Abstract
This paper presents a robust method to directly construct parametric representations of myocardial structure using a left ventricular (LV) finite element model customised to diffusion tensors derived from cardiac diffusion tensor magnetic resonance images (DTMRI). This method avoids the need to solve the eigenvector problem, and therefore avoids issues due to ambiguous eigenvector directions, and the non-uniqueness of eigenvectors in regions of isotropic diffusion. Finite element parameters describing the fibre orientations of a geometric model of the LV are directly fitted to diffusion tensors using non-linear least squares optimisation. The method was tested using ex vivo DTMRI data from a Wistar-Kyoto rat and compared against the conventional eigenvector analysis. Close agreement was found in most regions, except at some boundary locations, and in regions with low fractional anisotropy.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Finite element (FE) model-based descriptions of cardiac geometry and microstructure have become key components in personalised heart modelling frameworks designed for investigating the electrical [1–3], biomechanical [4–7], and energetic function of the heart [8, 9]. These models enable the integration of structural and functional data acquired using various imaging modalities, together with other measurements such as haemodynamic or electrophysiological recordings, to analyse the electro-mechanics of the heart on a subject-specific basis.
It is well established that cardiac shape and microstructural tissue organisation are important determinants of biomechanical function of the heart. While in vivo measurements of cardiac geometry are readily available via computed tomography (CT), magnetic resonance imaging (MRI) or ultrasound (US), in vivo microstructural measurements from the whole heart remain sparse and difficult to quantify. Diffusion tensor MRI (DTMRI) exploits the Brownian motion of the water molecules within myocardial tissue to determine local anisotropic diffusion in isolated heart preparations [10]. The direction of maximum water diffusion, represented by the primary eigenvector of the derived local diffusion tensor, has been found to correlate well with the local histologically-measured myofibre orientation [11, 12], and is often represented as an elevation angle (or fibre angle) with respect to the short-axis plane of the heart. There are several approaches for estimating fibre orientations from the diffusion tensor data. For example, shape predictors [13] or analyses of shape-based transformations [14] have been used to personalise fibre orientations. Some studies have proposed probabilistic streamlines incorporating uncertainty [15] or geodesic tractography [16], others have proposed descriptions based on the data from Streeter et al. [17, 18]. One of the most common techniques is the deterministic eigenanalysis, which estimates the fibre angle from the orientation of the primary eigenvector of the diffusion tensor. Spatial distributions of fibre angles are typically interpolated within FE models [3, 5, 6] although some studies have directly interpolated spatial distributions of the diffusion tensors in the log-Euclidean space to ensure that the interpolated tensors are positive-definite [4, 19] (which is not guaranteed when independently interpolating the tensor components). In either case, eigenanalyses of diffusion tensors are required to construct local microstructural material axes [4].
The use of the primary eigenvector alone to derive myocardial fibre angles has two main disadvantages. Firstly, water can diffuse equally in opposite directions, but the primary eigenvector arbitrarily represents just one of these directions. This representation can lead to large discontinuities in fibre angle distributions (see Fig. 1(a)), which can cause problems with FE interpolation of these data. In order to interpolate these spatially discontinuous data within FE models, the eigenvectors are usually phase-unwrapped according to a pre-defined smooth fibre field variation [6], however this method requires prior knowledge of the expected field, and does not eliminate all of the problems with these spatial discontinuities.
The second main issue with eigenanalysis of DTMRI is that in regions of apparent near-isotropic diffusion, the primary eigenvector may not reliably represent the local fibre orientation, as indicated in the inset of Fig. 1(b). Anisotropy in a diffusion process is generally quantified by fractional anisotropy (FA) which ranges from 0 (isotropic diffusion) to 1 (diffusion only along one axis) [20]. Low FA can arise from several factors, including low signal-to-noise ratio in the images [21], or that a single preferred direction does not adequately represent the local tissue microstructure. This can occur, for example, in some regions of myocardial infarct [22], or if there are multiple families of crossing fibres present in the tissue, such as near the inter-ventricular junction of the heart. In the case of crossing fibres, the assumption of a single direction to represent the fibres would need to be re-addressed. On the other hand, if regions of low FA are localised zones of poor image quality, then one could reduce the influence of these data during the fitting of fibre field parameters by incorporating a specialised weighting scheme, such as scaling the fitting error in each voxel by the FA value.
In this study, we present a workflow for parameterising myocardial fibre fields directly from the diffusion tensors without the need to compute eigenvectors or FA values for fibre orientation fitting. This framework not only circumvents issues associated with phase-unwrapping of eigenvectors prior to fibre fitting, but also helps to ensure that the interpolated fibre angles in regions with high FA are better representations of the diffusion tensors in those regions.
2 Methods
2.1 Experimental Data
The experimental study was approved by the Animal Ethics Committee of the University of Auckland and conforms to the National Institutes of Health Guide for the Care and Use of Laboratory Animals (NIH Publication No. 85–23).
The heart from a Wistar-Kyoto (WKY) rat was excised, perfused with St. Thomas’ cardioplegic solution to relax the heart, and then fixed using Bouin’s solution in an approximate end-diastolic state. One week later, DTMRI was performed using a 3D fast spin-echo pulse sequence on a Varian 4.7T MRI scanner. Each image set consisted of 11 or 12 short-axis slices with a thickness of 1.5 mm, and no gap between slices. Each slice contained one non-diffusion weighted anatomical image, and 30 diffusion weighted images. Other imaging parameters were as follows: TE = 15 ms; TR = 3 s; number of averages = 6; field of view (FOV) = 20 mm \(\times \) 20 mm \(\times \) 16 mm; in-plane resolution = 128 \(\times \) 64 voxels (zero-pad interpolated to 128 \(\times \) 128 voxels); in-plane voxel dimensions = 156 \(\upmu \)m \(\times \) 156 \(\upmu \)m.
2.2 Fibre Field Parameterisation Workflow
The following workflow (Fig. 2) was developed to parameterise a spatially varying myocardial fibre field directly from the diffusion tensors throughout the LV myocardium.
Step 1: Diffusion tensor calculation. From the non-diffusion weighted signals (\(S_0\)) and the diffusion weighted signals (\(S_k\)) captured in 30 non-collinear directions (i.e. \(k=1...30\)), a diffusion tensor, denoted \(\mathbf {D}\) (a symmetric tensor with six independent components), was estimated for each voxel by solving the logarithm of the diffusion equation (Eq. 1) proposed by Basser et al. [10].
where b is the diffusion weighting factor (1462 s/mm\(^2\) for this study). The outer-product \(\mathbf {Q}_{(k)}\) of the diffusion gradient direction (\(\mathbf {g}_{(k)}\)) was pre-calculated before the system of equations was solved using least-squares to determine the six independent components of \(\mathbf {D}\) at each voxel.
Step 2: Image segmentation. The analysis was limited to the LV for this study. The entire endocardial and epicardial surfaces of the LV were manually segmented from the non-diffusion images using MATLABFootnote 1. During the segmentation, papillary muscles and trabeculae were excluded. Three landmark points (the centroids of the LV base, LV apex, and RV base) were selected to construct an orthogonal cardiac coordinate system with the origin located one-third of the distance from base to apex along the long-axis of the LV, with which the x-axis was aligned. The y-axis was directed from the LV to RV centroids, and the z-axis was directed orthogonal to the x-axis and y-axis from the anterior to posterior of the LV.
Step 3: LV FE geometric model construction. To obtain a model representation of the LV geometry, a prolate ellipsoid-shaped 16-element (4 circumferential, 4-longitudinal and 1-transmural) hexahedral FE model was customised to the surface contours obtained from Step 2. The endocardial and epicardial surfaces of the model were simultaneously fitted using non-linear least squares to best match the corresponding surface data.
Step 4: Field-based parameterisation of LV fibre orientation. In order to parameterise the myofibre orientation field within the LV FE geometric model, we developed a new approach to estimate a smoothly continuous fibre field that best aligned with the diffusion tensors at all voxels within the LV. Firstly, we initialised the FE fibre field by setting the fibre angles (\(\theta _{(n)}\)) to be \(+60^\circ \) for the endocardial nodes, and \(-70^\circ \) for the epicardial nodes. Initial imbrication angles (\(\varphi _{(n)}\)) of all nodes were set to be \(0^\circ \). These angles were interpolated over the FE model using tricubic Hermite basis functions. Secondly, for each voxel (v), we determined its FE local coordinates within the LV geometric model, and, at that location, we computed the myofibre orientation (\(\mathbf {f}_{(v)}\)) defined by Euler angle rotation (using interpolated \(\theta \) and \(\varphi \)) of vectors initially aligned with the local element axes [23]. Thirdly, a scalar objective function (\(\varDelta \)) was constructed using Eq. 2. At a given voxel, if \(\mathbf {f}_{(v)}\) and the direction of maximal diffusion were perfectly aligned, then \(\delta _{(v)}\), defined as the dot product of \(\mathbf {f}_{(v)}\) with the projection of the diffusion tensor (\(\mathbf {D}_{(v)}\)) in the direction of \(\mathbf {f}_{(v)}\), would be maximal. Conversely, \(\delta _{(v)}\) would be minimal if \(\mathbf {f}_{(v)}\) was aligned with the direction of minimal diffusion. Finally, this objective function was maximised using nonlinear optimisation (least-squares quasi-Newton) by modifying the nodal parameters (\(\theta _{(n)}\) and \(\varphi _{(n)}\)). This procedure was implemented using the open-source Cmgui software packageFootnote 2 [24, 25].
3 Results
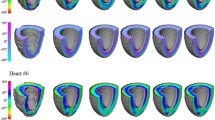
Based on the optimal fibre field fitted to the diffusion tensor data, we extracted the fibre orientations at all voxels from which the original tensors were calculated. Figure 3(a) shows the estimated fibre orientations colour-coded using the interpolated fibre angles. The resultant fibre field exhibits the expected smooth spatial variations throughout the LV, from negative angles (with respect to the short-axis plane) at the epicardium, to positive angles at the endocardium.
To quantitatively assess the ability of the new workflow to reconstruct local myofibre orientations, we performed eigenanalysis of the diffusion tensor for every voxel and examined the alignment between the estimated fibre orientation (\(\mathbf {f}_{(v)}\)) from our new workflow and the corresponding primary eigenvectors (\(\mathbf {e}_{1(v)}\)) on a voxel basis. When an estimated fibre vector is in perfect alignment with the primary eigenvector, their dot product is one. Conversely, if the two vectors are orthogonal, then the dot product is zero. We took the absolute value of the dot product as we were only interested in how well the vectors were aligned to each other regardless of their direction. To assess the overall degree of alignment throughout the model, we evaluated a normalised dot product (nDP; see Eq. 3), defined as the ratio of the sum of the dot products scaled by their associated FA values (\(\text {FA}_{(v)}\)) over all voxels (v), to the sum of FA values over all voxels. nDP is in the range from 0 to 1, with 1 representing best agreement of orientations. FA values were incorporated into the calculation of the nDP index to account for the differing degrees of confidence in the calculated eigenvectors (since an eigenvector in a voxel with low FA may not represent the underlying fibre orientation).
(a) Fitted fibre orientations colour-coded by the interpolated fibre angles at all voxels within the LV myocardium. (b) A mid-ventricular slice of dot product (DP) values between interpolated fibre orientations and primary eigenvectors of the associated diffusion tensors (see text for details). (c) Map of the FA values for the same mid-ventricular slice. The dotted squares indicate areas of low correlation between the estimated fibre orientations and primary eigenvectors.
The resulting nDP index was close to one, suggesting a high correlation between the estimated fibre orientations and primary eigenvectors of the diffusion tensors. Figure 3(b) shows a map of dot products for all voxels in a mid-ventricular slice. The alignment was generally good, except for some small localised regions in the interior, and near the boundaries of the LV. Boundary errors may be caused by the diffusion signals near the endocardial and epicardial surfaces being contaminated by partial-volume imaging artifacts, which may lead to unreliable diffusion tensor calculations near the ventricular surfaces. Internal regions with large disagreement between field and eigenvectors were generally associated with low FA, as highlighted in Fig. 3(c).
4 Discussion
We have developed a new method to construct a continuous fibre field within an FE model of the LV by directly fitting to cardiac diffusion tensor data. This approach was motivated by issues encountered with the eigenanalysis of diffusion tensors that can potentially lead to misrepresentation of the local myofibre orientation. Such misinterpretation can have a significant impact on electrophysiological and mechanical modelling studies since the orientations are used to construct local microstructural material axes from which electrophysiological, passive and contractile constitutive properties are derived.
We demonstrated that the fibre orientations estimated using this method agree well with the standard eigenvectors in regions with high FA, within which the primary eigenvector has been shown to reliably represent the local myofibre orientation. However, in regions with low FA, this method will still define continuously varying fibre orientations. If the main contributor to low FA is noise, then maintaining continuity in fibre field despite low FA is an important advantage of our method. On the other hand, if the image quality is high in regions with low FA, then the structural model should be carefully considered, and it may be appropriate to take the FA data into account when constructing biomechanical or electrophysiological constitutive models in these regions.
In previous studies that have explored direct interpolation of diffusion tensors [3, 4, 19], the tensors were first transformed into logarithmic space, and the six independent components of the logarithmic form were fitted as 3D scalar fields using FE interpolation. Fitted values were then transformed back to the Euclidean space to construct interpolated tensors throughout the models. To construct local fibre vectors, the primary eigenvectors were calculated from these interpolated tensors. Given that the tensors were interpolated using continuous functions, then this approach may improve the continuity of eigenvector directions, although this is not guaranteed. Furthermore, the above issues regarding variations in fractional anisotropy remain.
5 Conclusions
A field-based parameterisation scheme was developed to analyse ex vivo diffusion tensor MRI data. This scheme does not require the conventional calculation of primary eigenvectors of the diffusion tensors. Instead, myocardial fibre fields are fitted directly to spatial distributions of diffusion tensors. Results showed that the field-fitted fibre orientations correlated well with the primary eigenvectors in regions with high FA values. This workflow could be adapted to construct fibre fields using in vivo cardiac imaging data and associated geometric FE models for individualised analyses of heart mechanics.
Notes
- 1.
The MathWorks, Inc., Natick, Massachusetts, United States.
- 2.
OpenCMISS-Cmgui application, www.opencmiss.org.
References
Sermesant, M., Chabiniok, R., Chinchapatnam, P., Mansi, T., Billet, F., Moireau, P., Peyrat, J.M., Wong, K., Relan, J., Rhode, K., et al.: Patient-specific electromechanical models of the heart for the prediction of pacing acute effects in CRT: a preliminary clinical validation. Med. Image Anal. 16(1), 201–215 (2012)
Sermesant, M., Delingette, H., Ayache, N.: An electromechanical model of the heart for image analysis and simulation. IEEE Trans. Med. Imaging 25(5), 612–625 (2006)
Vadakkumpadan, F., Gurev, V., Constantino, J., Arevalo, H., Trayanova, N.: Modeling of whole-heart electrophysiology and mechanics: toward patient-specific simulations. In: Kerckhoffs, R.C. (ed.) Patient-Specific Modeling of the Cardiovascular System, pp. 145–165. Springer, New York (2010)
Krishnamurthy, A., Villongco, C.T., Chuang, J., Frank, L.R., Nigam, V., Belezzuoli, E., Stark, P., Krummen, D.E., Narayan, S., Omens, J.H., et al.: Patient-specific models of cardiac biomechanics. J. Comput. Phys. 244, 4–21 (2013)
Walker, J.C., Ratcliffe, M.B., Zhang, P., Wallace, A.W., Hsu, E.W., Saloner, D.A., Guccione, J.M.: Magnetic resonance imaging-based finite element stress analysis after linear repair of left ventricular aneurysm. J. Thorac. Cardiovasc. Surg. 135(5), 1094–1102 (2008)
Wang, V.Y., Lam, H., Ennis, D.B., Cowan, B.R., Young, A.A., Nash, M.P.: Modelling passive diastolic mechanics with quantitative MRI of cardiac structure and function. Med. Image Anal. 13(5), 773–784 (2009)
Xi, J., Lamata, P., Niederer, S., Land, S., Shi, W., Zhuang, X., Ourselin, S., Duckett, S.G., Shetty, A.K., Rinaldi, C.A., et al.: The estimation of patient-specific cardiac diastolic functions from clinical measurements. Med. Image Anal. 17(2), 133–146 (2013)
Niederer, S.A., Smith, N.P.: The role of the frank-starling law in the transduction of cellular work to whole organ pump function: a computational modeling analysis. PLoS Comput. Biol. 5(4), e1000371 (2009)
Wang, V.Y., Ennis, D.B., Cowan, B.R., Young, A.A., Nash, M.P.: Myocardial contractility and regional work throughout the cardiac cycle using FEM and MRI. In: Camara, O., Konukoglu, E., Pop, M., Rhode, K., Sermesant, M., Young, A. (eds.) STACOM 2011. LNCS, vol. 7085, pp. 149–159. Springer, Heidelberg (2012)
Basser, P.J., Mattiello, J., LeBihan, D.: Estimation of the effective self-diffusion tensor from the NMR spin echo. J. Magn. Reson. Ser. B 103(3), 247–254 (1994)
Hsu, E., Muzikant, A., Matulevicius, S., Penland, R., Henriquez, C.: Magnetic resonance myocardial fiber-orientation mapping with direct histological correlation. Am. J. Physiol. Heart Circ. Physiol. 274(5), H1627–H1634 (1998)
Scollan, D.F., Holmes, A., Winslow, R., Forder, J.: Histological validation of myocardial microstructure obtained from diffusion tensor magnetic resonance imaging. Am. J. Physiol. Heart Circ. Physiol. 275(6), H2308–H2318 (1998)
Lekadir, K., Hoogendoorn, C., Pereanez, M., Albà, X., Pashaei, A., Frangi, A.F.: Statistical personalization of ventricular fiber orientation using shape predictors. IEEE Trans. Med. Imaging 33(4), 882–890 (2014)
Toussaint, N., Stoeck, C.T., Schaeffter, T., Kozerke, S., Sermesant, M., Batchelor, P.G.: In vivo human cardiac fibre architecture estimation using shape-based diffusion tensor processing. Med. Image Anal. 17(8), 1243–1255 (2013)
Jones, D.K., Pierpaoli, C.: Confidence mapping in diffusion tensor magnetic resonance imaging tractography using a bootstrap approach. Magn. Reson. Med. 53(5), 1143–1149 (2005)
Jbabdi, S., Bellec, P., Toro, R., Daunizeau, J., Pélégrini-Issac, M., Benali, H.: Accurate anisotropic fast marching for diffusion-based geodesic tractography. J. Biomed. Imaging 2008, 2 (2008)
Bayer, J., Blake, R., Plank, G., Trayanova, N.: A novel rule-based algorithm for assigning myocardial fiber orientation to computational heart models. Ann. Biomed. Eng. 40(10), 2243–2254 (2012)
Streeter, D.D., Spotnitz, H.M., Patel, D.P., Ross, J., Sonnenblick, E.H.: Fiber orientation in the canine left ventricle during diastole and systole. Circ. Res. 24(3), 339–347 (1969)
Vadakkumpadan, F., Arevalo, H., Prassl, A.J., Chen, J., Kickinger, F., Kohl, P., Plank, G., Trayanova, N.: Image-based models of cardiac structure in health and disease. Wiley Interdisc. Rev. Syst. Biol. Med. 2(4), 489–506 (2010)
Basser, P.J., Pierpaoli, C.: Microstructural and physiological features of tissues elucidated by quantitative-diffusion-tensor MRI. J. Magn. Reson. 111, 209–219 (1996)
Farrell, J.A., Landman, B.A., Jones, C.K., Smith, S.A., Prince, J.L., van Zijl, P., Mori, S.: Effects of signal-to-noise ratio on the accuracy and reproducibility of diffusion tensor imaging-derived fractional anisotropy, mean diffusivity, and principal eigenvector measurements at 1.5 t. J. Magn. Reson. Imaging 26(3), 756–767 (2007)
Fomovsky, G.M., Rouillard, A.D., Holmes, J.W.: Regional mechanics determine collagen fiber structure in healing myocardial infarcts. J. Mol. Cell. Cardiol. 52(5), 1083–1090 (2012)
LeGrice, I.J., Hunter, P.J., Smaill, B.: Laminar structure of the heart: a mathematical model. Am. J. Physiol. Heart Circ. Physiol. 272, H2466–H2476 (1997)
Christie, G., Bullivant, D., Blackett, S., Hunter, P.J.: Modelling and visualising the heart. Comput. Vis. Sci. 4(4), 227–235 (2002)
Bradley, C., Bowery, A., Britten, R., Budelmann, V., Camara, O., Christie, R., Cookson, A., Frangi, A., Gamage, T., Heidlauf, T., Krittian, S., Ladd, D., Little, C., Mithraratne, K., Nash, M., Nickerson, D., Nielsen, P., Nordbø, T., Omholt, S., Pashaei, A., Paterson, D., Rajagopal, V., Reeve, A., Röhrle, O., Safaei, S., Sebastián, R., Steghfer, M., Wu, T., Yu, T., Zhang, H., Hunter, P.: OpenCMISS: a multi-physics & multi-scale computational infrastructure for the VPH/Physiome project. Prog. Biophys. Mol. Biol. 107(1), 32–47 (2011)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this paper
Cite this paper
Freytag, B. et al. (2015). Field-Based Parameterisation of Cardiac Muscle Structure from Diffusion Tensors. In: van Assen, H., Bovendeerd, P., Delhaas, T. (eds) Functional Imaging and Modeling of the Heart. FIMH 2015. Lecture Notes in Computer Science(), vol 9126. Springer, Cham. https://doi.org/10.1007/978-3-319-20309-6_17
Download citation
DOI: https://doi.org/10.1007/978-3-319-20309-6_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-20308-9
Online ISBN: 978-3-319-20309-6
eBook Packages: Computer ScienceComputer Science (R0)