Abstract
This chapter focuses on two major host responses recently found to be involved in CHIKV infection: autophagy and apoptosis. For each process, we first present molecular pathways and associated signalling, then we highlight the diverse strategies developed by host cells to prevent viral replication and virus-induced cell death, as well as by the virus to fight and hijack these host cell defence pathways.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Viral Replication
- Endoplasmic Reticulum Stress
- Unfold Protein Response
- Effector Caspases
- Autophagic Flux
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
This chapter focuses on two major host responses recently found to be involved in CHIKV infection: autophagy and apoptosis. For each process, we first present molecular pathways and associated signalling, then we highlight the diverse strategies developed by host cells to prevent viral replication and virus-induced cell death, as well as by the virus to fight and hijack these host cell defence pathways.
Autophagy Pathways and CHIKV
Autophagy Pathway
Autophagy is an intracellular degradative process highly conserved among eukaryotic cells that allows cells to recycle existing organelles and cytosolic components (Kuma and Mizushima 2010). It is required for cell development and survival of eukaryotes and has an impact on cell homeostasis, tumorigenesis, neurodegeneration, cancer, diabetes, and infection (Choi et al. 2013). It represents the primordial form of eukaryotic innate immunity against invading microorganisms (Deretic et al. 2013).
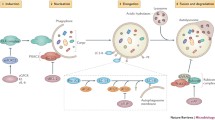
The autophagic process is initiated by the formation of a double-membrane vesicle surrounding cytosolic materials to be degraded, including proteins and organelles, to form an autophagosome. Then, fusion of the autophagosome with the endo-lysosomal compartment leads to an autophagolysosome. This process consists of three different steps, which require autophagy-related genes (Atgs) and organelles and involves complex interactions between dedicated protein machinery and subcellular organelles (Lamb et al. 2013; Fig. 1). The molecular machinery includes more than 30 Atgs, discovered in yeast, at least 18 of which are required for mammalian autophagy (Mizushima et al. 2011). The first step, called initiation, corresponds to the formation of the autophagic isolation membrane or phagophore (Mizushima 2010; Chan 2009). The second step includes the elongation and expansion of the phagophore that occur from multiple membrane sources (Lamb et al. 2013; Hamasaki et al. 2013) through an unknown process but likely by vesicular delivery, followed by the closure and completion of a double-membrane autophagosome. The elongation and closure are controlled by members of the Atg8 ubiquitin-like protein family (Geng and Klionsky 2008). The Atg8 ubiquitin-like protein family includes LC3 (LC3A, LC3B (referred to as LC3 henceforth), LC3C) and GABARAP subfamilies (GABARAP, GABARAPL1, and GABARAPL2). The soluble form of LC3 (referred to as LC3 hereafter) is termed LC3-I and the conjugated form LC3-PE as LC3-II. The LC3 conversion is widely used as a marker of autophagy flux (Klionsky et al. 2012). The last step is the maturation where the newly formed autophagosome fuses with endosomal compartment and/or with lysosomes to form the autophagolysosome.
Schematic representation of the nonselective and selective autophagy process. Autophagy is an evolutionarily conserved catabolic process in which intracellular material can be sequestered within double-membrane vesicles and targeted for degradation to lysosomes. Although autophagosomes can sequester cytosolic material nonspecifically in response to starvation (a), there is increasing evidence for selective autophagic degradation of various cellular structures, including protein aggregates, mitochondria, and pathogens (b). The selective autophagy process implicates autophagy receptors that mediate the docking of cargo to autophagosomes
Autophagy was previously described as a nonselective process but cumulative evidence has demonstrated its selectivity in recycling organelles, removing protein aggregates, and clearing specific viral proteins. Upon selective autophagy, autophagy receptors and the ubiquitination of the target are critical (Kirkin et al. 2009). Autophagy receptors are adaptor proteins, generally containing an ubiquitin-binding association domain (UBA) and an LC3-interaction region (LIR). Autophagy receptors can mediate the docking of ubiquitinated cargo to autophagosomes, thereby ensuring their selective degradation. The main autophagic receptors include p62 (SQSTM1), NBR1 (neighbour of BRCA1 gene 1), NDP52 (nuclear dot protein 52 kDa), and optineurin (Behrends and Fulda 2012). p62 is the best-characterized autophagy receptor and has been shown to target bacteria as well as viruses (Orvedahl et al. 2010; Mostowy and Cossart 2012).
Since the early reports, further studies have investigated the interplay between autophagy and viral infection and described that the autophagic process can be a host defence mechanism that clears intracytoplasmic viral products. However, viruses are able to subvert the autophagy machinery to favour their replication and release (Chiramel et al. 2013). Components of the autophagy machinery can therefore exert both an anti- or a pro-viral role, depending on the virus and the cell type considered (Dong and Levine 2013).
CHIKV Activates Autophagy
The evidence for the implication of the autophagy machinery during CHIKV infection, in cell cultures and in vivo, has been reported by several groups (Krejbich-Trotot et al. 2011; Judith et al. 2013; Joubert et al. 2012).
CHIKV infection induces autophagy as measured by the increased number of autophagosomes in infected human kidney epithelial cells (Krejbich-Trotot et al. 2011). Subsequent studies conducted by Judith et al. and by Joubert et al. showed that CHIKV infection triggers the conversion from LC3-I to LC3-II, a hallmark of the autophagy process, in primary and immortalised human cells as well as in mouse cells (Judith et al. 2013; Joubert et al. 2012). Analysis of the autophagy flux in the presence of lysosomal inhibitor and identification of autophagosomes and autolysosomes have proven evidence that CHIKV infection induces de novo autophagosome formation and that autophagosomes can fuse with lysosomes in CHIKV infected cells (Joubert et al 2012). Moreover, CHIKV infection decreases the level of p62, an autophagy receptor used as a marker for autophagic flux, providing evidence that CHIKV activates a complete autophagic response ending by the lysosomal degradation of the autophagic vesicle contents (Judith et al. 2013).
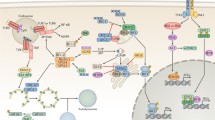
Although some viruses induce viral replication-independent autophagy, in most cases, autophagy induction by viruses is replication dependent, and initiated by a signal triggered either by viral replication steps, including entry and replication, or by accumulation of viral components or replication intermediates during the viral cycle. Indeed, this is the active CHIKV replication that induces autophagy, as it is not induced in cells treated with UV-inactivated CHIKV (Joubert et al. 2012). CHIKV promotes autophagy both by induction of endoplasmic reticulum (ER) stress and increase of reactive oxygen species (ROS) production (Joubert et al. 2012). ER stress is increased during viral infection and activates the unfolded protein response (UPR), which in turn induces autophagy. The UPR involves three different signalling pathways controlled by three integral ER membrane proteins: PERK, IRE-1α, and ATF6 (Hetz 2012). During CHIKV infection, accumulation of viral proteins in the ER may be the cause of ER stress, via an IRE1α- and XBP1s-mediated signalling pathway. ROS accumulation is a well-characterised host response to viral infections and free ROS are known to induce autophagy (Filomeni et al. 2014). CHIKV- induced ROS production induces autophagy through the inhibition of mTORC1. Both stress pathways act in an interdependent manner to enhance autophagic flux in CHIKV-infected cells (Joubert et al. 2012).
Antiviral Effect of Autophagy on CHIKV Infection
Xenophagy is a type of autophagy characterised by degradation of intracellular pathogens, helping to reduce their replication and spread. This type of autophagy involves selective recognition of pathogens that is ensured by particular autophagy receptors, such as p62 and NDP52 (Mostowy and Cossart 2012).
Judith et al. established direct antiviral roles for autophagy against CHIKV both in human and mouse cells. They found that CHIKV engages the molecular machinery of autophagy in a selective manner to protect infected cells (Fig. 2). By studying the implication of p62 in CHIKV infection, they found that the depletion of p62 significantly increased viral replication providing evidence that p62-mediated autophagy limits viral replication. They demonstrated that CHIKV capsid exhibits a cytotoxic effect and that the clearing of CHIKV capsid by p62 likely decreases its cellular toxicity, thereby limiting virus-induced cell death. They showed that by binding to LC3B, p62 recruit CHIKV capsid to the autophagosome in an ubiquitin-dependent manner and a SMURF1-independent manner, which degrade CHIKV capsid upon their fusion with lysosomes. Similarly, an earlier study was able to demonstrate the involvement of xenophagy during Sindbis virus (SINV) infection (Orvedahl et al. 2010). It has been reported that p62 delivered SINV capsids to degradation in autophagosome. However, even if SINV belongs to the same alphavirus genus as CHIKV, the signal recognition for the targeting of its capsids remains uncertain because, as opposed to CHIKV, it was reported to occur in an ubiquitin-independent manner but SMURF1-dependent mechanism. These observations raise questions regarding the status of CHIKV capsid (i.e., protein monomers or aggregates or assembled capsids), which is selectively targeted for autophagic degradation.
Antiviral and pro-viral effects of the autophagy machinery upon CHIKV infection. Viral replication upon CHIKV infection induces both oxidative and endoplasmic reticulum stress leading to the induction of the autophagy process. The autophagy process can play either an anti- or a pro-viral role upon CHIKV infection. The antiviral role of the autophagy process involves the autophagy receptor p62 and the autophagic protein LC3B. By targeting to degradation the toxic CHIKV-capsid, p62 facilitates its clearance by the autophagy process leading to the limitation of cell death. The pro-viral role of the autophagy process involves the autophagic receptor NDP52 and the autophagic protein LC3C. By binding to LC3C and the CHIKV-nsP2, NDP52 promotes viral infection and limits cell death
Pro-Viral Effect of the Autophagy Machinery on CHIKV Infection
Krejbich-Trotot et al. investigated the effect of CHIKV-induced autophagy on viral replication and found that overall it promotes CHIKV viral replication in human kidney epithelial cells. They showed that impairment of the autophagy machinery reduces CHIKV replication whereas its induction enhances it (Krejbich-Trotot et al. 2011). The same phenotype is observed in HeLa cells, where depletion of canonical mediators of autophagy, Beclin1 and Atg7, decreases CHIKV replication (Judith et al. 2013). During CHIKV infection, nonstructural CHIKV proteins (nsPs) bind to viral RNA to form replicative complexes (RC). Among them, CHIKV nsP2 has been shown by high-throughput yeast two-hybrid (HT-Y2H) assay (Bourai et al. 2012) to interact with NDP52, and the depletion of NDP52, similarly to that of canonical mediators of autophagy, decreases CHIKV replication (Judith et al. 2013; Fig. 2). This suggests that CHIKV nsP2 may engage the autophagy machinery to help virus replication through the binding of NDP52, in human cells. Further studies have shown that NDP52 associated with both LC3C and CHIKV nsP2, localizes to the trans-Golgi network-associated RCs that contain the other nsPs and double-stranded (ds)RNA replicative intermediate, in the vicinity of de novo protein synthesis (Judith et al. 2013). These observations suggest that NDP52 binding to CHIKV nsP2 and LC3C allows the anchorage of RCs to the TGN membrane.
However, one important result to consider is that mouse NDP52, in contrast to its human orthologue, is unable to bind to CHIKV nsP2, and LC3C is not expressed in mouse cells, accounting for the absence of promoting effect of the autophagy machinery on CHIKV infection in mouse cultured cells. The pro-viral role mediated by NDP52 is revealed by introducing human NDP52 and human LC3C in mouse cells, providing evidence of the species specificity of the pro-viral role of autophagy on CHIKV infection (Judith et al. 2013).
Apoptosis Pathway and CHIKV
Apoptosis Pathway
Apoptosis is highly conserved through evolution and is involved in the regulation of embryogenesis, development, and homeostasis by eliminating superfluous cells along these processes. Apoptosis can also be activated by a large number of stimuli as cell cycle perturbation, lack of nutrients, and viral infection. It is characterised by specific morphological features notably condensation and fragmentation of the nucleus, fragmentation of the mitochondrial network, and appearance of membrane blebs and apoptotic bodies (Taylor et al. 2008; Kerr et al. 1972).
The apoptosis process relies on the activation of cysteine aspartyl proteases known as caspases. Caspases are a conserved family of enzyme essential for initiation and execution of the apoptosis process. Caspases are central players in apoptosis because they catalyse many steps in the death pathway by irreversible cleavage of their substrates after aspartic acid residues. They are present as catalytically inactive proenzymes that are coordinately activated by caspase-specific cleavage. Two general classes of apoptotic caspases exist: initiator caspases including caspases 2, 8, 9, and 10, and effector caspases, which include caspases 3, 6, and 7. The initiator caspases are autoactivated under apoptotic condition, whereas effector caspases are activated in cascade through cleavage by initiator caspases. Effector caspases cleave a number of specific substrates, including structural components and regulatory proteins, leading to the destruction of cell–cell interactions and of the nuclear structure, reorganisation of the cytoskeleton, and inhibition of DNA synthesis (Kurokawa and Kornbluth 2009).
Apoptosis can be activated either by extrinsic or intrinsic stimuli. The extrinsic pathway is mediated by death receptors such as TNF receptors. Binding of the ligand to its death receptors induces a conformational change in the intracellular receptor domain that leads to the recruitment of apoptotic proteins to form the DISC (death inducing signalling complex, downstream of FASL/TRAIL) or complex I (downstream of TNFR). The inactive initiator caspase-8 is recruited to the DISC and subsequently activated, leading to the initiation of the apoptosis process (Wilson et al. 2009).
The intrinsic pathway, also called mitochondrial-dependent apoptosis, is triggered by intracellular signals such as UPR, DNA damage, hypoxia, and viral infection. The main actors of the intrinsic pathway are proteins of the Bcl2 family, which include subfamilies of antiapoptotic, pro-apoptotic, and BH3-only proteins. In response to stress signals, members of the BH3-only proteins are activated and stimulate the assembly of pro-apoptotic effector, notably BAX and BAK into oligomers. These oligomers form a pore into the mitochondrial membrane that leads to the release of apoptotic factors into the cytosol, in particular cytochrome C. The cytochrome C associates within the apoptosome, a multiprotein complex, and initiates apoptosis via the recruitment of the inactive initiator caspase-9. Caspase 9 cleaves and activates effector caspases, caspase 3, and caspase 7, leading to apoptosis. This cascade can be alternatively activated through the upstream caspase-8 in response to an extrinsic signal (Kroemer et al. 2007).
Many viral proteins disturb normal cell physiology and deliver upstream signals that end up in a death response by apoptosis. Apoptosis is an integral part of the host defence against invading intracellular pathogens, in particular viruses, which serves to limit pathogen replication (Upton and Chan 2014; Li and Stollar 2004). However, viral genomes often encode apoptosis inhibitors in order to impair apoptosis and as such promote their replication and persistence (Everett and McFadden 1999). On the contrary, viruses can use apoptosis to kill infected host cells at the end of the viral replication cycle to increase the dissemination of their progeny and limit inflammatory responses. Due to the packing of the entire cellular content into apoptotic bodies, viruses or viral material can be rapidly taken up by surrounding cells (Kepp et al. 2009).
As CHIKV is highly cytopathic for mammalian cells, numerous studies have been conducted to define the type of cell death responsible for the cytopathic effect in CHIKV-infected cells.
CHIKV Activates Apoptosis
In vitro studies have shown that death of human infected cells is associated with the presence of a marker of apoptosis: active cleaved form of caspase-3 (Sourisseau et al. 2007). CHIKV-infected cells display a mitochondrial relocalisation of Bax, as well as the presence of cleaved PARP in infected cells, a well-known target of the effector caspases. It has also been shown, by using pharmacological inhibitors of apoptosis, as well as cells unable to engage the apoptotic pathway, that the main form of CHIKV-induced cell death is caspase-mediated apoptosis (Joubert et al. 2012; Krejbich-Trotot et al. 2011). To define whether the intrinsic or extrinsic pathways are triggered upon CHIKV infection, the cleavage of two specific caspases, caspase-9 (intrinsic pathway) and caspase-8 (extrinsic pathway), has been analysed. CHIKV-induced apoptosis is triggered through an early caspase-9 intrinsic pathway, followed by a caspase-8 extrinsic dependent pathway. Moreover, CHIKV-induced apoptosis requires viral replication, as UV-inactivated CHIKV fails to cause apoptosis (Joubert et al. 2012; Krejbich-Trotot et al. 2011).
Pro-Viral Function of Apoptosis
Krejbich-Trotot et al. have reported that the apoptotic process promotes CHIKV dissemination in human cells (Krejbich-Trotot et al. 2011). They demonstrated that apoptosis inhibition decreases CHIKV infection by using drugs preventing apoptosis cell fragmentation, and that apoptosis contributes to perpetuate virus spreading through the formation of apoptotic bodies. Actually, CHIKV hijacks the apoptotic process through the formation and release of apoptotic blebs enclosing viral materials protected into membrane vesicles, promoting the infection of neighbouring cells (Fig. 3).
Dual effect of apoptosis on CHIKV infection. CHIKV infection induces two apoptotic pathways, the intrinsic and extrinsic pathway. This induction of apoptosis can play either a pro- or an antiviral function. Apoptosis plays an antiviral role by promoting cell death limiting viral propagation. By forming apoptotic blebs containing viral components, apoptosis plays a pro-viral role. The apoptotic blebs disseminate the infection by infecting the neighbouring cells
This mechanism was first reported for the SINV (Rosen et al. 1995). This process also limits the inflammatory response and thereby favours infection spreading in the infected host. Viral particles or materials enclosed within apoptotic vesicles are also protected from inactivation by host antibodies and proteases.
Overall Effects of Autophagy and Apoptosis on Cell Survival and Infection
CHIKV, by subverting the autophagy machinery, protects human infected cells against cell death and favours its replication (Munz 2013). Cell death is essential in many biological processes, and apart from apoptosis, there is an increased recognised role of other death modalities such as necroptosis and autophagic cell death in host response to infection (Tait et al. 2014).
Joubert et al. have shown, in CHIKV-infected mouse cells, a relationship between autophagy and apoptosis. By using cells unable to engage either the autophagy or the apoptotic pathway, they provided evidence that autophagy in CHIKV-infected cells promotes cell survival and delays apoptosis upon infection (Joubert et al. 2012). Moreover, mice with reduced autophagy, Atg16LHM mice (Cadwell et al. 2008), display higher susceptibility and higher lethality to CHIKV infection (Joubert et al. 2012). In human cells, the depletion of canonical mediators of autophagy, Beclin1 and Atg7, increases virus-induced cell death, indicating that autophagy also plays essentially a pro-survival role upon CHIKV infection in human cells (Judith et al. 2013; Fig.2). Two other autophagy mediators, p62 and NDP52, play a pro-survival role in CHIKV-infected human cells: p62 facilitates the clearance of CHIKV capsid, whereas NDP52 binds to CHIKV nsP2 in the cytosol and restricts transcriptional shutoff and apoptosis. Nuclear nsP2 indeed serves as a trigger for transcriptional shutoff and induction of apoptosis in SINV- and CHIKV-infected cells and these functions are assigned to its carboxy-terminal domain (Garmashova et al. 2006, 2007; Bourai et al. 2012). Thus, binding to NDP52 in the TGN-derived membranes retains nsP2 in the cytoplasm and restricts its migration in the nucleus, limiting transcriptional shutoff and cell death (Judith et al. 2013).
By facilitating the clearance of CHIKV capsid, autophagy plays an antiviral role, and limits infection-associated cell death. However, the cytoprotective role of autophagy, in addition to the fact that it is beneficial for the cell, can also be advantageous at the host level for the virus, as viral replication requires a living host cell. Premature cell death has also been considered as an anti-viral host mechanism that limits viral propagation (Fig. 3).
In conclusion, studies on CHIKV replication and the discovery that autophagy and apoptosis pathways are triggered by infection illustrate the intimate interconnection between these pathways in host response to infection.
References
Behrends C, Fulda S (2012) Receptor proteins in selective autophagy. Int J Cell Biol 2012:673290. doi:10.1155/2012/673290
Bourai M, Lucas-Hourani M, Gad HH, Drosten C, Jacob Y, Tafforeau L et al (2012) Mapping of chikungunya virus interactions with host proteins identified nsP2 as a highly connected viral component. J Virol 86(6):3121–3134. doi:10.1128/JVI.06390-11
Cadwell K, Liu JY, Brown SL, Miyoshi H, Loh J, Lennerz JK et al (2008) A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456(7219):259–263. doi:10.1038/nature07416
Chan EY (2009) mTORC1 phosphorylates the ULK1-mAtg13-FIP200 autophagy regulatory complex. Sci Signal 2(84), e51. doi:10.1126/scisignal.284pe51
Chiramel AI, Brady NR, Bartenschlager R (2013) Divergent roles of autophagy in virus infection. Cells 2(1):83–104. doi:10.3390/cells2010083
Choi AM, Ryter SW, Levine B (2013) Autophagy in human health and disease. N Engl J Med 368(19):1845–1846. doi:10.1056/NEJMc1303158
Deretic V, Saitoh T, Akira S (2013) Autophagy in infection, inflammation and immunity. Nat Rev Immunol 13(10):722–737. doi:10.1038/nri3532
Dong X, Levine B (2013) Autophagy and viruses: adversaries or allies? J Innate Immun 5(5):480–493. doi:10.1159/000346388
Everett H, McFadden G (1999) Apoptosis: an innate immune response to virus infection. Trends Microbiol 7(4):160–165
Filomeni G, De Zio D, Cecconi F (2014) Oxidative stress and autophagy: the clash between damage and metabolic needs. Cell Death Differ 22(3):377–388. doi:10.1038/cdd.2014.150
Garmashova N, Gorchakov R, Frolova E, Frolov I (2006) Sindbis virus nonstructural protein nsP2 is cytotoxic and inhibits cellular transcription. J Virol 80(12):5686–5696. doi:10.1128/JVI.02739-05
Garmashova N, Gorchakov R, Volkova E, Paessler S, Frolova E, Frolov I (2007) The Old World and New World alphaviruses use different virus-specific proteins for induction of transcriptional shutoff. J Virol 81(5):2472–2484. doi:10.1128/JVI.02073-06
Geng J, Klionsky DJ (2008) The Atg8 and Atg12 ubiquitin-like conjugation systems in macroautophagy. ‘Protein modifications: beyond the usual suspects’ review series. EMBO Rep 9(9):859–864. doi:10.1038/embor.2008.163
Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N et al (2013) Autophagosomes form at ER-mitochondria contact sites. Nature 495(7441):389–393. doi:10.1038/nature11910
Hetz C (2012) The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Biol 13(2):89–102. doi:10.1038/nrm3270
Joubert PE, Werneke SW, de la Calle C, Guivel-Benhassine F, Giodini A, Peduto L et al (2012) Chikungunya virus-induced autophagy delays caspase-dependent cell death. J Exp Med 209(5):1029–1047. doi:10.1084/jem.20110996
Judith D, Mostowy S, Bourai M, Gangneux N, Lelek M, Lucas-Hourani M et al (2013) Species-specific impact of the autophagy machinery on chikungunya virus infection. EMBO Rep 14(6):534–544. doi:10.1038/embor.2013.51
Kepp O, Senovilla L, Galluzzi L, Panaretakis T, Tesniere A, Schlemmer F et al (2009) Viral subversion of immunogenic cell death. Cell Cycle 8(6):860–869
Kerr JF, Wyllie AH, Currie AR (1972) Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br J Cancer 26(4):239–257
Kirkin V, McEwan DG, Novak I, Dikic I (2009) A role for ubiquitin in selective autophagy. Mol Cell 34(3):259–269. doi:10.1016/j.molcel.2009.04.026
Klionsky DJ, Abdalla FC, Abeliovich H, Abraham RT, Acevedo-Arozena A, Adeli K et al (2012) Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8(4):445–544
Krejbich-Trotot P, Gay B, Li-Pat-Yuen G, Hoarau JJ, Jaffar-Bandjee MC, Briant L et al (2011) Chikungunya triggers an autophagic process which promotes viral replication. Virol J 8:432. doi:10.1186/1743-422X-8-432
Kroemer G, Galluzzi L, Brenner C (2007) Mitochondrial membrane permeabilization in cell death. Physiol Rev 87(1):99–163. doi:10.1152/physrev.00013.2006
Kuma A, Mizushima N (2010) Physiological role of autophagy as an intracellular recycling system: with an emphasis on nutrient metabolism. Semin Cell Dev Biol 21(7):683–690. doi:10.1016/j.semcdb.2010.03.002
Kurokawa M, Kornbluth S (2009) Caspases and kinases in a death grip. Cell 138(5):838–854. doi:10.1016/j.cell.2009.08.021
Lamb CA, Yoshimori T, Tooze SA (2013) The autophagosome: origins unknown, biogenesis complex. Nat Rev Mol Cell Biol 14(12):759–774. doi:10.1038/nrm3696
Li ML, Stollar V (2004) Alphaviruses and apoptosis. Int Rev Immunol 23(1–2):7–24
Mizushima N (2010) The role of the Atg1/ULK1 complex in autophagy regulation. Curr Opin Cell Biol 22(2):132–139. doi:10.1016/j.ceb.2009.12.004
Mizushima N, Yoshimori T, Ohsumi Y (2011) The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol 27:107–132. doi:10.1146/annurev-cellbio-092910-154005
Mostowy S, Cossart P (2012) Bacterial autophagy: restriction or promotion of bacterial replication? Trends Cell Biol 22(6):283–291. doi:10.1016/j.tcb.2012.03.006
Munz C (2013) Macroautophagy—friend or foe of viral replication? EMBO Rep 14(6):483–484. doi:10.1038/embor.2013.55
Orvedahl A, MacPherson S, Sumpter R Jr, Talloczy Z, Zou Z, Levine B (2010) Autophagy protects against Sindbis virus infection of the central nervous system. Cell Host Microbe 7(2):115–127. doi:10.1016/j.chom.2010.01.007
Rosen A, Casciola-Rosen L, Ahearn J (1995) Novel packages of viral and self-antigens are generated during apoptosis. J Exp Med 181(4):1557–1561
Sourisseau M, Schilte C, Casartelli N, Trouillet C, Guivel-Benhassine F, Rudnicka D et al (2007) Characterization of reemerging chikungunya virus. PLoS Pathog 3(6), e89. doi:10.1371/journal.ppat.0030089
Tait SW, Ichim G, Green DR (2014) Die another way—non-apoptotic mechanisms of cell death. J Cell Sci 127(Pt 10):2135–2144. doi:10.1242/jcs.093575
Taylor RC, Cullen SP, Martin SJ (2008) Apoptosis: controlled demolition at the cellular level. Nat Rev Mol Cell Biol 9(3):231–241. doi:10.1038/nrm2312
Upton JW, Chan FK (2014) Staying alive: cell death in antiviral immunity. Mol Cell 54(2):273–280. doi:10.1016/j.molcel.2014.01.027
Wilson NS, Dixit V, Ashkenazi A (2009) Death receptor signal transducers: nodes of coordination in immune signaling networks. Nat Immunol 10(4):348–355. doi:10.1038/ni.1714
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Judith, D., Couderc, T., Lecuit, M. (2016). Chikungunya Virus-Induced Autophagy and Apoptosis. In: Okeoma, C. (eds) Chikungunya Virus. Springer, Cham. https://doi.org/10.1007/978-3-319-42958-8_9
Download citation
DOI: https://doi.org/10.1007/978-3-319-42958-8_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-42956-4
Online ISBN: 978-3-319-42958-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)