Abstract
Over the past decade, molecular cancer research studies have revealed microRNAs, a class of small, endogenous noncoding RNAs that negatively control the expression of target genes by cleaving mRNA or through translation repression. As critical gene regulators, microRNAs modulate a variety of cellular events that among others include growth, differentiation and apoptosis. The microRNAs cause cancerous diversity and complexity. Based on microRNA’s function, some microRNAs can be defined either tumor suppressor or oncogenic (oncomiR), and be focused as biomarkers or as therapy targets for oncology and clinical trial. In this review article, we have summarized recent available data on microRNAs playing an important role in the initiation, progression and drug resistance of colorectal cancer. More specifically, two of the most notorious miRNAs, tumor suppressor miR-145 and oncomiR miR-21, are discussed, since these miRNAs play a critical role in devolvement of resistance to chemotherapeutics in colorectal cancer by regulating cancer stem cell (CSC) growth and differentiation. Further, we have briefly described current miRNA-based therapeutic strategies. In conclusion, miRNAs and mRNA regulated network play key roles in cancer cell’s acquired ability of resistance to chemotherapeutics by regulating cancer stem cell growth and differentiation. The miRNA-based therapy targeting cancer stem cells shows the potential to overcome drug resistance.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
The state of our knowledge on the role of microRNAs (miRNAs) in therapeutic resistance of colon cancer is discussed in the context of cancer stem cells (CSCs) in this chapter.
2 MicroRNAs (miRNAs) Definition and Function
The miRNAs are a class of 18–24 nucleotide long endogenous noncoding RNAs that control gene expression through binding to the seed sequence at the 3′-UTR of target mRNAs, resulting in translational repression or mRNA degradation [1]. A given species of miR can perfectly or imperfectly base pair with multiple targets, allowing it to potentially regulate the translation of several mRNAs. It has been predicted that over 30 % of the human protein coding genes are post-transcriptionally regulated by this mechanism [2]. The miRNAs have emerged as critical gene regulators, which modulate a variety of cellular events that among others include growth, differentiation and apoptosis. Several hundred (940, http://www.mirbase.org/) miRNAs have been identified in human cells and the list is growing [3].
Based on miRNA’s localization in the genome, miRNAs can be intergenic, intronic, or exonic and can be transcribed as a single miRNA from its own promoter (monocistronic) or several miRNAs as a cluster from a shared promoter (polycistronic). Intergenic miRNAs are found in between genes in distinct transcription units. The miRNAs can be intronic of coding or noncoding genes, where they may be transcribed from the same promoter as the host gene. The exonic miRNAs are rare and are mainly found in exons of coding or noncoding genes [4]. Sevignani et al. have demonstrated that in the mouse, miRNA genes are frequently located near cancer susceptibility loci, which are often subjected to genomic alterations leading to activation by translocations or amplifications, or loss of function due to deletions, insertions, or mutations [5].
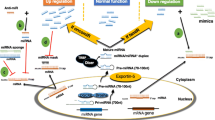
The miRNAs are transcribed by RNA polymerase II as long primary transcripts of variable sizes (pri-miRNA), which are processed into ~70 nucleotide long precursor miRNAs (pre-miR) by an RNAse-III-like enzyme, Drosha, together with DGCR8 (DiGeorge syndrome critical region gene 8), an RNA binding protein in the nucleus [6, 7]. Export of pre-miR to cytoplasm is mediated by Exportin-5 via GTP-dependent export, where it is further cleaved by another RNAse-III enzyme, called Dicer, into a mature dsRNA duplex. After strand-selection, mature miRNA is assembled into the RNA-induced silencing complex (RISC), which ultimately performs regulatory functions [8].
The miRNA-mRNA interactions are characterized by binding of the perfect or nearly perfect miRNA seed region (typically 2–8 bases) to the target mRNA. Each miRNA can control hundreds of target mRNAs just as a single mRNA can be targeted by multiple miRNAs. The targets of specific miRNA can be predicted by bioinformatics’ algorithms (Targetscan), but validation must be achieved through luciferase reporter assays, quantitative real-time PCR, and immunoblotting studies.
In addition to the canonical mechanisms of miRNA gene regulation through 3′ UTR interactions, other “noncanonical” miRNA-mediated mechanisms of modulation of mRNA expression are emerging [9]. Some miRNAs have been shown to bind to the open reading frame or to the 5′ UTR of the target genes and, in some cases, activate rather than inhibit gene expression. Garzon et al. have recently reported that miRNAs exhibit decoy activity and bind to ribonucleoproteins in a seed sequence in an RISC-independent manner and interfere with their RNA binding functions [9]. Few studies have reported that miRNAs can also regulate gene expression at the transcriptional level by binding directly to the DNA [9]. Overall, these data show the complexity and widespread regulation of gene expression by miRNAs that should be taken into consideration when developing miRNA-based therapies.
It is not surprising that miRNAs are involved in diverse biological processes, including cell differentiation, proliferation, and apoptosis, presumably through a myriad of targets. Deregulation of miRNAs contributes to human pathogenesis [3]. Aberrant expression of miRNAs, including miR-21, miR-17-92, miR-15, miR-16, and let-7, has been reported in cancer [10]. Furthermore, a substantial number of miRNA genes are located in the fragile sites in the genomic regions that are frequently amplified, deleted, or rearranged in cancer, providing plausible mechanisms of deregulated expression [11]. A miRNA can act as a tumor suppressor or an oncogene depending on its targets in different tissues and cell types [12].
Dysregulated miRNA expression is reported in various human diseases including colorectal cancer, suggesting an important role in its pathogenesis. Various studies have established that a subset of miRNAs plays a role in the initiation and progression of colorectal cancer. The purpose of this review article is to summarize the available data on miRNA profiling in light of developing therapeutic strategies specific for drug resistant colorectal cancer.
3 Cause of Colorectal Cancer: Multi-genetic Mutations
Human colorectal cancer, the third most common cancer in the US, is considered the end result of stepwise accumulation of genetic and epigenetic alterations (mutations) in oncogenes and tumor suppressor gene [13]. The process of tumor progression starting from its initiation to the development of malignant lesions as a result of acquiring a series of mutations over time, has been particularly well studied [14, 15] (Fig. 8.1).
Role of various mutations and miRNAs in colorectal cancer progression and pathogenesis. Colorectal cancer is the end result of stepwise accumulation of genetic and epigenetic alterations (mutations) in oncogenes and tumor suppressor gene affecting various signaling pathways. Selected miRNAs and their target genes are indicated in the scheme (This scheme is adapted from the genetic model for colorectal cancer highlighted by Fearon and Vogelstein [79] and Lao and Grady [80]). Abbreviations: APC adenomatous polyposis coli, CASP3 caspase 3, CTGF connective tissue growth factor, TSP1 thrombospondin1, EGFR epidermal growth factor receptor, P13K phosphoinositide-3 kinase, mTOR mammalian target of rapamycin, PTEN phosphatase and tensin homolog, DCC deleted in colorectal carcinoma, TGFβRI/II transforming growth factor β receptor I and II, SMAD2,3,4 mothers against decapentaplegic homolog 2,3,4, SIRT1 sirutin1, CDK4,6 cyclin dependent kinase 4,6, TIMP3 tissue inhibitor of metalloproteinase 3, PDCD4 programmed cell death 4, RECK reversion-inducing cysteine-rich protein with kazal motifs, uPAR plasminogen activator urokinase receptor, MMPs matrix metalloproteases, ECM extra cellular matrix, ZEB1/2 zinc finger E-box binding homeobox 1, EMT epithelial to mesenchymal transition, PRL3 Phosphatase of regenerating liver 3
The first, gatekeeping mutations occur in APC gene that makes normal epithelial cells outgrow and develop into a small adenoma. The APC protein acts as a “brake” on the accumulation of β-catenin protein. Without APC, β-catenin accumulates to high levels and translocates into the nucleus, binds to DNA, and activates the transcription of genes that are important for stem cell renewal and differentiation. While APC is mutated in most colon cancers, some studies have shown that mutations in β-catenin block its degradation and result in its increased levels [16]. Mutation(s) in other genes with functions analogous to APC such as AXIN1, AXIN2, TCF7L2, or NKD1 have also been reported to regulate β-catenin levels [17].
The second important mutation identified in colorectal cancer is in K-Ras gene, which unleashes an expansion of cell number[14]. K-Ras acts as a molecular on/off switch. Once it is turned on, it recruits and activates proteins necessary for the propagation of growth factors and other receptor signals such as c-Raf and PI3-kinase. Activation of EGFR signaling pathway is also involved at this step. The cells with mutation only in APC gene may persist, but their cell numbers are small compared to those that have mutations in both the genes. This process of mutation followed by clonal expansion continues, and mutations in genes such as PIK3CA, SMAD4, and TP53. Mutation in TP53 gene transforms the tissue from an adenoma into an invasive carcinoma and metastasis to lymph nodes and distant organs [18]. The tumor suppressor p53, produced by the TP53 gene, normally monitors cell division and selectively kills cells that have Wnt or EGFR pathway defects.
The mutations that confer a selective growth advantage to the tumor cell are called “driver” mutations. It has been estimated that each driver mutation provides only a small selective growth advantage to the cell, to the order of a 0.4 % increase between cell birth and cell death [19]. Over many years, however, this slight increase, compounded once or twice per week, can result in a large mass, containing billions of cells [18]. The mutations can be inherited or are acquired and most probably occur in the intestinal crypt stem cell. However, only about 4 % colorectal cancer cases show a family history, most of the mutations are acquired during the life span of an individual.
4 Colon Cancer Stem Cells (CSC): Adult Stem Cells with Accumulated Mutations
As mentioned above, colorectal cancer is generally a result of accumulated genetic and epigenetic mutations. Only long-lived cells may serve as reservoirs for such precancerous mutations. Colonic mucosa is a highly dynamic tissue. Mucosal surface epithelium cells are constantly replaced with cells derived from stem cells that are located at the base of the crypt. Although the origin of t colon cancer stem or stem-like cells (CSCs) is not fully known, they are thought to originate from stem cells that have acquired mutations. Considering that the appearance of CSCs might be one of the initial events in neoplastic transformation in solid tumors as well as in intestinal neoplasia, we investigated the status of CSCs in normal appearing colonic mucosa during aging in patients with adenomatous polyps. Colon CSCs, as evidenced by the expression of CSC markers (CD44, CD166 and Ep-CAM) were observed not only in premalignant adenomatous polyps, but also in normal appearing colonic mucosa, where expression increased with advancing age indicating increased risk of developing colorectal cancer during aging [20]. Additionally, we found the age-related increase in adenomatous polyps in the colon was associated with increased expression of colon CSC markers [38].
Over the last decade, the cancer stem cell model has become increasingly accepted as an explanation for cancer development, spread and recurrence. This model posits that a small subpopulation of tumor cells, termed cancer stem cells (CSCs), which are distinct from the bulk of the cells in the tumor, can self-renew, differentiate into multiple lineages, and drive tumor growth, metastasis and recurrence. Currently, most CSCs are identified by the use of a variety of cell surface markers, including CD44, CD166 and Ep-CAM. They can be isolated using FACS technology, and then tested by propagation in immune deficient mice [21].
5 Cancer Stem Cells (CSCs) and Drug Resistance
Although surgery and subsequent chemotherapy can cure over 75 % of colon cancer patients, more than 30 % of these patients develop new neoplastic polyps, and 10 % progress to frank second malignancy, underscoring the need for a better understanding of the underlying mechanisms of recurrence [22, 23].
The standard therapy for colorectal cancer includes surgery followed by chemotherapy or other effective therapeutic regimen to eliminate any cancer cells. However, most chemotherapy drugs only targets the rapidly dividing cells that form bulk of the tumor. The effectiveness of cancer therapeutics is evaluated by the reduction in tumor mass, resulting in elimination/killing of dividing differentiated or undifferentiated cells that form the bulk of the tumor. However, the slow growing CSCs/CSLCs remains untouched and may even be enriched resulting in relapse of the disease. Recently, we have shown that exposure of colon cancer HCT-116 or HT-29 cells to the combination of 5-FluoroUracil (FU) and Oxaliplatin (OX) [FUOX] inhibited their growth but led to the enrichment of CSC phenotype [24]. We have now generated FUOX-resistant HCT116 and HT29 cells that exhibit both enrichment of CSCs/CSLCs and elevated levels miR-21. We have further demonstrated that, miR-21 plays a determinant role in inducing stemness in colon cancer cells [25, 26].
CSCs/CSLCs show resistance to a number of conventional chemotherapies [27]. These include therapies targeting drug-efflux capabilities, anti-apoptotic mechanisms, and induction of differentiation as well as to other stem cell pathways resulting in chemotherapy-refractory tumors. Changes in these properties may explain why it is difficult to completely eradicate cancer and why recurrence is an ever-present threat. Thus, therapeutic strategies that specifically target colon CSCs are likely to be effective in eradicating tumors and in reducing the risk of relapse and metastasis.
6 Micro-RNAs Regulate Cancer Stem Cell Proliferation and Differentiation
As previously stated, miRNAs bind to target mRNA resulting in either its degradation or inhibition of translation. The miRNAs that normally down-regulate the expression of an oncogene and is often lost in tumor cells can be defined as a tumor suppressor. The lost expression of this miRNA by mutation, deletion, promoter methylation or by any other factor(s) that might result in an abnormal expression of the target oncogene, which subsequently contributes to tumor formation by inducing cell proliferation, invasion, angiogenesis or/and decreased cell death.
Alternatively, miRNAs that down-regulate tumor suppressor gene expression or other important genes are involved in differentiation, and could contribute to tumor formation by stimulating proliferation, angiogenesis and invasion. These miRNAs are defined as oncogenic miRNA or “oncomiR”, and are normally up-regulated in distinct types of human neoplasia and also often associated with distinct cytogenetic abnormalities. The miRNAs that function as tumor suppressors or oncogenes in colorectal cancer are listed in Table 8.1.
Two of the most notorious miRNAs, an oncomiR miR-21 and tumor suppressor miR-145 are discussed in detail in the following section, since these miRNAs play a critical role in recurring colorectal cancer by regulating stem cell growth and differentiation.
6.1 miR-21
The miRNA-21 is encoded by the MIR21 gene [28]. The human microRNA-21 gene is located on chromosome 17q23.2 within a coding gene TMEM49 (also called vacuole membrane protein). The stem–loop precursor of miR-21(pre-miR-21) resides between nucleotides 2445 and 2516 of pri-miR-21. Despite being located in intronic regions of a coding gene in the direction of transcription, it has its own promoter regions and forms a ~3433-nt long primary transcript of miR-21 (pri-miR-21), which is independently transcribed. The pri-miR-21 promoter has been partially characterized [29]. Fujita et al. described a promoter mapping −3,770 to −3,337 upstream to the miR-21 hairpin. This has several conserved enhancer elements including binding sites for activation protein 1 (AP-1; composed of Fos and Jun family proteins), Ets/PU.1, SRF, p53 and STAT3. Talotta et al. [30] have reported that the miR-21 is induced by AP-1 in response to Ras, and the tumor suppressors PTEN and PDCD4 are down-regulated by Ras in an AP-1- and miR-21-dependent fashion. They have demonstrated that PDCD4 is a negative regulator of AP-1. The miR-21-mediated down-regulation of PDCD4 is essential for the maximal induction of AP-1 activity in response to Ras. The data reveal a mechanism of positive auto-regulation of the AP-1 complex in Ras transformation and disclose the function of oncomiRs as critical targets and regulators of AP-1 in tumorigenesis [31].
In addition to the positive regulators of miR-21 transcription, several transcriptional suppressors have been reported. For example, miR-21 transcription was found to be repressed by NFI, C/EBPα [32]. In addition, Gfi1[33] and estrogen receptor[34] were also shown to negatively regulate miR-21 promoter activity.
The miR-21 expression is regulated at multiple levels, including transcription and post-transcriptional processing. Kern and colleagues showed that EGF/Ras efficiently induced the miR-21 primary transcript, but this does not rapidly and simply translate into higher mature miR-21 levels. Rather, induction of mature miR-21 by constitutive activation of this pathway is slow, is associated with only minimal activation of mitogen-activated protein kinase (MAPK), and may involve stimulation of post-transcriptional processing by mechanisms other than Dicer stabilization. Further, they identified Ets transcription factors as modifiers of miR-21 expression in colorectal cancer [35]. The effects of Ets factors on miR-21 expression involve both direct and indirect mechanisms and are cell context-dependent. The Ets factor Pea3 emerges from these studies as a consistent repressor of miR-21 transcription [35].
A number of targets for microRNA-21 have been experimentally validated and most of them are tumor suppressors. Notable targets include PTEN, PDCD4, Tropomyosin, Sprouty 1, Sprouty 2, RECK, TGFβRII, MEF2C, ANP32A, SMARCA4, RhoB, and hMSH2 [36, 37].
The miR-21 has been found to be overexpressed in most epithelial cancers such as lung, breast, stomach, prostate, colon, brain, head and neck, esophagus and pancreatic cancers. Data from own laboratory have demonstrated that miR-21 levels are markedly elevated in FUOX-resistant colon cancer cells [26] that are highly enriched in CSLCs/CSCs and exhibit increased drug-efflux property [38]. Studies show that knockdown of miR-21 impairs growth, induces apoptosis and reduces the migration and invasion of various cancer cells including those of colon cancer [32]. Induced expression of miR-21 leads to increased β-catenin activity, augmentation of c-Myc and Cyclin-D expression, increase in the number of cancer stem cells, and is accompanied by increased colonosphere forming ability in vitro and tumor formation in SCID mice [26]. Therefore, miR-21 is believed to play a pivotal role in the progression of many malignancies and has been called an “oncomiR”.
6.2 miR-145
The miR-145 is located on chromosome 5 (5q32-33) within a 4.09 kb region, and is co-transcribed with miR-143 by RNA polymerase II into pri-miRNA, which is processed to ~88 bp long pre-miRNA involving RNA splicing and exporting, and finally to mature miR-145.
The miR-145 is a p53-regulated gene. Several reports suggest that miR-145 can be induced transcriptionally by p53 in response to stress such as serum starvation or anticancer drugs [39, 40]. The tumor suppressor, p53, also enhances the post-transcriptional maturation of miR-143 and miR-145 in response to DNA damage by interacting with the Drosha processing complex [41]. A recent study has demonstrated that activated Ras can suppress miR-143/145 cluster through Ras-responsive element-binding protein (RREB1), which represses the miR-143/145 promoter [42].
Down-regulation of miR-145 has been found in multiple tumors including colon, breast, prostate, pancreas etc. [39, 43]. In fact, miR-145 has been well documented as a tumor suppressor gene because it negatively regulates multiple oncogenes such as Myc, K-Ras, IRS-1, ERK5[39, 44]. Moreover, miR-145 negatively regulates junctional cell adhesion molecule (JAM-A), fascin and MUC1 and suppresses breast cancer cell motility and invasiveness [45, 46]. miR-145 also inhibits colon cancer cells’ proliferation and sensitizes them to 5-fluorouracil by targeting oncogenic FLI1 [47]. We have recently observed FUOX-resistant colon cancer HCT-116 cells which have been stably transfected with miR-145 loses their ability to form colonospheres in vitro or tumor in SCID mice (unpublished observations). In FUOX-resistant cells, over-expression of miR-145 causes a marked increase in CK-20, indicating induction of differentiation (unpublished observation). Whether the latter renders them more susceptible to chemotherapy or other types of therapeutics remains to be determined.
Several other targets of miR-145 that participate in stem cell growth and dedifferentiation have also been identified. Xu et al. have shown that miR-145 is induced during differentiation, and it directly silences the stem cell self-renewal and pluripotency by suppressing multiple pluripotent genes such as OCT4, SOX2 and KLF4 [48]. Moreover, clinical studies show that down-regulation of miR-145 is frequently associated with cancers and in smaller adenomas, suggesting a negative role for miR-145 in the initiation of tumor development by regulating cancer stem cell proliferation and differentiation [49]. Kamatani et al. recently reported that the decreased expression of miR-143 and -145 frequently occurred before APC gene aberrations [50]. The down-regulation of miR-143 and -145 is thus an important genetic event for the initiation step in colorectal tumor development.
7 miRNA and Chemotherapy Resistance in Colorectal Cancer
In cancer, as a result of multiple genetic and epigenetic alteration events, multiple genes, protein and their network are abnormally expressed. The current chemotherapy regimen such as 5-FU based FOLFOX (5-FU plus Oxaliplatin and Folinic acid), the backbone of colorectal cancer chemotherapeutics, interferes with and halts the growth of rapidly dividing cancer cells, but with limited success. While differentiated or differentiating cells that form the bulk of the tumor, are sensitive to chemo or other therapeutic agent(s), CSCs/CSLCs are resistant to conventional chemotherapy. Therefore, targeting CSC directly and/or inducing their differentiation rendering them susceptible to therapy could be a novel strategy to overcome drug resistance in colorectal cancer. Indeed, we have demonstrated that induction of differentiation of FUOX-resistant colon CSLCs/CSCs by Schlafen-3, a gene that induces differentiation, renders them highly susceptible to the combination of 5-FU and Oxaliplatin [38, 51].
Dysregulation of miRNA profiles has been reported in cancer cells and tissues. As mentioned above, a single miRNA regulates many different signaling pathways and orchestrates integrated responses in normal cells and tissues. It is reasonable to think that miRNAs play key roles in coordinating cancerous networks and cause their diversity and complexity. Developing therapeutic strategies to restore homeostasis by modifying miRNA expression may prove to be more comprehensive and successful than targeting individual genes or proteins, since only a few specific deregulated miRNAs modulate large target gene expression and multiple signaling pathways in cancer cells. For example, expression of miR-21 is higher in colon adenocarcinomas than normal mucosa and is associated with decreased overall survival [52]. A number of genes, including PTEN, TPM1, PDCD4, Spry1, and Spry2, have been reported to be targets of miR-21, suggesting its potential function in regulating cell proliferation, apoptosis, and invasion [36, 37]. Furthermore, over-expression of miR-21 has been shown to dramatically reduce the therapeutic efficacy of 5-FU [53]. In addition, increased Ras signaling activity by miR-21 mediated by Ras-responsive element-binding protein (RREB1) represses expression of the miR-143/145 cluster [42, 54], which induces differentiation of CSCs. Thus, targeting miRNAs may be a worthwhile therapeutic strategy for CSCs enriched recurrent cancer.
8 miRNA-Based Therapeutic Strategies
There are two main strategies to target miRNA expression in cancer: (a) block the expression of an oncogenic miRNA or (b) restore the expression of a tumour suppressor miRNA. The methods include (a) direct use of oligonucleotides or virus-based constructs to either block the expression of an oncogenic miRNA or to substitute for the loss of expression of a tumour suppressor miRNA, (b) indirect use of the drugs to modulate miRNA expression by targeting their transcription and their processing.
8.1 Blocking Oncogenic miRNAs
This can be achieved by the use of antisense oligonucleotides, miRNA sponges, and small RNA inhibitors. Antisense oligonucleotides work as competitive inhibitors of miRNAs, presumably by annealing to the mature miRNA guide strand and inducing degradation or stoichiometric duplex formation. We have reported that transfection of FUOX-resistant colon cancer cells with anti-sense miR-21 induces differentiation, as evidenced by the stimulation of alkaline phosphates activity and increased expression of CK-20 [55]. Additionally, we reported that these differentiating/differentiated cells become highly susceptible to difluorinated curcumin (CDF), a synthetic analog of curcumin, which we have shown to induce apoptosis of FUOX-resistant cells [55, 56].
Chemically modified antisense oligonucleotides such as locked nucleic acid (LNA), 2′–O-methyl and phosphorothioate show increased stability, binding affinity and specificity [9]. Krutzfeldt et al. reported the synthesis of 2′–O-methyl-modified cholesterol-conjugated single-stranded RNA analogues, with phosphorothiolate (phosphor-orthothiolate) linkages, named “antagomirs” complementary to miR-122 and miR-16. When these antagomirs were injected into mice, silencing of endogenous miR-122 and miR-16 was observed; the antagomirs were stable and the effects were still observed 23 days after injections [57]. We have observed that the growth of xenograft of FUOX-resistant colon cancer cells in SCID mice could be greatly inhibited by administration of antagomir-21 (unpublished observation). Residual colon tumors from antagomir-21-treated SCID mice showed suppression of colon CSC marker CD44, indicating decrease in CSCs in the tumor (unpublished observation).
Locked nucleic acids (LNA) are a class of high-affinity RNA analogs, in which the ribose ring is “locked” by a methylene bridge between the 2′–O and 4′–C atoms. As a result, LNA oligonucleotides display unprecedented thermal stability when hybridized to a complementary RNA or DNA strand. In addition, they display excellent mismatch discrimination and high aqueous solubility. Elmen and colleagues reported that combining LNA anti-miR with phosphorothiolate modifications markedly improved delivery of the compounds and silenced target miR-122 more efficiently compared with the antagomirs [58]. Ebert and colleagues reported another miRNA inhibitor: miRNA sponges [59]. These competitive inhibitors are transcripts expressed from strong promoters, containing multiple, tandem binding sites to a miRNA of interest. When vectors encoding these sponges are transfected into cultured cells, sponges de-repress miRNA targets at least as strongly as chemically modified antisense oligonucleotides. They specifically inhibit miRNAs with a complementary heptameric seed, such that a single sponge can be used to block an entire miRNA seed family.
8.2 Restoring Expression of Tumour-Suppressor miRNAs in Cancer
The loss or down-regulation of a tumor-suppressor miRNAs could be overcome by introducing synthetic oligonucleotides, known as miRNA mimics or miRNA precursor (pre-miRNA). The miRNA mimics are usually small, double stranded and may be chemically modified (2′–O-methyl with phosphorothioate modifications). However, Ibrahim and colleagues reported that when unmodified miR-145 and miR-33a were delivered into mouse xenograft tumors, it caused profound antitumor effects [44]. As stated above, we also noted a similar phenomenon. miR-145 delivery reduced tumor proliferation and increased apoptosis, with concomitant repression of c-Myc and ERK5 as novel regulatory target of miR-145 [40]. Similarly, antitumor effects of miR-33a were validated in this model. miRNA with chemical modifications have been demonstrated to enhance stability and delivery to tissues. However, care has to be taken to avoid off-target effects due to extensive modifications.
As for the in vivo delivery strategies, a number of approaches have been attempted such as the use of adeno-associated virus (AAV) vectors. Kota and colleagues first reported the use of adenovirus-associated (AAV) vectors to deliver miR-26a in a mouse model of hepatocellular carcinoma [60]. The authors cloned mir-26 into an AAV vector and viral particles were tested in an established Myc-dependent liver cancer mouse model. Intravenous injection of viral miRNA26a resulted in the suppression of tumorigenicity by repressing cell growth and by inducing tumour apoptosis without signs of toxicity.
Compared with a number of approaches for the in vivo delivery, the advantage in AAV vector is its small size and self-complementary genome, which enhances therapeutic gene expression. In addition, they are eliminated efficiently with minimal toxicity, as shown in Phase I and Phase II clinical trials[61]. Another advantage of AAV vectors is the efficient transduction of target cells. The development of self-complementary genome and non-human primate AAV serotype allows more than 90 % transduction efficiency of hepatocytes and long-term gene expression without toxicity, following a single systemic administration of recombinant virus [61].
Many factors contribute to colorectal cancer specific chemo-resistance mechanism. The miRNAs and mRNA regulated network have been shown to play major roles in cancer stem cell acquired ability of self-renewal and resistance to chemotherapeutics agents. We are now in the middle of a critical and exciting time in miRNA-CSC based cancer research and anticancer therapeutic development. The miRNA-based therapy targeting cancer stem cells shows the potential to overcome drug resistance and be highly beneficial to patients.
9 Conclusion
Many factors contribute to colorectal cancer specific chemo-resistance mechanisms. Based on the available data it can be concluded that miRNAs and miRNA regulated networks play major roles in cancer stem cells’ acquired ability of self-renewal and resistance to chemotherapeutic agents. oncomiRNA21 and tumor suppressor miR145 have emerged as critical gene regulators, which modulate multiple signaling pathways. Up-regulation of miRNA21, down-regulation of miR145 and their positive feedback in chemo-resistant colon cancer cells suggest that they are important players in the process of chemo-resistance. Targeted therapies that can modulate these and other miRNAs involved in regulating chemoresistance in colorectal cancer will be highly beneficial in overcoming drug-resistance in patients with recurrent colorectal cancer.
References
Bartel D (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Lewis BP, Burge CB, Bartel DP (2005) Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120(1):15–20
Kloosterman WP, Plasterk RH (2006) The diverse functions of microRNAs in animal development and disease. Dev Cell 11(4):441–450
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A (2004) Identification of mammalian microRNA host genes and transcription units. Genome Res 14(10A):1902–1910
Sevignani C, Calin GA, Nnadi SC, Shimizu M, Davuluri RV, Hyslop T, Demant P, Croce CM, Siracusa LD (2007) MicroRNA genes are frequently located near mouse cancer susceptibility loci. Proc Natl Acad Sci U S A 104(19):8017–8022
Lee Y, Jeon K, Lee JT, Kim S, Kim VN (2002) MicroRNA maturation: stepwise processing and subcellular localization. EMBO J 21(17):4663–4670
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Radmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425(6956):415–419
Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (2005) Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123(4):631–640
Garzon R, Marcucci G, Croce CM (2010) Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov 9(10):775–789
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435(7043):834–838
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, Rassenti L, Kipps T, Negrini M, Bullrich F, Croce CM (2002) Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A 99(24):15524–15529
Calin GA, Croce CM (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6(11):857–866
Arends JW (2000) Molecular interactions in the Vogelstein model of colorectal carcinoma. J Pathol 190(4):412–416
Fearon ER, Vogelstein B (1990) A genetic model for colorectal tumorigenesis. Cell 61(5):759–767
Vogelstein B, Fearon ER, Hamilton SR, Kern SE, Preisinger AC, Leppert M, Nakamura Y, White R, Smits AM, Bos JL (1988) Genetic alterations during colorectal-tumor development. N Engl J Med 319(9):525–532
Rubinfeld B, Robbins P, El-Gamil M, Albert I, Porfiri E, Polakis P (1997) Stabilization of beta-catenin by genetic defects in melanoma cell lines. Science 275(5307):1790–1792
Markowitz SD, Bertagnolli MM (2009) Molecular origins of cancer: molecular basis of colorectal cancer. N Engl J Med 361(25):2449–2460
Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA Jr, Kinzler KW (2013) Cancer genome landscapes. Science 339(6127):1546–1558
Bozic I, Antal T, Ohtsuki H, Carter H, Kim D, Chen S, Karchin R, Kinzler KW, Vogelstein B, Nowak MA (2010) Accumulation of driver and passenger mutations during tumor progression. Proc Natl Acad Sci U S A 107(43):18545–18550
Patel BB, Yu Y, Du J, Levi E, Phillip PA, Majumdar AP (2009) Age-related increase in colorectal cancer stem cells in macroscopically normal mucosa of patients with adenomas: a risk factor for colon cancer. Biochem Biophys Res Commun 378(3):344–347
Sanders MA, Majumdar APN (2011) Colon cancer stem cells: implications in carcinogenesis. Front Biosci Landmark 16:1651–1662
Kindler HL, Shulman KL (2001) Metastatic colorectal cancer. Curr Treat Options Oncol 2(6):459–471
Coutinho AK, Rocha Lima CM (2003) Metastatic colorectal cancer: systemic treatment in the new millennium. Cancer Control 10(3):224–238
Patel BB, Sengupta R, Qazi S, Vachhani H, Yu Y, Rishi AK, Majumdar AP (2008) Curcumin enhances the effects of 5-fluorouracil and oxaliplatin in mediating growth inhibition of colon cancer cells by modulating EGFR and IGF-1R. Int J Cancer 122(2):267–273
Yu Y, Kanwar SS, Patel BB, Nautiyal J, Sarkar FH, Majumdar AP (2009) Elimination of colon cancer stem-like cells by the combination of curcumin and FOLFOX. Transl Oncol 2(4):321–328
Yu Y, Kanwar SS, Patel BB, Oh PS, Nautiyal J, Sarkar FH, Majumdar AP (2012) MicroRNA-21 induces stemness by downregulating transforming growth factor beta receptor 2 (TGFbetaR2) in colon cancer cells. Carcinogenesis 33(1):68–76
Dean M, Fojo T, Bates S (2005) Tumour stem cells and drug resistance. Nat Rev Cancer 5(4):275–284
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T (2001) Identification of novel genes coding for small expressed RNAs. Science 294(5543):853–858
Fujita S, Ito T, Mizutani T, Minoguchi S, Yamamichi N, Sakurai K, Iba H (2008) miR-21 gene expression triggered by AP-1 is sustained through a double-negative feedback mechanism. J Mol Biol 378(3):492–504
Talotta F, Cimmino A, Matarazzo MR, Casalino L, De Vita G, D’Esposito M, Di Lauro R, Verde P (2009) An autoregulatory loop mediated by miR-21 and PDCD4 controls the AP-1 activity in RAS transformation. Oncogene 28(1):73–84
Zhang Z, Zha Y, Hu W, Huang Z, Gao Z, Zang Y, Chen J, Dong L, Zhang J (2013) The auto-regulatory feedback loop of microRNA-21/Programmed cell death protein 4/Activation Protein-1 (miR-21/PDCD4/AP-1) as a driving force for hepatic fibrosis development. J Biol Chem 288:37082–37093
Kumarswamy R, Volkmann I, Thum T (2011) Regulation and function of miRNA-21 in health and disease. RNA Biol 8(5):706–713
Velu CS, Baktula AM, Grimes HL (2009) Gfi1 regulates miR-21 and miR-196b to control myelopoiesis. Blood 113(19):4720–4728
Wickramasinghe NS, Manavalan TT, Dougherty SM, Riggs KA, Li Y, Klinge CM (2009) Estradiol downregulates miR-21 expression and increases miR-21 target gene expression in MCF-7 breast cancer cells. Nucleic Acids Res 37(8):2584–2595
Kern HB, Niemeyer BF, Parrish JK, Kerr CA, Yaghi NK, Prescott JD, Gutierrez-Hartmann A, Jedlicka P (2012) Control of MicroRNA-21 expression in colorectal cancer cells by oncogenic epidermal growth factor/Ras signaling and Ets transcription factors. DNA Cell Biol 31(8):1403–1411
Asangani IA, Rasheed SA, Nikolova DA, Leupold JH, Colburn NH, Post S, Allgayer H (2008) MicroRNA-21 (miR-21) post-transcriptionally downregulates tumor suppressor Pdcd4 and stimulates invasion, intravasation and metastasis in colorectal cancer. Oncogene 27(15):2128–2136
Sayed D, Rane S, Lypowy J, He M, Chen IY, Vashistha H, Yan L, Malhotra A, Vatner D, Abdellatif M (2008) MicroRNA-21 targets Sprouty2 and promotes cellular outgrowths. Mol Biol Cell 19(8):3272–3282
Oh PS, Patel VB, Sanders MA, Kanwar SS, Yu Y, Nautiyal J, Patel BB, Majumdar AP (2011) Schlafen-3 decreases cancer stem cell marker expression and autocrine/juxtacrine signaling in FOLFOX-resistant colon cancer cells. Am J Physiol Gastrointest Liver Physiol 301(2):G347–G355
Sachdeva M, Zhu S, Wu F, Wu H, Walia V, Kumar S, Elble R, Watabe K, Mo YY (2009) p53 represses c-Myc through induction of the tumor suppressor miR-145. Proc Natl Acad Sci U S A 106(9):3207–3212
Spizzo R, Nicoloso MS, Lupini L, Lu Y, Fogarty J, Rossi S, Zagatti B, Fabbri M, Veronese A, Liu X, Davuluri R, Croce CM, Mills G, Negrini M, Calin GA (2010) miR-145 participates with TP53 in a death-promoting regulatory loop and targets estrogen receptor-alpha in human breast cancer cells. Cell Death Differ 17(2):246–254
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K (2009) Modulation of microRNA processing by p53. Nature 460(7254):529–533
Kent OA, Chivukula RR, Mullendore M, Wentzel EA, Feldmann G, Lee KH, Liu S, Leach SD, Maitra A, Mendell JT (2010) Repression of the miR-143/145 cluster by oncogenic Ras initiates a tumor-promoting feed-forward pathway. Genes Dev 24(24):2754–2759
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R, Ramirez N, Abajo A, Navarro A, Moreno I, Monzo M, Garcia-Foncillas J (2006) Identification by Real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 5:29
Ibrahim AF, Weirauch U, Thomas M, Grunweller A, Hartmann RK, Aigner A (2011) MicroRNA replacement therapy for miR-145 and miR-33a is efficacious in a model of colon carcinoma. Cancer Res 71(15):5214–5224
Gotte M, Mohr C, Koo CY, Stock C, Vaske AK, Viola M, Ibrahim SA, Peddibhotla S, Teng YH, Low JY, Ebnet K, Kiesel L, Yip GW (2010) miR-145-dependent targeting of junctional adhesion molecule A and modulation of fascin expression are associated with reduced breast cancer cell motility and invasiveness. Oncogene 29(50):6569–6580
Sachdeva M, Mo YY (2010) MicroRNA-145 suppresses cell invasion and metastasis by directly targeting mucin 1. Cancer Res 70(1):378–387
Zhang J, Guo H, Zhang H, Wang H, Qian G, Fan X, Hoffman AR, Hu JF, Ge S (2011) Putative tumor suppressor miR-145 inhibits colon cancer cell growth by targeting oncogene Friend leukemia virus integration 1 gene. Cancer 117(1):86–95
Xu N, Papagiannakopoulos T, Pan G, Thomson JA, Kosik KS (2009) MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 137(4):647–658
Akao Y, Nakagawa Y, Hirata I, Iio A, Itoh T, Kojima K, Nakashima R, Kitade Y, Naoe T (2010) Role of anti-oncomirs miR-143 and -145 in human colorectal tumors. Cancer Gene Ther 17(6):398–408
Kamatani A, Nakagawa Y, Akao Y, Maruyama N, Nagasaka M, Shibata T, Tahara T, Hirata I (2013) Downregulation of anti-oncomirs miR-143/145 cluster occurs before APC gene aberration in the development of colorectal tumors. Med Mol Morphol 46(3):166–171
Patel VB, Yu Y, Das JK, Patel BB, Majumdar AP (2009) Schlafen-3: a novel regulator of intestinal differentiation. Biochem Biophys Res Commun 388:752–756
Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N, Yuen ST, Chan TL, Kwong DL, Au GK, Liu CG, Calin GA, Croce CM, Harris CC (2008) MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 299(4):425–436
Valeri N, Gasparini P, Braconi C, Paone A, Lovat F, Fabbri M, Sumani KM, Alder H, Amadori D, Patel T, Nuovo GJ, Fishel R, Croce CM (2010) MicroRNA-21 induces resistance to 5-fluorouracil by down-regulating human DNA MutS homolog 2 (hMSH2). Proc Natl Acad Sci U S A 107(49):21098–103
Hatley ME, Patrick DM, Garcia MR, Richardson JA, Bassel-Duby R, van Rooij E, Olson EN (2010) Modulation of K-Ras-dependent lung tumorigenesis by MicroRNA-21. Cancer Cell 18(3):282–293
Yu Y, Sarkar FH, Majumdar AP (2013) Down-regulation of miR-21 induces differentiation of chemoresistant colon cancer cells and enhances susceptibility to therapeutic regimens. Transl Oncol 6(2):180–186
Kanwar SS, Yu Y, Nautiyal J, Patel BB, Padhye S, Sarkar FH, Majumdar AP (2011) Difluorinated-curcumin (CDF): a novel curcumin analog is a potent inhibitor of colon cancer stem-like cells. Pharm Res 28(4):827–838
Krutzfeldt J, Rajewsky N, Braich R, Rajeev KG, Tuschl T, Manoharan M, Stoffel M (2005) Silencing of microRNAs in vivo with ‘antagomirs’. Nature 438(7068):685–689
Elmen J, Lindow M, Silahtaroglu A, Bak M, Christensen M, Lind-Thomsen A, Hedtjarn M, Hansen JB, Hansen HF, Straarup EM, McCullagh K, Kearney P, Kauppinen S (2008) Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Res 36(4):1153–1162
Ebert MS, Neilson JR, Sharp PA (2007) MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods 4(9):721–726
Kota J, Chivukula RR, O’Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, Chang TC, Vivekanandan P, Torbenson M, Clark KR, Mendell JR, Mendell JT (2009) Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137(6):1005–1017
Michelfelder S, Trepel M (2009) Adeno-associated viral vectors and their redirection to cell-type specific receptors. Adv Genet 67:29–60
Zhou J, Zhou Y, Yin B, Hao W, Zhao L, Ju W, Bai C (2010) 5-Fluorouracil and oxaliplatin modify the expression profiles of microRNAs in human colon cancer cells in vitro. Oncol Rep 23(1):121–128
Nishida N, Yamashita S, Mimori K, Sudo T, Tanaka F, Shibata K, Yamamoto H, Ishii H, Doki Y, Mori M (2012) MicroRNA-10b is a prognostic indicator in colorectal cancer and confers resistance to the chemotherapeutic agent 5-fluorouracil in colorectal cancer cells. Ann Surg Oncol 19(9):3065–3071
Cekaite L, Rantala JK, Bruun J, Guriby M, Agesen TH, Danielsen SA, Lind GE, Nesbakken A, Kallioniemi O, Lothe RA, Skotheim RI (2012) MiR-9, -31, and -182 deregulation promote proliferation and tumor cell survival in colon cancer. Neoplasia 14(9):868–879
Shen K, Liang Q, Xu K, Cui D, Jiang L, Yin P, Lu Y, Li Q, Liu J (2012) MiR-139 inhibits invasion and metastasis of colorectal cancer by targeting the type I insulin-like growth factor receptor. Biochem Pharmacol 84(3):320–330
Takaoka Y, Shimizu Y, Hasegawa H, Ouchi Y, Qiao S, Nagahara M, Ichihara M, Lee JD, Adachi K, Hamaguchi M, Iwamoto T (2012) Forced expression of miR-143 represses ERK5/c-Myc and p68/p72 signaling in concert with miR-145 in gut tumors of Apc(Min) mice. PLoS One 7(8):e42137
Liu L, Chen L, Xu Y, Li R, Du X (2010) microRNA-195 promotes apoptosis and suppresses tumorigenicity of human colorectal cancer cells. Biochem Biophys Res Commun 400(2):236–240
Nagel R, le Sage C, Diosdado B, van der Waal M, Oude Vrielink JA, Bolijn A, Meijer GA, Agami R (2008) Regulation of the adenomatous polyposis coli gene by the miR-135 family in colorectal cancer. Cancer Res 68(14):5795–5802
Rossi L, Bonmassar E, Faraoni I (2007) Modification of miR gene expression pattern in human colon cancer cells following exposure to 5-fluorouracil in vitro. Pharmacol Res 56(3):248–253
Kurokawa K, Tanahashi T, Iima T, Yamamoto Y, Akaike Y, Nishida K, Masuda K, Kuwano Y, Murakami Y, Fukushima M, Rokutan K (2012) Role of miR-19b and its target mRNAs in 5-fluorouracil resistance in colon cancer cells. J Gastroenterol 47(8):883–895
Yan HJ, Liu WS, Sun WH, Wu J, Ji M, Wang Q, Zheng X, Jiang JT, Wu CP (2012) miR-17-5p inhibitor enhances chemosensitivity to gemcitabine via upregulating Bim expression in pancreatic cancer cells. Dig Dis Sci 57(12):3160–3167
Tsuchida A, Ohno S, Wu W, Borjigin N, Fujita K, Aoki T, Ueda S, Takanashi M, Kuroda M (2011) miR-92 is a key oncogenic component of the miR-17-92 cluster in colon cancer. Cancer Sci 102(12):2264–2271
Amodeo V, Bazan V, Fanale D, Insalaco L, Caruso S, Cicero G, Bronte G, Rolfo C, Santini D, Russo A (2013) Effects of anti-miR-182 on TSP-1 expression in human colon cancer cells: there is a sense in antisense? Expert Opin Ther Targets 17(11):1249–1261
Wang CJ, Stratmann J, Zhou ZG, Sun XF (2010) Suppression of microRNA-31 increases sensitivity to 5-FU at an early stage, and affects cell migration and invasion in HCT-116 colon cancer cells. BMC Cancer 10:616
Valastyan S, Reinhardt F, Benaich N, Calogrias D, Szasz AM, Wang ZC, Brock JE, Richardson AL, Weinberg RA (2009) A pleiotropically acting microRNA, miR-31, inhibits breast cancer metastasis. Cell 137(6):1032–1046
Roy S, Levi E, Majumdar AP, Sarkar FH (2012) Expression of miR-34 is lost in colon cancer which can be re-expressed by a novel agent CDF. J Hematol Oncol 5:58
Yamakuchi M, Ferlito M, Lowenstein CJ (2008) miR-34a repression of SIRT1 regulates apoptosis. Proc Natl Acad Sci U S A 105(36):13421–13426
Liu C, Cheng H, Shi S, Cui X, Yang J, Chen L, Cen P, Cai X, Lu Y, Wu C, Yao W, Qin Y, Liu L, Long J, Xu J, Li M, Yu X (2013) MicroRNA-34b inhibits pancreatic cancer metastasis through repressing Smad3. Curr Mol Med 13(4):467–478
Fearon ER (2011) Molecular genetics of colorectal cancer. Ann Rev Pathol 6:479–507
Lao VV, Grady WM (2011) Epigenetics and colorectal cancer. Nat Rev Gastroenterol Hepatol 8(12):686–700
Acknowledgements
This work was supported by grants to Dr. Majumdar from the National Institutes of Health/National Institute on Aging (AG014343) and the Department of Veterans Affairs.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Yu, Y., Nangia-Makker, P., Majumdar, A.P.N. (2014). Overcoming Drug Resistance in Colorectal Cancer by MicroRNAs. In: Sarkar, F. (eds) MicroRNA Targeted Cancer Therapy. Springer, Cham. https://doi.org/10.1007/978-3-319-05134-5_8
Download citation
DOI: https://doi.org/10.1007/978-3-319-05134-5_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-05133-8
Online ISBN: 978-3-319-05134-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)