responses to perturbations in translation elongation and proteasome activity. PLoS Genet 7(6), e1002119
Ewald CY, Landis JN, Porter Abate J, Murphy CT, Blackwell TK (2015) Dauer-independent insulin/IGF-1-signalling implicates collagen remodelling in longevity. Nature 519(7541):97–101
White JG, Southgate E, Thomson JN, Brenner S (1986) The structure of the nervous system of the nematode C. elegans. Philos Trans R Soc Lond B Biol Sci 314(1165):1–340
Jeong DE, Artan M, Seo K, Lee SJ (2012) Regulation of lifespan by chemosensory and thermosensory systems: findings in invertebrates and their implications in mammalian aging. Front Genet 3:218
Apfeld J, Kenyon C (1999) Regulation of lifespan by sensory perception in C. elegans. Nature 402(6763):804–809
Lee SJ, Kenyon C (2009) Regulation of the longevity response to temperature by thermosensory neurons in C. elegans. Curr Biol 19(9):715–722
Alcedo J, Kenyon C (2004) Regulation of C. elegans longevity by specific gustatory and olfactory neurons. Neuron 41(1):45–55
Lans H, Jansen G (2007) Multiple sensory G proteins in the olfactory, gustatory and nociceptive neurons modulate longevity in C. elegans. Dev Biol 303(2):474–482
Artan M, Jeong DE, Lee D, Kim YI, Son HG, Husain Z, Kim J, Altintas O, Kim K, Alcedo J, Lee SJ (2016) Food-derived sensory cues modulate longevity via distinct neuroendocrine insulin-like peptides. Genes Dev 30(9):1047–1057
Ailion M, Inoue T, Weaver CI, Holdcraft RW, Thomas JH (1999) Neurosecretory control of aging in C. elegans. Proc Natl Acad Sci U S A 96(13):7394–7397
Lanjuin A, Sengupta P (2002) Regulation of chemosensory receptor expression and sensory signaling by the KIN-29 Ser/Thr kinase. Neuron 33(3):369–381
Lee BH, Ashrafi K (2008) A TRPV channel modulates C. elegans neurosecretion, larval starvation survival, and adult lifespan. PLoS Genet 4(10), e1000213
Hahm JH, Kim S, Paik YK (2009) Endogenous cGMP regulates adult longevity via the insulin signaling pathway in C. elegans. Aging Cell 8(4):473–483
Riera CE, Huising MO, Follett P, Leblanc M, Halloran J, Van Andel R, de Magalhaes CD, Merkwirth C, Dillin A (2014) TRPV1 pain receptors regulate longevity and metabolism by neuropeptide signaling. Cell 157(5):1023–1036
Maier W, Adilov B, Regenass M, Alcedo J (2010) A neuromedin U receptor acts with the sensory system to modulate food type-dependent effects on C. elegans lifespan. PLoS Biol 8(5), e1000376
Gaglia MM, Jeong DE, Ryu EA, Lee D, Kenyon C, Lee SJ (2012) Genes that act downstream of sensory neurons to influence longevity, dauer formation, and pathogen responses in C. elegans. PLoS Genet 8(12), e1003133
Zhang B, Xiao R, Ronan EA, He Y, Hsu AL, Liu J, Xu XZ (2015) Environmental temperature differentially modulates C. elegans longevity through a thermosensitive TRP channel. Cell Rep 11(9):1414–1424
Duret L, Guex N, Peitsch MC, Bairoch A (1998) New insulin-like proteins with atypical disulfide bond pattern characterized in C. elegans by comparative sequence analysis and homology modeling. Genome Res 8(4):348–353
Gregoire FM, Chomiki N, Kachinskas D, Warden CH (1998) Cloning and developmental regulation of a novel member of the insulin-like gene family in C. elegans. Biochem Biophys Res Commun 249(2):385–390
Kawano T, Ito Y, Ishiguro M, Takuwa K, Nakajima T, Kimura Y (2000) Molecular cloning and characterization of a new insulin/IGF-like peptide of the nematode C. elegans. Biochem Biophys Res Commun 273(2):431–436
Pierce SB, Costa M, Wisotzkey R, Devadhar S, Homburger SA, Buchman AR, Ferguson KC, Heller J, Platt DM, Pasquinelli AA (2001) Regulation of DAF-2 receptor signaling by human insulin and ins-1, a member of the unusually large and diverse C. elegans insulin gene family. Genes Dev 15(6):672–686
Li W, Kennedy SG, Ruvkun G (2003) daf-28 encodes a C. elegans insulin superfamily member that is regulated by environmental cues and acts in the DAF-2 signaling pathway. Genes Dev 17(7):844–858
Husson SJ, Mertens I, Janssen T, Lindemans M, Schoofs L (2007) Neuropeptidergic signaling in the nematode C. elegans. Prog Neurobiol 82(1):33–55
Hua Q-x, Nakagawa SH, Wilken J, Ramos RR, Jia W, Bass J, Weiss MA (2003) A divergent INS protein in Caenorhabditis elegans structurally resembles human insulin and activates the human insulin receptor. Genes Dev 17(7):826–831
Murphy CT, Lee S-J, Kenyon C (2007) Tissue entrainment by feedback regulation of insulin gene expression in the endoderm of C. elegans. Proc Natl Acad Sci 104(48):19046–19050
Lin CHA, Tomioka M, Pereira S, Sellings L, Iino Y, van der Kooy D (2010) Insulin signaling plays a dual role in C. elegans memory acquisition and memory retrieval. J Neurosci 30(23):8001–8011
Cornils A, Gloeck M, Chen Z, Zhang Y, Alcedo J (2011) Specific insulin-like peptides encode sensory information to regulate distinct developmental processes. Development 138(6):1183–1193
Matsunaga Y, Gengyo-Ando K, Mitani S, Iwasaki T, Kawano T (2012) Physiological function, expression pattern, and transcriptional regulation of a C. elegans insulin-like peptide, INS-18. Biochem Biophys Res Commun 423(3):478–483
Matsunaga Y, Nakajima K, Gengyo-Ando K, Mitani S, Iwasaki T, Kawano T (2012) A C. elegans insulin-like peptide, INS-17: its physiological function and expression pattern. Biosci Biotechnol Biochem 76(11):2168–2172
Kulalert W, Kim DH (2013) The unfolded protein response in a pair of sensory neurons promotes entry of C. elegans into dauer diapause. Curr Biol 23(24):2540–2545
Leinwand SG, Chalasani SH (2013) Neuropeptide signaling remodels chemosensory circuit composition in C. elegans. Nat Neurosci 16(10):1461–1467
Chen Z, Hendricks M, Cornils A, Maier W, Alcedo J, Zhang Y (2013) Two insulin-like peptides antagonistically regulate aversive olfactory learning in C. elegans. Neuron 77(3):572–585
Hung WL, Wang Y, Chitturi J, Zhen M (2014) A C. elegans developmental decision requires insulin signaling-mediated neuron-intestine communication. Development 141(8):1767–1779
Malone EA, Inoue T, Thomas JH (1996) Genetic analysis of the roles of daf-28 and age-1 in regulating C. elegans Dauer formation. Genetics 143(3):1193–1205
Ohta A, Ujisawa T, Sonoda S, Kuhara A (2014) Light and pheromone-sensing neurons regulates cold habituation through insulin signalling in C. elegans. Nature Commun 5:4412
Chen Y, Baugh LR (2014) Ins-4 and daf-28 function redundantly to regulate C. elegans L1 arrest. Dev Biol 394(2):314–326
de Abreu DAF, Caballero A, Fardel P, Stroustrup N, Chen Z, Lee K, Keyes WD, Nash ZM, López-Moyado IF, Vaggi F (2014) An insulin-to-insulin regulatory network orchestrates phenotypic specificity in development and physiology. PLoS Genet 10(3), e1004225
Ritter AD, Shen Y, Bass JF, Jeyaraj S, Deplancke B, Mukhopadhyay A, Xu J, Driscoll M, Tissenbaum HA, Walhout AJ (2013) Complex expression dynamics and robustness in C. elegans insulin networks. Genome Res 23(6):954–965
Nelson DW, Padgett RW (2003) Insulin worms its way into the spotlight. Genes Dev 17(7):813–818
Michaelson D, Korta DZ, Capua Y, Hubbard EJA (2010) Insulin signaling promotes germline proliferation in C. elegans. Development 137(4):671–680
Apfeld J, Kenyon C (1998) Cell nonautonomy of C. elegans daf-2 function in the regulation of diapause and life span. Cell 95(2):199–210
Wolkow CA, Kimura KD, Lee M-S, Ruvkun G (2000) Regulation of C. elegans life-span by insulin-like signaling in the nervous system. Science (New York, NY) 290(5489):147–150
Masse I, Molin L, Billaud M, Solari F (2005) Lifespan and dauer regulation by tissue-specific activities of C. elegans DAF-18. Dev Biol 286(1):91–101
Libina N, Berman JR, Kenyon C (2003) Tissue-specific activities of C. elegans DAF-16 in the regulation of lifespan. Cell 115(4):489–502
Corsi AK, Wightman B, Chalfie M (2015) A transparent window into biology: a primer on C. elegans. WormBook: Online Rev C elegans Biol:1–31
Lithgow GJ, White TM, Melov S, Johnson TE (1995) Thermotolerance and extended life-span conferred by single-gene mutations and induced by thermal stress. Proc Natl Acad Sci U S A 92(16):7540–7544
Gems D, Sutton AJ, Sundermeyer ML, Albert PS, King KV, Edgley ML, Larsen PL, Riddle DL (1998) Two pleiotropic classes of daf-2 mutation affect larval arrest, adult behavior, reproduction and longevity in C. elegans. Genetics 150(1):129–155
Larsen PL (1993) Aging and resistance to oxidative damage in C. elegans. Proc Natl Acad Sci U S A 90(19):8905–8909
Vanfleteren JR (1993) Oxidative stress and ageing in C. elegans. Biochem J 292(Pt 2):605–608
Honda Y, Honda S (1999) The daf-2 gene network for longevity regulates oxidative stress resistance and Mn-superoxide dismutase gene expression in C. elegans. FASEB J 13(11):1385–1393
Lamitina ST, Strange K (2005) Transcriptional targets of DAF-16 insulin signaling pathway protect C. elegans from extreme hypertonic stress. Am J Physiol Cell Physiol 288(2):C467–C474
Scott BA, Avidan MS, Crowder CM (2002) Regulation of hypoxic death in C. elegans by the insulin/IGF receptor homolog DAF-2. Science (New York, NY) 296(5577):2388–2391
Mabon ME, Scott BA, Crowder CM (2009) Divergent mechanisms controlling hypoxic sensitivity and lifespan by the DAF-2/insulin/IGF-receptor pathway. PLoS ONE 4(11), e7937
Barsyte D, Lovejoy DA, Lithgow GJ (2001) Longevity and heavy metal resistance in daf-2 and age-1 long-lived mutants of C. elegans. FASEB J 15(3):627–634
Murakami S, Johnson TE (1996) A genetic pathway conferring life extension and resistance to UV stress in C. elegans. Genetics 143(3):1207–1218
Morley JF, Brignull HR, Weyers JJ, Morimoto RI (2002) The threshold for polyglutamine-expansion protein aggregation and cellular toxicity is dynamic and influenced by aging in C. elegans. Proc Natl Acad Sci U S A 99(16):10417–10422
Burkewitz K, Choe K, Strange K (2011) Hypertonic stress induces rapid and widespread protein damage in C. elegans. Am J Physiol Cell Physiol 301(3):C566–C576
Mueller MM, Castells-Roca L, Babu V, Ermolaeva MA, Muller RU, Frommolt P, Williams AB, Greiss S, Schneider JI, Benzing T, Schermer B, Schumacher B (2014) DAF-16/FOXO and EGL-27/GATA promote developmental growth in response to persistent somatic DNA damage. Nat Cell Biol 16(12):1168–1179
McColl G, Rogers AN, Alavez S, Hubbard AE, Melov S, Link CD, Bush AI, Kapahi P, Lithgow GJ (2010) Insulin-like signaling determines survival during stress via posttranscriptional mechanisms in C. elegans. Cell Metab 12(3):260–272
Essers MA, de Vries-Smits LM, Barker N, Polderman PE, Burgering BM, Korswagen HC (2005) Functional interaction between beta-catenin and FOXO in oxidative stress signaling. Science (New York, NY) 308(5725):1181–1184
Landis JN, Murphy CT (2010) Integration of diverse inputs in the regulation of C. elegans DAF-16/FOXO. Dev Dyn 239(5):1405–1412
Curran SP, Wu X, Riedel CG, Ruvkun G (2009) A soma-to-germline transformation in long-lived C. elegans mutants. Nature 459(7250):1079–1084
Garsin DA, Villanueva JM, Begun J, Kim DH, Sifri CD, Calderwood SB, Ruvkun G, Ausubel FM (2003) Long-lived C. elegans daf-2 mutants are resistant to bacterial pathogens. Science (New York, NY) 300(5627):1921
Kerry S, TeKippe M, Gaddis NC, Aballay A (2006) GATA transcription factor required for immunity to bacterial and fungal pathogens. PLoS ONE 1, e77
Kim DH, Feinbaum R, Alloing G, Emerson FE, Garsin DA, Inoue H, Tanaka-Hino M, Hisamoto N, Matsumoto K, Tan MW, Ausubel FM (2002) A conserved p38 MAP kinase pathway in C. elegans innate immunity. Science (New York, NY) 297(5581):623–626
Troemel ER, Chu SW, Reinke V, Lee SS, Ausubel FM, Kim DH (2006) p38 MAPK regulates expression of immune response genes and contributes to longevity in C. elegans. PLoS Genet 2(11), e183
Evans EA, Chen WC, Tan MW (2008) The DAF-2 insulin-like signaling pathway independently regulates aging and immunity in C. elegans. Aging Cell 7(6):879–893
Singh V, Aballay A (2006) Heat-shock transcription factor (HSF)-1 pathway required for C. elegans immunity. Proc Natl Acad Sci U S A 103(35):13092–13097
Papp D, Csermely P, Soti C (2012) A role for SKN-1/Nrf in pathogen resistance and immunosenescence in C. elegans. PLoS Pathog 8(4), e1002673
Wang D, Ruvkun G (2004) Regulation of C. elegans RNA interference by the daf-2 insulin stress and longevity signaling pathway. Cold Spring Harb Symp Quant Biol 69:429–431
Schott DH, Cureton DK, Whelan SP, Hunter CP (2005) An antiviral role for the RNA interference machinery in C. elegans. Proc Natl Acad Sci U S A 102(51):18420–18424
Wilkins C, Dishongh R, Moore SC, Whitt MA, Chow M, Machaca K (2005) RNA interference is an antiviral defence mechanism in C. elegans. Nature 436(7053):1044–1047
Felix MA, Ashe A, Piffaretti J, Wu G, Nuez I, Belicard T, Jiang Y, Zhao G, Franz CJ, Goldstein LD, Sanroman M, Miska EA, Wang D (2011) Natural and experimental infection of C. nematodes by novel viruses related to nodaviruses. PLoS Biol 9(1), e1000586
Faber PW, Alter JR, MacDonald ME, Hart AC (1999) Polyglutamine-mediated dysfunction and apoptotic death of a C. elegans sensory neuron. Proc Natl Acad Sci U S A 96(1):179–184
Link CD (1995) Expression of human beta-amyloid peptide in transgenic C. elegans. Proc Natl Acad Sci U S A 92(20):9368–9372
Kraemer BC, Zhang B, Leverenz JB, Thomas JH, Trojanowski JQ, Schellenberg GD (2003) Neurodegeneration and defective neurotransmission in a C. elegans model of tauopathy. Proc Natl Acad Sci U S A 100(17):9980–9985
Lakso M, Vartiainen S, Moilanen AM, Sirvio J, Thomas JH, Nass R, Blakely RD, Wong G (2003) Dopaminergic neuronal loss and motor deficits in C. elegans overexpressing human alpha-synuclein. J Neurochem 86(1):165–172
Wang J, Farr GW, Hall DH, Li F, Furtak K, Dreier L, Horwich AL (2009) An ALS-linked mutant SOD1 produces a locomotor defect associated with aggregation and synaptic dysfunction when expressed in neurons of C. elegans. PLoS Genet 5(1), e1000350
Moronetti Mazzeo LE, Dersh D, Boccitto M, Kalb RG, Lamitina T (2012) Stress and aging induce distinct polyQ protein aggregation states. Proc Natl Acad Sci U S A 109(26):10587–10592
Knight AL, Yan X, Hamamichi S, Ajjuri RR, Mazzulli JR, Zhang MW, Daigle JG, Zhang S, Borom AR, Roberts LR, Lee SK, DeLeon SM, Viollet-Djelassi C, Krainc D, O’Donnell JM, Caldwell KA, Caldwell GA (2014) The glycolytic enzyme, GPI, is a functionally conserved modifier of dopaminergic neurodegeneration in Parkinson’s models. Cell Metab 20(1):145–157
Cohen E, Bieschke J, Perciavalle RM, Kelly JW, Dillin A (2006) Opposing activities protect against age-onset proteotoxicity. Science (New York, NY) 313(5793):1604–1610
Florez-McClure ML, Hohsfield LA, Fonte G, Bealor MT, Link CD (2007) Decreased insulin-receptor signaling promotes the autophagic degradation of beta-amyloid peptide in C. elegans. Autophagy 3(6):569–580
Rosen DR, Siddique T, Patterson D, Figlewicz DA, Sapp P, Hentati A, Donaldson D, Goto J, O’Regan JP, Deng HX et al (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362(6415):59–62
Li J, Huang KX, Le WD (2013) Establishing a novel C. elegans model to investigate the role of autophagy in amyotrophic lateral sclerosis. Acta Pharmacol Sin 34(5):644–650
David DC, Ollikainen N, Trinidad JC, Cary MP, Burlingame AL, Kenyon C (2010) Widespread protein aggregation as an inherent part of aging in C. elegans. PLoS Biol 8(8), e1000450
Walther DM, Kasturi P, Zheng M, Pinkert S, Vecchi G, Ciryam P, Morimoto RI, Dobson CM, Vendruscolo M, Mann M, Hartl FU (2015) Widespread proteome remodeling and aggregation in aging C. elegans. Cell 161(4):919–932
Pan CL, Peng CY, Chen CH, McIntire S (2011) Genetic analysis of age-dependent defects of the C. elegans touch receptor neurons. Proc Natl Acad Sci U S A 108(22):9274–9279
Tank EM, Rodgers KE, Kenyon C (2011) Spontaneous age-related neurite branching in C. elegans. J Neurosci 31(25):9279–9288
Kauffman AL, Ashraf JM, Corces-Zimmerman MR, Landis JN, Murphy CT (2010) Insulin signaling and dietary restriction differentially influence the decline of learning and memory with age. PLoS Biol 8(5), e1000372
Tazearslan C, Cho M, Suh Y (2012) Discovery of functional gene variants associated with human longevity: opportunities and challenges. J Gerontol Ser A Biol Sci Med Sci 67(4):376–383
Acknowledgments
We thank the members of Lee laboratory for critical comments on the manuscript. This work was supported by the National Research Foundation of Korea (NRF) Grant funded by the Korean Government (MSIP) (NRF-2013R1A1A2014754) to S.-J.V.L.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing Switzerland
About this chapter
Cite this chapter
An, S.W.A., Artan, M., Park, S., Altintas, O., Lee, SJ.V. (2017). Longevity Regulation by Insulin/IGF-1 Signalling. In: Olsen, A., Gill, M. (eds) Ageing: Lessons from C. elegans. Healthy Ageing and Longevity. Springer, Cham. https://doi.org/10.1007/978-3-319-44703-2_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-44703-2_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-44701-8
Online ISBN: 978-3-319-44703-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)