Abstract
Cancer is a most common and lethal disease in which certain cells in our body grow in an uncontrolled way. The growth of new vascular network is essential for sustained growth of tumor to adequately supply nutrients and oxygen. These requirements are mainly fulfilled by formation of fresh blood vessels. Angiogenesis plays a pivotal role in various physiological conditions within the human body, including, embryonic development and tissue repair after trauma or surgery. Angiogenesis is a hallmark feature of cancer, inflammatory diseases, and wound healing. Different growth factors and vascular genes mediate the angiogenic process, which is regulated by epigenetic states of gene especially through small RNAs. Epigenetic modification of tumor cells includes diverse reinforcing and converging signals, including histone modifications, DNA methylation, and non-coding RNAs. This chapter is focused on highlighting the role of angiogenesis in vasculogenesis process and epigenetic modifications in cancer progression. Moreover, we reported the importance of various angiogenic factors and their epigenetic modifications together with a novel therapeutic window towards the treatment of most common cancers using epigenetic regulators.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
7.1 Introduction
Conrad Waddington was the first to coin the term “epigenetics” in 1939, and defined as “any heritable changes in a cellular phenotype without altering the DNA sequence” [1]. Epigenetic is the phenomenon which describes the event eventually involving chromatin mediated regulatory process of DNA-template. Moreover, highly regulated machineries are involved in the process of DNA and histones’ modification and their removal by chromatin-modifying enzymes while, DNA modifications are of four different types [2, 3], histone modification s have 16 classes [4]. These modifications can alter non covalent interactions within and between nucleosomes and thus lead to modify the chromatin structure. These altered chromatin structures act as docking sites for particular proteins with unique domains that specifically identify these modifications.
Epigenetic modification plays an integral role in the regulation of all DNA-based processes, including DNA replication, transcription, and DNA repair. Furthermore, altered genome or irregular expression patterns of chromatin regulators may trigger different tumor cells to transform into malignant cells. Interestingly, epigenetics process like histone modification , DNA methylation , nucleosome remodeling, and RNA-mediated processing trigger various biological activities and play profound regulatory roles in mammalian gene expression . Normal methylation state in the promoter region is involved in the regulation of gene expression; however, altered promoter methylation levels are molecular hallmarks for severe conditions ranging from cancers to psychiatric disorders. Additionally, epigenetic process is also involved in the regulation of vascular genes, and growth factors mediated angiogenic process. Angiogenesis is responsible for new growth in the vascular network since the proliferation along with the metastatic spread, of cancer cells is governed by an adequate supply of oxygen and nutrients and the removal of waste products. Expression levels of angiogenic factors reveal the hostility of tumor cells. Epigenetic state of the vascular endothelial growth factor-A (VEGF-A) promoter activity can be changed by using small RNAs and this result in either increased or decreased VEGF-A expression.
This epigenetic change in VEGF-A expression could be possible through changes in the histone code. Further, DNA methylation has no major impact on the epigenetic alterations of VEGF-A gene [5]. Epigenetic alteration plays an important role in signal transduction and gene-gene interaction. For instance, fibroblast growth factor receptor-2b (FGFR-2b) and Estrogen controls the activity of pituitary tumors by the activity of melanoma-associated antigen (MAGE-A3). This regulatory role is played by both DNA and histone modification and therefore, making an association between epigenetic alterations in vascular cells to angiogenesis [6]. Further, there are several genes present in the epigenetic state of cancers and exert their role in cell-proliferation, migration, DNA repairing, and cell cycle regulation. A comprehensive list of genes and their associated functions in various cancers has been described in Table 7.1.
7.1.1 DNA Methylation, Acetylation and Histone Modification
The methylation of the 5-carbon on cytosine residues in CpG dinucleotides has been reported to be involved in covalent modification of DNA and the most extensive alteration of chromatin . DNA methylation is predominantly identified in centromeres, inactive X-chromosomes, telomeres, and repeated sequences. Furthermore, an epigenetic study in alterations of cancer cell due to methylation is mainly occurred within CpG island promoters. Methylation of CpG Island plays a central role in transcriptional regulation and is altered during malignant transformation. NGS (Next-Generation Generation Sequencing) has provided the genome maps of CpG methylation. NGS has recognized that between 5 and 10 % of normally, unmethylated CpG promoter islands become unusually methylated in several cancer genomes. It has also demonstrated that CpG hypermethylation of promoters not only changed the expression of mRNAs but also the expression of several noncoding RNAs, some of which in turn lead to progression of malignant transformation [2].
In higher eukaryotes, three well-known active DNA methyltransferases have been recognized viz. DNMT1 which is responsible for methyltransferase process identifies hemimethylated DNA during DNA replication [7], DNMT3a and DNMT3b, which are also able to methylate hemimethylated DNA [8]. DNA methylation makes a platform for various methyl-binding proteins, which contains different DNA binding domains such as MBD1, MBD2, MBD3, and MeCP2. These domains are responsible for recruiting histone-modifying enzymes to organize the chromatin-template processes [9]. Mutations in MDB proteins and DNA methyltransferases have been also well-known that contribute to the onset of cancers. Importantly, these mutations are always heterozygous and predicted to interrupt the catalytic activity of such enzymes and thus lead to growing abnormalities [10].
In 1964, Vincent Allfrey prophetically introduced that histone modifications have a functional impact on the regulation of transcription [11]. However, histone modifications occur in distinctive histone proteins, including histone residues such as arginine, lysine and serine and histone variants. These modifications as well involve several chemical groups, including phosphate, methyl and acetyl and have various degrees of methylation such as tri-methylation, di-methylation, and mono-methylation. Methylation and acetylation of histones have direct impacts on a variety of nuclear processes, including DNA repair, DNA replication, gene transcription, and organization of chromosomes. Importantly, histone acetylation is often linked with transcriptional activation whereas histone methylation depends on amino acid sequence and its position in the histone tail [12, 13].
In the promoter regions of CpG islands , hypermethylation leads to progression of tumor-suppressor genes in tumor and is mainly associated with histone markers. It also promotes loss of H3K4 tri-methylation, deacetylation of histones H3, H4, and gain of H3K9 methylation [14, 15]. The occurrence of the hypermethylated and hypo-acetylated histones H3 and H4 silences certain genes, which lead to reducing progression of tumor. Interestingly, in human cancer cells, modifications of histone H4 require a reduced monoacetylated and trimethylated forms [16]. However, monoacetylated Lys16 and trimethylated Lys20 residues of histone H4 is also associated with hypomethylated repetitive DNA sequences, commonly found in breast and liver cancer [17, 18]. Table 7.2 shown below presents a comprehensive list of histone modifying enzyme and their associated functions in various cancers .
7.1.2 Interaction Between Epigenetics and miRNA
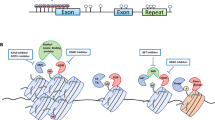
The miRNA is short, 22-nucleotide noncoding RNA, which plays a pivotal role in gene expression through sequence-specific base pairing in the 3′-untranslated region (3′-UTR) of the targeted mRNA. However, outcome of mRNA degradations is tightly regulated and simultaneously plays an essential role in cell proliferation, differentiation, and apoptosis [19]. Most of the human genes lose their binding activity of miRNA to the untranslated regions of the polypeptide chain that helps in quick growth mRNA. Recent studies have revealed that miRNA expression profile differs between normal tissues, cancerous tissues and among various types of tumor [20]. In tumor cell DNA hypermethylation in miRNA 5′-regulatory region causes downregulation of miRNA expression [21]. However, in colon-cancer cells with damaged DNMTs, hypermethylation of the CpG Island does not occur in miRNAs. In other studies, it has been confirmed that methylation silencing of miR-124a also provokes the CDK6 activity in the cell-cycle which is the general feature of epigenetic wound in tumors [21] (Fig. 7.1).
7.1.3 Relation Between Epigenetics and Cancer
Gene expression and DNA methylation studies highlighted the link between epigenetic process and the pathology of cancer [22]. These early studies were purely correlative, which provided possible interaction between cancer and epigenetic cycle. However, these observations have been significantly highlighted by recent results from the International Cancer Genome Consortium (ICGC). The genome sequencing of tumor cells have also given a list of recurrent somatic mutations in several epigenetic regulators [23]. Moreover, the principle theory behind analyzing these cancer genome sequences is the recognition of “driver” mutations. These driver mutations are frequently found in a number of cancers, and they are often present at high primacy in a particular tumor type. For instance, in most of the cases follicular lymphoma is caused due to result of recurrent mutations occurring in histone methyltransferase MLL2, similarly another histone modifying enzyme histone demethylase UTX is mutated in nearly 12 different cancers [24, 25]. Accumulation of such mutated epigenetic regulators in tumor cells describe the involvement of two major epigenetic modifications such as histone methylation and acetylation, which widely affects the epigenetic cycle.
Mapping of Chromatin modifications by deep sequencing technologies have also started to focus in the beginning of epigenetic defects in cancer. Nevertheless, DNA methylation profiles in human cancer with different data, including CHIP-Seq, histone modification and binding of chromatin regulators have raised interesting concerns between cancer associated hypermethylation and genes marked with “bivalent” histone modification in multipotent cells [26]. Although, these bivalent genes are highlighted by active and repressive histone modifications and often act as transcriptionally poised genes, which play a crucial role in development and lineage commitment [27].
In cancerous cells, Most of the genes are mainly targeted on DNA methylation process. In contrast, recent comparisons between normal tissues and malignant from of the same individuals are equally fascinating. These comparisons explain about wide domains in malignant cells that bring major changes in DNA methylation , which is mainly associated with late-replicating regions of the genome with the nuclear lamina [28]. Global alterations in the epigenetic landscape and genomic abrasions in chromatin modifiers show a vital role in cancer and also provide potentiate therapeutics against cancer. However, various numbers of small-molecule inhibitors have been already developed for chromatin regulators. They have various stages of development, which includes Janus kinase 2 (JAK2), histone deacetylase (HDACs), histone acetyl transferase (HAT), and DNA (cytosine-5-)-methyltransferase (DNMTs). These findings reveal that epigenetic pathways are crucial for drug discovery targets over the past decade and thus provide potential therapeutic approaches against a broad spectrum of cancers [29]. Table 7.3 shown below represents a comprehensive list of drugs that affects the acetylation process in various cancers.
7.2 Angiogenesis and Epigenetics
Angiogenesis is a well-defined phenomenon of formation of new blood vessels from pre-existing ones while vasculogenesis is the de novo formation of endothelial cells from mesodermal progenitors. Bone marrow derived endothelial progenitor cells are the lead participants of angiogenesis. Angiogenesis is a crucial determinant of normal human physiology, which plays a central role during reproduction, embryonic growth, development, wound healing and tissue repair following trauma. It is a multistep process; mediated by “on” and “off” triggers, which are regulated by a sophisticated network of interaction between angiogenic factors, tumor cells, phagocytes and their secreted factors, stromal cell cancer stem cells, components of the extracellular matrix (ECM) and most importantly the endothelial cells in a spatio-temporal manner. These factors Angiogenesis is a well-defined phenomenon of formation of new blood vessels from pre-existing ones while vasculogenesis is the de novo formation of endothelial cells from mesodermal progenitors. Bone marrow derived endothelial progenitor cells are the lead participants of angiogenesis. Angiogenesis is a crucial determinant of normal human physiology, which plays a central role during reproduction, embryonic growth, development, wound healing and tissue repair following trauma. It is a multistep process; mediated by “on” and “off” triggers, which are regulated by a sophisticated network of interaction between angiogenic factors, tumor cells, phagocytes and their secreted factors, stromal cell cancer stem cells, components of the extracellular matrix (ECM) and most importantly the endothelial cells in a spatio-temporal manner. These factors taken together have a significant impact on vessel dynamics; and as a certain consequence, uncontrolled or aberrant angiogenesis leads to baffling angiogenic disorders such as vascular insufficiency (myocardial or critical limb ischemia) and vascular overgrowth (hemangiomas, vascularized tumors, and retinopathies), and inflammatory diseases. In addition, uncontrolled angiogenesis marks a significant event during tumor propagation, invasiveness and metastasis thus resulting in aggressive tumor behavior [30–36]. Since, various vascular genes and growth factors are involved in the angiogenic process and thus its regulation is an important for maintaining its normal functions. Epigenetic process is involved in the regulation of these vascular genes and growth factors. The expression of the VEGF gene can be changed by epigenetic processes using small RNAs. The role of small RNAs in epigenetic regulation is crucial for targeting the promoter of VEGF gene thereby leading to alteration of histone code. VEGF bears its effects, mostly via two receptors, VEGFR1 and VEGFR2, and their expression is also controlled by promoter DNA methylation in various cancer cells [5]. These findings advocate about the significance of epigenetic mechanisms in the regulation of vascular genes and growth factors involved in the angiogenic process. together have a significant impact on vessel dynamics; and as a certain consequence, uncontrolled or aberrant angiogenesis leads to baffling angiogenic disorders such as vascular insufficiency (myocardial or critical limb ischemia) and vascular overgrowth (hemangiomas, vascularized tumors, and retinopathies), and inflammatory diseases. In addition, uncontrolled angiogenesis marks a significant event during tumor propagation, invasiveness and metastasis thus resulting in aggressive tumor behavior [30–36]. Since, various vascular genes and growth factors are involved in the angiogenic process and thus its regulation is an important for maintaining its normal functions. Epigenetic process is involved in the regulation of these vascular genes and growth factors. The expression of the VEGF gene can be changed by epigenetic processes using small RNAs. The role of small RNAs in epigenetic regulation is crucial for targeting the promoter of VEGF gene thereby leading to alteration of histone code. VEGF bears its effects, mostly via two receptors, VEGFR1 and VEGFR2, and their expression is also controlled by promoter DNA methylation in various cancer cells [5]. These findings advocate about the significance of epigenetic mechanisms in the regulation of vascular genes and growth factors involved in the angiogenic process.
7.2.1 Background of Angiogenesis
See Fig. 7.2
7.2.2 Types of Angiogenesis?
Angiogenesis; the mechanism by which new blood vessels “sprout” from the pre-existing ones, occurs primarily by endothelial sprouting or via non-sprouting mechanisms.
Sprouting angiogenesis: It was the first identified form of angiogenesis. Initially, it is facilitated via local degradation of the basement membrane at the site of dilated peritumoral postcapillary venule that is at a close proximity to the site of the angiogenic stimulus. Later, the inter-endothelial contacts are loosened, and the endothelial cells (ECs) subsequently migrate into the connective tissue. A solid core of ECs is then formed, and finally; lumen formation takes place proximal to the migratory front and reconnection of the adjacent tubular sprouts occurs in order to form functional capillaries. Formation of basement membrane and recruitment of pericytes occurs simultaneously during the final step of the process [37, 38].
Intussusceptive (splitting) angiogenesis: It was first observed in neonatal rats. This mode of angiogenesis operates in order to expand the capillary bed in size and complexity (intussusceptive microvascular growth). The process starts with the initial formation of a zone of contact between two capillary walls. Following which endothelial cell junctions are reorganized, and the vessel bilayer is perforated in order to facilitate the entry of growth factors and cells into the capillary lumen. Then, a core is formed at the zone of contact between the two vessels that is charged with pericytes and myofibroblasts. Thereafter, these cells start laying collagen fibers in the core area to lay the platform for an extracellular matrix (ECM) necessary for the growth into the vessel lumen. It is interesting to mention here that intussusceptions allow a significant increase in capillaries without a proportional increase in the number of ECs [39, 40].
7.2.3 Mechanisms of Vessel Formation
Apart from sprouting and spitting angiogenesis, nascent blood vessels are formed through several alternative mechanisms under the control of distinct arterio-venous differentiation signals. Table 7.4 briefly summarizes mechanisms for blood vessel generation. It is to be noted here that vascular co-option and vascular mimicry are exclusively used by tumor cells in order to form capillaries.
7.2.4 Roles of Endothelial Cells (ECs) in Angiogenesis?
Endothelial cells play a central role in the control of vascular function. Endothelial cells (ECs) arise from the splanchnopleuric mesoderm and internally line the blood vessels thereby erecting an anticoagulant barrier between the capillary wall and blood. It is known to have diverse functionalities comprising of both basal and inducible metabolic and synthetic functions. Indeed, ECs can respond to a wide repertoire of both physical and chemical stimuli and can critically modulate homeostasis, vasomotor tone, immune and inflammatory responses, cellular adhesion, thrombo resistance, smooth muscle cell proliferation. In addition, ECs are central to the process of angiogenesis and vasculogenesis and commencement of sprouting during vessel formation prerequisites the specification of ECs into tip and stalk cells with different morphological and functional parameters [38, 41–43].
7.2.4.1 Tip, Stalk and the Phalanx Phenotype
Tip endothelial cells are found to operate at the leading edge of an emerging vessel and are the first specialized ECs type within a sprouting vessel. In contrast, stalk cells generally trail behind the tip cells during the sprouting phenomenon and readily form the stalk of the sprout. Notch signals arising out of the tip cells readily attenuate the vascular endothelial growth factor (VEGF)-induced expression of Dll4 on stalk cells thus allowing the tip cells to maintain their forward post during active sprouting. Tip cells are polarized and have the ability to migrate but proliferate minimally, in comparison to endothelial stalks, which has been optimal proliferating capacity. Tip cells have numerous filopodia that serve to guide the newly formed capillary toward the direction of an angiogenic stimulus in comparison to the stalk cells, which produce fewer filopodia, form tubes and branches and proliferate rapidly during the extension of the sprout in order to form the nascent vascular lumen cell. Tip cells express increased levels of Dll-4; platelet derived growth factor-b (PDGF-b), receptors for axon guidance cues, such as the Netrin receptor unc-5 homolog b (UNC5b), CXCR4, neuropilin-1 VEGF receptor-2 (VEGFR-2), and VEGFR-3/Flt-4, while maintaining minimal notch signaling activity. Ang-2 receptor and Tie-2 is expressed in endothelial stalks but not found to be expressed in tip cells. Interestingly, endothelial tip cells can readily adopt a unique “phalanx” phenotype that resembles the phalanx formation as used by the earlier century Greek soldiers. The “phalanx” phenotype is normally observed during the transition from conditions of active sprouting to quiescence tip cells. The cells displaying a “phalanx” phenotype (cobblestone-shaped morphology) are lumenized, non-proliferating, and immobile, which promotes vessel integrity and stabilizes the newly formed vasculature through increased cell adhesion and diminished response to VEGF. Phalanx cells display higher levels of soluble and membrane-bound Flt-1. Flt-1 is a well known to mitigate the pro-angiogenic signals of VEGF thus enabling the phalanx population to maintain a stable morphology. In addition, the expression of VE-cadherin, which tightens the EC-to-EC adhesions, also aids in the phalanx cells in adopting a more quiescent behavior [38, 44, 45].
7.2.5 Growth Factors Involved in Angiogenesis
7.2.5.1 VEGF (Vascular Endothelial Growth Factor)
Vascular endothelial growth factor (VEGF) also known as a vascular permeability factor (VPF) was originally reported as an endothelial cell-specific mitogen. It is secreted by a panel of cells such as tumor cells, macrophages, platelets, keratinocytes, and renal mesangial cells. VEGF also plays critical roles during bone formation, hematopoiesis, wound healing, and development. VEGF family comprises mainly of functionally non-redundant members; of which, the central component VEGF-A participates in the signal transduction and activates the phenomenon of angiogenesis by interaction with VEGFR-2 (FLK1). NRP1 and NRP2 operate independently and act as co-receptors of VEGF signalling. VEGF isoforms facilitates vessel enlargement, whereas matrix-bound isoforms stimulate branching. VEGF (autocrine) released by ECs maintains vascular homeostasis whereas VEGF (paracrine) secreted by tumors, myeloid or stromal cells lead to increased vessel branching and form vessels with a disturbed architecture and dynamics. VEGF-C, acts as a ligand for VEGFR-2 and VEGFR-3 activity thus initiates tip cells. It moderates the formation of the vasculature during early embryogenesis, and later acts as a regulator of lymphamogenesis; the formation of new lymphatic vessels from pre-existing ones. VEGF-B has limited angiogenic capabilities in certain tissues such as the heart, but it can facilitate neuronal survival and induce metabolic effects. VEGF-B also has an articulate role in pathological angiogenesis, where it promotes the growth of cardiac vessels [46, 47].
VEGF receptors VEGFR-1, VEGFR-2 and VEGFR-3 belong to the tyrosine-kinase receptor family and are activated by all the VEGF isoforms. Both the receptors VEGFR-1 and VEGFR-2 are glycosylated. However, only the final glycosylated form of VEGFR-2 undergoes autophosphorylation in response to VEGF signals. These receptors form a subfamily characterized by the presence of seven immunoglobulin-like loops in their extracellular domain and a split tyrosine-kinase domain in their intracellular architecture VEGFR-2 and VEGFR-1 receptors are normally expressed in endothelial cells, but other cells could also express these receptors; VEGFR-1 is expressed in trophoblasts whereas VEGFR-2, in hematopoietic stem cells, megakaryocytes, and retinal progenitor cells. In addition, cancer cells also have the innate tendency to express VEGFR-1 or VEGFR-2. Both the receptors can participate in signal transduction mediated by other growth factors belonging to the VEGF family, but only the VEGF isoforms are equipped to bind to VEGFR-1 and VEGFR-2 [48–50].
7.2.5.2 PDGF (Platelet Derived Growth Factor)
PDGF, a 30 kDa dimer composed of an A- and/or B-chain, was initially extracted and studied as a potent mitogen and chemotactic factor for fibroblasts and all cells of mesenchymal origin, including chondrocytes and mesenchymal stem cells. Genes located on chromosomes 7, 22, 4, and 11 codes for each chain of PDGF. Interestingly, all four PDGF chains contain a highly conserved growth factor domain of approximately 100 amino acids in parallel to the VEGF family. PDGF can act as a mediator of meniscal cell proliferation and migration. It is secreted primarily by platelets but other cells like endothelium and smooth muscle also can act as PDGF sources. Human PDGF comprises of several dimeric forms that are produced from PDGF genes -A, -B, and most recently elucidated C and -D. PDGF plays critical roles during embryonic development and also acts as a stimulator of wound healing. The loss of any PDGF ligand or receptor gene is extremely lethal, and it has recently been reported that mice lacking PDGF receptor expression are vulnerable to traumatic defects in lungs, kidneys, vessels, placenta, brain, and skeleton. In addition, PDGF over expression can lead to several fibrotic disorders and malignancies. PDGF is produced in response to external stimuli like, exposure to low oxygen tension, thrombin, or stimulation by other cytokines and growth factors. PDGF can also function as an autocrine stimulator of tumor cells, a regulator of interstitial fluid pressure, and most importantly as a dynamic modulator of angiogenesis [51–55].
PDGF receptors belong to the type III receptor-tyrosine kinase (RTK) family of receptors. The receptor comprises of five extracellular immunoglobulin (Ig) loops and a split intracellular tyrosine kinase (TK) domain. Other members of the family with a similar structural prototype include c-KIT, c-Fms, FLT3, receptors for CSF-1, SCF, and Flt3-ligand and the macrophage-colony-stimulating factor receptor. PDGF receptors α and β, are encoded from two highly homologous genes. These receptors have specific binding preferences; the isoform binds to all the ligands barring PDGF-D, whereas, PDGFRβ binds to PDGF-B and -D alone [52, 56].
7.2.5.3 FGF (Fibroblast Growth Factor)
The last of the major growth factors involved in angiogenesis is the fibroblast growth factor (FGF). Fibroblast growth factors (FGFs) belong to a family of structurally related polypeptides that are essential during embryonic development and which, operates postnatally as homoeostatic factors, which enables wound healing following trauma. In addition, FGFs also plays a critical role in cellular proliferation, survival, migration, and differentiation. FGFs have been identified in metazoans, but their presence in unicellular organism is a matter of debate. Human FGF comprises of approximately 22 members (Fgf-1-Fgf-23) except Fgf-15, which has not been characterized in humans. bFGF was among the first discovered angiogenic factors just like FGF1, can modulate angiogenesis and arteriogenesis. In addition, FGF9 can also stimulate angiogenesis during bone repair FGFs can have intracrine, paracrine and endocrine functionalities. The paracrine and endocrine functionalities are mediated via FGF receptors (FGFRs), which are expressed on the cell surface. However, the intracrine module of FGF operates independent of the receptors [46, 57, 58].
Besides these all growth factors, Epidermal Growth Factor (EGF) and Transforming Growth Factor-β1 (TGF-β1) also induce angiogenesis. EGF is responsible for Growth, migration, and tube formation to evaluate the direct and indirect effect in the angiogenic process. VEGF has protective role on endothelial cells from apoptosis ; whereas, TGF-β1 induces apoptosis. Thus it signifies TGF-β1 has been opposing effect on endothelial cells. TGF-β1 mediated angiogenesis needs a rapid and transient apoptotic effect, facilitated by VEGF/VEGFR2. These major growth factors are involved in the angiogenesis process through different signalling cascade. The major signalling pathway and their associated growth factors have been listed in Table 7.5.
7.2.6 Molecular Signature of Angiogenesis
In addition to the growth factors discussed above, there is a participation of wide repertoire of molecules, which collaborate in order to accomplish the angiogenic program. Table 7.6 shown below presents a comprehensive list of those angiogenic determinants and their associated functionality.
7.2.7 Mechanism of Angiogenesis
As mentioned earlier, the developing embryo forms a primary vascular plexus initially by the mechanism of vasculogenesis, thereafter; both sprouting and non-sprouting angiogenesis becomes operational to generate blood vessels and the entire functional adult circulatory system [59]. Herein, we discuss the sequential steps involved during the phenomenon of vessel branching [46, 60–62].
Endothelial cells maintain the stage of quiescence: Quiescent endothelial cells are able to maintain longer half-lives in a healthy adult, owing to the operation of an autocrine signaling cascade involving VEGF, Notch, angiopoietin-1 (ANG-1), and fibroblast growth factors (FGFs). In addition, the endothelial mass express oxygen sensors and hypoxia-inducible factors viz. prolyl hydroxylase domain 2 (PHD2) and hypoxia-inducible factor-2α (HIF-2α), which respond to various signals (environmental and physiological) and accordingly allow the vessels to modify their architecture in order to maintain optimal blood flow. During the quiescent stage, streamlined ECs adopt a phalanx phenotype, maintain their pericytic coverage and are interconnected by junction adhesion molecules VE-cadherin and claudins. Pericytes retard EC proliferation, in addition, release cell-survival signals (VEGF and ANG-1), and together with the monolayer of ECs comprise the basement membrane during the stage of quiescence. The basement membrane is 100–200 μm thick and is located immediately below the monolayer of endothelial cells in the arterial intima. Major components of the basement membrane include laminins, type-IV collagen, type-VIII collagen, and proteoglycans.
Sensing the angiogenic signals and endothelial sprouting: Quiescent endothelial cells are capable of sensing a panel of factors (see Table 7.3), which act as angiogenic triggers and in response to those stimuli, ECs enter into a stage of active sprouting categorized by high mitotic index, increased migratory potential and matrix degradation. In addition, pericytes detach themselves from the basement membrane (specifically in response to ANG-2) by proteolytic degradation assisted by matrix metalloproteinases (MMPs). Afterwards, the tight junctions, adherens junctions, and gap junctions present between the intimal ECs and perivascular cells are lost thus allowing the ECs to stalk into the basement membrane and the surrounding milieu. In fact, VEGF augments the permeability of the endothelial cell membrane, which leads to extravasation of several plasma proteins in order to erect a temporary extracellular matrix (ECM). Endothelial cells once liberated from the capillary intima, proliferate, migrate, and are routed in the direction of the angiogenic stimulus in a 3-D extracellular environment thereby giving rise to fresh angiogenic sprouts. Several proteases operate within the ambit of the ECM in order to facilitate the release of angiogenic variables, which are accountable for remodelling the matrix in order to provide an optimal angiogenic ambience.
Endothelial cells split into tip and stalk phenotypes: In order to form a perfused tube with appropriate vessel dynamics, it is therefore, critical for the endothelial mass not to progress the angiogenic signal in unison. However, to prevent this activity from occurring, one endothelial cell is chosen amid the populace to act as leading or tip cell. The cells bordering the tip cell assume secondary positions and are known as stalk cells; these cells divide repeatedly to elongate the stalk and thereby form the vessel lumen. Tip cells are directionally guided by ephrins and semaphorins whereas stalk cells release EGFL7, which is compulsory for stock elongation. Myeloid cells facilitate the fusion of the newly formed vessel with another branch vessel, which is necessary for initiation of blood flow.
Lumen and Tube formation: Lumenogenesis (formation of the lumen) and Tubulogenesis (formation of tubes) are significant phenomenon observed during the process of angiogenesis. The ECs are genetically capable of building luminal compartments, which allows the flow of blood from pre-existing to the newly formed vasculature. The most widely investigated mechanism for the same is intracellular vacuolization (or intracellular canalization); a phenomenon mediated via α2β1 integrin and members of the Rho GTPase family. ECs activate pinocytic mechanisms in order to form a number of intracellular vacuoles; these vacuoles fuse together to form one large intracellular lumen. The protein caveolin-1, which is known to be a key player during receptor-mediated endocytosis and patterning of the caveolae (invagination of the cell which regularly occurs before pinocytosis and subsequent vacuole formation).
Transition to quiescent state and adopting the phalanx phenotype: Once the condition of active sprouting is over and new vessels are formed, the endothelial cells driven by signals (see Table 7.3) revert to the phalanx stage. Platelet-derived growth factor B (PDGF-B), ANG-1, transforming growth factor-β (TGF-β), ephrin-B2 and NOTCH act together in order to render a pericytic covering to the endothelial cells, following which, tissue inhibitors of metalloproteinases (TIMPs) and plasminogen activator operate to lay the basement membrane and reestablish the dismantled junctions to ensure proper vessel dynamics and flow. Vessels, which lack proper perfusion, normally regress.
7.2.8 Epigenetic Modifications of Major Angiogenic Growth Factors
Vascular endothelial growth factor A (VEGF-A) is crucial for the differentiation of endothelial cells. Moreover, its importance in various pathological states such as cancer, retinopathies, inflammation, and arthritis is well documented. VEGF-A activities are mediated through two tyrosine kinase receptors, including VEGFR1 (Flt-1) and VEGFR2 (KDR). The expression of VEGF-A gene is firmly regulated at multiple levels. Recently, it has been revealed that VEGFR1 and VEGFR2 are regulated by an epigenetic mechanism in most of the cancer, including stomach, colon, and hepatocellular carcinoma. Epigenetic state of the VEGF-A promoter can be manipulated by using promoter-targeted small RNAs and this result in either increased or decreased VEGF-A expression. This epigenetic change in VEGF-A could be possible, mostly through changes in the histone code rather than DNA methylation . VEGF-A also induces epigenetic reprogramming of the promoter regions of Rex1 and Oct4 genes. Rex1 gene is important for proliferation, differentiation and exhibits gene control in developing embryos via its epigenetic control on genes, for instance, PEG3, which has been found to play a key role in fetal growth. On the other hand, Oct4 expression is associated with an undifferentiated phenotype and tumors. Upon treatment with VEGF-A, methylation patterns in promoter of both genes (Rex1 and Oct-4) were diminished in endothelial progenitor cells. VEGF-A expression can therefore lead to epigenetic modifications in promoters of these genes [5]. It has been revealed that VEGF-mediated reduction in miR-101 expression causes pro-angiogenic effects that are mediated through reduced repression by miR-101 of the histone-methyltransferase EZH2 . This results into increasing methylation of histone H3 at lysine7 and transcriptome alterations. Furthermore, in the tumor vasculature, increase in endothelial histone-methyltransferase EZH2 is a direct result of VEGF stimulation by a paracrine circuit that stimulates angiogenic states by methylating and silencing vasohibin1 (vash1). This Vash1 gene expression is mainly associated with colorectal cancer [63].
Epigenetic alteration plays an important role in signal transduction and gene-gene interaction. For instance, FGFR2b and Estrogen regulate the activity of melanoma-associated antigen (MAGE-A3), which then controls p53 and p21 activity in pituitary tumors. This regulation is occurred via both DNA and histone modification . Further, FGFR2 allows fibroblast growth factors (FGFs) to transmit signals, which are involved in cell differentiation and proliferation, and is down-regulated by both epigenetic and genetic mechanisms in breast cancer . This is happened due to the presence of Loss of Heterozygosity (LOH) and DNA methylation in specific regions of FGFR2 which determine breast cancer progression by loss of the gene or by limiting transcriptional process. Importantly, epigenetic changes and genetic sequence alterations can simultaneously lead to cancer and other diseases. Fibroblast growth factor receptors (FGFRs) are also dysregulated in a number of developmental and neoplastic conditions. Genome-wide association studies have identified single nucleotide polymorphisms (SNPs) within intron 2 of FGFR2 as a locus is associated with increased risk of breast cancer [6]. Recently, Rao and colleagues have reported that ischemic injury also provokes DNA methylation process in several genes, which is critical for angiogenesis and endothelial cell survival. This DNA methylation process is sensed by MBD2 protein, which mediates transcriptional repression of genes involved in angiogenesis and endothelial cell survival. This report makes a connection between epigenetic alterations in vascular cells to angiogenesis and paving the way for new therapies that could ameliorate perfusion in patients with vascular disease.
7.2.9 Major Signalling Pathways Involved in Angiogenesis
7.2.9.1 ANG/TIE Signaling
Angiopoietins are approximately 70 kDa-secreted secreted glycoproteins, which can be structurally characterized by the presence of an amino-terminal half with coiled. However, these coil domains are necessary for ligand oligomerization. The most widely studied angiopoietins are Ang-1 and Ang-2. Ang-1 and Ang-2 can bind to Tie-2 (for tyrosine kinase with immunoglobulin and EGF-like domains) receptors expressed on the EC surface; however, the binding of Ang-1 alone facilitates angiogenesis and induces the restructuring and stabilisation of blood vessels. In addition, Ang-1 critically modulates vessel maturation, migration, adhesion and survival of endothelial cells, in contrast; Ang-2 acts to destabilise the connections between the endothelium and perivascular cells and promotes cell death and vascular regression and acts as a natural antagonist to Ang-1. However, Ang-2 can also promote neo vascularisation in union with VEGF. Therefore, clearly angiopoietins exert dominant effects in the angiogenic switch and up regulated expression Ang-2 in comparison to Ang-1 has been shown to associate strongly with tumor progression and poor prognosis.
As mentioned earlier, angiopoietins bind to Tie receptors, which are specifically expressed on the vascular endothelial surface and on macrophages deployed during angiogenesis. Tie receptors (Tie-1 and Tie-2) are tyrosine kinase receptors, which have critical functionalities in vascular maturation during development, and both during physiological and pathological angiogenesis. Ang 1-4 are bonafide ligands of the Tie-2 receptor, as a subtle coincidence, Tie-1 remains an orphan receptor. Tie-2 contributes by heterodimerizing with Tie-2 and thereby facilitating Tie-2 signal transduction. Binding of angiopoietins to Tie-2 is facilitated by fibrinogen-related domain (FReD), which is located in close proximity to a 20 residues short linker sequence. FReD comprises of three sub domains namely A, B and P; of which, P mediates the interaction of FReD containing proteins viz. Fibrinogen, tachylectin 5A and angiopoietins with their respective ligands.
Angiopoietin-like proteins (Angptls) have structural similarity with angiopoietins and are coded from seven genes, Angptls 1-7, with all inheriting N-terminal coiled-coil domains and a C-terminal fibrinogen-like domain, which are characteristic angiopoietin signatures. Interestingly, Angptls do not bind to either of the Tie receptors and therefore, remain orphan ligands. Angptls 1, 2, 3, 4, and Angptl6/angiopoietin-related growth factors have critical roles in modulating angiogenesis and regulating lipid, glucose, and energy metabolism independently of angiogenic effects [61, 64–66].
7.2.9.2 Notch Signalling
Notch signaling operates as a crucial angiogenic switch during development, wound healing and pregnancy (physiological angiogenesis), tumor angiogenesis (pathological angiogenesis), and in embryonic vascular development and tumor growth. Notch signalling cascade, therefore has been a subject of pristine interest in recent years towards formulating novel anti angiogenic cancer therapies. Delta-like ligand 4 (DLL4) is a ligand of the notch signalling cascade that operates as a negative regulator of tumor angiogenesis via the Notch-Dll-4 signalling axis. Notch-Dll4 axis serves as an intersection point between pro angiogenic and metabolic signalling prototypes in endothelial cells. Notch directly regulates the expression prototype of the VEGFR transcript and in EC cell lines with activated Notch signaling display a fall in expression level of VEGFR-2 (Kdr). In contrast, DLL-4 haploinsufficient neonatal mice display augmented levels of VEGFR-2 in the retinal vasculature. The angiogenic behavior of endothelial cells within developing blood vessel sprouts is limited by VEGF activity, which in turn is regulated via Notch. Furthermore, Notch signaling cascade is reiteratively utilized during the entire duration of angiogenesis. During the process of angiogenesis in tumors, DLL4 provides a negative feedback on VEGFR2 activity. Recent evidences gathered from various cancer studies predominantly highlight the role of Notch-1 in regulating tumor angiogenesis. However, Notch-3 also can mediate vascular development, remodeling and maturation and most recently it was observed that Notch-3 and the monomeric C-reactive protein (mCRP) together operate via the PI3K/Akt pathway in order to promote angiogenesis. Experiments on both zebrafish and mouse suggest that VEGF-2 and Notch-3 collaborate significantly in order to modulate cell behavior during the process of angiogenesis. In fact, this can be attributed to the ability of ECs within a developing blood vessel to sprout and utilize the Notch cascade in order to modulate cellular phenotypes during angiogenesis. Notch receptors are particularly expressed in “connector” cells and on cells lining the stable blood vessels such as the dorsal aorta. Notch cascade is triggered in these cells through interaction with DLL-4 expressed on the surface of tip cells. As a consequence, aberration in Notch signaling or loss in DLL-4 expression results in an increase in endothelial cells with tip cell signature phenotype in both mouse and zebrafish embryos. Interestingly, cells in zebrafish with activated Notch signaling reportedly do not contribute to vessel formation during angiogenesis. These observations indicates that cells with activated Notch signaling remain in the patent vessel, whereas, cells displaying a dampened Notch activity have the tendency to leave the patent vessel [33, 34, 67–72].
Interestingly, Notch functions as a “double faced angiogenic switch” and recent evidence strongly supports the hypothesis that the cascade has both the pro and anti angiogenic effects. Fatty acid binding protein 4 (FABP4) is a known mediator of VEGFA effects. Notch-DLL4 signalling can directly activate FABP4 independent of VEGFA participation and thereby display pro angiogenic effects. However, the activation of FABP4 following stimulation by VEGFA and/or Notch-Dll4 cascade, requires the involvement of the transcription factor FOXO1. It is therefore, evident that Notch signalling and FOXO1 mediated control of endothelial gene transcription are crucial determinants of angiogenesis. Notch signalling pathway also has the contrasting ability to limit sprouting angiogenesis by reducing the expression level of the pro angiogenic mediator Sox-17. Sox-17 is known to facilitate endothelial migration by destabilizing endothelial junctions and rearranging cytoskeletal structure and by upregulating several tip cell specific genes. The angiogenic effects of Notch signalling activation in GBM (Glioblastoma multiforms) stem cells is associated with a reduction in their growth and migratory potential, downregulation of neural stem cell transcription factors (ASCL1, OLIG2, SOX2) and upregulation of HEY1/2, KLF9, and SNAI2 transcription factors. Notch intracellular domain (NICD) expression in GBM stem cells facilitates the expression of pericyte cell markers NG2, PDGFRβ and α-smooth muscle actin (αSMA) and the induction of angiogenic factors such as HB-EGF, IL8, PLGF, MMP9, VCAM-1, ICAM-1, and ITGA9. Epidermal growth factor-like domain 7 (EGFL7) in parallel with secreted and crucial angiogenic factors such as VEGF and FGF-2, is suitably expressed in endothelial cells (ECs). Probable theories suggest that EGFL7 exerts its angiogenic functionalities by interfering with the Notch pathway and/or by discrete interaction with miR-126. DLL4 overexpression in human endothelial cells (ECs) ablates the effect of IFNγ in those cells. IFNγ is a cytokine and a key mediator of angiostatic response and the observed angiostatic effects of IFNγ could be due to dampening of the DLL4/Notch signal. Thus, the role of IFNγ in ECs is crucial for maintaining tumor angiostasis, which critically involves DLL4 down regulation [73–78]. Regulation of endothelial cell biology by the signaling pathway (Notch) is also necessary for vascular development, homeostasis, and sprouting angiogenesis. Krüppel-like factor (Klf) family of transcription factors act as modulators of endothelial cell biology. Krüppel-like factor 4 (KLF4) regulates sprouting angiogenesis via regulation of Notch activity by retarding the process of cleavage and subsequent Notch-1 activation and by differentially modulating the expression prototype of Notch components (receptor, ligands and target genes). In addition, Notch signaling necessarily mediates critical aspects of tumor angiogenesis as triggered by the vascular endothelial growth factor (VEGF). DLL4 has a major role in the process, and it was suggested that blocking DLL4 inhibits tumor growth and consequently, increases endothelial cell sprouting, but vessels formed lack perfusion. Indeed, this could be due to a significant reduction in the endothelial nitric oxide level caused primarily due to inhibition of the DLL4-Notch signalling module and can be a probable factor behind reduced vessel integrity [79, 80].
7.2.9.3 αv Integrin Signaling
Integrins are transmembrane receptors, which have the ability to bind to extracellular matrix proteins, or other adhesion receptors expressed on neighboring cells. Substrate binding specificity of integrin receptors is facilitated via heterodimeric pairing of α and β subunits. In fact, the αv subunit pairs with β1, β3, β5, β6, and β8. These pairings can preferentially bind a single ligand (αvβ5 for vitronectin) or recognize a number of ligands (αvβ3 binds vitronectin, fibronectin, vWF, tenascin, osteopontin, fibrillin, fibrinogen, and thrombospondin). Endothelial cells undergoing active angiogenesis and remodelling and pathological tissues most commonly display integrin αvβ3 on their surface. A transcriptional activator, Hox D3, mediates the expression of αvβ3 on these cells. However, the efficacy of αvβ3 as a marker of angiogenic endothelial cells is not limited to cancer as this is a common feature associated with the activation of ECs and therefore, activated EC population are highly sensitive to inhibition of αvβ3 most commonly during wound repair, arthritis and proliferative diabetic retinopathy. It is interesting to note here that despite the appearance of αvβ3 on all angiogenic endothelial cells, the participation of αvβ3 in other angiogenic signalling cross talk is not necessarily generic. Angiogenesis induced by basic fibroblast growth factor (bFGF) or tumor necrosis factor α (TNF-α) requires the involvement of αvβ3, whereas integrin αvβ5 is required for processes induced by vascular endothelial growth factor (VEGF) or transforming growth factor α (TGF-α). The striking difference in the functioning of β3 and β5 can be attributed to the ability of these integrins to differentially activate the Ras/Raf/MEK/Erk pathway in capillaries.
αvβ5 integrin pathway operating downstream of VEGF leads to the activation of focal adhesion kinase (FAK) and Src kinase, whereas αvβ3 pathway activates p21-activated kinase (PAK). αvβ5 also plays a major role in endothelial cell protection from apoptosis (intrinsic pathway), independently of MEK-1. This apoptotic protective module in EC results from αvβ5 mediated activation of Raf on serines 338/339, which leads to Raf-1 mitochondrial translocation. Conversely, αvβ3 protect the EC from extrinsic mediated apoptosis by activating Raf on tyrosines 340/341. However, αvβ3 signal activation requires the participation of MEK-1 [81].
7.2.9.4 Wnt Signaling
Cellular proliferation and polarity, apoptosis , branching morphogenesis, inductive processes, and the maintenance of stem cells in an undifferentiated, proliferative state are key cellular prototypes under the control of Wnt signaling. A Wnt/Frizzled signaling pathway also modulates vascular growth, endothelial proliferation, survival and migration in mammals and hence controls the phenomenon of angiogenesis. Wnt signals are propagated by the transcriptional activity of β-catenin and in the absence of ligand-receptor interaction; the cytoplasmic β-catenin is subjected to degradation by the phosphodestruction complex formed by adenomatous polyposis coli (APC), GSK3β and few other proteins thus resulting in repression of Wnt downstream targets. However, in the event of Wnt binding to the receptor Frizzled, β-catenin is preserved from degradation by the complex. The stabilized β-catenin is trafficked into the nucleus where it interacts with T-cell factor (TCF)/Lef transcription factors and activates transcription of target genes. β-catenin can also stabilize cell-to-cell adhesion and tissue integrity by binding to VE- and N-cadherins at the cellular junctions.
β-catenin initiates the transcription of DLL4, which in turn, activates Notch-1 and -4; this interaction plays a central role during embryonic vascular development. Moreover, β-catenin overexpression is associated with alterations in vascular morphology characterized by endothelial cell arterialization and lack of venous specificity. Notch and Wnt pathway operate synergistically in regulating EC differentiation and vascular morphogenesis. This is facilitated via the formation of a transcription complex comprising of β-Catenin, RBP-J and intracellular domain of Notch (NICD); the complex in upregulates the expression of arterial genes. Nrarp is a transcription factor activated via Notch signaling in the retina vasculature. Nrarp can inhibit Notch activation and at the same time increase Wnt signaling by stimulating the transcription factor Lef-1. Th pro angiogenic transcription factor Sox-17 which is required for Norrin/Frizzled-4/Lrp-mediated angiogenesis in a 3D matrix gel is a target of both the Notch and Wnt signaling pathways. Gain-of-function mutation of Frizzled-4 results in diminished levels of Sox-17.
However, the exact mechanism of Sox-17-β-catenin mediated transcriptional activity during angiogenesis in vivo is still a subject of immense curiosity. HIF-1α is a potent angiogenic trigger of vascular endothelial growth factor (VEGF) activity. HIF-1α competes with TCF4 in order to bind to β-catenin. Β-Catenin/HIF1α interaction occurs at the promoter regions of HIF1α target genes thereby increasing their expression. In addition, Rspo1-Wnt-Vegfc-Vegfr3 signaling pathway plays a central role in developmental angiogenesis. R-spondin1 (rspo1) is a Wnt signaling regulator and the catalogued pro-angiogenic effects of Rspo1/Wnt signaling are mediated by Vegfc/Vegfr3 (Flt4) signaling. Actually, Vegfc expression is dependent on Rspo1 and Wnt, and Vegfc and Vegfr3 acts as a mediator of angiogenesis downstream of Rspo1-Wnt axis. All these molecules are active during the dynamic stage of endothelial sprouting and Rspo1-Wnt-VegfC-Vegfr3 signaling cascade plays a crucial role as an endothelial-autonomous permissive cue for developmental angiogenesis. These observations altogether strongly suggest that Wnt transcriptional activity is controlled in a cell-context manner and the interaction of Wnt pathway components with other angiogenic transcriptional factors indeed determines the functional phenotype of the cell during the entire duration of neo-vasculation [82–84].
7.2.9.5 Shh Cascade as an Indirect Angiogenic Switch
The last of our discussion on major angiogenic signaling pathway focuses on the Shh (sonic hedgehog) cascade. The entire panel of Hh (hedgehog) proteins viz. Sonic hedgehog (Shh), Indian hedgehog (Ihh), and Desert hedgehog (Dhh) act as morphogens for a wide range of cells during embryonic development. Shh cascade is signalled via the interaction between Shh and the receptor Patched-1 (Ptch-1); on activation, Ptch-1 consequently, represses the Ptch-1-mediated inhibition of Smoothened (Smo). This leads to the activation of Gli transcription factors, which subsequently control the expression of Shh downstream targets. Shh cascade is known to act as an indirect activator of EC function’s evidence supporting the direct involvement of the cascade in angiogenesis is lacking.
Nevertheless, the role of Shh in vaculogenesis has been investigated, and it was observed that transgenic over expression of Shh in the dorsal neural tube of mouse embryos leads to hypervascularization of the neuroectoderm. Zebrafish deficient in Shh levels, display muddled endothelial precursors, which are incompetent to form the dorsal aorta or axial vein, whereas, Shh deficient mice develop lungs with the improper vasculature. The embryonic Shh cascade becomes operational in adults in response to ischemic injury, including hind limb-ischemia and myocardial infarction, and it was most recently observed that delivery of Shh into the traumatized cells promote neovascularization of ischemic tissue both by promoting angiogenesis and by controlling endothelial progenitor cell adhesion, migration, and proliferation. Shh can facilitate the release of pro-angiogenic factors VEGF and ANG-1 from Shh induced fibroblasts and cardiomyocytes, thereby highlighting the indirect role of Shh on vascular cells. Ptch-1 is not over expressed in corneal neovessel ECs and human umbilical vein endothelial cells (HUVECs) on Shh mediated activation; however, Shh augments capillary morphogenesis in both HUVECs and murine brain capillary ECs. Shh can activate the PI3K/Akt pathway in ECs, and in turn, cyclic AMP/protein kinase A axis can modulate the activity of Shh downstream transcription factor Gli-1. In addition, Shh also interacts with other crucial signaling components such as chicken ovalbumin upstream promoter transcription factor II (COUP TFII) in mouse embryonic carcinoma cells, extracellular signal-regulated kinases 1/2 (ERK 1/2) and protein kinase Cδ (PKCδ) in fibroblasts, Rho/Rho kinase (ROCK) pathway in neuronal cells, matrix metalloproteinase 9 (MMP-9), and osteopontin (OPN). Taken together, the function of Shh activity in mature endothelial cells and the intricate role of non-classical signalling pathways in Shh-mediated angiogenesis remain an obscure subject for active debate in future [85, 86].
7.3 Conclusion
The future of cancer therapies depends on the researchers’ ability to adjust actions to circumstances and has a clear projection relating to the aberrant mechanisms that ultimately decide the fate of the pathogenesis and henceforth degeneration through various processes. Despite great advancement in the field of cancer therapies, still the future of such therapies hangs on morbid conjecture and fragile hopes. The distinct signalling axis presents us an interesting opportunity to exploit in pathology of cancers; furthermore, interrupting critical interactions of the pathways by using different epigenetic regulators can solve the “targeted therapy crisis” problem in cancer pathogenesis .
Angiogenesis plays a crucial role in cell proliferation, metastasis and thus leads to cancer pathogenesis . Since, epigenetic modification plays a key role in the regulation of all DNA-based processes, including DNA replication, transcription, and DNA repair. Thus, altered genome or irregular expression patterns in chromatin regulators can have intense results, which may lead to the trigger various tumor cells and also maintain the tumor cells. Here in this book chapter, we have also discussed about various genes which is mainly involved in epigenetic mediated regulatory process and also highlighted about several histone modifying enzymes, which have a potent role in transcription repression, DNA damage repairing, and also have a tumor suppressor like activity. At last, we have focused on various drugs such as butyrate, curcumin, and several others, which are critical for acetylation process in epigenetics. They mainly target HAT, HDAC, and SIRT activity in different cancers. However, further research is needed in order to make drugs/compounds in cancer therapeutics a blatant reality.
References
Berger SL, Kouzarides T, Shiekhattar R, Shilatifard A. An operational definition of epigenetics. Genes Dev. 2009;23:781–3.
Baylin SB, Jones PA. A decade of exploring the cancer epigenome—biological and translational implications. Nat Rev Cancer. 2011;11:726–34.
Wu H, Zhang Y. Mechanisms and functions of Tet protein-mediated 5-methylcytosine oxidation. Genes Dev. 2011;25:2436–52.
Tan M, Luo H, Lee S, Jin F, Yang JS, Montellier E, Buchou T, Cheng Z, Rousseaux S, Rajagopal N, et al. Identification of 67 histone marks and histone lysine crotonylation as a new type of histone modification. Cell. 2011;146:1016–28.
Mikko PT, Seppo YH. Epigenetic regulation of key vascular genes and growth factors. Cardiovasc Res. 2011;90:441–6.
Zhu X, Wetta H. Genetics and epigenetics in tumorigenesis: acting separately or linked? Austin J Clin Med. 2014;1:1016.
Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell. 1992;69:915–26.
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57.
Klose RJ, Bird AP. Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci. 2006;31:89–97.
Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J, Van Vlierberghe P, Dolgalev I, Thomas S, Aminova O, et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med. 2012;366:1079–89.
Allfrey VG, Faulkner R, Mirsky AE. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci U S A. 1964;51:786–94.
Bernstein BE, Meissner A, Lander ES. The mammalian epigenome. Cell. 2007;128:669–81.
Mack GS. Epigenetic cancer therapy makes headway. J Natl Cancer Inst. 2006;98:1443–4.
Ballestar E, Paz MF, Valle L, et al. Methyl-CpG binding proteins identify novel sites of epigenetic inactivation in human cancer. EMBO J. 2003;22:6335–45.
Jones PA, Baylin SB. The epigenomics of cancer. Cell. 2007;128:683–92.
Fraga MF, Ballestar E, Villar-Garea A, et al. Loss of acetylation at Lys16 and trimethylation at Lys20 of histone H4 is a common hallmark of human cancer. Nat Genet. 2005;37:391–400.
Pogribny IP, Ross SA, Tryndyak VP, Pogribna M, Poirier LA, Karpinets TV. Histone H3 lysine 9 and H4 lysine 20 trimethylation and the expression of Suv4-20h2 and Suv-39h1 histone methyltransferases in hepatocarcinogenesis induced by methyl deficiency in rats. Carcinogenesis. 2006;27:1180–6.
Tryndyak VP, Kovalchuk O, Pogribny IP. Loss of DNA methylation and histone H4 lysine 20 trimethylation in human breast cancer cells is associated with aberrant expression of DNA methyltransferase 1, Suv4-20h2 histone methyltransferase and methyl-binding proteins. Cancer Biol Ther. 2006;5:65–70.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Lujambio A, Ropero S, Ballestar E, et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 2007;67:1424–9.
Feinberg AP, Tycko B. The history of cancer epigenetics. Nat Rev Cancer. 2004;4:143–53.
Forbes SA, Bindal N, Bamford S, Cole C, Kok CY, Beare D, Jia M, Shepherd R, Leung K, Menzies A, et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 2011;39(Database issue):D945–50.
Morin RD, Mendez-Lago M, Mungall AJ, Goya R, Mungall KL, Corbett RD, Johnson NA, Severson TM, Chiu R, Field M, et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature. 2011;476:298–303.
van Haaften G, Dalgliesh GL, Davies H, Chen L, Bignell G, Greenman C, Edkins S, Hardy C, O’Meara S, Teague J, et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat Genet. 2009;41:521–3.
Easwaran H, Johnstone SE, Van Neste L, Ohm J, Mosbruger T, Wang Q, Aryee MJ, Joyce P, Ahuja N, Weisenberger D, et al. A DNA hypermethylation module for the stem/progenitor cell signature of cancer. Genome Res. 2012;22:837–49.
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–26.
Berman BP, Weisenberger DJ, Aman JF, Hinoue T, Ramjan Z, Liu Y, Noushmehr H, Lange CP, van Dijk CM, Tollenaar RA, et al. Regions of focal DNA hypermethylation and long-range hypomethylation in colorectal cancer coincide with nuclear lamina-associated domains. Nat Genet. 2012;44:40–6.
Minucci S, Pelicci PG. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer. 2006;6:38–51.
Dimova I, Popivanov G, Djonov V. Angiogenesis in cancer general pathways and their therapeutic implications. J BUON. 2014;19:15–21.
Flamme I, Frölich T, Risau W. Molecular mechanisms of vasculogenesis and embryonic angiogenesis. J Cell Physiol. 1997;173:206–10.
Katoh M. Therapeutics targeting angiogenesis: genetics and epigenetics, extracellular miRNAs and signaling networks. Int J Mol Med. 2013;32:763–7.
Liu J, Deutsch U, Jeong J, Lobe CG. Constitutive notch signaling in adult transgenic mice inhibits bFGF-induced angiogenesis and blocks ovarian follicle development. Genesis. 2014;52:809–16.
Liu Z, Fan F, Wang A, Zheng S, Lu Y. Dll4-Notch signaling in regulation of tumor angiogenesis. J Cancer Res Clin Oncol. 2014;140:525–36.
Risau W, Flamme I. Vasculogenesis. Annu Rev Cell Dev Biol. 1995;11:73–91.
Yoo SY, Kwon SM. Angiogenesis and its therapeutic opportunities (2013). Mediators Inflamm. 2013;2013:127170.
Eilken HM, Adams RH. Dynamics of endothelial cell behavior in sprouting angiogenesis. Curr Opin Cell Biol. 2010;22:617–25.
Ribatti D, Crivellato E. “Sprouting angiogenesis”, a reappraisal. Dev Biol. 2012;372:157–65.
Burri PH, Hlushchuk R, Djonov V. Intussusceptive angiogenesis: its emergence, its characteristics, and its significance. Dev Dyn. 2004;231:474–88.
De Spiegelaere W, Casteleyn C, Van den Broeck W, Plendl J, Bahramsoltani M, Simoens P, Djonov V, Cornillie P. Intussusceptive angiogenesis: a biologically relevant form of angiogenesis. J Vasc Res. 2012;49:390–404.
Deanfield JE, Halcox JP, Rabelink TJ. Endothelial function and dysfunction: testing and clinical relevance. Circulation. 2007;115:1285–95.
Michiels C. Endothelial cell functions. J Cell Physiol. 2003;196:430–43.
Sumpio BE, Riley JT, Dardik A. Cells in focus: endothelial cell. Int J Biochem Cell Biol. 2002;34:1508–12.
Bautch VL. Endothelial cells form a phalanx to block tumor metastasis. Cell. 2009;136:810–2.
Blancas AA, Wong LE, Glaser DE, McCloskey KE. Specialized tip/stalk-like and phalanx-like endothelial cells from embryonic stem cells. Stem Cells Dev. 2013;22:1398–407.
Carmeliet P, Jain RK. Molecular mechanisms and clinical applications of angiogenesis. Nature. 2011;473:298–307.
Duffy AM, Bouchier-Hayes DJ, Harmey JH (2000). Vascular endothelial growth factor (VEGF) and its role in non-endothelial cells: autocrine signalling by VEGF. In: Madame Curie bioscience database [Internet]. Austin: Landes Bioscience.
Moreira IS, Fernandes PA, Ramos MJ. Vascular endothelial growth factor (VEGF) inhibition critical review. Anticancer Agents Med Chem. 2007;7:223–45.
Neufeld G, Cohen T, Gengrinovitch S, Poltorak Z. Vascular endothelial growth factor (VEGF) and its receptors. FASEB J. 1999;13:9–22.
Shibuya M. Vascular endothelial growth factor (VEGF) and its receptor (VEGFR) signaling in angiogenesis: a crucial target for anti- and pro-angiogenic therapies. Genes Cancer. 2011;2:1097–105.
Alvarez RH, Kantarjian HM, Cortes JE. Biology of platelet-derived growth factor and its involvement in disease. Mayo Clin Proc. 2006;81:1241–57.
Demoulin JB, Montano-Almendras CP. Platelet-derived growth factors and their receptors in normal and malignant hematopoiesis. Am J Blood Res. 2012;2:44–56.
Raica M, Cimpean AM. Platelet-derived growth factor (PDGF)/PDGF receptors (PDGFR) axis as target for antitumor and antiangiogenic therapy. Pharmaceuticals. 2010;3:572–99.
Schmidt MB, Chen EH, Lynch SE. A review of the effects of insulin-like growth factor and platelet derived growth factor on in vivo cartilage healing and repair. Osteoarthritis Cartilage. 2006;14:403–12.
Tennant M, McGeachie JK. Platelet-derived growth factor and its role in atherogenesis: a brief review. Aust N Z J Surg. 1991;61:482–8.
Andrae J, Gallini R, Betsholtz C. Role of platelet-derived growth factors in physiology and medicine. Genes Dev. 2008;22:1276–312.
Itoh N, Ornitz DM. Fibroblast growth factors: from molecular evolution to roles in development, metabolism and disease. J Biochem. 2011;149:121–30.
Turner N, Grose R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer. 2010;10:116–29.
Risau W. Mechanisms of angiogenesis. Nature. 1997;386:671–4.
Adair TH, Montani JP. Overview of angiogenesis. San Rafael: Morgan & Claypool Life Sciences; 2010.
Bisht M, Dhasmana DC, Bist SS. Angiogenesis: future of pharmacological modulation. Indian J Pharmacol. 2010;42:2–8.
Ucuzian AA, Gassman AA, East AT, Greisler HP. Molecular mediators of angiogenesis. J Burn Care Res. 2010;31:158–75.
Smits M, Mir SE, Nilsson RJ, et al. Down-regulation of miR-101 in endothelial cells promotes blood vessel formation through reduced repression of EZH2. PLoS One. 2011;6:e16282.
Fagiani E, Christofori G. Angiopoietins in angiogenesis. Cancer Lett. 2013;328:18–26.
Hato T, Tabata M, Oike Y. The role of angiopoietin-like proteins in angiogenesis and metabolism. Trends Cardiovasc Med. 2008;18:6–14.
Hansen TM, Singh H, Tahir TA, Brindle NPJ. Effects of angiopoietins-1 and -2 on the receptor tyrosine kinase Tie2 are differentially regulated at the endothelial cell surface. Cell Signal. 2010;22:527–32.
Boras E, Slevin M, Alexander MY, Aljohi A, Gilmore W, Ashworth J, et al. Monomeric C-reactive protein and Notch-3 co-operatively increase angiogenesis through PI3K signalling pathway. Cytokine. 2014;69:165–79.
Hu GH, Liu H, Lai P, Guo ZF, Xu L, Yao XD, et al. Delta-like ligand 4 (Dll4) predicts the prognosis of clear cell renal cell carcinoma, and anti-Dll4 suppresses tumor growth in vivo. Int J Clin Exp Pathol. 2014;7:2143–52.
Murtas D, Piras F, Minerba L, Maxia C, Ferreli C, Demurtas P, et al. Activated Notch1 expression is associated with angiogenesis in cutaneous melanoma. Clin Exp Med. 2014;15(3):351–60.
Siekmann AF, Lawson ND. Notch signalling and the regulation of angiogenesis. Cell Adh Migr. 2007;1:104–6.
Zhang E, Feng X, Liu F, Zhang P, Liang J, Tang X. Roles of PI3K/Akt and c-jun signaling pathways in human papillomavirus type 16 oncoprotein-induced HIF-1α, VEGF, and IL-8 expression and in vitro angiogenesis in non-small cell lung cancer cells. PLoS One. 2014;9:e103440.
Zhang JP, Li N, Bai WZ, Qiu XC, Ma BA, Zhou Y, et al. Notch ligand Delta-like 1 promotes the metastasis of melanoma by enhancing tumor adhesion. Braz J Med Biol. 2014;47:299–306.
Deng J, Liu X, Rong L, Ni C, Li X, Yang W, et al. IFNγ-responsiveness of endothelial cells leads to efficient angiostasis in tumors involving down-regulation of Dll4. J Pathol. 2014;233:170–82.
Guichet PO, Guelfi S, Teigell M, Hoppe L, Bakalara N, Bauchet L, et al. Notch1 stimulation induces a vascularization switch with pericyte-like cell differentiation of glioblastoma stem cells. Stem Cells. 2015;33:21–34.
Harjes U, Bridges E, McIntyre A, Fielding BA, Harris AL. Fatty acid binding protein 4, a point of convergence for angiogenic and metabolic signalling pathways in endothelial cells. J Biol Chem. 2014;289:23168–76.
Lee C, Jia Z, Rahmatpanah F, Zhang Q, Zi X, McClelland M, Mercola D. Role of the adjacent stroma cells in prostate cancer development and progression: synergy between TGF-β and IGF signaling. Biomed Res Int. 2014;2014:502093.
Lee SH, Lee S, Yang H, Song S, Kim K, Saunders TL, et al. Notch pathway targets proangiogenic regulator sox17 to restrict angiogenesis. Circ Res. 2014;115:215–26.
Takeuchi K, Yanai R, Kumase F, Morizane Y, Suzuki J, Kayama M, et al. EGF-like-domain-7 is required for VEGF-induced Akt/ERK activation and vascular tube formation in an ex vivo angiogenesis assay. PLoS One. 2014;9:e91849.
Hale AT, Tian H, Anih E, Recio FO, Shatat MA, Johnson T, et al. Endothelial Kruppel-like factor 4 regulates angiogenesis and the Notch signaling pathway. J Biol Chem. 2014;289:12016–28.
Patenaude A, Fuller M, Chang L, Wong F, Paliouras G, Shaw R, et al. Endothelial-specific Notch blockade inhibits vascular function and tumor growth through an eNOS-dependent mechanism. Cancer Res. 2014;74:2402–11.
Weis SM, Cheresh DA. αV integrins in angiogenesis and cancer. Cold Spring Harb Perspect Med. 2011;1:a006478.
Dejana E. The role of wnt signaling in physiological and pathological angiogenesis. Circ Res. 2010;107:943–52.
Gore AV, Swift MR, Cha YR, Lo B, McKinney MC, Li W, Castranova D, Davis A, Mukouyama YS, Weinstein BM. Rspo1/Wnt signaling promotes angiogenesis via Vegfc/Vegfr3. Development. 2011;138:4875–86.
Zerlin M, Julius MA, Kitajewski J. Wnt/Frizzled signaling in angiogenesis. Angiogenesis. 2008;11:63–9.
Pola R, Ling LE, Silver M, Corbley MJ, Kearney M, Blake Pepinsky R, Shapiro R, Taylor FR, Baker DP, Asahara T, Isner JM. The morphogen Sonic hedgehog is an indirect angiogenic agent upregulating two families of angiogenic growth factors. Nat Med. 2001;7:706–11.
Renault MA, Roncalli J, Tongers J, Thorne T, Klyachko E, Misener S, Volpert OV, Mehta S, Burg A, Luedemann C, Qin G, Kishore R, Losordo DW. Sonic hedgehog induces angiogenesis via Rho kinase-dependent signaling in endothelial cells. J Mol Cell Cardiol. 2010;49:490–8.
Virmani AK, Rathi A, Sathyanarayana UG, et al. Aberrant methylation of the adenomatous polyposis coli (APC) gene promoter 1A in breast and lung carcinomas. Clin Cancer Res. 2001;7:1998–2004.
Kawakami K, Brabender J, Lord RV, et al. Hypermethylated APC DNA in plasma and prognosis of patients with esophageal adenocarcinoma. J Natl Cancer Inst. 2000;92:1805–11.
Dobrovic A, Simpfendorfer D. Methylation of the BRCA1 gene in sporadic breast cancer. Cancer Res. 1997;57:3347–50.
Chan KY, Ozcelik H, Cheung AN, et al. Epigenetic factors controlling the BRCA1 and BRCA2 genes in sporadic ovarian cancer. Cancer Res. 2002;62:4151–6.
Sanchez-Cespedes M, Esteller M, Wu L, et al. Gene promoter hypermethylation in tumors and serum of head and neck cancer patients. Cancer Res. 2000;60:892–5.
Villuendas R, Sanchez-Beato M, Martinez JC, et al. Loss of p16/INK4A protein expression in non-Hodgkin’s lymphomas is a frequent finding associated with tumor progression. Am J Pathol. 1998;153:887–997.
Evron E, Dooley WC, Umbricht CB, et al. Detection of breast cancer cells in ductal lavage fluid by methylation-specific PCR. Lancet. 2001;357:1335–6.
Harden SV, Tokumaru Y, Westra WH, et al. Gene promoter hypermethylation in tumors and lymph nodes of stage I lung cancer patients. Clin Cancer Res. 2003;9:1370–5.
Graff JR, Greenberg VE, Herman JG, et al. Distinct patterns of E-cadherin CpG island methylation in papillary, follicular, Hurthle’s cell, and poorly differentiated human thyroid carcinoma. Cancer Res. 1998;58:2063–6.
Waki T, Tamura G, Tsuchiya T, et al. Promoter methylation status of E-cadherin, hMLH1, and p16 genes in nonneoplastic gastric epithelia. Am J Pathol. 2002;161:399–403.
Yang X, Yan L, Davidson NE. DNA methylation in breast cancer. Endocr Relat Cancer. 2001;8:115–27.
Li LC, Chui R, Nakajima K, et al. Frequent methylation of estrogen receptor in prostate cancer: correlation with tumor progression. Cancer Res. 2000;60:702–6.
Lee WH, Morton RA, Epstein JI, et al. Cytidine methylation of regulatory sequences near the pi-class glutathione S-transferase gene accompanies human prostatic carcinogenesis. Proc Natl Acad Sci U S A. 1994;91:11733–7.
Esteller M, Corn PG, Urena JM, et al. Inactivation of glutathione S-transferase P1 gene by promoter hypermethylation in human neoplasia. Cancer Res. 1998;58:4515–8.
Kondo E, Furukawa T, Yoshinaga K, et al. Not hMSH2 but hMLH1 is frequently silenced by hypermethylation in endometrial cancer but rarely silenced in pancreatic cancer with microsatellite instability. Int J Oncol. 2000;17:535–41.
Esteller M, Garcia-Foncillas J, Andion E, et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med. 2000;343:1350–4.
Dominguez G, Carballido J, Silva J, et al. P14ARF promoter hypermethylation in plasma DNA as an indicator of disease recurrence in bladder cancer patients. Clin Cancer Res. 2002;8:980–5.
Melki JR, Vincent PC, Clark SJ. Concurrent DNA hypermethylation of multiple genes in acute myeloid leukemia. Cancer Res. 1999;59:3730–40.
Garcia MJ, Martinez-Delgado B, Cebrian A, et al. Different incidence and pattern of p15INK4b and p16INK4a promoter region hypermethylation in Hodgkin’s and CD30-Positive non-Hodgkin’s lymphomas. Am J Pathol. 2002;161:1007–13.
Zou HZ, Yu BM, Wang ZW, et al. Detection of aberrant p16 methylation in the serum of colorectal cancer patients. Clin Cancer Res. 2002;8:188–91.
Silva JM, Dominguez G, Garcia JM, et al. Presence of tumor DNA in plasma of breast cancer patients: clinicopathological correlations. Cancer Res. 1999;59:3251–6.
Morrissey C, Martinez A, Zatyka M, et al. Epigenetic inactivation of the RASSF1A 3p21.3 tumor suppressor gene in both clear cell and papillary renal cell carcinoma. Cancer Res. 2001;61:7277–81.
Kwong J, Lo KW, To KF, et al. Promoter hypermethylation of multiple genes in nasopharyngeal carcinoma. Clin Cancer Res. 2002;8:131–7.
Gonzalez-Gomez P, Bello MJ, Alonso ME, et al. CpG island methylation status and mutation analysis of the RB1 gene essential promoter region and protein-binding pocket domain in nervous system tumors. Br J Cancer. 2003;88:109–14.
Pietersen AM, Horlings HM, Hauptmann M, Langerod A, Ajouaou A, Cornelissen-Steijger P, Wessels LF, Jonkers J, van de Vijver MJ, van Lohuizen M. EZH2 and BMI1 inversely correlate with prognosis and TP53 mutation in breast cancer. Breast Cancer Res. 2008;10:R109.
Kim JH, Yoon SY, Jeong SH, Kim SY, Moon SK, Joo JH, Lee Y, Choe IS, Kim JW. Overexpression of Bmi-1 oncoprotein correlates with axillary lymph node metastases in invasive ductal breast cancer. Breast. 2004;13:383–8.
Chang MJ, Wu H, Achille NJ, Reisenauer MR, Chou CW, Zeleznik-Le NJ, Hemenway CS, Zhang W. Histone H3 lysine 79 methyltransferase Dot1 is required for immortalization by MLL oncogenes. Cancer Res. 2010;70:10234–42.
Tatum D, Li S. Evidence that the histone methyltransferase Dot1 mediates global genomic repair by methylating histone H3 on lysine 79. J Biol Chem. 2011;286:17530–5.
Kleer CC, Cao Q, Varambally S, Shen R, Ota I, Tomlins SA, Ghosh D, Sewalt RG, Otte AP, Hayes DF, Sabel MS, Livant D, Weiss SJ, Rubin MA, Chinnaiyan AM. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc Natl Acad Sci U S A. 2003;100:11606–11.
Suvà ML, Riggi N, Janiszewska M, Radovanovic I, Provero P, Stehle JC, Baumer K, Le Bitoux MA, Marino D, Cironi L. EZH2 is essential for glioblastoma cancer stem cell maintenance. Cancer Res. 2009;69:9211.
Tapia C, Zlobec I, Schneider S, Kilic E, Güth U, Bubendorf L, Kim S. Deletion of the inhibitor of growth 4 (ING4) tumor suppressor gene is prevalent in human epidermal growth factor 2 (HER2)-positive breast cancer. Hum Pathol. 2011;42:983.
Gunduz M, Nagatsuka H, Demircan K, Gunduz E, Cengiz B, Ouchida M, Tsujigiwa H, Yamachika E, Fukushima K, Beder L. Frequent deletion and down-regulation of ING4, a candidate tumor suppressor gene at 12p13, in head and neck squamous cell carcinomas. Gene. 2005;356:109–17.
Liu G, Bollig-Fischer A, Kreike B, van de Vijver MJ, Abrams J, Ethier SP, Yang ZQ. Genomic amplification and oncogenic properties of the GASC1 histone demethylase gene in breast cancer. Oncogene. 2009;28:4491–500.
Vinatzer U, Gollinger M, Mullauer L, Raderer M, Chott A, Streubel B. Mucosa-associated lymphoid tissue lymphoma: novel translocations including rearrangements of ODZ2, JMJD2C, and CNN3. Clin Cancer Res. 2008;14:6426–31.
Xiang Y, Zhu Z, Han G, Lin H, Xu L, Chen CD. JMJD3 is a histone H3K27 demethylase. Cell Res. 2007;17:850–7.
Lim S, Janzer A, Becker A, Zimmer A, Schule R, Buettner R, Kirfel J. Lysine-specific demethylase 1 (LSD1) is highly expressed in ER-negative breast cancers and a biomarker predicting aggressive biology. Carcinogenesis. 2010;31:512–20.
Wang Y, Zhang H, Chen Y, Sun Y, Yang F, Yu W, Liang J, Sun L, Yang X, Shi L. LSD1 is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell. 2009;138:660–72.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD, Sallan SE, Lander ES, Golub TR, Korsmeyer SJ. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet. 2002;30:41–7.
Corral J, Lavenir I, Impey H, Warren AJ, Forster A, Larson TA, Bell S, McKenzie AN, King G, Rabbitts TH. An Mll-AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: a method to create fusion oncogenes. Cell. 1996;85:853–61.
Taketani T, Taki T, Nakamura H, Taniwaki M, Masuda J, Hayashi Y. NUP98-NSD3 fusion gene in radiation-associated myelodysplastic syndrome with t(8;11)(p11;p15) and expression pattern of NSD family genes. Cancer Genet Cytogenet. 2009;190:108–12.
Wang GG, Allis CD, Chi P. Chromatin remodeling and cancer, part I: covalent histone modifications. Trends Mol Med. 2007;13:363–72.
Ceol CJ, Houvras Y, Jane-Valbuena J, Bilodeau S, Orlando DA, Battisti V, Fritsch L, Lin WM, Hollmann TJ, Ferré F. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature. 2011;471:513–7.
Balasubramanyam K, Swaminathan V, Ranganathan A, Kundu TK. Small molecule modulators of histone acetyltransferase p300. J Biol Chem. 2003;278:19134–40.
Sun Y, Jiang X, Chen S, Price BD. Inhibition of histone acetyltransferase activity by anacardic acid sensitizes tumor cells to ionizing radiation. FEBS Lett. 2006;580:4353–6.
Candido EP, Reeves R, Davie JR. Sodium butyrate inhibits histone deacetylation in cultured cells. Cell. 1978;14:105–13.
Sealy L, Chalkley R. The effect of sodium butyrate on histone modification. Cell. 1978;14:115–21.
Heltweg B, Gatbonton T, Schuler AD, Posakony J, Li H, Goehle S, Kollipara R, Depinho RA, Gu Y, Simon JA, Bedalov A. Antitumor activity of a small-molecule inhibitor of human silent information regulator 2 enzymes. Cancer Res. 2006;66:4368–77.
Balasubramanyam K, Varier RA, Altaf M, Swaminathan V, Siddappa NB, Ranga U, Kundu TK. Curcumin, a novel p300/CREB-binding proteinspecific inhibitor of acetyltransferase, represses the acetylation of histone/nonhistone proteins and histone acetyltransferase-dependent chromatin transcription. J Biol Chem. 2004;279:51163–71.
Balasubramanyam K, Altaf M, Varier RA, Swaminathan V, Ravindran A, Sadhale PP, Kundu TK. Polyisoprenylated benzophenone, garcinol, a natural histone acetyltransferase inhibitor, represses chromatin transcription and alters global gene expression. J Biol Chem. 2004;279:33716–26.
Kang J, Chen J, Shi Y, Jia J, Zhang Y. Curcumin induced histone hypoacetylation: the role of reactive oxygen species. Biochem Pharmacol. 2005;69:1205–13.
Bianchini F, Vainio H. Allium vegetables and organosulfur compounds: do they help prevent cancer? Environ Health Perspect. 2001;109:893–902.
Nian H, Delage B, Ho E, Dashwood RH. Modulation of histone deacetylase activity by dietary isothiocyanates and allyl sulfides: studies with sulforaphane and garlic organosulfur compounds. Environ Mol Mutagen. 2009;50:213–21.
Olaharski AJ, Rine J, Marshall BL, Babiarz J, Zhang L, Verdin E, Smith MT. The flavoring agent dihydrocoumarin reverses epigenetic silencing and inhibits sirtuin deacetylases. PLoS Genet. 2005;1:e77.
Li Y, Li X, Guo B. Chemopreventive agent 3,3′-diindolylmethane selectively induces proteasomal degradation of class I histone deacetylases. Cancer Res. 2010;70:646–54.
Nandakumar V, Vaid M, Katiyar SK. (−)-Epigallocatechin-3-gallate reactivates silenced tumor suppressor genes, Cip1/p21 and p16INK4a, by reducing DNA methylation and increasing histones acetylation in human skin cancer cells. Carcinogenesis. 2011;32:537–44.
Padhye S, Ahmad A, Oswal N, Sarkar FH. Emerging role of Garcinol, the antioxidant chalcone from Garcinia indica Choisy and its synthetic analogs. J Hematol Oncol. 2009;2:38.
Kikuno N, Shiina H, Urakami S, Kawamoto K, Hirata H, Tanaka Y, Majid S, Igawa M, Dahiya R. Genistein mediated histone acetylation and demethylation activates tumor suppressor genes in prostate cancer cells. Int J Cancer. 2008;123:552–60.
Myzak MC, Dashwood WM, Orner GA, Ho E, Dashwood RH. Sulforaphane inhibits histone deacetylase in vivo and suppresses tumorigenesis in Apc-minus mice. FASEB J. 2006;20:506–8.
Presta LG, Chen H, O’Connor SJ, Chisholm V, Meng YG, Krummen L, Winkler M, Ferrara N. Humanization of an anti-vascular endothelial growth factor monoclonal antibody for the therapy of solid tumors and other disorders. Cancer Res. 1997;57:4593–9.
Chen PC, Cheng HC, Wang J, Wang SW, Tai HC, Lin CW, Tang CH. Prostate cancer-derived CCN3 induces M2 macrophage infiltration and contributes to angiogenesis in prostate cancer microenvironment. Oncotarget. 2014;5:1595–608.
Jeong E, Koo JE, Yeon SH, Kwak MK, Hwang DH, Lee JY. PPARδ deficiency disrupts hypoxia-mediated tumorigenic potential of colon cancer cells. Mol Carcinog. 2014;53:926–37.
Abolhassani A, Riazi GH, Azizi E, Amanpour S, Muhammadnejad S, Haddadi M, Zekri A, Shirkoohi R. FGF10: Type III epithelial mesenchymal transition and invasion in breast cancer cell lines. J Cancer. 2014;5:537–47.
Salgia R. Fibroblast growth factor signaling and inhibition in non-small cell lung cancer and their role in squamous cell tumors. Cancer Med. 2014;3:681–92.
Corn PG, Wang F, McKeehan WL, Navone N. Targeting fibroblast growth factor pathways in prostate cancer. Clin Cancer Res. 2013;19:5856–66.
Schulze D, Plohmann P, Höbel S, Aigner A. Anti-tumor effects of fibroblast growth factor-binding protein (FGF-BP) knockdown in colon carcinoma. Mol Cancer. 2011;10:144.
Ren W, Liu Y, Wan S, Fei C, Wang W, Chen Y, Zhang Z, Wang T, Wang J, Zhou L, Weng Y, He T, Zhang Y. BMP9 inhibits proliferation and metastasis of HER2-positive SK-BR-3 breast cancer cells through ERK1/2 and PI3K/AKT pathways. PLoS One. 2014;9:e96816.
Wang LK, Hsiao TH, Hong TM, Chen HY, Kao SH, Wang WL, Yu SL, Lin CW, Yang PC. MicroRNA-133a suppresses multiple oncogenic membrane receptors and cell invasion in non-small cell lung carcinoma. PLoS One. 2014;9:e96765.
Xu Y, Pasche B. TGF-beta signaling alterations and susceptibility to colorectal cancer. Hum Mol Genet. 2007;1:R14–20.
Li H, Zhang B, Liu Y, Yin C. EBP50 inhibits the migration and invasion of human breast cancer cells via LIMK/cofilin and the PI3K/Akt/mTOR/MMP signalling pathway. Med Oncol. 2014;31:162.
Scagliotti GV, Selvaggi G, Novello S, Hirsch FR. The biology of epidermal growth factor receptor in lung cancer. Clin Cancer Res. 2004;10:4227s–32.
Bhat FA, Sharmila G, Balakrishnan S, Arunkumar R, Elumalai P, Suganya S, Raja Singh P, Srinivasan N, Arunakaran J. Quercetin reverses EGF-induced epithelial to mesenchymal transition and invasiveness in prostate cancer (PC-3) cell line via EGFR/PI3K/Akt pathway. J Nutr Biochem. 2014;S0955-2863:00149-1.
Ien GS, Wu MS, Bien MY, Chen CH, Lin CH, Chen BC. Epidermal growth factor stimulates nuclear factor-κB activation and heme oxygenase-1 expression via c-Src, NADPH oxidase, PI3K, and Akt in human colon cancer cells. PLoS One. 2014;9:e104891.
Karamysheva AF. Mechanisms of angiogenesis. Biochemistry (Mosc). 2008;73:751–62.
Klagsbrun M, Moses MA. Molecular angiogenesis. Chem Biol. 1999;6:R217–24.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Jha, N.K. et al. (2016). Epigenetics and Angiogenesis in Cancer. In: Mishra, M., Bishnupuri, K. (eds) Epigenetic Advancements in Cancer. Springer, Cham. https://doi.org/10.1007/978-3-319-24951-3_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-24951-3_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24949-0
Online ISBN: 978-3-319-24951-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)