Abstract
In vivo, mitochondria display a high degree of connectivity and mobility. Within the cell, mitochondrial fusion and fission machineries tightly control the dynamics and distribution of the mitochondrial network. Due to their key energetic role, the localization of mitochondria at intracellular sites of high-energy demand is crucial to maintain cell energy metabolism. Neurons are metabolically active cells with high-energy demands at locations distant from the cell body (see Chaps. 8 and 9). Consequently, they are particularly dependent on mitochondrial distribution and function. Accordingly, new evidence identifies defective mitochondrial dynamics as a central pathological event underpinning a number of early and late-onset neurodegenerative disorders. Mutations in genes encoding proteins playing central roles in mitochondrial dynamics and functions have been identified in patients with peripheral neuropathies such as Charcot-Marie-Tooth (CMT) and dominant inherited optic atrophy. Moreover, defects of mitochondrial dynamics have recently been associated with common neurodegenerative diseases such as Parkinson’s, Alzheimer’s, and Huntington’s diseases. Understanding the regulation of mitochondrial dynamics in neurons may open new avenues for the development of therapies in neurodegenerative diseases.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Mitochondria are organelles originally observed in spermatocytes more than a century ago [1]. The name mitochondrion is based on their microscopic appearance and is derived from the Greek language for the words thread (mitos) and granules (chondros). Mitochondria are double membrane bound: the inner membrane delimits the matrix and intermembrane space (IMS), whereas the outer membrane separates the IMS from the cytosol. The inner mitochondrial membrane has the highest density of protein in the cell and can be dissociated in two domains: the boundary region, which constitutes flattened membranes that are in close proximity to the outer membrane, and the cristae membranes, which are lamellar invaginations with highly curved edges. The cristae invaginations house the oxidative phosphorylation system (OXPHOS) [2–4]. This system is composed of two functional entities, i.e., the respiratory/electron transport chain (RC) and the phosphorylation system, which includes the ATP synthase and membrane carriers, such as the ATP/ADP carrier (ANT) and the phosphate carrier (PiC). The RC is historically defined as consisting of mobile electron carriers: coenzyme Q and cytochrome c and four complexes, denoted complex I–IV, which perform substrate oxidation to drive proton extrusion from the mitochondrial matrix to the IMS. The proton electrochemical potential difference across the inner membrane (ΔP) is then used by the ATP synthase to drive ATP synthesis thus coupling proton transport to ATP production [5] (see Chap. 1 for details).
Interestingly, complexes I, III, and IV of the RC and ATP synthase are under dual genetic control. The mitochondrial genome (mtDNA) only encodes 13 proteins that are all components of the OXPHOS system, and nuclear genes encode the remaining mitochondrial proteins. The nuclear genome encodes the mitochondrial proteome required for the maintenance and expression of mtDNA [6], protein synthesis [7], import and degradation [8, 9], iron-sulfur cluster synthesis [10], citric acid and urea cycles, fatty acid oxidation, and additional metabolic pathways.
Mitochondria form a dynamic network inside the cell, and specialized transport machineries ensure their mobility and proper subcellular localizations. Due to their key energetic role, mitochondria are often positioned at intracellular sites of high-energy demand. In the muscle, mitochondria are embedded between myofibrils that consume ATP during contraction. Likewise, in neurons mitochondria are transported to and accumulate in synapses to provide the energy required to maintain and regulate neurotransmission. Thus, proper control of mitochondrial subcellular localization and network morphology is an absolute requisite to maintain energy homeostasis and cell functions. This chapter will review recent findings on the energetic relevance of mitochondrial dynamics and its implication in neurodegenerative disorders.
2 Mitochondrial Dynamics
“Mitochondrial dynamics” describes the continuous changes in the position, size, and shape of mitochondria within cells. In eukaryotic cells, mitochondria are arranged in a wide variety of shapes, ranging from long interconnected tubules to individual spheres [11–13]. Mitochondrial morphology is highly plastic and dynamic, and mitochondria can travel long distances on the cytoskeletal track. Depending on the cell type, mitochondrial mobility can be mediated through either the microtubules or the actin cytoskeleton [14, 15]. In neurons, mitochondrial transport occupies a key role in the delivery and renewal of axonal and synaptic mitochondria [16]. Interestingly, while the cytoskeleton is important to ensure proper mitochondrial intracellular distribution and transport, it is not required to maintain mitochondrial network morphology [17–20]. Mitochondrial network morphology is in fact the result of balanced fusion and fission controlled by protein members of the dynamin-related protein (DRP) family. Recent microscopic, structural, and biochemical studies have led to the characterization of the core machinery of mitochondrial fusion and fission.
2.1 Mitochondrial Fusion
The identification of fuzzy onion (FZO) as an essential protein mediating mitochondrial fusion events occurring during spermatogenesis in drosophila drove rapid advances in our knowledge of the mitochondrial dynamic machinery [21]. The characterization of this evolutionary conserved GTPase protein in budding yeast (FZO1) showed that this outer mitochondrial membrane protein functions in mitochondrial fusion [22, 23]. Genetic screens, performed in yeast, led to the identification of the protein machineries involved in mitochondrial fusion and fission [24]. This approach revealed that the core of the yeast mitochondrial fusion machinery is composed of three proteins. FZO1 and MGM1 (Mitochondrial Genome Maintenance 1) [25–27], respectively, control fusion of the outer and inner mitochondrial membrane, whereas UGO1 (UGO is Japanese for fusion) is proposed to be a two membrane-spanning protein mediating the interaction between FZO1 and MGM1 [28–30]. In contrast to UGO1, both FZO1 and MGM1 present mammalian homologues known as mitofusin 1 and mitofusin 2 (MFN1 and MFN2) [31] and OPA1 (optic atrophy 1) [32, 33], respectively (Fig. 7.1). MFN1 and MFN2 show 81 % similarity to each other and are 52 % similar to drosophila FZO [19]. Both MFNs are ubiquitously expressed in mammals, although their mRNA and protein levels strongly differ according to the tissues [17, 31, 34, 35]. The posttranscriptional and posttranslational mechanisms regulating MFNs tissue-specific expression remain largely unknown. Like FZO1, MFNs are anchored to the outer mitochondrial membrane by two transmembrane segments and contain one GTPase and multiple predicted coiled-coil-forming domains which are exposed and face toward the cytosol [17, 36]. The C terminal coiled-coil domain has been shown to mediate MFN1 and MFN2 antiparallel homotypic and heterotypic complexes [19, 37]. Even though both MFN1 and MFN2 are expressed in mouse embryonic fibroblasts, each protein is essential to maintain mitochondrial network morphology [19]. However, despite their high similarity, MFN1 and MFN2 exhibit different GTPase and membrane-tethering capacities [38]. These functional disparities could explain why the mitochondrial network morphology aberration in Mfn2 knockout MEFs can be efficiently rescued by MFN1 overexpression, whereas, overexpression of MFN2 only mildly rescues the diffracted mitochondrial network of Mfn1 knockout MEFs [19]. Interestingly, activity or degradation of FZO1/MFNs proteins is highly regulated by phosphorylation and ubiquitination at distinct lysine residues [39–42].
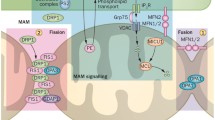
The mammalian machineries involved in mitochondrial fusion. Heterotypic interaction between the mitochondrial outer membrane proteins MFN1 and MFN2 is depicted. OPA1 is located in the intermembrane space and associated with the inner mitochondrial membrane. A red square highlights protein implicated in diseases
In mammals, OPA1 is the main actor controlling the fusion of the mitochondrial inner membrane. MGM1/OPA1 presents multiple isoforms that are located in the IMS or associated with the inner membrane [25–27, 43]. The proteolytic processing of OPA1 generates so-called non-cleaved or long OPA1 (L-OPA1) and cleaved or short OPA1 (S-OPA1) isoforms. The regulated cleavage of OPA1 isoforms by OMA1 (metallopeptidase that exerts activities overlapping with the m-AAA protease) [44, 45] or YME1L (mammalian orthologue of the yeast Yme1) [46–48] results in the loss of the transmembrane domain of the protein and controls the role of OPA1 in mitochondrial fusion. Furthermore, the alternative splicing of OPA1 pre-mRNA introduces additional complexity and yields a total of eight isoforms presenting one or two processing sites and expressed in a tissue-specific manner [49]. However, in contrast with the proteolytic processing, which is common to both orthologues OPA1 and MGM1, the alternative splicing is uniquely involved in the generation of mammalian OPA1 isoforms.
2.2 Mitochondrial Fission
Yeast genetic screens for an extragenic suppressor of fusion mutants led to the identification of key regulators of mitochondrial fission. The key component of the fission machinery DNM1 (dynamin-1) is a cytosolic protein which can be recruited into punctuate structures on the outer mitochondrial membrane [50, 51]. According to the most recent model, this DNM1p recruitment to the outer mitochondrial membrane is mediated by FIS1 (mitochondrial fission protein 1) [52] and causes membrane constriction through its interaction with adaptor proteins MDV1 (mitochondrial division 1) [53–58] and CAF4 (CCR4-associated factor 4) [59, 60]. After its recruitment, DNM1 forms extended spirals [61], which undergo conformational change upon GTP hydrolysis leading to the constriction and division of mitochondria [62].
Extension of these studies to mammalian cells led to the identification of dynamin-related protein 1 (DRP1), also called DLP1 (dynamin-like protein 1) in humans, and FIS1 as components of the mammalian fission machinery [63–65] (Fig. 7.2). However, no orthologues of MDV1 and CAF4 were found in mammals; instead, a growing body of evidence indicates that the mitochondrial fission factor, MFF [66], and the mitochondrial elongation factors, MID49 and MID51 (mitochondrial dynamic proteins of 49 and 51 kDa) [67–71], are further components of the fission machinery. In addition, GDAP1 (ganglioside-induced differentiation-associated protein 1) is a tail-anchored protein of the mitochondrial outer membrane that regulates mitochondrial fission [72, 73]. Whereas GDAP1 recessive mutations are associated with decreased mitochondrial fission activity, dominant mutations result in impairment of mitochondrial fusion [74]. Phylogenetic and structural analyses suggest that GDAP1 belongs to a subfamily of glutathione-S-transferases (GSTs). However, no functional GST activity associated to this protein has been found so far [75]. Unlike other proteins involved in mitochondrial fusion and fission containing GTPase and dynamin domains, GDAP1 sequence analysis does not suggest any involvement in mitochondrial dynamics.
Mammalian mitochondrion under fission. The scheme illustrates DRP1 recruitment to the outer membrane constriction sites by adaptor proteins: FIS1, MFF1, MiD49/51. GDAP1 is located at the mitochondrial outer membrane to mediate mitochondrial fission. A red square highlights protein implicated in diseases
Mitochondrial fission is highly regulated and controlled. For instance, DRP1 undergoes several posttranslational modifications such as phosphorylation [76–79], S-nitrosylation [80, 81], ubiquitination [82–84], and sumoylation [85–87]. These modifications control the activity and subcellular localization of DRP1 [19]. Furthermore, in contrast to mitochondrial fusion, the core of the mitochondrial fission machinery plays a similar role in peroxisomes [65, 88, 89].
3 What Are the Physiological and Bioenergetic Roles of Mitochondrial Dynamics?
Mitochondrial fusion plays a critical role in controlling mitochondrial OXPHOS activity through maintenance of the mitochondrial genome. The complete loss of mitochondrial fusion is associated with a loss of the mitochondrial genome in yeast and with a partial loss of mtDNA in mammals [22, 23, 90–92]. Moreover, the mtDNA maintenance defect observed in mitochondrial fusion deficient skeletal muscle is associated with an accumulation of mitochondrial point mutations and deletions [92]. Remarkably, where emerging data indicate that the majority of mitochondrial fission sites are located in close proximity to mtDNA molecules [93, 94] and endoplasmic reticulum contact sites [95, 96], the loss of mitochondrial fission has no deleterious effect on mtDNA levels [51, 97, 98]. Understanding the role of mitochondrial dynamics in the maintenance and protection of mtDNA continues to attract great scientific interest. Intriguingly, partial impairment of mitochondrial fusion caused by the loss of MFN2 in different mammalian tissues or cultured cells affects mitochondrial bioenergetics without drastically affecting OXPHOS subunits or mtDNA levels [99–101].
Despite their high similarity and their common role in mitochondrial fusion through their physical interaction, MFN1 and MFN2 seem to functionally differ. Ubiquitous knockout of the Mfn1 or Mfn2 genes results in embryo lethality in mid-gestation [19], due to placental dysfunction [19, 102]. Remarkably, by using a conditional knockout allele in conjunction with cre-recombinase expression only in the embryo, Mfn1 or Mfn2 knockout mice are born alive [102]. Mice ubiquitously lacking MFN1 are apparently healthy, whereas loss of MFN2 causes mouse lethality in the early postnatal period and triggers cerebellar atrophy causing severe defects in movement and balance [102]. Despite its well-established role in mitochondrial fusion, a growing body of evidence suggests that MFN2 has additional functions, such as tethering mitochondria with the endoplasmic reticulum (ER) [103], lipid droplets [104], and with the Miro/kinesin system [105, 106]. However, the role of MFN2 in mediating ER-mitochondrial tethering has been recently revised [107, 108]. The different functions of MFN2 are supported by in vivo studies in mice, showing that MFN2 is required for normal glucose homeostasis [99], steroidogenesis [104, 109], is essential for cerebellum development and function [102], and for the axonal projections of dopaminergic neurons [110, 111]. Interestingly, the loss of MFN2 in Purkinje or dopaminergic neurons is associated with a loss of complex IV activity [102, 110]. In contrast, loss of MFN2 in the liver, skeletal muscle, and heart is associated with mild OXPHOS dysfunctions, which are not due to the loss of OXPHOS protein complex levels or activities [99, 112]. Similarly, loss of function of MFN2 in mouse and human fibroblasts cultivated with glucose and high serum shows no gross bioenergetic defects [19, 108, 113]. Interestingly, respiratory chain deficiency observed in MEFs cultivated under low-serum conditions and in mouse heart lacking MFN2 is caused by a deficiency of the terpenoid synthesis pathway affecting the levels of coenzyme Q [101]. This interesting observation demonstrates that bioenergetic defects associated with loss of mitochondrial fusion are not only related to mitochondrial genetic impairments but could originate from anabolic pathway dysfunctions.
Mitochondrial bioenergetics and mitochondrial fusion are heavily interdependent since mitochondrial fusion requires the membrane potential generated across the inner mitochondrial membrane by the respiratory chain [18, 114–116]. Moreover, bioenergetic defects impair mitochondrial dynamics [117] through the proteolytic processing of OPA1 [118]. The strong connection between mitochondrial fusion and bioenergetics could explain the mitochondrial network remodeling observed when cells are metabolizing non-fermentable [119, 120] or high-energy demanding carbon sources [121, 122]. In addition, downregulation of DRP1 activity under starvation has been shown to increase mitochondrial fusion to prevent the degradation of mitochondria through autophagy [123, 124]. In contrast, DRP1 upregulation [77, 85] and MFN2 downregulation [39] under apoptotic conditions promote mitochondrial fragmentation and the following release of proapoptotic factors from mitochondria mediated by BAK [BCL2-antagonist/killer] and BAX [BCL2-associated protein X] [125].
4 Mitochondrial Dynamics and Neurodegeneration
Mitochondrial dynamics hold a central role in maintaining proper mitochondrial distribution and function. Therefore, it is not surprising that alterations in mitochondrial fusion and fission significantly impair neuronal functions. The importance of mitochondrial fusion and fission proteins in the biology of neurons is underscored by the occurrence of several neurodegenerative diseases that result from mutation of these genes. For example, mutations in Mfn2 are associated with Charcot-Marie-Tooth disease type 2A, and Opa1 mutations cause autosomal dominant atrophy characterized by progressive degeneration of the optic nerve.
4.1 Charcot-Marie-Tooth Neuropathy
Charcot-Marie-Tooth [CMT] diseases represent a group of clinically and genetically heterogeneous inherited neuropathies affecting motor and/or sensory neurons. With a prevalence between 1 and 8 cases per 10,000 people, CMT is the most prevalent inherited neuropathy [126–128]. CMT type 2A accounts for 22 % of autosomal dominant neuropathies. Typical clinical symptoms of CMT2A are progressive distal limb muscle weakness and/or atrophy, stepping gait, distal sensory loss, and mobility impairment, which can lead to wheelchair dependency. In addition, optic atrophy can be associated with CMT2A presenting an unusually severe phenotype with an early age of onset [129]. However, while affecting a highly specific set of neurons, i.e., neurons with the longest axons [peripheral sensory and motor neurons], the clinical and electrophysiological phenotypes of CMT2A are very diverse. A major step in the comprehension of the disease was made when several studies identified pathogenic mutations in the Mfn2 gene [129–131], and to date about 60 different Mfn2 mutations have been identified in CMT2A patients [132]. Remarkably, homozygous expression of mutated MFN2T105M in mouse motor neurons recapitulates the key clinical signs of CMT2A. As previously observed in CMT2A neurons [133, 134], affected motor neurons exhibit improper mitochondrial distribution [135]. However, since MFN2 is important for the maintenance of mitochondrial bioenergetics and mitochondrial transport in different types of neuron [102, 110], the molecular role of MFN2 in axonal integrity maintenance remains to be elucidated.
MFN2 is not the only protein involved in mitochondrial dynamics to be associated with the Charcot-Marie-Tooth disease. Unlike other Charcot-Marie-Tooth disease-linked genes, the various GDAP1-associated mutations are associated with demyelinating [136], axonal, or mixed forms of CMT disease with recessive or dominant modes of inheritance, showing a wide range of severity and onset of disease [137–139]. CMT diseases caused by autosomal recessive GDAP1 mutations are generally severe and lead to aggressive disorders appearing during early childhood. The rapid progression of these disorders leads to functional disability. However, the phenotypic presentations of patients carrying GDAP1 mutations are heterogeneous. In most cases patients can present with vocal cord paresis, diaphragmatic paralysis, and facial weakness. In contrast, dominant inherited GDAP1 mutations are less severe and associated with a later onset. Interestingly, GDAP1 knockout mice present with an age-related hypomyelinating peripheral neuropathy. Furthermore, loss of GDAP1 impairs mitochondrial morphology and axonal transport in peripheral neurons [140, 141].
4.2 Autosomal Dominant Optic Atrophy
Autosomal dominant optic atrophy (ADOA) is a hereditary optic neuropathy characterized by a bilateral degeneration of optic nerves causing symmetrical visual loss, typically starting during the first decade of life. The disease primarily affects the retinal ganglion cells (RGCs) and their axons which form the optic nerve [142]. ADOA is mainly linked to OPA1 mutations [32, 49] and is a relatively common form of inherited optic neuropathy; its prevalence is around 3 cases in 100,000 people [143, 144]. In 2007, two independent groups generated mice models expressing a different mutated version of OPA1. These studies showed that OPA1 was required during early mouse embryonic development. Interestingly, heterozygous OPA1 mutants are viable, but exhibit an age-dependent loss of RGCs that eventually progresses to a severe degeneration of ganglion cells and nerve fiber layer [145, 146]. These works demonstrate that the phenotype was not caused by a specific OPA1 proteolytic processing defect, but was associated with a concerted decrease of both L-OPA1 and S-OPA1 levels. In addition, loss of OPA1 was associated with abnormal mitochondrial cristae morphology, mitochondrial swelling, and a reduced number of optic nerve axons. These mouse models showed that mitochondrial defects caused by the loss of OPA1 affect high-energy glutamatergic synapses which lead to dendritic degeneration of ganglion cells [147].
4.3 Neuropathy Linked to DLP1
Human diseases linked to DRP1/DLP1 mutations are extremely rare. In 2007, a de novo mutation in one DLP1 allele was identified in a neonate patient presenting microcephaly, abnormal brain development, optic atrophy, and elevated plasma concentration of lactic acid as well as very long-chain fatty acids causing their death at 37 days of age [148]. Despite the drastic elongation of the mitochondrial and peroxisomal networks, OXPHOS activities were not affected in skin fibroblasts and skeletal muscle biopsies. This mutation was later reported to prevent higher-order assembly of DLP1, thus precluding organelle fission [149]. Interestingly, independent mouse knockouts showed that DRP1 was required for embryonic and brain development. Neuron-specific Drp1 knockout presents aberrant mitochondrial morphology and distribution preventing proper dendritic and axonal development [98, 150]. Loss of DRP1 in dopaminergic neurons affects mitochondrial mobility and depletes axonal mitochondria. These mitochondrial defects cause degeneration of synaptic terminals and cell loss [151].
The pathological spectrum associated with disturbed mitochondrial fission has recently expanded to include Parkinson’s, Huntington’s, and Alzheimer’s diseases. Pathogenic mutations involved in Parkinson’s and Huntington’s diseases have been associated with an increased recruitment of DRP1 to mitochondria resulting in excessive mitochondrial fragmentation [152, 153]. Furthermore, DRP1 has also been shown to interact with a Parkinson disease-related protein, LRRK2 (leucine-rich repeat kinase 2) [154]. This interaction enhances mitochondrial translocation of DRP1 to promote mitochondrial fragmentation. In the case of Alzheimer’s disease, mitochondrial fragmentation is progressively increased during the progression of the disease both in patients and in transgenic mouse models [155–157]. However, the molecular mechanism involved is still unclear. Interestingly, it has been reported that DRP1 posttranslational modifications such as S-nitrosylation and phosphorylation hold a crucial role in Alzheimer’s disease [80, 158].
5 Conclusion
Overall, it is clear that perturbations in mitochondrial dynamics and activity are directly or indirectly involved in neurodegenerative diseases. However, although in vitro studies have provided crucial information regarding mutations, structure, and molecular response, almost nothing is known about the in vivo regulation of mitochondrial dynamics in healthy adult axons or synapses. Furthermore, it is becoming increasingly clear that multiple factors can affect mitochondrial dynamics, and the strong interdependence between mitochondrial activity, dynamics, and transport introduces additional complexity. A future challenge will be to unravel the molecular nature of these interactions. To this end, availability of established disease models will allow rapid progress in our understanding of the physiological regulation of mitochondrial dynamics and will open promising new avenues to test both in vitro and in vivo efficacy of potential new agents to find treatment options for these progressive neurodegenerative diseases.
References
Benda C. Ueber die Spermatogenese der Vertebraten und höherer Evertebraten, II. Theil: Die Histiogenese der Spermien. Arch Anat Physiol. 1989;73:393–8.
Gilkerson RW, Selker JML, Capaldi RA. The cristae membrane of mitochondria is the principal site of oxidative phosphorylation. FEBS Lett. 2003;546(2–3):355–8.
Appelhans T, Richter CP, Wilkens V, Hess ST, Piehler J, Busch KB. Nanoscale organization of mitochondrial microcompartments revealed by combining tracking and localization microscopy. Nano Lett. 2012;12(2):610–6.
Busch KB, Deckers-Hebestreit G, Hanke GT, Mulkidjanian AY. Dynamics of bioenergetic microcompartments. Biol Chem. 2013;394(2):163–88.
Mitchell P. Coupling of phosphorylation to electron and hydrogen transfer by a chemi-osmotic type of mechanism. Nature. 1961;191:144–8.
Falkenberg M, Larsson N-G, Gustafsson CM. DNA replication and transcription in mammalian mitochondria. Annu Rev Biochem. 2007;76:679–99.
Rorbach J, Soleimanpour-Lichaei R, Lightowlers RN, Chrzanowska-Lightowlers ZMA. How do mammalian mitochondria synthesize proteins? Biochem Soc Trans. 2007;35(Pt 5):1290–1.
Bolender N, Sickmann A, Wagner R, Meisinger C, Pfanner N. Multiple pathways for sorting mitochondrial precursor proteins. EMBO Rep. 2008;9(1):42–9.
Tatsuta T, Langer T. Quality control of mitochondria: protection against neurodegeneration and ageing. EMBO J. 2008;27(2):306–14.
Lill R, Mühlenhoff U. Maturation of iron-sulfur proteins in eukaryotes: mechanisms, connected processes, and diseases. Annu Rev Biochem. 2008;77:669–700.
Koning AJ, Lum PY, Williams JM, Wright R. DiOC6 staining reveals organelle structure and dynamics in living yeast cells. Cell Motil Cytoskeleton. 1993;25(2):111–28.
Bereiter-Hahn J, Vöth M. Dynamics of mitochondria in living cells: shape changes, dislocations, fusion, and fission of mitochondria. Microsc Res Tech. 1994;27(3):198–219.
Chen H, Chan DC. Mitochondrial dynamics in mammals. Curr Top Dev Biol. 2004;59:119–44.
Morris RL, Hollenbeck PJ. Axonal transport of mitochondria along microtubules and F-actin in living vertebrate neurons. J Cell Biol. 1995;131(5):1315–26.
Grafstein B, Forman DS. Intracellular transport in neurons. Physiol Rev. 1980;60(4):1167–283.
Hollenbeck PJ, Saxton WM. The axonal transport of mitochondria. J Cell Sci. 2005;118(Pt 23):5411–9.
Rojo M, Legros F, Chateau D, Lombès A. Membrane topology and mitochondrial targeting of mitofusins, ubiquitous mammalian homologs of the transmembrane GTPase Fzo. J Cell Sci. 2002;115(Pt 8):1663–74.
Legros F, Lombès A, Frachon P, Rojo M. Mitochondrial fusion in human cells is efficient, requires the inner membrane potential, and is mediated by mitofusins. Mol Biol Cell. 2002;13(12):4343–54.
Chen H, Detmer SA, Ewald AJ, Griffin EE, Fraser SE, Chan DC. Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J Cell Biol. 2003;160(2):189–200.
Mattenberger Y, James DI, Martinou JC. Fusion of mitochondria in mammalian cells is dependent on the mitochondrial inner membrane potential and independent of microtubules or actin. FEBS Lett. 2003;538(1–3):53–9.
Hales KG, Fuller MT. Developmentally regulated mitochondrial fusion mediated by a conserved, novel, predicted GTPase. Cell. 1997;90(1):121–9.
Hermann GJ, Thatcher JW, Mills JP, Hales KG, Fuller MT, Nunnari J, et al. Mitochondrial fusion in yeast requires the transmembrane GTPase Fzo1p. J Cell Biol. 1998;143(2):359–73.
Rapaport D, Brunner M, Neupert W, Westermann B. Fzo1p is a mitochondrial outer membrane protein essential for the biogenesis of functional mitochondria in Saccharomyces cerevisiae. J Biol Chem. 1998;273(32):20150–5.
Okamoto K, Shaw JM. Mitochondrial morphology and dynamics in yeast and multicellular eukaryotes. Annu Rev Genet. 2005;39:503–36.
Wong ED, Wagner JA, Gorsich SW, McCaffery JM, Shaw JM, Nunnari J. The dynamin-related GTPase, Mgm1p, is an intermembrane space protein required for maintenance of fusion competent mitochondria. J Cell Biol. 2000;151(2):341–52.
Herlan M, Vogel F, Bornhövd C, Neupert W, Reichert AS. Processing of Mgm1 by the rhomboid-type protease Pcp1 is required for maintenance of mitochondrial morphology and of mitochondrial DNA. J Biol Chem. 2003;278(30):27781–8.
Griparic L, van der Wel NN, Orozco IJ, Peters PJ, van der Bliek AM. Loss of the intermembrane space protein Mgm1/OPA1 induces swelling and localized constrictions along the lengths of mitochondria. J Biol Chem. 2004;279(18):18792–8.
Sesaki H, Jensen RE. UGO1 encodes an outer membrane protein required for mitochondrial fusion. J Cell Biol. 2001;152(6):1123–34.
Sesaki H, Jensen RE. Ugo1p links the Fzo1p and Mgm1p GTPases for mitochondrial fusion. J Biol Chem. 2004;279(27):28298–303.
Coonrod EM, Karren MA, Shaw JM. Ugo1p is a multipass transmembrane protein with a single carrier domain required for mitochondrial fusion. Traffic. 2007;8(5):500–11.
Eura Y, Ishihara N, Yokota S, Mihara K. Two mitofusin proteins, mammalian homologues of FZO, with distinct functions are both required for mitochondrial fusion. J Biochem. 2003;134(3):333–44.
Delettre C, Lenaers G, Griffoin JM, Gigarel N, Lorenzo C, Belenguer P, et al. Nuclear gene OPA1, encoding a mitochondrial dynamin-related protein, is mutated in dominant optic atrophy. Nat Genet. 2000;26(2):207–10.
Alexander C, Votruba M, Pesch UE, Thiselton DL, Mayer S, Moore A, et al. OPA1, encoding a dynamin-related GTPase, is mutated in autosomal dominant optic atrophy linked to chromosome 3q28. Nat Genet. 2000;26(2):211–5.
Santel A, Frank S, Gaume B, Herrler M, Youle RJ, Fuller MT. Mitofusin-1 protein is a generally expressed mediator of mitochondrial fusion in mammalian cells. J Cell Sci. 2003;116(Pt 13):2763–74.
Kawalec M, Zabłocka B, Kabzińska D, Neska J, Beręsewicz M. Mitofusin 2 expression dominates over mitofusin 1 exclusively in mouse dorsal root ganglia – a possible explanation for peripheral nervous system involvement in Charcot-Marie-Tooth 2A. Folia Neuropathol. 2014;52(4):436–42.
Fritz S, Rapaport D, Klanner E, Neupert W, Westermann B. Connection of the mitochondrial outer and inner membranes by Fzo1 is critical for organellar fusion. J Cell Biol. 2001;152(4):683–92.
Koshiba T, Detmer SA, Kaiser JT, Chen H, McCaffery JM, Chan DC. Structural basis of mitochondrial tethering by mitofusin complexes. Science. 2004;305(5685):858–62.
Ishihara N, Eura Y, Mihara K. Mitofusin 1 and 2 play distinct roles in mitochondrial fusion reactions via GTPase activity. J Cell Sci. 2004;117(Pt 26):6535–46.
Leboucher GP, Tsai YC, Yang M, Shaw KC, Zhou M, Veenstra TD, et al. Stress-induced phosphorylation and proteasomal degradation of mitofusin 2 facilitates mitochondrial fragmentation and apoptosis. Mol Cell. 2012;47(4):547–57.
Anton F, Dittmar G, Langer T, Escobar-Henriques M. Two deubiquitylases Act on mitofusin and regulate mitochondrial fusion along independent pathways. Mol Cell. 2013;49(3):487–98.
Cohen MMJ, Leboucher GP, Livnat-Levanon N, Glickman MH, Weissman AM. Ubiquitin-proteasome-dependent degradation of a mitofusin, a critical regulator of mitochondrial fusion. Mol Biol Cell. 2008;19(6):2457–64.
Park Y-Y, Cho H. Mitofusin 1 is degraded at G(2)/M phase through ubiquitylation by MARCH5. Cell Div. 2012;7(7:25):1–6.
Olichon A, Emorine LJ, Descoins E, Pelloquin L, Brichese L, Gas N, et al. The human dynamin-related protein OPA1 is anchored to the mitochondrial inner membrane facing the inter-membrane space. FEBS Lett. 2002;523(1–3):171–6.
Ehses S, Raschke I, Mancuso G, Bernacchia A, Geimer S, Tondera D, et al. Regulation of OPA1 processing and mitochondrial fusion by m-AAA protease isoenzymes and OMA1. J Cell Biol. 2009;187(7):1023–36.
Head B, Griparic L, Amiri M, Gandre-Babbe S, van der Bliek AM. Inducible proteolytic inactivation of OPA1 mediated by the OMA1 protease in mammalian cells. J Cell Biol. 2009;187(7):959–66.
Griparic L, Kanazawa T, van der Bliek AM. Regulation of the mitochondrial dynamin-like protein Opa1 by proteolytic cleavage. J Cell Biol. 2007;178(5):757–64.
Song Z, Chen H, Fiket M, Alexander C, Chan DC. OPA1 processing controls mitochondrial fusion and is regulated by mRNA splicing, membrane potential, and Yme1L. J Cell Biol. 2007;178(5):749–55.
Anand R, Wai T, Baker MJ, Kladt N, Schauss AC, Rugarli E, et al. The i-AAA protease YME1L and OMA1 cleave OPA1 to balance mitochondrial fusion and fission. J Cell Biol. 2014;204(6):919–29.
Delettre C, Griffoin JM, Kaplan J, Dollfus H, Lorenz B, Faivre L, et al. Mutation spectrum and splicing variants in the OPA1 gene. Hum Genet. 2001;109(6):584–91.
Otsuga D, Keegan BR, Brisch E, Thatcher JW, Hermann GJ, Bleazard W, et al. The dynamin-related GTPase, Dnm1p, controls mitochondrial morphology in yeast. J Cell Biol. 1998;143(2):333–49.
Bleazard W, McCaffery JM, King EJ, Bale S, Mozdy A, Tieu Q, et al. The dynamin-related GTPase Dnm1 regulates mitochondrial fission in yeast. Nat Cell Biol. 1999;1(5):298–304.
Mozdy AD, McCaffery JM, Shaw JM. Dnm1p GTPase-mediated mitochondrial fission is a multi-step process requiring the novel integral membrane component Fis1p. J Cell Biol. 2000;151(2):367–80.
Tieu Q, Nunnari J. Mdv1p is a WD repeat protein that interacts with the dynamin-related GTPase, Dnm1p, to trigger mitochondrial division. J Cell Biol. 2000;151(2):353–66.
Cerveny KL, McCaffery JM, Jensen RE. Division of mitochondria requires a novel DMN1-interacting protein, Net2p. Mol Biol Cell. 2001;12(2):309–21.
Tieu Q, Okreglak V, Naylor K, Nunnari J. The WD repeat protein, Mdv1p, functions as a molecular adaptor by interacting with Dnm1p and Fis1p during mitochondrial fission. J Cell Biol. 2002;158(3):445–52.
Cerveny KL, Jensen RE. The WD-repeats of Net2p interact with Dnm1p and Fis1p to regulate division of mitochondria. Mol Biol Cell. 2003;14(10):4126–39.
Karren MA, Coonrod EM, Anderson TK, Shaw JM. The role of Fis1p-Mdv1p interactions in mitochondrial fission complex assembly. J Cell Biol. 2005;171(2):291–301.
Naylor K, Ingerman E, Okreglak V, Marino M, Hinshaw JE, Nunnari J. Mdv1 interacts with assembled dnm1 to promote mitochondrial division. J Biol Chem. 2006;281(4):2177–83.
Schauss AC, Bewersdorf J, Jakobs S. Fis1p and Caf4p, but not Mdv1p, determine the polar localization of Dnm1p clusters on the mitochondrial surface. J Cell Sci. 2006;119(Pt 15):3098–106.
Guo Q, Koirala S, Perkins EM, McCaffery JM, Shaw JM. The mitochondrial fission adaptors Caf4 and Mdv1 are not functionally equivalent. PLoS One. 2012;7(12):e53523.
Ingerman E, Perkins EM, Marino M, Mears JA, McCaffery JM, Hinshaw JE, et al. Dnm1 forms spirals that are structurally tailored to fit mitochondria. J Cell Biol. 2005;170(7):1021–7.
Mears JA, Lackner LL, Fang S, Ingerman E, Nunnari J, Hinshaw JE. Conformational changes in Dnm1 support a contractile mechanism for mitochondrial fission. Nat Struct Mol Biol. 2011;18(1):20–6.
James DI, Parone PA, Mattenberger Y, Martinou JC. hFis1, a novel component of the mammalian mitochondrial fission machinery. J Biol Chem. 2003;278(38):36373–9.
Smirnova E, Griparic L, Shurland DL, van der Bliek AM. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol Biol Cell. 2001;12(8):2245–56.
Yoon Y, Krueger EW, Oswald BJ, McNiven MA. The mitochondrial protein hFis1 regulates mitochondrial fission in mammalian cells through an interaction with the dynamin-like protein DLP1. Mol Cell Biol. 2003;23(15):5409–20.
Gandre-Babbe S, van der Bliek AM. The novel tail-anchored membrane protein Mff controls mitochondrial and peroxisomal fission in mammalian cells. Mol Biol Cell. 2008;19(6):2402–12.
Palmer CS, Osellame LD, Laine D, Koutsopoulos OS, Frazier AE, Ryan MT. MiD49 and MiD51, new components of the mitochondrial fission machinery. EMBO Rep. 2011;12(6):565–73.
Zhao J, Liu T, Jin S, Wang X, Qu M, Uhlén P, et al. Human MIEF1 recruits Drp1 to mitochondrial outer membranes and promotes mitochondrial fusion rather than fission. EMBO J. 2011;30(14):2762–78.
Losón OC, Song Z, Chen H, Chan DC. Fis1, Mff, MiD49, and MiD51 mediate Drp1 recruitment in mitochondrial fission. Mol Biol Cell. 2013;24(5):659–67.
Losón OC, Liu R, Rome ME, Meng S, Kaiser JT, Shan S-O, et al. The mitochondrial fission receptor MiD51 requires ADP as a cofactor. Structure. 2014;22(3):367–77.
Losón OC, Meng S, Ngo H, Liu R, Kaiser JT, Chan DC. Crystal structure and functional analysis of MiD49, a receptor for the mitochondrial fission protein Drp1. Protein Sci. 2015;24(3):386–94.
Niemann A, Ruegg M, La Padula V, Schenone A, Suter U. Ganglioside-induced differentiation associated protein 1 is a regulator of the mitochondrial network: new implications for Charcot-Marie-Tooth disease. J Cell Biol. 2005;170(7):1067–78.
Pedrola L, Espert A, Wu X, Claramunt R, Shy ME, Palau F. GDAP1, the protein causing Charcot-Marie-Tooth disease type 4A, is expressed in neurons and is associated with mitochondria. Hum Mol Genet. 2005;14(8):1087–94.
Niemann A, Wagner KM, Ruegg M, Suter U. GDAP1 mutations differ in their effects on mitochondrial dynamics and apoptosis depending on the mode of inheritance. Neurobiol Dis. 2009;36(3):509–20.
Shield AJ, Murray TP, Board PG. Functional characterisation of ganglioside-induced differentiation-associated protein 1 as a glutathione transferase. Biochem Biophys Res Commun. 2006;347(4):859–66.
Taguchi N, Ishihara N, Jofuku A, Oka T, Mihara K. Mitotic phosphorylation of dynamin-related GTPase Drp1 participates in mitochondrial fission. J Biol Chem. 2007;282(15):11521–9.
Cribbs JT, Strack S. Reversible phosphorylation of Drp1 by cyclic AMP-dependent protein kinase and calcineurin regulates mitochondrial fission and cell death. EMBO Rep. 2007;8(10):939–44.
Chang C-R, Blackstone C. Drp1 phosphorylation and mitochondrial regulation. EMBO Rep. 2007;8(12):1088–9, author reply 1089–90.
Cereghetti GM, Stangherlin A, Martins de Brito O, Chang CR, Blackstone C, Bernardi P, et al. Dephosphorylation by calcineurin regulates translocation of Drp1 to mitochondria. Proc Natl Acad Sci U S A. 2008;105(41):15803–8.
Cho D-H, Nakamura T, Fang J, Cieplak P, Godzik A, Gu Z, et al. S-nitrosylation of Drp1 mediates beta-amyloid-related mitochondrial fission and neuronal injury. Science. 2009;324(5923):102–5.
Nakamura T, Cieplak P, Cho D-H, Godzik A, Lipton SA. S-nitrosylation of Drp1 links excessive mitochondrial fission to neuronal injury in neurodegeneration. Mitochondrion. 2010;10(5):573–8.
Nakamura N, Kimura Y, Tokuda M, Honda S, Hirose S. MARCH-V is a novel mitofusin 2- and Drp1-binding protein able to change mitochondrial morphology. EMBO Rep. 2006;7(10):1019–22.
Karbowski M, Neutzner A, Youle RJ. The mitochondrial E3 ubiquitin ligase MARCH5 is required for Drp1 dependent mitochondrial division. J Cell Biol. 2007;178(1):71–84.
Horn SR, Thomenius MJ, Johnson ES, Freel CD, Wu JQ, Coloff JL, et al. Regulation of mitochondrial morphology by APC/CCdh1-mediated control of Drp1 stability. Mol Biol Cell. 2011;22(8):1207–16.
Wasiak S, Zunino R, McBride HM. Bax/Bak promote sumoylation of DRP1 and its stable association with mitochondria during apoptotic cell death. J Cell Biol. 2007;177(3):439–50.
Braschi E, Zunino R, McBride HM. MAPL is a new mitochondrial SUMO E3 ligase that regulates mitochondrial fission. EMBO Rep. 2009;10(7):748–54.
Figueroa-Romero C, Iñiguez-Lluhí JA, Stadler J, Chang C-R, Arnoult D, Keller PJ, et al. SUMOylation of the mitochondrial fission protein Drp1 occurs at multiple nonconsensus sites within the B domain and is linked to its activity cycle. FASEB J. 2009;23(11):3917–27.
Koch A, Thiemann M, Grabenbauer M, Yoon Y, McNiven MA, Schrader M. Dynamin-like protein 1 is involved in peroxisomal fission. J Biol Chem. 2003;278(10):8597–605.
Li X, Gould SJ. The dynamin-like GTPase DLP1 is essential for peroxisome division and is recruited to peroxisomes in part by PEX11. J Biol Chem. 2003;278(19):17012–20.
Jones BA, Fangman WL. Mitochondrial DNA maintenance in yeast requires a protein containing a region related to the GTP-binding domain of dynamin. Genes Dev. 1992;6(3):380–9.
Shepard KA, Yaffe MP. The yeast dynamin-like protein, Mgm1p, functions on the mitochondrial outer membrane to mediate mitochondrial inheritance. J Cell Biol. 1999;144(4):711–20.
Chen H, Vermulst M, Wang YE, Chomyn A, Prolla TA, McCaffery JM, et al. Mitochondrial fusion is required for mtDNA stability in skeletal muscle and tolerance of mtDNA mutations. Cell. 2010;141(2):280–9.
Murley A, Lackner LL, Osman C, West M, Voeltz GK, Walter P, et al. ER-associated mitochondrial division links the distribution of mitochondria and mitochondrial DNA in yeast. Elife. 2013;2:e00422.
Ban-Ishihara R, Ishihara T, Sasaki N, Mihara K, Ishihara N. Dynamics of nucleoid structure regulated by mitochondrial fission contributes to cristae reformation and release of cytochrome c. Proc Natl Acad Sci U S A. 2013;110(29):11863–8.
Meeusen S, Nunnari J. Evidence for a two membrane-spanning autonomous mitochondrial DNA replisome. J Cell Biol. 2003;163(3):503–10.
Kornmann B, Currie E, Collins SR, Schuldiner M, Nunnari J, Weissman JS, et al. An ER-mitochondria tethering complex revealed by a synthetic biology screen. Science. 2009;325(5939):477–81.
Sesaki H, Jensen RE. Division versus fusion: Dnm1p and Fzo1p antagonistically regulate mitochondrial shape. J Cell Biol. 1999;147(4):699–706.
Ishihara N, Nomura M, Jofuku A, Kato H, Suzuki SO, Masuda K, et al. Mitochondrial fission factor Drp1 is essential for embryonic development and synapse formation in mice. Nat Cell Biol. 2009;11(8):958–66.
Sebastián D, Hernández-Alvarez MI, Segalés J, Sorianello E, Muñoz JP, Sala D, et al. Mitofusin 2 (Mfn2) links mitochondrial and endoplasmic reticulum function with insulin signaling and is essential for normal glucose homeostasis. Proc Natl Acad Sci U S A. 2012;109(14):5523–8.
Loiseau D, Chevrollier A, Verny C, Guillet V, Gueguen N, Pou de Crescenzo M-A, et al. Mitochondrial coupling defect in Charcot-Marie-Tooth type 2A disease. Ann Neurol. 2007;61(4):315–23.
Mourier A, Motori E, Brandt T, Lagouge M, Atanassov I, Galinier A, et al. Mitofusin 2 is required to maintain mitochondrial coenzyme Q levels. J Cell Biol. 2015;208(4):429–42.
Chen H, McCaffery JM, Chan DC. Mitochondrial fusion protects against neurodegeneration in the cerebellum. Cell. 2007;130(3):548–62.
de Brito OM, Scorrano L. Mitofusin 2 tethers endoplasmic reticulum to mitochondria. Nature. 2008;456(7222):605–10.
Wasilewski M, Semenzato M, Rafelski SM, Robbins J, Bakardjiev AI, Scorrano L. Optic atrophy 1-dependent mitochondrial remodeling controls steroidogenesis in trophoblasts. Curr Biol. 2012;22(13):1228–34.
Misko A, Jiang S, Wegorzewska I, Milbrandt J, Baloh RH. Mitofusin 2 is necessary for transport of axonal mitochondria and interacts with the Miro/Milton complex. J Neurosci. 2010;30(12):4232–40.
Misko AL, Sasaki Y, Tuck E, Milbrandt J, Baloh RH. Mitofusin2 mutations disrupt axonal mitochondrial positioning and promote axon degeneration. J Neurosci. 2012;32(12):4145–55.
Cosson P, Marchetti A, Ravazzola M, Orci L. Mitofusin-2 independent juxtaposition of endoplasmic reticulum and mitochondria: an ultrastructural study. PLoS One. 2012;7(9):e46293.
Filadi R, Greotti E, Turacchio G, Luini A, Pozzan T, Pizzo P. Mitofusin 2 ablation increases endoplasmic reticulum-mitochondria coupling. Proc Natl Acad Sci USA. 2015;112(17):2174–81.
Duarte A, Poderoso C, Cooke M, Soria G, Cornejo Maciel F, Gottifredi V, et al. Mitochondrial fusion is essential for steroid biosynthesis. PLoS One. 2012;7(9):e45829.
Lee S, Sterky FH, Mourier A, Terzioglu M, Cullheim S, Olson L, et al. Mitofusin 2 is necessary for striatal axonal projections of midbrain dopamine neurons. Hum Mol Genet. 2012;21(22):4827–35.
Pham AH, Meng S, Chu QN, Chan DC. Loss of Mfn2 results in progressive, retrograde degeneration of dopaminergic neurons in the nigrostriatal circuit. Hum Mol Genet. 2012;21(22):4817–26.
Papanicolaou KN, Khairallah RJ, Ngoh GA, Chikando A, Luptak I, O’Shea KM, et al. Mitofusin-2 maintains mitochondrial structure and contributes to stress-induced permeability transition in cardiac myocytes. Mol Cell Biol. 2011;31(6):1309–28.
Amiott EA, Lott P, Soto J, Kang PB, McCaffery JM, Dimauro S, et al. Mitochondrial fusion and function in Charcot-Marie-Tooth type 2A patient fibroblasts with mitofusin 2 mutations. Exp Neurol. 2008;211(1):115–27.
Meeusen S, McCaffery JM, Nunnari J. Mitochondrial fusion intermediates revealed in vitro. Science. 2004;305(5691):1747–52.
Malka F, Guillery O, Cifuentes-Diaz C, Guillou E, Belenguer P, Lombès A, et al. Separate fusion of outer and inner mitochondrial membranes. EMBO Rep. 2005;6(9):853–9.
Sauvanet C, Duvezin-Caubet S, di Rago J-P, Rojo M. Energetic requirements and bioenergetic modulation of mitochondrial morphology and dynamics. Semin Cell Dev Biol. 2010;21(6):558–65.
Benard G, Bellance N, James D, Parrone P, Fernandez H, Letellier T, et al. Mitochondrial bioenergetics and structural network organization. J Cell Sci. 2007;120(Pt 5):838–48.
Duvezin-Caubet S, Jagasia R, Wagener J, Hofmann S, Trifunovic A, Hansson A, et al. Proteolytic processing of OPA1 links mitochondrial dysfunction to alterations in mitochondrial morphology. J Biol Chem. 2006;281(49):37972–9.
Egner A, Jakobs S, Hell SW. Fast 100-nm resolution three-dimensional microscope reveals structural plasticity of mitochondria in live yeast. Proc Natl Acad Sci U S A. 2002;99(6):3370–5.
Jimenez L, Laporte D, Duvezin-Caubet S, Courtout F, Sagot I. Mitochondrial ATP synthases cluster as discrete domains that reorganize with the cellular demand for oxidative phosphorylation. J Cell Sci. 2014;127(Pt 4):719–26.
Rossignol R, Gilkerson R, Aggeler R, Yamagata K, Remington SJ, Capaldi RA. Energy substrate modulates mitochondrial structure and oxidative capacity in cancer cells. Cancer Res. 2004;64(3):985–93.
Mishra P, Carelli V, Manfredi G, Chan DC. Proteolytic cleavage of Opa1 stimulates mitochondrial inner membrane fusion and couples fusion to oxidative phosphorylation. Cell Metab. 2014;19(4):630–41.
Gomes LC, Di Benedetto G, Scorrano L. During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat Cell Biol. 2011;13(5):589–98.
Rambold AS, Kostelecky B, Elia N, Lippincott-Schwartz J. Tubular network formation protects mitochondria from autophagosomal degradation during nutrient starvation. Proc Natl Acad Sci U S A. 2011;108(25):10190–5.
Cassidy-Stone A, Chipuk JE, Ingerman E, Song C, Yoo C, Kuwana T, et al. Chemical inhibition of the mitochondrial division dynamin reveals its role in Bax/Bak-dependent mitochondrial outer membrane permeabilization. Dev Cell. 2008;14(2):193–204.
Skre H. Genetic and clinical aspects of Charcot-Marie-Tooth disease. Clin Genet. 1974;6(2):98–118.
Emery AE. Population frequencies of inherited neuromuscular diseases – a world survey. Neuromuscul Disord. 1991;1(1):19–29.
Hoyer H, Braathen GJ, Busk OL, Holla OL, Svendsen M, Hilmarsen HT, et al. Genetic diagnosis of Charcot-Marie-Tooth disease in a population by next-generation sequencing. Biomed Res Int. 2014;2014:210401.
Chung KW, Kim SB, Park KD, Choi KG, Lee JH, Eun HW, et al. Early onset severe and late-onset mild Charcot-Marie-Tooth disease with mitofusin 2 (MFN2) mutations. Brain. 2006;129(Pt 8):2103–18.
Züchner SS, Mersiyanova IVI, Muglia MM, Bissar-Tadmouri NN, Rochelle JJ, Dadali ELE, et al. Mutations in the mitochondrial GTPase mitofusin 2 cause Charcot-Marie-Tooth neuropathy type 2A. Nat Genet. 2004;36(5):449–51.
Kijima K, Numakura C, Izumino H, Umetsu K, Nezu A, Shiiki T, et al. Mitochondrial GTPase mitofusin 2 mutation in Charcot-Marie-Tooth neuropathy type 2A. Hum Genet. 2005;116(1–2):23–7.
Cartoni R, Martinou JC. Role of mitofusin 2 mutations in the physiopathology of Charcot-Marie-Tooth disease type 2A. Exp Neurol. 2009;218(2):268–73.
Vallat J-M, Ouvrier RA, Pollard JD, Magdelaine C, Zhu D, Nicholson GA, et al. Histopathological findings in hereditary motor and sensory neuropathy of axonal type with onset in early childhood associated with mitofusin 2 mutations. J Neuropathol Exp Neurol. 2008;67(11):1097–102.
Verhoeven K, Claeys KG, Züchner S, Schröder JM, Weis J, Ceuterick C, et al. MFN2 mutation distribution and genotype/phenotype correlation in Charcot-Marie-Tooth type 2. Brain. 2006;129(Pt 8):2093–102.
Detmer SA, Vande Velde C, Cleveland DW, Chan DC. Hind limb gait defects due to motor axon loss and reduced distal muscles in a transgenic mouse model of Charcot-Marie-Tooth type 2A. Hum Mol Genet. 2008;17(3):367–75.
Cuesta A, Pedrola L, Sevilla T, García-Planells J, Chumillas MJ, Mayordomo F, et al. The gene encoding ganglioside-induced differentiation-associated protein 1 is mutated in axonal Charcot-Marie-Tooth type 4A disease. Nat Genet. 2002;30(1):22–5.
Baxter RV, Ben Othmane K, Rochelle JM, Stajich JE, Hulette C, Dew-Knight S, et al. Ganglioside-induced differentiation-associated protein-1 is mutant in Charcot-Marie-Tooth disease type 4A/8q21. Nat Genet. 2002;30(1):21–2.
Claramunt R, Pedrola L, Sevilla T, López de Munain A, Berciano J, Cuesta A, et al. Genetics of Charcot-Marie-Tooth disease type 4A: mutations, inheritance, phenotypic variability, and founder effect. J Med Genet. 2005;42(4):358–65.
Senderek J, Bergmann C, Ramaekers VT, Nelis E, Bernert G, Makowski A, et al. Mutations in the ganglioside-induced differentiation-associated protein-1 (GDAP1) gene in intermediate type autosomal recessive Charcot-Marie-Tooth neuropathy. Brain. 2003;126(Pt 3):642–9.
Niemann A, Huber N, Wagner KM, Somandin C, Horn M, Lebrun-Julien F, et al. The Gdap1 knockout mouse mechanistically links redox control to Charcot-Marie-Tooth disease. Brain. 2014;137(Pt 3):668–82.
Barneo-Muñoz M, Juárez P, Civera-Tregón A, Yndriago L, Pla-Martin D, Zenker J, et al. Lack of GDAP1 induces neuronal calcium and mitochondrial defects in a knockout mouse model of Charcot-Marie-Tooth neuropathy. PLoS Genet. 2015;11(4):e1005115.
Lenaers G, Hamel C, Delettre C, Amati-Bonneau P, Procaccio V, Bonneau D, et al. Dominant optic atrophy. Orphanet J Rare Dis. 2012;7:46.
Thiselton DL, Alexander C, Taanman J-W, Brooks S, Rosenberg T, Eiberg H, et al. A comprehensive survey of mutations in the OPA1 gene in patients with autosomal dominant optic atrophy. Invest Ophthalmol Vis Sci. 2002;43(6):1715–24.
Yu-Wai-Man P, Sitarz KS, Samuels DC, Griffiths PG, Reeve AK, Bindoff LA, et al. OPA1 mutations cause cytochrome c oxidase deficiency due to loss of wild-type mtDNA molecules. Hum Mol Genet. 2010;19(15):3043–52.
Davies VJ, Hollins AJ, Piechota MJ, Yip W, Davies JR, White KE, et al. Opa1 deficiency in a mouse model of autosomal dominant optic atrophy impairs mitochondrial morphology, optic nerve structure and visual function. Hum Mol Genet. 2007;16(11):1307–18.
Alavi MV, Bette S, Schimpf S, Schuettauf F, Schraermeyer U, Wehrl HF, et al. A splice site mutation in the murine Opa1 gene features pathology of autosomal dominant optic atrophy. Brain. 2007;130(Pt 4):1029–42.
Williams PA, Piechota M, von Ruhland C, Taylor E, Morgan JE, Votruba M. Opa1 is essential for retinal ganglion cell synaptic architecture and connectivity. Brain. 2012;135(Pt 2):493–505.
Waterham HR, Koster J, van Roermund CWT, Mooyer PAW, Wanders RJA, Leonard JV. A lethal defect of mitochondrial and peroxisomal fission. N Engl J Med. 2007;356(17):1736–41.
Chang C-R, Manlandro CM, Arnoult D, Stadler J, Posey AE, Hill RB, et al. A lethal de novo mutation in the middle domain of the dynamin-related GTPase Drp1 impairs higher order assembly and mitochondrial division. J Biol Chem. 2010;285(42):32494–503.
Wakabayashi J, Zhang Z, Wakabayashi N, Tamura Y, Fukaya M, Kensler TW, et al. The dynamin-related GTPase Drp1 is required for embryonic and brain development in mice. J Cell Biol. 2009;186(6):805–16.
Berthet A, Margolis EB, Zhang J, Hsieh I, Zhang J, Hnasko TS, et al. Loss of mitochondrial fission depletes axonal mitochondria in midbrain dopamine neurons. J Neurosci. 2014;34(43):14304–17.
Shirendeb U, Reddy AP, Manczak M, Calkins MJ, Mao P, Tagle DA, et al. Abnormal mitochondrial dynamics, mitochondrial loss and mutant huntingtin oligomers in Huntington’s disease: implications for selective neuronal damage. Hum Mol Genet. 2011;20(7):1438–55.
Costa V, Giacomello M, Hudec R, Lopreiato R, Ermak G, Lim D, et al. Mitochondrial fission and cristae disruption increase the response of cell models of Huntington’s disease to apoptotic stimuli. EMBO Mol Med. 2010;2(12):490–503.
Wang X, Yan MH, Fujioka H, Liu J, Wilson-Delfosse A, Chen SG, et al. LRRK2 regulates mitochondrial dynamics and function through direct interaction with DLP1. Hum Mol Genet. 2012;21(9):1931–44.
Su B, Wang X, Bonda D, Perry G, Smith M, Zhu X. Abnormal mitochondrial dynamics – a novel therapeutic target for Alzheimer’s disease? Mol Neurobiol. 2010;41(2–3):87–96.
Wang X, Su B, Lee H-G, Li X, Perry G, Smith MA, et al. Impaired balance of mitochondrial fission and fusion in Alzheimer’s disease. J Neurosci. 2009;29(28):9090–103.
Calkins MJ, Manczak M, Mao P, Shirendeb U, Reddy PH. Impaired mitochondrial biogenesis, defective axonal transport of mitochondria, abnormal mitochondrial dynamics and synaptic degeneration in a mouse model of Alzheimer’s disease. Hum Mol Genet. 2011;20(23):4515–29.
Wang S, Song J, Tan M, Albers KM, Jia J. Mitochondrial fission proteins in peripheral blood lymphocytes are potential biomarkers for Alzheimer’s disease. Eur J Neurol. 2012;19(7):1015–22.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing
About this chapter
Cite this chapter
Mourier, A. (2016). Mitochondrial Dynamics and Neurodegeneration. In: Reeve, A., Simcox, E., Duchen, M., Turnbull, D. (eds) Mitochondrial Dysfunction in Neurodegenerative Disorders. Springer, Cham. https://doi.org/10.1007/978-3-319-28637-2_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-28637-2_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-28635-8
Online ISBN: 978-3-319-28637-2
eBook Packages: MedicineMedicine (R0)