Abstract
The clinical use of microRNAs (miRNAs) as diagnostic tools for example for tumor classification or as prognostic markers is becoming increasingly established. In addition, recent studies demonstrated that miRNAs could be used as new therapeutic approach in anticancer treatment including the highly interesting aspect that is regulated by miRNAs: resistance to chemo- and radiotherapy. This chapter aims to elucidate the impact of miRNAs on drug resistance from a clinical point of view, and to highlight their potential role as predictors or modifiers of resistance towards chemotherapeutics and radiotherapy. Therefore, we selected exemplary two different tumor types that present either high or low resistance to chemotherapeutic treatment: esophageal cancer (highly therapy resistant tumor) and ovarian cancer (quite therapy sensitive tumor).
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
MicroRNAs (miRNAs) are a novel class of regulatory molecules that control translation and stability of mRNAs on a post-transcriptional level. MiRNAs are involved in almost all physiological processes such as cell development or differentiation. So far, more over 1,000 human miRNAs have been identified (Kozomara and Griffith-Jones 2011), and each single miRNA can target hundreds of mRNAs (Li and Yang 2013). However, most importantly from a clinical point of view, miRNAs are highly involved in the initiation and progression of cancer by regulating for example metastasis and angiogenesis amongst others. Interesting in this context is that hundreds of miRNAs map to regions of the human genome that are known to be altered in cancer (Calin et al. 2004). Esquela-Kerschner et al. even established the term “oncomiRs” for miRNAs with oncogenic function, implying that abnormalities in miRNA expression might directly result in the de-differentiation of cells, allowing tumor formation to occur (Esquela-Kerschner and Slack 2006).
1.1 General Considerations: miRNAs and Their Clinical Use in Cancer Diagnostics and Treatment
Most oncological studies in the recent past investigated miRNA expression pattern in fresh frozen samples such as tumor biopsies or resection specimens, but miRNAs can also be detected and extracted from other sample types such as paraffin-embedded tissues (Iorio and Croce 2012). Most interestingly for clinicians, miRNAs can further be found as so called “Circulating miRNA” in a variety of human body fluids in healthy volunteers and cancer patients (Fang et al. 2012; Weber et al. 2010; Xiao et al. 2013; Allegra et al. 2012; Iorio and Croce 2012). These circulating miRNAs, which are surprisingly stable (Mo et al. 2012; Kim and Reitmair 2013) and can be detected even in 10-year-old human serum samples or in un-refrigerated dried serum blots (Cortez et al. 2011), are tissue-specific, stable, reproducible and consistent among individuals in the same species (Fang et al. 2012). In cancer patients, circulating miRNAs are thought to be mainly related to apoptosis and necrosis of cancer cells in the tumor microenvironment (Kim and Reitmaier 2013), and miRNA expression profiles of primary tumors and metastases seem very similar (Rosenwald et al. 2010).
With this in mind, the clinical use of miRNAs as diagnostic tools for example for tumor classification or as prognostic markers seems logical and feasible. In addition, latest research provides first very promising data that miRNAs could be used as new therapeutic approach in the fight against cancer as for example knockdown of oncomirs can affect tumor growth. In this context, only recently the focus has been turned on another highly interesting aspect that is regulated by miRNAs: resistance to chemo- and radiotherapy.
This chapter aims to elucidate the impact of miRNAs on drug resistance from a clinical point of view, and to highlight their potential role as predictors or modifiers of resistance towards chemotherapeutics and radiotherapy. Therefore, we selected exemplary two different tumor types that present either high or low resistance to chemotherapeutic treatment: (a) esophageal cancer, as a highly therapy resistant tumor with a complete response to neoadjuvant therapy in only about 13–25 % of all patients (Courrech Staal et al. 2010) and (b) ovarian cancer, as a quite therapy sensitive tumor with complete response to adjuvant therapy in about 80–90 % of the patients (Ozols 2005).
1.2 High Versus Low Resistant Tumors: On Overview About the Clinical Background
1.2.1 Esophageal Cancer
Esophageal cancer (EC) is characterized by a poor overall prognosis. Because of the high incidence of advanced disease at the time of diagnosis, the 5-year survival rate remains below 15 % and only about 15–20 % of patients finally qualify for curative surgical resection. In an attempt to improve outcome of patients after surgery and to potentially increase the number of patients who qualify for surgery by downstaging of the tumor, neoadjuvant therapy including chemotherapy and radiotherapy has been demonstrated to potentially advance overall survival for both histological subtypes adenocarcinoma (AdenoCA) and squamous cell carcinoma (SCC) (Urschel and Vasan 2003; Fiorica et al. 2004; Sjoquist et al. 2011). However, a complete pathologic response as determined by the “tumor regression grade TRG” can only be achieved in about 13–25 % of all patients (Courrech Staal et al. 2010).
There are several miRNAs that have been reported to be differentially expressed in esophageal cancer, with (prognostic and diagnostic) associations to tumor stage (Lin et al. 2012), histological differentiation (Hummel et al. 2011a; Lin et al. 2012), distant lymph node metastasis (Hummel et al. 2011a; Liu et al. 2012a), vascular invasion (Komatsu et al. 2011), overall and disease-free survival (Komatsu et al. 2012; Takeshita et al. 2013; Zhang et al. 2011) and tumor recurrence (Komatsu et al. 2011; Hummel et al. 2011a). Additionally, a number of serum miRNAs were described as potential diagnostic biomarkers (Komatsu et al. 2011; Takeshita et al. 2013; Zhang et al. 2011; Liu et al. 2012a; Cai et al. 2012; Wang and Zhang 2012; Zhang et al. 2010, 2012; Revilla-Nuin et al. 2013).
1.2.2 Ovarian Cancer
Ovarian cancer is one of the most aggressive female reproductive tract tumors. The prognosis depends on the stage of the disease and on histological and molecular characteristics. Platinum based chemotherapy agents, namely cisplatin and carboplatin, are widely used for the treatment of ovarian cancer. For advanced-stage disease, taxanes (e.g. paclitaxel, docetaxel) are often supplemented (van Jaarsveld et al. 2010). Despite the fact that a complete clinical response can be achieved in 80–90 and 50 % of patients with early-stage or advanced-stage disease, respectively, (Ozols 2005), ovarian cancer patients frequently develop resistance to chemotherapy, often resulting in a poor overall 5-year survival of only 30 % (Moss and Kaye 2002).
Several studies showed aberrantly expressed miRNAs in ovarian cancer and established a connection to histological subtypes (Lee et al. 2012; Zaman et al. 2012; Iorio et al. 2007), tumor stage or grade (Yang et al. 2012), primary or recurrent tumors (Hu et al. 2009), and survival (Hong et al. 2013; Peng et al. 2012; Lee et al. 2012; Marchini et al. 2011; Nam et al. 2008). Again, miRNAs could also be identified as diagnostic and prognostic biomarkers in serum (Xu et al. 2013; Resnick et al. 2009; Chang et al. 2012; Peng et al. 2012).
2 Clinical Application of miRNAs as Diagnostic Tools: MicroRNAs as Predictors of Response to Conventional Treatments
As outlined above, one highly promising clinical application of miRNAs as diagnostic tools involves their potential to predict response to conventional treatment such as chemotherapy and radiotherapy. If it might be possible to identify responders and non-responders before the start of neoadjuvant or adjuvant treatment, cancer therapy could be tailored more individually. Patients who do not benefit from chemotherapy or irradiation would not have to undergo this toxic treatment, and could be referred immediately to curative surgical resection. The use of miRNAs as predictors of therapy response implicates however that chemotherapy or radiotherapy resistant tumors exhibit distinct miRNA expression pattern that distinguishes them from sensitive tumors. These differences in miRNA expression would be necessary to allocate patients into the responder and non-responder groups. And indeed, there is growing evidence that chemo- and radiotherapy resistant tumors show specific pattern of miRNA deregulation, both in vitro and in vivo.
2.1 Experimental In Vitro Data
We found only one study that investigated the direct effect of chemotherapy treatment on miRNA expression in one adenocarcinoma and one squamous cell carcinoma cell lines after treatment with either cisplatin or 5-fluorouracil for 24/72 h. The authors could show that 13 miRNAs (miR-199a-5p, miR-302f, miR-320a, miR-342-3p, miR-425, miR-455-3p, miR-486-3p, miR-519c-5p, miR-548d-5p, miR-617, miR-758, miR-766, miR-1286) were deregulated after short-term or long-term treatment in either of the cell lines (Hummel et al. 2011b). However, a number of studies were published that report different miRNA expression pattern between drug resistant and sensitive cells. For example, comparing two cisplatin resistant human esophageal squamous cell lines (that were generated via exposure of sensitive cells to the chemotherapeutic drug) with controls, Sugimura et al. identified a total of 365 miRNAs to be differentially expressed between resistant and sensitive cells, with more than 1.7-fold changes in expression of 128 respectively 177 miRNAs. Most interestingly, 15 miRNAs showed an overlap between the two resistant cell lines with regards to their deregulation: miR-135a, miR-96, miR-141, miR-101, miR-146a, miR-489 and miR-545 were up-regulated, whereas miR-99a, let-7b, miR-204, let-7c, miR-202, miR-10a, miR-136 and miR-145 were down-regulated in both cisplatin-resistant cell limes (Sugimura et al. 2012). Another study reported that miR-141, miR-21, miR-19b, miR-200a, miR-19a, miR-27a, miR-20a and miR-20b were expressed at significantly higher levels, and miR-205 and miR-224 at significantly lower levels in cells with increasing resistance towards cisplatin. In this context, most profound deregulation was found for miR-141 (Imanaka et al. 2011). Concerning altered miRNA profiles in radioresistant cells, Zheng et al. compared a radioresistant squamous cell carcinoma cell line with controls and found 35 miRNAs to be deregulated: 10 miRNAs (miR-1539, miR-1237, miR-92b, etc.) were up-regulated, and 25 miRNAs (miR-185, miR-18b, miR-17, etc.) were down-regulated (Zheng et al. 2011).
For ovarian cancer, there are a number of reports published on deregulated miRNAs in chemotherapy resistant cell lines. For example, Kumar et al. compared sensitive human ovarian cancer cells and their cisplatin-resistant counterparts and found changes in the expression of 11 miRNAs out of 1,500 miRNAs, with miRplus-F1064, miR-300, miR-193b, miR-642 and miR-1299 being up-regulated and miR-625, miR-20b, miRPlus-F1147, let-7c, miRPlus-F1231 and miR-542-3p being down-regulated (Kumar et al. 2011). Van Jaarsveld reported 27 miRNAs to be differentially expressed in cells with increasing resistance towards cisplatin: miR-214, miR-412, miR-645, miR-17, miR-106a, miR-199a-5p, miR-215, miR-199a/b-3p, miR-335, miR-338-5p, miR-493, miR-135b, miR-130a, miR-186, miR-942, miR-18b, miR-20b, miR-196a, miR-10b, miR-19a, miR-421, miR-19b, miR-518e, miR-631, miR-222, miR-141 and miR-200c (van Jaarsveld et al. 2012). Another study identified diversely expressed miRNAs in resistant cell lines using one cisplatin-resistant and three paclitaxel-resistant cell lines: let-7e, miR-30c, miR-125b, miR-130a and miR-335. Interestingly, let-7e was up-regulated in one of the paclitaxel resistant cell line, while it was down-regulated in the other resistant cell lines. The opposite phenomenon was described for miR-125b, which was down-regulated in one paclitaxel resistant cell line and up-regulated in the other ones. MiR-30c, miR-130a and miR-335 were down-regulated in all the resistant cell lines (Sorrentino et al. 2008). Yang et al. detected 79 differently expressed miRNAs in a cisplatin-resistant cell line, including miR-130a associated with MDR1/P-gp-mediated drug resistance (Yang et al. 2012).
These experimental in vitro data clearly support the hypothesis that chemotherapy or radiotherapy resistant tumors present unique miRNA expression pattern that might allow identification of therapy responders based on profiling information in tumor cells. With regard to either high or low resistant tumors, 53 miRNAs were described in esophageal cancer to correlate with resistance, and 124 miRNAs were associated with response to therapy in ovarian cancer cells. Most interestingly, 18 miRNAs were identified in more than one study to present altered expression between resistant and sensitive tumors in vitro, with a few of them even presenting similar deregulation pattern when comparing esophageal (high resistant) and ovarian (low resistant) cancer. Table 15.1 provides an overview about these 18 miRNAs that seem to highly impact on drug resistance.
2.2 First Clinical Data
Odenthal et al. examined 80 patients with esophageal cancer (AdenoCA and SCC), who underwent multimodal therapy. Comprehensive miRNA profiling identified a number of miRNAs in pretherapeutic biopsies that were significantly differently expressed between major and minor responders. The pretherapeutic intratumoral expression of miR-192 and miR-194 was significantly associated with the histopathologic response of esophageal squamous cell carcinoma to neoadjuvant treatment (Odenthal et al. 2013). Also using pretreatment biopsy specimen of 25 patients who underwent irinotecan/cisplatin based chemotherapy and radiotherapy followed by surgical treatment, 71 miRNAs were found to be significantly differently expressed between pathologic complete responders and non-responders. Five of these miRNAs had a greater than two-fold difference in expression: HAS-240, miR-296, miR-141, miR-31 and miR-217. Comparison of post-treatment biopsies of responders versus non-responder patients further revealed that 52 miRNAs were significantly up-regulated or down-regulated after induction therapy, and nine of these had a greater than two-fold change in expression: miR-1238, miR-938, HS_228.1, HS_282, miR-200a, miR-200b, miR-429 and miR-141 amongst others. Patients with high levels of miR-135b or miR-145 in the posttreatment biopsy specimens had a significantly shorter median disease-free survival compared to patients with low levels (11.5 versus 5.1 months; p = 0.04; 11.5 versus 2.8 months; p = 0.03) (Ko et al. 2012). Furthermore, miR-31 expression was found to be significantly reduced in patients presenting a poor histomorphologic response to neoadjuvant therapy. In addition, Lynam-Lennon et al. could demonstrate an influence of miR-31 on the modulation of radioresistance (Lynam-Lennon et al. 2012). Other studies evaluated the impact of miRNA expression on therapy response by defining therapy response as longer survival. Hamano et al. investigated the expression of 9 miRNAs (let-7a, let-7 g, miR-21, miR-134, miR-145, miR-155, miR-200c, miR-203 and miR-296) in esophageal cancer patients who had received preoperative chemotherapy with cisplatin and 5-FU followed by surgery. The expression of miR-200c correlated inversely and significantly with the response to chemotherapy. Furthermore, the overexpression of miR-200c and miR-21 respectively the underexpression of miR-145 correlated significantly with shorter overall survival (Hamano et al. 2011). Additionally and highly interesting regarding a potential clinical use of miRNAs as response predictors, miRNA-21 levels in serum were shown to be significantly reduced in esophageal squamous cell carcinoma patients who responded to chemotherapy (Kurashige et al. 2012). Another study demonstrated an inverse correlation between expression levels of miR-483 and miR-214 and overall survival (Zhou et al. 2013). Finally, low expression of let-7c correlated with poor prognosis and was able to predict response to cisplatin-based chemotherapy (Sugimura et al. 2012).
A similar situation regarding miRNA expression and its correlation to response to treatment can be observed in ovarian cancer patients. Eitan et al. compared the miRNA profile of surgically treated ovarian cancer patients that received either solely platinum based chemotherapy (n = 21), or paclitaxel with carboplatin (n = 34) and cyclophosphamid with cisplatin as first line treatment. Based on outcome, the authors divided the patients into platinum-sensitive vs. platinum-resistant patients. Seven miRNAs were identified to be significantly differently expressed between the two groups: miR-27a, miR-23a, miR-30c miR-7 g and miR-199-3p were increased in platinum-resistant patients, and miR-378 and miR-625 were increased in platinum-sensitive patients (Eitan et al. 2009). Lu et al. investigated let-7a expression in ovarian cancer patients receiving an adjuvant platinum based chemotherapy with or without paclitaxel in addition to surgical debulking. The let-7a expression did not correlate with disease stage, tumor grade, histology or debulking results. However, the authors could demonstrate that patients who responded well to platinum based chemotherapy combined with paclitaxel presented significantly lower let-7a levels. Conversely, survival analyses showed that patients with high let-7a levels presented a better survival compared to those with low levels (Lu et al. 2011). In another study, let-7i expression was found to be significantly reduced in chemotherapy resistant patients treated with paclitaxel and platinum (Yang et al. 2008). Most interestingly, patients who experienced a relapse of the disease showed a down-regulation of let-7 in samples collected after chemotherapy compared to pretherapeutic samples. Furthermore, a decrease in the expression of let-7 after chemotherapy negatively correlated with disease-free time before recurrence (Boyerinas et al. 2012). In this context, low levels of miR-199a may be another predictor for chemoresistance in recurrent tumors (Nam et al. 2008). Several studies further looked at different histological subtypes (serous versus non-serous) of ovarian cancer. For example, a signature of 23 miRNAs was associated with chemoresistance in patients with serous ovarian cancer treated with carboplatin and taxol, and PCR-based validation confirmed that three miRNAs were able to predict chemoresistance of these tumors: miR-484, miR-642 and miR-217 (Vecchione et al. 2013). Van Jaarsveld found higher miR-141 expression in patients with serous ovarian tumors who did not response to platinum-based chemotherapy (van Jaarsveld et al. 2012). Furthermore, low miR-376c expression was detected in patients with serous ovarian tumors who responded well to cisplatin based chemotherapy (Ye et al. 2011). Finally, another clinical study compared the expression of miR-21 and miR-214 in ascites and omental metastasis of patients with ovarian cancer treated with carboplatin. Malignant cells in ascites showed greater cell viability when treated with carboplatin compared to omental metastasis. Additionally, there was a significant up-regulation of miRNA-21 and miRNA-214 in tumor cells from ascites (Frederick et al. 2013).
Table 15.2 presents an overview about the data available so far on miRNAs as potential clinical predictors of chemotherapy resistance in esophageal and ovarian cancers. These first in vivo data clearly supports the results from the in vitro experiments and prove that, at least in the isolated patient populations of the respective publications, miRNAs can help separating responders from non-responders based on profiling information obtained from tumor specimen. However, the final clinical impact of miRNAs as potential response predictors remains to be determined, as the heterogeneity of the different treatment protocols in the studies, the respective experimental setups including the clinical response evaluation, and finally the obtained results in the different studies does not allow a definitive statement yet. For example, miR-31 was reported on the one hand to be up-regulated (Lynam-Lennon et al. 2012), and on the other hand to be down-regulated in esophageal cancer patients that respond to therapy (Ko et al. 2012). However, in summary these data are highly promising regarding a potential clinical benefit of miRNAs as response predictors in the future.
3 Clinical Application of miRNAs as Therapeutic Tools: MicroRNAs as Modifiers of Response to Conventional Treatments
Another possibly even more interesting approach for the clinical use of miRNAs might be their application as potential modifiers of chemotherapy. As the resistance of tumors to conventional treatment such as chemotherapy or radiotherapy represents a major obstacle in the fight against cancer, identification of a way to reverse drug resistance is one of the most important challenges for researchers all over the world. A solution to this problem might be a key breakthrough in the treatment of malignant diseases. If resistance to chemo- and radiotherapy can be overcome, toxicity of these treatments could be minimized by for example lowering the doses of chemotherapeutics, while achieving the same antitumor effects. And in fact, there is growing evidence that modulation of miRNA expression can affect resistance of various tumors to treatment. However, as this field of research is still very young, results are somewhat limited and refer so far only to in vitro experiments. But this does not affect its promising clinical significance.
A number of authors addressed the question whether miRNA modulation affects chemotherapy resistance in esophageal cancer. Zhang et al. for example demonstrated that down-regulation of miR-27a in esophageal cancer cells could significantly decrease the expression of P-glycoprotein [a well known drug efflux pump that influences on drug resistance (Wen et al. 2009)], Bcl-2 (an anti-apoptotic protein which is involved in tumor cell apoptosis and response to chemotherapy) (Kang and Reynolds 2009; Ballesta et al. 2013; Asakura and Ohkawa 2004; Ohkawa et al. 2004), and the transcription of the multidrug resistance gene. This conferred sensitivity towards P-glycoprotein related chemotherapeutics such as etoposide, doxorubicin and vinblastine (Zhang et al. 2010). Hong et al. discovered that decreased miR-296 expression improved the response of tumor cells to 5-FU and Ciplatin, probably due to changes in Bcl-2 and Bax levels, finally leading to an increase in apoptosis and decreased MDR-1 expression (Hong et al. 2010). Furthermore, miR-141 was shown to play an important role in the development of cisplatin resistance in esophageal squamous cell carcinoma by down-regulation of YAP1, which is known to have a crucial role in apoptosis induced by DNA-damaging agents (Imanaka et al. 2011). Another group found miR-148a to sensitize esophageal cancer cell lines to cisplatin and, to a lesser extent, to 5-flurouracil, and to attenuate resistance in chemotherapy-resistant variants (Hummel et al. 2011b). In a variety of tumors other than esophageal cancer, expression of miR-148a has been shown to negatively affect tumor growth, cell motility, invasion, migration and metastasis. One of the targets that can explain the effects of miR-148a modulation might be mitogen- and stress-activated kinase 1 (MSK1), which was identified in prostate cancer cells (Fujita et al. 2010). Other potential resistance-relevant targets of miR-148a include DNA methyltransferase-3B (DNMT3B) and DNA methyltransferase-1 (DNMT-1) (Merkerova et al. 2010; Duursma et al. 2009). Sugimura and colleagues demonstrated that let-7 modulated the chemosensitivity to cisplatin in esophageal cancer through the regulation of IL-6/STAT3 pathway (Sugimura et al. 2012). IL-6 is an inflammatory cytokine, which is released from macrophages and T-lymphocytes as well as from cancer cells. It modulates various cell functions (e.g., inflammatory reactions), and is a major activator of the JAK/STAT3 and PI3K/AKT signaling pathways. Various studies already demonstrated an association between IL-6 and resistance to chemotherapy for example in ovarian cancer (Wang et al. 2010), breast cancer (Iliopoulos et al. 2009) or gastrointestinal cancer (Chen et al. 2013). Additionally, with regards to esophageal cancer, one recent study showed that increasing intracellular IL-6 expression after cisplatin exposure is associated with reduced sensitivity to cisplatin treatment, and that knockdown of IL-6 expression restored sensitivity to cisplatin treatment (Chen et al. 2013). Finally, Wu et al. could show that overexpression of miRNA-200b, miR-200c and miR-429 correlated with resistance to cisplatin treatment. Chemotherapeutic drugs such as cisplatin induce expression of endogenous AP-2α, which contributes to chemosensitivity by enhancing therapy-induced apoptosis (Wu et al. 2011).
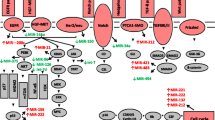
Again, a similar picture is found for ovarian cancer. For example, overexpression of miR-200c, miR-200a and miR-141 (the miR-200 family is known as the main suppressor of the epithelial-to-mesenchymal transition, EMT) was reported to enhance sensitivity to microtubule-targeting drugs (e.g. paclitaxel, vincristine, epothilone B) in ovarian cancer lines by repressing the class III β-tubulin TUBB3 (Cochrane et al. 2009; Prislei et al. 2013; Leskelä et al. 2010; Mateescu et al. 2011). TUBB3 is well known as a prominent mechanism of drug resistance found in a variety of solid tumors, but particularly in lung and ovarian cancer where it is associated with a perturbation in microtubule dynamics (Mariani et al. 2011). Accordingly, up-regulation of miR-200c levels in an ovarian cancer cell line increased the sensitivity towards micro-targeting drugs up to 85 % (Cochrane et al. 2010). Also, the family members miR-141/200c showed a correlation with cisplatin sensitivity in the NCI-60 panel. The NCI-60 cancer cell panel consists of 60 cancer cell lines of various histological origins, of which miRNA expression and drug sensitivity data can be obtained from a public database (Blower et al. 2007). Overexpression of miR-141 resulted in enhanced resistance to cisplatin in ovarian cancer cell lines as it directly targeted KEAP1, and induced cisplatin resistance via affection of the NF-κB pathway (van Jaarsveld et al. 2012). Another miRNA, miR-199a, was demonstrated to significantly increase the chemosensitivity of ovarian cancer-initiating cells to cisplatin, paclitaxel and adriamycin, and to reduce mRNA expression of the multidrug resistance gene ABCG2 (Chen et al. 2012). Further in vitro assays with knockdown of let-7i led to a decreased cisplatin-induced cell death in ovarian cancer cell lines (Yang et al. 2008). Let-7 g selectively affected the sensitivity of a drug resistant ovarian cancer cell line towards taxanes by targeting IMP-1, which in turn caused destabilization of MDR1 at the mRNA and protein level. This finally increased sensitivity of the multidrug resistant ovarian cancer cells to taxanes (Boyerinas et al. 2012). Furthermore, miR-125b targeted Bak1, a gene of the Bcl-2 protein family. Down-regulation of Bak1 resulted in an increased resistance to cisplatin by suppressing cisplatin-induced apoptosis (Kong et al. 2011). Down-regulation of miR-130b promoted the development of multidrug resistant ovarian cancer cells partly by targeting CSF-1, and silencing of miR-130b was demonstrated to be potentially mediated by DNA methylation. At the same time, low levels of miR-130b were associated with FIGO III-IV clinical stages in ovarian cancer patients and poor histological differentiation (Yang et al. 2012). MiR-152 and miR-185, which targets DNMT1 (Xiang et al. 2013), were also reported to be involved in chemotherapy resistance. DNMT1, the principal DNA methyltransferase, controls DNA methylation. In addition, miR-214 was shown to target PTEN, a regulator of cell proliferation, via the PI3K-Akt pathway. Knockdown of miR-214 reduced cell survival at around 20 % in cisplatin resistant ovarian cancer cells (Yang et al. 2008). MiR-376c was described to target al.K7 (a member of the TGF family that inhibits proliferation and induces apoptosis of epithelial ovarian cancer cells), and overexpression of miR-376c was found to block cisplatin-induced cell death, whereas anti-miR-376c treatment enhanced the effect of cisplatin (Ye et al. 2011). Another highly promising miRNA that is potentially involved in chemosensitivity is miR-21. As Liu et al. (2012b), found that berberine could inhibit miR-21 expression in several cancer cell lines. Subsequently, these authors investigated the influence of berberine on chemosensitivity of ovarian cancer cells to cisplatin. Interestingly, berberine could inhibit miR-21 expression and thereby modulate the sensitivity of cisplatin via regulating of the miR-21/PDCD4 axis (Liu et al. 2013). Other miRNAs involved in therapy resistance were miR-484 (target: vascular endothelial growth factor VEGFB and VEGFR2 pathways) (Vecchione et al. 2013), miR-23b, miR-27b, miR-424, and miR-503 (target: ALDH1) (Park et al. 2013), miR-106a (target: BCL10 and caspase-7) (Huh et al. 2013), miR-591 (target: ZEB1) (Huh et al. 2013), miR-31 (target: MET) (Mitamura et al. 2013), miR-130a (target: pro-metastatic and chemoresistance associated M-CSF) (Sorrentino et al. 2008), miR-484, miR-642 and miR-217 were described to be able to predict chemoresistance (Vecchione et al. 2013). Figure 15.1 shows a graphic presentation of a selection of resistance-relevant miRNAs and their downstream targets.
Despite the limitation of the in vitro character of these data, these experiments clearly demonstrate that miRNAs affect chemotherapy resistance on a cellular level. If these data can be reproduced in in vivo animal experiments, this would mean a major step towards a miRNA based new therapeutic approach for cancer patients. This new treatment option could be used either as additive treatment in conjunction with conventional therapies such as chemo- or radiotherapy, or maybe even as first line treatment if proven toxic to tumors in vivo. However, further experiments are highly warranted in order to unravel potential systemic side effects of miRNA based therapies, and to prove their success in complex organisms.
4 Conclusion and Perspectives
The presented in vitro data and (so far limited) in vivo data draw a most promising picture of miRNAs as potential clinical predictors and modifiers of response to chemo- and (to a lesser extent) radiotherapy. Overall, these results are highly encouraging and outline the enormous clinical impact that might arise from the use of miRNAs in the near future. However, as hopeful as these data are, their limitations have definitively to be considered: as outlined in this chapter, data are still very limited and sometimes even somewhat contradictory. Furthermore, conclusive in vivo data on miRNA based therapies are missing to this very date in general, so that information about possible complex interactions of a systemic therapy and potential toxic side effects for the patients have to be investigated before bringing these molecules into the clinic. However, the data presented in this chapter highlight the enormous potential of miRNAs for clinical application, and we are very confident that soon first reports on the clinical (diagnostic or therapeutic) use of miRNAs in the context of chemotherapy resistance will be available.
References
Allegra A, Alonci A, Campo S, Penna G, Petrunago A, Gerace D et al (2012) Circulating microRNAs: new biomarkers in diagnosis, prognosis and treatment of cancer (review). Int J Oncol 41(6):1897–1912
Asakura T, Ohkawa K (2004) Chemotherapeutic agents that induce mitochondrial apoptosis. Curr Cancer Drug Targets 4(7):577–590
Ballesta A, Lopez J, Popgeorgiev N, Gonzalo P, Doumic M, Gillet G (2013) Data-driven modeling of SRC control on the mitochondrial pathway of apoptosis: implication for anticancer therapy optimization. PLoS Comput Biol 9(4):e1003011
Blower PE, Verducci JS, Lin S, Park JK, Dai Z, Liu CG et al (2007) MicroRNA expression profiles for the NCI-60 cancer cell panel. Mol Cancer Ther 6(5):1483–1491
Boyerinas B, Park SM, Murmann AE, Gwin K, Montag AG, Zillhardt M et al (2012) Let-7 modulates acquired resistance of ovarian cancer to Taxanes via IMP-1-mediated stabilization of multidrug resistance 1. Int J Cancer 130(8):1787–1797
Cai EH, Gao YX, Wei ZZ, Chen WY, Yu P, Li K (2012) Serum miR-21 expression in human esophageal squamous cell carcinomas. Asian Pac J Cancer Prev 13(4):1563–1567
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S et al (2004) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci U S A 101(9):2999–3004
Chang H, Zhou X, Wang ZN, Song YX, Zhao F, Gao P et al (2012) Increased expression of miR-148b in ovarian carcinoma and its clinical significance. Mol Med Rep 5(5):1277–1280
Chen F, Chen C, Yang S, Gong W, Wang Y, Cianflone K et al (2012) Let-7b inhibits human cancer phenotype by targeting cytochrome P450 epoxygenase 2 J2. PLoS One 7(6):e39197
Chen MF, Chen PT, Lu MS, Lin PY, Chen WC, Lee KD (2013) IL-6 expression predicts treatment response and outcome in squamous cell carcinoma of the esophagus. Mol Cancer 12:26
Cochrane DR, Howe EN, Spoelstra NS, Richer JK (2010) Loss of miR-200c: a marker of aggressiveness and chemoresistance in female reproductive cancers. J Oncol 2010:821717
Cochrane DR, Spoelstra NS, Howe EN, Nordeen SK, Richer JK (2009) MicroRNA-200c mitigates invasiveness and restores sensitivity to microtubule-targeting chemotherapeutic agents. Mol Cancer Ther 8(5):1055–1066
Cortez MA, Bueso-Ramos C, Ferdin J, Lopez-Berestein G, Sood AK, Calin GA (2011) MicroRNAs in body fluids–the mix of hormones and biomarkers. Nat Rev Clin Oncol 8(8):467–477
Courrech Staal EF, Aleman BM, Boot H, van Velthuysen ML, van Tinteren H, van Sandick JW (2010) Systematic review of the benefits and risks of neoadjuvant chemoradiation for oesophageal cancer. Br J Surg 97(10):1482–1496
Duursma AM, Kedde M, Schrier M, le Sage C, Agami R (2009) miR-148 targets human DNMT3b protein coding region. RNA 14(5):872–877
Eitan R, Kushnir M, Lithwick-Yanai G, David MB, Hoshen M, Glezerman M et al (2009) Tumor microRNA expression patterns associated with resistance to platinum based chemotherapy and survival in ovarian cancer patients. Gynecol Oncol 114(2):253–259
Esquela-Kerscher A, Slack FJ (2006) Oncomirs – microRNAs with a role in cancer. Nat Rev Cancer 6(4):259–269
Fang Y, Fang D, Hu J (2012) MicroRNA and its roles in esophageal cancer. Med Sci Monit 18(3):RA22–RA30
Fiorica F, Di Bona D, Schepis F, Licata A, Shahied L, Venturi A et al (2004) Preoperative chemoradiotherapy for oesophageal cancer: a systematic review and meta-analysis. Gut 53(7):925–930
Frederick P, Green H, Huang J, Egger M, Frieboes H, Grizzle W et al (2013) Chemoresistance in ovarian cancer linked to expression of microRNAs. Biotech Histochem 88(7):403–409
Fujita Y, Kojima K, Ohhashi R, Hamada N, Nozawa Y, Kitamoto A et al (2010) MiR-148a attenuates paclitaxel resistance of hormone-refractory, drug-resistant prostate cancer PC3 cells by regulating MSK1 expression. J Biol Chem 285(25):19076–19084
Hamano R, Miyata H, Yamasaki M, Kurokawa Y, Hara J, Moon JH et al (2011) Overexpression of miR-200c induces chemoresistance in esophageal cancers mediated through activation of the Akt signaling pathway. Clin Cancer Res 17(9):3029–3038
Hong L, Han Y, Zhang H, Li M, Gong T, Sun L et al (2010) The prognostic and chemotherapeutic value of miR-296 in esophageal squamous cell carcinoma. Ann Surg 251(6):1056–1063
Hong F, Li Y, Xu Y, Zhu L (2013) Prognostic significance of serum microRNA-221 expression in human epithelial ovarian cancer. J Int Med Res 41(1):64–71
Hu X, Macdonald DM, Huettner PC, Feng Z, El Naqa IM, Schwarz JK et al (2009) A miR-200 microRNA cluster as prognostic marker in advanced ovarian cancer. Gynecol Oncol 114(3):457–464
Huh JH, Kim TH, Kim K, Song JA, Jung YJ, Jeong JY et al (2013) Dysregulation of miR-106a and miR-591 confers paclitaxel resistance to ovarian cancer. Br J Cancer 109(2):452–461
Hummel R, Hussey DJ, Michael MZ, Haier J, Bruewer M, Senninger N et al (2011a) MiRNAs and their association with locoregional staging and survival following surgery for esophageal carcinoma. Ann Surg Oncol 18(1):253–260
Hummel R, Wang T, Watson DI, Michael MZ, Van der Hoek M, Haier J et al (2011b) Chemotherapy-induced modification of microRNA expression in esophageal cancer. Oncol Rep 26(4):1011–1017
Iliopoulos D, Hirsch HA, Struhl K (2009) An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell 139(4):693–706
Imanaka Y, Tsuchiya S, Sato F, Shimada Y, Shimizu K, Tsujimoto G (2011) MicroRNA-141 confers resistance to cisplatin-induced apoptosis by targeting YAP1 in human esophageal squamous cell carcinoma. J Hum Genet 56(4):270–276
Iorio MV, Croce CM (2012) MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 4(3):143–159
Iorio MV, Visone R, Di Leva G, Donati V, Petrocca F, Casalini P et al (2007) MicroRNA signatures in human ovarian cancer. Cancer Res 67(18):8699–8707
Kang MH, Reynolds CP (2009) Bcl-2 inhibitors: targeting mitochondrial apoptotic pathways in cancer therapy. Clin Cancer Res 15(4):1126–1132
Kim T, Reitmair A (2013) Non-coding RNAs: functional aspects and diagnostic utility in oncology. Int J Mol Sci 14(3):4934–4968
Ko MA, Zehong G, Virtanen C, Guindi M, Waddell TK, Keshavjee S et al (2012) MicroRNA expression profiling of esophageal cancer before and after induction chemoradiotherapy. Ann Thorac Surg 94(4):1094–1102
Komatsu S, Ichikawa D, Takeshita H, Konishi H, Nagata H, Hirajima S et al (2012) Prognostic impact of circulating miR-21 and miR-375 in plasma of patients with esophageal squamous cell carcinoma. Expert Opin Biol Ther 12(Suppl 1):53–59
Komatsu S, Ichikawa D, Takeshita H, Tsujiura M, Morimura R, Nagata H et al (2011) Circulating microRNAs in plasma of patients with oesophageal squamous cell carcinoma. Br J Cancer 105(1):104–111
Kong F, Sun C, Wang Z, Han L, Weng D, Lu Y, Chen G (2011) miR-125b confers resistance of ovarian cancer cells to cisplatin by targeting pro-apoptotic Bcl-2 antagonist killer 1. J Huazhong Univ Sci Technolog Med Sci 31(4):543–549
Kozomara A, Griffith-Jones S (2011) miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 39(Database issue):D152–D157
Kumar S, Kumar A, Shah PP, Rai SN, Panguluri SK, Kakar SS (2011) MicroRNA signature of cis-platin resistant vs. cis-platin sensitive ovarian cancer cell lines. J Ovarian Res 4(1):17
Kurashige J, Kamohara H, Watanabe M, Tanaka Y, Kinoshita K, Saito S et al (2012) Serum microRNA-21 is a novel biomarker in patients with esophageal squamous cell carcinoma. J Surg Oncol 106(2):188–192
Lee H, Park CS, Deftereos G, Morihara J, Stern JE, Hawes SE et al (2012) MicroRNA expression in ovarian carcinoma and its correlation with clinicopathological features. World J Surg Oncol 10:174
Leskelä S, Leandro-García LJ, Mendiola M, Barriuso J, Inglada-Pérez L, Muñoz I et al (2010) The miR-200 family controls beta-tubulin III expression and is associated with paclitaxel-based treatment response and progression-free survival in ovarian cancer patients. Endocr Relat Cancer 18(1):85–95
Li H, Yang BB (2013) Friend or foe: the role of microRNA in chemotherapy resistance. Acta Pharmacol Sin 34(7):870–879
Lin RJ, Xiao DW, Liao LD, Chen T, Xie ZF, Huang WZ et al (2012) MiR-142-3p as a potential prognostic biomarker for esophageal squamous cell carcinoma. J Surg Oncol 105(2):175–182
Liu R, Liao J, Yang M, Shi Y, Peng Y, Wang Y, Pan E et al (2012a) Circulating miR-155 expression in plasma: a potential biomarker for early diagnosis of esophageal cancer in humans. J Toxicol Environ Health A 75(18):1154–1162
Liu S, Fang Y, Shen H, Xu W, Li H (2012b) Berberine sensitizes ovarian cancer cells to cisplatin through miR-21/PDCD4 axis. Acta Biochim Biophys Sin (shanghai) 45(9):756–762
Liu S, Fang Y, Shen H, Xu W, Li H (2013) Berberine sensitizes ovarian cancer cells to cisplatin through miR-21/PDCD4 axis. Acta Biochim Biophys Sin (Shanghai) 45(9):756–762
Lu L, Schwartz P, Scarampi L, Rutherford T, Canuto EM, Yu H et al (2011) MicroRNA let-7a: a potential marker for selection of paclitaxel in ovarian cancer management. Gynecol Oncol 122(2):366–371
Lynam-Lennon N, Reynolds JV, Marignol L, Sheils OM, Pidgeon GP, Maher SG (2012) MicroRNA-31 modulates tumour sensitivity to radiation in oesophageal adenocarcinoma. J Mol Med 90(12):1449–1458
Marchini S, Cavalieri D, Fruscio R, Calura E, Garavaglia D, Nerini IF et al (2011) Association between miR-200c and the survival of patients with stage I epithelial ovarian cancer: a retrospective study of two independent tumour tissue collections. Lancet Oncol 12(3):273–285
Mariani M, Shahabi S, Sieber S, Scambia G, Ferlini C (2011) Class III β-tubulin (TUBB3): more than a biomarker in solid tumors? Curr Mol Med 11(9):726–731
Mateescu B, Batista L, Cardon M, Gruosso T, de Feraudy Y, Mariani O et al (2011) MiR-141 and miR-200a act on ovarian tumorigenesis by controlling oxidative stress response. Nat Med 17(12):1627–1635
Merkerova M, Vasikova A, Belickova M, Bruchova H (2010) Micro-RNA expression profiles in umbilical cord blood cell lineages. Stem Cells Dev 19(1):17–26
Mitamura T, Watari H, Wang L, Kanno H, Hassan MK, Miyazaki M et al (2013) Downregulation of miRNA-31 induces taxane resistance in ovarian cancer cells through increase of receptor tyrosine kinase MET. Oncogenesis 2(3):e40
Mo MH, Chen L, Fu Y, Wang W, Fu SW (2012) Cell-free circulating miRNA biomarkers in cancer. J Cancer 3:432–448
Moss C, Kaye SB (2002) Ovarian cancer: progress and continuing controversies in management. Eur J Cancer 38(13):1701–1707
Nam EJ, Yoon H, Kim SW, Kim H, Kim YT, Kim JH et al (2008) MicroRNA expression profiles in serous ovarian carcinoma. Clin Cancer Res 14(9):2690–2695
Odenthal M, Bollschweiler E, Grimminger PP, Schröder W, Brabender J, Drebber U et al (2013) MicroRNA profiling in locally advanced esophageal cancer indicates a high potential of miR-192 in prediction of multimodality therapy response. Int J Cancer 133(10):2454–2463
Ohkawa K, Asakura T, Aoki K, Shibata S, Minami J, Fujiwara C et al (2004) Establishment and some characteristics of epoxomicin (a proteasome inhibitor) resistant variants of the human squamous cell carcinoma cell line, A431. Int J Oncol 24(2):425–433
Ozols RF (2005) Treatment goals in ovarian cancer. Int J Gynecol Cancer 15(Suppl 1):3–11
Park YT, Jeong JY, Lee MJ, Kim KI, Kim TH, Kwon YD, Lee C et al (2013) MicroRNAs overexpressed in ovarian ALDH1-positive cells are associated with chemoresistance. J Ovarian Res 6(1):18
Peng DX, Luo M, Qiu LW, He YL, Wang XF (2012) Prognostic implications of microRNA-100 and its functional roles in human epithelial ovarian cancer. Oncol Rep 27(4):1238–1244
Prislei S, Martinelli E, Mariani M, Raspaglio G, Sieber S, Ferrandina G et al (2013) MiR-200c and HuR in ovarian cancer. BMC Cancer 13:72
Resnick KE, Alder H, Hagan JP, Richardson DL, Croce CM, Cohn DE (2009) The detection of differentially expressed microRNAs from the serum of ovarian cancer patients using a novel real-time PCR platform. Gynecol Oncol 112(1):55–59
Revilla-Nuin B, Parrilla P, Lozano JJ, de Haro LF, Ortiz A, Martínez C, Munitiz V et al (2013) Predictive value of MicroRNAs in the progression of Barrett esophagus to adenocarcinoma in a long-term follow-up study. Ann Surg 257(5):886–893
Rosenwald S, Gilad S, Benjamin S (2010) Validation of a microRNA-based qRT-PCR test for accurate identification of tumor tissue origin. Mod Pathol 23(6):814–823
Sjoquist KM, Burmeister BH, Smithers BM, Zalcberg JR, Simes RJ, Barbour A et al (2011) Survival after neoadjuvant chemotherapy or chemoradiotherapy for resectable oesophageal carcinoma: an updated meta-analysis. Lancet Oncol 12(7):681–692
Sorrentino A, Liu CG, Addario A, Peschle C, Scambia G, Ferlini C (2008) Role of microRNAs in drug-resistant ovarian cells. Gynecol Oncol 111(3):478–486
Sugimura K, Miyata H, Tanaka K, Hamano R, Takahashi T, Kurokawa Y et al (2012) Let-7 expression is a significant determinant of response to chemotherapy through the regulation of IL-6/STAT3 pathway in esophageal squamous cell carcinoma. Clin Cancer Res 18(18):5144–5153
Takeshita N, Hoshino I, Mori M, Akutsu Y, Hanari N, Yoneyama Y et al (2013) Serum microRNA expression profile: miR-1246 as a novel diagnostic and prognostic biomarker for oesophageal squamous cell carcinoma. Br J Cancer 108(3):644–652
Urschel JD, Vasan H (2003) A meta-analysis of randomized controlled trials that compared neoadjuvant chemoradiation and surgery to surgery alone for resectable esophageal cancer. Am J Surg 185(6):538–543
van Jaarsveld MT, Helleman J, Berns EM, Wiemer EA (2010) MicroRNAs in ovarian cancer biology and therapy resistance. Int J Biochem Cell Biol 42(8):1282–1290
van Jaarsveld MT, Helleman J, Boersma AW, van Kuijk PF, van Ijcken WF, Despierre E (2012) miR-141 regulates KEAP1 and modulates cisplatin sensitivity in ovarian cancer cells. Oncogene 32(36):4284–4293
Vecchione A, Belletti B, Lovat F, Volinia S, Chiappetta G, Giglio S et al (2013) A microRNA signature defines chemoresistance in ovarian cancer through modulation of angiogenesis. Proc Natl Acad Sci 110(24):9845–9850
Wang B, Zhang Q (2012) The expression and clinical significance of circulating microRNA-21 in serum of five solid tumors. J Cancer Res Clin Oncol 138(10):1659–1666
Wang Y, Niu XL, Qu Y, Wu J, Zhu YQ, Sun WJ et al (2010) Autocrine production of interleukin-6 confers cisplatin and paclitaxel resistance in ovarian cancer cells. Cancer Lett 295(1):110–123
Weber JA, Baxter DH, Zhang S (2010) The microRNA spectrum in 12 body fluids. Clin Chem 56(11):1733–1741
Wen J, Zheng B, Hu Y, Zhang X, Yang H, Luo KJ et al (2009) Establishment and biological analysis of the EC109/CDDP multidrug-resistant esophageal squamous cell carcinoma cell line. Oncol Rep 22(1):65–71
Wu Y, Xiao Y, Ding X, Zhuo Y, Ren P, Zhou C et al (2011) A miR-200b/200c/429-binding site polymorphism in the 3′ untranslated region of the AP-2 α gene is associated with cisplatin resistance. PLoS One 6(12):e29043
Xiang Y, Ma N, Wang D, Zhang Y, Zhou J, Wu G et al (2013) MiR-152 and miR-185 co-contribute to ovarian cancer cells cisplatin sensitivity by targeting DNMT1 directly: a novel epigenetic therapy independent of decitabine. Oncogene. doi: 10.1038/onc.2012.575 [Epub ahead of print]
Xiao YF, Yong X, Fan YH, Lü MH, Yang SM, Hu CJ (2013) MicroRNA detection in feces, sputum, pleural effusion and urine: novel tools for cancer screening (Review). Oncol Rep 30(2):535–544
Xu YZ, Xi QH, Ge WL, Zhang XQ (2013) Identification of serum microRNA-21 as a biomarker for early detection and prognosis in human epithelial ovarian cancer. Asian Pac J Cancer Prev 14(2):1057–1060
Yang N, Kaur S, Volinia S, Greshock J, Lassus H, Hasegawa K et al (2008) MicroRNA microarray identifies Let-7i as a novel biomarker and therapeutic target in human epithelial ovarian cancer. Cancer Res 68(24):10307–10314
Yang C, Cai J, Wang Q, Tang H, Cao J, Wu L et al (2012) Epigenetic silencing of miR-130b in ovarian cancer promotes the development of multidrug resistance by targeting colony-stimulating factor 1. Gynecol Oncol 124(2):325–334
Ye G, Fu G, Cui S, Zhao S, Bernaudo S, Bai Y et al (2011) MicroRNA 376c enhances ovarian cancer cell survival by targeting activin receptor-like kinase 7: implications for chemoresistance. J Cell Sci 124(Pt 3):359–368
Zaman MS, Maher DM, Khan S, Jaggi M, Chauhan SC (2012) Current status and implications of microRNAs in ovarian cancer diagnosis and therapy. J Ovarian Res 5(1):44
Zhang C, Wang C, Chen X, Yang C, Li K, Wang J et al (2010) Expression profile of microRNAs in serum: a fingerprint for esophageal squamous cell carcinoma. Clin Chem 56(12):1871–1879
Zhang T, Wang Q, Zhao D, Cui Y, Cao B, Guo L et al (2011) The oncogenetic role of microRNA-31 as a potential biomarker in oesophageal squamous cell carcinoma. Clin Sci 121(10):437–447
Zhang T, Zhao D, Wang Q, Yu X, Cui Y, Guo L et al (2012) MicroRNA-1322 regulates ECRG2 allele specifically and acts as a potential biomarker in patients with esophageal squamous cell carcinoma. Mol Carcinog 52(8):581–590
Zheng ZF, Su HF, Zou Y, Peng Z, Wu SX (2011) Expression profiles of microRNAs in radioresistant esophageal cell line. Zhonghua Yi Xue Za Zhi 91(9):639–642
Zhou L, Yang ZX, Song WJ, Li QJ, Yang F, Wang DS et al (2013) MicroRNA-21 regulates the migration and invasion of a stem-like population in hepatocellular carcinoma. Int J Oncol 43(2):661–669
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Lindner, K., Haier, J., Hummel, R. (2014). MicroRNAs and Their Clinical Impact on Resistance to Anticancer Treatment. In: Babashah, S. (eds) MicroRNAs: Key Regulators of Oncogenesis. Springer, Cham. https://doi.org/10.1007/978-3-319-03725-7_15
Download citation
DOI: https://doi.org/10.1007/978-3-319-03725-7_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-03724-0
Online ISBN: 978-3-319-03725-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)