Abstract
Colorectal cancer (CRC) is one of the most common types of human cancer with high cancer-related morbidity and mortality rates. The development and clinical validation of novel therapeutic avenues have improved the clinical outcome, but metastatic CRC still remains an incurable disease in most cases. The interest in discovering novel pathophysiological drivers in CRC is intensively ongoing and the search for novel biomarkers for early diagnosis, for patient’s stratification for prognostic purposes or for predicting treatment response are warranted. microRNAs are small RNA molecules that regulate the expression of larger messenger RNA species by different mechanisms with the final consequence to provide a fine tuning tool for global gene expression patterns. First discovered in worms, around 15 years ago it became clear that microRNAs are also existing in humans and that they are widely involved in human carcinogenesis. Within the last years, tremendous progress in the understanding of microRNAs and their role in CRC carcinogenesis has been developed. In this book chapter, several examples of previously identified microRNAs and how they influence colorectal carcinogenesis will be discussed. The information starting at the underlying molecular mechanisms towards clinical applications will be depicted and an overview what great potential these small molecules might carry in future colorectal cancer medicine, will be discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
In their seminal review paper “The hallmarks of Cancer,” D. Hanahan and R. Weinberg described the traits that most forms of cancer have in common [1]. Colorectal cancer (CRC) is determined by these hallmarks, which were later expanded upon in 2011 to include the so-called emerging hallmark capabilities. The exact role of the regulatory circuits regulated by epigenetic mechanisms including microRNAs (miRNAs) in the context of the eight hallmarks has yet to be determined. However, at present, dozens of regulatory miRNAs have been explored and discovered and almost all of them demonstrate differential expression and effects that are involved in the pathogenesis of CRC [2].
In general, CRC is the third most commonly diagnosed cancer and the second leading cause of cancer-related death. The high mortality rates are mainly related to the development of metastatic disease, which is associated with only a 12 % 5-year survival rate [3]. Therapeutic decision-making mainly relies on the tumor stage, ranging from surgical treatment alone for localized stages to a multimodal treatment strategy in advanced forms. For several years now, the therapeutic backbone of all current treatment schedules has contained 5-fluorouracil and it is frequently used in combination with oxaliplatin or irinotecan [4], both classical cytotoxic chemotherapeutic drugs. In addition, within the last 10 years monoclonal antibodies against the vascular endothelial growth factor-A (bevacizumab) or the epidermal growth factor receptor (cetuximab and panitumumab), as well as protein-traps (aflibercept) or small molecules (regorafnib) against other soluble or membrane-located angiogenic factors have extended the progression-free and overall survival time for CRC patients [5–7].
In the following pages, we summarize the involvement of miRNAs in the pathogenesis of CRC, relating them to the hallmarks of cancer and highlighting their potential clinical utility in CRC patients. There are different modes of miRNA functioning described in the literature. Apart from the basic approach of a single miRNAs -mRNA interaction, there is the broad approach of one miRNA influencing many mRNAs. Additionally, not only are miRNAs able to target transcripts, but they can also interact with other functionally related miRNAs to obtain certain effects. Adding to their complexity, miRNAs can belong to a cluster of miRNAs which regulates similar transcription factors and target several proteins within a defined pathway [8]. Knowing that miRNAs function in a network explains the involvement of one miRNA in many pathways as well as its ability to cause various effects as demonstrated in the different following topics. In the following sections, our main goal is to exemplify the involvement of miRNAs in CRC; however, these examples are by no means exhaustive.
MiRNAs and Their Relationship to the Hallmarks of Cancer
Regulation of Cell Proliferation and Evasion of Growth Suppressors
The most fundamental characterization of cancer cells is their ability of uncontrolled proliferation. As the growth of normal cells is tightly controlled, healthy cells are able to influence cell growth and antiproliferative signals through strict cell cycle regulations in order to guarantee a normal tissue architecture and function. Malfunction of this fundamental process can happen when, for example, tumor suppressor genes become deactivated. MiRNAs can function as tumor suppressor-targeting oncomirs. Among other oncogenes, K-ras is the most commonly dysregulated proto-oncogene in CRC patients and influences cell proliferation, but it is also prone as a target for let-7a, mir-143, and mir-133 [9]. Another example of a K-ras-targeting miRNA, which thereby influences cell growth in CRC, is miR-96-5p [10]. Elevated miR-21 expression leads to increased cell proliferation and is one of the most important oncogenic miRNAs that is relevant in most cancer types [11]. DHFR, directly targeted by miR-24, miR-192, and miR-215, is an S-phase-specific enzyme, converting dihydrofolate to tetrahydrofolate. This is essential for the synthesis of purin and thymidylate and partly influences cell proliferation [12]. MiR-145 affects cell growth and its tumor suppressor activity is mediated through different targets. These include the inhibition of the oncogenic insulin receptor substrate-1 (IRS-1), c-Myc, Yamaguchi sarcoma viral oncogene homolog 1 (YES1), signal transducer and activator of transcription (STAT1) and, as recently reported, Friend leukemia integration 1 (FLI1) [13].
MiR-30a-5p suppresses tumor growth in colon carcinoma by affecting denticleless homolog (DTL) [14]. By influencing macrophage migration inhibitory factor (MIF), upregulated miR-451 leads to reduced proliferation in gastrointestinal cancer [15]. MiR-675 influences CRC tumor growth via downregulation of its target pRB [16]. In summary, miRNAs that influence this hallmark can be useful as prognostic factors or as therapeutic targets to improve clinical outcomes in CRC patients.
Evasion of Apoptosis
Apoptosis is the process of programmed cell death that occurs in multicellular organisms, regulated by pro- and anti-apoptotic factors. Cancer cells can interrupt this controlled process of cell death and thereby promote cell survival. The central apoptosis-regulating gene in the miRNA-protein-coding gene network of CRC is the anti-apoptotic factor Bcl-2. Bcl-2 is directly targeted by the tumor suppressors miR-195, miR-129, miR-365, and miR-143 [17–20]. Other suppressors of Bcl-2 include miR-34a and BAG3 (Bcl-2-associated athanogene 3), which is linked with miR-345 and miR-491 and which is also promoted by the frequently overexpressed BCL-xl [21–23]. The central miRNAs for apoptosis are considered to be those of the miR-17-92 cluster, particularly oncomir-1, which has wide influence in CRC apoptosis processes. MiR-17 and miR-92 take part in a negative feedback loop with the E2F1/3 transcription factors, including c-Myc. Also, the BcL-2 like 11 gene, the proapoptotic factor BNIP2 and BIM are putative targets of miR-17-92 and influence apoptosis in CRC [24–26]. The expression of the oncomir miR-21 is supposed to be induced by CD24 via Src signaling and the programmed cell death protein 4 (PDCD4) influences CRC cells in apoptosis-related processes [27, 28]. MiR-30a-5p induces apoptosis by binding to the mRNA of denticleless protein homolog (DTL) [14].
MiR-218 promotes apoptosis by downregulating BMI-1 and was found to be significantly lower in CRC when compared to adjacent normal tissue [29]. In addition, overexpression of miR-429 and miR-96 represses apoptosis by directly targeting SOX2 and the transcription factor CHES1, respectively [30].
Regulating Immortalization or Senescence
Immortalization is one of the first steps a normal cell takes towards malignancy and requires at least two to three genetic events, typically ones that affect the p53 and/or pRb pathway. More specifically, replicative senescence is modulated by key regulatory pathways. These pathways can be categorized as follows: pRb/p53 activity, telomerase maintenance, epigenetic modulation, miRNAs regulation, and oxidative stress response [31]. A direct target of p53 is the activation of transcription of the miR-34 family. Overexpression of miR-34a represses SIRT1, which deacetylates p53 and allows for increased p53 activity. Additionally, during B-RAF-induced senescence, which is independent of p53, a member of E-twenty-six oncogene family of transcription factors (ELK1) activates miR-34a expression. MiR-34 again represses Myc, which provokes senescence [32, 33]. Specific to CRC, transient introduction of miR-34a causes complete suppression of cell proliferation and induces senescence-like growth arrest through modulation of the E2F signaling pathway [34].
Induction of Angiogenesis
Once a primary tumor has reached a certain size or cancer cells have spread to distant organs, they require and therefore induce the generation of new blood vessels to ensure the supply of oxygen and nutrients which are necessary for further tumor growth. In this process, many pro- and antiangiogenic factors are involved, which are again targets of certain miRNAs as several studies have indicated [35]. Hypoxia is one of the dominant drivers for neovascularization in tumor cells and HIF-1 protein plays a central role in hypoxia-activated gene expression. In CRC, miR-145 and miR-107 regulate HIF-1 by targeting p70S6K1 [36, 37]. MiR-17-92 and miR-194 were found to promote angiogenesis by regulating p53 in colon cancer. Both miRNAs repress TSP-1 (thrombospondin-1), which acts as a barrier to neovascularization in CRC [38, 39]. Additionally, the polycistronic miR-17-92 cluster produces six mature miRNAs (mir-17, mir-18a, mir-19a, mir-20a, mir-19b-1, and mir-92-1) and can coordinate multiple functions in tumorigenesis. Among other methods, it is activated by the Myc oncogene and stimulates angiogenesis by inhibiting the TGF-β pathway [40].
MiR-27b plays an important role in angiogenesis and controls the fine balance between stimulators and suppressors of endothelial cell proliferation, migration, and differentiation by targeting the endogenous angiogenesis inhibitor SEMA6A or by controlling EfnB2, EfnB4, Flt1, and Flt4 [9, 41]. MiR-126, an endothelial cell-restricted miRNA, mediates developmental angiogenesis in vivo and enhances the pro-angiogenic actions of VEGF and FGF. In addition, by repressing the intracellular inhibitor of the angiogenic signaling Spred-1, miR-126 promotes blood vessel formation [42].
Migration Invasion and Metastasis
The major cause of death in CRC patients arises from migration, invasion, and metastasis of the primary tumor cells. This two-step system is followed by proliferation and colonization of the tumor cells into their new secondary site. Leading mechanisms associated with these two steps and the mechanism of metastasis are the epithelial-mesenchymal transition (EMT) and mesenchymal to epithelial transition (MET). Vimentin, β-Catenin, TCF8-ZEB1, E-Cadherin, Snail, and Slug are established EMT markers [43]. In CRC, miR-21 and miR-31 act as downstream effectors in the TGF-β /Wnt signaling pathway, which is one of the prominent pathways in EMT [44] MiR-574-5p and miR-17 downregulate Qki6/7 and P130, which inversely correlate with β-Catenin [45]. In CRC, also a decrease in expression of the miR-200 family (for instance mir-200a and mir-200c), a master regulator of the epithelial phenotype, represses EMT by targeting ZEB1/2. ZEB1/2 upregulates Vimentin and downregulates E-cadherin [46–48]. MiR-147 was found to induce MET through the TGF-β signaling pathway [49]. The elevated expression of miR-103/107 is responsible for local invasion and liver metastasis and these miRNAs induce their effects through DAPK and KLF4 [50]. Prospero Homebox 1 (PROX1) inhibits E-cadherin via miR-9 [51]. Mucin-1, which is frequently upregulated in CRC patients, is a metastasis gene associated with cell invasion and metastasis and is targeted by miR-145 [52].
Further studies which investigate the role of miRNAs in metastasis are supported and are highly desired, as currently about 90 % of patients diagnosed with metastatic CRC die due to metastases [9, 53, 54].
Promotion of Genomic Instability
Calin et al. have shown that miRNAs are frequently clustered in fragile sites of chromosomes or regions of genomic instability . These areas are often associated with various human cancers, including CRC. This association between the location of miRNAs and chromosomal aberrations is significant, as it leads to chromosomal abnormalities, which in turn results in the disruption of miRNA expression. It is currently known that the abnormal expression of miRNAs plays a central role in cancer progression [55–57]. The first described miRNAs in CRC influenced by chromosomal rearrangements belong to the miR-145/142 cluster, which is located on 5q32 and is affected by a deletion of the 5q32 band [58]. Other changes involve miR-21 (7q23.2, 3′ÚTR of the vacuole membrane protein 1) and mirR-155 (21q21.3) [59]. Besides the fact that miRNAs are frequently affected by chromosomal aberration, they can influence chromosomal instability on their own by targeting different protein-coding genes (see also to an excellent review recently published) [60].
Reprogramming of Energy Metabolism
Particularly in gastrointestinal malignancies, significant malnutrition accompanies malignant processes in approximately 30–50 % of deaths. Cancer cachexia is a complex syndrome characterized by progressive tissue depletion, increased metabolic expenditure, and dysfunctional metabolic processes [61]. “The Warburg effect” may be an approach to explain these dramatic symptoms. This metabolic phenotype in cancer cells allows the cells, even in the presence of oxygen, to shift their ATP generation from oxidative phosphorylation to glycolysis. Contrary to normal cells, cancer cells predominantly generate energy from aerobic glycolysis. Incoming glucose is converted to lactate rather than being metabolized through oxidative phosphorylation. This reprogramming of energy metabolism in the cell allows ATP to be generated more rapidly in comparison to oxidative phosphorylation, but production is less efficient in respect to the amount of ATP produced per unit of glucose consumed. Consequently, cancer cells need an extremely high rate of glucose uptake to fulfill their energy demand [2, 62]. Recent studies have shown that miRNAs play important roles in CRC energy metabolism. MiR-26a regulates glucose metabolism of CRC cells by targeting the pyruvate dehydrogenase protein X component (PDHX), which inhibits the conversion of pyruvate to acetyl CoA in the tricarboxylic acid cycle [63]. The adaptor protein p66Shc, also found in CRC, has the potential to respond to energy status changes and regulate mitogenic signaling [64].
Evasion of Immune System and Inflammation
Inflammation-associated CRC is mainly colitic cancer and develops in patients with inflammatory bowel disease (IBD) or celiac disease [65, 66]. Most of the changes in miRNA expression observed in inflammatory tissues are likely the result of immune cells participating in hematopoietic tumorigenesis. For this reason, it is important to examine changes in miRNA, in immune and epithelial cells when investigating the various miRNA functions in inflammatory-associated cancer development. During inflammation, miRNA expression can be altered in epithelial cells and miRNAs can adopt a tumor suppressor function. In CRC, genomic locations of lethal-7 (let-7) family members, miRNAs targeting the RAS family and c-MYC family, are frequently deleted [55]. Also, IL-6, a cytokine frequently produced in cancers cells, is directly inhibited by let-7 [67]. Recent studies have shown a direct link between high levels of miR-155 and the development of gastric and colon cancers. MiR-155 is associated with hematopoiesis and the regulation of lymphocyte homeostasis and tolerance, which is substantially influenced by a bic/microRNA-155 interaction in B and T cells. In addition, some genes involved in normal immune functions are regulated by miR-155, including cytokines (IL-4), chemokines (CCL5) and transcription factors (c-Maf) [68, 69].
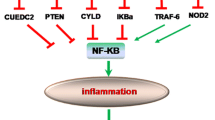
A positive feedback loop underlying the epigenetic switch which links inflammation to cancer was observed in CRC. STAT3, a transcription factor and downstream target of IL-6, activates miR-21 and miR-181b-1. Both miRNAs inhibit tumor suppressors (PTEN and CYLD), leading to increased NF-κB activity, which is required to support the transformed switch [70]. MiR-31 is associated with the stepwise transformation from IBD to IBD-related neoplasia by regulating the expression of factor inhibiting hypoxia inducible factor 1 [71]. Further studies will be required to improve our understanding of the role of tumor-related miRNAs in inflammation-associated cancer development [72].
Clinical Implications of miRNAs in Colorectal Cancer
MiRNAs as Diagnostics
Circulating miRNAs
The carcinoembryonic antigen (CEA) and carbohydrate antigen 19-9 (CA19-9) are widely-used blood-based biomarkers for CRC detection. Nevertheless, for early-stage CRC, the sensitivity and specificity seems to be insufficient [73]. Therefore, novel noninvasive biomarkers are urgently required for the advancement of diagnostics in CRC. Recent studies have shown a high amount of circulating miRNAs in the blood which are able to withstand adverse and labile conditions (pH, temperature, multiple freeze/thaw cycles) and still show consistent expression levels [74]. It has been hypothesized that most extracellular miRNAs are part of complexes with the Argonaute2 protein , derived from dead or dying cells. In this form, they could serve as a reflection of the underlying disease or help monitor pathological changes during the clinical course of disease. Due to the relatively easy access to plasma and serum, circulating miRNAs are one of the most promising biomarkers for cancer detection and prognosis [75].
Currently, there are several ways of taking advantage of circulating miRNAs. They have been used either alone or in a set of miRNAs as well as in combination with well-established biomarkers to enhance the predictive accuracy of prognostic factors/models . Almost 5 years ago, the overrepresentation of miRNAs in preoperative serum samples of CRC patients was described for the first time and a sensitivity of 89 % and specificity of 70 % was reported for miR-92. One year later, an independent group confirmed these results of miR-92 as a potential noninvasive biomarker for CRC diagnosis with even higher sensitivity and specificity values [76, 77]. In another study, the levels of miR-23a and miR-1246 in exosomes showed a high sensitivity for stage I CRC samples of 95 % and 90 %, respectively, in comparison to the inferior sensitivities of CA19-9 (10 %) and CEA (15 %) for stage I samples [78]. Biomarkers detecting TNM stage I or II CRC may bear the highest potential for reducing the mortality and overall health burden, as tumors in these stages can be removed by surgery alone and early detection will lead to high cure rates.
Another impressive clinical finding was that miR-200c levels were significantly higher in stage IV than in stage I–III CRCs and showed a positive correlation with lymph node metastases, distant metastases, and tumor recurrence [79]. MiR-29a levels in serum have been proposed as a powerful tool to detect liver metastases, reaching a sensitivity and specificity of 75 % [80].
Wang et al. established a biomarker profile based on six serum miRNAs. In this panel, miR-21 and let-7 g were described as upregulated markers and miR-181b, miR-92a, and miR-203 as downregulated markers. Interestingly, this panel was more accurate for CRC diagnosis (sensitivity and specificity of 93 % and 91 %, respectively) than CEA and CA19-9 (sensitivity and specificity of 35 % and 23 %, respectively) using the same serum samples [81].
Another study showed that in combination with CEA levels, miR-141 enhanced and supplemented the ability to diagnose a subset of stage IV colon cancer patients [82]. One limitation of miRNAs as blood-based diagnostic markers for screening purposes is the high costs and long latency time of serum miRNAs [83].
Mucosal Colon Wash Fluid
Conventional colonoscopy is considered the gold standard for detecting CRC by visual and histological sampling. The wash fluid used in this procedure could possibly serve as an additional specimen for further diagnostic tests. Kamimae et al. collected DNA from the mucosal wash fluid of patients undergoing colonoscopy screening. They assessed methylation levels of miR-34b/c and found that this DNA fragment had the greatest correlation with the invasion depth of tumors (sensitivity 83 %, specificity 76 %). Further studies for optimization will be necessary; however the combination of endoscopy and DNA methylation analysis based on miR-34b/c levels may facilitate accurate preoperative staging of CRC and support the decision-making process to help avoid unnecessary surgery [84].
Feces
The fecal occult blood test (FOBT) detects hidden blood in patient stool. A positive result is not necessarily associated with CRC and may result from any bleeding occurring in the gastrointestinal system. On the other hand, a false negative test result is frequently the consequence of the low sensitivity of the assay. Although the sensitivity and specificity is not very high, the FOBT is widely used for colorectal tumor screening [85].
To improve the sensitivity, Yamazaki et al. investigated the usefulness of testing fecal miRNAs out of FOBT residuum and evaluated the best technical conditions for sufficient fecal miRNA extraction from FOBT for PCR analysis. They found that storage at 4C for 5 days is feasible without the loss of RNA quality for miRNA analysis [86] In general, it was determined that stool-based miRNAs are relatively stable and show highly reproducible detection rates. Various studies have reported different miRNAs which are differentially expressed in CRC patients in comparison to healthy volunteers. These reports include miR-143, miR-145 [87], miR-34a, miR-34b/c [88], miR-144* [89], and miR-92a [90] among the most promising stool-based biomarkers. Other potential stool-based miRNA biomarkers for the early detection of CRC include miR-4478 and miR-1295b-3p, as their expression levels are significantly lower when compared to healthy controls [91].
In addition to the discrimination power of detecting tumors, miR-135b, miR-221, and miR-18a were increased significantly in stool samples of advanced-stage CRC [92, 93].
Stool-based miRNAs can either be used as individual biomarkers or be integrated into currently existing marker panels. It is most likely the case that the earliest neoplastic changes in the expression pattern of specific miRNAs might be detected in the feces rather than in the blood, considering the increased number of exfoliated cancer cells shed in the colon from CRC patients. Although the listed results need further validation, fecal miRNAs might provide a promising and noninvasive method for the diagnosis of early colorectal neoplasia and thereby serve as diagnostic markers [94, 95].
microRNAs as Therapeutics
The applicability of miRNA-based therapy can be explained as a double-edged sword. On the one hand, modulation of a single miRNA offers the opportunity to target multiple genes and regulatory networks simultaneously. On the other hand, caution and careful design are necessary to prevent any unwanted off-target effects [75]. However, applying miRNAs to anticancer therapy could be efficient and the off-target effects might even be helpful. As in several preclinical models previously shown, there are at least two main strategies for miRNA-based therapy: the restoration of tumor suppressor miRNAs or the inhibition of oncogenic miRNAs. A good example of this, although not in cancer patients, is the anti-miR-122 drug , which has progressed to phase II clinical trials to treat the hepatitis C virus in humans [96].
Drug resistance leads to therapy failure, cancer relapse, and poor prognosis. In this context, miRNAs can also predict individual chemotherapy response and drugs targeting specific miRNAs can influence chemosensitivity in CRC patients. MiR-20, miR-130, miR-145, miR-216, and miR-372 serum levels are significantly upregulated in oxaliplatin chemotherapy-resistant CRC patients in comparison to chemosensitive patients [97]. MiR-215, miR-99a*, miR-196b, miR-450b-5p, and let-7e are associated with neoadjuvant chemoradiotherapy response [98]. MiR-10b has been involved as a predictor of 5-flourouracil-based chemotherapy resistance [99]. The promising results of preclinical models led to the initiation of the first miRNA-based clinical trial. Recently, it was reported that the data from a phase I clinical trial of a novel drug (MRX34) targeting miR-34 proposed a manageable safety profile in patients with advanced primary liver cancer (hepatocellular carcinoma), other solid tumors with liver metastasis and hematological malignancies (source: www.mirnarx.com ).
All the miRNAs described above, as well as many not referred to here, appear to form a network to coordinate and influence the regulation of colorectal carcinogenesis.
Many underlying mechanisms still remain largely unknown. Further research regarding the identification of novel miRNAs, their target genes, new drugs, or CRC characteristics will enhance our knowledge and will be another step towards a more individual and efficient addition to therapy for CRC patients.
References
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
Schmoll HJ, Van Cutsem E, Stein A, Valentini V, Glimelius B, Haustermans K, et al. ESMO Consensus Guidelines for management of patients with colon and rectal cancer. a personalized approach to clinical decision making. Ann Oncol. 2012;23:2479–516.
Stiegelbauer V, Perakis S, Deutsch A, Ling H, Gerger A, Pichler M. MicroRNAs as novel predictive biomarkers and therapeutic targets in colorectal cancer. World J Gastroenterol. 2014;20:11727–35.
Linnekamp JF, Wang X, Medema JP, Vermeulen L. Colorectal cancer heterogeneity and targeted therapy: a case for molecular disease subtypes. Cancer Res. 2015;75:245–9.
Ahmed S, Johnson K, Ahmed O, Iqbal N. Advances in the management of colorectal cancer: from biology to treatment. Int J Colorectal Dis. 2014;29:1031–42.
Hayes J, Peruzzi PP, Lawler S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol Med. 2014;20:460–9.
Xuan Y, Yang H, Zhao L, Lau WB, Lau B, Ren N, et al. MicroRNAs in colorectal cancer: small molecules with big functions. Cancer Lett. 2014;360(2):89–105.
Ress AL, Stiegelbauer V, Winter E, Schwarzenbacher D, Kiesslich T, Lax S, et al. MiR-96-5p influences cellular growth and is associated with poor survival in colorectal cancer patients. Mol Carcinog. 2014. doi:10.1002/mc.22218.
Selcuklu SD, Donoghue MT, Spillane C. miR-21 as a key regulator of oncogenic processes. Biochem Soc Trans. 2009;37:918–25.
McGuire JJ. Anticancer antifolates: current status and future directions. Curr Pharm Des. 2003;9:2593–613.
Schetter AJ, Okayama H, Harris CC. The role of microRNAs in colorectal cancer. Cancer J. 2012;18:244–52.
Baraniskin A, Birkenkamp-Demtroder K, Maghnouj A, Zollner H, Munding J, Klein-Scory S, et al. MiR-30a-5p suppresses tumor growth in colon carcinoma by targeting DTL. Carcinogenesis. 2012;33:732–9.
Bandres E, Bitarte N, Arias F, Agorreta J, Fortes P, Agirre X, et al. microRNA-451 regulates macrophage migration inhibitory factor production and proliferation of gastrointestinal cancer cells. Clin Cancer Res. 2009;15:2281–90.
Tsang WP, Ng EK, Ng SS, Jin H, Yu J, Sung JJ, et al. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis. 2010;31:350–8.
Karaayvaz M, Zhai H, Ju J. miR-129 promotes apoptosis and enhances chemosensitivity to 5-fluorouracil in colorectal cancer. Cell Death Dis. 2013;4:659.
Liu L, Chen L, Xu Y, Li R, Du X. microRNA-195 promotes apoptosis and suppresses tumorigenicity of human colorectal cancer cells. Biochem Biophys Res Commun. 2010;400:236–40.
Nie J, Liu L, Zheng W, Chen L, Wu X, Xu Y, et al. microRNA-365, down-regulated in colon cancer, inhibits cell cycle progression and promotes apoptosis of colon cancer cells by probably targeting Cyclin D1 and Bcl-2. Carcinogenesis. 2012;33:220–5.
Akao Y, Nakagawa Y, Hirata I, Iio A, Itoh T, Kojima K, et al. Role of anti-oncomirs miR-143 and -145 in human colorectal tumors. Cancer Gene Ther. 2010;17:398–408.
Tang JT, Wang JL, Du W, Hong J, Zhao SL, Wang YC, et al. MicroRNA 345, a methylation-sensitive microRNA is involved in cell proliferation and invasion in human colorectal cancer. Carcinogenesis. 2011;32:1207–15.
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007;26:745–52.
Nakano H, Miyazawa T, Kinoshita K, Yamada Y, Yoshida T. Functional screening identifies a microRNA, miR-491 that induces apoptosis by targeting Bcl-X(L) in colorectal cancer cells. Int J Cancer. 2010;127:1072–80.
Woods K, Thomson JM, Hammond SM. Direct regulation of an oncogenic micro-RNA cluster by E2F transcription factors. J Biol Chem. 2007;282:2130–4.
Diosdado B, van de Wiel MA, Terhaar Sive Droste JS, Mongera S, Postma C, Meijerink WJ, et al. MiR-17-92 cluster is associated with 13q gain and c-myc expression during colorectal adenoma to adenocarcinoma progression. Br J Cancer. 2009;101:707–14.
Chai H, Liu M, Tian R, Li X, Tang H. miR-20a targets BNIP2 and contributes chemotherapeutic resistance in colorectal adenocarcinoma SW480 and SW620 cell lines. Acta Biochim Biophys Sin. 2011;43:217–25.
Muppala S, Mudduluru G, Leupold JH, Buergy D, Sleeman JP, Allgayer H. CD24 induces expression of the oncomir miR-21 via Src, and CD24 and Src are both post-transcriptionally downregulated by the tumor suppressor miR-34a. PLoS One. 2013;8, e59563.
Leupold JH, Yang HS, Colburn NH, Asangani I, Post S, Allgayer H. Tumor suppressor Pdcd4 inhibits invasion/intravasation and regulates urokinase receptor (u-PAR) gene expression via Sp-transcription factors. Oncogene. 2007;26:4550–62.
He X, Dong Y, Wu CW, Zhao Z, Ng SS, Chan FK, et al. MicroRNA-218 inhibits cell cycle progression and promotes apoptosis in colon cancer by downregulating BMI1 polycomb ring finger oncogene. Mol Med. 2012;18:1491–8.
Li J, Du L, Yang Y, Wang C, Liu H, Wang L, et al. MiR-429 is an independent prognostic factor in colorectal cancer and exerts its anti-apoptotic function by targeting SOX2. Cancer Lett. 2013;329:84–90.
Leal JA, Feliciano A, Lleonart ME. Stem cell microRNAs in senescence and immortalization: novel players in cancer therapy. Med Res Rev. 2013;33:112–38.
Christoffersen NR, Shalgi R, Frankel LB, Leucci E, Lees M, Klausen M, et al. p53-independent upregulation of miR-34a during oncogene-induced senescence represses MYC. Cell Death Differ. 2010;17:236–45.
Yamakuchi M, Ferlito M, Lowenstein CJ. miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci U S A. 2008;105:13421–6.
Tazawa H, Tsuchiya N, Izumiya M, Nakagama H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci U S A. 2007;104:15472–7.
Hur K. MicroRNAs: promising biomarkers for diagnosis and therapeutic targets in human colorectal cancer metastasis. BMB Rep. 2015;48(4):217–22.
Xu Q, Liu LZ, Qian X, Chen Q, Jiang Y, Li D, et al. MiR-145 directly targets p70S6K1 in cancer cells to inhibit tumor growth and angiogenesis. Nucleic Acids Res. 2012;40:761–74.
Yamakuchi M, Lotterman CD, Bao C, Hruban RH, Karim B, Mendell JT, et al. P53-induced microRNA-107 inhibits HIF-1 and tumor angiogenesis. Proc Natl Acad Sci U S A. 2010;107:6334–9.
Lawler J. Thrombospondin-1 as an endogenous inhibitor of angiogenesis and tumor growth. J Cell Mol Med. 2002;6:1–12.
Sundaram P, Hultine S, Smith LM, Dews M, Fox JL, Biyashev D, et al. p53-responsive miR-194 inhibits thrombospondin-1 and promotes angiogenesis in colon cancers. Cancer Res. 2011;71:7490–501.
Dews M, Fox JL, Hultine S, Sundaram P, Wang W, Liu YY, et al. The myc-miR-17 ~ 92 axis blunts TGF{beta} signaling and production of multiple TGF{beta}-dependent antiangiogenic factors. Cancer Res. 2010;70:8233–46.
Melo SA, Kalluri R. Angiogenesis is controlled by miR-27b associated with endothelial tip cells. Blood. 2012;119:2439–40.
Wang S, Aurora AB, Johnson BA, Qi X, McAnally J, Hill JA, et al. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev Cell. 2008;15:261–71.
Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Invest. 2009;119:1420–8.
Cottonham CL, Kaneko S, Xu L. miR-21 and miR-31 converge on TIAM1 to regulate migration and invasion of colon carcinoma cells. J Biol Chem. 2010;285:35293–302.
Ji S, Ye G, Zhang J, Wang L, Wang T, Wang Z, et al. miR-574-5p negatively regulates Qki6/7 to impact beta-catenin/Wnt signalling and the development of colorectal cancer. Gut. 2013;62:716–26.
Davalos V, Moutinho C, Villanueva A, Boque R, Silva P, Carneiro F, et al. Dynamic epigenetic regulation of the microRNA-200 family mediates epithelial and mesenchymal transitions in human tumorigenesis. Oncogene. 2012;31:2062–74.
Pichler M, Ress AL, Winter E, Stiegelbauer V, Karbiener M, Schwarzenbacher D, et al. MiR-200a regulates epithelial to mesenchymal transition-related gene expression and determines prognosis in colorectal cancer patients. Br J Cancer. 2014;110:1614–21.
Chen ML, Liang LS, Wang XK. miR-200c inhibits invasion and migration in human colon cancer cells SW480/620 by targeting ZEB1. Clin Exp Metastasis. 2012;29:457–69.
Lee CG, McCarthy S, Gruidl M, Timme C, Yeatman TJ. MicroRNA-147 induces a mesenchymal-to-epithelial transition (MET) and reverses EGFR inhibitor resistance. PLoS One. 2014;9, e84597.
Chen HY, Lin YM, Chung HC, Lang YD, Lin CJ, Huang J, et al. miR-103/107 promote metastasis of colorectal cancer by targeting the metastasis suppressors DAPK and KLF4. Cancer Res. 2012;72:3631–41.
Lu MH, Huang CC, Pan MR, Chen HH, Hung WC. Prospero homeobox 1 promotes epithelial-mesenchymal transition in colon cancer cells by inhibiting E-cadherin via miR-9. Clin Cancer Res. 2012;18:6416–25.
Sachdeva M, Mo YY. MicroRNA-145 suppresses cell invasion and metastasis by directly targeting mucin 1. Cancer Res. 2010;70:378–87.
Muhammad S, Kaur K, Huang R, Zhang Q, Kaur P, Yazdani HO, et al. MicroRNAs in colorectal cancer: role in metastasis and clinical perspectives. World J Gastroenterol. 2014;20:17011–9.
Zhou JJ, Zheng S, Sun LF, Zheng L. MicroRNA regulation network in colorectal cancer metastasis. World J Biol Chem. 2014;5:301–7.
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A. 2004;101:2999–3004.
Calin GA, Croce CM. MicroRNAs and chromosomal abnormalities in cancer cells. Oncogene. 2006;25:6202–10.
Calin GA, Croce CM. Chromosomal rearrangements and microRNAs: a new cancer link with clinical implications. J Clin Invest. 2007;117:2059–66.
Michael MZ, O’Connor SM, van Holst Pellekaan NG, Young GP, James RJ. Reduced accumulation of specific microRNAs in colorectal neoplasia. Mol Cancer Res. 2003;1:882–91.
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A. 2006;103:2257–61.
Vincent K, Pichler M, Lee GW, Ling H. MicroRNAs, genomic instability and cancer. Int J Mol Sci. 2014;15:14475–91.
Palesty JA, Dudrick SJ. What we have learned about cachexia in gastrointestinal cancer. Digestive Dis. 2003;21:198–213.
Cairns RA, Harris IS, Mak TW. Regulation of cancer cell metabolism. Nat Rev Cancer. 2011;11:85–95.
Chen B, Liu Y, Jin X, Lu W, Liu J, Xia Z, et al. MicroRNA-26a regulates glucose metabolism by direct targeting PDHX in colorectal cancer cells. BMC Cancer. 2014;14:443.
Lebiedzinska-Arciszewska M, Oparka M, Vega-Naredo I, Karkucinska-Wieckowska A, Pinton P, Duszynski J, et al. The interplay between p66Shc, reactive oxygen species and cancer cell metabolism. Eur J Clin Invest. 2015;45 Suppl 1:25–31.
Bernstein CN, Blanchard JF, Kliewer E, Wajda A. Cancer risk in patients with inflammatory bowel disease: a population-based study. Cancer. 2001;91:854–62.
Elfstrom P, Granath F, Ye W, Ludvigsson JF. Low risk of gastrointestinal cancer among patients with celiac disease, inflammation, or latent celiac disease. Clin Gastroenterol Hepatol. 2012;10:30–6.
Iliopoulos D, Hirsch HA, Struhl K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell. 2009;139:693–706.
Tili E, Croce CM, Michaille JJ. miR-155: on the crosstalk between inflammation and cancer. Int Rev Immunol. 2009;28:264–84.
Rodriguez A, Vigorito E, Clare S, Warren MV, Couttet P, Soond DR, et al. Requirement of bic/microRNA-155 for normal immune function. Science. 2007;316:608–11.
Iliopoulos D, Jaeger SA, Hirsch HA, Bulyk ML, Struhl K. STAT3 activation of miR-21 and miR-181b-1 via PTEN and CYLD are part of the epigenetic switch linking inflammation to cancer. Mol Cell. 2010;39:493–506.
Olaru AV, Selaru FM, Mori Y, Vazquez C, David S, Paun B, et al. Dynamic changes in the expression of MicroRNA-31 during inflammatory bowel disease-associated neoplastic transformation. Inflamm Bowel Dis. 2011;17:221–31.
Chiba T, Marusawa H, Ushijima T. Inflammation-associated cancer development in digestive organs: mechanisms and roles for genetic and epigenetic modulation. Gastroenterology. 2012;143:550–63.
Duffy MJ, van Dalen A, Haglund C, Hansson L, Holinski-Feder E, Klapdor R, et al. Tumour markers in colorectal cancer: European Group on Tumour Markers (EGTM) guidelines for clinical use. Eur J Cancer. 2007;43:1348–60.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008;105:10513–8.
Ajit SK. Circulating microRNAs as biomarkers, therapeutic targets, and signaling molecules. Sensors. 2012;12:3359–69.
Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–81.
Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127:118–26.
Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, et al. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS One. 2014;9, e92921.
Toiyama Y, Hur K, Tanaka K, Inoue Y, Kusunoki M, Boland CR, et al. Serum miR-200c is a novel prognostic and metastasis-predictive biomarker in patients with colorectal cancer. Ann Surg. 2014;259:735–43.
Wang LG, Gu J. Serum microRNA-29a is a promising novel marker for early detection of colorectal liver metastasis. Cancer Epidemiol. 2012;36:e61–7.
Wang J, Huang SK, Zhao M, Yang M, Zhong JL, Gu YY, et al. Identification of a circulating microRNA signature for colorectal cancer detection. PLoS One. 2014;9, e87451.
Cheng H, Zhang L, Cogdell DE, Zheng H, Schetter AJ, Nykter M, et al. Circulating plasma MiR-141 is a novel biomarker for metastatic colon cancer and predicts poor prognosis. PLoS One. 2011;6, e17745.
Brase JC, Wuttig D, Kuner R, Sultmann H. Serum microRNAs as non-invasive biomarkers for cancer. Mol Cancer. 2010;9:306.
Kamimae S, Yamamoto E, Yamano HO, Nojima M, Suzuki H, Ashida M, et al. Epigenetic alteration of DNA in mucosal wash fluid predicts invasiveness of colorectal tumors. Cancer Prev Res (Phila). 2011;4:674–83.
Burch JA, Soares-Weiser K, St John DJ, Duffy S, Smith S, Kleijnen J, et al. Diagnostic accuracy of faecal occult blood tests used in screening for colorectal cancer: a systematic review. J Med Screen. 2007;14:132–7.
Yamazaki N, Koga Y, Yamamoto S, Kakugawa Y, Otake Y, Hayashi R, et al. Application of the fecal microRNA test to the residuum from the fecal occult blood test. Jpn J Clin Oncol. 2013;43:726–33.
Li JM, Zhao RH, Li ST, Xie CX, Jiang HH, Ding WJ, et al. Down-regulation of fecal miR-143 and miR-145 as potential markers for colorectal cancer. Saudi Med J. 2012;33:24–9.
Wu XD, Song YC, Cao PL, Zhang H, Guo Q, Yan R, et al. Detection of miR-34a and miR-34b/c in stool sample as potential screening biomarkers for noninvasive diagnosis of colorectal cancer. Med Oncol. 2014;31:894.
Kalimutho M, Del Vecchio BG, Di Cecilia S, Sileri P, Cretella M, Pallone F, et al. Differential expression of miR-144* as a novel fecal-based diagnostic marker for colorectal cancer. J Gastroenterol. 2011;46:1391–402.
Wu CW, Ng SS, Dong YJ, Ng SC, Leung WW, Lee CW, et al. Detection of miR-92a and miR-21 in stool samples as potential screening biomarkers for colorectal cancer and polyps. Gut. 2012;61:739–45.
Ghanbari R, Mosakhani N, Asadi J, Nouraee N, Mowla SJ, Poustchi H, et al. Decreased expression of fecal miR-4478 and miR-1295b-3p in early-stage colorectal cancer. Cancer Biomark. 2014;15(2):189–95.
Yau TO, Wu CW, Dong Y, Tang CM, Ng SS, Chan FK, et al. microRNA-221 and microRNA-18a identification in stool as potential biomarkers for the non-invasive diagnosis of colorectal carcinoma. Br J Cancer. 2014;111:1765–71.
Wu CW, Ng SC, Dong Y, Tian L, Ng SS, Leung WW, et al. Identification of microRNA-135b in stool as a potential noninvasive biomarker for colorectal cancer and adenoma. Clin Cancer Res. 2014;20:2994–3002.
Link A, Balaguer F, Shen Y, Nagasaka T, Lozano JJ, Boland CR, et al. Fecal MicroRNAs as novel biomarkers for colon cancer screening. Cancer Epidemiol Biomarkers Prev. 2010;19:1766–74.
Menendez P, Villarejo P, Padilla D, Menendez JM, Montes JA. Diagnostic and prognostic significance of serum microRNAs in colorectal cancer. J Surg Oncol. 2013;107:217–20.
Lanford RE, Hildebrandt-Eriksen ES, Petri A, Persson R, Lindow M, Munk ME, et al. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science. 2010;327:198–201.
Zhang J, Zhang K, Bi M, Jiao X, Zhang D, Dong Q. Circulating microRNA expressions in colorectal cancer as predictors of response to chemotherapy. Anticancer Drugs. 2014;25:346–52.
Svoboda M, Sana J, Fabian P, Kocakova I, Gombosova J, Nekvindova J, et al. MicroRNA expression profile associated with response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer patients. Radiat Oncol. 2012;7:195.
Nishida N, Yamashita S, Mimori K, Sudo T, Tanaka F, Shibata K, et al. MicroRNA-10b is a prognostic indicator in colorectal cancer and confers resistance to the chemotherapeutic agent 5-fluorouracil in colorectal cancer cells. Ann Surg Oncol. 2012;19:3065–71.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Ress, A.L., Perakis, S., Pichler, M. (2015). microRNAs and Colorectal Cancer. In: Santulli, G. (eds) microRNA: Cancer. Advances in Experimental Medicine and Biology, vol 889. Springer, Cham. https://doi.org/10.1007/978-3-319-23730-5_6
Download citation
DOI: https://doi.org/10.1007/978-3-319-23730-5_6
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-23729-9
Online ISBN: 978-3-319-23730-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)