Abstract
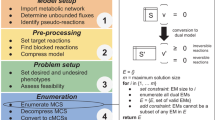
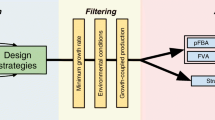
Under the realm of in silico Metabolic Engineering, pathway analysis approaches to strain optimization have shown a large potential as tools capable of providing an unbiased view over metabolic models. Most of these methods were difficult or impossible to use due to their heavy computational needs, since they are based in the calculation of elementary modes/minimal cut sets in large networks. However, a recent method (MCSEnumerator) has enabled the application of these approaches to genome-scale metabolic models. This work proposes a new software tool where this method is implemented in a novel Java library, that provides support for a plugin for the OptFlux metabolic engineering platform. Together, these tools implement the routines necessary for the calculation of minimal cut sets and their use to provide strain optimization methods. The aim is to provide an open-source software tool that includes an intuitive graphical user interface, thus facilitating its use by the community.
Access provided by Autonomous University of Puebla. Download to read the full chapter text
Chapter PDF
Similar content being viewed by others
Keywords
References
Stephanopoulos, G.: Metabolic Fluxes and Metabolic Engineering 11 (1999)
Patil, K.R., Åkesson, M., Nielsen, J.: Use of genome-scale microbial models for metabolic engineering. Curr. Opin. Biotechnol. 15(1), 64–69 (2004)
Szallasi, Z., Stelling, J., Periwal, V.: System Modeling in Cell Biology (2010)
Varma, A., Palsson, B.O., Arbor, A., Varma, A.: Stoichiometric Flux Balance Models Quantitatively Predict. Appl. Environ. Microbiol. 60(10), 3724–3731 (1994)
Segrè, D., Vitkup, D., Church, G.M.: Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. U. S. A. 99(23), 15112–15117 (2002)
Shlomi, T., Berkman, O., Ruppin, E.: Regulatory on/off minimization of metabolic flux changes after genetic perturbations. Proc. Natl. Acad. Sci. U. S. A. 102(21), 7695–7700 (2005)
Maia, P., Rocha, M., Rocha, I.: In silico constraint-based strain optimization methods: the quest for optimal cell factories. Microbiology and Molecular Biology Reviews 80(1), 45–67 (2016)
Burgard, A.P., Pharkya, P., Maranas, C.D.: Optknock: A bilevel programming framework for identifying gene knockout strategies for microbial strain optimization. Biotechnol. Bioeng. 84(6), 647–657 (2003)
Patil, K.R., Rocha, I., Förster, J., Nielsen, J.: Evolutionary programming as a platform for in silico metabolic engineering. BMC Bioinformatics 6(1), 308 (2005)
Klamt, S., Gilles, E.D.: Minimal cut sets in biochemical reaction networks. Bioinformatics 20(2), 226–234 (2004)
von Kamp, A., Klamt, S.: Enumeration of Smallest Intervention Strategies in Genome-Scale Metabolic Networks. PLoS Comput. Biol. 10(1), e1003378 (2014)
Schuster, S., Hilgetag, C.: On Elementary Flux Modes in Biochemical Reaction Systems At Steady State. J. Biol. Syst. 02(02), 165–182 (1994)
Hädicke, O., Klamt, S.: Computing complex metabolic intervention strategies using constrained minimal cut sets. Metab. Eng. 13(2), 204–213 (2011)
Ballerstein, K., von Kamp, A., Klamt, S., Haus, U.U.: Minimal cut sets in a metabolic network are elementary modes in a dual network. Bioinformatics 28(3), 381–387 (2012)
de Figueiredo, L.F., Podhorski, A., Rubio, A., Kaleta, C., Beasley, J.E., Schuster, S., Planes, F.J.: Computing the shortest elementary flux modes in genome-scale metabolic networks. Bioinformatics 25(23), 3158–3165 (2009)
Rocha, I., Maia, P., Evangelista, P., Vilaça, P., Soares, S., Pinto, J.P., Nielsen, J., Patil, K.R., Ferreira, E.C., Rocha, M.: OptFlux: an open-source software platform for in silico metabolic engineering. BMC Syst. Biol. 4(1), 45 (2010)
Rocha, M., Maia, P., Mendes, R., Pinto, J.P., Ferreira, E.C., Nielsen, J., Patil, K., Rocha, I.: Natural computation meta-heuristics for the in silico optimization of microbial strains. BMC Bioinformatics 9(1), 499 (2008)
Feist, A.M., Henry, C.S., Reed, J.L., Krummenacker, M., Joyce, A.R., Karp, P.D., Broadbelt, L.J., Hatzimanikatis, V., Palsson, B.Ø.: A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Biol. 3(121), 1–18 (2007)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this paper
Cite this paper
Vieira, V., Maia, P., Rocha, I., Rocha, M. (2016). Development of an Integrated Framework for Minimal Cut Set Enumeration in Constraint-Based Models. In: Saberi Mohamad, M., Rocha, M., Fdez-Riverola, F., Domínguez Mayo, F., De Paz, J. (eds) 10th International Conference on Practical Applications of Computational Biology & Bioinformatics. PACBB 2016. Advances in Intelligent Systems and Computing, vol 477. Springer, Cham. https://doi.org/10.1007/978-3-319-40126-3_20
Download citation
DOI: https://doi.org/10.1007/978-3-319-40126-3_20
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-40125-6
Online ISBN: 978-3-319-40126-3
eBook Packages: EngineeringEngineering (R0)