Abstract
Sulfite reductase (SiR) catalyzes the reduction of sulfite to sulfide by using six electrons transported from ferredoxin (Fd) for eventual sulfur assimilation. As efficient electron flows are ensured by forming a productive Fd:SiR complex, detailed characterization of a Fd:SiR complex in solution is of particular importance. Here, we show that acidic residues of Fd play essential roles in forming an electron transfer complex with SiR by using attractive electrostatic interactions with putative basic residues of SiR. The thermodynamic approach using calorimetry revealed a favorable electrostatic contribution to form the Fd:SiR complex at the molecular level. Solution-state nuclear magnetic resonance (NMR) spectroscopy on 15N-labeled Fd in the presence of SiR showed large perturbations in NMR signals of acidic residues of Fd. The addition of NaCl diminished overall perturbations of NMR signals of SiR-bound Fd which resulted from the decrease in interprotein affinity. However, acidic residues at both termini still showed relatively large peak perturbation. These results at the residue level suggested that intermolecular interactions between Fd and SiR are electrostatic in nature and the electrostatic interaction is a dominant contributor to form the Fd:SiR complex. We suggest that a combination of calorimetry and NMR is a powerful approach to investigating protein-protein interactions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Nuclear Magnetic Resonance
- Nuclear Magnetic Resonance Spectroscopy
- Isothermal Titration Calorimetry
- Nuclear Magnetic Resonance Signal
- Sulfite Reductase
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
A variety of life phenomena such as electron transfer, signal transduction, and protein homeostasis, are conducted by intermolecular interactions of multiple proteins. Protein surfaces generally consist of both charged/polar hydrophilic and apolar hydrophobic regions. The intermolecular interactions, achieved by the electrostatic and hydrophobic forces, occur in an energetically favorable way. Several physicochemical studies have been reported on the molecular interaction between electron transfer proteins and their partner enzymes. Cytochrome c uses the highly positive electrostatic patches to interact with acidic residues of the subunit II of cytochrome c oxidase for electron transfer (Maneg et al. 2004). Similarly, positive residues of cytochrome f interact with negative residues of plastocyanin (Gross and Pearson 2003). In chloroplasts, Fd , which possesses clusters of acidic regions, acts as a multiple electron donor for various redox proteins. Biochemical and biophysical studies have suggested that various Fd-dependent enzymes such as nitrite reductase (NiR), sulfite reductase (SiR ), and ferredoxin-NADP+ reductase (FNR ) mainly interact with Fd using their positive residues on surfaces (Akashi et al. 1999; Nakayama et al. 2000; Kurisu et al. 2001; Saitoh et al. 2006; Lee et al. 2011; Sakakibara et al. 2012).
SiR reduces sulfite to sulfide using six electrons transferred from Fd . Linked sequential metabolic reactions with other enzymes produce amino acid residues such as methionine and cysteine using sulfide. Electrons are transferred from a [2Fe-2S] cluster of Fd to a [4Fe-4S] cluster of SiR and then from a [4Fe-4S] to siroheme intramolecularly. We have reported biochemical analyses of interprotein interactions between Fd and SiR to form the electron transfer complex (Akashi et al. 1999; Nakayama et al. 2000; Saitoh et al. 2006). In this study, we first applied isothermal titration calorimetry (ITC) to investigate interactions between Fd and SiR (Fig. 1). By measuring gradual heat changes attributed to the binding of SiR and Fd under a titration of increasing concentrations of Fd in the presence or absence of 100 mM NaCl, thermodynamic parameters, changes in enthalpy (ΔH), entropy (ΔS) and binding affinity (K d) with binding stoichiometry (n), were determined using the equations as below.
Isothermal calorimetric titration of Fd with SiR . Binding reactions between Fd and SiR were initiated by adding Fd (1.49 mM) in the syringe to SiR (40 μM) in the reaction cell using VP-ITC instrument (GE-Healthcare, USA) at 30 °C in 50 mM Tris/HCl buffer (pH 7.5) in the absence (a) and presence (b) of 100 mM NaCl. Titration consisted of 38 injections spaced at intervals of 300 s. The injection volume was 7 μL for each, and the cell was continuously stirred at 307 rpm. The corresponding heat of dilution of Fd titrated to the buffer was used to correct data. Thermograms and binding isotherms are shown in upper and lower panels, respectively. For clarity, the thermogram with 100 mM was magnified in inset of B. Solid lines in lower panels are theoretical curves
Observed ΔH for binding and the dissociation constant (K d) were directly calculated from the integrated heats using the two-sets (Fig. 1a, lower panel) or one-set (Fig. 1b, lower panel) of an independent binding site mode using Eq. 1:
where Q is the change in heat, V 0 is the effective volume of the calorimeter cell (~1.43 ml), L R is the ratio of the total SiR concentration to total Fd concentration ([P]t) at any given point during titrations, and n is the binding stoichiometry of Fd per a binding site on SiR. By using the values of ΔH and K d, the change in Gibbs free energy (ΔG) and ΔS were calculated using thermodynamic relationships (Eqs. 2 and 3) as follows:
where R is the gas constant and T is the temperature in Kelvin. Schematic representation of binding states of Fd and SiR are shown in the upper panels.
A large exothermic reaction was observed during a successive titration of Fd to SiR without NaCl and thus the formation of the Fd:SiR complex was assumed to be driven with negative ΔH (ΔH < 0; Fig. 1a). As the molar ratio of Fd/SiR increased, the extent of binding heat decreased and saturation was reached, indicating that all SiR was occupied with Fd. Interestingly, there were two binding phases: a high binding affinity phase at the low molar ratio of Fd/SiR (0 to ~2) was followed by a low binding affinity phase at the high molar ratio (~2 to ~7). The analyses of these isotherms provided that the values of ΔH, K d, and n for a high affinity binding reaction were −4.4 kcal/mol, 0.2 μM, and ~1, respectively, and, for a low affinity binding reaction, ΔH of several negative kcal/mol, K d of several hundred μM, and n of several values (~3 to ~5) (Table 1): the value of “n” indicates the number of Fd which binds to SiR. Negative ΔH clearly revealed, regardless of the binding affinity and site, an exothermic nature of the Fd:SiR complex formation, which suggested the favorable contribution of interprotein electrostatic interactions. Furthermore, it was shown that when concentrations of Fd were lower than those of SiR, one Fd bound to one SiR (n = ~1) with a strong affinity (K d = 0.2 μM; Table 1) at a high affinity binding site (Fig. 2). However, as excess amounts of Fd existed, several Fds further bound to SiR (n = ~3 to ~5) with a weak affinity (K d of several hundred μM) at a low affinity binding site. Considering the difference in the molecular size of small Fd (~11 kDa) and large SiR (~65 kDa), multiple bindings of Fd to SiR are possible.
Schematic representation of the effect of the NaCl addition on interprotein interactions between Fd and SiR . Green and red spheres represent SiR and Fd, respectively. Red ellipses and orange spheres indicate the siroheme and the [4Fe-4S] cluster in SiR, respectively. Yellow spheres signify the [2Fe-2S] cluster in Fd. In the absence of NaCl, a high affinity binding site of SiR accommodates one Fd as evidenced by the n value (n = ~1; Table 1) with high affinity and specificity. Several Fds (n = ~3 to ~5) interact with SiR at a low affinity binding site in a lesser specific way. In the presence of 100 mM NaCl, SiR holds one Fd just at a high affinity binding site with relatively high affinity and specificity
In order to further reveal characteristics of the interaction between Fd and SiR , ITC measurements were performed in the presence of 100 mM NaCl (Fig. 1b). If electrostatic interactions are dominant, binding affinity should decrease upon addition of NaCl. In contrast, in the case of hydrophobic interactions playing a major role, reinforced hydrophobic interactions by increasing NaCl concentration stabilize the complex. Although exothermic heat was generated from the binding of the two proteins, the extent of heat was found to be much smaller than that without NaCl. Decreases in the degree of exothermic heat were interpreted as dropping of attractive electrostatic interactions between negative charges of Fd and positive charges of SiR due to the screening effects of NaCl and/or direct binding of the counter ions (Na+ and Cl−) to charged/polar residues. Accordingly, ΔH decreased in magnitude from −4.4 to −0.9 kcal/mol. K d and ΔG also increased to 23 μM and −6.5 kcal/mol, respectively, indicative of decreases in affinity between proteins. Intriguingly, adding NaCl apparently abolished weak interprotein interactions even at excess amounts of Fd (Fig. 2), which suggested that the origin of such weak interactions was also electrostatic in nature.
The entropy term (TΔS) exhibited positive values in the absence (4.9 kcal/mol) and presence (5.6 kcal/mol) of NaCl (Table 1). Large energetic costs (3.5 kcal/mol) coming from the increase in ΔH by adding NaCl were still compensated for by energetic gains from the positive entropy change. These indicated that entropy changes were also an important driving force to form the Fd :SiR complex which may be a result of dehydration of water upon the complex formation: decreasing ΔG by increasing TΔS favors the binding reaction (Eq. 3).
Consequently, ITC results showed that intermolecular electrostatic interactions are a main driving force to form the Fd :SiR complex by using enthalpy. However, it is worth noting that a favorable hydrophobic contribution to complexation is also plausible since hydrophobic interactions gain entropy (i.e., ΔS > 0) from dehydration on interfacial packing among hydrophobic residues.
Solution-state NMR spectroscopy is useful to investigate intermolecular interactions at the atomic/residue level. NMR further probes weak and/or dynamic features of intermolecular interactions (Mizushima et al. 2014). We first carried out the measurement of 1H-15N heteronuclear single-quantum coherence correlation (HSQC), using 15N-labeled Fd without SiR and NaCl (Fig. 3a). Clear and sharp NMR signals of free Fd were obtained with good dispersion ranging from 6 to 11 ppm in the transverse axis for proton (1H). One peak in the HSQC spectrum corresponds to one amino acid residue of the backbone in Fd. Based on the previous assignment information (Saitoh et al. 2006), we assigned the NMR peaks to the corresponding residues. The addition of SiR changed the position of several NMR peak signals of Fd (i.e., chemical shift), indicating that a set of Fd residues was involved in SiR binding. The number of assigned peaks (78) was the same as that without SiR, suggestive of a fast exchange regime in terms of the NMR time scale between free and complex states, thereby showing an averaged one peak between these two conformational states. A binding system, which shows a fast exchange regime has suggested (relatively) weak intermolecular interactions over a nanomolar order of K d. Thus, a system with a fast exchange often shows K d in a micromolar order. A slow exchange regime produces two separated peaks which correspond individually to free and complex states because intermolecular interactions are significantly strong. Typically, K d in a nanomolar order shows slow exchange regimes. Therefore, the binding affinity of Fd for SiR is judged to be relatively strong based on a fast exchange, consistent with ITC results which showed the submicro molar K d value.
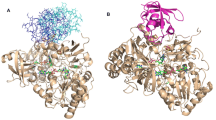
NMR spectra of SiR -(un)bound Fd without NaCl and binding interfaces of Fd for SiR. (a) 1H-15N HSQC spectra of 15N uniformly-labeled Fd without (red) and with (blue) SiR were obtained using an AVANCE II-800 spectrometer equipped with a cryogenic probe (Bruker, Germany) in 50 mM Tris/HCl buffer (pH 7.4) containing 10 % D2O at 25 °C. Data were processed by NMRPipe and analyzed by Sparky. Stable isotope-labeled maize leaf Fd with 15 N was expressed using E. coli cultured in minimal media containing [15N]-NH4Cl as a nitrogen source and purified as previously described (Kurisu et al. 2001; Saitoh et al. 2006; Sakakibara et al. 2012). Maize leaf SiR was prepared as described in our previous study (Saitoh et al. 2006). The protein concentrations for NMR measurements were 100 μM for Fd and 50 μM for SiR. The assigned peaks are shown with the one letter amino acid code and residue number. The NMR signal stemmed from the side chain of tryptophan is shown (W73). (b) The values of chemical shift differences (CSD) were plotted against residue numbers. Mainly acidic clusters are shown by the red rectangles. The regions where NMR peaks are invisible due to the paramagnetic relaxation effect (PRE) from iron are displayed with the green rectangle. (c) Mapping of residues which showed CSD on the crystal structure of Fd (PDB ID: 1GAQ) (Kurisu et al. 2001). The degrees of CSD are shown by the color code: red > 0.06 ppm, 0.04 < orange < 0.06 ppm, and 0.02 < yellow < 0.04 ppm. NMR invisible regions due to PRE are represented with green colors. N- and C-terminal parts are indicated
The degree of shifts in the peak position (i.e., chemical shift difference, CSD) in NMR peaks with and without SiR (Fig. 3a) were calculated by using Eq. 4:
where Δδ HN and Δδ N are changes in 1H and 15 N chemical shifts in ppm, respectively. The weighting factor of 0.158 was used to adjust the relative magnitudes of the amide nitrogen chemical shift range and the amide proton chemical shift range.
By using CSD information, we could identify the important residues of Fd responsible for SiR binding (Fig. 3b). The largely perturbed residues which showed CSD over 0.03 were distributed in the N-terminal region (I24, A28, E29, E30, D34, L35, and Y37), the central region (G49, Q61, and L64), and the C-terminal region (E92, E93, and E94), respectively (Fig. 3b, c). Together with a few hydrophobic residues (I24, L35, and L64), it was obvious that many negatively charged residues were responsible for forming a complex with SiR.
To clarify the effects of salts on Fd :SiR interactions, 1H-15N HSQC spectrum of SiR-unbound Fd at 100 mM NaCl was obtained (Fig. 4a). The spectrum also exhibited sharp peaks with broad dispersion as observed without NaCl and SiR. 1H-15N HSQC spectrum was then obtained with SiR: adding SiR shifted many peak positions with a fast exchange regime but to a lesser extent than those without NaCl (Fig. 4b). The overall direction of peak shifts in the presence of SiR was similar at 0 and 100 mM NaCl, which suggested the similarity of an overall binding mode between Fd and SiR at two distinct salt concentrations. The CSD analysis displayed an overall decrease in CSD of SiR-bound Fd (Fig. 4b) compared to that without NaCl (Fig. 3b). This indicated the decrease in the population of a Fd:SiR complex due mainly to the dropping of interprotein affinity by the disruption of attractive electrostatic interactions by NaCl, in good accordance with our ITC results which showed the increase in K d values from 0.2 to 22 μM (Table 1).
NMR spectra of SiR -(un)bound Fd with NaCl and binding interfaces of Fd for SiR. (a) 1H-15N HSQC spectra of 15N uniformly-labeled Fd with (cyan) and without (magenta) SiR were obtained using the identical condition and procedure which are described in Fig. 3 except the presence of 100 mM NaCl. (b) CSD in NMR peaks with and without SiR was calculated and plotted against residue numbers. Mainly acidic clusters are shown by the red rectangles. The regions where NMR peaks are invisible due to PRE are displayed with green rectangles. (c) Mapping of residues which showed CSD on the crystal structure of Fd (PDB ID: 1GAQ) (Saitoh et al. 2006). The degree of CSD is shown by the color code: red > 0.04 ppm, 0.03 < orange < 0.04 ppm and 0.02 < yellow < 0.03 ppm. NMR invisible regions due to PRE are represented with green colors and both termini are indicated
The analysis further revealed that acidic residues at the N- and C terminal parts and the central part (I24, A28, E29, E30, D34, L35, Y37, G49, Q61, L64, E92, E93, and E94) showed larger CSD values than those in the other parts (Fig. 4b, c). These acidic residues were almost the same as those without NaCl although the degree of change in CSD for each residue was different. Furthermore, considering the fact that acidic residues of Fd in both termini were also used for interactions with FNR (Kurisu et al. 2001; Saitoh et al. 2006) and NiR (Sakakibara et al. 2012), a favorable electrostatic force may be a common feature in interprotein interactions between Fd and its redox partners. Meanwhile, it should be noted that hydrophobic residues, L35 and L64 also exhibited large CSD values, which implied the contribution of hydrophobic interactions to the complex formation.
Taken all together, it was concluded that attractive electrostatic interactions act as a main stabilizer for forming a Fd :SiR complex and NaCl attenuates interprotein interactions between electrically-charged residues. On the one hand, although favorable electrostatic interactions for complexation are evident, hydrophobic contributions are still unclear. Favorable hydrophobic interactions enhanced by adding of 100 mM NaCl might promote the complex formation to a much lesser extent than opposite contribution of electrostatic interactions decreased by 100 mM NaCl. Thus, it appears that only electrostatic contributions may appear as the dominant contributor to complexation. Further detailed study will require the complete understanding of interprotein interactions between Fd and SiR by considering hydrophobic interactions such as the residues detected here (I24, L35, and L64) and the hydrophobic region around the [2Fe-2S] clusters that is NMR -invisible due to PRE. Site-directed mutagenesis combined with SiR activity assays and direct structural studies based on X-ray crystal and solution-state NMR approaches should prove promising in obtaining a clue or solution for this issue. Finally, current results imply that subtle changes in salt concentration, which might reflect conditions in the chloroplast, control electrostatic interaction for the electron transfer complex.
References
Akashi T, Matsumura T, Ideguchi T, Iwakiri KI, Kawakatsu T, Taniguchi I, Hase T (1999) Comparison of the electrostatic binding sites on the surface of ferredoxin for two ferredoxin-dependent enzymes, ferredoxin-NADP+ reductase and sulfite reductase. J Biol Chem 274:29399–29405
Gross EL, Pearson DC (2003) Brownian dynamics simulations of the interaction of chlamydomonas cytochrome f with plastocyanin and cytochrome c6. Biophys J 85:2055–2068
Kurisu G, Kusunoki M, Katoh E, Yamazaki T, Teshima K, Onda Y, Kimata-Ariga Y, Hase T (2001) Structure of the electron transfer complex between ferredoxin and ferredoxin-NADP+ reductase. Nat Struct Biol 8:117–121
Lee YH, Ikegami T, Standley DM, Sakurai K, Hase T, Goto Y (2011) Binding energetics of ferredoxin-NADP+ reductase with ferredoxin and its relation to function. ChemBioChem 12:2062–2070
Maneg O, Malatesta F, Ludwig B, Drosou V (2004) Interaction of cytochrome c with cytochrome oxidase: two different docking scenarios. Biochim Biophys Acta 1655:274–281
Mizushima R, Kim JY, Suetake I, Tanaka H, Takai T, Kamiya N, Takano Y, Mishima Y, Tajima S, Goto Y, Fukui K, Lee YH (2014) NMR characterization of the interaction of the endonuclease domain of MutL with divalent metal ions and ATP. PLoS One 9:e98554
Nakayama M, Akashi T, Hase T (2000) Plant sulfite reductase: molecular structure, catalytic function and interaction with ferredoxin. J Inorg Biochem 82:27–32
Saitoh T, Ikegami T, Nakayama M, Teshima K, Akutsu H, Hase T (2006) NMR study of the electron transfer complex of plant ferredoxin and sulfite reductase. J Biol Chem 281:10482–10488
Sakakibara Y, Kimura H, Iwamura A, Saitoh T, Ikegami T, Kurisu G, Hase T (2012) A new structural insight into differential interaction of cyanobacterial and plant ferredoxins with nitrite reductase as revealed by NMR and X-ray crystallographic studies. J Biochem 151:483–492
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Kim, J.Y., Ikegami, T., Goto, Y., Hase, T., Lee, YH. (2015). Investigation of Protein-Protein Interactions of Ferredoxin and Sulfite Reductase Under Different Sodium Chloride Concentrations by NMR Spectroscopy and Isothermal Titration Calorimetry. In: De Kok, L., Hawkesford, M., Rennenberg, H., Saito, K., Schnug, E. (eds) Molecular Physiology and Ecophysiology of Sulfur. Proceedings of the International Plant Sulfur Workshop. Springer, Cham. https://doi.org/10.1007/978-3-319-20137-5_17
Download citation
DOI: https://doi.org/10.1007/978-3-319-20137-5_17
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-20136-8
Online ISBN: 978-3-319-20137-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)