Abstract
DNA origami is a robust assembly technique that folds a single-stranded DNA template into a target structure by annealing it with hundreds of short ‘staple’ strands1,2,3,4. Its guiding design principle is that the target structure is the single most stable configuration5. The folding transition is cooperative4,6,7 and, as in the case of proteins, is governed by information encoded in the polymer sequence8,9,10,11. A typical origami folds primarily into the desired shape, but misfolded structures can kinetically trap the system and reduce the yield2. Although adjusting assembly conditions2,12 or following empirical design rules12,13 can improve yield, well-folded origami often need to be separated from misfolded structures2,3,14,15,16. The problem could in principle be avoided if assembly pathway and kinetics were fully understood and then rationally optimized. To this end, here we present a DNA origami system with the unusual property of being able to form a small set of distinguishable and well-folded shapes that represent discrete and approximately degenerate energy minima in a vast folding landscape, thus allowing us to probe the assembly process. The obtained high yield of well-folded origami structures confirms the existence of efficient folding pathways, while the shape distribution provides information about individual trajectories through the folding landscape. We find that, similarly to protein folding, the assembly of DNA origami is highly cooperative; that reversible bond formation is important in recovering from transient misfoldings; and that the early formation of long-range connections can very effectively enforce particular folds. We use these insights to inform the design of the system so as to steer assembly towards desired structures. Expanding the rational design process to include the assembly pathway should thus enable more reproducible synthesis, particularly when targeting more complex structures. We anticipate that this expansion will be essential if DNA origami is to continue its rapid development1,2,3,17,18,19 and become a reliable manufacturing technology20.
Similar content being viewed by others
Main
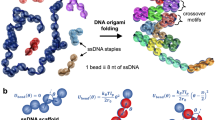
This study is based on a simplified version of the archetypal origami tile1 and, in particular, on the distribution of observed folds of a ‘dimer’ variant which contains two copies of the template sequence in head-to-tail repeat. The ‘monomer’ tile (Fig. 1) is created by annealing a 2,646-nucleotide (nt) circular template with 90 staples, each designed to hybridize to one or more 15- or 16-base domains of the template. 76 of the staples mediate interactions between pairs of non-contiguous template domains, as follows: 66 U-shaped ‘body’ staples form short-range contacts between domains that are relatively close in the primary sequence of the template; and 5 pairs of ‘seam’ staples form long-range contacts, bridging between positions where the template folds back on itself to form a central seam1. Unlike the interactions between amino acid residues that stabilize a protein, staples mediate interactions between template domains that are highly specific: each staple can be considered to bind stably only to complementary domains of the template. The designed fold of the monomer tile corresponds to an absolute minimum in the free energy landscape. This origami folds with high yield to form discrete rectangular tiles of approximately 80 nm × 40 nm (Fig. 1c); approximately 80% of tiles appear to be well folded.
a, 66 body staples (blue) and 5 pairs of seam staples (brown) each hybridize to two non-contiguous domains of the circular template. Edge staples (grey) fill gaps at the top and bottom of the structure. Hybridization of body and seam staples pins the corresponding domains of the template together, determining the unique stable, rectangular fold of this simplified origami tile as indicated in b. c, Atomic force micrograph of the monomer tile (scale bar, 50 nm).
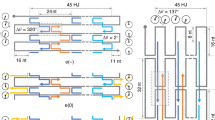
The ‘dimer’ template is also circular. It contains two identical copies of the monomer joined head-to-tail and can therefore bind two copies of each staple (Fig. 2). Each pair of body and seam staples can bind in one of two configurations (Fig. 2a) to form either an internal link within each copy of the monomer sequence or a pair of cross-links between the two copies. The total number of possible domain pairings is 276 ≈ 1023. Although many of these configurations are sterically inaccessible, it is clear that the result of reducing the specificity of staple binding is that, as in the case of protein folding, the number of possible states of the system is overwhelmingly greater than the number of well-folded structures. However, in contrast to proteins (and to conventional origami structures) there is more than one ‘well-folded’ state (Fig. 2): not one but a handful of well-folded states occupy discrete energy minima in a vast configurational landscape. Remarkably, when the dimer origami is annealed by cooling from 95 °C, a small set of well-folded shapes are formed with good yield: each consists of a pair of rectangular tiles attached on one edge (Fig. 2b, c). The probability of finding well-folded structures by random search of configuration space is negligible8, therefore efficient folding pathways must exist21,22. As in protein folding, assembly is constrained such that the system is highly likely to discover free-energy minima that correspond to well-formed final states.
a, Left, the base sequence of the green section of the template is the same as that of the pink section, so the dimer template can fully hybridize to two copies of each staple. Arrows indicating staple binding and unbinding transitions are annotated with reaction rates used in a model of assembly. Two identical staples can bind to the template in one of two configurations, binding together pairs of domains within each section or connecting domains in different sections. This gives a vast number of possible configurations, a handful of which are well-folded (shown right, and defined in Fig. 3 and Extended Data Figs 1 and 2). Ordered folds of the dimer template comprise two linked rectangular tiles with a characteristic offset on the long or short edges, for example, b or c, respectively (in each case, an AFM image with a scale bar of 50 nm is shown alongside a schematic of the fold).
The dimer origami tile has 22 template routings that correspond to well-folded configurations in which all staple binding sites are occupied and in which the tile is expected to be planar and unstrained. These give 6 unique shapes, each with a characteristic offset between two linked rectangular components which have essentially the same structure as the monomer tile (Fig. 3a–c and Extended Data Fig. 1). These shapes can be grouped into classes according to the contacts made by the seam staples: fold m:n has m pairs of seam staples that connect domains within each half of the template and n pairs of seam staples that form connections between domains in opposite halves (Fig. 3a–c). Folds m:n and n:m are related by symmetry and are therefore not distinguished in our experiments or analysis (Extended Data Fig. 1b). A set of non-planar folds adds a seventh shape to the six defined above and a further 52 template routings (Fig. 3d and Extended Data Fig. 2). Fink and Ball23 have estimated the maximum number of distinct, compact configurations that can be encoded into a single polymer sequence: for a polymer of 168 unique domain types on a square lattice23,24 the theoretical limit is 13. A major factor in allowing the large number of folds in our system is the extensive re-use of structural motifs within distinct folds, a possibility not considered by Fink and Ball.
a–c, Folds of the dimer origami tile can be classified by the pattern of interactions mediated by the seam staples: these contacts are shown schematically in diagrams in which the dimer template is represented as a circle. Fold m:n has m seam contacts between domains within each copy of the monomer sequence and n seam contacts that form connections between the two copies (that is, connecting template domains of different colour). The fold 5:0 can be further divided into shapes that differ in the offset along the long edge of the two tiles (5:0i, 5:0ii and so on: c). d, The set of legal folds allowed by the model includes the configurations shown in a–c and an additional set of non-planar configurations (NP), one of which is shown (see Extended Data Fig. 2 for the complete set). e, Tiles observed in folding experiments can be classified according to the fractional offset along the short or long edge of the tile (w/W and l/L respectively). f, Seven unique shapes corresponding to well-folded configurations (see also Extended Data Fig. 1). g, Gallery of shapes observed by AFM in a typical experiment with measured fractional offsets (each image is 300 × 300 nm). A bin size of 0.1 is used in histograms of fitted fractional offsets (Fig. 4).
Atomic force microscopy (AFM) enables us to distinguish different configurations of the template and this provides a unique opportunity to study folding pathways. Samples of annealed origami were imaged by AFM. Most observed shapes are consistent with the classification scheme shown in Fig. 3, and the outlines of 44% of objects identified as candidate dimer tiles were successfully fitted to measure the offset between the two component monomer tiles (Fig. 3e and Extended Data Fig. 3).
The distribution of tile shapes was compared to predictions made using a Markov chain model of folding in which each transition corresponds to binding or unbinding of a single staple domain (Fig. 2, Methods section ‘Folding model’). An unbound staple at concentration c binds to the template with a rate k+c (where k+ = 106 M−1 s−1, ref. 25). After one half of a staple has bound, the second half can bind with a rate (k+ceff) that depends on its effective concentration, ceff, at the corresponding template domain. The effective concentration depends on the proximity of the template domain which, in turn, depends on the contacts between template domains already established by hybridization of other staples. We expect folding to be dominated by short-range interactions because staples are more likely to connect two template domains that are spatially close, either because they are closely spaced along the template or because the previous binding of other staples is holding them together. To determine the effective concentration, the shortest path through the part-assembled origami that connects the complementary template and staple domains is identified. This connection is modelled as a heterogeneous freely jointed chain with double-stranded (ds) and single-stranded (ss) DNA components. The effective concentration of the part-bound staple at the complementary template domain is related to the probability that the ends of the chain lie spontaneously within a (short) interaction range. Unbinding of a staple domain is treated as a two-state transition, with a configuration-independent rate: k− = k+ exp{ΔG0,duplex/RT}(1 M) where ΔG0,duplex is the change in standard free energy on forming the duplex at standard concentrations of 1 M. In order to represent steric constraints on folding, the state space of the model is restricted to patterns of staple binding in which each segment of the partially folded origami occurs in one of a set of pre-defined, well-ordered folds.
The histograms in Fig. 4 show distributions of offset values, measured by fitting AFM data (Extended Data Figs 3 and 4), and the corresponding distributions between the discrete shapes shown in Fig. 3 that are predicted by the model. Figure 4a corresponds to the staple set described above (see Fig. 1): structures with each of the seam configurations 5:0, 4:1 and 3:2 are observed. The model suggests that the folding pathway depends on competition between body and seam staples. If local interactions mediated by body staples were to form first and dominate the outcome, the system would prefer the 5:0i fold (see Fig. 3f for nomenclature) in which all body staples are bound to two domains that are as close as possible along the template. In this fold, no staples link the two halves of the template. However, strong seam connections that are inserted early in the folding pathway favour a more uniform distribution between all possible seam configurations: for example, once the part-folded structure 1:1 has formed, the 5:0 fold is inaccessible unless at least one seam connection is broken (Extended Data Fig. 5). With the staple set shown in Fig. 4a, each seam contact is bridged by two staples. The cooperative binding of seam staple pairs offsets the increased entropic cost of forming long-range contacts, with the result that seam staples are incorporated at a similar temperature to body staples in both model (Fig. 4a) and experiment (Extended Data Fig. 6). Consequently the model predicts that all seam configurations should be observed, consistent with experimental observations.
The reference staple set (a) folds to give a distribution of shapes that are characterized by the fractional offset between the two component tiles along the long or short edge. Modifications to the reference staple set (b–e) were designed to fold into specific target shapes. Left-hand panels show the staple configurations and the seam-staple contacts in the target structures. The top left rectangle of each target shape is used to highlight modified staples in red. The distance between the two template domains linked by a staple depends on fold: in the bottom right rectangle, staples are grouped and coloured according to the distance spanned (see key in a: short-range body staples are blue, seam staples are brown to yellow; lighter shades indicate larger distances). Graphs in the central panels show the calculated fraction of contacts formed for each staple group (whether or not as part of the target structure) as a function of temperature during assembly. The right-hand panels show the distribution of shapes predicted and observed for each set: a histogram representing the continuous distribution of fitted offset values is plotted above the distribution between discrete shapes predicted by the model (indicated by silhouettes). The number of fitted shapes, N, and the yield (shapes fitted as a percentage of candidate structures identified) are shown in the top right corner of each histogram.
We predict that the folding pathway can be changed by altering the relative strengths of short- and long-range interactions. Breaking in half one of each pair of seam staples (Fig. 4b), so that the pairs no longer bind cooperatively, weakens these long-range bonds, causing them to form later in the folding pathway (Fig. 4b, central panel and Extended Data Fig. 6) and to break and reform in alternative configurations more frequently (Extended Data Fig. 7). With weakened long-range interactions, we expect folding to be governed primarily by local interactions. The model predicts that the distribution of shapes is shifted strongly towards the 5:0i fold, in which all body staples span the smallest possible distances along the template (the same distances as in the monomer tile), and this is confirmed by experiment (Fig. 4b). The thermodynamic cost of breaking every other seam staple is approximately equal in each well-folded state and therefore this change should not affect their equilibrium populations. We have changed the distribution between folds not by changing the relative stability of the final states but by deliberately controlling the stabilities of crucial intermediate states, thus shaping the folding pathway.
The importance of stable, long-range interactions in determining the folding pathway is revealed by the evolving correlations between seam staples in the model. Characteristic patterns of correlation can be used to predict the final fold even before seam staple occupancy has reached 50% (Extended Data Figs 8 and 9).
The influence of seam staples on folding is similar to that of disulphide bonds in Anfinsen’s experiment on protein folding9. If long-range bonds are allowed to form first and, effectively, irreversibly, then folding is kinetically trapped. If they are weakened and permitted to rearrange then folding can be controlled by weaker short-range interactions.
Figure 4c shows an alternative staple set incorporating extended staples that form particularly strong short-range connections and therefore bind to the template early in the folding process (Fig. 4c, central panel). Without interference from other staples, these contacts are most likely to form between the pairs of template domains with the smallest separation along the template. These preferred contacts occur in the 3:2 and 5:0 folds but not the 4:1 fold (in the 4:1 fold, one extended staple forms a long-range contact between the two halves of the template). Experimental results confirm the model prediction that the 4:1 fold is strongly suppressed (Fig. 4c). As with the broken seam staples (Fig. 4b), this modification guides the folding pathway without imposing an energetic penalty on alternative folds.
We can control the fold of the dimer very effectively by engineering both the folding pathway and the stability of the chosen target structure. The 3:2 configuration can be favoured by weakening the original seams (as in Fig. 4b) and adding new seam staples that bridge between the monomer tiles without distortion only in the 3:2 configuration (Fig. 4d). This modification guides folding by increasing the stability of 3:2 relative to other folds. Similarly, a long staple in the bottom right corner of the monomer tile (Fig. 4e) biases folding towards the 5:0iv shape by decreasing the stability of other folds, which would require introduction of a sharp bend within the long staple. (The model does not include any penalty for bending and so fails to predict the engineered bias in this case.)
By showing that an origami tile with a duplicated template can be annealed to produce a high yield of well-folded structures from among ∼1023 disordered alternative staple configurations, our results confirm that, as in the case of proteins, efficient folding pathways exist and that folding is highly cooperative. We infer that the folding of all DNA origami is shaped by similar pathways. Manipulation of the folding pathway validates our simple folding model, which successfully predicts the dominant folding pathways observed in experiments. We anticipate that this tool will prove more generally useful, to establish how to change the relative strengths of local and long-range staple interactions to rationally steer the folding pathway towards desired target structures.
Methods
Experimental methods
Plasmid pUC19 cut with HindIII and EcoRI was amplified by PCR with the primers TGACCTAATCCTCAGCAATTCACTGGCCGTCGTTTTACAA and ACGGACGCGCTGAGGAGCTTGGCGTAATCATGGTCATAG in order to trim the template to the desired length and introduce a unique BbvCI site. The PCR product was cut with BbvCI and ligated to generate pKD1 (2,646 bp). A typical monomer plasmid preparation contains a small amount (∼1%) of plasmid dimer. The dimer plasmid was obtained by nicking a monomer plasmid preparation with Nt.BbvCI (in order to resolve monomer and dimer more easily), purifying the nicked dimer band from a 0.7% TAE agarose gel, then transforming the purified nicked dimer into the recA host DH5α. The template sequence is given in Supplementary Information.
Single-stranded template was prepared by sequential reaction of either monomer or dimer pKD1 with Nt.BspQI at 50 °C and ExoIII at 37 °C to digest the non-template strand and leave a covalently-closed single-stranded template26. Enzymes were removed by phenol:chloroform extraction and the template was recovered by ethanol precipitation; its concentration was then determined by measuring ultraviolet absorbance at 260 nm.
DNA origami was designed using caDNAno27 and was assembled by cooling template at 4–10 nM with a ∼10-fold excess of staples from 95 °C to 25 °C at 1 °C per minute in a buffer containing 40 mM Tris-acetate (pH 8.3) and 12.5 mM magnesium acetate. Excess staples were removed using an S-300 size exclusion spin column28. Staple sequences for the standard design and variations are given in Supplementary Information.
Atomic force microscopy images were acquired using either an Agilent 5500 AFM with Olympus TR400-PSA probes (Figs 1, 2, 3, 4a) or a Veeco Dimension 3100 with Bruker SNL-10 probes (all other figures). A few microlitres of sample were added to freshly cleaved mica and the sample was imaged in tapping mode in an imaging buffer containing 12.5 mM magnesium acetate, 4 mM NiCl2, 1 mM EDTA and 40 mM Tris-acetate pH 8.0–8.3 (the imaging buffer for Fig. 1c lacked NiCl2, the imaging buffer for Fig. 2c lacked EDTA).
Folding model
Our domain-level description of origami assembly is intended to reproduce some aspects of cooperativity. In particular, it accounts for the increase in incorporation rate for a staple when its target domains on the template are held more closely together as a result of the earlier binding of other staples. This effect is most noticeable in the seam where the binding of the first of a pair of seam staples greatly accelerates, and is stabilized by, the binding of the second. The model incorporates a physically reasonable approximation of the entropic cost of closing loops by staple binding, but is far from a complete description of the physics of assembly. It is useful in guiding, and providing insights into, the effects of significant changes to the origami design.
We model the folding of an isolated template in the presence of an excess of staples as an inhomogeneous continuous-time Markov chain. Each transition between states corresponds to the binding or unbinding of a single staple domain. Transition rates between two states are chosen according to an estimate of the free energy difference between the two, in a manner that would reproduce the correct Boltzmann distribution if this free energy difference were calculated exactly. The temperature is updated once per second of simulated time which allows us to use an event-based Gillespie simulation algorithm29 with transition rates fixed over one second intervals. Data on folding processes are collected by simulating multiple folding trajectories (typically 1,600 per experiment).
Subsequent sections contain more detailed descriptions of the folding model.
State space
We consider the possible configurations of staples hybridized to the template with domain-level resolution: a domain is either fully hybridized or unhybridized. A staple is called half-bound if only one of its two domains is hybridized to the template and fully bound if both domains are bound. In the model, a staple domain can only hybridize to the complementary template domain; we ignore weaker interactions that result from inevitable partial sequence complementary between other pairs of domains.
For each type of two-domain staple (and the corresponding two pairs of complementary template domains) there are 34 distinct patterns of domain binding (states) with between zero and four copies of the staple bound to the dimer template. One is an empty state. When one staple is bound to the template there are four states in which the staple is half-bound and four states in which the staple is fully bound. When two staples are bound to the template there are six states in which both staples are half-bound, eight states with one half-bound and one fully bound staple, and two states with two fully bound staples. There are four states with three half-bound staples and another four states with one fully bound and two half-bound staples. Finally there is the possibility that four half-bound staples are attached to the template. For a single-domain staple and the associated pair of template domains there are just four states. There are therefore 34 x × 4 y states of the dimer template with staples, including part-folded states, where x is the number of two-domain staples and y is the number of single-domain staples. Of these, 2 x states consist exclusively of fully-bound staples. Formally, the state space S is given by p0 × p1 ×… × pk−1 where pi denotes the set of possible states for staple i as described above and k is the total number of staples.
Exclusion algorithm
Two template domains hybridized to a single two-domain staple are held within a few tenths of a nanometre of each other at the staple crossover: many of the folds in S cannot meet this constraint. We provide an algorithm that provides an approximate representation of steric constraints, preventing the model from accessing unrealistic states. This method provides an approximation to the real steric constraints: it does not guarantee that each legal state satisfies the constraints or that all states that satisfy the steric constraints are legal.
We define a connected segment of an origami as a set of hybridized domains such that each domain can be reached from each other domain without leaving the set. Two template domains hybridized to the same staple are defined to be connected, as are two adjacent template domains hybridized to different staples. A partially folded segment of origami is considered stress-free (is legal) when it occurs in one of the set of well-ordered, two-dimensional folds shown in Extended Data Figs 1 or 2. These pre-defined folds satisfy the constraints imposed by finite staple length and steric exclusion.
More formally, we can represent the physical origami in partially folded state s ∈ S as an abstract graph G(s) = (V, E) such that each boundary between adjacent domains is a vertex v ∈ V and each template domain and staple crossover is an edge e ∈ E between the appropriate vertices. Each edge has a labelling function f: E → {single-stranded, double-stranded, crossover} that assigns an appropriate status. We can draw subgraphs consisting of connected hybridized segments of the graph: for the origami to be in a legal (stress free) state, each of these subgraphs must be present in a single well-ordered fold from the set shown in Extended Data Figs 1 and 2.
Misfolds occur in the model when at least two connected segments would be incapable of satisfying the constraints were they to become connected to each other. At that point, folding cannot advance unless one of the segments unfolds, allowing another to expand. Extended Data Fig. 3c shows a misfolded dimer that has three connected parts that cannot be joined to form a stress-free state. When simulating assembly using the staple set corresponding to Fig. 4a, about half of the simulations end in a misfolded state; for the weakened-seam variant (Fig. 4b) there are only ∼1% misfolds.
Rates model
We develop a kinetic model of folding based on standard reaction models for hybridization and a method to estimate the effective local concentration of the unhybridized domain of a half-bound staple at its complementary template domain.
Consider complementary strands A and B that can bind reversibly to form duplex AB. Under the assumptions of mass action kinetics, the concentration [AB] is described by

for rate constants k+ and k−. The rate constants are constrained by the requirement that the equilibrium concentrations {A}, {B} and {AB} are consistent with  , the standard change in Gibbs free energy on duplex formation:
, the standard change in Gibbs free energy on duplex formation:

where R denotes the molar gas constant, T temperature.
For staples within a partially folded origami, binding and unbinding rates are similarly constrained by the difference in free energy between states. We approximate the difference in free energy between partially folded states s,s′ that differ by the hybridization of a single template domain as

where  is the standard free energy change corresponding to the formation or dissociation of an equivalent isolated duplex and
is the standard free energy change corresponding to the formation or dissociation of an equivalent isolated duplex and  represents the change in entropy corresponding to the geometric constraints on the template that arise when two-domain staples connect non-contiguous template domains (‘looping constraints’)6,30,31. ΔGshape quantifies cooperative effects: when a single staple domain binds or unbinds, ΔGshape depends on the pattern of binding of other staples.
represents the change in entropy corresponding to the geometric constraints on the template that arise when two-domain staples connect non-contiguous template domains (‘looping constraints’)6,30,31. ΔGshape quantifies cooperative effects: when a single staple domain binds or unbinds, ΔGshape depends on the pattern of binding of other staples.
Consider a single, isolated origami in partially folded state s00 and let staple p bind to the template by a single domain, resulting in state s01. The rate for this reaction is taken to be equal to that for duplex formation between isolated strands:

where σ(s,s′) is the rate of transition from state s to s′. The unbinding rate is then determined by a thermodynamic constraint analogous to equation (2):

We have set  because transitions s01 ↔ s00 do not create or destroy loops in the template. (We do not take into account other ways in which hybridization of a single staple domain affects the free energy of the partly-folded origami, for example, by changing the mechanical properties and thus the free-energy cost of any pre-existing loop of which it forms part.) For the second domain of the staple, once the first domain is bound, we again fix the unbinding rate to be that of the corresponding isolated duplex. This rate does not depend on the change in entropy that results from the removal of a looping constraint30,31 because, immediately after unbinding, the conformation of the template is unchanged:
because transitions s01 ↔ s00 do not create or destroy loops in the template. (We do not take into account other ways in which hybridization of a single staple domain affects the free energy of the partly-folded origami, for example, by changing the mechanical properties and thus the free-energy cost of any pre-existing loop of which it forms part.) For the second domain of the staple, once the first domain is bound, we again fix the unbinding rate to be that of the corresponding isolated duplex. This rate does not depend on the change in entropy that results from the removal of a looping constraint30,31 because, immediately after unbinding, the conformation of the template is unchanged:

where s11 denotes the state in which the staple is bound to the template with both domains. The binding rates of the second domain of the staple, once the first domain is bound, can then be found from the thermodynamic constraint

The free energy penalty  , that corresponds to the additional geometric constraints associated with the binding of the second staple domain, thus determines the binding rate for the second domain.
, that corresponds to the additional geometric constraints associated with the binding of the second staple domain, thus determines the binding rate for the second domain.
Looping constraints
We approximate  , where
, where  corresponds to the entropic penalty of closing the new loop that forms in the template when the second domain of a staple binds. For other transitions, no loop forms and we take ΔGshape = 0. ΔGloop quantifies the difference between the entropic penalties for pinning the template into a loop so that the second staple and template domains can bind and for bringing together two domains unconnected by a loop in a hypothetical ideal system at standard conditions (1 M concentration)32. ΔGloop is thus related to the ratio between the probabilities of bringing two domains into contact in the looped system and in the ideal unconnected system:
corresponds to the entropic penalty of closing the new loop that forms in the template when the second domain of a staple binds. For other transitions, no loop forms and we take ΔGshape = 0. ΔGloop quantifies the difference between the entropic penalties for pinning the template into a loop so that the second staple and template domains can bind and for bringing together two domains unconnected by a loop in a hypothetical ideal system at standard conditions (1 M concentration)32. ΔGloop is thus related to the ratio between the probabilities of bringing two domains into contact in the looped system and in the ideal unconnected system:

Here,  is the probability that the origami adopts a confirmation in which the unbound staple arm and the template domain are spontaneously within an interaction radius r0 of each other, where r0 is an unspecified small distance necessary for closure of the loop.
is the probability that the origami adopts a confirmation in which the unbound staple arm and the template domain are spontaneously within an interaction radius r0 of each other, where r0 is an unspecified small distance necessary for closure of the loop.  is the probability that two unconnected molecules would be within r0 in a hypothetical ideal system of v0 = 1/NA litres, NA being Avogadro’s number. The rate of hybridization of a second staple domain is therefore given by
is the probability that two unconnected molecules would be within r0 in a hypothetical ideal system of v0 = 1/NA litres, NA being Avogadro’s number. The rate of hybridization of a second staple domain is therefore given by

so  denotes the effective concentration of the opposing domain.
denotes the effective concentration of the opposing domain.
As a first approximation we treat the loop of DNA as a freely-jointed chain comprising two types of link, double-stranded DNA and single-stranded DNA (dsDNA and ssDNA respectively). Let P(r) be the probability density for the end-to-end extension of the chain r. Then  is the probability that the two domains are separated by at most r0.
is the probability that the two domains are separated by at most r0.
The end-to-end distance distribution P(r) of a freely-jointed chain, in the limit of a large number of segments, is

where E[r2] is the mean squared distance between the two ends. The result for a single segment type is a classic result of statistical physics33,34. The following argument shows that the result also holds for a chain with heterogeneous segments. From the central limit theorem, for a large number of segments we expect a Gaussian distribution over the x, y and z components of r. Equation (10) is the only Gaussian distribution that also satisfies the symmetry conditions E[x] = E[y] = E[z] = 0, and E[xy] = E[xz] = E[yz] = 0.
The internal association rate is therefore given by:

where we have assumed  in the second step.
in the second step.
The loop that is closed by the insertion of a staple into a part-folded origami has, in general, a complex structure comprising multiply connected domains of single- and double-stranded DNA. We approximate this loop by a single path through the origami, the loop with the smallest expected square end-to-end distance E[r2]. This path represents the most important constraint that leads to the enhancement of the effective local concentration of one end of the loop at the other, and thus provides the most significant enhancement of σ (s01, s11). In order to identify the dominant loop, each edge e ∈ E in the implied graph G(s) = (V, E) of the partially folded origami is assigned a weight equal to the contribution to E[r2] in the freely jointed chain approximation. Dijkstra’s shortest path algorithm35 is used to determine a loop that minimizes E[r2] and hence determines σ (s01, s11).
For the seam staples, which are paired, the loop closed by hybridization of the second staple is particularly small: it consists only of the crossover link. The predictions of the model remain physically sensible: a second staple binding to a seam has an overall ΔG which is ∼4.4 kcal mol−1 less favourable (at T = 60°C) than a continuous duplex. This destabilization is equal to that expected from a 5-nt bulge within a duplex30. We note that for the broken seam variant, the model predicts incorporation temperatures for the unbroken staple that are lower than the regular case by 2.0 °C, compared to 2.2 °C measured in experiment (Extended Data Fig. 6). It is therefore clear that we do not overestimate the cooperative stabilization of seam staples.
The approximations made in estimating the change in free energy when a staple domain binds or unbinds are not thermodynamically self-consistent: the value assigned to the difference in free energy between states depends, in general, on the path taken between them. Models of this kind will be presented in a companion paper, in which they are compared to thermodynamically self-consistent approaches for simpler systems (F.D. et al., submitted).
Parameterization of the model
Compared to unbinding rates, the rate of binding of an isolated duplex is known to be weakly dependent on duplex stability36. We assume k+ to be independent of temperature, domain sequence, and folding state, and we set k+ = 106 M−1 s−1 (refs 25, 36, 37).
The free energy change when each domain binds to its complement, ΔG0,duplex, is taken to be that of a 16-bp DNA duplex averaged over all possible sequences38. Buffer conditions of 40 mM [Tris] and 12.5 mM [Mg2+] are assumed, giving an additional entropic penalty (in units of cal mol−1 K−1) for duplex formation of:38,39,40

where N is the number of phosphates in the duplex. For ssDNA we use a contour length of Lc,ss = 0.6 nm per base and a Kuhn length of λss = 1.8 nm: a single-stranded domain of 16 bases thus has a contour length of 16 × 0.6 nm41,42,43,44,45. For dsDNA we use a contour length of Lc,ds = 0.34 nm per base46 and make the approximation that the persistence length is much longer47 than any relevant duplex: a double-stranded domain of 16 bases thus corresponds to a single rigid link of length λds = 16 × 0.34 nm. A crossover link between the two template domains hybridized to a single staple is treated as a single segment of length λss.
Example rate calculations
Consider the half-bound staple shown in Extended Data Fig. 10a that is hybridized to an otherwise empty template. A seam staple, labelled A, is used as an example here. Its second domain can hybridize to either of two sites: the closer is connected by a 448-nt ssDNA chain (E[r2] = 480 nm2) and the further by a composite chain comprising a 2,208-nt single-stranded chain and one rigid 16-bp double stranded segment (E[r2] = 2,400 nm2). Following the calculation outlined above, we find that for the closer site the effective local concentration of the opposing domain ceff = 51 µM, the loop cost ΔGloop = 6.5 kcal mol−1 (at T = 60 °C) and the hybridization rate σ = 50 s−1. For the further site: ceff = 4.6 µM, ΔGloop = 8.1 kcal mol−1, and σ = 4.5 s−1. The staple is 11 times more likely to bind to the closer domain.
Binding of one staple affects the binding of others by changing the characteristics of the template (or partly-formed origami) that links their two binding domains. We now compute the hybridization rate, loop cost and local concentration for a second seam staple, staple B, in the presence or absence of staple A. In the absence of staple A, the shorter of the two loops that connect two binding domains of the second staple consists of a 864-nt ssDNA chain: E[r2] = 980 nm2, ceff = 18 µM, ΔGloop = 7.2 kcal mol−1, σ = 18 s−1. In the presence of staple A, the loop passes through the link formed by staple A and comprises 384 nt ssDNA, 3 rigid 16-bp dsDNA segments and a staple crossover modelled as a single segment of length λss (Extended Data Fig. 10b): for this shortened loop, E[r2] = 520 nm2, ceff = 46 μM, ΔGloop = 6.6 kcal mol−1 and σ = 46 s−1. Insertion of staple A increases the rate of hybridization of the second domain of staple B by a factor of 2.6 by shortening the distance between its binding sites.
Code availability
The code used to implement the folding model is freely available via https://github.com/fdannenberg/dna.
References
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006)
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009)
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009)
Sobczak, J. P. J., Martin, T. G., Gerling, T. & Dietz, H. Rapid folding of DNA into nanoscale shapes at constant temperature. Science 338, 1458–1461 (2012)
Seeman, N. C. DNA in a material world. Nature 421, 427–431 (2003)
Arbona, J.-M., Aimé, J.-P. & Elezgaray, J. Cooperativity in the annealing of DNA origamis. J. Chem. Phys. 138, 015105 (2013)
Song, J. et al. Direct visualization of transient thermal response of a DNA origami. J. Am. Chem. Soc. 134, 9844–9847 (2012)
Levinthal, C. How to fold graciously. Mössbauer spectroscopy in biological systems. Univ. Illinois Bull. 67, 22–24 (1969)
Anfinsen, C. B. Principles that govern the folding of protein chains. Science 181, 223–230 (1973)
Dobson, C. M. Protein folding and misfolding. Nature 426, 884–890 (2003)
Baker, D. A surprising simplicity to protein folding. Nature 405, 39–42 (2000)
Martin, T. G. & Dietz, H. Magnesium-free self-assembly of multi-layer DNA objects. Nature Commun. 3, 1103 (2012)
Ke, Y., Bellot, G., Voigt, N. V., Fradkov, E. & Shih, W. M. Two design strategies for enhancement of multilayer-DNA-origami folding: underwinding for specific intercalator rescue and staple-break positioning. Chem. Sci. 3, 2587–2597 (2012)
Castro, C. E. et al. A primer to scaffolded DNA origami. Nature Methods 8, 221–229 (2011)
Douglas, S. M., Bachalet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012)
Perrault, S. D. & Shih, W. M. Virus-inspired membrane encapsulation of DNA nanostructures to achieve in vivo stability. ACS Nano 8, 5132–5140 (2014)
Han, D. et al. DNA gridiron nanostructures based on four-arm junctions. Science 339, 1412–1415 (2013)
Ke, Y., Ong, L. L., Shih, W. M. & Yin, P. Three-dimensional structures self-assembled from DNA bricks. Science 338, 1177–1183 (2012)
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008)
Kershner, R. J. et al. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nature Nanotechnol. 4, 557–561 (2009)
Wolynes, P. G., Onuchic, J. N. & Thirumalai, D. Navigating the folding routes. Science 267, 1619–1620 (1995)
Onuchic, J. N., Wolynes, P. G., Lutheyschulten, Z. & Socci, N. D. Towards an outline of the topography of a realistic protein-folding funnel. Proc. Natl Acad. Sci. USA 92, 3626–3630 (1995)
Fink, T. M. A. & Ball, R. C. How many conformations can a protein remember? Phys. Rev. Lett. 87, 198103 (2001)
Jacobsen, J. L. & Kondev, J. Field theory of compact polymers on a square lattice. Nucl. Phys. B 532, 635–688 (1998)
Zhang, D. Y. & Winfree, E. Control of DNA strand displacement kinetics using toehold exchange. J. Am. Chem. Soc. 131, 17303–17314 (2009)
Zhang, P. H. et al. Engineering BspQI nicking enzymes and application of N.BspQI in DNA labeling and production of single-strand DNA. Protein Expr. Purif. 69, 226–234 (2010)
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009)
Wickham, S. F. J. et al. Direct observation of stepwise movement of a synthetic molecular transporter. Nature Nanotechnol. 6, 166–169 (2011)
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977)
SantaLucia, J., Jr & Hicks, D. The thermodynamics of DNA structural motifs. Annu. Rev. Biophys 33, 415–440 (2004)
Jacobson, H. & Stockmayer, W. H. Intramolecular reaction in polycondensations. I. The theory of linear systems. J. Chem. Phys. 18, 1600–1606 (1950)
Ouldridge, T. E., Louis, A. A. & Doye, J. P. K. Extracting bulk properties of self-assembling systems from small simulations. J. Phys. Condens. Matter 22, 104102 (2010)
Rayleigh On the problem of random vibrations, and of random flights in one, two or three dimensions. Phil. Mag. 37, 321–347 (1919)
Chandrasekhar, S. Stochastic problems in physics and astronomy. Rev. Mod. Phys. 15, 1–89 (1943)
Dijkstra, E. A note on two problems in connexion with graphs. Numer. Math. 1, 269–271 (1959)
Morrison, L. E. & Stols, L. M. Sensitive fluorescence-based thermodynamic and kinetic measurements of DNA hybridization in solution. Biochemistry 32, 3095–3104 (1993)
Gao, Y., Wolf, L. K. & Georgiadis, R. M. Secondary structure effects on DNA hybridization kinetics: a solution versus surface comparison. Nucleic Acids Res. 34, 3370–3377 (2006)
SantaLucia, J., Jr A unified view of polymer, dumbbell, and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl Acad. Sci. USA 95, 1460–1465 (1998)
Peryet, N. Prediction of Nucleic Acid Hybridisation: Parameters and Algorithms. PhD thesis, Wayne State Univ. (2000)
Owczarzy, R., Moreira, B. G., You, Y., Behlke, M. A. & Walder, J. A Predicting stability of DNA duplexes in solutions containing magnesium and monovalent cations. Biochemistry 47, 5336–5353 (2008)
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996)
Rivetti, C., Walker, C. & Bustamante, C. Polymer chain statistics and conformational analysis of DNA molecules with bends or sections of different flexibility. J. Mol. Biol. 280, 41–59 (1998)
Mills, J. B., Vacano, E. & Hagerman, P. J. Flexibility of single-stranded DNA: use of gapped duplex helices to determine the persistence lengths of poly(dT) and poly(dA). J. Mol. Biol. 285, 245–257 (1999)
Murphy, M. C., Rasnik, I., Chang, W., Lohman, T. M. & Ha, T. Probing single-stranded DNA conformational flexibility using fluorescence spectroscopy. Biophys. J. 86, 2530–2537 (2004)
Chen, H. et al. Ionic strength-dependent persistence lengths of single-stranded RNA and DNA. Proc. Natl Acad. Sci. USA 109, 799–804 (2012)
Saenger, W. Principles of Nucleic Acid Structure (Springer, 1984)
Hagerman, P. J. Flexibility of DNA. Annu. Rev. Biophys. Biophys. Chem. 17, 265–286 (1988)
Acknowledgements
We thank K.V. Gothelf, M. Dong, A.L.B. Kodal, S. Helmig and S. Zhang (Department of Chemistry and Interdisciplinary Nanoscience Centre iNano, Aarhus, Denmark) for assistance with AFM imaging. This research was supported by Engineering and Physical Sciences Research Council grants EP/G037930/1 and EP/P504287/1, a Human Frontier Science Program grant RGP0030/2013, a Microsoft Research PhD Scholarship (F.D.), the ERC Advanced Grant VERIWARE (F.D. and M.K.) and a Royal Society–Wolfson Research Merit Award (A.J.T.).
Author information
Authors and Affiliations
Contributions
K.E.D. performed the experimental work, F.D. and T.E.O. developed the folding model, J.B. and A.J.T. devised the experimental strategy. All authors contributed to experimental design, interpretation of the data and preparation of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 The set of well-folded, planar states.
a, Well-folded, planar states can be considered as two adjacent monomer tiles linked by a single reciprocal template crossing at any of the locations marked with a triangle and numbered (centre). This gives a set of 6 unique shapes, as indicated (periphery). b, With the exception noted below, there are four ways to make each of these shapes, distinguished by the nucleotide sequence at the template crossing but not resolved by AFM imaging. In the example shown, crossings made at positions 5 and 8 correspond to the fold 4:1, and crossings made at 17 and 20 correspond to fold 1:4, as indicated in the circle diagrams (left). All give the same shape with fractional short edge offset w/W of 3/6 (right). The exception is that there are only two variants of state 5:0i as configurations formed by linking tiles at positions 1 and 24 are not distinguishable, nor are links at positions 12 and 13. c, Detailed view of the connection between monomer tiles in this case, for which the long-edge offset is not precisely defined (it can range from 0 to 2/7 depending on the conformation of the long edge staple). For the purpose of predicting geometry for model configurations, we take an average value of l/L = 1/7. The set of 22 well-folded, planar states thus consists of two folds for the shape shown in c and four folds for each of the other five shapes.
Extended Data Figure 2 Well-folded, non-planar states and an illegal fold.
a, b, The set of legal folds permitted by the model consists of the 22 planar folds defined in Extended Data Fig. 1 and an additional 52 non-planar folds, four for each of the 13 shapes shown here in a, b. Shapes in a are formed by allowing three reciprocal crossings between two tiles, those in b are formed by allowing 5 reciprocal crossings. These non-planar folds form only rarely in simulation. c, An example of a misfolded shape: the part-folded domains are, individually, well-formed but cannot be joined to give a legal fold.
Extended Data Figure 3 Fitting the shapes of origami tiles observed by AFM.
a, AFM images were flattened by line-by-line subtraction of a second-order polynomial. Image processing and fitting were performed using custom MATLAB programs. Image 1.2 × 1.2 µm. b, A histogram of pixel heights was used to set the threshold for the generation of a binary image. The threshold was found by calculating the average of the means of the two peaks corresponding to background and tiles; if this failed because the image was noisy the threshold was set manually. c, Well-separated objects in the binary image which have the approximate area of a dimer tile were flagged for fitting (numbered). d, Tile outlines were generated using a Sobel edge-finding filter. e, Representative fitted outlines (two equal, offset parallelograms) were used to classify dimer tiles as described in the text (compare Fig. 3d).
Extended Data Figure 4 AFM data.
Panels a–e show a 1.5 µm field of view containing structures folded from each of the five staple sets of Fig. 4a–e. Shapes that were flagged for fitting are marked with a dot, green if the shape was successfully fitted and red otherwise. The fitted outlines are superimposed on the image. f, Examples of structures that were either not flagged for fitting or not successfully fitted. AFM images are shown alongside the outline of a suggested structure. The collection of shapes that were not successfully fitted includes crowded areas where shapes are touching and shapes where the two component monomer tiles are distorted, perhaps during deposition on the mica surface, but can be clearly assigned to one of the predicted shapes. Part-folded (or damaged) shapes are also observed, often with one well-folded monomer attached to a part-folded monomer; sometimes a portion of unfolded template can be observed.
Extended Data Figure 5 Strong seam connections influence the folding pathway.
The structure labelled 1:1 is a part-folded intermediate in which four pairs of seam staples are bound. If the seam staples remain in place this intermediate could progress to a fully folded structure with seam configuration 4:1 or 3:2 (as indicated by arrows to the right), but fold 5:0 is inaccessible unless two pairs of seam staples dissociate. Circle diagrams in the upper panel show seam connections corresponding to the structures below.
Extended Data Figure 6 Monitoring origami assembly using fluorescence.
Assembly of a monomer tile (Fig. 1) was monitored using fluorescently labelled staples. The positions of the labelled strands in the folded tile are shown in a: the seam staple was labelled with 5′ Cy3 and 3′ Black Hole Quencher 2, and the body staple with 5′ Cy5 and 3′ Black Hole Quencher 2. Reactions containing the monomer template at 50 nM and staples at 100 nM in a buffer containing 12.5 mM MgCl2, 10 mM Tris-HCl and 0.5 mM EDTA pH 8.0 were held at 96 °C for 10 min, cooled from 96 °C to 25 °C at 0.3 °C min−1, held at 25 °C for 10 min then heated to 96 °C at 0.3 °C min−1. The fluorescence signal for Cy3 and Cy5 was recorded at 0.3 °C intervals during cooling and heating cycles. Staple binding increases the separation between fluorophore and quencher and therefore increases the fluorescence intensity. b, Fluorescence intensities (F) and c, their derivatives (dF/dT) as functions of temperature during origami annealing and melting. Sharp transitions, corresponding to narrow ranges of staple incorporation temperatures, are consistent with cooperative origami assembly. In the case of the unmodified tile the seam staple is incorporated into the tile at the same temperature as the body staple. Hysteresis (marked *) is consistent with the cooperative binding of the seam staple. When one half of the seam is broken the hysteresis observed for seam staple binding is reduced and the seam staple is incorporated at a lower temperature than the body staple. Weakening the seam has little effect on the incorporation of the body staple.
Extended Data Figure 7 Rearrangement of staples during folding.
a–e, Heat maps showing the predictions of the model for the number of reconfiguration events during assembly for each of the staple sets shown in Fig. 4a–e. A ‘reconfiguration event’ occurs when a contact between two template domains is released and replaced by an alternative contact. Domains omitted from the map are those which would generate an illegal fold if reconfigured.
Extended Data Figure 8 Evolving correlations between seam staples in the model during folding.
a, The original staple set (Fig. 4a); b, the broken-seam variant (Fig. 4b). In each case, average data from 1,600 simulations are presented (‘all’) together with subsets sorted by final fold (5:0, 4:1, 3:2 and misfold). The simulation count for each subset is indicated below each panel. Simulations resulting in well-folded, non-planar structures (NP) are included in ‘all’ but not presented separately: such structures occurred 65 times in a and 5 times in b. Circular icons with internal connections of different lengths represent links across the seam (‘seam links’) connecting points on the template spanning (that is, that are separated by) 28, 56, 84, 112 and 140 template domains (as in the ‘circle’ diagrams of Fig. 3). A ‘seam link’ represents a connection across the seam mediated by at least one seam staple (with the original staple set, a, it may also represent a pair of staples). Data are presented at seven different temperatures as the system is cooled. Correlations between seam links are represented graphically by three 5 × 5 blocks. Each pixel represents a correlation between a pair of seam links which are identified by two icons. The orientation of the icons has the same significance as in the ‘circle’ diagrams: two icons related by 180° rotation represent one internal link in each of the two halves of the template; a 90° rotation represents one internal link and one cross link. Only relative orientation is significant so, for example, fully folded state m:n is not distinguished from n:m (Extended Data Fig. 1b). Each pixel represents the average number of pairs of links present with the specified spans and relative orientations (range 0–8; colour coded, key at top right). The bar on the right of the figure, labelled ‘B’, represents the average occupancy of body staples (range 0–2). For staple set a, folding is substantially complete at 62 °C: at this temperature the patterns of correlation that are characteristic of the fully-folded structures can be seen clearly. For example, the presence of the longest (140–domain) link with no cross-link to the other half of the template is characteristic of fold 5:0. (A 140-domain link with a cross-link only occurs in misfolds and NP structures.) A 112-domain link with a 28-domain cross-link is characteristic of 4:1, and the presence of two 56-domain links including a cross-link is characteristic of 3:2. These and other correlations that are characteristic of the final folds are already visible in the averaged correlation maps (when simulations are sorted by final fold) at very early stages of folding. The pattern of seam staples at an early stage of folding is therefore predictive of the final fold (Extended Data Fig. 9). For the broken-seam staple set b, intact seam staples are incorporated later in the folding pathway (the 50% incorporation temperature for seam staples is 64.2 °C for a, 62.3 °C for b). The 50% body staple incorporation temperature is unchanged (63.9 °C for a, 64.0 °C for b). The same characteristic patterns of seam staples that, with the full seam, are associated with different final folds are also visible at high temperatures for the broken-seam staples. However, 90% of broken-seam simulations result in fold 5:0, as designed. Additional evidence for the influence of strong seam contacts on the folding pathway in the model is provided by the dramatically different yields of misfolds: 52% for full-seam staples a, 1% for broken-seam staples b. Stable incorporation of incompatible seam staples in a prevents the formation of well-folded structures.
Extended Data Figure 9 Seam–staple correlations at early stages of folding are predictive of the final fold.
Data shown correspond to the original staple set (see Extended Data Fig. 8a and Fig. 4a). Three tests were applied at the temperature at which, on average, half of all seam staples are incorporated (64.2 °C). These tests were designed to discriminate between patterns of seam staples characteristic of different final folds. For simulations that satisfy each test, the table records the distribution between final folds. Test 1: a 140-domain seam link with no cross-link to the other half of the template (characteristic of fold 5:0). Test 2: a 112-domain link with a 28-domain cross-link (characteristic of fold 4:1). Test 3: two 56-domain links, including one internal link and one cross-link between halves of the template (characteristic of fold 3:2). Highlighted entries correspond to the fold that each test was designed to predict. The last row of the table records the final distribution between folds of all 1,600 simulations.
Extended Data Figure 10 Example calculations of staple hybridization rates.
See Methods section ‘Example rate calculations’ for the worked examples. a, A half-bound seam staple (brown) can bind to one of two sites on the template (green). Distances along the template to each of the two possible binding sites for the second domain of the staple, measured in nucleotides and base pairs, are marked on the template. In the example shown, the closer binding site is connected by a 448-nt ssDNA chain and the further by a composite chain comprising a 2,208-nt single-stranded chain and one rigid 16-bp double stranded segment. The local concentration of the closer domain at the half-bound staple is estimated to be 11 times higher than that of the more distant domain with a correspondingly greater hybridization rate. b, The previous incorporation of staples changes the physical properties of the loops connecting staple binding sites and thus staple incorporation rates. In the absence of staple A, the shortest path between the binding domains of staple B shown consists of a 864-nt ssDNA chain. In the presence of staple A the path is shortened: it passes through the link formed by staple A and comprises 384 nt ssDNA, 3 rigid 16-bp dsDNA segments and a staple crossover. The effect of the previous insertion of staple A, shortening the link between the two binding sites, is to accelerate the hybridization of the second domain of staple B by a factor of 2.6.
Supplementary information
Supplementary Information
This file contains the nucleotide sequences. (PDF 185 kb)
Rights and permissions
About this article
Cite this article
Dunn, K., Dannenberg, F., Ouldridge, T. et al. Guiding the folding pathway of DNA origami. Nature 525, 82–86 (2015). https://doi.org/10.1038/nature14860
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14860
- Springer Nature Limited
This article is cited by
-
Pattern recognition in the nucleation kinetics of non-equilibrium self-assembly
Nature (2024)
-
Synthetic molecular switches driven by DNA-modifying enzymes
Nature Communications (2024)
-
Isothermal self-assembly of multicomponent and evolutive DNA nanostructures
Nature Nanotechnology (2023)
-
In silico induction of missense mutation in NNRTI protein: computational modelling and stability study of modelled proteins
Journal of Mathematical Chemistry (2023)
-
A reversibly gated protein-transporting membrane channel made of DNA
Nature Communications (2022)