Abstract
Proteins are highly perfected natural molecular machines, owing their properties to the complex tertiary structures with precise spatial positioning of different functional groups that have been honed through millennia of evolutionary selection. The prospects of designing new molecular machines and structural scaffolds beyond the limits of natural proteins make design of new protein folds a very attractive prospect. However, de novo design of new protein folds based on optimization of multiple cooperative interactions is very demanding. As a new alternative approach to design new protein folds unseen in nature, folds can be designed as a mathematical graph, by the self-assembly of interacting polypeptide modules within the single chain. Orthogonal coiled-coil dimers seem like an ideal building module due to their shape, adjustable length, and above all their designability. Similar to the approach of DNA nanotechnology, where complex tertiary structures are designed from complementary nucleotide segments, a polypeptide chain composed of a precisely specified sequence of coiled-coil forming segments can be designed to self-assemble into polyhedral scaffolds. This modular approach encompasses long-range interactions that define complex tertiary structures. We envision that by expansion of the toolkit of building blocks and design strategies of the folding pathways protein origami technology will be able to construct diverse molecular machines.
*Author contributed equally with all other contributors.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Protein origami
- Protein design
- Coiled-coil
- Building blocks
- Protein assemblies
- Modular topological folds
- Oligomerization domains
2.1 Protein Architecture
It is no coincidence that proteins, more than any other type of biomolecule, are the workhorses of life. They are responsible for executing and in most cases also regulating the vast majority of cellular processes. The reason nature has chosen proteins for this vast, varied, and very dynamic range of tasks lies in their versatility. The range of 20 amino acids with their chemically diverse side chains offers no less than 8000 possible permutations for a single tripeptide (compared to 64 for a trinucleotide). We must add to that the much greater conformational freedom of the polypeptide backbone in comparison with nucleic acids or saccharides, and the number of possible configurations becomes staggering. Given this enormous variability in sequence and structure, proteins appear to have a nearly unlimited functional potential under conditions conducive to life (aqueous solution, temperatures in the −20–+100 °C range, pressure on the ~1 atm scale). With all this variability, the problem then becomes how to find the sequence that will selectively produce the desired structure and perform the desired function. Nature solved this problem using evolutionary selection, which required hundreds of millions of years. In order to harness the tremendous potential of proteins beyond natural proteins or their close homologues and to endow them with new interesting features we need a more efficient design strategy than trial-and-error. While protein engineering has been used since the dawn of the recombinant DNA technology, researchers have only recently started to design new protein folds and design polypeptide-based structures based on the rational design principles. Protein structure modeling techniques demonstrated important progress, supporting accurate modeling of protein-protein interactions and enabling assembly of proteins into larger complexes. For the design of entirely new protein folds, on the other hand, a topological modular design approach has been presented, also called protein origami for resembling the approach used to design DNA nanostructures .
Protein function is inevitably linked to its structure, whether the latter is stable or dynamic [1]. The first step in any protein functional design strategy is therefore to consider what structure(s) would serve the desired function and how to make the protein assume such a structure. This is a difficult problem and still not fully resolved after more than 50 years of research [2–4]. Essentially, native protein structure is determined by a multitude and cooperativity of individually weak interactions among the residues of a particular polypeptide sequence, as well as interactions between the residues and the solvent (water along with ions and other cosolutes). All these interactions depend on the specific conformation of the protein and given the vast number of conformational degrees of freedom available to a protein, it is not feasible to simply calculate the energy of every possible conformation and select the most stable one. The complexity of the protein design problem therefore stems from the same abundance of variability that confers on proteins their versatility in performing a variety of tasks. Despite the difficulties involved in understanding protein structure, the basic design principles of naturally occurring proteins have by now been established.
Natural proteins are largely stabilized by the hydrophobic effect that is exclusion of non-polar surfaces from water. This stems from the fact that interactions between non-polar side chains and water are weaker than the sum of interactions of non-polar groups among themselves and water molecules among themselves [5]. To optimize the energy of these interactions, hydrophobic portions of a protein will tend to cluster together at the core of the structure, removing themselves from the contact with water, thus stabilizing a compact structure. The hydrophobic effect is not specific to proteins and applies very generally to non-polar surfaces in contact with water [6]. Accordingly, such non-specific interactions cannot be relied upon to produce a single, well-defined designed tertiary structure. For that sort of specificity, proteins must turn to more selective interactions like charge-charge interactions and hydrogen bonds that preferentially form between specific groups in specific geometries, rather than being driven by their aversion to water. These interactions are responsible for maintaining the protein’s secondary structure. Because there are only a few possible backbone configurations that allow efficient hydrogen bonding between the peptide groups of the protein backbone (primarily α-helices and β-sheets) [7, 8], these structural motifs are reused over and over in many different proteins. The way these secondary structure elements combine into a three-dimensional structure depends on the order in which they are arranged in the primary sequence (which will limit the number of possible arrangements), the polarity of their side chains (as non-polar residues will tend to turn away from water), and finally on the more specific interactions between their polar and/or charged side chains. The latter are very strong, yet they do not make a dominant contribution to stabilizing the structure compared to the unfolded state, since they can be complemented by water and counter ions when exposed to the solvent. They do, however, confer great specificity to the structure, because the few conformations that allow all their polar groups to make proper contacts (without exposing large non-polar surfaces to water) will be much more stable than any conformation that leaves some of these strong interactions unformed [9]. In this context it also bears mentioning that constraining the protein to just a single conformation out of a myriad possible ones is entropically unfavorable, meaning that it is statistically unlikely to happen without any help from specific stabilizing forces. The attractive interactions that hold the protein structure together must therefore be strong enough to overcome this intrinsic tendency of molecules to sample all possible conformations. Protein structure is thus the end result of a fine balance between conformational entropy, topological constraints, the hydrophobic effect, and the need to maximize the number of favorable interactions of polar groups among themselves or with water [10–13].
As a result of these considerations, naturally occurring proteins have evolved a considerable but finite number of folds that satisfy all the requirements of a stable structure. To date, almost 1400 distinct protein folds have been registered among the experimentally determined protein structures in the PDB [14], with very few new ones being discovered in recent years. This apparent saturation of the number of natural folds indicates that a limited number of folds had evolved through the evolution. The common features of natural protein folds are a hydrophobic core of the protein, where a large fraction of the non-polar residue side chains are packed and excluded from water, and where backbone hydrogen bonds are satisfied by strict adherence to regular secondary structures. The loops that connect these elements of secondary structure are located at the surface of the protein, where missing hydrogen bonds can be complemented with water. The same applies to most polar amino acid residues, although a few of these also make contacts in the protein core; these tend to be important for maintaining a specific, non-degenerate structure because breaking them requires either a lot of energy or replacing them with water, thereby also exposing the hydrophobic core [9, 15, 16]. The elements of secondary structure that make up the protein core can be arranged in different topological configurations, depending on the order in which they appear in the amino acid sequence. Formation of a compact hydrophobic core brings together residues that are far apart in the primary sequence. While forming such long-range interactions is critical to establishing a compact protein fold, it also appears to be the rate-limiting step in the process of protein folding. The prevalence of long-range contacts, also called contact order, correlates with lower folding rates [17, 18]. For protein stability, on the other hand, no such simple correlations have been found, beyond the fact that in general larger proteins tend to be more stable due to the larger number of intramolecular contacts [19, 20]. That is not to say that all proteins of the same size display the same stability, but rather that the differences between them are hidden in the myriad of individual intramolecular contacts that are specific to each protein and not easily predicted.
2.1.1 Designed Protein Assemblies
With respect to designing stable protein structures, the main obstacle is ensuring specificity of the folded conformation, because this requires placing strongly interacting residues in the right positions so that they can interact in the desired structure, but not in any undesired alternative conformations. We lack the precision and computing power to predict and design the balance of specific interactions from scratch, so the most efficient current methods are based on naturally occurring fragments of protein structure that are known to interact specifically and using those as building elements to engineer specific interactions into our systems [21, 22].
2.1.2 Tethered Oligomerizing Protein Assemblies
Interactions between protein domain subunits can be introduced by designing ligand binding sites (in particular metal binding sites) or disulfide bridges in order to obtain new protein assemblies. Such design problems are computationally more difficult than genetic fusion, but generally simpler than de novo design of protein-protein interaction surfaces, since fewer interactions need to be accounted for. Using binding sites for small ligands allows creation of smart bionanomaterials by regulating the assembly and disassembly. Self-assembly of the fusion protein composed of dimerization domain gyrase B and trimerization domain can be driven by the addition of a small molecule. Coumermycin, a gyrase B dimeric ligand induces formation of the periodic protein lattice with nanopores of defined size. The disassembly of the lattice is achieved by the subsequent addition of monomeric novobiocin which competes for the same gyrase B sites as the coumermycin [21].
Metal-mediated protein interaction is geometrically specific with tunable interaction strength, making it an attractive alternative for building protein assemblies. The strength of binding can be controlled by the concentration of the transition metal or by the local pH. By using metal template interface redesign it is possible to convert naturally occurring monomeric proteins into oligomeric assemblies [23]. The reverse, converting a self-assembling protein into a protein that assembles only in the presence of transition metals is also possible [24]. Using this approach, chemically tunable three dimensional protein arrays have been designed [25]. The structure of the assemblies is very dependent on the growth conditions such as pH and metal to protein ratio. Binding of metal can also induce conformational changes. A homotrimeric coiled coils has been designed, that forms fibrils in the presence of cadmium ions and separate trimeric units in the absence of metal ions [26]. The coil was designed to have blunt ends when the helices are in register. The binding of Cd+2 ions shifts the relative orientation of the helices, creating staggered ends enabling assembly of fibrils. Fletcher et al. [27] reported self-assembling cages from a coiled coil homotrimeric (CC-Tri3) and a heterodimeric peptide (CC-Di-AB). CC-Tri3 and CC-Di-A or CC-Di-B were connected via disulfide bonds to form two types of triangular hubs (A and B). Mixing hubs A and B formed closed spherical cages of approximately 100 nm presumably due to the intrinsic curvature of building blocks .
2.1.3 Oligomerizing Protein Domain Fusion Strategies
Many natural oligomerization domains , typically containing 100–200 amino acid residues, can non-covalently self-assemble into larger, often symmetric, superstructures. Fusion strategies are based on (genetically) linking two or more natural oligomerization domains, thereby generating a molecule with two or more interaction surfaces. This was one of the earliest ideas for generating protein assemblies, as extensive design of new interaction surfaces was not needed. In order to form precise geometrically defined structures, the orientation of the two subunits must be tightly controlled.
A pioneering study fused a dimer and a trimer promoting domain with a continuous alpha helix [28]. Twelve copies of the fused protein were designed to assemble into a tetrahedral cage. Introduction of two mutations enabled the production of homogeneous cage-like particles 16 nm in diameter with a 5 nm cavity. Significant deviations from the idealized predicted tetrahedral model were observed, which were mostly a consequence of the bending and twisting of the helical linker [29]. The study thus outlined the need for developing more rigid linkers. If higher orders of symmetry are present in oligomers, then the linker can be aligned along an axis of symmetry, bypassing the need for a rigid linker. This approach was used to successfully create filaments and 2D crystals [30]. The largest finite cage-like assembly generated using the genetic fusion strategy is currently a 24 component 22.5 nm large cube with a 13 nm diameter inner cavity. Crystals of the structure were obtained after prolonged incubation (longer than half a year). The crystal structure matched the design very well with a backbone RMSD of only 1.2 Å. However in solution significant populations of 12-mers and 18-meres and a detectable amount of trimers were also present. The authors speculate that the heterogeneity may be due to the flexibility of linkers, although kinetic effects of assembly of different states cannot be excluded.
2.1.4 Designing New Interaction Surfaces for Assemblies Based on Oligomerizing Domains
The most general design strategy based on multiple folded protein domains involves designing de novo interactions surfaces. This method has become possible due to powerful computer programs for designing weak non-covalent interaction surfaces. King and et al. [31] designed particles with octahedral and tetrahedral symmetry, starting from natural trimeric building blocks . The design was accomplished in two stages: first natural homomeric trimers were docked and screened to identify candidates with “designable interfaces”. The score function used for screening favored interfaces with a high density of contacting residues in well-anchored regions. Due to the high symmetry of the final assembly, only two degrees of freedom for the orientation of the trimers had to be scanned. In the second stage, the interfaces were designed at atomic detail. For both stages the Rosetta [32] framework was used. Eight tetrahedral and thirty-three octahedral sequences were tested experimentally and one from each category was successfully crystalized. The octahedral particle matched the design very well, with backbone RMSD smaller than 1 Å. The tetrahedral particles matched the design with RMSD better than 5 Å. Based on the crystal structure, an improved tetrahedron was designed and tested. The improved tetrahedron matched the design very well. In another study King et al. [33] designed and characterized five tetrahedral 24 subunit cage-like particles. The particles were composed either of two kinds of trimers (T33 symmetry) or of trimers and dimers (T32 symmetry). 30 T32 and 27 T33 sequences were experimentally tested and 1 T32 and 4 T33 were crystalized and shown to match the design very well, with backbone RMSD ranging from 1 to 2.6 Å. Most recently, the same strategy has been used to construct a highly stable designed icosahedral cage with a 25 nm diameter and a large (approximately 3000 nm3) central cavity [34].
Lanci et al. [35] designed a 3D protein crystal with a P6 symmetry, which has “honeycomb-like” channels that span the entire structure. This approach involved designing backbone structures with consistent target symmetry, screening them for “designability” and finally designing the sequences for the identified structures. In the study a de novo designed homotrimeric coiled coil was used. A single low-energy sequence (P6-a) was identified and experimentally tested. P6-a formed diffraction-quality crystals overnight, but crystallized in the P321 space group with neighboring proteins in antiparallel arrangement, instead of the intended P6 with the parallel alignment of all subunits. Five further sequences at different parts of the sequence-structure energy map were tested. One of the structures was successfully crystalized. The crystal structure (with P6 symmetry) was nearly identical to the design with backbone RMSD of 0.45 Å.
In principle the design of new interaction surfaces is not bound by symmetry restraints, however the use of highly symmetrical design confers the advantage of having to design fewer new interactions and to increase the robustness.
2.1.5 Repeat Domain Proteins
An alternative approach to modular protein structure design is provided by repeat proteins . These are naturally occurring proteins composed of repeating structural motifs/domains that can be stacked one after the other as modules to create single chain proteins with predictable structures and a considerable range of lengths, stabilities, and even folding energy landscapes [36–38]. The down side of this approach is the bulkiness of the structural backbone, as well as the limited range of the complexity of protein structures that can be built using modules that only interact with their nearest neighbor modules. Another way of addressing the protein design problem is to use smaller, more independent building blocks , such as individual elements of the secondary structure. β strands tend to interact non-specifically with each other, so they are difficult to assemble into well-defined structures. Instead, they have been engineered to form assemblies such as fibrils or hydrogels that possess certain interesting properties [39–41]. For a more precise control of structure and function, α helix-based design has proven most promising. Helices allow construction of both multimeric assemblies as well as novel single-chain protein folds, and are discussed in detail in the following sections as implemented in protein origami.
2.1.6 The Importance of Long Range Interactions to Define Complex Shapes
As we noted above, specific long-range interactions are essential for assembling chains of biological polymers into specific tertiary structures. As an example, consider the difference between the linear B-DNA structure which results from contacts between the neighboring nucleotides in the sequence, and DNA origami, where long-range contacts facilitated by stapler oligonucleotides enable the design of a multitude of intricate 3D structures [42]. The same is true for proteins and most naturally occurring proteins have quite complex contact maps (Fig. 2.1). By contrast, interactions in a repeat protein are localized to nearest neighbor repeats which allow only longitudinal stabilization along the length of the sequence to this type of proteins, restricting them to linear or circular/helical structures. Protein origami, in analogy to DNA origami, introduces long-range interactions into the polypeptide chain in a rational, controlled manner, allowing design of complex three-dimensional structures. This technology holds great promise but also presents a number of challenges that will need to be addressed in order to unlock its full potential. For example, any effort to engineer precise assembly of complex structures will require a sufficiently large set of specifically interacting (orthogonal) structural elements. Folding kinetics can also be complicated by the introduction of numerous long-range contacts and its control would be highly desirable. While genetic fusion protein assemblies composed of natural protein domains and novel interaction surfaces are based on the symmetric oligomerizing domains, another quite novel approach to modular protein assembly in the spirit of synthetic biology is to design topological polypeptide folds, based on concatenated coiled-coil interacting domains. The concepts, challenges, and successes of designed protein origami are discussed in the remainder of the chapter.
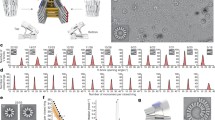
Long range contacts as determinants of protein tertiary structure. A comparison is drawn between the three-dimensional structures of three classes of proteins and the maps of residue-residue contacts (cutoff = 8 Å) that stabilize these structures. In most natural proteins like GFP (PDB ID: 1GFL), contacts between residues that are far apart in the amino acid sequence (off-diagonal on the contact map) allow the protein structure to close itself up and protect its hydrophobic core from water. In repeat proteins like the ankyrin repeat protein Asb11 (PDB ID: 4UUC), structural modules only interact with their nearest neighbors, which results in an overall linear or twisted structure. The hydrophobic core is limited to the contact interfaces between the repeats. A topological fold a.k.a. designed protein origami (Model structure from [90]) uses coiled coil units as building modules, where dimeric interactions occur only along a narrow hydrophobic spine running along the length of each helix-helix interface. Despite similarly modular composition as repeat proteins, a complex three-dimensional fold can be achieved by ordering interacting segments in such a sequence that coiled coil pairing defines the fold via discrete long-range interactions
2.2 Coiled-Coils as Versatile Building Blocks
2.2.1 Basic Structure of Coiled-Coils
Coiled coils are dimers or higher oligomers composed of α-helices. In addition to different oligomerization states, coiled-coils may differ in the orientation of chains, namely they can oligomerize either in the parallel or antiparallel direction (Fig. 2.2), in comparison to the canonical antiparallel double helix of nucleic acids.
Representation of parallel and antiparallel coiled coil dimers. (a) Side view of a parallel homodimer (2zta). (b) Side view of an antiparallel heterodimer (3qo5). The a and d sites in the heptad repeat are represented with violet and blue spheres. (c) and (d) A schematic representation of the structure of the abcdef heptad repeat and its interaction surface in the parallel and antiparallel orientation
This extends the range of assemblies that can be designed and will be described in the section devoted to the topology of designed protein origami . While natural coiled-coil dimers are often homodimers, which requires only a single coding polypeptide chain, the heterodimers offer a higher degree of control in the assembly.
Coiled coils are stabilized by a characteristic ‘knobs-into-holes’ packing, where sidechains (knobs) of one helix fit between the four residues (a hole) on the other helix [43]. Such regular packing requires periodicity, which is not possible with a pitch of 3.6 residues per turn of regular α-helices. By coiling right hand α-helices into a left hand superhelix, the value of residues per turn is reduced to 3.5 (periodicity 7/2), giving rise to the hallmark heptad repeat per two turns of the helix. The amino acids in the heptad repeat are labeled as abcdefg as shown in Fig. 2.2. In coiled-coil dimers and trimers sites a and d usually contain hydrophobic residues (such as Val, Leu, Ile), which confer stability due to packing of hydrophobic residues, while sites e and g are typically occupied by complementary charged residues (for example a Lys and Glu pair), which confer specify of binding through electrostatic complementarity. The b, c and f sites do not directly participate in interactions with the other helix in dimers and can therefore be used to modulate the desired properties of the peptide, for example by introducing residues for specific interactions with other molecules.
The structure of coiled coils represents one of the few protein folds that can be described mathematically. The parametric description of the structure was proposed in as early as 1953 by Crick [44] and Pauling [45]. Several excellent reviews are available on the topic of coiled coils structure [46–49].
Although coiled coils may seem deceptively simple to build complex tertiary structures, the fold represents extremely versatile building blocks. The structural motif represents at least 2 % of encoded residues in most organisms and 8 % of the residues in the human proteome [50].
2.2.2 Functional Role of Coiled-Coils in Nature
Due to their elongated shape and rigid structure, coiled coils make excellent scaffolds, levers and rods [51]. The coiled-coil motif was first discovered in mechanically rigid fibrous proteins such as keratin and fibrinogen. As efficient spacers coiled coil domains are present in all classes of cytoskeletal motor proteins (myosins, kinesins and dyneins) [47]. The longest known coiled coil (protein PUMA1 [52]) spans an amazing 1750 amino acid residues (or 250 nm) and is involved in the organization of the mitotic spindle.
The biological role of coiled coils is not limited to their structural role as rigid rods but are also involved in the molecular recognition and in fact represent one of the most common dimerization motifs. Many transcription factors, including one of the largest family transcription factors in humans, the basic region-leucine zipper (b/ZIP) family, contain a coiled coil dimerization domain, which is responsible for specific and controlled homo- or hetero-dimerization. In fact it was the b/ZIP yeast activator GCN4 [53, 54], that refocused the direction of research from long and fibrous to shorter coiled coil domains. GCN4 remains one of the most studied coiled coil systems, but considerable progress has been made in elucidating the interaction network of other members of the bZIP family [55, 56]. Coiled-coil interactions also play an important role in membrane trafficking and fusion, where recognition is based on the dynamic formation of a four-helix coiled coil bundle. The target membrane contributes three helices (one from SNARE protein and two from SNAP25 protein) while the vesicle membrane contributes the final helix (synaptobrevin) [57]. Finally the assembly of coiled coils can be regulated by pH [58, 59], phosphorylation [60] and interactions with ions [61].
2.2.3 Engineered Coiled-Coils
Coiled coils are the most well understood protein structure motifs and have proved very useful in protein design and engineering [63]. The first rationally designed coiled coil was an analogue of tropomyosin [63]. The field rapidly expanded with the design of a “peptide velcro” [64], a leucine zipper based on GCN4 and the Fos/Jun transcription factors. An antiparallel variant followed [65], establishing rules for setting the orientation of coiled coil dimers using a polar Asn introduced at a and d sites. One research direction pursued building bundles with ever more alpha helices. As the rules governing oligomerization states were elucidated [66], first trimers [67] and then tetramers [68] were developed and even a seven-helix coiled coil [69]. A database of coiled coil tertiary structures [70], as well as classification of coiled coils packing, termed “A Periodic Table of Coiled-Coil Protein Structures” is available [71]. The affinity of coiled coils can be readily tuned, giving rise to interesting applications, such as temperature biosensors [72], or probes for tumor markers [73].
2.2.4 Engineering Coiled-Coil Orthogonality
Modular and orthogonal components have been regularly used in other engineering fields, such as the design of cars, computers and software. Modularity offers flexibility, a shorter learning time due to abstraction of complexity, and the ability to extend the functionality by the addition of other modules. The net result is a reduction of cost in design and manufacture of products. Modular assembly utilizing polypeptide domains requires either high degree of symmetry of the assembly or utilization of a larger number of orthogonal modules, which is required for the complex assemblies.
Several small set of orthogonal coiled coil dimers have been reported. Reinke et al. [74] measured the interactions between 48 synthetic and 7 human bZIP coiled coils using peptide microarrays. From the interaction matrix only a set of two parallel heterodimeric coiled coils was identified, therefore the rational design of the orthogonal building modules seems to be more productive. In designing orthogonal toolkits, where binding specificity is as important as the binding affinity, both positive and negative design principles must be used [75]. Positive design refers to optimizing binding interaction with the desired target partner, while negative design involves the destabilization of undesired states, such as binding to other sequences in the toolkit or trimer formation. In short, the designed sequences must have a preference for binding the target partner over all other undesired off-target states. Bromley at al. [76] used a reduced set of amino acids at the adgf positions and a scoring matrix based on bCIPA to design three pairs of short parallel coiled coil dimers. Gradišar et al. [77] used the principles governing the selectivity and stability of coiled-coil segments to design four pairs of parallel coiled coil dimers comprising four heptads. The orthogonality of peptide pairs was confirmed using circular dichroism (CD) spectroscopy. The design of an orthogonal parallel CC dimer set was based on the combinatorial variation of the heptad patterns, using two different types of heptads based on the EK electrostatic pattern between positions e and g within the heptad and introduction on an Asn residues into the a position, versus the Ile residues, while the d position was kept as the invariant Leu residue. The heptad patterns used in the design are presented in Table 2.1. This set was used for the design of self-assembling single-chain tetrahedron as described later.
Negron et al. [78] used a computational approach to design three pairs of antiparallel coiled coil homodimers. The orientation and orthogonality of the designs was tested using disulfide exchanges and CD spectroscopy.
2.2.5 Computational Tools for the CC Design
Several tools, most of them available as free web applications, are available to assist in the rational design of coil coiled structures and sequences. Many algorithms have been proposed for predicting the coiled coil motif and its oligomerization state from the amino acid sequence, such as SCORER 2.0 [79] and ProCoil [80], that can classify a sequence with assigned heptad registers as either parallel dimers or trimers. RFCoil [81] improves these predictions given the same input data. Multicoil2 [82] can assign heptad registers and distinguish between dimers and trimers. LOGICOIL [83] can predict oligomeric states up to tetramers (including antiparallel dimers) and heptad registers given sequence information alone.
Temperature melting points for the bZIP family of coiled coils (parallel dimers) can be estimated using bCIPA [84] using only sequence information with assigned registers. Given a 3D structural model, the COILCHECK [85] webserver can be used to obtain interaction energies between two helices in a coiled coil bundle. SOCKET [43] is program that identifies coiled coils in 3D structures by finding the characteristic knobs-into-holes packing between helices. Since structural information, along with the most basic feature of coiled coils is used, the algorithm represents the most reliable method for identifying coiled coils. SOCKET also enabled the development of the CC+ database of all know 3D structures of coiled coils [70].
CCBuilder [86] is a web-based application for building 3D model structures of coiled coil bundles given the Crick backbone parameters and a sequence with assigned heptad registers. Bundles with arbitrary number of coils and orientations can be built. The basic interface enables construction of more than 96 % of coiled coil types in the CC+ database, while an advanced mode enables even more unusual coiled coils to be constructed. TWISTER [87] and CCCP [88] are programs for extracting the Crick backbone parameters from 3D structures. TWISTER was written to work primarily with parallel orientations in mind, while CCCP can obtain also parameters for antiparallel alignments such as the Z-shift.
2.2.6 Attractive Features of CC Dimers
Several features make the coiled coil motif one of the most attractive elements for protein engineering . Perhaps the most attractive feature is the fold’s simplicity. The sequence/structure relationship of coiled coil structures is quite well understood. Several rules-of-thumb have been devised that allow specifying the oligomerization state and orientation of alpha helices in a coiled coil bundle [62]. The parametric description of the coiled coil backbone enables efficient exploration of conformational states, vastly simplifying computer assisted design [88]. Despite the apparent simplicity, coiled coils are very versatile and widely used building blocks. Efficient spacers, scaffolds, rods and levers can be made, as a coiled coil dimer requires only 14 amino acids per nanometer of distance. Coiled coils can also oligomerize with an affinity and specificity than can be easily tuned. Coiled coil dimers obtain a stable structure above 25 residues and are thus smaller than typical globular dimerization domains which start at about 70 residues. A smaller number of amino acids translate into smaller genes that are easier to manipulate, clone and express.
2.3 Designed Protein Origami – Modular Topological Protein Fold
While nucleic acids are able to fold into compact tertiary structures defined by the cooperative weak interactions between nucleotides similar to protein folds the large majority of DNA exists in form of a DNA duplex based on complementary AT, GC pairs. This straightforward complementarity allows design of orthogonal sequences that discriminate strongly between the correct and incorrect pairs, providing an almost unlimited set of orthogonal pairs. Combinations of nucleotide sequences that share complementary segments allowed formation of cruciform Holliday junctions that gave rise to the field of DNA nanotechnology three decades ago. The key components of designed DNA nanostructures are orthogonal long-range pairwise interactions between concatenated interacting modules. This approach developed several strategies, mainly based on the self-assembly from many short or long DNA strands comprising at least two complementary segments to make versatile tertiary structures. Nowadays DNA nanotechnology can make almost any selected 3D shape such as different polyhedra, lattices, arbitrary shapes as well as molecular machines able to perform logic functions as well as locomotion. While DNA nanostructures have been functionalized to bind different molecules and implement chemical reactivity introducing functionality, the ideal designed molecules should combine the designability of shapes of DNA nanostructures with the versatility of side chains of proteins (Fig. 2.3).
Inspired by the spectacular demonstration of the complex molecular self-assembly achieved by the DNA nanotechnology we decided to explore the implementation of a similar concept into the polypeptide-based designed nanostructures using coiled-coil dimers as the modular building blocks . We reasoned that orthogonal coiled-coil forming peptides concatenated into a single chain are potentially more suitable as building blocks compared to much larger natural oligomerizing protein domains. This assumption also enables the precise control of the assembly geometry and allows self-assembling of the asymmetric polyhedral nanostructures . The advantage of the modular protein self-assembly in comparison to native protein folds or combinations of folded protein domains is that it should be much easier to design new folds. Additionally this new type of protein folds, unseen in nature, might provide proteins with new interesting properties.
The key component of designed protein origami are the concatenated coiled-coil dimer forming segments that selectively pair to another segment within this or another chain. In this respect this strategy resembles very much the idea of DNA nanostructures. The basic requirement is to have available the set of orthogonal coiled-coil dimers that direct the fold of the polypeptide chain. The coiled-coil modules are concatenated to each other by the flexible peptide linkers that act as hinges that assemble the scaffold of the rigid coiled-coil dimers (Fig. 2.4).
The three-dimensional polyhedra are constructed by coiled-coil dimers as the rigid edges, while the flexible hinges converge at the vertices of the polyhedra. Therefore the problem of designing the polypeptide-based polyhedron can be abstracted into the trail along the graph, where vertices are connected by a double path therefore each edge must be crossed by the polypeptide chain exactly twice. Therefore the polypeptide polyhedron represents a molecular embodiment of a mathematical concept. As described in the next section, mathematical topology can provide firm proofs on the possible solutions to the problems of the coiled-coil module based assembly. Selection of coiled-coil dimers as the building blocks turned out to be particularly appropriate as we can and must use both parallel and antiparallel coiled-coil dimers for the construction of the single-chain tetrahedron. The required building blocks for the construction of a tetrahedron are six orthogonal coiled-coil dimers that form six edges of the tetrahedron. Each of those coiled-coil forming segments is, in isolation, unstructured and forms a coiled-coil only when it independently dimerizes with the corresponding complementary segment. Therefore 12 coiled-coil segments were concatenated into a single polypeptide chain with flexible tetrapeptide linkers between each segment. The role of those segments was to break the helix-forming segments, provide the kink in the direction of the chain and sufficient flexibility to allow assembly of the edges onto the final fold. The required angle between the edges in the selected polyhedron is defined only by the length of the edges, following the mathematical requirements to define the shape of the polyhedron by the length of all of its edges.
In comparison to native protein folds the topological polyhedra do not have a hydrophobic core to anchor the elements of the secondary structure. The hydrophobic interactions are restricted to the well-understood and designable interactions between the coiled-coil dimers, while the global fold is defined by the topology of the interacting segments. The order of coiled-coil segments uniquely defines the global fold in a similar way as the order of amino acid residues defines the fold of native proteins. Scrambling the order of coiled-coil forming segments prevents correct assembly. Order does not restrict the selection of specific segments but rather that e.g. the first segment must for an antiparallel dimer with the fifth segment, the second segment must form a parallel dimer with the eighth segment etc. Consequently many permutations are possible, however only a small fraction of the possible orders of segments is able to fold into a correct structure. This type of the fold is therefore not just a new fold unseen in nature but it represents a new type of protein folds, defined by the topology of the chain rather than by packing of the hydrophobic protein core.
For the first demonstration of this new type of protein design we selected a tetrahedron composed of a combination of designed and natural coiled-coil dimers, comprising both homodimers as well as heterodimers (Fig. 2.5). Three of the parallel pairs were selected from the designed orthogonal coiled-coil forming set [77], each composed of four heptads and designed based on the known coiled-coil stability and selectivity principles. In addition to the designed parallel heterodimers, one parallel homodimer based on the natural GCN4 and two antiparallel homodimers [89, 90] were used. The tetrapeptide Ser-Gly-Pro-Gly was selected as the flexible linker to connect the consecutive coiled-coil forming segments.
Modular topological design of a protein fold from a single chain. (a) The designed shape of a polyhedron is decomposed into the edges, which are composed of rigid coiled-coil dimers. (b) Building blocks for coiled-coil dimeric edges are selected from a tool box of orthogonal coiled-coil dimers. The polypeptide path is threaded through the edges of a tetrahedron traversing each edge exactly twice, so that the path interlocks the structure into a stable shape stabilized by the six coiled-coil dimers, where four of them have to be parallel and two antiparallel. Coiled-coil forming segments are concatenated in a defined order into a single polypeptide chain with flexible peptide linker hinges. (Reproduced by permission from the Current Opinion in Chemical Biology [21])
As described later, only three different topologies are available for the designed tetrahedron, one combining four parallel and two antiparallel dimers and two different topologies combining three parallel and three antiparallel dimers. The important advantage of protein nanostructures in comparison to the DNA nanostructures is that the protein can be produced in large amounts using biotechnological methods. The synthetic gene was assembled to encode the designed polypeptide sequence encoding for the tetrahedron which allowed its production in E. coli and purification. The recombinant protein did not assemble correctly in bacteria and had to be isolated and refolded by annealing in slow dialysis, similar as DNA nanostructures.
The self-assembled protein nanostructures were investigated by atomic force microscopy and electron microscopy which verified the correct shape and size according to the design, additionally gauging the size by the gold nanobeads, coupled to the C-terminus. Polypeptide self-assembled into a stable nanoscale tetrahedral structure whose edges measure around 5 nm, as confirmed by the DLS and MALS analysis. According to the mathematical rules underlying the trail of the graph, the beginning and end must coincide in the same vertex, which we demonstrated by the reconstitution of the split fluorescent protein, genetically linked to both ends of the tetrahedral (TET12) polypeptide.
In comparison to designed protein assemblies based on oligomerization domains the designed protein origami is not symmetric and each of its edges or vertices may be addressed separately. The polypeptide scaffold occupies much lower fraction of the volume than the assemblies composed of folded protein domains.
The cavity of the tetrahedral fold could be augmented by using longer modules, i.e. the number of heptads of peptide segments must be increased. This fact significantly limits the set of orthogonal coiled-coil pairs therefore expansion of the number of available modules is needed. Another way to prepare the structures with larger cavity is to design higher polyhedra, such as a trigonal bipyramid.
2.4 Mathematical Abstraction of Modeling of the Topology of Protein Origami
2.4.1 String as an Abstract Model
Our abstract model assumes we are designing one or more directed strands (polypetide chains), composed of segments, connected by flexible linkers. Furthermore we assume that each segment of the collection of strands attaches to a unique segment of the system, thus forming a dimer. Finally we assume that after completion of all attachments a single stable polyhedron is formed with dimers as edges. A dimer may be parallel or anti-parallel. We will represent each segment by a symbol and each strand by a string. A prime example of a single strand self-assembly is TET12, designed by Gradišar et al. [91] and described in the previous section. Their segments were originally named:
Three of the dimers were heterodimers: P3-P4, P5-P6, P7-P8 and three were homodimers APH-APH, BCR-BCR, GCNsh – GCNsh. Furthermore four dimers were parallel and two were anti-parallel: APH-APH, BCR-BCR.
By ignoring the information about the hetero-homo nature of dimers, and using capital letter or exponent −1 to represent the anti−parallelism, we may use the following transformations:
Our abstract encoding:
contains sufficient information for a computer to recreate the self-assembled tetrahedron. In the case of TET12 the string contains 12 characters. Mathematically, it represents an oriented fundamental polygon of a closed surface, see Fijavž et al. [92]. Any of the 12 cyclic permutations of the string yields topologically the same self-assembly. In practice this means that the original strand may be modified in such a way that it is cut in two pieces and the order of the two pieces is interchanged in the design of the new strand. In (*) we are using standard encoding. This means we use consecutive letters of the alphabet, starting with a. An uppercase letter appear only after the corresponding lower case letter has been used.
The reflection of the original string, say (*),
represents the same fundamental polygon with the reverse orientation, yielding again the same self-assembled structure. Note that (**) is not written in the standard form but can be easily rewritten in a standard encoding.
Standard encoding has some advantages but also disadvantages. Two strings are equivalent if and only if they have the same standard form. Standard form thus represents a canonical labeling of a string. On the other hand by changing the labeling from (**) to standard (***) we also relabeled the edges of the tetrahedron.
In addition to 12 cyclic rotations that will generate the same tetrahedron, we may add also 12 reflections, obtained by forming a sequence in the reverse order of segments. All these 24 strands will self-assemble into the same topological form: the tetrahedron. A natural question is: how many different topologies are there? How many strands will self-assemble into the same polyhedral shape? In Gradišar et al. [91] it was shown that there are three non-equivalent topologies forming tetrahedron. Each of them is equivalent to its reflection after some rotation. By choosing lexicographically the first string from the equivalents we obtain the following three cases:
The first one has two antiparallel dimers while the other two have three anti-parallel dimers. The first and the second have indeed 12 different strings each. The third one has three symmetries, hence it has only 12/3 = 4 distinct strings. This means that there are 12 strings with two anti-parallel dimers and 16 strings with three anti-parallel dimers.
2.4.2 Trigonal Bipyramid
The situation is quite different in the case of trigonal bipyramid. There are 30 distinct directed fundamental polygons, 12 of them being equivalent under the reversal of orientations and 18 cases obtained by 9 pairs with opposite orientation. Out of 30 cases 10 have two anti-parallel dimers, 4 have 3 anti-parallel dimers, 1 has 4 anti-parallel dimers, 6 have 5 anti-parallel dimers and 9 have six anti-parallel dimers.
Table 2.2 presents the complete analysis for the trigonal bipyramid. In total there are 468 non-equivalent strands that will self-assemble into a trigonal bipyramid. Note that the bipyramid has 5 vertices and 9 edges. It has two types of vertices, three lying in the equator and the other two on poles. It also has two types of edges, three on the equator and 6 having one end-vertex at the pole. In total there are 12 symmetries of the solid: 6 permutations of vertices 1, 2, 3 (Fig. 2.6), each of them may be followed by the swap of vertices 4 and 5. There are 6 orientation preserving and 6 orientation reversing symmetries (Fig. 2.6).
Trigonal bipyramid (left) and a stable single-strand double trace in the Schlegel diagram of the solid (right) corresponding to the grey entry in Table 2.1 having six symmetries and six anti-parallel dimers. Vertex-figures are depicted in red
2.4.3 Extension and Limits of Topological Single-Chain Polyhedra
We have proven that any polyhedron whose edges are composed of pairs of segments (or double traces) can be formed from a single strand, which is quite reassuring for the potentials of this type of molecular structures. The limit for the efficient assembly of structures may however be imposed by the order of formation of edges, which reflects the kinetics of folding molecules. We would like to exclude the folding pathways, where a certain formed segment needs to be unfolded before a new pair is formed, as this would likely represent a kinetic barrier. This can only be ensured if at least one end of the strand can remain free until the final structure is formed and therefore allow threading of the free end, which would not be possible if both ends already contain the structured segments. We can show that this is indeed possible for any type of the polyhedron, which is an additional support of mathematical topology for the design of complex modular polypeptide-based polyhedra.
2.5 Future Opportunities and Challenges in Designed Protein Origami
2.5.1 Expansion of the of Designed Polyhedral Shapes
Topological analysis of designed polyhedra composed of dimeric edges demonstrated that in principle any type of a polyhedron could be assembled from a single chain using concatenated dimerizing modules. Assembly from several polypeptide chains rather than from a single chain would makes this strategy even simpler, as demonstrated by DNA nanostructures that have been almost exclusively assembled from multiple, sometimes even hundreds of chains. Construction of more complex shapes will require an expanded orthogonal coiled-coil dimer set, which should deserve significant attention in the near future. Application of coiled-coil segments of different lengths additionally extends the accessible shapes of polyhedra. Natural coiled-coil segments differ in length from several up to 50 nm. Design of long orthogonal coiled-coil dimers is also lagging behind in comparison to typically 3–4 heptad segments reported so far. The problem in designing longer orthogonal coiled-coil dimers is that the difference in free energy gap between the correct and most stable misfolded structures decreases with the increasing sequence lengths.
2.5.2 In Vivo Folding of Protein Origami
The first designed protein tetrahedron formed aggregates in bacterial cells that were not correctly folded and had to be solubilized in the denaturing agents and slowly refolded by a dialysis from the denaturing solution or by the slow temperature annealing at low concentrations. This is similar to the large majority of DNA nanostructures that had to be self-assembled over an extended time. In vivo folding ability of designed protein origami structures would however be highly valuable, for its in vivo biological and medical role, as well as for the more efficient manufacturing of designed nanomaterials . The task of designing in vivo foldable sequences should include the topological considerations, in order to avoid formation of topological knots that may prevent folding. The importance of topological considerations has recently been demonstrated by the construction of a highly knotted single-chain DNA pyramid that folds quickly and efficiently by conforming to the “free end” design rule. By contrast, the folding of alternative designs that use the same segments but have a higher propensity to form topologically trapped intermediates was kinetically hindered [93]. Selection of the distribution of stability of building elements opens another challenge for modeling with the final goal of designing the folding pathway of modular topological proteins. This type of engineering is not feasible for the native proteins, due to their complex interplay of long range noncovalent interactions and cooperativity. The similarity between DNA- and polypeptide-based modular structures may allow translation of the design principles to engineer folding pathways from DNA to polypeptide-based modular structures. Although the design of the folding pathway of DNA nanostructures is still in its infancy, DNA may provide a very suitable prototyping material to design the folding pathway as the orthogonality and stability of DNA segments is much more reliable to predict than it is for polypeptide-based modules.
2.5.3 Regulation of the Protein Origami (Dis)Assembly
Interaction between the polypeptide strands of a coiled-coil dimer can be regulated by different physicochemical parameters, such as the temperature, chemical denaturants, pH, metal ions or presence of competing binding peptides. This could represent a range of different ways to regulate the assembly or disassembly of polypeptide nanostructures, providing in principle a broader range of adjustable parameters than for the nucleic acids. Regulated assembly/disassembly provides the possibility to regulate the stepwise assembly, encapsulation or release of the trapped molecules from the internal cavity of the polyhedra, which could be particularly useful for the drug delivery or for enzymatic reactions.
2.5.4 Functionalization of Designed Protein Origami
Besides the simplicity of the nucleic acid complementarity in comparison to the coiled-coil dimers the most important difference between DNA and protein origami is that polypeptides are composed of 20 residues with chemically very different properties, which enable formation of versatile catalytic and binding sites of proteins. The structure of designed coiled-coil dimers is to a large degree specified by 4 out of the 7 residues of the heptad repeats, leaving positions b, c and f for the introduction of residues with desired properties. This provides the possibility to introduce different functionalities into the polypeptide scaffold such as the binding or catalytic sites with numerous potential applications in areas including medicine, biotechnology and chemistry (Fig. 2.7).
2.5.5 Extension of Strategies of DNA Nanotechnology for Polypeptide-Based Nanostructures
DNA origami [94], based on a one very long strand and numerous shorter staple oligonucleotides, represented a great step ahead for the ability to make numerous different 2D or 3D nanoscale shapes. It is conceivable that a similar principle might be applied also for protein-based structures. Assembly of 2D or 3D shapes can also be achieved from a set of short DNA oligonucleotide building bricks, where each brick is comprised of 4 interacting segments [95]. Currently the main limitation preventing implementation of this strategy for designed polypeptides is the availability of the orthogonal coiled-coil segments. Toehold replacement of DNA-based nanostructures appeared as a very powerful strategy for the dynamic assemblies, allowing tuning kinetics of assemblies and construction of molecular machines, such as different molecular walkers and implementation of different logical functions in complex solutions of nucleotide oligomers in the solution [96]. Key feature of the toehold strategy is to replace one strand in the dimer with another strand that has higher stability due to the longer region of complementarity. This strategy is useful only when the dissociation rates occur at much slower time scale than the intended time scale for the displacement, typically within at least minutes, which means typically subnanomolar affinity. Toehold displacement has not been demonstrated yet in coiled-coil dimers, although there are no fundamental limitations that would prevent the same approach, given the availability of appropriate designed (or natural) coiled-coil building blocks .
In summary, the technology of designed protein origami or designed topological modular protein folds opens an exciting range of possibilities of designing new protein folds.
References
Turoverov KK, Kuznetsova IM, Uversky VN (2010) The protein kingdom extended: ordered and intrinsically disordered proteins, their folding, supramolecular complex formation, and aggregation. Prog Biophys Mol Biol 102:73–84. doi:10.1016/j.pbiomolbio.2010.01.003
Fersht AR (2008) From the first protein structures to our current knowledge of protein folding: delights and scepticisms. Nat Rev Mol Cell Biol 9:650–654. doi:10.1038/nrm2446
Dill KA, MacCallum JL (2012) The protein-folding problem, 50 years on. Science (80-) 338:1042–1046. doi:10.1126/science.1219021
Dill KA, Ozkan SB, Shell MS, Weikl TR (2008) The protein folding problem. Annu Rev Biophys 37:289–316. doi:10.1146/annurev.biophys.37.092707.153558
Chandler D (2005) Interfaces and the driving force of hydrophobic assembly. Nature 437:640–647. doi:10.1038/nature04162
Berne BJ, Weeks JD, Zhou R (2009) Dewetting and hydrophobic interaction in physical and biological systems. Annu Rev Phys Chem 60:85–103. doi:10.1146/annurev.physchem.58.032806.104445
Pauling L, Corey RB, Branson HR (1951) The structure of proteins: two hydrogen-bonded helical configurations of the polypeptide chain. Proc Natl Acad Sci U S A 37:205–211
Ramakrishnan C, Ramachandran GN (1965) Stereochemical criteria for polypeptide and protein chain conformations. Biophys J 5:909–933. doi:10.1016/S0006-3495(65)86759-5
Kumar S, Nussinov R (2002) Close-range electrostatic interactions in proteins. ChemBioChem 3:604–617.doi:10.1002/1439-7633(20020703)3:7<604::AID-CBIC604>3.0.CO;2-X
Makhatadze GI, Privalov PL (1995) Energetics of protein structure. Adv Protein Chem 47:307–425
Murphy KP (2001) Stabilization of protein structure. Methods Mol Biol 168:1–16. doi:10.1385/1-59259-193-0:001
Baldwin RL (2007) Energetics of protein folding. J Mol Biol 371:283–301. doi:http://dx.doi.org/10.1016/j.jmb.2007.05.078
Guo W, Shea J-E, Berry RS (2006) The physics of the interactions governing folding and association of proteins. Ann NY Acad Sci 1066:34–53. doi:10.1196/annals.1363.025
Berman HM, Westbrook J, Feng Z et al (2000) The protein data bank. Nucleic Acids Res 28:235–242. doi:10.1093/nar/28.1.235
Sindelar CV, Hendsch ZS, Tidor B (1998) Effects of salt bridges on protein structure and design. Protein Sci 7:1898–1914. doi:10.1002/pro.5560070906
Sinha N, Smith-Gill SJ (2002) Electrostatics in protein binding and function. Curr Protein Pept Sci 3:601–614
Plaxco KW, Simons KT, Baker D (1998) Contact order, transition state placement and the refolding rates of single domain proteins. J Mol Biol 277:985–994. doi:http://dx.doi.org/10.1006/jmbi.1998.1645
Ivankov DN, Garbuzynskiy SO, Alm E et al (2003) Contact order revisited: influence of protein size on the folding rate. Protein Sci 12:2057–2062. doi:10.1110/ps.0302503
Robertson AD, Murphy KP (1997) Protein structure and the energetics of protein stability. Chem Rev 97:1251–1267
Ghosh K, Dill KA (2009) Computing protein stabilities from their chain lengths. Proc Natl Acad Sci U S A 106:10649–10654. doi:10.1073/pnas.0903995106
Božič S, Doles T, Gradišar H, Jerala R (2013) New designed protein assemblies. Curr Opin Chem Biol 17:940–945. doi:10.1016/j.cbpa.2013.10.014
King NP, Lai Y-T (2013) Practical approaches to designing novel protein assemblies. Curr Opin Struct Biol 23:632–638. doi:10.1016/j.sbi.2013.06.002
Salgado EN, Ambroggio XI, Brodin JD et al (2010) Metal templated design of protein interfaces. Proc Natl Acad Sci 107:1827–1832. doi:10.1073/pnas.0906852107
Huard DJE, Kane KM, Tezcan FA (2013) Re-engineering protein interfaces yields copper-inducible ferritin cage assembly. Nat Chem Biol 9:169–176. doi:10.1038/nchembio.1163
Brodin JD, Ambroggio XI, Tang C et al (2012) Metal-directed, chemically tunable assembly of one-, two- and three-dimensional crystalline protein arrays. Nat Chem 4:375–382. doi:10.1038/nchem.1290
Anzini P, Xu C, Hughes S et al (2013) Controlling self-assembly of a peptide-based material via metal-Ion induced registry shift. J Am Chem Soc 135:10278–10281. doi:10.1021/ja404677c
Fletcher JM, Harniman RL, Barnes FRH, et al (2013) Self-assembling cages from coiled-coil peptide modules. Science (80-) 340:595–599. doi:10.1126/science.1233936
Padilla JE, Colovos C, Yeates TO (2001) Nanohedra: using symmetry to design self assembling protein cages, layers, crystals, and filaments. Proc Natl Acad Sci 98:2217–2221. doi:10.1073/pnas.041614998
Lai Y-T, Tsai K-L, Sawaya MR et al (2013) Structure and flexibility of nanoscale protein cages designed by symmetric self-assembly. J Am Chem Soc 135:7738–7743. doi:10.1021/ja402277f
Sinclair JC, Davies KM, Vénien-Bryan C, Noble MEM (2011) Generation of protein lattices by fusing proteins with matching rotational symmetry. Nat Nanotechnol 6:558–562. doi:10.1038/nnano.2011.122
King NP, Sheffler W, Sawaya MR et al (2012) Computational design of self-assembling protein nanomaterials with atomic level accuracy. Science (80-) 336:1171–1174. doi:10.1126/science.1219364
Leaver-Fay A, Tyka M, Lewis SM et al (2011) Rosetta3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol 487:545–574. doi:10.1016/B978-0-12-381270-4.00019-6
King NP, Bale JB, Sheffler W et al (2014) Accurate design of co-assembling multi-component protein nanomaterials. Nature 510:103–108. doi:10.1038/nature13404
Hsia Y, Bale JB, Gonen S et al (2016) Design of a hyperstable 60-subunit protein icosahedron. Nature 535:136–139. doi:10.1038/nature18010
Lanci CJ, MacDermaid CM, Kang S et al (2012) Computational design of a protein crystal. Proc Natl Acad Sci 109:7304–7309. doi:10.1073/pnas.1112595109
Barrick D, Ferreiro DU, Komives EA (2008) Folding landscapes of ankyrin repeat proteins: experiments meet theory. Curr Opin Struct Biol 18:27–34. doi:10.1016/j.sbi.2007.12.004
Cortajarena AL, Mochrie SGJ, Regan L (2011) Modulating repeat protein stability: the effect of individual helix stability on the collective behavior of the ensemble. Prot Sci 20:1042–1047. doi:10.1002/pro.638
Parmeggiani F, Huang P-S, Vorobiev S et al (2015) A general computational approach for repeat protein design. J Mol Biol 427:563–575. doi:10.1016/j.jmb.2014.11.005
De la Paz ML, Goldie K, Zurdo J et al (2002) De novo designed peptide-based amyloid fibrils. Proc Natl Acad Sci U S A 99:16052–16057. doi:10.1073/pnas.252340199
Haines LA, Rajagopal K, Ozbas B et al (2005) Light-activated hydrogel formation via the triggered folding and self-assembly of a designed peptide. J Am Chem Soc 127:17025–17029. doi:10.1021/ja054719o
Yanlian Y, Ulung K, Xiumei W et al (2009) Designer self-assembling peptide nanomaterials. Nano Today 4:193–210. doi:10.1016/j.nantod.2009.02.009
Saaem I, LaBean TH (2013) Overview of DNA origami for molecular self-assembly. WIREs Nanomed Nanobiotechnol 5:150–162. doi:10.1002/wnan.1204
Walshaw J, Woolfson DN (2001) SOCKET: a program for identifying and analysing coiled-coil motifs within protein structures. J Mol Biol 307:1427–1450. doi:10.1006/jmbi.2001.4545
Crick FHC (1953) The packing of α-helices: simple coiled-coils. Acta Crystallogr 6:689–697. doi:10.1107/S0365110X53001964
Pauling L, Corey RB (1953) Compound helical configurations of polypeptide chains: structure of proteins of the α-keratin type. Nature 171:59–61. doi:10.1038/171059a0
Lupas A (1996) Coiled coils: new structures and new functions. Trends Biochem Sci 21:375–382. doi:10.1016/S0968-0004(96)10052-9
Burkhard P, Stetefeld J, Strelkov SV (2001) Coiled coils: a highly versatile protein folding motif. Trends Cell Biol 11:82–88. doi:10.1016/S0962-8924(00)01898-5
Lupas AN, Gruber M (2005) The structure of α-helical coiled coils. In: Parry DAD, John MS (eds). Academic Press, San Diego, CA, pp 37–38
Gruber M, Lupas AN (2003) Historical review: another 50th anniversary – new periodicities in coiled coils. Trends Biochem Sci 28:679–685. doi:10.1016/j.tibs.2003.10.008
Rose A, Schraegle SJ, Stahlberg EA, Meier I (2005) Coiled-coil protein composition of 22 proteomes – differences and common themes in subcellular infrastructure and traffic control. BMC Evol Biol 5:66. doi:10.1186/1471-2148-5-66
Rose A, Meier I (2004) Scaffolds, levers, rods and springs: diverse cellular functions of long coiled-coil proteins. Cell Mol Life Sci C 61:1996–2009. doi:10.1007/s00018-004-4039-6
Esteban MR, Giovinazzo G, de la Hera A, Goday C (1998) PUMA1: a novel protein that associates with the centrosomes, spindle and centromeres in the nematode parascaris. J Cell Sci 111:723–735
O’Shea EK, Klemm JD, Kim PS, Alber T (1991) X-ray structure of the GCN4 leucine zipper, a two-stranded, parallel coiled coil. Science (80-) 254:539–544
Steinmetz MO, Jelesarov I, Matousek WM et al (2007) Molecular basis of coiled-coil formation. Proc Natl Acad Sci 104:7062–7067. doi:10.1073/pnas.0700321104
Grigoryan G, Reinke AW, Keating AE (2009) Design of protein-interaction specificity gives selective bZIP-binding peptides. Nature 458:859–864. doi:10.1038/nature07885
Grigoryan G, Keating AE (2008) Structural specificity in coiled-coil interactions. Curr Opin Struct Biol 18:477–483. doi:10.1016/j.sbi.2008.04.008
Sutton RB, Fasshauer D, Jahn R, Brunger AT (1998) Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 Å resolution. Nature 395:347–353. doi:10.1038/26412
Bullough PA, Hughson FM, Skehel JJ, Wiley DC (1994) Structure of influenza haemagglutinin at the pH of membrane fusion. Nature 371:37–43. doi:10.1038/371037a0
Dutta K, Alexandrov A, Huang H, Pascal SM (2001) pH-induced folding of an apoptotic coiled coil. Protein Sci 10:2531–2540
Liang W, Warrick HM, Spudich JA (1999) A structural model for phosphorylation control of dictyostelium myosin II thick filament assembly. J Cell Biol 147:1039–1048. doi:10.1083/jcb.147.5.1039
Farah CS, Reinach FC (1999) Regulatory properties of recombinant tropomyosins containing 5-hydroxytryptophan: Ca2 + -binding to troponin results in a conformational change in a region of tropomyosin outside the troponin binding site. Biochemistry 38:10543–10551. doi:10.1021/bi982813u
Woolfson DN (2005) The design of coiled-coil structures and assemblies. In: Parry DAD, John MS (eds). Academic Press, San Diego, CA, pp 79–112
Hodges RS, Saund AK, Chong PC et al (1981) Synthetic model for two-stranded alpha-helical coiled-coils. Design, synthesis, and characterization of an 86-residue analog of tropomyosin. J Biol Chem 256:1214–1224
O’Shea EK, Lumb KJ, Kim PS (1993) Peptide “Velcro”: design of a heterodimeric coiled coil. Curr Biol 3:658–667. doi:10.1016/0960-9822(93)90063-T
Oakley MG, Kim PS (1998) A buried polar interaction can direct the relative orientation of helices in a coiled coil. Biochemistry 37:12603–12610. doi:10.1021/bi981269m
Harbury PB, Zhang T, Kim PS, Alber T (1993) A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants. Science 262:1401–1407
Nautiyal S, Woolfson DN, King DS, Alber T (1995) A designed heterotrimeric coiled coil. Biochemistry 34:11645–11651. doi:10.1021/bi00037a001
Fairman R, Chao HG, Lavoie TB et al (1996) Design of heterotetrameric coiled coils: evidence for increased stabilization by Glu(−)-Lys(+) ion pair interactions. Biochemistry 35:2824–2829. doi:10.1021/bi952784c
Liu J, Zheng Q, Deng Y et al (2006) A seven-helix coiled coil. Proc Natl Acad Sci U S A 103:15457–15462. doi:10.1073/pnas.0604871103
Testa OD, Moutevelis E, Woolfson DN (2009) CC+: a relational database of coiled-coil structures. Nucleic Acids Res 37:D315–D322. doi:10.1093/nar/gkn675
Moutevelis E, Woolfson DN (2009) A periodic table of coiled-coil protein structures. J Mol Biol 385:726–732. doi:10.1016/j.jmb.2008.11.028
Naik RR, Kirkpatrick SM, Stone MO (2001) The thermostability of an alpha-helical coiled-coil protein and its potential use in sensor applications. Biosens Bioelectron 16:1051–1057. doi:10.1016/j.ijfoodmicro.2010.12.030
Sharma VA, Logan J, King DS et al (1998) Sequence-based design of a peptide probe for the APC tumor suppressor protein. Curr Biol 8:823–830. doi:10.1016/S0960-9822(98)70324-0
Reinke AW, Grant RA, Keating AE (2010) A synthetic coiled-coil interactome provides heterospecific modules for molecular engineering. J Am Chem Soc 132:6025–6031. doi:10.1021/ja907617a
Havranek JJ, Harbury PB (2003) Automated design of specificity in molecular recognition. Nat Struct Mol Biol 10:45–52. doi:10.1038/nsb877
Bromley EHC, Sessions RB, Thomson AR, Woolfson DN (2008) Designed α-helical tectons for constructing multicomponent synthetic biological systems. J Am Chem Soc 131:928–930. doi:10.1021/ja804231a
Gradišar H, Jerala R (2011) De novo design of orthogonal peptide pairs forming parallel coiled-coil heterodimers. J Pept Sci 17:100–106. doi:10.1002/psc.1331
Negron C, Keating AE (2014) A set of computationally designed orthogonal antiparallel homodimers that expands the synthetic coiled-coil toolkit. J Am Chem Soc 136:16544–16556. doi:10.1021/ja507847t
Armstrong CT, Vincent TL, Green PJ, Woolfson DN (2011) SCORER 2.0: an algorithm for distinguishing parallel dimeric and trimeric coiled-coil sequences. Bioinformatics 27:1908–1914. doi:10.1093/bioinformatics/btr299
Mahrenholz CC, Abfalter IG, Bodenhofer U et al (2011) Complex networks govern coiled-coil oligomerization – predicting and profiling by means of a machine learning approach. Mol Cell Proteomics 10:M110.004994. doi:10.1074/mcp.M110.004994
Li C, Wang X-F, Chen Z et al (2015) Computational characterization of parallel dimeric and trimeric coiled-coils using effective amino acid indices. Mol Biosyst 11:354–360. doi:10.1039/C4MB00569D
Trigg J, Gutwin K, Keating AE, Berger B (2011) Multicoil2: predicting coiled coils and their oligomerization states from sequence in the twilight zone. PLoS One 6:e23519. doi:10.1371/journal.pone.0023519
Vincent TL, Green PJ, Woolfson DN (2013) LOGICOIL—multi-state prediction of coiled-coil oligomeric state. Bioinformatics 29:69–76. doi:10.1093/bioinformatics/bts648
Hagemann UB, Mason JM, Müller KM, Arndt KM (2008) Selectional and mutational scope of peptides sequestering the Jun–Fos coiled-coil domain. J Mol Biol 381:73–88. doi:10.1016/j.jmb.2008.04.030
Sowdhamini R, Alva V, Syamala Devi D (2008) COILCHECK: an interactive server for the analysis of interface regions in coiled coils. Protein Pept Lett 15:33–38. doi:10.2174/092986608783330314
Wood CW, Bruning M, Ibarra AÁ et al (2014) CCBuilder: an interactive web-based tool for building, designing and assessing coiled-coil protein assemblies. Bioinformatics 30:3029–3035. doi:10.1093/bioinformatics/btu502
Strelkov SV, Burkhard P (2002) Analysis of α-helical coiled coils with the program TWISTER reveals a structural mechanism for stutter compensation. J Struct Biol 137:54–64. doi:10.1006/jsbi.2002.4454
Grigoryan G, DeGrado WF (2011) Probing designability via a generalized model of helical bundle geometry. J Mol Biol 405:1079–1100. doi:10.1016/j.jmb.2010.08.058
Gurnon DG, Whitaker JA, Oakley MG (2003) Design and characterization of a homodimeric antiparallel coiled coil. J Am Chem Soc 125:7518–7519. doi:10.1021/ja0357590
Taylor CM, Keating AE (2005) Orientation and oligomerization specificity of the Bcr coiled-coil oligomerization domain†. Biochemistry 44:16246–16256. doi:10.1021/bi051493t
Gradišar H, Božič S, Doles T et al (2013) Design of a single-chain polypeptide tetrahedron assembled from coiled-coil segments. Nat Chem Biol 9:362–366. doi:10.1038/nchembio.1248
Fijavž G, Pisanski T, Rus J (2015) Strong traces model of self-assembly polypeptide structures. MATCH Commun Math Comput Chem 71:199–212
Kočar V, Schreck JS, Čeru S, Gradišar H, Bašić N, Pisanski T, Doye JP, Jerala R (2016) Design principles for rapid folding of knotted DNA nanostructures. Nat Commun 7:10803. doi:10.1038/ncomms10803
Rothemund PWK (2006) Folding DNA to create nanoscale shapes and patterns. Nature 440:297–302. doi:10.1038/nature04586
Ke Y, Ong LL, Shih WM, Yin P (2012) Three-dimensional structures self-assembled from DNA bricks. Science 338:1177–1183. doi:10.1126/science.1227268
Zhang DY, Winfree E (2009) Control of DNA strand displacement kinetics using toehold exchange. J Am Chem Soc 131:17303–17314. doi:10.1021/ja906987s
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Drobnak, I., Ljubetič, A., Gradišar, H., Pisanski, T., Jerala, R. (2016). Designed Protein Origami. In: Cortajarena, A., Grove, T. (eds) Protein-based Engineered Nanostructures. Advances in Experimental Medicine and Biology, vol 940. Springer, Cham. https://doi.org/10.1007/978-3-319-39196-0_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-39196-0_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-39194-6
Online ISBN: 978-3-319-39196-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)