Abstract
The cancer cells obtain their invasion potential not only by genetic mutations, but also by changing their cellular biophysical and biomechanical features and adapting to the surrounding microenvironments. The extracellular matrix, as a crucial component of the tumor microenvironment, provides the mechanical support for the tissue, mediates the cell-microenvironment interactions, and plays a key role in cancer cell invasion. The biomechanics of the extracellular matrix, particularly collagen, have been extensively studied in the biomechanics community. Cell migration has also enjoyed much attention from both the experimental and modeling efforts. However, the detailed mechanistic understanding of tumor cell-ECM interactions, especially during cancer invasion, has been unclear. This chapter reviews the recent advances in the studies of ECM biomechanics, cell migration, and cell-ECM interactions in the context of cancer invasion.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Extracellular matrix
- Cell-ECM interactions
- Cell migration
- Mathematical models
- Collagen
- Mechanotransduction
- Cancer invasion
4.1 Introduction
The tumor microenvironment is created by proliferating tumor cells and dominated by tumor-induced interactions [112]. It has been well accepted that the tumor microenvironment plays a significant role in disease progression, but the precise function of each constituent remains unclear. The tissue microenvironment of a developing tumor can be broken down into three categories: the biological, the chemical, and the biophysical/biomechanical. The biological environment is comprised of the cellular constituents that surround the malignant cancer cells. A variety of infiltrating immune cells [112], cancer-associated fibroblasts [100], and angiogenic endothelial cells [110] perform critical functions in sustaining cell proliferation, evading growth suppressors, promoting survival, activating invasion and metastasis, as well as reprogramming energy metabolism. The chemical environment refers to the abnormal distribution of oxygen, nutrients, wastes, and cytokines, as well as many growth factors and inhibitors. For example, excess growth of the tumor cells leads to a hypoxic environment [55], elevated oxidative stress [22], and consequently, the accumulation of lactic acid due to anaerobic metabolism [44] and up-regulation growth factor production (e.g., VEGF). The biophysical and biomechanical aspect of the tumor is both the physical and geometrical constraints from the tissue structure, and the mechanical interactions between the tumor and surrounding environment, most importantly the extracellular matrix (ECM). This category of the microenvironment has only recently begun to receive an increased level of attention, including studies on the hydrostatic stress from interstitial fluid [12], substrate topography [61, 66, 82], and the biomechanics of the extracellular matrix [71]. In this chapter, we focus on the emerging concepts in the contribution of ECM heterogeneity and remodeling to tumor growth and invasion.
It has been appreciated for some time that the extracellular matrix (ECM) plays an important role in all the stages of cancer development. In breast cancer, dense breast tissue on mammography is associated with increased collagen content. Women with more than 75 % dense regions have been shown to have an increased risk of breast cancer by up to five fold in comparison to women with less than 5 % density [13, 80]. Breast density is common, heritable, and has been postulated to account for up to one third of breast cancers [80]. In mouse models of breast cancer, it has been shown that increasing either the density or the crosslinking of collagen promotes invasiveness and, to a lesser extent, the formation of breast cancer [74, 87, 90], confirming the critical role of extracellular matrix (ECM) in assisting tumor progression.
Moving beyond correlative data and understanding the underlying mechanisms is difficult using traditional experimentation alone, since ECM affects many aspects of both host and tumor cell behavior, such as migration, differentiation, invasion, and proliferation. Furthermore, the properties of ECM itself are also complex, with diverse topographies and mechanical properties possibly depending on density, alignment, polymerization, and crosslinking. Because tumor invasion and growth are emergent outcomes of the complex interactions between cells and ECM, computational and mathematical models are becoming necessary tools to help dissect this complexity. Here, we will review the recent advances in the understanding of how cell-ECM interactions help to regulate cancer invasion, focusing on the biomechanical effects.
4.2 ECM in Cancer Invasion
The ECM, a fibrous macromolecular network outside cells, plays a crucial role in tissue environments, providing mechanical structures [94] as well as promoting cell phenotype change [75]. Through direct or indirect means, the ECM regulates almost all cellular behavior and is indispensable for developmental processes [78]. Recent experimental evidence has suggested that cancer cells interact with ECM fibers during invasion, condensing [106], remodeling [106], and aligning [89] fibers. Using in vitro mouse breast cancer models, Provenzano and coworkers discovered three tumor-associated collagen signatures (TACS): TACS-1 with dense collagen near the tumor, TACS-2 with stretched collagen fibers encasing the tumor, and TACS-3 with aligned collagen fibers normal to the tumor boundary. Despite the fact that the mechanisms are still unclear, it has been well accepted that breast tumors are associated with dense breast tissue, notably dense collagen [89]. At the early stage of cancer, collagen fibers condense near the tumor, interacted with growing cancer cells (Fig. 4.1A). As cancer progresses, cancer cells invade outward. Migrating cells supposedly pull on the surrounding ECM fibers, producing stretched fibers, but the mechanism for the aligned fibers normal to the tumor boundary is still unclear (Fig. 4.1B) [89]. A recent review summarizes remodeled ECM as an anomaly that deregulates behavior of stromal cells, facilitates tumor angiogenesis and inflammation, and leads to a tumorigenic microenvironment [79].
The ECM in cancer invasion. Multiphoton microscopy images of mouse breast tumor: (a) dense collagen, and (b) aligned collagen fibers (From Provenzano etal. [89] with permission). Yellow outline in (a) is a tumor boundary. Single (c) and multiple (d) U87 glioblastoma cells modified collagen fiber structures 10 h after gel polymerization (From Vader et al. [106] with permission), cell nuclei are green, and collagen fibers are red. Multiphoton intravital microscopy images of a tip cell of invasion into a mouse dermis (e) and the multicellular core (f) (From Alexander et al. [3] with permission)
The remodeling of collagen fibers by invasive cancer cells has also been observed in vitro, where collagen fibers condense near single glioblastoma cells (Fig. 4.1C) and aligned fiber tracks appear between multiple migrating glioblastoma cells (Fig. 4.1D) [106]. Cancer invasion in vivo is more complicated. Using intravital microscopy, Alexander et al. [3] observed melanoma cells invading into the mouse dermis. They suggested that heterogeneous connective tissue, in particular the porous 3D ECM network, provides a guidance or track for invasive cancer cells (Fig. 4.1E). They also observed that, in addition to individual migrating cells, cancer cells often invade collectively as a multicellular unit with cell-cell junctions retained [37]. This suggests that the leader cell searches for a pore space in the ECM fiber network and squeezes itself through the space, whereas the following cells collectively invade using the track (Fig. 4.1F) [3].
Both in vitro and in vivo evidence shows substantial ECM remodeling associated with proliferating and invading cancer cells. However, because of the complexity of the microenvironment, many other factors could potentially contribute to ECM remodeling, including tumor associated fibroblasts [100] that can produce or degrade the ECM. In order to understand how mechanical cell-ECM interactions contribute to the remodeling, theoretical and computational models have been developed to further investigate the mechanical properties of ECM fibers and their interactions with cells. A two-dimensional (2D) discrete fiber network model using the finite element method simulated ECM fiber remodeling by contractile force from a single cell and between two cells [1]. Anisotropic contractile forces produce ECM fiber patterns (Fig. 4.2a, b) resembling the experimental observations [1]. More recently, a three-dimensional (3D) elastic fiber network model using a bead-and-spring fiber representation with elastic crosslinking simulated tensile and shear tests for random and aligned fiber networks [71] (Fig. 4.2c, d). Their simulations show that aligned fiber network structure is stiffer than the random network, while both structures showed nonlinear strain-stiffening. The stress-strain curve of a random fiber network illustrates how the matrix responds to external strain. Upon small strain, the network first responds with minimal stress, like a fluid. As the external strain increases, the stress increases slowly until the strain reaches about 10 %, when the fibers start to align. Between 10 and 30 % strain, the stress of the fiber network increases linearly, indicating that the network behaves like an elastic material. At 30 % strain, the fiber alignment reaches 70 % [93], after which the fibers will be stretched to show a much stiffer bulk response. The residual stress distributions, depicted as force vectors, after a shear test and a local displacement showed the nonaffine deformation of the network and the accumulation of stress at the boundary of displacement (Fig. 4.2e, f). Feng et al. [32] explored the role of fiber alignment in a fiber network using a Landau-type theory for the nonlinear elasticity with the order parameter taking into account the kinematic order of fibers. Comparing the theory and simulation of a disordered lattice model, they concluded that the nonlinear elastic behavior of biopolymer gels arises from strain-induced fiber alignment, suggesting that it explained contact guidance of cell motility.
Computational models of ECM. A two-dimensional ECM fiber model showing configurations upon anisotropic contraction from (a) a single cell and (b) two cells (From Abhilash et al. [1] with permission). A three-dimensional elastic bead-spring fiber network model for random (c), and pre-aligned structure (d) Black lines are fibers, and red lines are crosslinkers. The residual stress distribution for a random fiber network upon a shear strain (e) and a local box displacement at the center of the fiber network, mimicking a local deformation imposed by a migrating cell (f) (From Lee et al. [71] with permission)
4.3 Cell-ECM Interactions
Regulation of cell motility and invasiveness in the ECM is complex. On one hand, deposition of fibrillar collagen appears to promote tumor cell motility by providing one-dimensional or two-dimensional “tracks” for cell movement [31, 38]. Further crosslinking of collagen fibrils by enzymes such as lysyl oxidase may increase the alignment and rigidity of those tracks, aiding cell invasiveness [2, 74]. Increasing collagen density may also inhibit cell migration and require proteolytic activity to allow tumor cell migration [113, 115]. On the other hand, cellular machinery that recognizes not only the biochemical diversity of the ECM, but also its physical and topographical characteristics, such as rigidity, dimensionality and ligand spacing is critical for the response of cells to ECM.
It has been increasing clear that the cellular response to environmental signaling goes far beyond the ability of chemically sensing specific ECM ligands and encompass a wide range of physical cues and the adhesive interface [47]. More attempts on understanding cell migration during tumor invasion are focusing on the interplay of multiscale mechanotransduction, which is comprised of how the cell sense and react to internally generated and externally applied signals [45]. Current understanding of the biomechanics of cell-matrix interactions is based primarily on in vitro studies of the cell leading edge of migration (focal adhesion and membrane remodeling), and intra-cellular cytoskeletal activities (actin protrusion, actomyosin contraction, and cell motility signaling pathway). All these elements need to work in concert to regulate cell migration speed, directionality, and cell migration plasticity. We discuss these elements of cell motility below.
4.3.1 Focal Adhesion
Focal adhesions are integrin-based structures that mediate strong cell-substrate adhesion and transmit information between the extracellular matrix and the cytoplasm [46, 48]. During the formation of focal adhesion, a subset of adhesion components with actin nucleates the nascent adhesion, which is stabilized by its association with integrin to form stable focal adhesion assembly [70, 72, 98]. Increasing the strength and longevity of integrin binding and integrin clustering is a crucial step in this adhesion process. Active integrin complexes promote recruitment of cytoskeletal components, activate signaling molecules, and enhance adhesive force [23]. In particular, integrin activation regulates microtubule dynamics and helps to stabilize microtubules at the cell cortex [14]. Integrins connect the ECM to the cytoskeleton and provide cells with mechanical anchorages and signaling platforms. At the molecular level, force-induced strengthening of cell adhesion [4, 19, 42] has been explained in terms of recruitment of integrins and cytoskeletal proteins [92] and/or ligand-integrin catch bonds [39]. Furthermore, cyclic mechanical reinforcement [68] is found to be a more effective regulatory mechanism than the catch bond, as it prolongs the bond lifetime for fibronectin and integrin-α5β1 [28]. While the short-lived integrin-ligand bonds may allow the cell to rapidly explore its environment, the long-lived integrin-ligand bonds are critical to adhesion maturation and downstream signaling, which takes tens of seconds to minutes [42]. The mechanically reinforced ligand-integrin bonds enable nascent adhesion to be stabilized by myosin-generated contractile forces.
Live-cell microscopy studies reveal four main stages in the “life cycle” of integrin-mediated adhesions, including nascent adhesions, focal complexes, focal adhesions, and fibrillar adhesions [116]. Nascent adhesions are submicron-sized, and are barely visible by means of ordinary fluorescence microscopy. The process of generating focal complexes is on a timescale of seconds and involves only a small number of integrin that triggers actin polymerization [119]. Measurements of mechanical tension across vinculin, a protein that connects integrins to actin filaments, also showed that vinculin recruitment to focal adhesions and force transmission to vinculin are regulated separately [49]. The subsequent strengthening of adhesions through myosin pulling is believed to lead to the recruitment of additional adhesive proteins, which promotes the growth of larger focal complexes. The growth processes depend on actomyosin-based stress fibers and also require the stress fibers to serve as physical contractile anchors [83]. The transformation of one form of adhesion into another is tightly regulated by the cellular signaling system and is also mediated by cues from ECM and intracellular structures (Fig. 4.3).
Molecular architecture of cell-ECM interactions centered around focal adhesion (From Wehrle-Haller [109] with permission)
Both ECM rigidity and ligand spacing have been found to influence focal adhesion, stress fiber assembly, cell spreading, cell migration speed, and adhesive forces [60]. Adhesive area is also found to strongly modulate adhesion strength, integrin binding, and vinculin and talin recruitment [43]. Interestingly, cells cannot integrate signals from integrin-ligand complexes spaced more than 58 nm from each other, as demonstrated in experiments of cells sitting on fibronectin nano-islands within non-adhesive background [16]. The minimal area of integrin-fibronectin clusters required for stable focal adhesion assembly and force transmission is not a predetermined value; it arises dynamically from the interaction between pathways controlling adhesive force, cytoskeletal tension, and the structural linkage that transmits these forces [23, 81].
4.3.2 Intracellular Mechanical Structures
The intracellular mechanical structures that play a key role in cell migration include actin microfilaments, intermediate filaments, lamin, nucleoskeleton and cytoskeleton linker, microtubules, and cell nucleus. The latter adds an additional layer of mechanical stability because of its significant stiffness [25, 50] and the possibility to physically divide the cytoplasm into forward and rear compartments [88]. These structures can be altered during cancer progression [9]; e.g., cell nucleus deformation can be a function of malignancy [27, 40].
Actin microfilaments provide the largest contribution to cell body stiffness when probed at adhesion sites [10, 11, 42, 58]. They are organized into different structures, including actin bundles and stress fibers. They stabilize cell architecture, including the formation of lamellipodia and filopodia, which play important roles in cell motility [57, 108]. Actin participates as an internal stabilizer and a dynamic mechanical structure in cells for migration and mechanosensing [9]. The inherent elastic features and the myosin-mediated contractility of actin fibers [10, 42, 59] and the linkage to ECM via focal adhesion [42, 57] together regulate cell-ECM interaction.
As the load–bearing element in the cell [59], the microtubule network provides internal structural support while contributing to the polarization and initiation of cell migration [64, 101]. The microtubules allowing the cell to polarize in response to ECM cues contribute to spatial organization and participate in initiating cell migration [9]. Large scale disruption of the microtubules has dramatic mechanical consequences on cell stiffness [10]. During cell migration, the microtubule depolymerization and the inhibition of the microtubule-associated molecular motors can effectively impair cell motility [63]. Focal adhesion is found to be necessary for the microtubule depolymerization and the microtubules are also required for focal adhesion disassembly and regulation [52, 62].
Intermediate filaments are the most diverse family of cytoskeletal components that exist as associated effectors of the cytoskeletal framework through connections with actin and the microtubules. The overexpression of intermediate filament proteins during transformation process is notably connected to carcinomas [21]. In most epithelia cells, intermediate filaments span the cell cortex and wind around the nucleus to form an interconnected network that provides a continuous link between focal adhesions, cell-cell adhesions and the nucleus through the linker of nucleoskeleton and cytoskeleton complex [26, 77].
The cell nucleus is the largest and the stiffest organelle with the ability to affect cell migration through nucleocytoskeletal connections [9]. The nucleoskeleton links directly to the cytoplasmic cytoskeleton through the linkers that connects the lamin network in the nucleus to actin and intermediate filaments [20]. Functionally, the nucleus sustains global deformation and changes in its sub-nuclear spatial organization when the cell is subjected to mechanical stress, indicating that the nucleus is also a mechanosensitive element participating in cell-ECM interactions. The features of the nucleus regulate cell migration, but the exact mechanisms are not clear. However, it has been found that the nucleus physically divides the cytoplasm into forward and rear pressure compartments when a human fibroblast migrates through a 3D ECM [88]. This finding suggests that the nucleus can act as a piston to increase the hydrostatic pressure between the nucleus and the leading edge of the cell in order to drive lamellipodia-independent 3D cell migration.
4.3.3 Cell Membrane Remodeling and Mechanotransduction Signaling Network
Cell membrane tension together with the pressure generated by intra-cellular structures and the focal adhesion strength are the forces that define the movement of cell membrane. The contact angle between substrate and membrane has been found to correlate with the load on actin polymerization and cell protrusion rate [41]. This result emphasizes the fundamental importance of membrane configuration for cellular force balance and the subcellular scale biophysical dynamics. In a model trying to explain the cell morphology of slime mold Dictyostelium during its chemotactic migration, an analogy was drawn between the interaction of pushing microtubules with the surface tension of the plasma membrane and the Marangoni effect that generates the tear drops of wine-covered glass (Fig. 4.4a) [97]. The protrusion results from a dynamic balance between the outward push from cytoskeletal fibers and membrane surface tension, while the wine tear pattern arises from the balance between gravity and the surface tension gradient between alcohol and water. Furthermore, the recent biochemical understanding of reaction-diffusion inside the cell has led to the similar results of cell morphology. For chemotactic cell migration, the local excitation global inhibition (LEGI) model [97] (Fig. 4.4) and its variations were proposed to explain the signaling responses of cells exposed to gradients of chemoattractant [86]. The response to a stimulus is mediated through the balance between a fast, local excitation and a slower, global inhibition process, both of which are controlled by receptor occupancy [30, 69, 73]. When stimulated by uniform concentration of chemoattractant, the faster local excitation rises with receptor occupancy, leading to an increase in the response. As the slower inhibition rises, the response subsides, ensuring perfect adaptation. When in a gradient, local excitation mirrors receptor occupancy, and hence, chemoattractant concentration gradients. The inhibition process integrates the global signal, leading to an inhibitory signal that is equivalent to the average level of receptor occupancy in the cell. This model has satisfactorily explained Ras, PTEN and PI3K activation during amoeboid Dictyostelium cell migration [118]. Thus far the LEGI models seem most promising in providing a plausible mechanism for chemotactic migration, possibly applicable to more generic cell migration as well.
Many studies of the molecular mechanisms of cell motility signaling pathways have focused on the Rho family of small GTPases that regulate the cytoskeleton-dependent processes. The complexity of the interactions among the Rho family of proteins, their regulators, and effectors (Fig. 4.4d) is challenging to both experimental and mathematical studies. Integrating the cell motility pathway to mechanotransduction network alone is difficult. Moreover, the spatiotemporal reaction-diffusion dynamics of the signaling molecules in the cytosol and on cell membrane are thought to be the key determinants of cell migration plasticity [85].
4.3.4 Cell Migration Modes
Cell migration plasticity refers to the cell’s ability to switch between different cell migration modes. The migration modes were originally classified based on the cell morphology alone, but have since been extended to describe the multi-scale properties of cell migration, including cell shape, cell migration speed, and organization of intracellular structures. The main categories are individual (amoeboid and mesenchymal), and collective (as cohesive multicellular units) migration [34]. The amoeboid migration refers to the movement of round or ellipsoid cells that lack mature focal adhesions and stress fibers [38, 85] with blebby membrane dynamics and faster migration speed. Mesenchymal migration is characterized by a spindle-like elongated cell shape, actin-rich filopodia and more focalized cell–matrix interactions; mesenchymal movement resembles the migration of a fibroblast [103], whereby the cell entangles with the ECM [38].
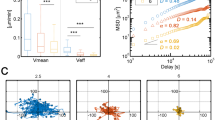
Individual cell migration modes differ depending on cell type, developmental stage, local environment, and disease state [111]. Cell migration mode can be dynamically changed by the strength of adhesion, physical confinement (e.g., squeezed between two surfaces), contractility, and chemical cues [76, 95]. With low adhesion and strong confinement, slow mesenchymal cells can switch to fast amoeboid migration, suggesting that no specific genetic alteration is necessary for tumor cells to escape the primary tumors [29]. In vitro evidence shows that, in confined 3D ECM, the intrinsic fluctuation in cortical contraction is sufficient to trigger the switch from embryonic progenitor cells to prototypic amoeboid migration mode [95]. Cancer cell migration persistence and local membrane protrusion persistence have been measured in vivo [102] (Fig. 4.5a). However, because of the lack of local measurements of ECM dynamics concurrently, cellular and subcellular imaging has not been able to offer comprehensive understanding of the cell-ECM interaction. Recent advances in combination of live-cell imaging, molecular manipulation and force measurement have revealed multiscale cell migration details with extraordinary precision that allowed for mechanistic mathematical modeling. An ideal 2D experimental cell migration model has been the fish epithelial keratocytes for investigating cell shape determination [6, 65]. Individual keratocytes maintain nearly constant shape, speed, and direction over many cell lengths of migration, with considerable heterogeneity within a population of keratocyte [65] (Fig. 4.5b). Several mathematical models have been developed to take advantage of such data and explained the detailed intracellular signaling and mechanical interactions leading to the specific cell shape during migration (Fig. 4.5c, d).
Collective cell migration plays a crucial role in many biological processes, including embryonic development, wound healing, as well as cancer invasion and metastasis [35, 36]. During collectively cell migration, the enhanced migration is led by a subset of “leader cells” that extend filopodia at the leading edge of the cell cluster [105]. The “invasion-competent” malignant cells induced the collective invasion of otherwise “invasion incompetent” epithelial cells, and that these two cell types consistently exhibited distinct leader and follower roles during invasion. Analysis of extracellular matrix (ECM) microarchitecture revealed that malignant cell invasion was accompanied by extensive ECM remodeling including matrix alignment and proteolytic track making [15].
Physical characteristics of ECM strongly modulate cell migration by outside-in signaling from microenvironment, while morphological properties of cell and intracellular dynamics feedback to ECM by inside-out signaling [91]. Current knowledge about the focal adhesion, cell migration, mechano-signaling, and cytoskeletal function is derived primarily from studies on planar 2D tissue culture substrates. The 2D substrate may induce artificial polarity between the basal and apical surfaces of the normally nonpolar cells, e.g., fibroblastic cells [24]. It also may exclude ECM-dependent regulators of 3D cell migration, including ECM porosity, ECM compliance, collagen fiber size, and collagen concentration [114]. The microarchitecture of 3D scaffolds has been found to influence cell migration behavior via junction interactions [54]. The pore size of collagen-glycosaminoglycan scaffolds influences the fibroblast migration: the migration speed decreases as pore size increases across a range from 90 to 150 μm [84]. Importantly, ECM density, stiffness and alignment also contributes to cell migration speed and persistence differently; ECM density and stiffness influences cell speed, but ECM alignment does not change cell speed; instead, alignment increases cell migration persistence[93].
Signaling, cytoskeletal dynamics, and cell shape. (a) The interaction of pushing microtubules (\textit{red arrows}) with the surface tension (blue arrows) of the plasma membrane (left) resembles the balance of gravitational pull (red arrow) and alcohol-dependent surface tension (blue arrows) along the edge of a wine-covered glass (right). (b) Activator-inhibitor system of an autocatalytically activated RTK. The high curvature at the tip of a protrusion facilitates initial RTK activation by effectively exposing the receptors to more extracellular space. The faster diffusing phosphatase limits spreading of autocatalytic activation by lateral inhibition. (c) (Left) The LEGI-BEN model: in the local excitation global inhibition (LEGI) model, a stimulus (S) turns on excitation (E) and inhibition (I) processes that act in parallel on a response regulator RR, which activates the biased excitable network (BEN), consisting of autocatalytic activity (X) that activates its own inhibitor (Y). (Right) Activity of X at different times after initial exposure to an extracellular chemotactic gradient (From Schmick and Bastiaens [97] with permission). (d) Interactions among the components of signaling pathways involved in the MAT/AMT transitions of cells in a 3D environment. The inhibition of the activity of the proteins highlighted in red was shown to trigger amoeboid to mesenchymal transitions. Inactivation of the proteins depicted in green induces a conversion from the mesenchymal to the amoeboid mode of invasiveness (From Pankova et al. [85] with permission)
Cell shape and membrane dynamics during migration. (a) Kymographs of an HT1080 cell on tissue culture-treated dishes showing membrane protrusion dynamics (From Sung et al. [102] with permission). (b) Phase contrast images of keratocytes crawling at low (left), intermediate (center), and high (right) adhesion strengths (From Barnhart et al. [6] with permission). (c) Simulations of membrane dynamics for keratocyte migration: membrane flow, velocity and tension (From Fogelson and Mogilner [33] with permission). (d) Steady-state maps of actin flow and substrate stress for keratocyte migration (From Shao et al. [99] with permission)
4.4 2D Cell Migration Models
Computational and mathematical modeling has benefited from the availability of new data with combination of live-cell imaging, molecular manipulation and force measurement. To date, most models in cell-ECM interactions focus on cell shape and cell motility. These models have treated implicit or explicit focal-adhesion, motility related diffusion-reaction of molecules, cytoskeletal dynamics, intracellular flows, and cell morphology related protrusion and contraction.
A rule-based model was developed for cell migration, in which the underlying mechano-chemical events are incorporated implicitly using rules describing the evolution of cell shapes and regulatory signals [96]. The main rules are about the local/global feedbacks and deterministic/stochastic signaling regulations. A cell is modeled using a collection of perimeter points and a center. The perimeter points can move according to the balance between protrusion signal and retraction signal. The local protrusion signal propagates and decays, with a stochastic positive feedback loop that accounts for both “local stimulation” and generation of random noises. Focal adhesion is a probabilistic event with a fixed average halftime. This simple model was capable of generating the dynamic shapes and persistence of amoeboid cells migration without the chemo-attractants.
Using the keratocyte migration as a model, a whole series of mathematical models explained the keratocyte cell shape [5, 6, 32, 94, 99]. Actin fibers polymerize pushing on the cell membrane from within, generating membrane tension that rapidly equilibrates. The membrane tension in turn exerts a constant force on the actin network. The spatiotemporal dynamics of adhesion-dependent actin polymerization, retrograde flow, myosin distribution, and traction forces together provide a more complete understanding of distribution of cell motility molecules and cell shape [33, 56] (Fig. 4.5c).
As the cell morphology adapts to the local forces from focal adhesion, actin flow, and myosin activities, the macromolecular distribution inside the cell presents a moving boundary reaction-diffusion problem for modeling. Because of the computational complexity, few models have integrated or implemented this problem. A recent model used the phase-field method to integrate the adhesion dynamics with the dynamics of the actin filaments modeled as a viscous network, and to solve for the moving boundary with membrane tension. The model included a reaction-diffusion model for the actin-myosin machinery and discrete adhesion sites that can be in a “gripping” or “slipping” mode. This model suggested the pattern of the actin flow inside the cell, the cell velocity, and the cell morphology are determined by the integration of actin polymerization, myosin contraction, adhesion forces, and membrane forces (Fig. 4.5d) [99].
The interaction between migrating cells and the ECM has also become a focal point of modeling in the past decade. In the context of angiogenesis, Bauer et al. developed a 2D model based on the cellular Potts model to study the effects of ECM topography on the collective migration morphology of endothelial cells [7]. They varied the density and alignment of the matrix fibers to simulate different tissue environments and to explore the possibility of manipulating the extracellular matrix to achieve pro- and anti-angiogenic effects. The ECM in this model only provided contact guidance, without mechanical interactions with the cells. Van Oers et al. coupled a 2D cellular Potts model with a finite element model for the ECM substrate, to simulate the mechanical interaction between cells and the ECM [107]. They showed that the resulting matrix strain could in turn mediate the interaction between cells and promote collective migration (Fig. 4.6a). The effect of ECM geometry on cell migration mode determination was studied by Tozluoglu et al. [104], using a 2D hybrid agent-based/finite element model of cell blebbing migration. The model integrated actin-polymerization-based protrusion, actomyosin contractility, and membrane blebbing due to actin-plasma membrane linkage, cell-ECM adhesion and varied matrix geometries (Fig. 4.6d–f) [104]. Actomyosin cortex and cell membrane were agents, with local levels of actin cortex density, myosin concentration, cortex-membrane linker proteins recorded at each agent. The model predicted the optimal migration strategies with different matrix geometries.
Simulated cell-ECM interaction. (a) Traction forces (black arrow) and resulting matrix strains (blue line segments) generated in the hybrid cellular Potts and finite element model (From van Oers et al. [107] with permission). Cell invasion into ECM fiber network with pore sizes of (b) 0.5 and (c) 1.5 \( \mu m \) (From Kim et al. [67] with permission). Simulations of cell moving through different matrix geometries show different optimal migration strategies. (d) Cell crawling on a surface. (e) Actin-protrusion-based solution within confined continuous environments. (f) Blebbing-driven solution for cells with 50 % more overall contractility (From Tozluoglu [104] with permission)
4.5 3D Cell-ECM Model
Most cells encounter a 3D matrix environment during processes such as wound healing or cancer metastasis. Increasing evidence from literature suggests that 2D ECM models are inherently limited in their scope to capture the ability of cells to form adhesions in three dimensions. Therefore, it is important that we use 3D systems to study cell-matrix interactions to gain more physiologically-relevant insights. The 3D matrix structure, focal adhesion, cell signaling, and cell morphology are more complex. But with the advance of imaging tools, such as multi-photon microscopy for imaging the ECM, and lattice light-sheet microscopy to image both cell and ECM with very high spatial and temporal resolutions [17], the hope is high for a more complete understanding of 3D cell-ECM interactions in the near future.
A phenomenological 3D model of single cell migration through cell-ECM interaction is developed taking into account the ECM deposition, cell protrusion, adhesion detachment and MMP activities [18] [53]. In this model, cells can degrade, deposit, or pull local fibers, depending on the fiber density around each cell. The cells can also move within the 3D matrix. The model produced results consistent with the current understanding: in low density environments, cells deposit more collagen to increase fibril fraction; in higher density environments, the less invasive cell line reduced the fibril fraction as compared to the highly invasive phenotype. Riching et al. showed another 3D cell-ECM interaction model (in the supporting materials in [93]). In this model, the cells migrate in 2D but interact with a 3D ECM environment they are embeded in. The cells send out protrusion vectors around their perimeters, the magnitudes of these protrusion vectors are determined by its interaction with the local matrix stiffness, alignment and ligand density. Mechanics was only considered implicitly through the coefficient of matrix rigidity. The simple cell-ECM model was able to qualitatively agree with experiments in concluding that matrix alignment does not change cell speed but increases cell migration persistence [93].
Borau et al. [8] developed a probabilistic, cell voxel-based finite element for 3D cell-ECM interactions in a microfluidic environment. A cell is a collection of voxels, where stress, chemical concentration and fluid flow surrounding the cell drives cell migration by adding and removing voxels. Cell contains cortex, cytoplasm and nucleus. The nucleus is an elastic material that only plays a passive role during cell migration. The cortex and cytoplasm contractility depends on the mechanosensing of ECM stiffness, which is modeled implicitly. It provides a methodology for testing and designing experiments in microfluidic systems.
Kim et al. [67] reported a more biomechanically realistic cell-ECM interaction model, which accounted for intracellular mechanics of cellular and nuclear membranes, contractile actin stress fibers, focal adhesion dynamics, structural mechanics of ECM fiber networks, and reaction-diffusion mass transfers of seven biochemical concentrations associated with chemotaxis, proteolysis, haptotaxis, and degradation in ECM. Simulations of cell invasion into fiber networks with various pore sizes, such as 0.5 μm pore size (Fig. 4.6b) and 1.5 μm pore size (Fig. 4.6c), show that filopodia invaded more deeply in the large pore ECM fibers [67]. The results were successfully compared with experiments of 3D HUVEC migration for ECMs with different pore sizes and stiffness.
4.6 Modeling Collective Behavior of Cell Migration
Comparing to single cell modeling, less effort has been directed towards understanding how clusters of cells migrate collectively through microenvironments. A few models have been developed to explore how the cell-cell and cell-ECM interactions influence collective behavior of migrating cells. Models at this scale are usually complicated, but computationally less expensive than single cell level, because most cell details have been coarse-grained.
Most multi-cellular models contain three parts: single agent with simple properties to represent a cell, cell-cell adhesion, and heterogeneous environment. Guven et al. [51] developed a coarse-grained stochastic model of Dictyostelium cells using 2D self-propelled soft disks to study the influence of signal relay. Wynn et al. [117] developed an agent-based cell model that treats point-like cells with biased migration directionality and cell-ECM interactions, and modeled the leader-follower dynamic patterns of collective migration in neural crest cells. In this model the ECM is a passive substrate that can be degraded by cells to form tracks of less resistance. Zaritsky et al. [120] proposed a new analytical framework to explicitly detect and quantify cell clusters that move coordinately in a monolayer, and reported the finding of waves of coordinated migration in wound healing experiments. They explained the wave by Met activation with hepatocyte growth factor or scatter factor. The data and model suggested that collective migration emerges from spatial and temporal accumulation and directionality, which can be a basic cellular mechanism for long-term cell guidance during collective cell migration.
4.7 Summary
Cancer cell invasion into ECM is the first step of metastasis, the main difficulty in treating cancer. Biomechanical experiments and simulations of the ECM, cell, and interactions between the cell and ECM are necessary to better understand the invasion behavior of cancer cells. Recent technologies in microscopy, biomechanical rheology, image processing, and 2D and 3D computational modeling shed light on cancer invasion. We highlighted recent studies on tumor microenvironment, especially ECM, cell, and their interaction.
The importance of the mechanics of ECM and cell-ECM interactions in regulating and contributing to cancer invasion has been increasingly accepted. Moreover, it is necessary to combine these understandings into a unified framework of cancer invasion. The biophysical and biomechanical aspects of the microenvironment should be integrated with the biological and the biochemical aspects, to form a comprehensive description of the tumor microenvironment. The integrated understanding of the cell, ECM, and their interactions is required to better predict cancer invasion and possibly develop new tools to prevent or stop cancer. The strong interplay between cancer cell biology and the mechanical microenvironment suggests new possibilities of regulation and manipulation of cell behavior to alter the outcome of cancer.
References
Abhilash AS, Baker BM, Trappmann B, Chen CS, Shenoy VB (2014) Remodeling of fibrous extracellular matrices by contractile cells: predictions from discrete fiber network simulations. Biophys J 107(8):1829–1840. doi:10.1016/j.bpj.2014.08.029
Akiri G, Sabo E, Dafni H, Vadasz Z, Kartvelishvily Y, Gan N, Kessler O, Cohen T, Resnick M, Neeman M, Neufeld G (2003) Lysyl oxidase-related protein-1 promotes tumor fibrosis and tumor progression in vivo. Cancer Res 63(7):1657–1666
Alexander S, Weigelin B, Winkler F, Friedl P (2013) Preclinical intravital microscopy of the tumour-stroma interface: invasion, metastasis, and therapy response. Curr Opin Cell Biol 25(5):659–671. doi:10.1016/j.ceb.2013.07.001
Balaban NQ, Schwarz US, Riveline D, Goichberg P, Tzur G, Sabanay I, Mahalu D, Safran S, Bershadsky A, Addadi L, Geiger B (2001) Force and focal adhesion assembly: a close relationship studied using elastic micropatterned substrates. Nat Cell Biol 3(5):466–472
Barnhart E, Lee K-C, Allen GM, Theriot JA, Mogilner A (2015) Balance between cell−substrate adhesion and myosin contraction determines the frequency of motility initiation in fish keratocytes. Proc Natl Acad Sci 112(16):5045–5050
Barnhart EL, Lee KC, Keren K, Mogilner A, Theriot JA (2011) An adhesion-dependent switch between mechanisms that determine motile cell shape. PLoS Biol 9(5):e1001059. doi:10.1371/journal.pbio.1001059
Bauer AL, Jackson TL, Jiang Y (2009) Topography of extracellular matrix mediates vascular morphogenesis and migration speeds in angiogenesis. PLoS Comput Biol 5(7):e1000445. doi:10.1371/journal.pcbi.1000445
Borau C, Polacheck WJ, Kamm RD, García-Aznar JM (2014) Probabilistic Voxel-Fe model for single cell motility in 3D. In Silico Cell Tissue Sci 1(1):2
Bordeleau F, Ta A, Ca R-K (2014) Physical biology in cancer. 5. The rocky road of metastasis: the role of cytoskeletal mechanics in cell migratory response to 3D matrix topography. Am Physiol Cell Physiol 306(2):C110–C120
Bordeleau F, Bessard J, Marceau N, Sheng Y (2011) Measuring integrated cellular mechanical stress response at focal adhesions by optical tweezers. J Biomed Opt 16(9):095005
Bordeleau F, Bessard J, Sheng Y, Marceau N (2008) Keratin contribution to cellular mechanical stress response at focal adhesions as assayed by laser tweezers. Biochem Cell Biol 86(4):352–359
Boucher Y, Salehi H, Witwer B, Harsh GR, Jain RK (1997) Interstitial fluid pressure in intracranial tumours in patients and in rodents. Br J Cancer 75(6):829–836
Boyd NF, Guo H, Martin LJ, Sun L, Stone J, Fishell E, Jong RA, Hislop G, Chiarelli A, Minkin S, Yaffe MJ (2007) Mammographic density and the risk and detection of breast cancer. N Engl J Med 356(3):227–236. doi:10.1056/NEJMoa062790
Byron A, Ja A, Humphries JD, Jacquemet G, Koper EJ, Warwood S, Choi CK, Stroud MJ, Chen CS, Knight D, Humphries MJ (2015) A proteomic approach reveals integrin activation state-dependent control of microtubule cortical targeting. Nat Commun 6(6135):1–14
Carey SP, Starchenko A, McGregor AL, Reinhart-King CA (2013) Leading malignant cells initiate collective epithelial cell invasion in a three-dimensional heterotypic tumor spheroid model. Clin Exp Metastasis 30(5):615–630
Ea C-A, Micoulet A, Blümmel J, Auernheimer J, Kessler H, Spatz JP (2006) Lateral spacing of integrin ligands influences cell spreading and focal adhesion assembly. Eur J Cell Biol 85(3-4):219–224
Chen B-C, Legant WR, Wang K, Shao L, Milkie DE, Davidson MW, Janetopoulos C, Wu XS, Hammer JA, Liu Z, English BP, Mimori-Kiyosue Y, Romero DP, Ritter AT, Lippincott-Schwartz J, Fritz-Laylin L, Mullins RD, Mitchell DM, Bembenek JN, Reymann A-C, Bohme R, Grill SW, Wang JT, Seydoux G, Tulu US, Kiehart DP, Betzig E (2014) Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346(6208):1257998
Chisholm RH, Hughes BD, Landman KA, Zaman MH (2013) Analytic study of three-dimensional single cell migration with and without proteolytic enzymes. Cell Mol Bioeng 6(2):239–249. doi:10.1007/s12195-012-0261-8
Choquet D, Felsenfeld DP, Sheetz MP (1997) Extracellular matrix rigidity causes strengthening of integrin-cytoskeleton linkages. Cell 88(1):39–48
Chung BM, Rotty JD, Pa C (2013) Networking galore: intermediate filaments and cell migration. Curr Opin Cell Biol 25(5):600–612
Condeelis J, Segall JE (2003) Intravital imaging of cell movement in tumours. Nat Rev Cancer 3(12):921–930. doi:10.1038/nrc1231
Cook JA, Gius D, Wink DA, Krishna MC, Russo A, Mitchell JB (2004) Oxidative stress, redox, and the tumor microenvironment. Semin Radiat Oncol 14(3):259–266
Coyer SR, Singh A, Dumbauld DW, Calderwood DA, Craig SW, Delamarche E, Garcia AJ (2012) Nanopatterning reveals an ECM area threshold for focal adhesion assembly and force transmission that is regulated by integrin activation and cytoskeleton tension. J Cell Sci 125(21):5110–5123. doi:10.1242/jcs.108035
Cukierman E, Pankov R, Stevens DR, Yamada KM (2001) Taking cell-matrix adhesions to the third dimension. Science 294(5547):1708–1712. doi:10.1126/science.1064829
Dahl KN, Engler AJ, Pajerowski JD, Discher DE (2005) Power-law rheology of isolated nuclei with deformation mapping of nuclear substructures. Biophys J 89(4):2855–2864. doi:10.1529/biophysj.105.062554
Dahl KN, Ribeiro AJ, Lammerding J (2008) Nuclear shape, mechanics, and mechanotransduction. Circ Res 102(11):1307–1318. doi:10.1161/CIRCRESAHA.108.173989
Davidson PM, Sliz J, Isermann P, Denais C, Lammerding J (2015) Design of a microfluidic device to quantify dynamic intra-nuclear deformation during cell migration through confining environments. Integr Biol 7(12):1534–1546. doi:10.1039/c5ib00200a
Dembo M, Torney DC, Saxman K, Hammer D (1988) The reaction-limited kinetics of membrane-to-surface adhesion and detachment. Proc R Soc Lond B Biol Sci 234(1274):55–83
Diaz-Cano SJ (2012) Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci 13(2):1951–2011. doi:10.3390/ijms13021951
Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, Lafontaine I, De Montigny J, Marck C, Neuveglise C, Talla E, Goffard N, Frangeul L, Aigle M, Anthouard V, Babour A, Barbe V, Barnay S, Blanchin S, Beckerich JM, Beyne E, Bleykasten C, Boisrame A, Boyer J, Cattolico L, Confanioleri F, De Daruvar A, Despons L, Fabre E, Fairhead C, Ferry-Dumazet H, Groppi A, Hantraye F, Hennequin C, Jauniaux N, Joyet P, Kachouri R, Kerrest A, Koszul R, Lemaire M, Lesur I, Ma L, Muller H, Nicaud JM, Nikolski M, Oztas S, Ozier-Kalogeropoulos O, Pellenz S, Potier S, Richard GF, Straub ML, Suleau A, Swennen D, Tekaia F, Wesolowski-Louvel M, Westhof E, Wirth B, Zeniou-Meyer M, Zivanovic I, Bolotin-Fukuhara M, Thierry A, Bouchier C, Caudron B, Scarpelli C, Gaillardin C, Weissenbach J, Wincker P, Souciet JL (2004) Genome evolution in yeasts. Nature 430(6995):35–44. doi:10.1038/nature02579
Even-Ram S, Yamada KM (2005) Cell migration in 3D matrix. Curr Opin Cell Biol 17(5):524–532. doi:10.1016/j.ceb.2005.08.015
Feng J, Levine H, Mao X, Sander LM (2015) Alignment and nonlinear elasticity in biopolymer gels. Phys Rev E Stat Nonlin Soft Matter Phys 91(4):042710. doi:10.1103/PhysRevE.91.042710
Fogelson B, Mogilner A (2014) Computational estimates of membrane flow and tension gradient in motile cells. PLoS One 9(1):e84524. doi:10.1371/journal.pone.0084524
Friedl P (2004) Prespecification and plasticity: shifting mechanisms of cell migration. Curr Opin Cell Biol 16(1):14–23. doi:10.1016/j.ceb.2003.11.001
Friedl P, Gilmour D (2009) Collective cell migration in morphogenesis, regeneration and cancer. Nat Rev Mol Cell Biol 10(7):445–457. doi:10.1038/nrm2720
Friedl P, Hegerfeldt Y, Tusch M (2004) Collective cell migration in morphogenesis and cancer. Int J Dev Biol 48(5-6):441–449
Friedl P, Locker J, Sahai E, Segall JE (2012) Classifying collective cancer cell invasion. Nat Cell Biol 14(8):777–783. doi:10.1038/ncb2548
Friedl P, Wolf K (2010) Plasticity of cell migration: a multiscale tuning model. J Cell Biol 188(1):11–19
Friedland JC, Lee MH, Boettiger D (2009) Mechanically activated integrin switch controls α5β1 function. Science (New York) 323(5914):642–644
Fu Y, Chin LK, Bourouina T, Liu AQ, VanDongen AM (2012) Nuclear deformation during breast cancer cell transmigration. Lab Chip 12(19):3774–3778. doi:10.1039/c2lc40477j
Gabella C, Bertseva E, Bottier C, Piacentini N, Bornert A, Jeney S, Forro L, Sbalzarini IF, Meister JJ, Verkhovsky AB (2014) Contact angle at the leading edge controls cell protrusion rate. Curr Biol 24(10):1126–1132
Galbraith CG, Yamada KM, Sheetz MP (2002) The relationship between force and focal complex development. J Cell Biol 159(4):695–705. doi:10.1083/jcb.200204153
Gallant ND, García AJ (2005) Cell adhesion strengthening and focal adhesion assembly on micropatterned substrates. Mol Biol Cell 16(9):4329–4340
Gatenby RA, Gawlinski ET (2003) The glycolytic phenotype in carcinogenesis and tumor invasion: insights through mathematical models. Cancer Res 63(14):3847–3854
Geiger B, Bershadsky A (2002) Exploring the neighborhood: adhesion-coupled cell mechanosensors. Cell 110(2):139–142
Geiger B, Bershadsky A, Pankov R, Yamada KM (2001) Transmembrane crosstalk between the extracellular matrix–cytoskeleton crosstalk. Nat Rev Mol Cell Biol 2(11):793–805. doi:10.1038/35099066
Geiger B, Spatz JP, Bershadsky AD (2009) Environmental sensing through focal adhesions. Nat Rev Mol Cell Biol 10(1):21–33. doi:10.1038/nrm2593
Geiger B, Yamada KM (2011) Molecular architecture and function of matrix adhesions. Cold Spring Harb Perspect Biol 3(5):1–21. doi:10.1101/cshperspect.a005033
Grashoff C, Hoffman BD, Brenner MD, Zhou R, Parsons M, Yang MT, McLean MA, Sligar SG, Chen CS, Ha T, Schwartz MA (2010) Measuring mechanical tension across vinculin reveals regulation of focal adhesion dynamics. Nature 466(7303):263–266. doi:10.1038/nature09198
Guilak F, Tedrow JR, Burgkart R (2000) Viscoelastic properties of the cell nucleus. Biochem Biophys Res Commun 269(3):781–786. doi:10.1006/bbrc.2000.2360
Guven C, Rericha E, Ott E, Losert W (2013) Modeling and measuring signal relay in noisy directed migration of cell groups. PLoS Comput Biol 9(5):e1003041. doi:10.1371/journal.pcbi.1003041
Hamadi A, Bouali M, Dontenwill M, Stoeckel H, Takeda K, Ronde P (2005) Regulation of focal adhesion dynamics and disassembly by phosphorylation of FAK at tyrosine 397. J Cell Sci 118(19):4415–4425. doi:10.1242/jcs.02565
Harjanto D, Zaman MH (2013) Modeling extracellular matrix reorganization in 3D environments. PLoS One 8(1):e52509. doi:10.1371/journal.pone.0052509
Harley BA, Kim HD, Zaman MH, Yannas IV, Lauffenburger DA, Gibson LJ (2008) Microarchitecture of three-dimensional scaffolds influences cell migration behavior via junction interactions. Biophys J 95(8):4013–4024. doi:10.1529/biophysj.107.122598
Hockel M, Vaupel P (2001) Tumor hypoxia: definitions and current clinical, biologic, and molecular aspects. J Natl Cancer Inst 93(4):266–276
Holmes WR, Edelstein-Keshet L (2012) A comparison of computational models for eukaryotic cell shape and motility. PLoS Comput Biol 8(12):e1002793. doi:10.1371/journal.pcbi.1002793
Huveneers S, Danen EH (2009) Adhesion signaling – crosstalk between integrins, Src and Rho. J Cell Sci 122(Pt 8):1059–1069. doi:10.1242/jcs.039446
Icard-Arcizet D, Cardoso O, Richert A, Henon S (2008) Cell stiffening in response to external stress is correlated to actin recruitment. Biophys J 94(7):2906–2913. doi:10.1529/biophysj.107.118265
Ingber DE (2003) Tensegrity I. Cell structure and hierarchical systems biology. J Cell Sci 116(Pt 7):1157–1173
Jiang GY, Giannone G, Critchley DR, Fukumoto E, Sheetz MP (2003) Two-piconewton slip bond between fibronectin and the cytoskeleton depends on talin. Nature 424(6946):334–337. doi:10.1038/nature01805
Kaltenbrunner M, White MS, Glowacki ED, Sekitani T, Someya T, Sariciftci NS, Bauer S (2012) Ultrathin and lightweight organic solar cells with high flexibility. Nat Commun 3:770. doi:10.1038/ncomms1772
Kaverina I, Rottner K, Small JV (1998) Targeting, capture, and stabilization of microtubules at early focal adhesions. J Cell Biol 142(1):181–190. doi:10.1083/jcb.142.1.181
Kaverina I, Straube A (2011) Regulation of cell migration by dynamic microtubules. Semin Cell Dev Biol 22(9):968–974. doi:10.1016/j.semcdb.2011.09.017
Kaverina I, Straube A (2011) Regulation of cell migration by dynamic microtubules. Semin Cell Dev Biol 22(9):968–974
Keren K, Pincus Z, Allen GM, Barnhart EL, Marriott G, Mogilner A, Theriot JA (2008) Mechanism of shape determination in motile cells. Nature 453(7194):475–480. doi:10.1038/nature06952
Kim DH, Han K, Gupta K, Kwon KW, Suh KY, Levchenko A (2009) Mechanosensitivity of fibroblast cell shape and movement to anisotropic substratum topography gradients. Biomaterials 30(29):5433–5444. doi:10.1016/j.biomaterials.2009.06.042
Kim MC, Whisler J, Silberberg YR, Kamm RD, Asada HH (2015) Cell invasion dynamics into a three dimensional extracellular matrix fibre network. PLoS Comput Biol 11(10):e1004535. doi:10.1371/journal.pcbi.1004535
Kong F, Li Z, Parks WM, Dumbauld DW, Garcia AJ, Mould AP, Humphries MJ, Zhu C (2013) Cyclic mechanical reinforcement of integrin-ligand interactions. Mol Cell 49(6):1060–1068. doi:10.1016/j.molcel.2013.01.015
Krishnan J, Iglesias PA (2003) Analysis of the signal transduction properties of a module of spatial sensing in eukaryotic chemotaxis. Bull Math Biol 65(1):95–128. doi:10.1006/bulm.2002.0323
Kuo JC (2013) Mechanotransduction at focal adhesions: integrating cytoskeletal mechanics in migrating cells. J Cell Mol Med 17(6):704–712. doi:10.1111/jcmm.12054
Lee B, Zhou X, Riching K, Eliceiri KW, Keely PJ, Guelcher SA, Weaver AM, Jiang Y (2014) A three-dimensional computational model of collagen network mechanics. PLoS One 9(11):e111896. doi:10.1371/journal.pone.0111896
Legate KR, Wickstrom SA, Fassler R (2009) Genetic and cell biological analysis of integrin outside-in signaling. Genes Dev 23(4):397–418. doi:10.1101/gad.1758709
Levchenko A, Iglesias PA (2002) Models of eukaryotic gradient sensing: application to chemotaxis of amoebae and neutrophils. Biophys J 82(1 Pt 1):50–63. doi:10.1016/S0006-3495(02)75373-3
Levental KR, Yu H, Kass L, Lakins JN, Egeblad M, Erler JT, Fong SF, Csiszar K, Giaccia A, Weninger W, Yamauchi M, Gasser DL, Weaver VM (2009) Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139(5):891–906. doi:10.1016/j.cell.2009.10.027
Lin CQ, Bissell MJ (1993) Multi-faceted regulation of cell differentiation by extracellular matrix. FASEB J 7(9):737–743
Liu YJ, Le Berre M, Lautenschlaeger F, Maiuri P, Callan-Jones A, Heuze M, Takaki T, Voituriez R, Piel M (2015) Confinement and low adhesion induce fast amoeboid migration of slow mesenchymal cells. Cell 160(4):659–672. doi:10.1016/j.cell.2015.01.007
Lombardi ML, Lammerding J (2011) Keeping the LINC: the importance of nucleocytoskeletal coupling in intracellular force transmission and cellular function. Biochem Soc Trans 39(6):1729–1734. doi:10.1042/BST20110686
Lu P, Takai K, Weaver VM, Werb Z (2011) Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb Perspect Biol 3(12):a005058. doi:10.1101/cshperspect.a005058
Lu P, Weaver VM, Werb Z (2012) The extracellular matrix: a dynamic niche in cancer progression. J Cell Biol 196(4):395–406. doi:10.1083/jcb.201102147
McCormack VA, dos Santos SI (2006) Breast density and parenchymal patterns as markers of breast cancer risk: a meta-analysis. Cancer Epidemiol Biomark Prev 15(6):1159–1169. doi:10.1158/1055-9965.EPI-06-0034
Michael KE, Dumbauld DW, Burns KL, Hanks SK, Garcia AJ (2009) Focal adhesion kinase modulates cell adhesion strengthening via integrin activation. Mol Biol Cell 20(9):2508–2519. doi:10.1091/mbc.E08-01-0076
Mrksich M, Chen CS, Xia Y, Dike LE, Ingber DE, Whitesides GM (1996) Controlling cell attachment on contoured surfaces with self-assembled monolayers of alkanethiolates on gold. Proc Natl Acad Sci U S A 93(20):10775–10778
Oakes PW, Beckham Y, Stricker J, Gardel ML (2012) Tension is required but not sufficient for focal adhesion maturation without a stress fiber template. Dev Cell 196:3
Oryan A, Moshiri A, Sharifi P (2012) Advances in injured tendon engineering with emphasis on the role of collagen implants. Hard Tissue 1(2):12
Panková K, Rösel D, Novotný M, Brábek J (2010) The molecular mechanisms of transition between mesenchymal and amoeboid invasiveness in tumor cells. Cell Mol Life Sci: CMLS 67(1):63–71
Parent CA, Devreotes PN (1999) A cell’s sense of direction. Science 284(5415):765–770. doi:10.1126/science.284.5415.765
Paszek MJ, Zahir N, Johnson KR, Lakins JN, Rozenberg GI, Gefen A, Reinhart-King CA, Margulies SS, Dembo M, Boettiger D, Hammer DA, Weaver VM (2005) Tensional homeostasis and the malignant phenotype. Cancer Cell 8(3):241–254. doi:10.1016/j.ccr.2005.08.010
Petrie RJ, Koo H, Yamada KM (2014) Generation of compartmentalized pressure by a nuclear piston governs cell motility in a 3D matrix. Science 345(6200):1062–1065. doi:10.1126/science.1256965
Provenzano PP, Eliceiri KW, Campbell JM, Inman DR, White JG, Keely PJ (2006) Collagen reorganization at the tumor-stromal interface facilitates local invasion. BMC Med 4(1):38. doi:10.1186/1741-7015-4-38
Provenzano PP, Inman DR, Eliceiri KW, Knittel JG, Yan L, Rueden CT, White JG, Keely PJ (2008) Collagen density promotes mammary tumor initiation and progression. BMC Med 6:11. doi:10.1186/1741-7015-6-11
Provenzano PP, Keely PJ (2011) Mechanical signaling through the cytoskeleton regulates cell proliferation by coordinated focal adhesion and Rho GTPase signaling. J Cell Sci 124(8):1195–1205. doi:10.1242/jcs.067009
Puklin-Faucher E, Sheetz MP (2009) The mechanical integrin cycle. J Cell Sci 122(Pt 2):179–186. doi:10.1242/jcs.042127
Riching KM, Cox BL, Salick MR, Pehlke C, Riching AS, Ponik SM, Bass BR, Crone WC, Jiang Y, Weaver AM, Eliceiri KW, Keely PJ (2014) 3D collagen alignment limits protrusions to enhance breast cancer cell persistence. Biophys J 107(11):2546–2558. doi:10.1016/j.bpj.2014.10.035
Rosso F, Giordano A, Barbarisi M, Barbarisi A (2004) From cell-ECM interactions to tissue engineering. J Cell Physiol 199(2):174–180. doi:10.1002/jcp.10471
Ruprecht V, Wieser S, Callan-Jones A, Smutny M, Morita H, Sako K, Barone V, Ritsch-Marte M, Sixt M, Voituriez R, Heisenberg CP (2015) Cortical contractility triggers a stochastic switch to fast amoeboid cell motility. Cell 160(4):673–685. doi:10.1016/j.cell.2015.01.008
Satulovsky J, Lui R, Wang YL (2008) Exploring the control circuit of cell migration by mathematical modeling. Biophys J 94(9):3671–3683. doi:10.1529/biophysj.107.117002
Schmick M, Bastiaens PI (2014) The interdependence of membrane shape and cellular signal processing. Cell 156(6):1132–1138. doi:10.1016/j.cell.2014.02.007
Serrels B, Serrels A, Brunton VG, Holt M, McLean GW, Gray CH, Jones GE, Frame MC (2007) Focal adhesion kinase controls actin assembly via a FERM-mediated interaction with the Arp2/3 complex. Nat Cell Biol 9(9):1046–1056. doi:10.1038/ncb1626
Shao D, Levine H, Rappel WJ (2012) Coupling actin flow, adhesion, and morphology in a computational cell motility model. Proc Natl Acad Sci U S A 109(18):6851–6856. doi:10.1073/pnas.1203252109
Spaeth EL, Dembinski JL, Sasser AK, Watson K, Klopp A, Hall B, Andreeff M, Marini F (2009) Mesenchymal stem cell transition to tumor-associated fibroblasts contributes to fibrovascular network expansion and tumor progression. PLoS One 4(4):e4992. doi:10.1371/journal.pone.0004992
Stamenović D (2005) Microtubules may harden or soften cells, depending of the extent of cell distension. J Biomech 38(8):1728–1732
Sung BH, Ketova T, Hoshino D, Zijlstra A, Weaver AM (2015) Directional cell movement through tissues is controlled by exosome secretion. Nat Commun 13(6):2546–2558
Takahashi R, Nagayama S, Furu M, Kajita Y, Jin Y, Kato T, Imoto S, Sakai Y, Toguchida J (2014) AFAP1L1, a novel associating partner with vinculin, modulates cellular morphology and motility, and promotes the progression of colorectal cancers. Cancer Med 3(4):759–774. doi:10.1002/cam4.237
Tozluoglu M, Tournier AL, Jenkins RP, Hooper S, Bates PA, Sahai E (2013) Matrix geometry determines optimal cancer cell migration strategy and modulates response to interventions. Nat Cell Biol 15(7):751–762. doi:10.1038/ncb2775
Tse JM, Cheng G, Tyrrell JA, Wilcox-Adelman SA, Boucher Y, Jain RK, Munn LL (2012) Mechanical compression drives cancer cells toward invasive phenotype. Proc Natl Acad Sci U S A 109(3):911–916. doi:10.1073/pnas.1118910109
Vader D, Kabla A, Weitz D, Mahadevan L (2009) Strain-induced alignment in collagen gels. PLoS One 4(6):e5902. doi:10.1371/journal.pone.0005902
van Oers RF, Rens EG, LaValley DJ, Reinhart-King CA, Merks RM (2014) Mechanical cell-matrix feedback explains pairwise and collective endothelial cell behavior in vitro. PLoS Comput Biol 10(8):e1003774. doi:10.1371/journal.pcbi.1003774
Vicente-Manzanares M, Horwitz A (2011) Cell migration: an overview. Springer, Berlin
Wehrle-Haller B (2012) Structure and function of focal adhesions. Curr Opin Cell Biol 24(1):116–124. doi:10.1016/j.ceb.2011.11.001
Weidner N, Folkman J, Pozza F, Bevilacqua P, Allred EN, Moore DH, Meli S, Gasparini G (1992) Tumor angiogenesis: a new significant and independent prognostic indicator in early-stage breast carcinoma. J Natl Cancer Inst 84(24): 1875–1887
Welch MD (2015) Cell migration, freshly squeezed. Cell 160(4):581–582. doi:10.1016/j.cell.2015.01.053
Whiteside TL (2008) The tumor microenvironment and its role in promoting tumor growth. Oncogene 27(45):5904–5912. doi:10.1038/onc.2008.271
Wolf K, Mazo I, Leung H, Engelke K, von Andrian UH, Deryugina EI, Strongin AY, Brocker EB, Friedl P (2003) Compensation mechanism in tumor cell migration: mesenchymal-amoeboid transition after blocking of pericellular proteolysis. J Cell Biol 160(2):267–277. doi:10.1083/jcb.200209006
Wolf K, Te Lindert M, Krause M, Alexander S, TeRiet J, Willis AL, Hoffman RM, Figdor CG, Weiss SJ, Friedl P (2013) Physical limits of cell migration: control by ECM space and nuclear deformation and tuning by proteolysis and traction force. J Cell Biol 201(7):1069–1084. doi:10.1083/jcb.201210152
Wolf K, Wu YI, Liu Y, Geiger J, Tam E, Overall C, Stack MS, Friedl P (2007) Multi-step pericellular proteolysis controls the transition from individual to collective cancer cell invasion. Nat Cell Biol 9(8):893–904. doi:10.1038/ncb1616
Wolfenson H, Lavelin I, Geiger B (2013) Dynamic regulation of the structure and functions of integrin adhesions. Dev Cell 24(5):447–458. doi:10.1016/j.devcel.2013.02.012
Wynn ML, Rupp P, Trainor PA, Schnell S, Kulesa PM (2013) Follow-the-leader cell migration requires biased cell-cell contact and local microenvironmental signals. Phys Biol 10(3):035003. doi:10.1088/1478-3975/10/3/035003
Xiong Y, Huang CH, Iglesias PA, Devreotes PN (2010) Cells navigate with a local-excitation, global-inhibition-biased excitable network. Proc Natl Acad Sci U S A 107(40):17079–17086. doi:10.1073/pnas.1011271107
Yu CH, Law JBK, Suryana M, Low HY, Sheetz MP (2011) Early integrin binding to Arg-Gly-Asp peptide activates actin polymerization and contractile movement that stimulates outward translocation. Proc Natl Acad Sci U S A 108(51):20585–20590. doi:10.1073/pnas.1109485108
Zaritsky A, Kaplan D, Hecht I, Natan S, Wolf L, GovNS, Ben-Jacob E, Tsarfaty I (2014) Propagating waves of directionality and coordination orchestrate collective cell migration. PLoS Comput Biol 10(7):e1003747. doi:10.1371/journal.pcbi.1003747
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
He, X., Lee, B., Jiang, Y. (2016). Cell-ECM Interactions in Tumor Invasion. In: Rejniak, K. (eds) Systems Biology of Tumor Microenvironment. Advances in Experimental Medicine and Biology, vol 936. Springer, Cham. https://doi.org/10.1007/978-3-319-42023-3_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-42023-3_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-42021-9
Online ISBN: 978-3-319-42023-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)